Designing a European-Wide Crop Type Mapping Approach Based on Machine Learning Algorithms Using LUCAS Field Survey and Sentinel-2 Data
Abstract
:1. Introduction
- To evaluate the potential of using the LUCAS 2018 data in combination with S2 data for generating detailed LULC maps.
- To compare the classification performance of S2 and S1, using as [16] a reference study for S1.
- To compare the performance of RF and SVM classifiers.
- To study the impact on the classification accuracy due to autocorrelation in field data (considering data sampled from the LUCAS Copernicus polygon geometries).
2. Materials and Methods
2.1. Sentinel-2: Data Preparation, Spectral and Temporal Features
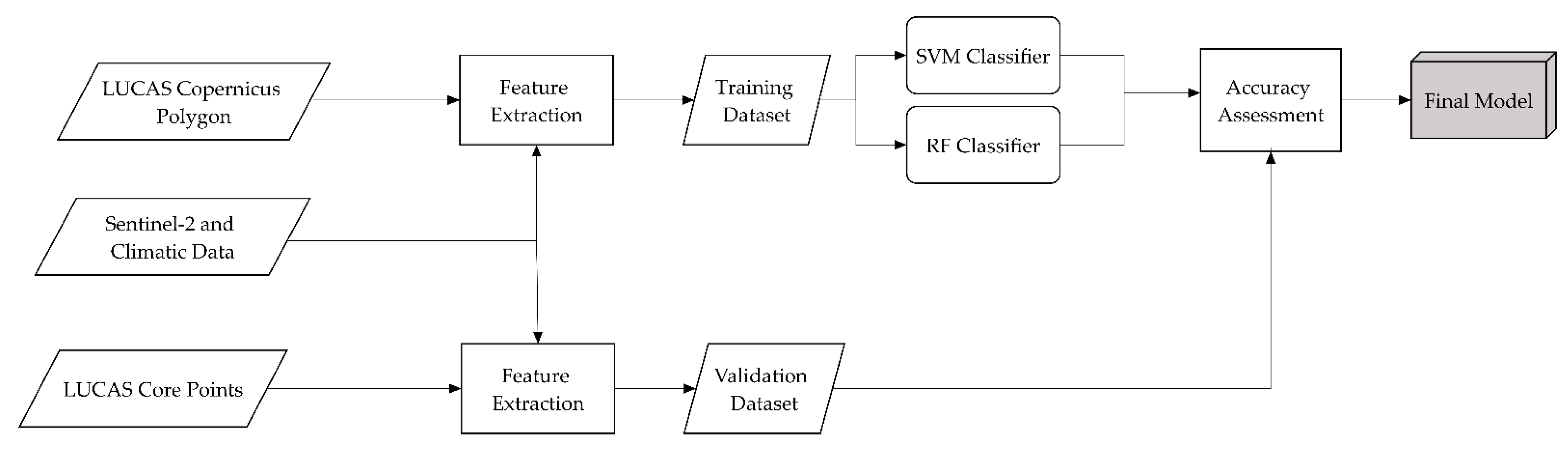
2.2. Preparation of the Training Data
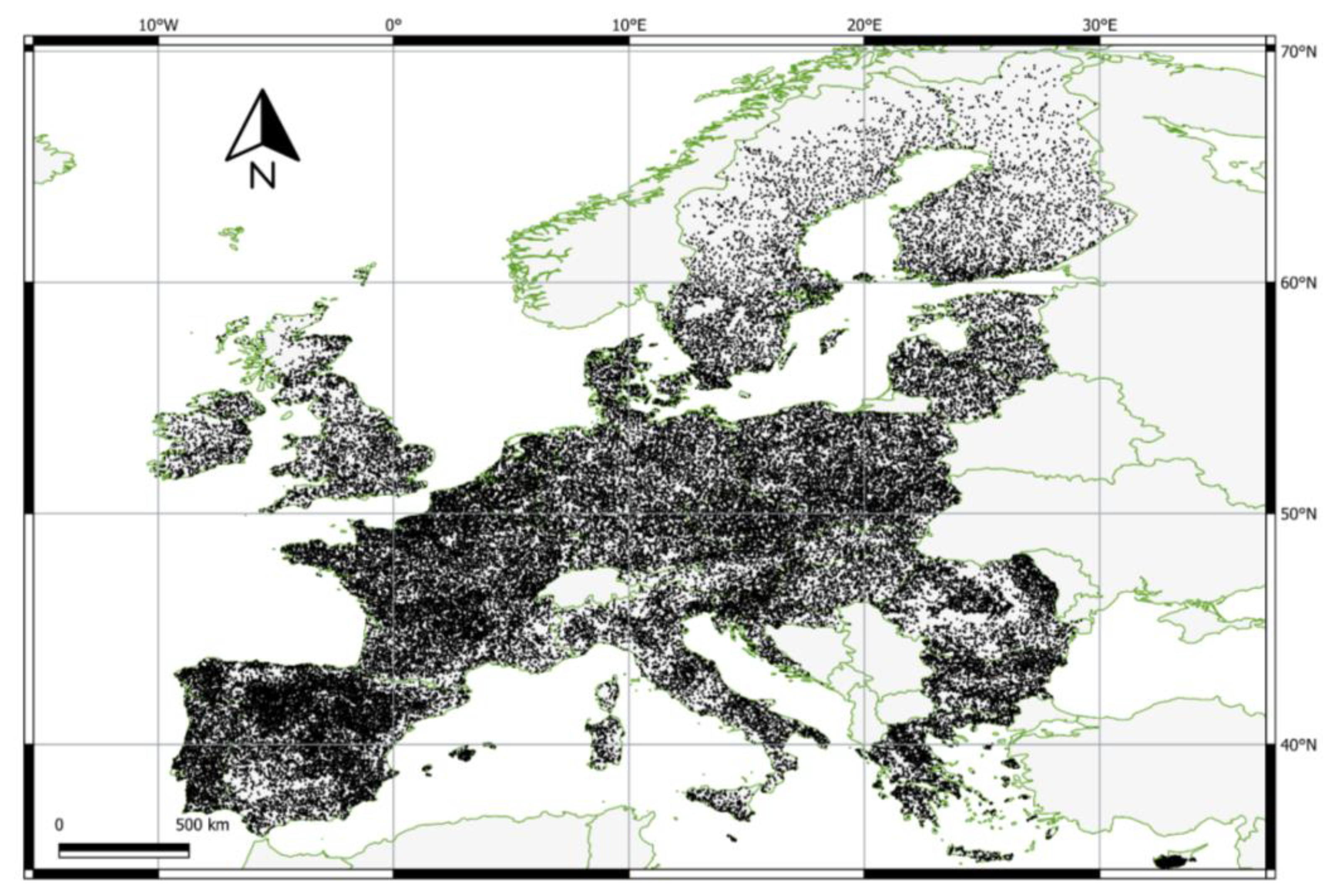
2.2.1. European Land Use and Cover Area Frame Survey (LUCAS) 2018 Data
2.2.2. LUCAS 2018 Data Refinement Due to Gaps in Sentinel-2 Data
2.2.3. LUCAS 2018 Data Refinement Related to the Classification Scheme
2.3. Classification Process
2.3.1. Classification Methods
2.3.2. Training Data Sub-Setting
- Proportional (One random sample from each polygon)
- Balanced
- Dissimilar
2.3.3. Feature Selection
2.3.4. Hyperparameter Tuning
2.4. Accuracy Assessment
2.4.1. Validation Data
2.4.2. Assessment Metrics
3. Results
3.1. Selecting the Best Classifier and Dataset
3.2. The Accuracy Assessment of the Best Classification Model
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Appendix A
| Code | 211 | 212 | 213 | 214 | 215 | 216 | 217 | 218 | 219 | 221 | 222 | 223 | 230 | 231 | 232 | 233 | 240 | 250 | 290 | 300 | 500 | Total | UA | F1-Score |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 211 | 3855 | 326 | 839 | 309 | 212 | 74 | 1 | 280 | 16 | 18 | 17 | 5 | 47 | 19 | 123 | 1 | 66 | 110 | 513 | 99 | 505 | 7435 | 51.8% | 62.9% |
| 212 | 51 | 224 | 37 | 1 | 19 | 1 | 0 | 3 | 0 | 0 | 0 | 1 | 1 | 1 | 1 | 0 | 10 | 11 | 18 | 5 | 25 | 409 | 54.8% | 34.4% |
| 213 | 388 | 112 | 1430 | 81 | 164 | 20 | 0 | 43 | 9 | 6 | 4 | 9 | 14 | 1 | 56 | 0 | 58 | 93 | 166 | 29 | 219 | 2902 | 49.3% | 49.4% |
| 214 | 41 | 3 | 28 | 135 | 20 | 1 | 0 | 59 | 1 | 0 | 0 | 0 | 1 | 0 | 11 | 0 | 1 | 12 | 13 | 2 | 25 | 353 | 38.2% | 26.5% |
| 215 | 17 | 2 | 36 | 9 | 54 | 0 | 0 | 2 | 0 | 0 | 0 | 0 | 1 | 0 | 1 | 0 | 4 | 12 | 9 | 2 | 25 | 174 | 31.0% | 12.3% |
| 216 | 45 | 4 | 31 | 10 | 11 | 3058 | 14 | 5 | 30 | 36 | 12 | 8 | 7 | 22 | 8 | 36 | 23 | 34 | 67 | 65 | 183 | 3709 | 82.4% | 85.4% |
| 217 | 0 | 0 | 0 | 0 | 0 | 0 | 19 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 19 | 100.0% | 54.3% |
| 218 | 0 | 0 | 1 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 2 | 0.0% | 0.0% |
| 219 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 8 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 9 | 88.9% | 11.8% |
| 221 | 0 | 0 | 4 | 1 | 0 | 9 | 0 | 1 | 0 | 230 | 6 | 0 | 7 | 4 | 0 | 8 | 10 | 5 | 6 | 0 | 6 | 297 | 77.4% | 68.7% |
| 222 | 2 | 0 | 7 | 0 | 0 | 4 | 0 | 0 | 7 | 8 | 507 | 12 | 5 | 7 | 3 | 0 | 14 | 1 | 3 | 2 | 10 | 592 | 85.6% | 84.4% |
| 223 | 0 | 0 | 0 | 0 | 0 | 0 | 2 | 0 | 0 | 0 | 1 | 5 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 9 | 55.6% | 9.8% |
| 230 | 0 | 1 | 0 | 0 | 1 | 2 | 2 | 0 | 0 | 0 | 0 | 0 | 88 | 0 | 2 | 1 | 11 | 1 | 4 | 0 | 2 | 115 | 76.5% | 44.3% |
