1. Introduction
Hyperspectral images (HSIs), acquired by advanced sensors on satellites, aerial vehicles, or drones [
1,
2,
3], provide continuous spectral curves for each pixel. This rich spectral information enables the precise identification of materials, making HSIs indispensable in mineral exploration [
4], urban planning [
5], precision agriculture [
6], environmental monitoring [
7], and military reconnaissance [
8]. However, the very advantage of HSIs—their high dimensionality—also introduces the “curse of dimensionality” [
9]. More critically, real-world HSI scenes are characterized by significant spatial–spectral heterogeneity, where the same material may exhibit varying spectral signatures due to environmental changes, and land covers often present irregular geometries and scale variations [
10]. Consequently, effectively utilizing this complex data for accurate classification remains a significant challenge.
Deep learning has revolutionized HSI classification by automatically learning hierarchical representations [
11,
12], moving beyond handcrafted features. Among these techniques, Convolutional Neural Networks (CNNs) have become the dominant backbone. Early works, such as Li et al. [
13], employed 1D CNNs to extract spectral signatures. Recognizing that HSIs are volumetric data, subsequent studies integrated spatial context. For instance, Zhang et al. [
14] and Xu et al. [
15] utilized dual-branch architectures to extract spectral and spatial features respectively. While these methods improved performance, they typically rely on standard convolution operations defined on a fixed, rigid grid. This inherent rigidity assumes that relevant features always lie within a regular rectangular neighborhood, ignoring the fact that object boundaries in HSIs are often curved or fragmented. As a result, standard CNNs often fail to capture the intrinsic geometric deformations of objects, leading to feature misalignment at boundaries [
16].
To address this, Li et al. [
17] partitioned HSIs into multiple 3D cubes and applied 3D CNNs to simultaneously perform convolutions along spatial and spectral dimensions. However, employing stacked 3D CNNs significantly increases parameter count and leads to gradient vanishing issues [
18,
19,
20]. Therefore, Roy et al. [
21] proposed a hybrid 3D-2D convolution approach to reduce network complexity. Zhong et al. [
22] introduced residual structures [
23] into 3D spectral and 2D spatial convolutions. To further address the geometric limitations of standard convolutions, Dai et al. [
24] introduced deformable convolutions by adding offsets to standard 2D convolutions, enabling adaptive sampling. Yu and Vladlen [
25] proposed an efficient method to enlarge convolutional receptive fields. While these techniques alleviate the constraints of fixed grids, modeling long-range dependencies remains a critical challenge for CNNs.
In recent years, Transformer architectures have been introduced into the field of computer vision and achieved impressive results [
26]. Unlike CNNs that primarily capture local features, Transformers can capture long-range dependencies between features with global features [
27]. In the context of HSI classification, Hong et al. [
28] reformulated the task as a sequence modeling problem and proposed a spectral Transformer, surpassing classical ViT. Similarly, Qing et al. [
29] exploited spectral attention and self-attention mechanisms, while Liu et al. [
30] designed a hierarchical Transformer with shifted windows to enable multi-scale feature extraction with reduced computational redundancy. Moreover, interactive learning frameworks, such as the Center Transformer [
31], have been developed to capture multi-scale spatial–spectral representations by interacting features from center to surrounding regions.
In addition, hybrid CNN-Transformer architectures have gained popularity for combining local and global feature modeling. For instance, Sun et al. [
32] proposed SSFTT, where a Transformer encoder processes spectral–spatial features extracted from hierarchical 3D and 2D convolutional blocks. Similarly, Fu et al. [
33] constructed parallel CNN and Transformer branches to integrate local and non-local features. Xu et al. [
34] developed a novel Transformer architecture that incorporates embedded convolution modules to adaptively fuse features from diverse receptive fields. Roy et al. [
35] designed learnable spectral and spatial morphological networks using morphological convolutions combined with attention mechanisms. Yang et al. [
36] proposed a two-stream CNN using 2D and 3D convolutions for local feature extraction, followed by a Transformer to model global dependencies. To alleviate the computational burden and overfitting of Transformers, Woo et al. [
37] introduced channel and spatial attention modules using convolution operations to emulate attention effects. Furthermore, Zhang et al. [
38] proposed a cascaded spatial cross-attention network that simultaneously captures local and global spatial contextual features via cross-attention. Beyond the aforementioned architectures, the HSI classification field has recently witnessed significant progress in several other advanced dimensions. For instance, enhanced multiscale feature fusion networks [
39] have been developed to capture robust spatial–spectral representations. To alleviate the reliance on massive labeled data, weakly supervised paradigms like the ITER framework [
40] have been explored to generate effective image-to-pixel representations. Furthermore, with the advent of large-scale deep learning, vision transformer-based foundation models, such as HyperSIGMA [
41], have emerged to unify HSI interpretation across diverse and complex scenes.
