Study of Potential Anti-Inflammatory Effects of Red Wine Extract and Resveratrol through a Modulation of Interleukin-1-Beta in Macrophages
Abstract
1. Introduction
2. Materials and Methods
2.1. Cells Culture
2.2. Preparation of Red Wine Extract
2.3. HPLC Analysis
2.4. Reagents
2.5. Experimental Protocol
2.6. Cell Proliferation Assay
2.7. Western Blotting
2.8. Imaging
2.9. IL-1β Secretion
2.10. Statistical Analysis
3. Results
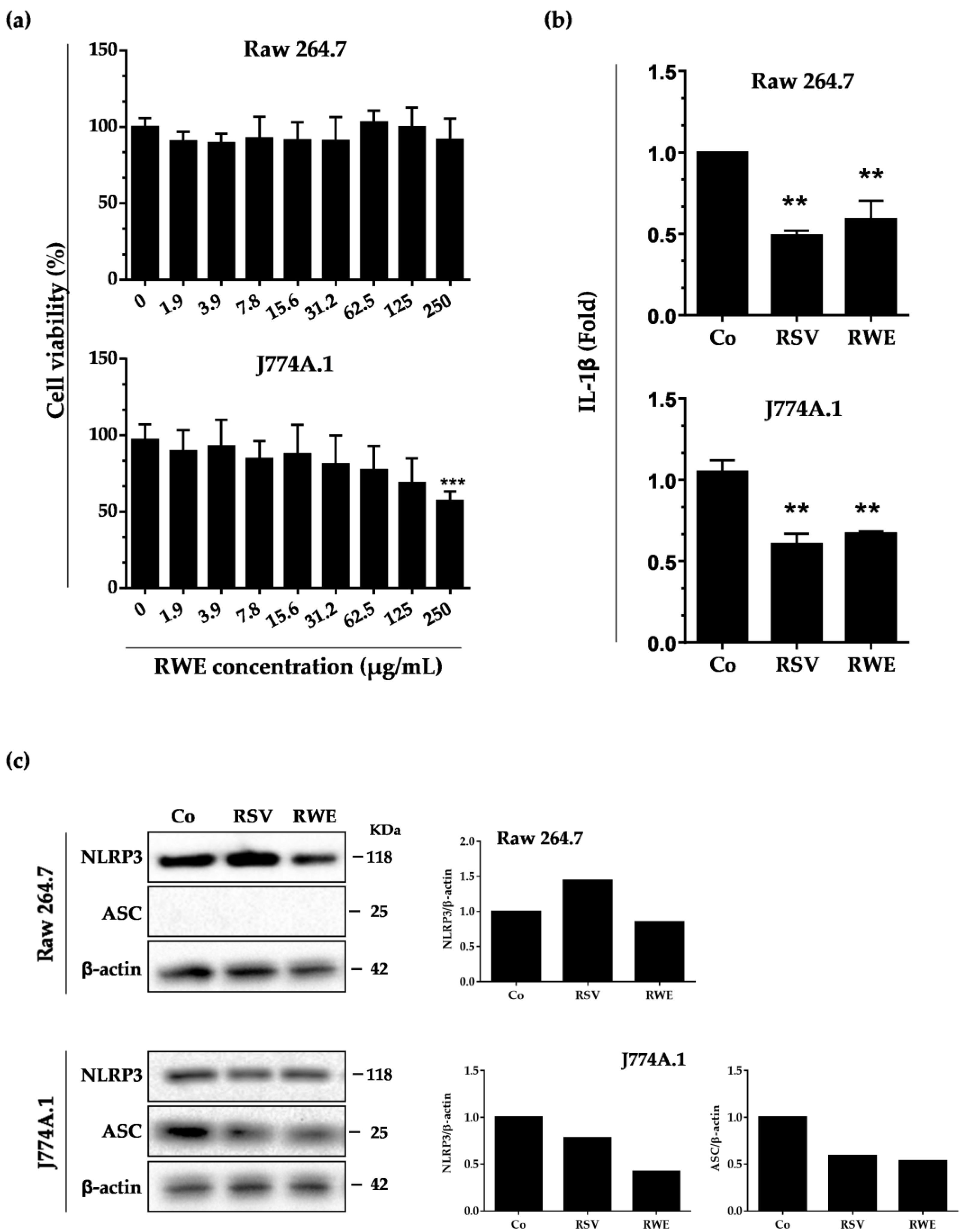
3.1. RWE Decreases IL-1β Secretion and NLRP3 Expression in Murine Macrophages without Toxicity
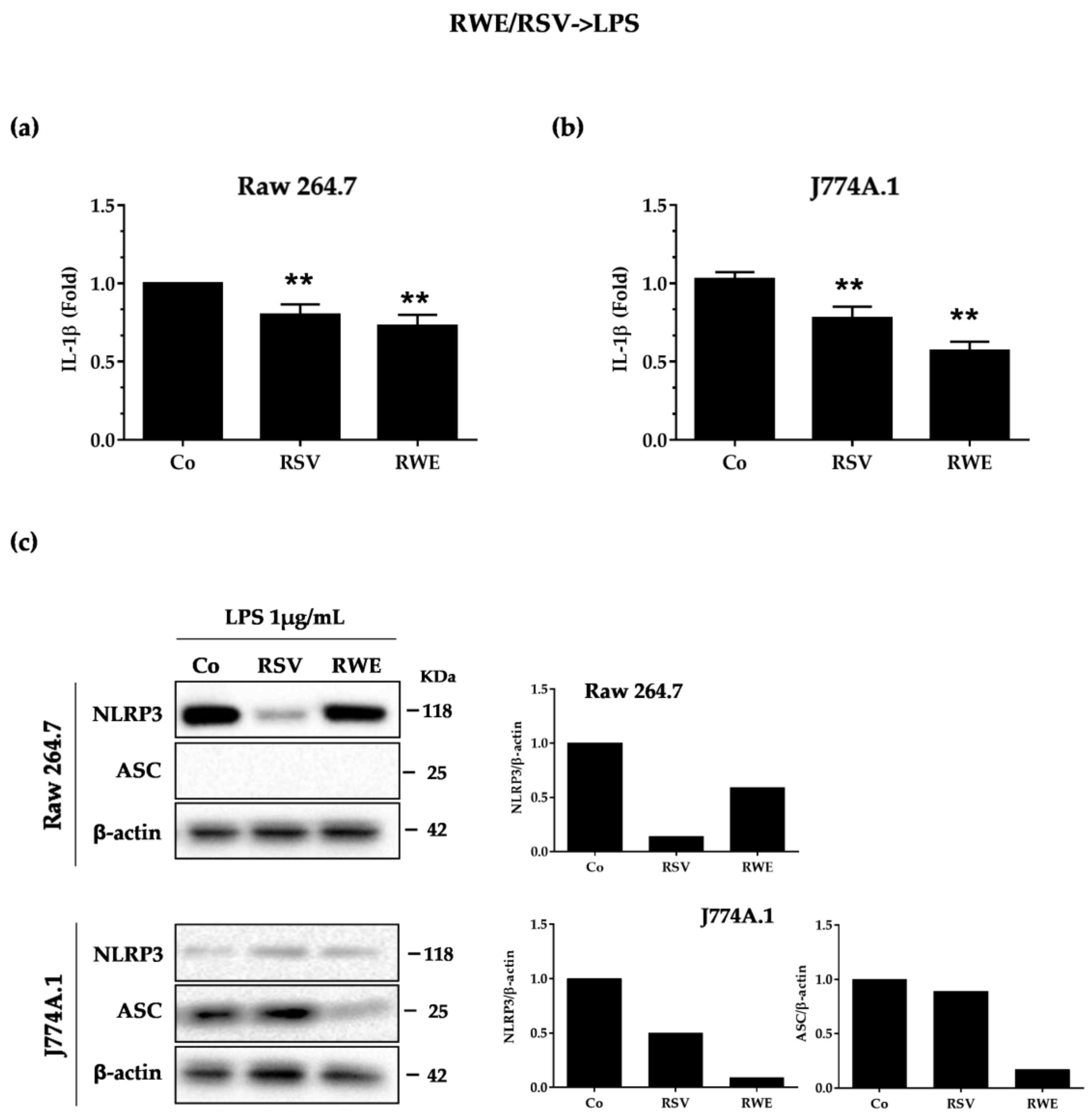
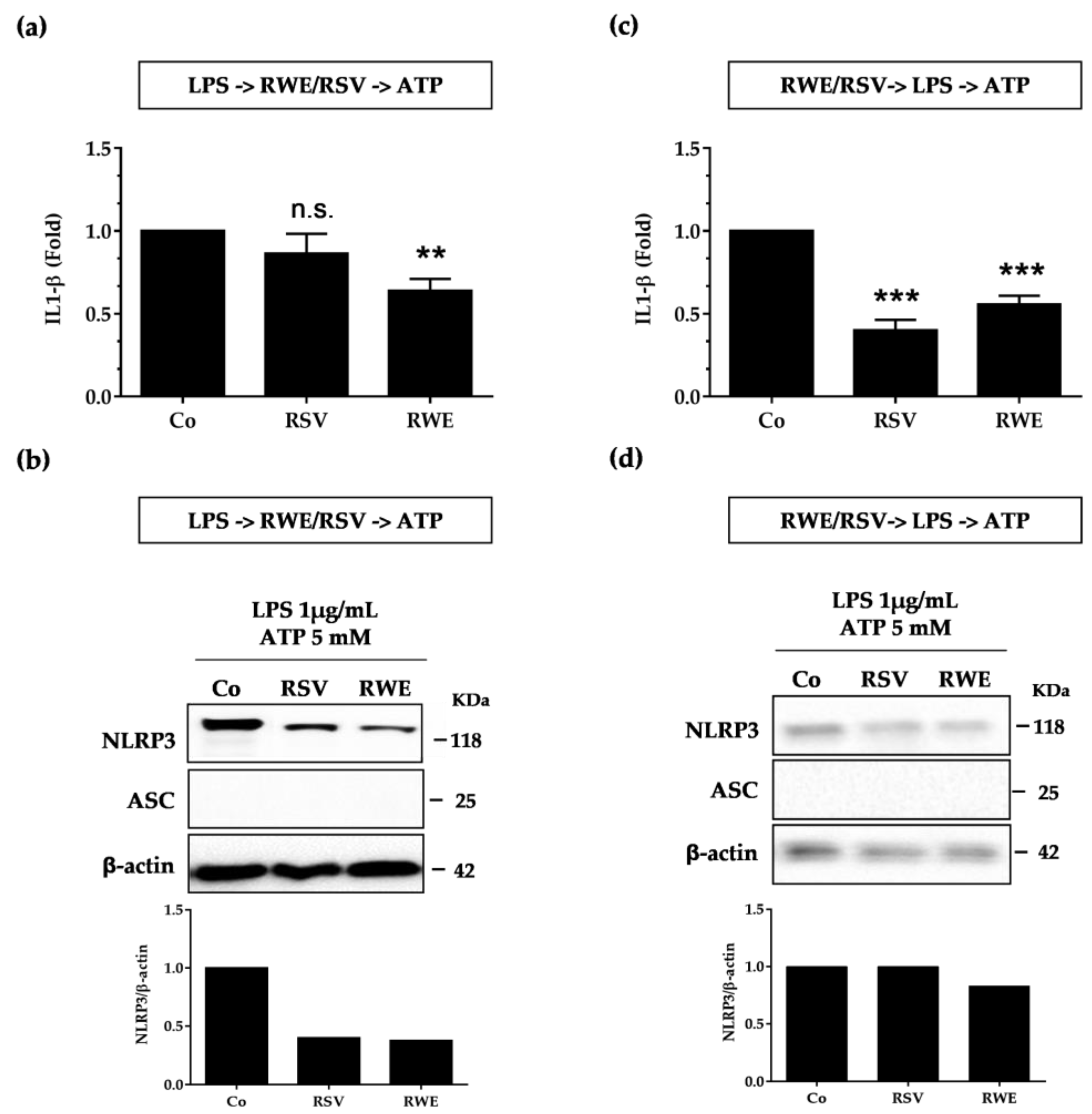
3.2. RWE Prevents Priming of NLRP3 Inflammasome from LPS in Macrophages
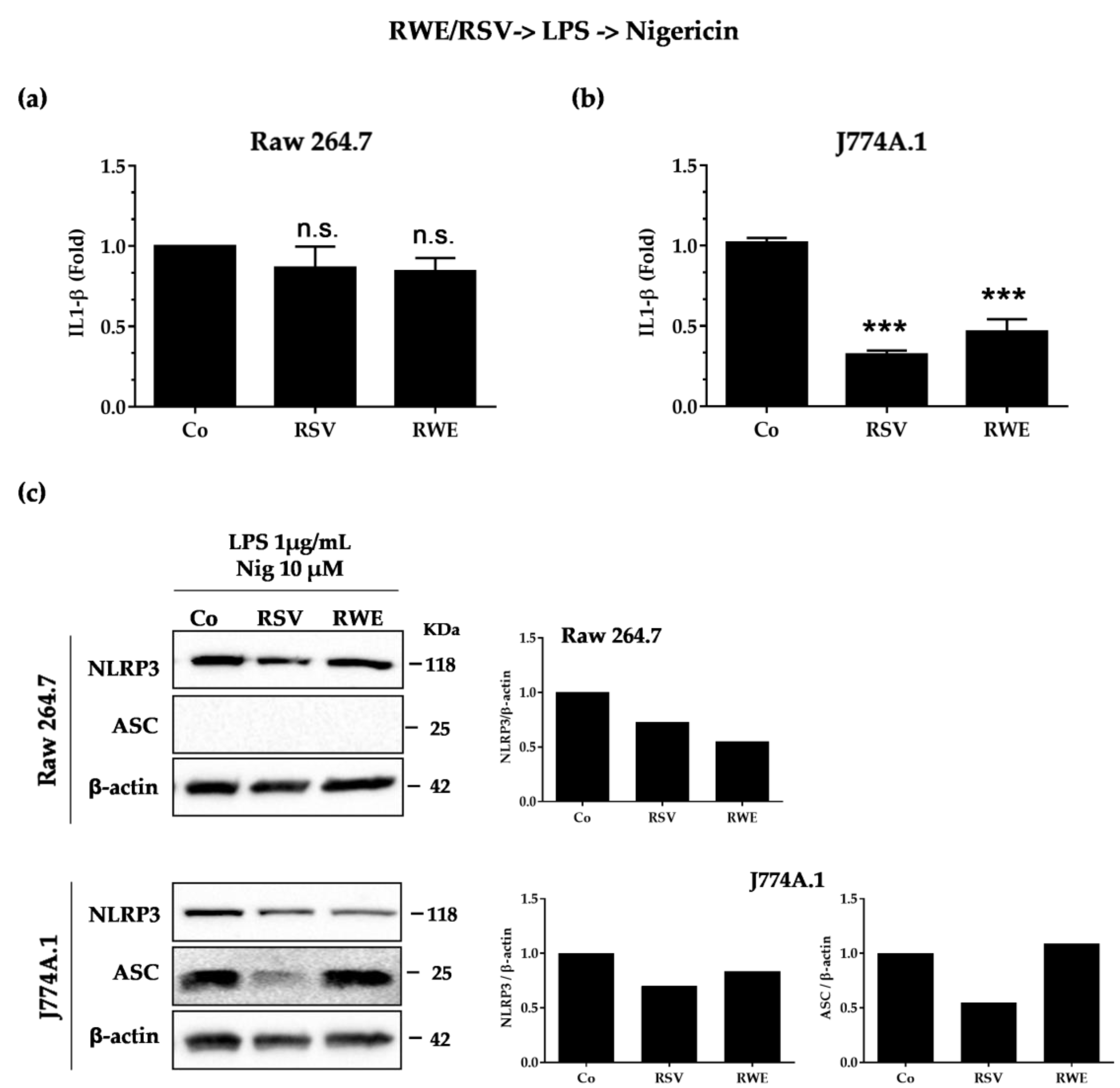
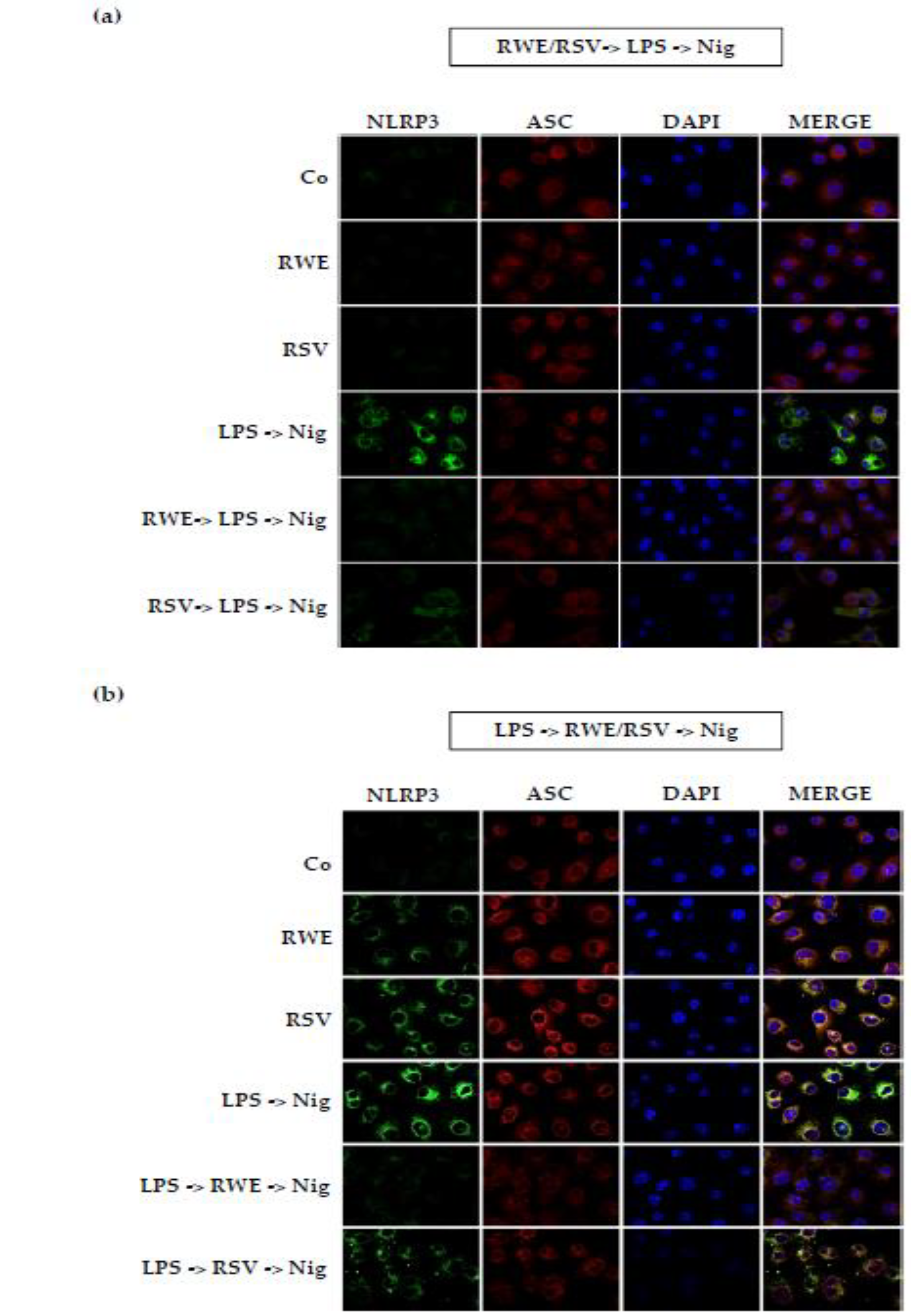
3.3. RWE Decreases IL-1β Secretion after Activation by LPS/Nigericin in Macrophages
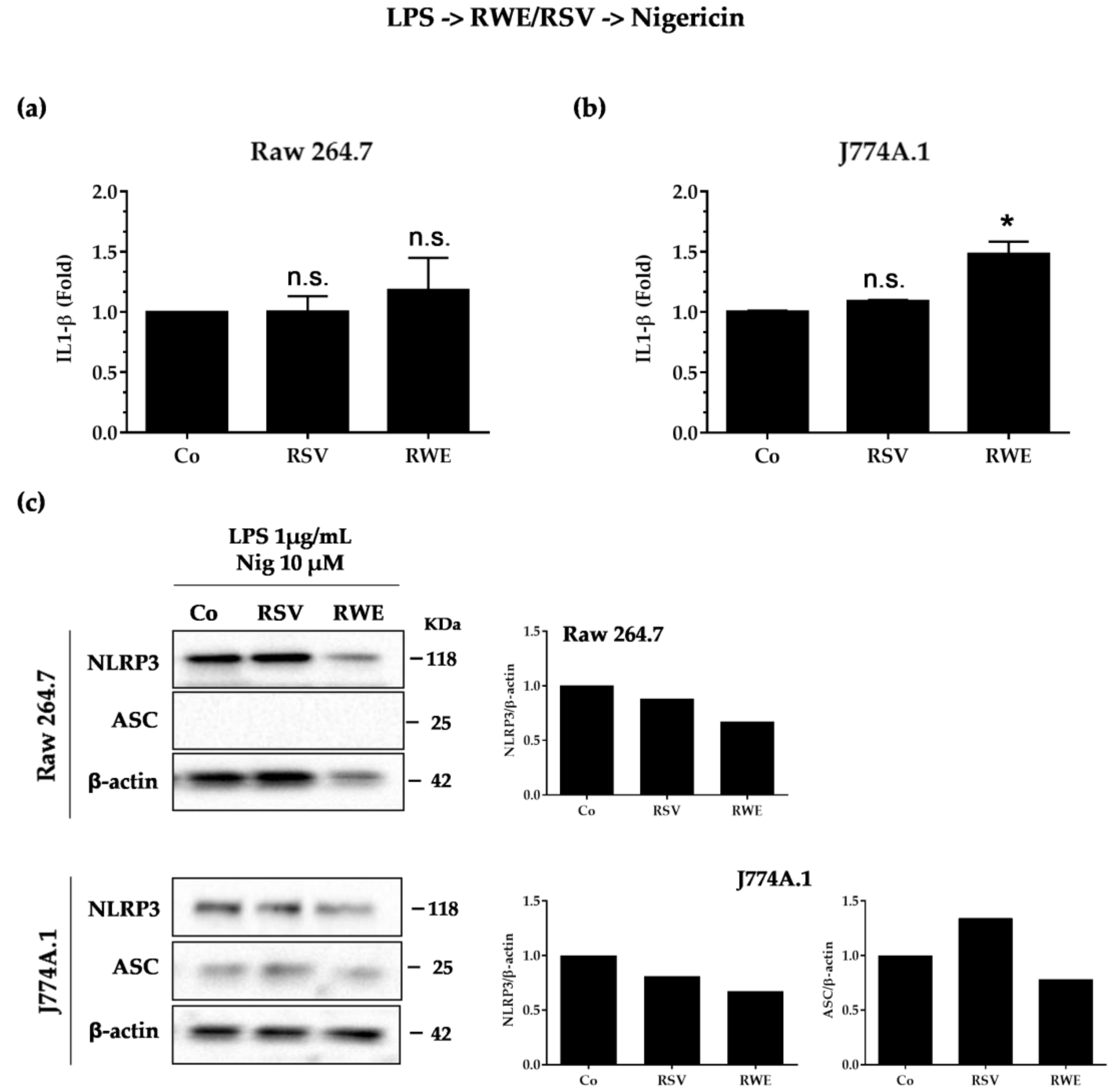
3.4. Crucial Choice of Activators for RWE and RSV Effect on IL-1β Secretion in Macrophages after LPS Priming
4. Discussion and Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Renaud, S.C.; Gueguen, R.; Schenker, J.; d’Houtaud, A. Alcohol and mortality in middle-aged men from eastern france. Epidemiology 1998, 9, 184–188. [Google Scholar] [CrossRef] [PubMed]
- Goldberg, D.M.; Soleas, G.J.; Levesque, M. Moderate alcohol consumption: The gentle face of janus. Clin. Biochem. 1999, 32, 505–518. [Google Scholar] [CrossRef]
- St Leger, A.S.; Cochrane, A.L.; Moore, F. Ischaemic heart-disease and wine. Lancet 1979, 1, 1294. [Google Scholar] [CrossRef]
