Feasibility of Combining Deep Learning and RGB Images Obtained by Unmanned Aerial Vehicle for Leaf Area Index Estimation in Rice
Abstract
1. Introduction
2. Materials and Methods
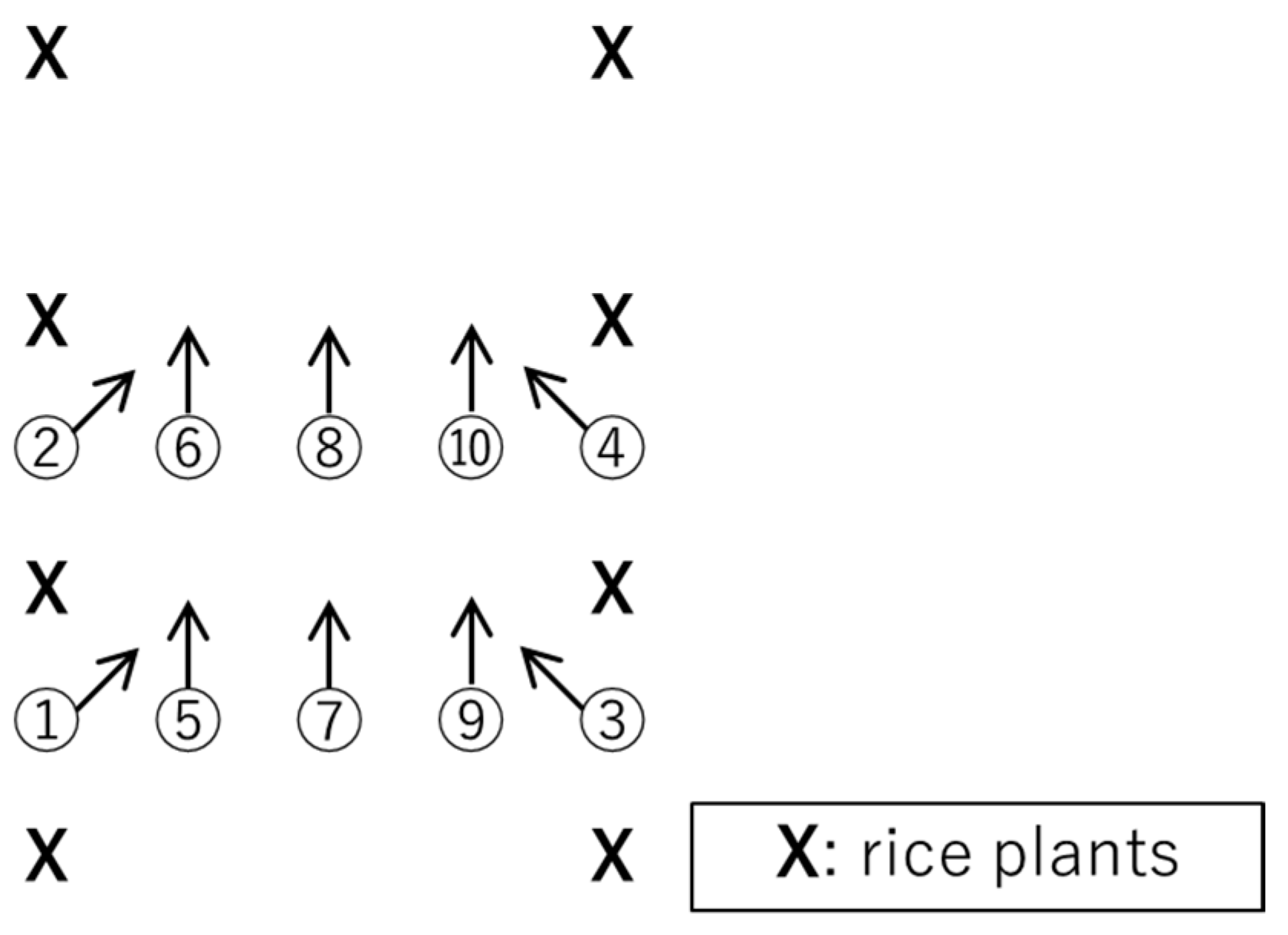
2.1. Experimental Design and Data Acquisition
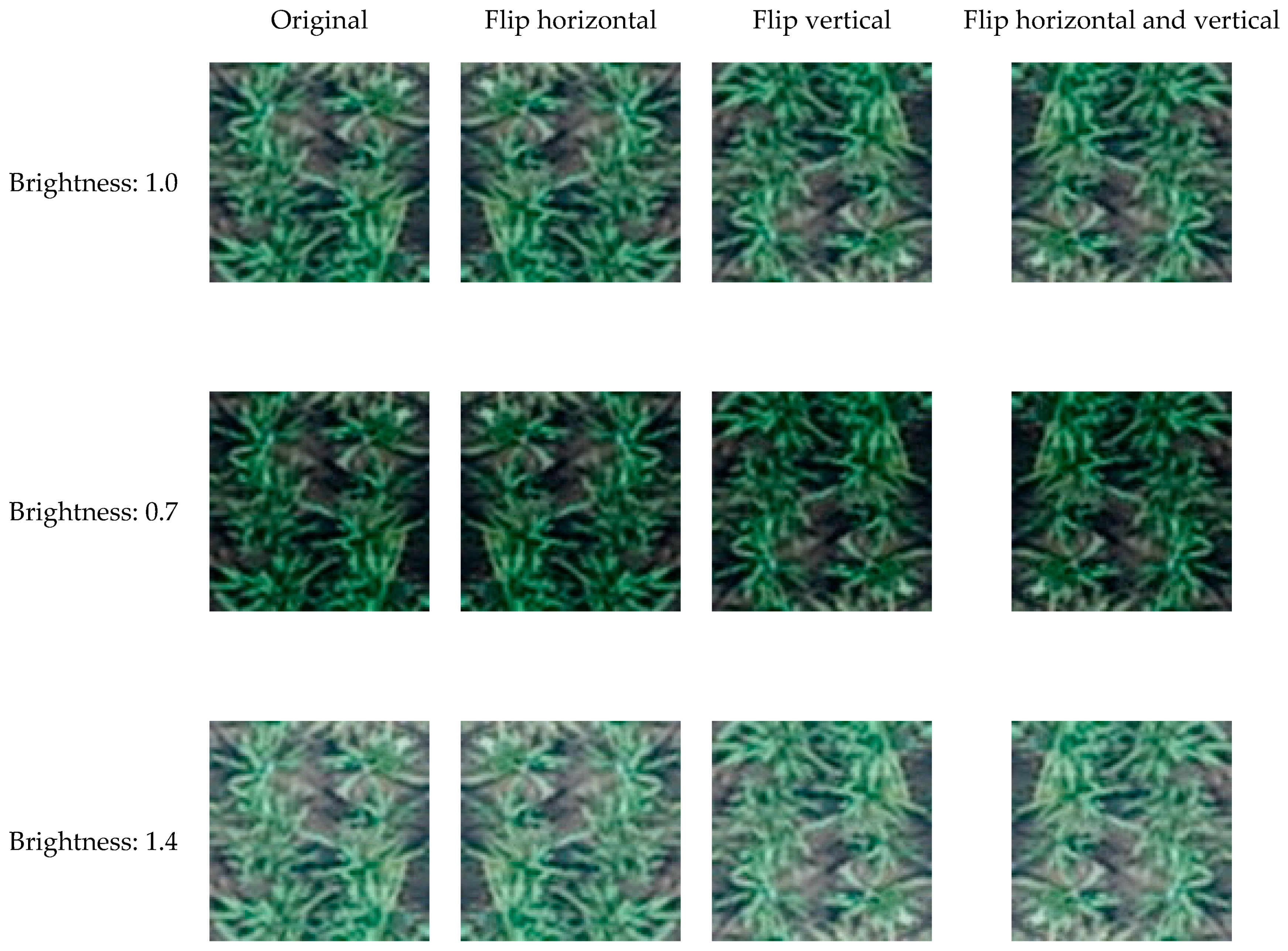
2.2. Image Processing
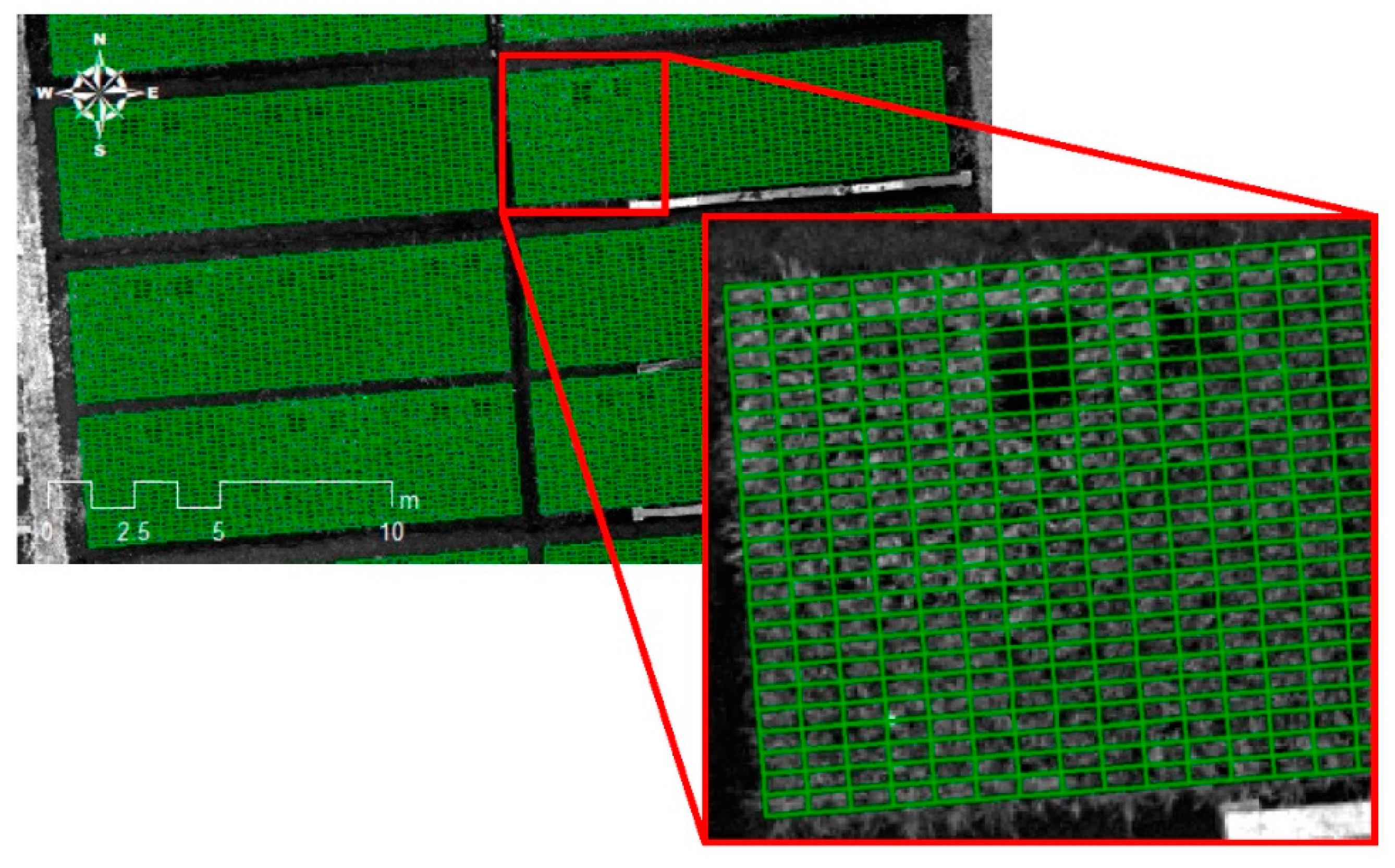
2.2.1. Generation of Ortho-Mosaic Images
2.2.2. Calculation of Vegetation Indices and Color Indices
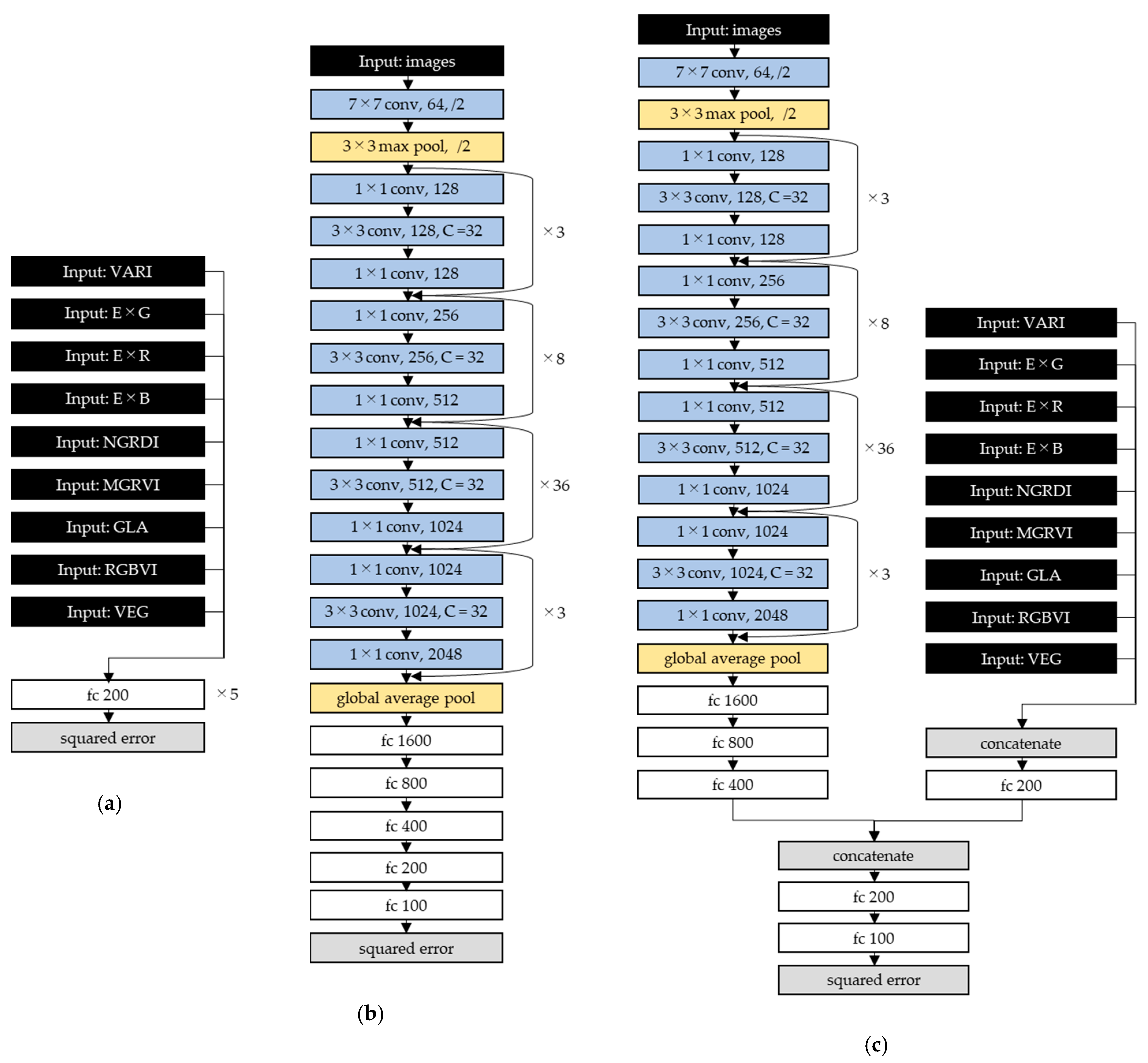
2.3. Estimation Model Development and Accuracy Assessment
3. Results
3.1. Variations of the Ground-Measured Leaf Area Index
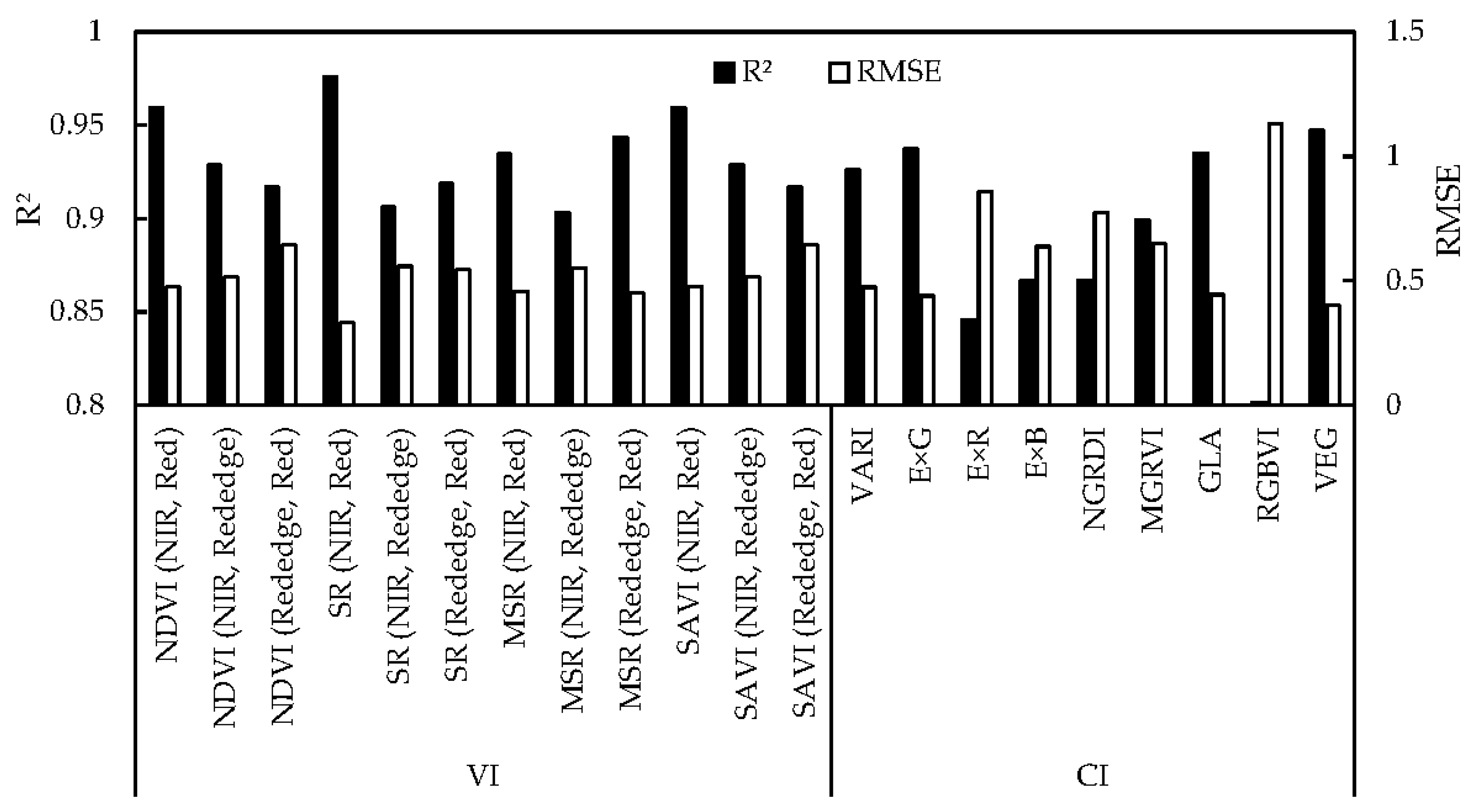
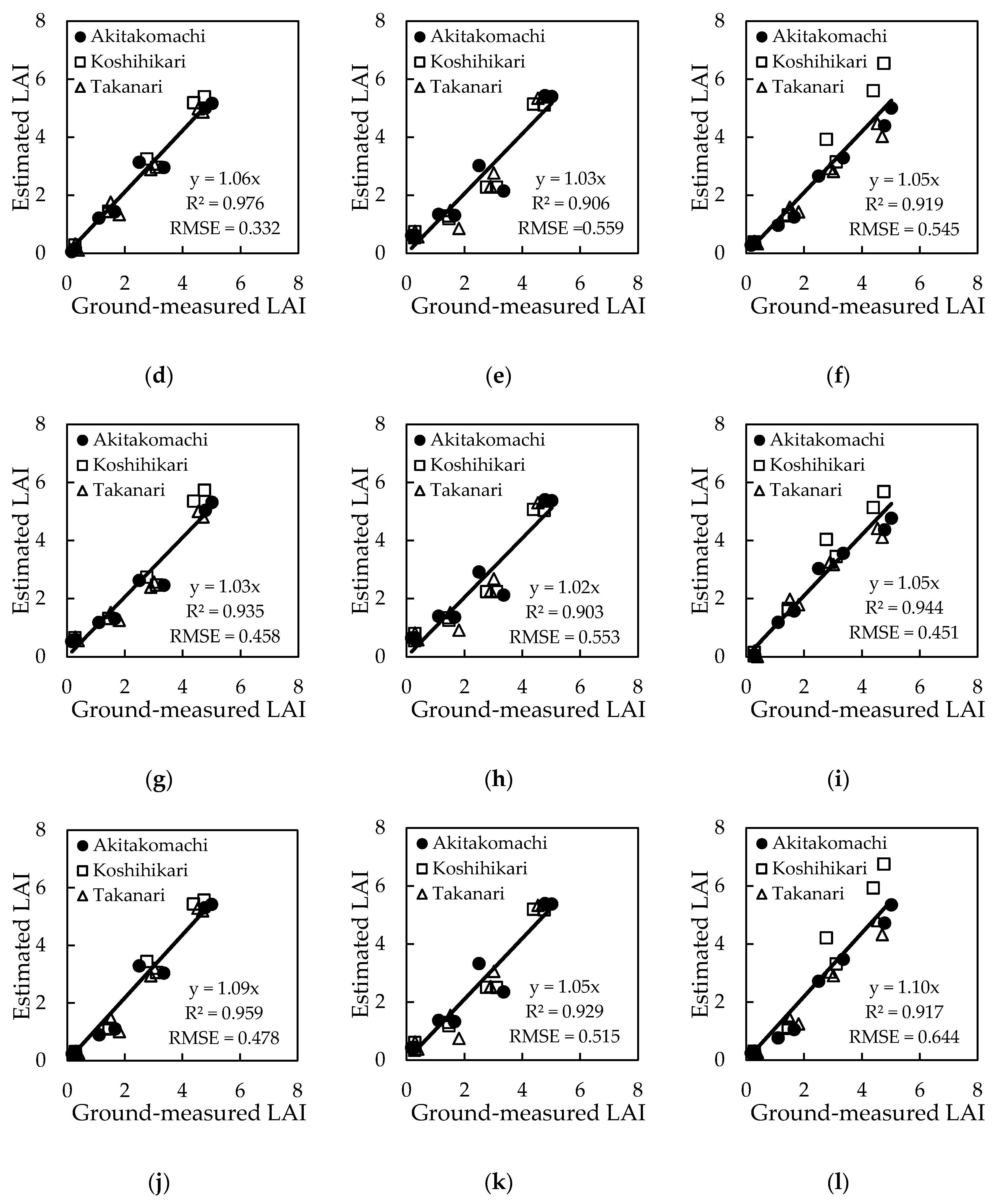
3.2. Regression Models Using Each of VIs and CIs
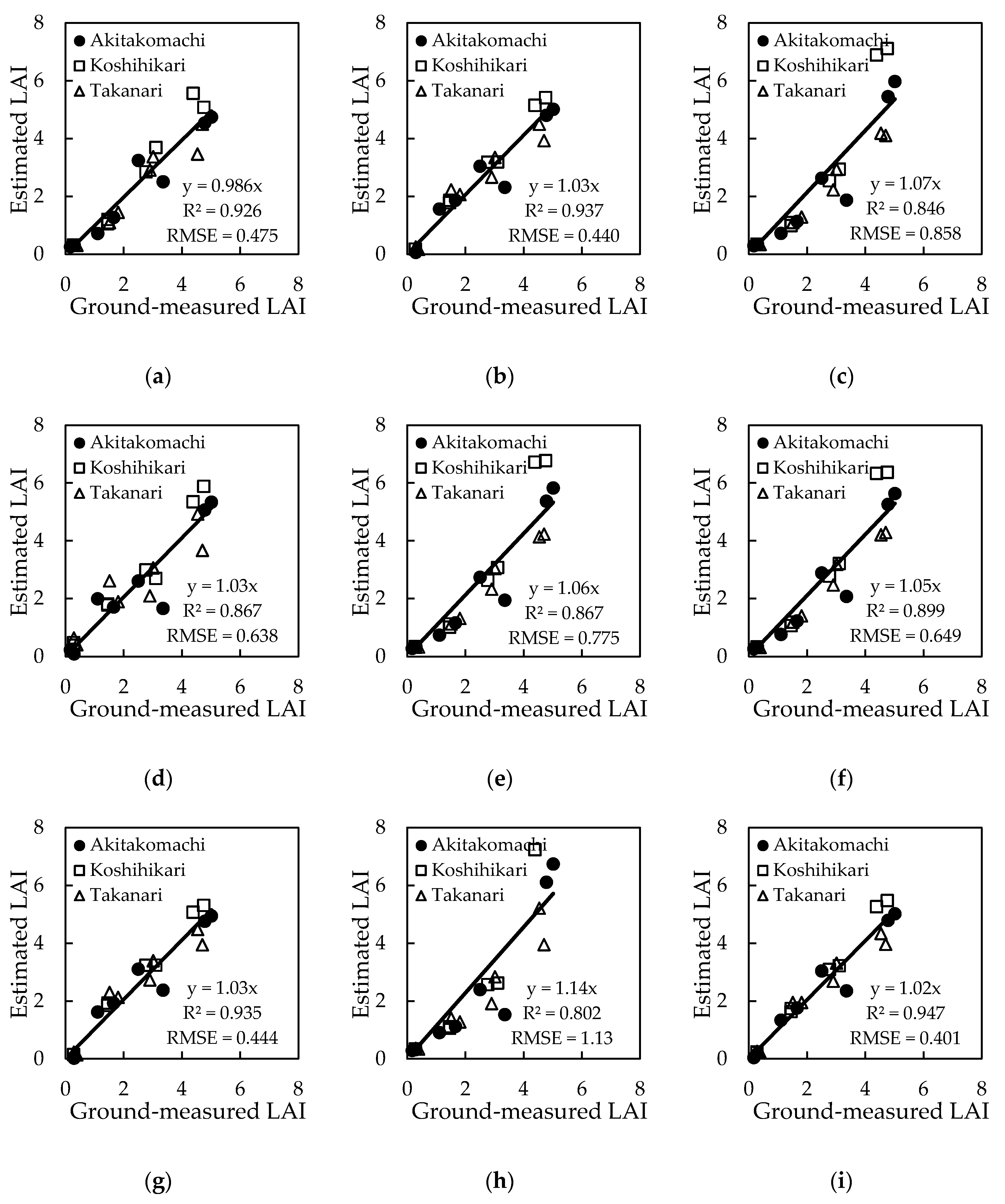
3.3. Estimation Models by Machine-Learning Algorithms Other Than Deep Learning
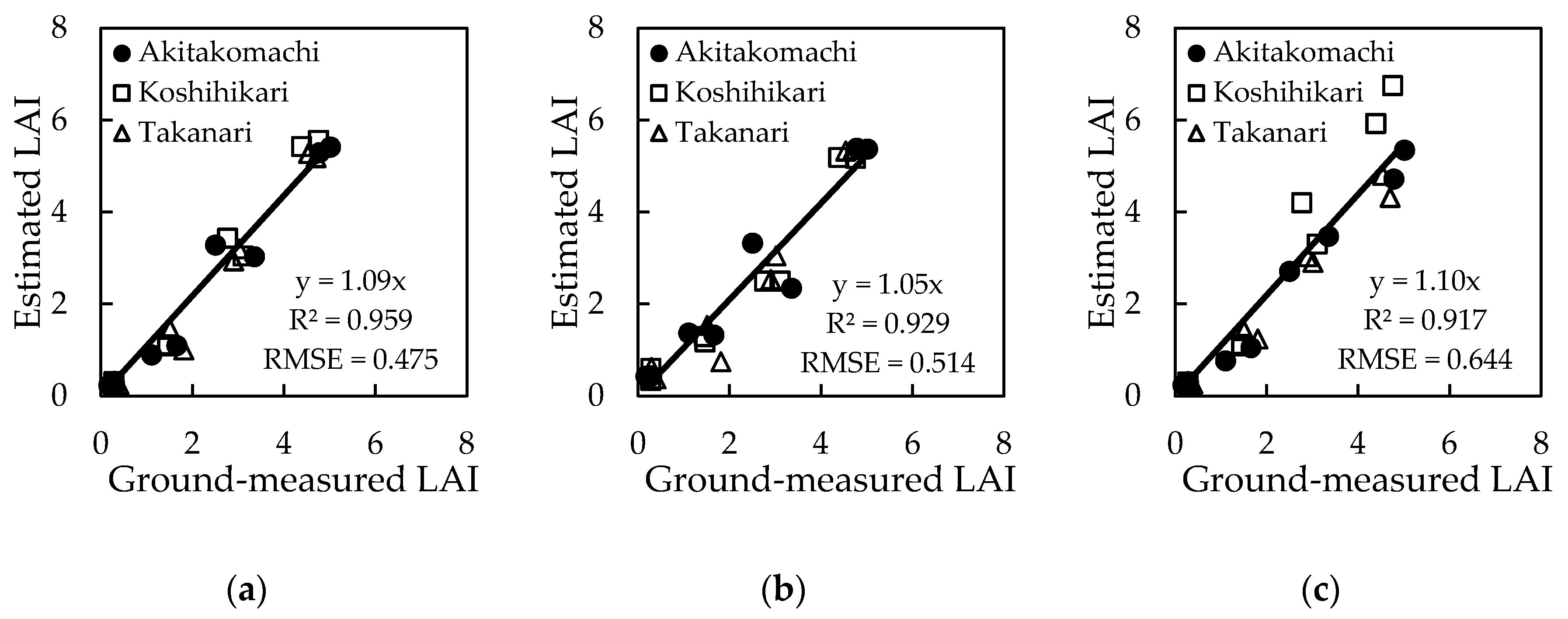
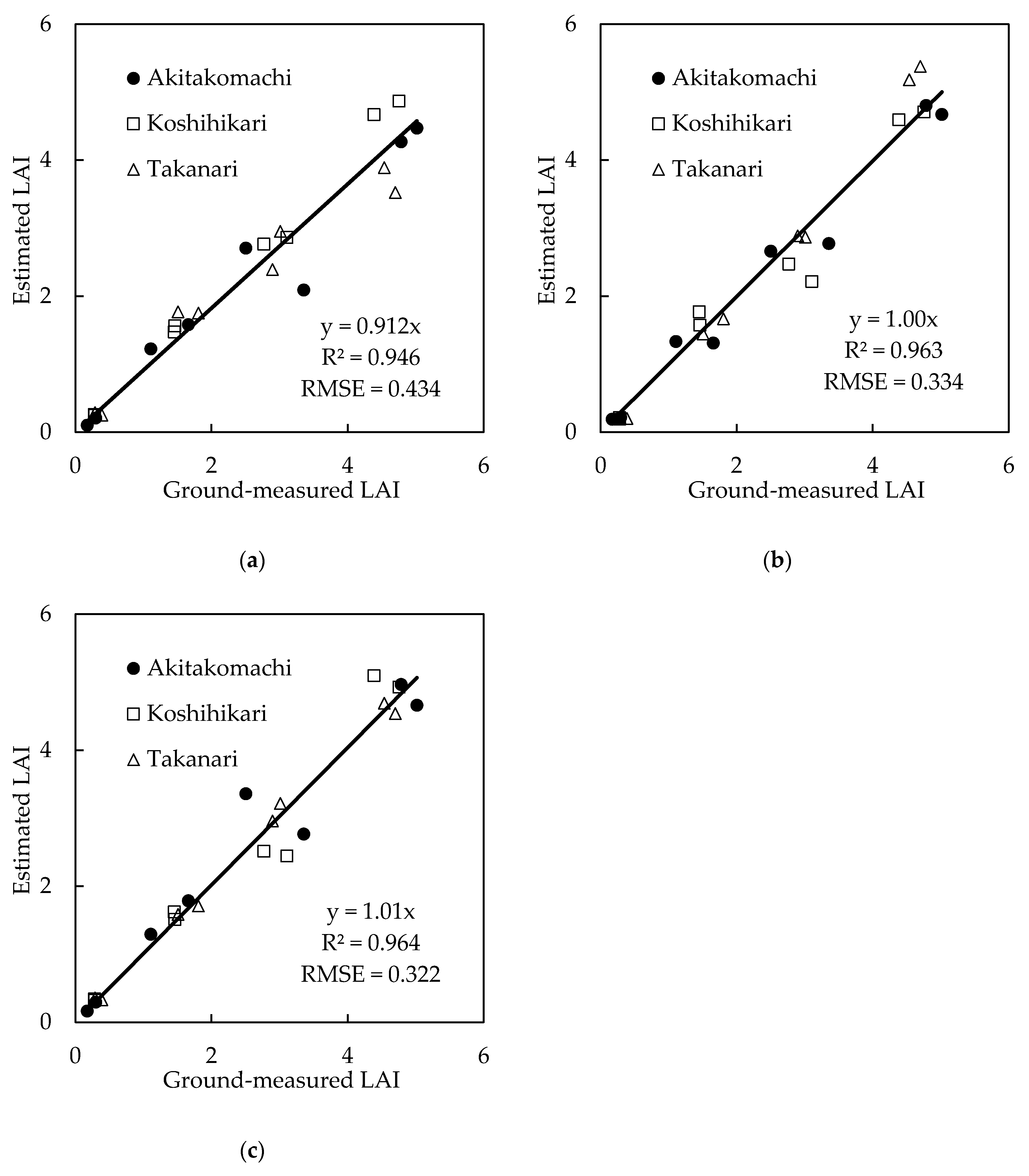
3.4. Estimation Models by Deep Learning
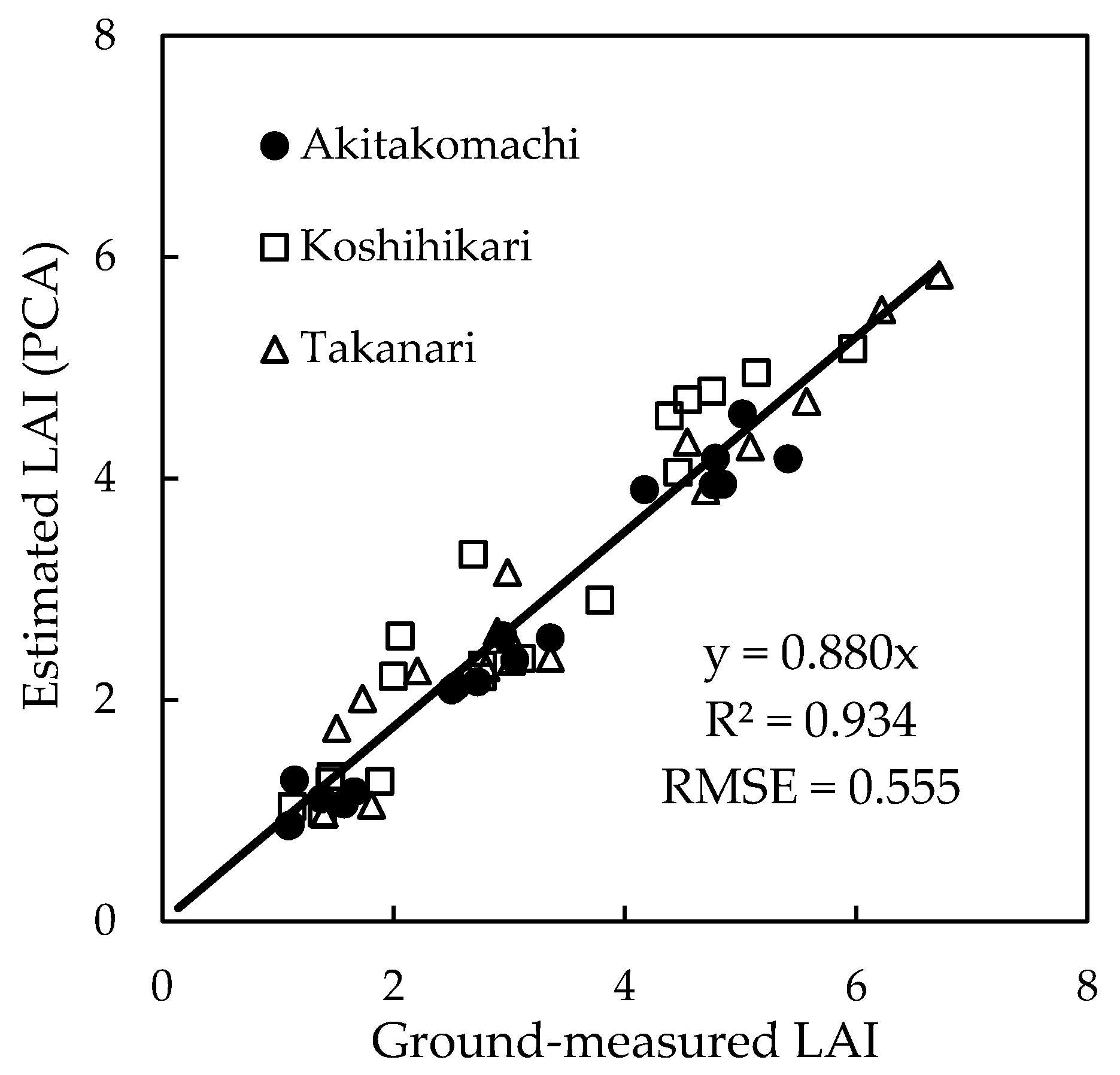
3.5. Plant Canopy Analyzer
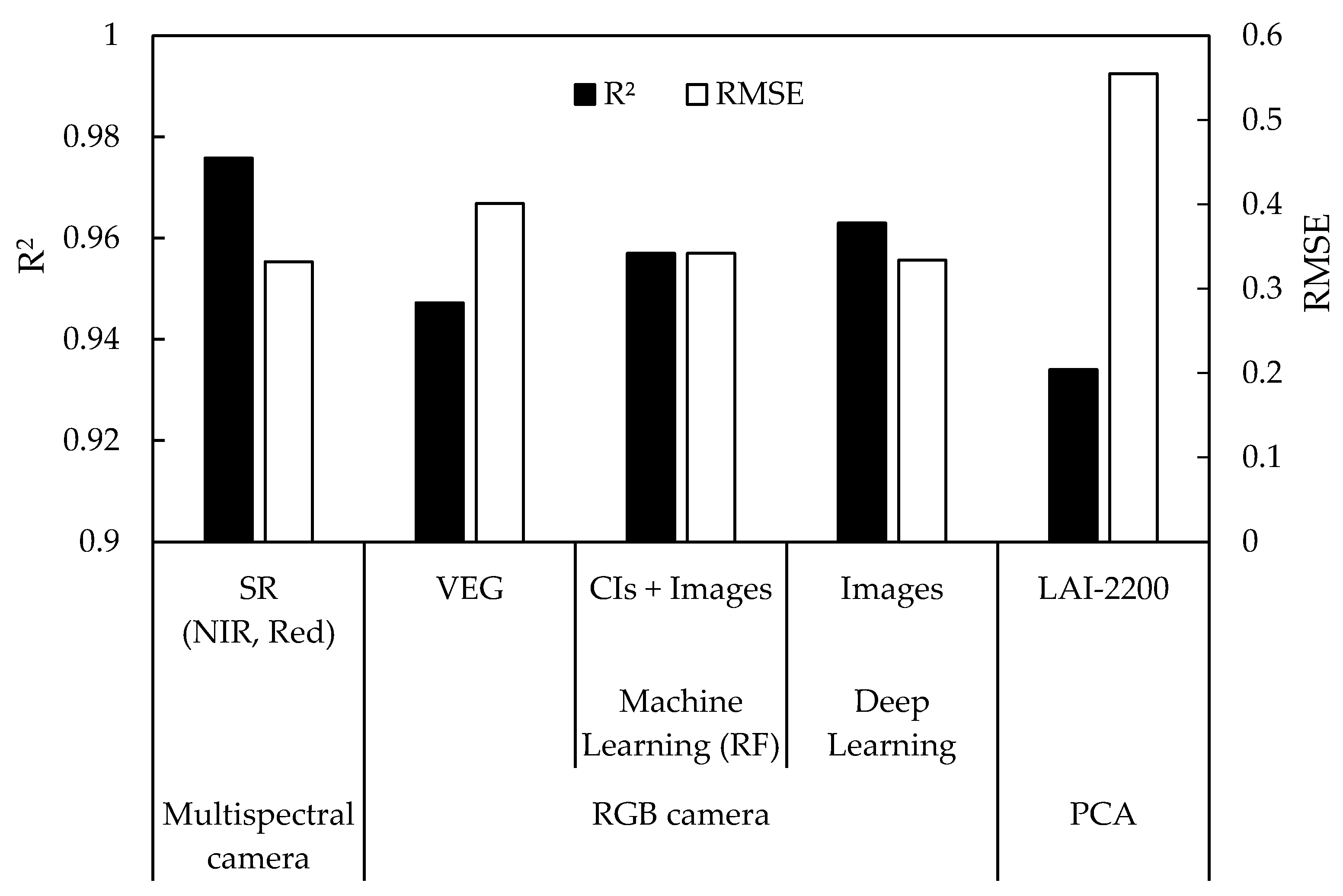
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Lahoz, W. Systematic Observation Requirements for Satellite-based Products for Climate, 2011 Update, Supplemental Details to the Satellite-Based Component of the Implementation Plan for the Global Observing System for Climate in Support of the UNFCCC (2010 Update); World Meteorological Organization (WMO): Geneva, Switzerland, 2011; p. 83. [Google Scholar]
- Asner, G.P.; Scurlock, J.M.O.; Hicke, J.A. Global synthesis of leaf area index observations. Glob. Chang. Biol. 2008, 14, 237–243. [Google Scholar] [CrossRef]