| 231 | 3 | 2 | 2 | 0 | 3 | 15 | 0 | 0 | 2 | 14 | 2 | 5 | 22 | 431 | 3 | 5 | 38 | 7 | 22 | 4 | 52 | 632 | 68.2% | 73.1% |
| 232 | 12 | 1 | 18 | 3 | 1 | 8 | 0 | 4 | 4 | 3 | 7 | 3 | 10 | 1 | 1105 | 3 | 20 | 4 | 100 | 8 | 62 | 1377 | 80.2% | 78.4% |
| 233 | 0 | 0 | 2 | 0 | 0 | 4 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 1 | 1 | 63 | 1 | 1 | 1 | 0 | 2 | 77 | 81.8% | 60.6% |
| 240 | 10 | 7 | 22 | 3 | 7 | 14 | 4 | 1 | 6 | 25 | 21 | 21 | 21 | 17 | 24 | 1 | 203 | 31 | 56 | 4 | 53 | 551 | 36.8% | 33.8% |
| 250 | 5 | 6 | 8 | 3 | 11 | 7 | 0 | 0 | 2 | 4 | 1 | 2 | 4 | 1 | 1 | 3 | 19 | 405 | 11 | 9 | 71 | 573 | 70.7% | 34.3% |
| 290 | 23 | 21 | 46 | 1 | 14 | 4 | 2 | 1 | 1 | 3 | 3 | 3 | 1 | 1 | 14 | 0 | 19 | 6 | 347 | 14 | 118 | 642 | 54.0% | 26.8% |
| 300 | 72 | 27 | 63 | 11 | 27 | 61 | 2 | 5 | 6 | 2 | 9 | 1 | 12 | 9 | 11 | 2 | 40 | 95 | 179 | 25,321 | 2715 | 28,670 | 88.3% | 90.4% |
| 500 | 304 | 158 | 315 | 96 | 162 | 167 | 5 | 35 | 35 | 23 | 20 | 18 | 40 | 33 | 79 | 8 | 112 | 963 | 428 | 1775 | 17,477 | 22,253 | 78.5% | 79.8% |
| Total | 4828 | 894 | 2889 | 664 | 706 | 3449 | 51 | 439 | 127 | 373 | 610 | 93 | 282 | 548 | 1443 | 131 | 649 | 1791 | 1943 | 27,339 | 21,551 | 70,800 | OA = 77.6% | |
| PA | 79.8% | 25.1% | 49.5% | 20.3% | 7.6% | 88.7% | 37.3% | 0.0% | 6.3% | 61.7% | 83.1% | 5.4% | 31.2% | 78.6% | 76.6% | 48.1% | 31.3% | 22.6% | 17.9% | 92.6% | 81.1% | |||
References
- Bonan, G.B. Forests and climate change: Forcings, feedbacks, and the climate benefits of forests. Science 2008, 320, 1444–1449. [Google Scholar] [CrossRef] [Green Version]
- Brovkin, V.; Sitch, S.; von Bloh, W.; Claussen, M.; Bauer, E.; Cramer, W. Role of land cover changes for atmospheric CO2 increase and climate change during the last 150 years. Glob. Chang. Biol. 2004, 10, 1253–1266. [Google Scholar] [CrossRef] [Green Version]
- Foley, J.A.; Defries, R.; Asner, G.P.; Barford, C.; Bonan, G.; Carpenter, S.R.; Chapin, F.S.; Coe, M.T.; Daily, G.C.; Gibbs, H.K.; et al. Global consequences of land use. Science 2005, 309, 570–574. [Google Scholar] [CrossRef] [Green Version]
- Malinowski, R.; Lewiński, S.; Rybicki, M.; Gromny, E.; Jenerowicz, M.; Krupiński, M.; Nowakowski, A.; Wojtkowski, C.; Krupiński, M.; Krätzschmar, E.; et al. Automated Production of a Land Cover/Use Map of Europe Based on Sentinel-2 Imagery. Remote Sens. 2020, 12, 3523. [Google Scholar] [CrossRef]
- Topaloğlu, R.H.; Sertel, E.; Musaoğlu, N. Assessment of Classification Accuracies of Sentinel-2 and Landsat-8 Data for Land Cover/Use Mapping. Int. Arch. Photogramm. Remote Sens. Spat. Inf. Sci. 2016, XLI-B8, 1055–1059. [Google Scholar] [CrossRef] [Green Version]
- Khatami, R.; Mountrakis, G.; Stehman, S.V. A meta-analysis of remote sensing research on supervised pixel-based land-cover image classification processes: General guidelines for practitioners and future research. Remote Sens. Environ. 2016, 177, 89–100. [Google Scholar] [CrossRef] [Green Version]
- Hansen, M.C.; Loveland, T.R. A review of large area monitoring of land cover change using Landsat data. Remote Sens. Environ. 2012, 122, 66–74. [Google Scholar] [CrossRef]
- Vuolo, F.; Neuwirth, M.; Immitzer, M.; Atzberger, C.; Ng, W.-T. How much does multi-temporal Sentinel-2 data improve crop type classification? Int. J. Appl. Earth Obs. Geoinf. 2018, 72, 122–130. [Google Scholar] [CrossRef]