- Rifler, J.P.; Lorcerie, F.; Durand, P.; Delmas, D.; Ragot, K.; Limagne, E.; Mazue, F.; Riedinger, J.M.; d’Athis, P.; Hudelot, B.; et al. A moderate red wine intake improves blood lipid parameters and erythrocytes membrane fluidity in post myocardial infarct patients. Mol. Nutr. Food Res. 2012, 56, 345–351. [Google Scholar] [CrossRef] [PubMed]
- Scalbert, A.; Johnson, I.T.; Saltmarsh, M. Polyphenols: Antioxidants and beyond. Am. J. Clin. Nutr. 2005, 81, 215S–217S. [Google Scholar] [CrossRef] [PubMed]
- Waterhouse, A.L. Wine phenolics. Ann. N. Y. Acad. Sci. 2002, 957, 21–36. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Zhang, K.; Zhang, Q.; Mei, J.; Chen, C.J.; Feng, Z.Z.; Yu, D.H. Effects of quercetin on the apoptosis of the human gastric carcinoma cells. Toxicol. In Vitro 2012, 26, 221–228. [Google Scholar] [CrossRef] [PubMed]
- Ebeler, S.E.; Brenneman, C.A.; Kim, G.S.; Jewell, W.T.; Webb, M.R.; Chacon-Rodriguez, L.; MacDonald, E.A.; Cramer, A.C.; Levi, A.; Ebeler, J.D.; et al. Dietary catechin delays tumor onset in a transgenic mouse model. Am. J. Clin. Nutr. 2002, 76, 865–872. [Google Scholar] [CrossRef] [PubMed]
- Angel-Morales, G.; Noratto, G.; Mertens-Talcott, S. Red wine polyphenolics reduce the expression of inflammation markers in human colon-derived ccd-18co myofibroblast cells: Potential role of microrna-126. Food Funct. 2012, 3, 745–752. [Google Scholar] [CrossRef] [PubMed]
- Kontou, N.; Psaltopoulou, T.; Soupos, N.; Polychronopoulos, E.; Xinopoulos, D.; Linos, A.; Panagiotakos, D. Alcohol consumption and colorectal cancer in a mediterranean population: A case-control study. Dis. Colon Rectum 2012, 55, 703–710. [Google Scholar] [CrossRef] [PubMed]
- Crockett, S.D.; Long, M.D.; Dellon, E.S.; Martin, C.F.; Galanko, J.A.; Sandler, R.S. Inverse relationship between moderate alcohol intake and rectal cancer: Analysis of the north carolina colon cancer study. Dis. Colon Rectum 2011, 54, 887–894. [Google Scholar] [CrossRef] [PubMed]