- Stark, S.C.; Leitold, V.; Wu, J.L.; Hunter, M.O.; de Castilho, C.V.; Costa, F.R.C.; Mcmahon, S.M.; Parker, G.G.; Shimabukuro, M.T.; Lefsky, M.A.; et al. Amazon forest carbon dynamics predicted by profiles of canopy leaf area and light environment. Ecol. Lett. 2012, 15, 1406–1414. [Google Scholar] [CrossRef] [PubMed]
- Duchemin, B.; Maisongrande, P.; Boulet, G.; Benhadj, I. A simple algorithm for yield estimates: Evaluation for semi-arid irrigated winter wheat monitored with green leaf area index. Environ. Model. Softw. 2008, 23, 876–892. [Google Scholar] [CrossRef]
- Fang, H.; Liang, S.; Hoogenboom, G. Integration of MODIS LAI and vegetation index products with the CSM-CERES-Maize model for corn yield estimation. Int. J. Remote Sens. 2011, 32, 1039–1065. [Google Scholar] [CrossRef]
- Brisson, N.; Gary, C.; Justes, E.; Roche, R.; Mary, B.; Ripoche, D.; Zimmer, D.; Sierra, J.; Bertuzzi, P.; Burger, P.; et al. An overview of the crop model STICS. Eur. J. Agron. 2003, 18, 309–332. [Google Scholar] [CrossRef]
- Stenberg, P.; Linder, S.; Smolander, H.; Flower-Ellis, J. Performance of the LAI-2000 plant canopy analyzer in estimating leaf area index of some Scots pine stands. Tree Physiol. 1994, 14, 981–995. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Nie, S.; Xi, X.; Luo, S.; Sun, X. Estimating the biomass of maize with hyperspectral and LiDAR data. Remote Sens. 2016, 9, 11. [Google Scholar] [CrossRef]
- Luo, S.; Wang, C.; Xi, X.; Nie, S.; Fan, X.; Chen, H.; Yang, X.; Peng, D.; Lin, Y.; Zhou, G. Combining hyperspectral imagery and LiDAR pseudo-waveform for predicting crop LAI, canopy height and above-ground biomass. Ecol. Indic. 2019, 102, 801–812. [Google Scholar] [CrossRef]
- Hashimoto, N.; Saito, Y.; Maki, M.; Homma, K. Simulation of reflectance and vegetation indices for unmanned aerial vehicle (UAV) monitoring of paddy fields. Remote Sens. 2019, 11, 2119. [Google Scholar] [CrossRef]