However, despite these advances, existing methods still face challenges. First, standard Transformers suffer from quadratic computational complexity (), which restricts efficiency and scalability. Second, in HSI scenarios with limited samples, they are prone to overfitting. Third, standard downsampling methods in these hierarchical networks often cause the irreversible loss of fine-grained details, leading to the disappearance of small-scale objects.
To overcome these challenges—specifically geometric rigidity, high computational complexity, and information loss during downsampling—we propose an innovative hierarchical framework named Multiscale Deformable Spectral–Spatial Sequence Network (MDS3-Net). Unlike previous methods, MDS3-Net introduces a synergistic design that balances local adaptivity and global efficiency. Specifically, we design a Multiscale Spectral-Deformable Convolution (MSDC) module to simultaneously extract discriminative spectral features and adaptively align spatial features with irregular object boundaries. To resolve the quadratic complexity of Transformers, we introduce a Spectral–Spatial Sequence (S3) Encoder based on a gated convolutional mechanism, which captures long-range dependencies with linear complexity (). Furthermore, a Dual-Path Feature Extraction (DPFE) module is proposed to perform dimension reduction while preserving salient spectral–spatial information.
The main contributions of this paper are summarized as follows:
- (1)
We propose MDS3-Net, a novel unified hierarchical framework that synergizes local geometric adaptability with efficient global modeling, achieving state-of-the-art HSI classification performance even under limited training samples.
- (2)
We design a MSDC module that decouples spectral and spatial feature extraction, enabling effective spectral discrimination and dynamic alignment with irregular object boundaries.
- (3)
We introduce a S3 Encoder that utilizes a gated large-kernel convolution mechanism to capture global long-range dependencies with linear computational complexity (), overcoming the heavy computational burden of traditional self-attention.
- (4)
We propose a DPFE module as a semantics-preserving downsampling mechanism, which performs dimensionality reduction via spatial attention and spectral reorganization to prevent the loss of fine-grained details.
The remainder of the paper is organized as follows.
Section 2 elaborates on the proposed MDS
3-Net framework and its core components.
Section 3 details the experimental setup, datasets, and comprehensive performance analysis.
Section 4 presents a further discussion on the experimental results. Finally,
Section 5 concludes the paper with a summary of findings and future directions.
2. Methodology
2.1. Overall Architecture
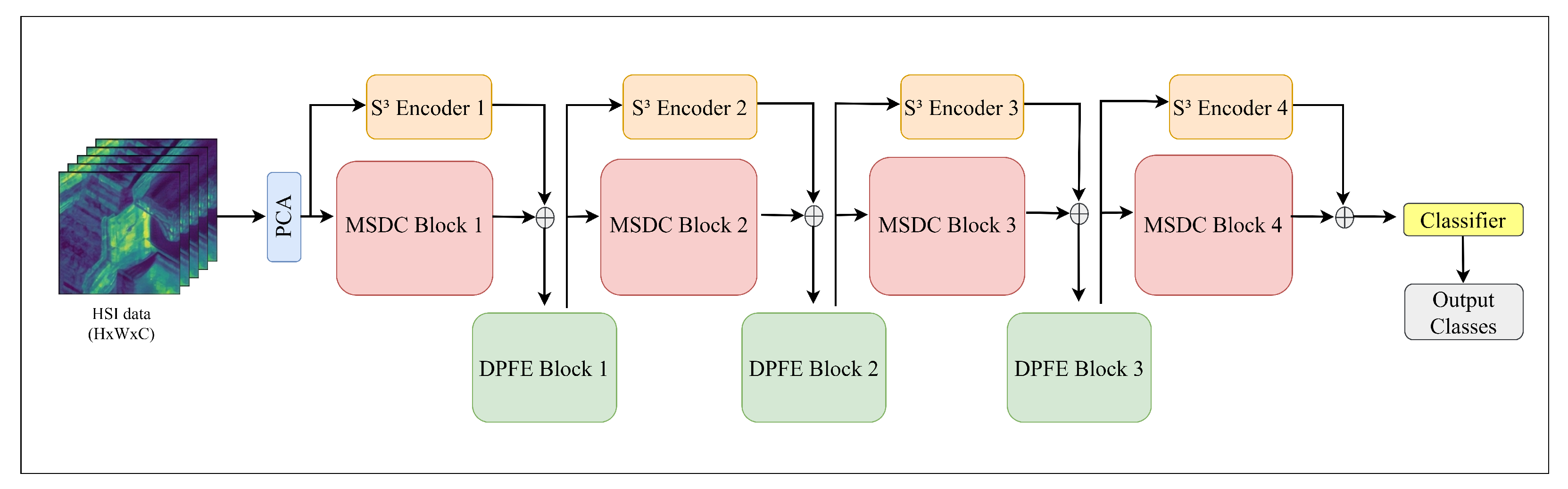