- Magalhaes, B.; Bastos, J.; Lunet, N. Dietary patterns and colorectal cancer: A case-control study from portugal. Eur. J. Cancer Prev. 2011, 20, 389–395. [Google Scholar] [CrossRef] [PubMed]
- Fira-Mladinescu, C.; Fira-Mladinescu, O.; Doroftei, S.; Sas, F.; Ursoniu, S.; Ionut, R.; Putnoky, S.; Suciu, O.; Vlaicu, B. Food intake and colorectal cancers; an ecological study in Romania. Rev. Med. Chir. Soc. Med. Nat. Iasi 2008, 112, 805–811. [Google Scholar] [PubMed]
- Andreatta, M.M.; Navarro, A.; Munoz, S.E.; Aballay, L.; Eynard, A.R. Dietary patterns and food groups are linked to the risk of urinary tract tumors in Argentina. Eur. J. Cancer Prev. 2010, 19, 478–484. [Google Scholar] [CrossRef] [PubMed]
- Chao, C.; Haque, R.; Caan, B.J.; Poon, K.Y.; Tseng, H.F.; Quinn, V.P. Red wine consumption not associated with reduced risk of colorectal cancer. Nutr. Cancer 2010, 62, 849–855. [Google Scholar] [CrossRef] [PubMed]
- Newcomb, P.A.; Nichols, H.B.; Beasley, J.M.; Egan, K.; Titus-Ernstoff, L.; Hampton, J.M.; Trentham-Dietz, A. No difference between red wine or white wine consumption and breast cancer risk. Cancer Epidemiol. Biomarkers Prev. 2009, 18, 1007–1010. [Google Scholar] [CrossRef] [PubMed]
- Colin, D.; Gimazane, A.; Lizard, G.; Izard, J.C.; Solary, E.; Latruffe, N.; Delmas, D. Effects of resveratrol analogs on cell cycle progression, cell cycle associated proteins and 5fluoro-uracil sensitivity in human derived colon cancer cells. Int. J. Cancer 2009, 124, 2780–2788. [Google Scholar] [CrossRef] [PubMed]