- Sakamoto, T.; Shibayama, M.; Kimura, A.; Takada, E. Assessment of digital camera-derived vegetation indices in quantitative monitoring of seasonal rice growth. ISPRS J. Photogramm. Remote Sens. 2011, 66, 872–882. [Google Scholar] [CrossRef]
- Fan, X.; Kawamura, K.; Guo, W.; Xuan, T.D.; Lim, J.; Yuba, N.; Kurokawa, Y.; Obitsu, T.; Lv, R.; Tsumiyama, Y.; et al. A simple visible and near-infrared (V-NIR) camera system for monitoring the leaf area index and growth stage of Italian ryegrass. Comput. Electron. Agric. 2018, 144, 314–323. [Google Scholar] [CrossRef]
- Lee, K.J.; Lee, B.W. Estimation of rice growth and nitrogen nutrition status using color digital camera image analysis. Eur. J. Agron. 2013, 48, 57–65. [Google Scholar] [CrossRef]
- Tanaka, Y.; Katsura, K.; Yamashita, Y. Verification of image processing methods using digital cameras for rice growth diagnosis. J. Japan Soc. Photogramm. Remote Sens. 2020, 59, 248–258. [Google Scholar]
- Maes, W.H.; Steppe, K. Perspectives for Remote Sensing with Unmanned Aerial Vehicles in Precision Agriculture. Trends Plant Sci. 2019, 24, 152–164. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Kovacs, J.M. The application of small unmanned aerial systems for precision agriculture: A review. Precis. Agric. 2012, 13, 693–712. [Google Scholar] [CrossRef]
- Kross, A.; McNairn, H.; Lapen, D.; Sunohara, M.; Champagne, C. Assessment of RapidEye vegetation indices for estimation of leaf area index and biomass in corn and soybean crops. Int. J. Appl. Earth Obs. Geoinf. 2015, 34, 235–248. [Google Scholar] [CrossRef]