- Xiong, J.; Thenkabail, P.; Tilton, J.; Gumma, M.; Teluguntla, P.; Oliphant, A.; Congalton, R.; Yadav, K.; Gorelick, N. Nominal 30-m Cropland Extent Map of Continental Africa by Integrating Pixel-Based and Object-Based Algorithms Using Sentinel-2 and Landsat-8 Data on Google Earth Engine. Remote Sens. 2017, 9, 1065. [Google Scholar] [CrossRef] [Green Version]
- Inglada, J.; Vincent, A.; Arias, M.; Tardy, B.; Morin, D.; Rodes, I. Operational High Resolution Land Cover Map Production at the Country Scale Using Satellite Image Time Series. Remote Sens. 2017, 9, 95. [Google Scholar] [CrossRef] [Green Version]
- Dostálová, A.; Lang, M.; Ivanovs, J.; Waser, L.T.; Wagner, W. European Wide Forest Classification Based on Sentinel-1 Data. Remote Sens. 2021, 13, 337. [Google Scholar] [CrossRef]
- Karra, K.; Kontgis, C.; Statman-Weil, Z.; Mazzariello, J.C.; Mathis, M.; Brumby, S.P. Global land use/land cover with Sentinel 2 and deep learning. In Proceedings of the 2021 IEEE International Geoscience and Remote Sensing Symposium IGARSS. IGARSS 2021, Brussels, Belgium, 11–16 July 2021; pp. 4704–4707. [Google Scholar]
- Zanaga, D.; van de Kerchove, R.; de Keersmaecker, W.; Souverijns, N.; Brockmann, C.; Quast, R.; Wevers, J.; Grosu, A.; Paccini, A.; Vergnaud, S.; et al. ESA WorldCover 10 m 2020 v100. 2021. Available online: https://doi.org/10.5281/zenodo.5571935 (accessed on 13 December 2021).
- Defourny, P.; Bontemps, S.; Bellemans, N.; Cara, C.; Dedieu, G.; Guzzonato, E.; Hagolle, O.; Inglada, J.; Nicola, L.; Rabaute, T.; et al. Near real-time agriculture monitoring at national scale at parcel resolution: Performance assessment of the Sen2-Agri automated system in various cropping systems around the world. Remote Sens. Environ. 2019, 221, 551–568. [Google Scholar] [CrossRef]
- Jiang, Y.; Lu, Z.; Li, S.; Lei, Y.; Chu, Q.; Yin, X.; Chen, F. Large-Scale and High-Resolution Crop Mapping in China Using Sentinel-2 Satellite Imagery. Agriculture 2020, 10, 433. [Google Scholar] [CrossRef]
- d’Andrimont, R.; Yordanov, M.; Martinez-Sanchez, L.; Eiselt, B.; Palmieri, A.; Dominici, P.; Gallego, J.; Reuter, H.I.; Joebges, C.; Lemoine, G.; et al. Harmonised LUCAS in-situ land cover and use database for field surveys from 2006 to 2018 in the European Union. Sci. Data 2020, 7, 352. [Google Scholar] [CrossRef] [PubMed]
- d’Andrimont, R.; Verhegghen, A.; Meroni, M.; Lemoine, G.; Strobl, P.; Eiselt, B.; Yordanov, M.; Martinez-Sanchez, L.; van der Velde, M. LUCAS Copernicus 2018: Earth-observation-relevant in situ data on land cover and use throughout the European Union. Earth Syst. Sci. Data 2021, 13, 1119–1133. [Google Scholar] [CrossRef]
- Close, O.; Benjamin, B.; Petit, S.; Fripiat, X.; Hallot, E. Use of Sentinel-2 and LUCAS Database for the Inventory of Land Use, Land Use Change, and Forestry in Wallonia, Belgium. Land 2018, 7, 154. [Google Scholar] [CrossRef] [Green Version]
- Pflugmacher, D.; Rabe, A.; Peters, M.; Hostert, P. Mapping pan-European land cover using Landsat spectral-temporal metrics and the European LUCAS survey. Remote Sens. Environ. 2019, 221, 583–595. [Google Scholar] [CrossRef]
- Weigand, M.; Staab, J.; Wurm, M.; Taubenböck, H. Spatial and semantic effects of LUCAS samples on fully automated land use/land cover classification in high-resolution Sentinel-2 data. Int. J. Appl. Earth Obs. Geoinf. 2020, 88, 102065. [Google Scholar] [CrossRef]