- Mazue, F.; Delmas, D.; Murillo, G.; Saleiro, D.; Limagne, E.; Latruffe, N. Differential protective effects of red wine polyphenol extracts (rwes) on colon carcinogenesis. Food Funct. 2014, 5, 663–670. [Google Scholar] [CrossRef] [PubMed]
- Walter, A.; Etienne-Selloum, N.; Brasse, D.; Khallouf, H.; Bronner, C.; Rio, M.C.; Beretz, A.; Schini-Kerth, V.B. Intake of grape-derived polyphenols reduces c26 tumor growth by inhibiting angiogenesis and inducing apoptosis. FASEB J. 2010, 24, 3360–3369. [Google Scholar] [CrossRef] [PubMed]
- Dolara, P.; Luceri, C.; De Filippo, C.; Femia, A.P.; Giovannelli, L.; Caderni, G.; Cecchini, C.; Silvi, S.; Orpianesi, C.; Cresci, A. Red wine polyphenols influence carcinogenesis, intestinal microflora, oxidative damage and gene expression profiles of colonic mucosa in F344 rats. Mutat. Res. 2005, 591, 237–246. [Google Scholar] [CrossRef] [PubMed]
- Caderni, G.; Remy, S.; Cheynier, V.; Morozzi, G.; Dolara, P. Effect of complex polyphenols on colon carcinogenesis. Eur. J. Nutr. 1999, 38, 126–132. [Google Scholar] [CrossRef] [PubMed]
- Chang, Y.P.; Ka, S.M.; Hsu, W.H.; Chen, A.; Chao, L.K.; Lin, C.C.; Hsieh, C.C.; Chen, M.C.; Chiu, H.W.; Ho, C.L.; et al. Resveratrol inhibits nlrp3 inflammasome activation by preserving mitochondrial integrity and augmenting autophagy. J. Cell Physiol. 2015, 230, 1567–1579. [Google Scholar] [CrossRef] [PubMed]
- Duewell, P.; Kono, H.; Rayner, K.J.; Sirois, C.M.; Vladimer, G.; Bauernfeind, F.G.; Abela, G.S.; Franchi, L.; Nunez, G.; Schnurr, M.; et al. Nlrp3 inflammasomes are required for atherogenesis and activated by cholesterol crystals. Nature 2010, 464, 1357–1361. [Google Scholar] [CrossRef] [PubMed]