- Liu, K.; Zhou, Q.B.; Wu, W.B.; Xia, T.; Tang, H.J. Estimating the crop leaf area index using hyperspectral remote sensing. J. Integr. Agric. 2016, 15, 475–491. [Google Scholar] [CrossRef]
- Gupta, R.K.; Vijayan, D.; Prasad, T.S. The relationship of hyper-spectral vegetation indices with leaf area index (LAI) over the growth cycle of wheat and chickpea at 3 nm spectral resolution. Adv. Sp. Res. 2006. [Google Scholar] [CrossRef]
- Lu, N.; Zhou, J.; Han, Z.; Li, D.; Cao, Q.; Yao, X.; Tian, Y.; Zhu, Y.; Cao, W.; Cheng, T. Improved estimation of aboveground biomass in wheat from RGB imagery and point cloud data acquired with a low-cost unmanned aerial vehicle system. Plant Methods 2019, 15, 1–16. [Google Scholar] [CrossRef]
- Houborg, R.; McCabe, M.F. A hybrid training approach for leaf area index estimation via Cubist and random forests machine-learning. ISPRS J. Photogramm. Remote Sens. 2018, 135, 173–188. [Google Scholar] [CrossRef]
- Li, S.; Yuan, F.; Ata-UI-Karim, S.T.; Zheng, H.; Cheng, T.; Liu, X.; Tian, Y.; Zhu, Y.; Cao, W.; Cao, Q. Combining color indices and textures of UAV-based digital imagery for rice LAI estimation. Remote Sens. 2019, 11, 63. [Google Scholar] [CrossRef]
- Verrelst, J.; Muñoz, J.; Alonso, L.; Delegido, J.; Rivera, J.P.; Camps-Valls, G.; Moreno, J. Machine learning regression algorithms for biophysical parameter retrieval: Opportunities for Sentinel-2 and -3. Remote Sens. Environ. 2012, 118, 127–139. [Google Scholar] [CrossRef]
- Wang, L.; Chang, Q.; Yang, J.; Zhang, X.; Li, F. Estimation of paddy rice leaf area index using machine learning methods based on hyperspectral data from multi-year experiments. PLoS ONE 2018, 13, e0207624. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.; Liu, K.; Liu, L.; Myint, S.W.; Wang, S.; Liu, H.; He, Z. Exploring the potential of world view-2 red-edge band-based vegetation indices for estimation of mangrove leaf area index with machine learning algorithms. Remote Sens. 2017, 9, 60. [Google Scholar] [CrossRef]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Delving Deep into Rectifiers: Surpassing Human-Level Performance on ImageNet Classification. In Proceedings of the IEEE International Conference on Computer Vision (ICCV), Santiago, Chile, 7–13 December 2015; pp. 1026–1034. [Google Scholar]
- Mohanty, S.P.; Hughes, D.P.; Salathé, M. Using Deep Learning for Image-Based Plant Disease Detection. Front. Plant Sci. 2016, 7, 1419. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Lin, Z.; Zhao, X.; Wang, G.; Gu, Y. Deep learning-based classification of hyperspectral data. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2014, 7, 2094–2107. [Google Scholar] [CrossRef]
- Reyes, A.K.; Caicedo, J.C.; Camargo, J. Fine-tuning Deep Convolutional Networks for Plant Recognition. CLEF Work. Notes 2015, 1391, 467–475. [Google Scholar]
- Xinshao, W.; Cheng, C. Weed seeds classification based on PCANet deep learning baseline. In Proceedings of the 2015 Asia-Pacific Signal and Information Processing Association Annual Summit and Conference, APSIPA ASC 2015, Hong Kong, China, 16–19 December 2015; pp. 408–415. [Google Scholar]
- Song, X.; Zhang, G.; Liu, F.; Li, D.; Zhao, Y.; Yang, J. Modeling spatio-temporal distribution of soil moisture by deep learning-based cellular automata model. J. Arid Land 2016, 8, 734–748. [Google Scholar] [CrossRef]
- Zhou, X.; Kono, Y.; Win, A.; Matsui, T.; Tanaka, T.S.T. Predicting within-field variability in grain yield and protein content of winter wheat using UAV-based multispectral imagery and machine learning approaches. Plant Prod. Sci. 2020. [Google Scholar] [CrossRef]