- Venter, Z.S.; Sydenham, M.A.K. Continental-Scale Land Cover Mapping at 10 m Resolution Over Europe (ELC10). Remote Sens. 2021, 13, 2301. [Google Scholar] [CrossRef]
- d’Andrimont, R.; Verhegghen, A.; Lemoine, G.; Kempeneers, P.; Meroni, M.; van der Velde, M. From parcel to continental scale –A first European crop type map based on Sentinel-1 and LUCAS Copernicus in-situ observations. Remote Sens. Environ. 2021, 266, 112708. [Google Scholar] [CrossRef]
- Lu, D.; Weng, Q. A survey of image classification methods and techniques for improving classification performance. Int. J. Remote Sens. 2007, 28, 823–870. [Google Scholar] [CrossRef]
- Thanh Noi, P.; Kappas, M. Comparison of Random Forest, k-Nearest Neighbor, and Support Vector Machine Classifiers for Land Cover Classification Using Sentinel-2 Imagery. Sensors 2017, 18, 18. [Google Scholar] [CrossRef] [Green Version]
- Vuolo, F.; Żółtak, M.; Pipitone, C.; Zappa, L.; Wenng, H.; Immitzer, M.; Weiss, M.; Baret, F.; Atzberger, C. Data Service Platform for Sentinel-2 Surface Reflectance and Value-Added Products: System Use and Examples. Remote Sens. 2016, 8, 938. [Google Scholar] [CrossRef] [Green Version]
- Bouhennache, R.; Bouden, T.; Taleb-Ahmed, A.; Cheddad, A. A new spectral index for the extraction of built-up land features from Landsat 8 satellite imagery. Geocarto Int. 2019, 34, 1531–1551. [Google Scholar] [CrossRef]
- Rikimaru, A.; Roy, P.S.; Miyatake, S. Tropical forest cover density mapping. Trop. Ecol. 2002, 43, 39–47. [Google Scholar]
- Xu, H. Modification of normalised difference water index (NDWI) to enhance open water features in remotely sensed imagery. Int. J. Remote Sens. 2006, 27, 3025–3033. [Google Scholar] [CrossRef]
- Zha, Y.; Gao, J.; Ni, S. Use of normalized difference built-up index in automatically mapping urban areas from TM imagery. Int. J. Remote Sens. 2003, 24, 583–594. [Google Scholar] [CrossRef]
- van Deventer, A.P.; Ward, A.D.; Gowda, P.H.; Lyon, J.G. Using thematic mapper data to identify contrasting soil plains and tillage practices. Photogramm. Eng. Remote Sens. 1997, 63, 87–93. [Google Scholar]
- Kriegler, F.J.; Malila, W.A.; Nalepka, R.F.; Richardson, W. Preprocessing transformations and their effects on multispectral recognition. Remote Sens. Environ. 1969, VI, 97–132. [Google Scholar]
- Fick, S.E.; Hijmans, R.J. WorldClim 2: New 1-km spatial resolution climate surfaces for global land areas. Int. J. Clim. 2017, 37, 4302–4315. [Google Scholar] [CrossRef]
- Weiss, M.; Baret, F. S2ToolBox Level 2 products: LAI, FAPAR, FCOVER, Version 1.1. In ESA Contract nr 4000110612/14/I-BG (p. 52); INRA: Avignon, France, 2016. [Google Scholar]
- d’Andrimont, R.; Verhegghen, A.; Meroni, M.; Lemoine, G.; Strobl, P.; Eiselt, B.; Yordanov, M.; Martinez-Sanchez, L.; van der Velde, M. LUCAS Copernicus 2018: Earth Observation relevant insitu data on land cover throughout the European Union. Earth Syst. Sci. Data Discuss. 2020, 2020, 1–19. [Google Scholar] [CrossRef]
- Breiman, L. Random Forests. Mach. Learn. 2001, 45, 5–32. [Google Scholar] [CrossRef] [Green Version]
- Immitzer, M.; Atzberger, C.; Koukal, T. Tree Species Classification with Random Forest Using Very High Spatial Resolution 8-Band WorldView-2 Satellite Data. Remote Sens. 2012, 4, 2661–2693. [Google Scholar] [CrossRef] [Green Version]