- Heneka, M.T.; Kummer, M.P.; Stutz, A.; Delekate, A.; Schwartz, S.; Vieira-Saecker, A.; Griep, A.; Axt, D.; Remus, A.; Tzeng, T.C.; et al. Nlrp3 is activated in alzheimer’s disease and contributes to pathology in app/ps1 mice. Nature 2013, 493, 674–678. [Google Scholar] [CrossRef] [PubMed]
- Bruchard, M.; Mignot, G.; Derangere, V.; Chalmin, F.; Chevriaux, A.; Vegran, F.; Boireau, W.; Simon, B.; Ryffel, B.; Connat, J.L.; et al. Chemotherapy-triggered cathepsin b release in myeloid-derived suppressor cells activates the nlrp3 inflammasome and promotes tumor growth. Nat. Med. 2013, 19, 57–64. [Google Scholar] [CrossRef] [PubMed]
- Scarpioni, R.; Obici, L. Renal involvement in autoinflammatory diseases and inflammasome-mediated chronic kidney damage. Clin. Exp. Rheumatol. 2018, 36, 54–60. [Google Scholar] [PubMed]
- Nunes, C.; Teixeira, N.; Serra, D.; Freitas, V.; Almeida, L.; Laranjinha, J. Red wine polyphenol extract efficiently protects intestinal epithelial cells from inflammation via opposite modulation of jak/stat and nrf2 pathways. Toxicol. Res. 2016, 5, 53–65. [Google Scholar] [CrossRef] [PubMed]
- Biasi, F.; Guina, T.; Maina, M.; Cabboi, B.; Deiana, M.; Tuberoso, C.I.; Calfapietra, S.; Chiarpotto, E.; Sottero, B.; Gamba, P.; et al. Phenolic compounds present in sardinian wine extracts protect against the production of inflammatory cytokines induced by oxysterols in caco-2 human enterocyte-like cells. Biochem. Pharmacol. 2013, 86, 138–145. [Google Scholar] [CrossRef] [PubMed]
- Guerrero, R.F.; Garcia-Parrilla, M.C.; Puertas, B.; Cantos-Villar, E. Wine, resveratrol and health: A review. Nat. Prod. Commun. 2009, 4, 635–658. [Google Scholar] [PubMed]