- Krizhevsky, A.; Sutskever, I.; Hinton, G.E. ImageNet Classification with Deep Convolutional Neural Networks. In Proceedings of the Advances in Neural Information Processing Systems 25, Neural Information Processing Systems Foundation, Inc. ( NIPS ), Lake Tahoe, NV, USA, 3–6 December 2012; pp. 1106–1114. [Google Scholar]
- Carl, F. Jordan Derivation of Leaf-Area Index from Quality of Light on the Forest Floor. Ecology 1969, 50, 663–666. [Google Scholar]
- Chen, J.M. Evaluation of vegetation indices and a modified simple ratio for boreal applications. Can. J. Remote Sens. 1996, 22, 229–242. [Google Scholar] [CrossRef]
- Huete, A. A soil-adjusted vegetation index (SAVI). Remote Sens. Environ. 1988, 25, 295–309. [Google Scholar] [CrossRef]
- Gitelson, A.A.; Kaufman, Y.J.; Stark, R.; Rundquist, D. Novel algorithms for remote estimation of vegetation fraction. Remote Sens. Environ. 2002, 80, 76–87. [Google Scholar] [CrossRef]
- Woebbecke, D.M.; Meyer, G.E.; Von Bargen, K.; Mortensen, D.A. Color Indices for Weed Identification Under Various Soil, Residue, and Lighting Conditions. Trans. ASAE 1995, 38, 259–269. [Google Scholar] [CrossRef]
- Meyer, G.E.; Neto, J.C. Verification of color vegetation indices for automated crop imaging applications. Comput. Electron. Agric. 2008, 63, 282–293. [Google Scholar] [CrossRef]
- Mao, W.; Wang, Y.; Wang, Y. Real-time Detection of Between-row Weeds Using Machine Vision. In Proceedings of the 2003 ASAE Annual Meeting, American Society of Agricultural and Biological Engineers (ASABE), Las Vegas, NV, USA, 27–30 July 2003; p. 1. [Google Scholar]
- Tucker, C.J. Red and photographic infrared linear combinations for monitoring vegetation. Remote Sens. Environ. 1979, 8, 127–150. [Google Scholar] [CrossRef]
