The significance levels of the factors included in the experiment and obtained by the ANOVA are presented in
Table 1. Treatment (inoculation), aging time, and the interaction between them showed a significant effect on proteolytic changes expressed as primary amino groups (-NH
2) and antiradical activity (ABTS) before and after each step of in vitro gastrointestinal digestion and simulated absorption of dry-cured loins.
2.1. Stability of Antiradical Activity during Long-Term Aging
The highest antiradical activity against ABTS (%) was noted with the protein fractions extracted from dry-cured loins after 180 days of aging (see
Table 2). However, the influence of either the strain used (LOCK, BAUER, or BB12) or the aging time on the inactivation ability of radicals generated from ABTS was ambiguous. Generally, the increase of antiradical activity of protein extracts up to 180 days of aging (expressed as percentage scavenging activity) was observed and followed by a systematic decrease during further aging stages. As far as WSF is concerned, at the first sampling point (after 28 days of aging), the highest antiradical activity (
p < 0.05) was noted for the LOCK sample (69.94%).
However, in the subsequent steps (90, 180, and 270 days), the Lactobacillus rhamnosus LOCK900 strain was less effective in generating antiradical components with no statistically significant differences between LOCK and C sample. The highest antiradical effects were achieved for loins with BAUER after 90 and 180 days of aging (77.53% and 83.89%, respectively) and BB12 in 270 days (74.12%). The varied effect of probiotic LAB strains on generating antiradical molecules during long-term aging was also observed for SSF. Lower (p < 0.05) antiradical values were observed for batches with LAB compared with the variant that underwent spontaneous fermentation (C) at day 28. After this time, the behavior of this parameter in inoculated samples was ambiguous during the rest of the aging period. The biggest differences in antiradical activity (%) were noted between the sample subjected to spontaneous fermentation (C) and BB12 after 28, 90, 180, and 270 days. The LOCK batches had statistically significantly lower antiradical properties (%) compared to C (p < 0.05) at 28, 270, and 360 days. BAUER batches had significantly lower values (p < 0.05) of radical scavenging activity after 28, 180, and 360 days of aging as compared to C batches.
Antiradical properties were also defined as the concentration of the sample required to inhibit 50% of the radical-scavenging effect (IC
50). Generally, the IC
50 values clearly indicated the higher antiradical properties of protein-released components during long-term aging from SSF compared to WSF (see
Table 2). With respect to the WSF, the antiradical activity of the hydrolysis products was stable between 28 and 180 day of aging (
p > 0.05) and then decreased in all samples. SSF fractions were characterized by greater fluctuations of this parameter especially between 28 and 180 days of aging (
p < 0.05).
Quantitative analysis was also used for investigating the correlation between antiradical activities and the content of protein degradation products in both fractions of muscle proteins. The correlation between the antiradical activity (expressed as percent inhibition and 1/IC
50 (not IC
50) showing parallelism with antiradical activity) and the primary amino groups content (µM/mL) was therefore determined [
20].
As shown in
Table 3, there was no positive correlation between the content of protein degradation products and the antiradical activity, which corresponds with other studies [
21,
22,
23]. However, the strong negative correlation between the antiradical activity of the WSF fraction (expressed as 1/IC
50) and the content of components with potential antiradical properties indicated the loss of bioactive properties during aging, which corresponds with the results presented in
Table 1.
Therefore, the progressive degradation of proteins by endogenous meat enzymes and exogenous microbial proteases during 360 days of aging results in the disappearance of biological activity of protein-related compounds at a later stage. This may be a result of an overly extensive hydrolysis of the peptide chains [
23].
2.2. Stability of Antiradical Activity during In Vitro Digestion
Determining the in vitro bioactivity to promote the beneficial effects of bioactive compounds should be carried out in the context of their immunity to digestive enzymes and to estimate their nutritional importance. In this context, hydrolysis of protein extracts obtained from dry-cured pork loin using gastrointestinal enzymes was accomplished using a two-step hydrolysis reaction. The first step was hydrolysis by pepsin (pH 2 at 37 °C for 2 hours) while the second step was the successive hydrolysis by pancreatin (pH 7 at 37 °C for 3 hours). The sequential digestion with pepsin and pancreatin provides a suitable model for evaluating peptides released in the intestinal tract. The effect of gastric in vitro (pepsin) and consecutive intestinal in vitro (pancreatin) digestion on protein extracts of dry-cured loins was discussed. The results are summarized in
Table 4 and
Table 5.