- Cortes, C.; Vapnik, V. Support-vector networks. Mach. Learn. 1995, 20, 273–297. [Google Scholar] [CrossRef]
- Vapnik, V. The Nature of Statistical Learning Theory; Springer Science & Business Media: Berlin/Heidelberg, Germany, 1999; ISBN 9780387987804. [Google Scholar]
- Sheykhmousa, M.; Mahdianpari, M.; Ghanbari, H.; Mohammadimanesh, F.; Ghamisi, P.; Homayouni, S. Support Vector Machine vs. Random Forest for Remote Sensing Image Classification: A Meta-Analysis and Systematic Review. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2020, 13, 6308–6325. [Google Scholar]
- Schultz, B.; Immitzer, M.; Formaggio, A.; Sanches, I.; Luiz, A.; Atzberger, C. Self-Guided Segmentation and Classification of Multi-Temporal Landsat 8 Images for Crop Type Mapping in Southeastern Brazil. Remote Sens. 2015, 7, 14482–14508. [Google Scholar] [CrossRef] [Green Version]
- Elgeldawi, E.; Sayed, A.; Galal, A.R.; Zaki, A.M. Hyperparameter Tuning for Machine Learning Algorithms Used for Arabic Sentiment Analysis. Informatics 2021, 8, 79. [Google Scholar] [CrossRef]
- Bergstra, J.; Bengio, Y. Random Search for Hyper-Parameter Optimization. J. Mach. Learn. Res. 2012, 13, 281–305. [Google Scholar]
- Cohen, J. A Coefficient of Agreement for Nominal Scales. Educ. Psychol. Meas. 1960, 20, 37–46. [Google Scholar] [CrossRef]
- Landis, J.R.; Koch, G.G. A One-Way Components of Variance Model for Categorical Data. Biometrics 1977, 33, 671. [Google Scholar] [CrossRef]
- Immitzer, M.; Neuwirth, M.; Böck, S.; Brenner, H.; Vuolo, F.; Atzberger, C. Optimal Input Features for Tree Species Classification in Central Europe Based on Multi-Temporal Sentinel-2 Data. Remote Sens. 2019, 11, 2599. [Google Scholar] [CrossRef] [Green Version]
- Vuolo, F.; Ng, W.-T.; Atzberger, C. Smoothing and gap-filling of high resolution multispectral time series: Example of Landsat data. Int. J. Appl. Earth Obs. Geoinf. 2017, 57, 202–213. [Google Scholar] [CrossRef]
- Inglada, J.; Vincent, A.; Arias, M.; Marais-Sicre, C. Improved Early Crop Type Identification By Joint Use of High Temporal Resolution SAR And Optical Image Time Series. Remote Sens. 2016, 8, 362. [Google Scholar] [CrossRef] [Green Version]
- Belgiu, M.; Guţ, L.; Strobl, J. Quantitative evaluation of variations in rule-based classifications of land cover in urban neighbourhoods using WorldView-2 imagery. ISPRS J. Photogramm. Remote Sens. 2014, 87, 205–215. [Google Scholar] [CrossRef] [Green Version]
- Millard, K.; Richardson, M. On the Importance of Training Data Sample Selection in Random Forest Image Classification: A Case Study in Peatland Ecosystem Mapping. Remote Sens. 2015, 7, 8489–8515. [Google Scholar] [CrossRef] [Green Version]
- Ørka, H.O.; Dalponte, M.; Gobakken, T.; Næsset, E.; Ene, L.T. Characterizing forest species composition using multiple remote sensing data sources and inventory approaches. Scand. J. For. Res. 2013, 28, 677–688. [Google Scholar] [CrossRef]
- Toscani, P.; Immitzer, M.; Atzberger, C. Wavelet-based texture measures for object-based classification of aerial images. PFG Photogramm. Fernerkund. Geoinf. 2013, 2013, 105–121. [Google Scholar] [CrossRef]