- Louhaichi, M.; Borman, M.M.; Johnson, D.E. Spatially located platform and aerial photography for documentation of grazing impacts on wheat. Geocarto Int. 2001, 16, 65–70. [Google Scholar] [CrossRef]
- Bendig, J.; Yu, K.; Aasen, H.; Bolten, A.; Bennertz, S.; Broscheit, J.; Gnyp, M.L.; Bareth, G. Combining UAV-based plant height from crop surface models, visible, and near infrared vegetation indices for biomass monitoring in barley. Int. J. Appl. Earth Obs. Geoinf. 2015, 39, 79–87. [Google Scholar] [CrossRef]
- Hague, T.; Tillett, N.D.; Wheeler, H. Automated crop and weed monitoring in widely spaced cereals. Precis. Agric. 2006, 7, 21–32. [Google Scholar] [CrossRef]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep residual learning for image recognition. In Proceedings of the IEEE Computer Society Conference on Computer Vision and Pattern Recognition; IEEE Computer Society, Las Vegas, NV, USA, 26 June–1 July 2016; pp. 770–778. [Google Scholar]
- Xie, S.; Girshick, R.; Dollár, P.; Tu, Z.; He, K. Aggregated residual transformations for deep neural networks. In Proceedings of the 30th IEEE Conference on Computer Vision and Pattern Recognition, CVPR 2017, Honolulu, HI, USA, 21–26 July 2017; pp. 5987–5995. [Google Scholar]
- Afonso, M.V.; Barth, R.; Chauhan, A. Deep learning based plant part detection in Greenhouse settings. In Proceedings of the 12th EFITA International Conference: Digitizing Agriculture, European Federation for Information Technology in Agriculture, Food and the Environment (EFITA), Rhodes Island, Greece, 19 July 2019; pp. 48–53. [Google Scholar]
- Liu, B.; Tan, C.; Li, S.; He, J.; Wang, H. A Data Augmentation Method Based on Generative Adversarial Networks for Grape Leaf Disease Identification. IEEE Access 2020, 8, 102188–102198. [Google Scholar] [CrossRef]