- He, Y.; Ou, Z.; Chen, X.; Zu, X.; Liu, L.; Li, Y.; Cao, Z.; Chen, M.; Chen, Z.; Chen, H.; et al. LPS/TLR4 signaling enhances TGF-beta response through downregulating bambi during prostatic hyperplasia. Sci. Rep. 2016, 6, 27051. [Google Scholar] [CrossRef] [PubMed]
- Py, B.F.; Kim, M.S.; Vakifahmetoglu-Norberg, H.; Yuan, J. Deubiquitination of nlrp3 by brcc3 critically regulates inflammasome activity. Mol. Cell. 2013, 49, 331–338. [Google Scholar] [CrossRef] [PubMed]
- Petrilli, V.; Papin, S.; Dostert, C.; Mayor, A.; Martinon, F.; Tschopp, J. Activation of the NALP3 inflammasome is triggered by low intracellular potassium concentration. Cell Death Differ. 2007, 14, 1583–1589. [Google Scholar] [CrossRef] [PubMed]
- Tschopp, J.; Schroder, K. NLRP3 inflammasome activation: The convergence of multiple signalling pathways on ros production? Nat. Rev. Immunol. 2010, 10, 210–215. [Google Scholar] [CrossRef] [PubMed]
- Schroder, K.; Sagulenko, V.; Zamoshnikova, A.; Richards, A.A.; Cridland, J.A.; Irvine, K.M.; Stacey, K.J.; Sweet, M.J. Acute lipopolysaccharide priming boosts inflammasome activation independently of inflammasome sensor induction. Immunobiology 2012, 217, 1325–1329. [Google Scholar] [CrossRef] [PubMed]
- Fernandes-Alnemri, T.; Kang, S.; Anderson, C.; Sagara, J.; Fitzgerald, K.A.; Alnemri, E.S. Cutting edge: Tlr signaling licenses IRAK1 for rapid activation of the NLRP3 inflammasome. J. Immunol. 2013, 191, 3995–3999. [Google Scholar] [CrossRef] [PubMed]
- Misawa, T.; Takahama, M.; Kozaki, T.; Lee, H.; Zou, J.; Saitoh, T.; Akira, S. Microtubule-driven spatial arrangement of mitochondria promotes activation of the NLRP3 inflammasome. Nat. Immunol. 2013, 14, 454–460. [Google Scholar] [CrossRef] [PubMed]