We present MDS
3-Net, an innovative hierarchical framework designed for hyperspectral image classification, with the detailed architecture depicted in
Figure 1. The MDS
3-Net architecture incorporates three key synergistic components: the MSDC module, the S
3 Encoder, and the DPFE module.
Prior to feature extraction, to mitigate the curse of dimensionality and spectral redundancy, we first perform Principal Component Analysis (PCA) on the raw hyperspectral imagery. Subsequently, we extract spatial neighborhoods from the dimension-reduced data, generating multiple 3D image patches denoted as , where represents the spatial dimensions and B is the number of spectral bands. These patches serve as the input for the subsequent stage-wise hierarchical processing.
The proposed framework is designed to progressively extract and integrate spectral–spatial information through a pyramidal structure. Within each processing stage, we adopt a dual-branch strategy to simultaneously capture local and global information. Specifically, the MSDC module serves as the primary extractor, employing a decoupled strategy that combines spectral convolution for discriminative spectral features with deformable convolution for geometric adaptability. Simultaneously, the S3 Encoder operates along a parallel residual path, sharing the same input as the MSDC module. It models long-range sequential dependencies via a gated convolutional mechanism, achieving global receptive fields with linear computational complexity. The local features from the MSDC and the global context from the S3 Encoder are then fused via element-wise addition. Subsequently, this fused representation is fed into the DPFE module, which serves as a semantics-preserving downsampling mechanism. By prioritizing salient information preservation during resolution reduction, the DPFE effectively bridges adjacent stages.
Overall, this architectural design adheres to the principle of complementary feature learning, wherein each component performs a distinct yet cooperative role to strengthen the joint spectral–spatial representation. The MSDC emphasizes both spectral fidelity and local structural adaptability; the S3 Encoder facilitates global contextual awareness while maintaining high computational efficiency; and the DPFE functions as a critical filtering mechanism to ensure salient information preservation during spatial and spectral dimension reduction. Through the progressive integration of these complementary cues, MDS3-Net achieves a balanced synergy between local detail preservation, global semantic understanding, and model efficiency.
2.2. MSDC
As the fundamental feature extraction unit of MDS
3-Net, the MSDC module is engineered to address the inherent spectral–spatial coupling and geometric complexity of hyperspectral data.