- Colditz, R. An Evaluation of Different Training Sample Allocation Schemes for Discrete and Continuous Land Cover Classification Using Decision Tree-Based Algorithms. Remote Sens. 2015, 7, 9655–9681. [Google Scholar] [CrossRef] [Green Version]
- Mellor, A.; Boukir, S.; Haywood, A.; Jones, S. Exploring issues of training data imbalance and mislabelling on random forest performance for large area land cover classification using the ensemble margin. ISPRS J. Photogramm. Remote Sens. 2015, 105, 155–168. [Google Scholar] [CrossRef]
- Immitzer, M.; Böck, S.; Einzmann, K.; Vuolo, F.; Pinnel, N.; Wallner, A.; Atzberger, C. Fractional cover mapping of spruce and pine at 1 ha resolution combining very high and medium spatial resolution satellite imagery. Remote Sens. Environ. 2018, 204, 690–703. [Google Scholar] [CrossRef] [Green Version]
- Ghosh, A.; Joshi, P.K. A comparison of selected classification algorithms for mapping bamboo patches in lower Gangetic plains using very high resolution WorldView 2 imagery. Int. J. Appl. Earth Obs. Geoinf. 2014, 26, 298–311. [Google Scholar] [CrossRef]


| Feature Name | Description | Counts | |
|---|---|---|---|
| Bio-geophysical Indicators | LAI, FAPAR, FCOVER | 12 monthly composites | |
| Spectral Bands | B2: Blue | B7: Red Edge 3 | |
| B3: Green | B8: NIR | ||
| B4: Red | B8A: NIR narrow | ||
| B5: Red Edge 1 | B11: SWIR 1 | ||
| B6: Red Edge 2 | B12: SWIR 2 | ||
| Climatic variables | RAIN: Mean rainfall value during the year 2018 | 1 yearly value | |
| TEMP: Mean temperature value during the year 2018 | |||
| Phenological variables | Day of maximum NDVI | ||
| Spectral Indices | BLFEI: | 12 monthly composites and 3 yearly centiles | |
| BSI: | |||
| MNDWI: | |||
| NDBI: | |||
| NDTI: | |||
| NDVI: | |||
| LUCAS 2018 Points | LUCAS Copernicus Points Training Data | LUCAS Core Points Validation Data | |
|---|---|---|---|
| In-situ | 238,014 | 63,364 | 174,650 |
| Office photo-interpreted | 99,803 | - | 99,803 |
| Others | 37 | - | 37 |
| Total | 337,854 | 63,364 | 274,490 |
| LUCAS Copernicus polygons | |||
| Remaining after exclusion of points due to geolocation problems in the construction of polygons | 63,287 | ||
| Remaining after exclusion of points due to missing level-3 information | 58,428 | ||
| Remaining after exclusion of points due to missing S2 data | 56,366 (1,901,627 pixels) | ||
| Remaining after exclusion of points due to missing S2 features in winter months | 43,013 (1,349,052 pixels) | ||
| Remaining after exclusion of points not covered by one of the 21 LC classes assessed in this study | 42,753 (1,344,885 pixels) | ||
| Month (2018) | Dec. | Jan. | Feb. | Mar. | Nov. | Apr. | Sep. | Aug. | May. | Jun. | Jul. | Oct. |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Missing values | 1,191,638 | 1,156,586 | 994,954 | 883,140 | 748,819 | 167,270 | 99,414 | 86,544 | 86,029 | 83,494 | 79,334 | 73,261 |
| % of missing values | 63% | 61% | 52% | 46% | 39% | 9% | 5% | 5% | 5% | 4% | 4% | 4% |
| Grouped Class Name | Code | Main Class Name | Class Descriptors in LUCAS Level-3 Land Cover |
|---|---|---|---|
| Cereals | 211 | Common wheat | B11-Common wheat |
| 212 | Durum wheat | B12-Durum wheat | |
| 213 | Barley | B13-Barley | |
| 214 | Rye | B14-Rye | |
| 215 | Oats | B15-Oats | |
| 216 | Maize | B16-Maize | |
| 217 | Rice | B17-Rice | |
| 218 | Triticale | B18-Triticale | |
| 219 | Other cereals | B19-Other cereals | |