- Baroja-Mazo, A.; Martin-Sanchez, F.; Gomez, A.I.; Martinez, C.M.; Amores-Iniesta, J.; Compan, V.; Barbera-Cremades, M.; Yague, J.; Ruiz-Ortiz, E.; Anton, J.; et al. The NLRP3 inflammasome is released as a particulate danger signal that amplifies the inflammatory response. Nat. Immunol. 2014, 15, 738–748. [Google Scholar] [CrossRef] [PubMed]
- Kanneganti, T.D.; Lamkanfi, M.; Kim, Y.G.; Chen, G.; Park, J.H.; Franchi, L.; Vandenabeele, P.; Nunez, G. Pannexin-1-mediated recognition of bacterial molecules activates the cryopyrin inflammasome independent of toll-like receptor signaling. Immunity 2007, 26, 433–443. [Google Scholar] [CrossRef] [PubMed]
- Nomura, J.; So, A.; Tamura, M.; Busso, N. Intracellular ATP decrease mediates NLRP3 inflammasome activation upon nigericin and crystal stimulation. J. Immunol. 2015, 195, 5718–5724. [Google Scholar] [CrossRef] [PubMed]
- Jhang, J.J.; Lu, C.C.; Ho, C.Y.; Cheng, Y.T.; Yen, G.C. Protective effects of catechin against monosodium urate-induced inflammation through the modulation of NLRP3 inflammasome activation. J. Agric. Food Chem. 2015, 63, 7343–7352. [Google Scholar] [CrossRef] [PubMed]
- Domiciano, T.P.; Wakita, D.; Jones, H.D.; Crother, T.R.; Verri, W.A., Jr.; Arditi, M.; Shimada, K. Quercetin inhibits inflammasome activation by interfering with asc oligomerization and prevents interleukin-1 mediated mouse vasculitis. Sci. Rep. 2017, 7, 41539. [Google Scholar] [CrossRef] [PubMed]
| mg/L of Wine | mg/L RWE | |
|---|---|---|
| Phenolic acids | ||
| Gallic acid | 21 | 5.04 |
| Caftaric acid | 53 | 12.74 |
| Coutaric acid | 20 | 4.80 |
| Caffeic acid | 10 | 2.40 |
| Stilbenes | ||
| Piceid | 1 | 0.24 |
| Resveratrol | 7 | 1.68 |
| Anthocyanidins | ||
| Delphinidin derivatives | 15 | 3.60 |
| Petunidin derivatives | 10 | 2.40 |