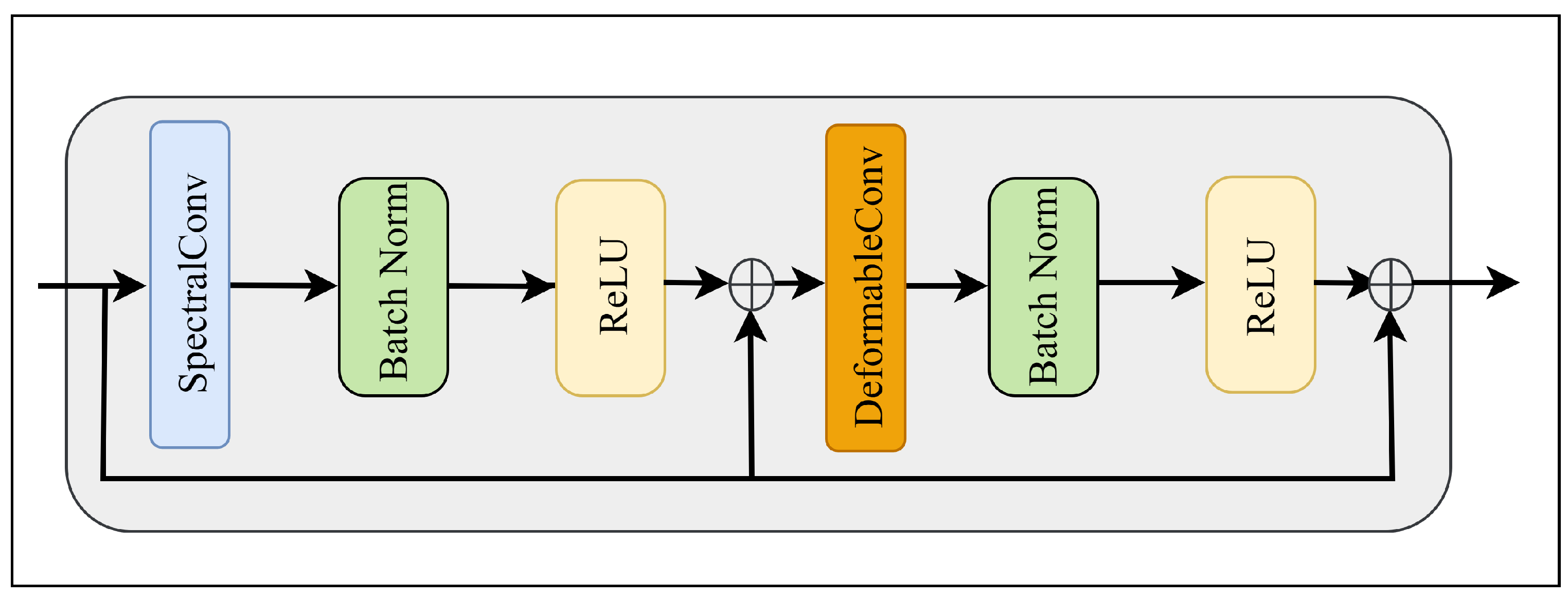
Figure 2 illustrates the internal structure of the MSDC block. It adopts a decoupled strategy combining spectral convolution and deformable spatial convolution, augmented with dual residual connections to facilitate feature reuse and gradient propagation.
The module first applies a spectral convolution with a kernel size of
, meaning the operation is performed exclusively in the spectral dimension to aggregate spectral information without altering the spatial structure. To preserve the original spectral fidelity and prevent network degradation, a residual connection is introduced. The intermediate output
is formulated as:
where
x is the input feature,
denotes the 3D convolution with a spatial kernel size of
, BN represents Batch Normalization [
42], and
denotes the ReLU activation function [
43].
Subsequently, the spatial features
are processed by a deformable convolution with a kernel size of
. Unlike standard convolutions that sample from a fixed grid, deformable convolution introduces learnable offsets to dynamically adjust the sampling positions [
24]:
where
denotes the regular sampling grid,
W represents the convolution weights, and
is the learnable offset. Similar to the first stage, a second residual connection is applied to the spatial branch. The final output of the MSDC block is obtained by:
where
and
denote the output and input feature maps of the residual connection, respectively.
represents the deformable convolution operation.
and
denote the operations defined previously. Notably, the symbol + denotes element-wise addition rather than feature concatenation. This operation acts as a standard residual connection to refine the representations while perfectly preserving the original channel dimensions, thereby avoiding the drastic increase in computational complexity that concatenation would cause in subsequent deep layers.
To capture features at varying scales and receptive fields, the MSDC module employs a hierarchical kernel configuration. Specifically, regarding the implementation details, in the shallow stages (Block 1 and Block 2), we utilize a smaller kernel size () to capture fine-grained texture and local spectral variations. Conversely, in the deeper stages (Block 3 and Block 4), the kernel size is increased () to expand the receptive field and encapsulate broader semantic context. This multiscale design enables the network to effectively recognize objects of various sizes, ranging from small targets to large homogeneous regions.