| Root crops | 221 | Potatoes | B21-Potatoes |
| 222 | Sugar beet | B22-Sugar beet | |
| 223 | Other root crops | B23-Other root crops | |
| Non-permanent industrial crops | 230 | Other non-permanent industrial crops | B34-Cotton |
| B35-Other fibre and oleaginous crops | |||
| B36-Tobacco | |||
| B37-Other non-permanent industrial crops | |||
| 231 | Sunflower | B31-Sunflower | |
| 232 | Rape and turnip rape | B32-Rape and turnip rape | |
| 233 | Soya | B33-Soya | |
| Dry pulses, vegetables, and flowers | 240 | Dry pulses, vegetables, and flowers | B41-Dry pulses |
| B42-Tomatoes | |||
| B43-Other fresh vegetables | |||
| B44-Floriculture and ornamental plants | |||
| B45-Strawberries | |||
| Fodder crops | 250 | Fodder crops | B51-Clovers |
| B52-Lucerne | |||
| B53-Other leguminous and mixtures for fodder | |||
| B54-Mixed cereals for fodder | |||
| Bare arable land | 290 | Bare arable land | F40-Other bare soil (only with U111/112/113 Land use) |
| Woodland and shrubland | 300 | Woodland and shrubland | B71-Apple fruit |
| B72-Pear fruit | |||
| B73-Cherry fruit | |||
| B74-Nuts trees | |||
| B75-Other fruit trees and berries | |||
| B76-Oranges | |||
| B77-Other citrus fruit | |||
| B81-Olive groves | |||
| B82-Vineyards | |||
| B83-Nurseries | |||
| B84-Permanent industrial crops | |||
| C10-Broadleaved woodland | |||
| C21-Spruce dominated coniferous woodland | |||
| C22-Pine dominated coniferous woodland | |||
| C23-Other coniferous woodland | |||
| C31-Spruce dominated mixed woodland | |||
| C32-Pine dominated mixed woodland | |||
| C33-Other mixed woodland | |||
| D10-Shrubland with sparse tree cover | |||
| D20-Shrubland without tree cover | |||
| Grassland | 500 | Grassland | B55-Temporary grasslands |
| E10-Grassland with sparse tree/shrub cover | |||
| E20-Grassland without tree/shrub cover | |||
| E30-Spontaneously vegetated surfaces |
| Class Name | All Data | Proportional | Balanced | Dissimilar |
|---|---|---|---|---|
| Woodland and shrubland | 562,564 | 16,708 | 4000 | 62,812 |
| Grassland | 338,977 | 12,300 | 4000 | 39,149 |
| Common wheat | 132,878 | 3749 | 4000 | 14,757 |
| Maize | 63,788 | 1893 | 4000 | 7102 |
| Barley | 54,881 | 1830 | 4000 | 6250 |
| Fodder crops | 28,882 | 997 | 4000 | 3316 |
| Rape and turnip rape | 30,388 | 887 | 4000 | 3393 |
| Bare arable land | 19,678 | 777 | 4000 | 2336 |
| Sunflower | 17,001 | 520 | 4000 | 1913 |
| Dry pulses, vegetables, and flowers | 14,241 | 487 | 4000 | 1619 |
| Rye | 16,934 | 483 | 4000 | 1885 |
| Oats | 13,778 | 474 | 4000 | 1579 |
| Durum wheat | 11,236 | 381 | 4000 | 1284 |
| Sugar beet | 10,752 | 327 | 4000 | 1209 |
| Triticale | 9573 | 284 | 4000 | 1070 |
| Potatoes | 6827 | 227 | 4000 | 777 |
| Other non-permanent industrial crops | 5363 | 178 | 4000 | 612 |
| Soya | 3234 | 116 | 3234 | 371 |
| Other cereals | 2187 | 71 | 2187 | 247 |
| Other roots crops | 1647 | 61 | 1647 | 189 |
| Rice | 76 | 3 | 76 | 9 |
| Sum | 1,344,885 | 42,753 | 75,144 | 151,879 |
| Parameters | Value Range in Random Search | Value Range in Grid Search | Tuned Values |
|---|---|---|---|
| Number of possible permutations | 1000 | 56 | |
| Number of assessed permutations | 100 | 56 | |
| n_estimators | [200:200:2000] | [600:100:1900] | 1100 |
| max_features | [‘sqrt’, ‘log2’] | [‘sqrt’] | [‘sqrt’] |