As noted earlier, the antiradical activity defined as percent inhibition and IC
50 values were uncorrelated. However, the higher antiradical activity (%) was determined for the WSF fraction, which was probably indirectly related to a higher content of primary amino groups. The IC
50 values unambiguously indicate SSF as a source of antiradical components and this tendency was maintained during in vitro gastrointestinal digestion (see
Table 4 and
Table 5) and simulated adsorption (see
Table 6).
The decrease in the antiradical activity of the dry-cured loin protein hydrolysates determined by the ABTS test (expressed as % and IC
50) was observed after gastric digestion (see
Table 4) and compared to the undigested samples, which was followed by an increase of biological activity after pancreatin treatment (see
Table 5). This observation corresponds with other authors’ findings [
24,
25,
26,
27]. The highest biological activity was achieved after 90 days of aging due to the hydrolytic degradation of proteins under the action of pepsin. During this period, an average of 83.60% was reported for WSF with significantly higher (
p < 0.05) values of biological activity obtained for BB12 batches (86.51%). With regard to SSF, the antiradical activity after 90 days of aging was at an average level of 69.34% for C and LOCK (
p > 0.05), while BAUER and BB12 batches was statistically significantly (
p < 0.05) higher (70.61% and 75.18%, respectively). Pancreatin digested samples (see
Table 5) showed values ranging from 85.94 for BAUER (28 days) to 98.56 for BB12 (90 days) for WSF and 72.45 for BAUER (28 days) to 95.86% for C (270 days) in the case of SSF. This suggests that fewer peptides with antiradical properties are associated with pepsin digestion than pancreatin. This may be due to the nature of the enzymes used. While pepsin can break down proteins and large peptides into smaller fragments by shielding the cleavage sites for further enzymes, pancreatin can primarily hydrolyze some of the peptides in smaller peptides and possibly amino acids. Pancreatin contains many enzymes including trypsin and additional proteases that give rise to hydrolysis activity, which leads to deeper breakdown of peptide chains. These results can be explained by the formation of a greater proportion of peptides and amino acids with hydrophilic properties during pancreatic digestion. Moreover, Zhu et al. [
28] reported that pepsin cleaves peptides into smaller fragments, which exposes the internal groups to the environment. While trypsin hydrolyzed peptides into smaller chains, it also produced more free amino acids due to its greater hydrolytic activity. Therefore, these amino acids have greater affinity with water. This is because the increase of hydrophobic properties of GI digestion after pepsin treatment makes them less likely to react with the water-soluble ABTS radical. However, the increase of the hydrophilic property of GI digestion after pancreatin treatment favors their trapping of the ABTS radical [
24].
After in vitro gastrointestinal digestion and simulated absorption, the antiradical properties of the hydrolysates increased compared to undigested proteins (see
Table 6).
As expected, the best antiradical properties were achieved after in vitro gastrointestinal digestion and simulated adsorption of WSF extracted after 28 days of aging. These results are not consistent with the radical scavenging activity expressed by the IC
50 for which the increase in antiradical properties attain the lowest biological activity (highest antiradical activity) on day 180, which was followed by a systematic decline until the end of the aging period was noted. This tendency was described earlier (see
Table 2). In this period (180 days of aging), the statistically significantly higher biological activity as an antiradical within the WSF (
p < 0.05) was shown for the BAUER and BB12 (IC
50 = 0.02 µM/mL both;
Table 6). With regard to SSF, the best antiradical properties (determined by % and IC
50) were recorded after simulated absorption of the digested product after 180 days of aging with the best properties reported (
p < 0.05) for the spontaneous fermentation (C) and BB12 batches. However, the differences between them were not statistically significant (
p > 0.05).
2.3. Stability of Antiradical Activity during In Vitro Digestion
Dry-cured meats constitute a specific group of products in which the proteolytic processes take place from raw material to finished product, which can take up to 24 months. Proteolytic degradation of proteins takes place through exogenous enzymes of meat as well as by exogenous enzymes derived from microorganisms primarily responsible for fermentation processes that occur on raw meat. The proteolytic activity attributed to bacteria is characteristic of a particular strain. Therefore, different peptide and amino acid profiles are predicted depending on the LAB strains used during production. These aspects are crucial for detecting the specific functions of potentially bioactive peptides especially when digested with enzymes of the human gastrointestinal tract. The digestive tract affects the release of peptides from parent proteins and modifies or degrades peptides that may exhibit antiradical properties. In fact, the specificity of the enzyme affects the amount, size, and composition of the peptides, which influences the biological activity of the digested samples [
25,
29] and the degree of their absorption through the intestinal membrane.