2.3. S3 Encoder
While the MSDC module excels at extracting local spectral–spatial features, it inherently lacks the ability to model global contextual dependencies due to its limited receptive field [
44]. To address this limitation, we introduce the S
3 Encoder. Unlike traditional Transformers that suffer from quadratic computational complexity with respect to token length, the S
3 Encoder models long-range interactions with linear complexity via a gated convolutional mechanism.
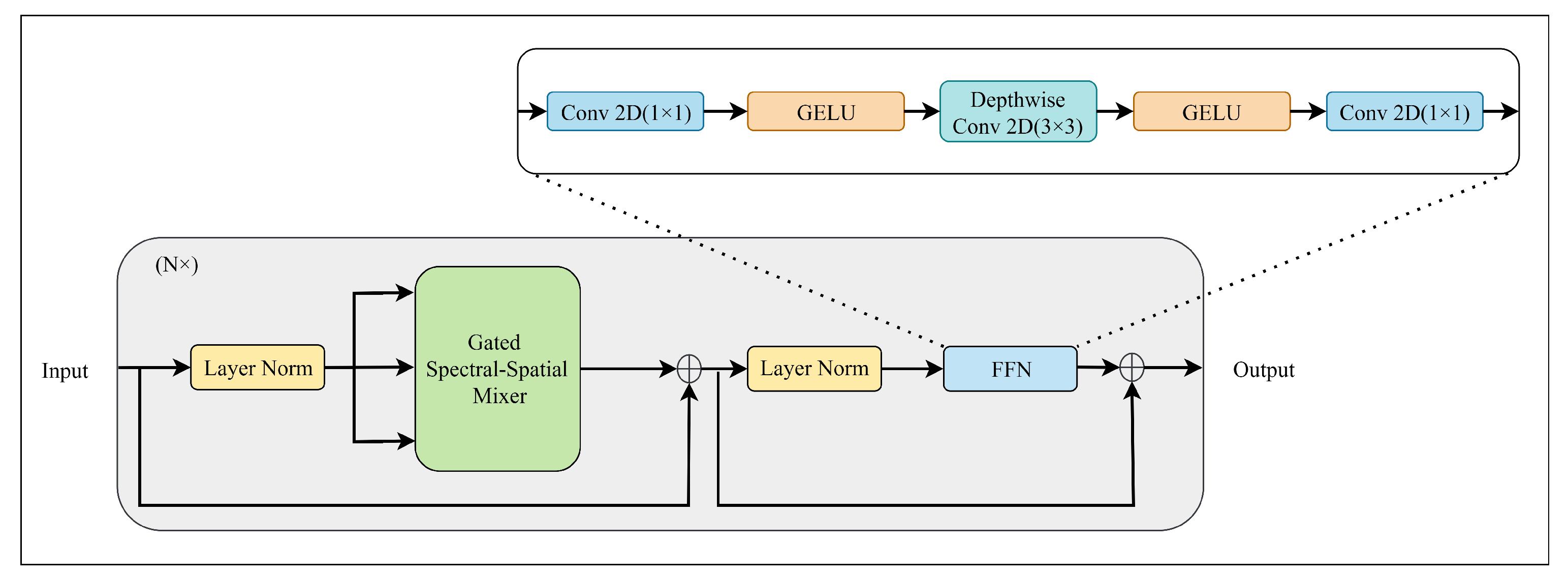
As shown in
Figure 3, the encoder block comprises two synergistic sub-modules: the Gated Spectral–Spatial Mixer (GS
2M) and the Feed-Forward Network (FFN). Layer Normalization (LN) [
45] is applied before each sub-module to normalize feature distributions, and residual connections are employed after each block. This design effectively alleviates the vanishing gradient problem during the training of deep networks.
2.3.1. GS2M
The GS
2M is specifically designed to replace the computationally intensive Multi-Head Self-Attention (MHSA). As depicted in
Figure 4, it adopts a large-kernel convolution combined with a gating mechanism to efficiently aggregate global context.
Given a normalized input feature map
, the module first projects it into a hidden representation using a
convolution. This representation is then split along the channel dimension into two parallel branches: the gating branch and the feature branch. The gating branch utilizes a depthwise convolution with a large kernel size (
) to capture broad spatial cues, followed by a GELU activation to generate a spatial attention map. Simultaneously, the feature branch retains the local spectral details. The attention map then modulates the feature branch via element-wise multiplication. The mathematical formulation is defined as:
where ⊙ denotes the element-wise multiplication and
is the output projection layer. This gating design allows the model to adaptively select spectral–spatial features based on global context while maintaining a linear computational complexity of
, where
N is the number of pixels.