| min_samples_split | [1:1:10] | [2:1:5] | 3 |
| min_samples_leaf | [1:1:5] | 1 | 1 |
| Parameters | Value Range in Random Search | Value Range in Grid Search | Tuned Values |
|---|---|---|---|
| Number of possible permutations | 1800 | 30 | |
| Number of assessed permutations | 180 | 30 | |
| C | [1:1:100] | [2:1:6] | 3 |
| kernel | [‘poly’, ‘rbf’, ‘sigmoid’] | [‘rbf’] | [‘rbf’] |
| degree | [2:1:4] | [2:1:4] | 2 |
| gamma | [‘scale’, ‘auto’] | [‘scale’, ‘auto’] | [‘auto’] |
| KC | <0.00 | 0.00–0.20 | 0.21–0.4 | 0.41–0.60 | 0.61–0.80 | 0.81–1.00 |
|---|---|---|---|---|---|---|
| Strength of agreement | Poor | Slight | Fair | Moderate | Substantial | Almost perfect |
| Sampling Method | Number of Samples | OOB for RF | OA for RF | OA for SVM |
|---|---|---|---|---|
| Proportional | 42,753 | 77.5% | 75.7% | 76.3% |
| Balanced | 75,144 | 91.4% | 72.6% | 71.2% |
| Dissimilar | 151,879 | 86.8% | 76.2% | 77.6% |
| All data | 1,344,885 | 98.2% | 76.8% | 77.8% |
| Comprehensive Class | Code | 210 | 220 | 230 | 240 | 250 | 290 | 300 | 500 | Total | UA | F1-score |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Cereals | 210 | 12,140 | 116 | 351 | 162 | 272 | 786 | 202 | 983 | 15,012 | 80.9% | 83.6% |
| Root crops | 220 | 37 | 769 | 35 | 24 | 6 | 9 | 2 | 16 | 898 | 85.6% | 77.9% |
| Non permanent industrial crops | 230 | 90 | 35 | 1736 | 70 | 13 | 127 | 12 | 118 | 2201 | 78.9% | 75.4% |
| Dry pulses, vegetables and flowers | 240 | 74 | 67 | 63 | 203 | 31 | 56 | 4 | 53 | 551 | 36.8% | 33.8% |
| Fodder crops | 250 | 42 | 7 | 9 | 19 | 405 | 11 | 9 | 71 | 573 | 70.7% | 34.3% |
| Bare arable land | 290 | 113 | 9 | 16 | 19 | 6 | 347 | 14 | 118 | 642 | 54.0% | 26.8% |
| Woodland & shrubland | 300 | 274 | 12 | 34 | 40 | 95 | 179 | 25,321 | 2715 | 28,670 | 88.3% | 90.4% |
| Grassland | 500 | 1277 | 61 | 160 | 112 | 963 | 428 | 1775 | 17,477 | 22,253 | 78.5% | 79.8% |
| Total | 14,047 | 1076 | 2404 | 649 | 1791 | 1943 | 27,339 | 21,551 | 70,800 | OA = 82.5% | ||
| PA | 86.4% | 71.5% | 72.2% | 31.3% | 22.6% | 17.9% | 92.6% | 81.1% | ||||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ghassemi, B.; Dujakovic, A.; Żółtak, M.; Immitzer, M.; Atzberger, C.; Vuolo, F. Designing a European-Wide Crop Type Mapping Approach Based on Machine Learning Algorithms Using LUCAS Field Survey and Sentinel-2 Data. Remote Sens. 2022, 14, 541. https://doi.org/10.3390/rs14030541
Ghassemi B, Dujakovic A, Żółtak M, Immitzer M, Atzberger C, Vuolo F. Designing a European-Wide Crop Type Mapping Approach Based on Machine Learning Algorithms Using LUCAS Field Survey and Sentinel-2 Data. Remote Sensing. 2022; 14(3):541. https://doi.org/10.3390/rs14030541
Chicago/Turabian StyleGhassemi, Babak, Aleksandar Dujakovic, Mateusz Żółtak, Markus Immitzer, Clement Atzberger, and Francesco Vuolo. 2022. "Designing a European-Wide Crop Type Mapping Approach Based on Machine Learning Algorithms Using LUCAS Field Survey and Sentinel-2 Data" Remote Sensing 14, no. 3: 541. https://doi.org/10.3390/rs14030541
APA StyleGhassemi, B., Dujakovic, A., Żółtak, M., Immitzer, M., Atzberger, C., & Vuolo, F. (2022). Designing a European-Wide Crop Type Mapping Approach Based on Machine Learning Algorithms Using LUCAS Field Survey and Sentinel-2 Data. Remote Sensing, 14(3), 541. https://doi.org/10.3390/rs14030541