Therefore, LC-MS/MS analysis was used to evaluate peptides (28, 90, and 180 days) and compare the peptide profile of the samples after in vitro digestion and absorption in the simulated gastrointestinal tract. Peptide sequences were identified and characterized by nano-LC-MS, which confirms the identification by exact mass determination with LTQ-Orbitrap. The cleavage of peptide bonds by digestive proteases leads to the release of peptides that have different lengths and free amino acids. The most typical well-known bioactive peptides are 200–1700 Da with fragment lengths from 2 to 14 amino acids so that they are able to easily pass through the gut lane and are capable of secreting nutritional value and bioactive functions [
30].
The peptides found in the digested samples have MW between 700 and 4232 Da and the length of the peptide fragments determined in the study ranged from 5 to 47 amino acids in a sequence, which is in agreement with other peptide profile studies obtained by digestion [
31,
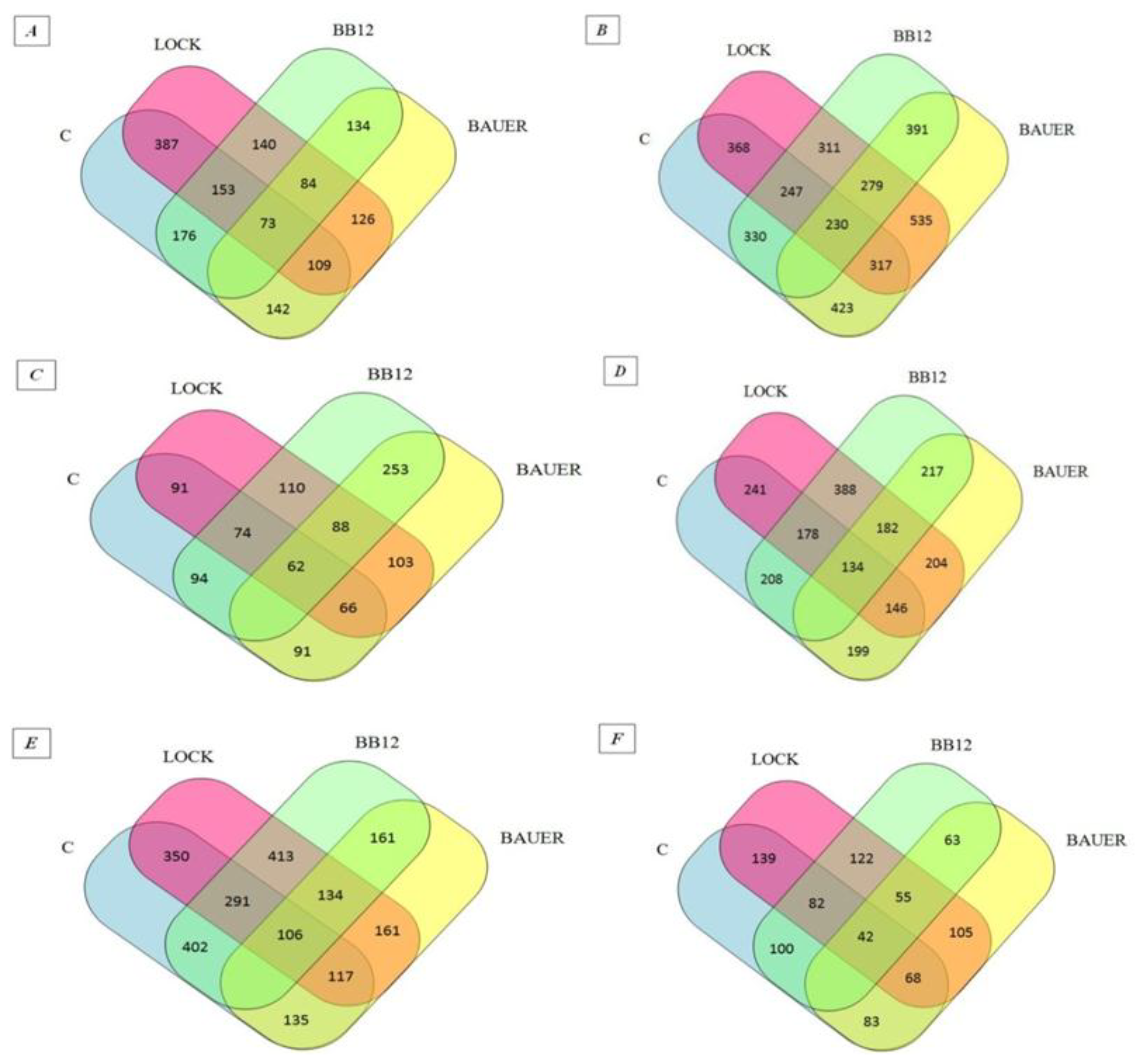
32]. This probably indicates that they may have biological effects. The influence of individual strains on the peptide profile of the analyzed samples after the digestion and absorption process was summarized in the Venn diagram (see
Figure 1) [
33]. Venn diagrams indicated an increase in the number of peptides identified as common for the WSF of batches inoculated with LAB (from 84 to 134 sequences obtained after in vitro digestion and adsorption. At the same time in relation to the SSF, a decrease from 279 to 55 common sequences has been noted. After 180 days of aging, 402 common peptide sequences were identified for the WSF of C and BB12 samples while only 135 identified peptides were common for the C and BAUER batches. By analogy, taking into account the SSF, the C sample had more common peptide sequences with the LOCK (139) and less when compared to the BAUER sample (83).
Due to the large number of peptides and, taking into account the highest antiradical activity of protein-released components extracted from the loins after 180 days of aging, the peptides derived from digestion and simulated absorption were identified by chromatographic methods and in silico analysis. The analyses were repeated twice and identical peptide sequences were selected for further analysis. In total, all peptides up to 480 sequences (38.53%) showed potency as antioxidants in the in silico study. In addition, the selected sequences were evaluated by rating capacity for bioactivity.
The peptide sequences with the highest
A parameter (i.e., above 0.4) are presented in