2.3.2. FFN
As illustrated in
Figure 3, the output of the GS
2M is subsequently processed by the FFN. Standard FFNs in Transformers typically operate in a pixel-wise manner (using two
convolutions), which may overlook local structural details. To mitigate this, our FFN integrates a
depthwise convolution within the expansion layer. This locality-enhanced design ensures that fine-grained texture information is preserved and refined during the channel mixing process. The FFN can be expressed as:
where
denotes the GELU activation. This modification effectively complements the global modeling capability of the GS
2M, creating a comprehensive feature encoder.
2.4. DPFE
Downsampling is a pivotal operation in hierarchical networks. However, standard pooling methods often result in the irreversible degradation of fine-grained details, leading to the disappearance of small-scale objects. To address this, we propose the DPFE module. Distinct from the S3 Encoder which focuses on global feature modeling, the DPFE functions as a downsampling mechanism dedicated to preserving key spectral and spatial information. It is explicitly designed to filter background noise and minimize the loss of salient features during spatial and spectral dimension reduction.
As illustrated in
Figure 5, the DPFE module operates through two parallel paths. The spectral reorganization path is designed to perform linear spectral transformation. It employs a
convolution to project the input spectral features onto the target dimension, followed by a
Average Pooling layer to perform spatial downsampling. This path ensures that essential spectral context is efficiently transferred during the spatial and spectral dimension reduction process.
Simultaneously, the spatial squeeze path functions as a global attention filter. It first reduces channel dimensionality via a
convolution to generate intermediate features
. These features are then processed by a large-kernel depthwise convolution (
) and a Sigmoid activation to generate a spatial attention map. This map modulates
via element-wise multiplication, effectively suppressing background noise and highlighting salient regions. Finally, the refined features undergo spatial downsampling identical to the spectral reorganization path. The outputs from both paths are fused via element-wise addition to integrate local spectral details with global salient semantics. The operation is summarized as:
where
and
denote the intermediate features generated by the spectral reorganization path and the spatial squeeze path, respectively. Pool represents the
Average Pooling operation, and
refers to the Sigmoid activation function. Specifically,
and
correspond to the distinct pointwise convolutions utilized in these two respective paths.
4. Discussion
4.1. Mechanism Analysis of Performance Superiority
The extensive experimental results in
Section 3 validate that MDS
3-Net outperforms current state-of-the-art methods. Beyond the numerical improvements, it is crucial to understand the underlying mechanisms driving this success. The superiority of MDS
3-Net primarily stems from its ability to synergistically address HSI classification challenges through three synergistic dimensions: the unified extraction of local spatial–spectral features, the efficient modeling of global long-range dependencies, and the preservation of salient information during dimensionality reduction.
First and foremost, the MSDC module serves as the core engine for joint spectral–spatial feature extraction. Unlike standard CNNs that utilize fixed square kernels and treat spatial–spectral dimensions rigidly, the MSDC module introduces a dual-enhancement mechanism. Regarding spatial adaptability, land cover objects—such as the complex urban structures in HS2013 or winding roads in Trento—often exhibit irregular shapes that do not conform to fixed grids. The deformable mechanism in MSDC decouples the receptive field, allowing sampling locations to dynamically align with object boundaries, thereby effectively reducing “mixed pixel” interference at edges. Concurrently, in terms of spectral discrimination, the MSDC employs a cascaded strategy. It first applies multiscale spectral convolution to extract discriminative spectral signatures across different local band ranges. These spectrally refined features are then sequentially processed by the spatial deformable convolution. This serial design ensures that the network distinguishes between materials with subtle spectral discrepancies before performing geometric alignment. Therefore, the high classification accuracy of MDS3-Net in heterogeneous regions is a direct result of this synergy—MSDC refines features spectrally and then aligns them spatially.