| Peonidin derivatives | 14 | 3.36 |
| Malvidin derivatives | 78 | 18.75 |
| Catechins | ||
| Catechin | 31 | 7.45 |
| Epicatechin | 8 | 1.92 |
| Procyanidin dimers | 42 | 10.09 |
| Flavonols | ||
| Quercetin derivatives | 4 | 0.96 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chalons, P.; Amor, S.; Courtaut, F.; Cantos-Villar, E.; Richard, T.; Auger, C.; Chabert, P.; Schni-Kerth, V.; Aires, V.; Delmas, D. Study of Potential Anti-Inflammatory Effects of Red Wine Extract and Resveratrol through a Modulation of Interleukin-1-Beta in Macrophages. Nutrients 2018, 10, 1856. https://doi.org/10.3390/nu10121856
Chalons P, Amor S, Courtaut F, Cantos-Villar E, Richard T, Auger C, Chabert P, Schni-Kerth V, Aires V, Delmas D. Study of Potential Anti-Inflammatory Effects of Red Wine Extract and Resveratrol through a Modulation of Interleukin-1-Beta in Macrophages. Nutrients. 2018; 10(12):1856. https://doi.org/10.3390/nu10121856
Chicago/Turabian StyleChalons, Pauline, Souheila Amor, Flavie Courtaut, Emma Cantos-Villar, Tristan Richard, Cyril Auger, Philippe Chabert, Valérie Schni-Kerth, Virginie Aires, and Dominique Delmas. 2018. "Study of Potential Anti-Inflammatory Effects of Red Wine Extract and Resveratrol through a Modulation of Interleukin-1-Beta in Macrophages" Nutrients 10, no. 12: 1856. https://doi.org/10.3390/nu10121856
APA StyleChalons, P., Amor, S., Courtaut, F., Cantos-Villar, E., Richard, T., Auger, C., Chabert, P., Schni-Kerth, V., Aires, V., & Delmas, D. (2018). Study of Potential Anti-Inflammatory Effects of Red Wine Extract and Resveratrol through a Modulation of Interleukin-1-Beta in Macrophages. Nutrients, 10(12), 1856. https://doi.org/10.3390/nu10121856