Table 7. Both sarcoplasmic and myofibrillar proteins have been described as precursors of bioactive peptides on the basis of in vitro assays. The amino acid composition, conformation, and hydrophobicity is correlated with antioxidant activity and likely determines the mechanism (transfer of hydrogen (HAT) or single electron (SET)) and the effectiveness of antioxidants [
34,
35]. It has been found that the peptides possibly containing substances and donating electrons are likely to react with the free radicals to terminate a radical reaction. Cys and Met residues, which contain nucleophilic sulphur side chains as well as Trp, Tyr, and Phe, have aromatic side chains and readily donate hydrogen atoms [
28]. The antioxidant activity of peptides with one or more residues of His, Pro, Cys, Tyr, Trp, Phe, or Met and the presence of hydrophobic amino acids might be enhanced [
36]. Peptides containing the amino acid residues Val, Leu, Ile, Ala, Phe, Lys, or Cys at the N-terminal and Trp, Tyr, His, or Pro in the sequence had been reported to show antioxidant activity [
36,
37]. This is caused by acidic or basic amino acid residues (Asp, Glu, His, Arg, or Lys) or hydrophilic amino acids (Ser, Thr) in this position. Moreover, as reported by Power et al. [
35], it is suggested that a second amino acid adhering to the C-terminal is a major factor influencing antioxidant activity. If this amino acid has a high hydrogen bond and steric and low hydrophobicity, it will increase its anti-oxidative potential.
Within the peptide sequences identified in the present study (see
Table 7), Leu (19.01%), Pro (15.85%), Lys (11.62%), Glu (7.39%), and His (7.04%) had the largest share in their amino acid composition, which corresponds to other authors [
17,
21]. Their presence is likely to determine the antiradical ability of the peptides due to their ability to quench unpaired electrons or radicals by supporting protons. Other authors also reported that the presence of these nonpolar amino acids such as Leu and Pro has been correlated to the antioxidant activity [
21,
38]. This contributes to the radical scavenging activity of peptides due to their special structural characteristics. Chen et al. [
39] demonstrated that peptides containing His can act as metal ion chelators, active-oxygen quenchers, and hydroxy radical scavengers. Escudero et al. [
17] reported about proline-rich peptide SAGNPN, which showed a high radical scavenging activity. Yet, many synthesized peptides like GGSILI, IAKLE, ALGGA, NVLVG, GLAGA, and NAAKL possessed Leu residues. The presence of Leu possibly contributed to the antioxidant activities of peptides [
17,
40].