Second, the ablation study (
Table 6) confirms the necessity of the S
3 Encoder. While MSDC excels at local spectral–spatial extraction, pure convolutional operations inherently struggle to capture long-range dependencies. The S
3 Encoder compensates for this by utilizing a gated large-kernel mechanism to model global sequential relationships across the spectral–spatial domain. Unlike traditional Transformers that rely on computationally intensive self-attention, the S
3 Encoder achieves this global modeling with linear complexity. Consequently, the architecture establishes a complementary hierarchy: both localized and adaptive feature extraction through MSDC, and efficient global context refinement through S
3 Encoder as a supplement.
Third, the DPFE module plays a critical role in maintaining feature integrity during downsampling. In conventional hierarchical networks, standard pooling operations often lead to the irreversible loss of fine-grained details, causing small-scale objects to disappear in deeper layers. The DPFE addresses this by employing a dual-path strategy: a spectral reorganization path to strictly preserve spectral context and a spatial squeeze path to highlight salient regions via spatial attention. By filtering background noise while retaining key semantic information during dimension reduction, the DPFE effectively bridges adjacent stages, ensuring that the network maintains high distinctiveness even for small targets or complex boundaries.
4.2. Architectural Efficiency and Practicality
In the realm of HSI classification, achieving a balance between high accuracy and low computational cost is a pivotal consideration for practical deployment. The efficiency of the proposed MDS3-Net stems from its strategic architectural design.
Specifically, the efficiency and practicality of MDS3-Net are realized through the layer-wise synergistic operation of its three core components. First, within each processing stage, the MSDC module and the S3 Encoder work in a complementary manner. The MSDC utilizes decoupled convolutions to efficiently extract dense local features. Simultaneously, the S3 Encoder, strategically embedded in the residual path, captures global context to rectify these local representations. Distinct from standard ViTs, our S3 Encoder avoids quadratic complexity (), ensuring that the overhead of global modeling remains manageable.
Second, connecting these stages is the DPFE module, which acts as a semantics-preserving compressor. Unlike standard pooling layers that indiscriminately discard information, the DPFE employs a dual-path strategy: a spectral reorganization path to linearly project spectral dimensions and a spatial squeeze path to filter background noise via spatial attention. This hierarchical architecture optimizes the allocation of computational resources. By progressively reducing feature resolution while preserving salient information through DPFE, the network ensures that deeper layers operate on compact, high-level semantic embeddings. This “coarse-to-fine” processing flow allows MDS3-Net to retain powerful global modeling capabilities without incurring the prohibitive costs associated with full-resolution processing, thereby achieving an optimal balance between inference speed and classification accuracy.
4.3. Limitations
Despite the superior classification performance and competitive efficiency achieved by MDS3-Net, there remain limitations regarding model complexity that warrant further discussion. Although MDS3-Net is significantly faster than standard Transformer-based methods, it inevitably incurs higher storage and computational costs compared to extremely lightweight CNNs (such as SSRN). Specifically, the calculation of learnable offsets in the MSDC module and the large-kernel depthwise convolutions in the S3 Encoder require more floating-point operations than simple static convolutions. This reflects a necessary trade-off to achieve high-precision classification in complex scenes. To address this, future work will focus on developing lightweight versions of MDS3-Net, thereby further reducing the resource overhead to enhance deployability on resource-constrained platforms.
5. Conclusions
In this paper, we have proposed MDS3-Net, a novel hierarchical framework designed to address the dual challenges of geometric rigidity and computational inefficiency in HSI classification. By synergizing the MSDC with the S3 Encoder, our method effectively unifies the adaptive extraction of local spectral–spatial features and the modeling of global long-range dependencies with linear computational complexity. Additionally, the DPFE module functions as a critical bridge between stages, facilitating semantics-preserving dimensionality reduction through spectral reorganization and spatial attention mechanisms.
Extensive experiments on four benchmark datasets (UP, HS2013, LK, and UT) demonstrate that MDS3-Net consistently achieves state-of-the-art classification performance, particularly in scenes characterized by complex geometric boundaries and significant spectral variability. Quantitative comparisons and ablation studies further validate the necessity of each synergistic component—MSDC for joint spectral discrimination and geometric alignment, S3 Encoder for efficient global context modeling, and DPFE for semantics-preserving dimensionality reduction. Moreover, the complexity analysis confirms that MDS3-Net attains an optimal trade-off between accuracy and efficiency, significantly outperforming standard Transformer-based models in terms of training speed while maintaining superior precision.
In future work, we intend to focus on developing lightweight versions of MDS3-Net. This will aim to reduce the parameter count and computational overhead identified in the limitations, thereby further facilitating the deployment of high-performance HSI classification models on resource-constrained edge devices.