- Yoshida, H.; Horie, T.; Katsura, K.; Shiraiwa, T. A model explaining genotypic and environmental variation in leaf area development of rice based on biomass growth and leaf N accumulation. F. Crop. Res. 2007, 102, 228–238. [Google Scholar] [CrossRef]
- Rasmussen, J.; Ntakos, G.; Nielsen, J.; Svensgaard, J.; Poulsen, R.N.; Christensen, S. Are vegetation indices derived from consumer-grade cameras mounted on UAVs sufficiently reliable for assessing experimental plots? Eur. J. Agron. 2016, 74, 75–92. [Google Scholar] [CrossRef]
- Inoue, Y.; Guérif, M.; Baret, F.; Skidmore, A.; Gitelson, A.; Schlerf, M.; Darvishzadeh, R.; Olioso, A. Simple and robust methods for remote sensing of canopy chlorophyll content: A comparative analysis of hyperspectral data for different types of vegetation. Plant Cell Environ. 2016, 39, 2609–2623. [Google Scholar] [CrossRef] [PubMed]
- Liang, L.; Di, L.; Zhang, L.; Deng, M.; Qin, Z.; Zhao, S.; Lin, H. Estimation of crop LAI using hyperspectral vegetation indices and a hybrid inversion method. Remote Sens. Environ. 2015, 165, 123–134. [Google Scholar] [CrossRef]
- Broge, N.H.; Leblanc, E. Comparing prediction power and stability of broadband and hyperspectral vegetation indices for estimation of green leaf area index and canopy chlorophyll density. Remote Sens. Environ. 2001, 76, 156–172. [Google Scholar] [CrossRef]
- Dong, T.; Liu, J.; Shang, J.; Qian, B.; Ma, B.; Kovacs, J.M.; Walters, D.; Jiao, X.; Geng, X.; Shi, Y. Assessment of red-edge vegetation indices for crop leaf area index estimation. Remote Sens. Environ. 2019, 222, 133–143. [Google Scholar] [CrossRef]
- Lecun, Y.; Bengio, Y.; Hinton, G. Deep learning. Nature 2015, 521, 436–444. [Google Scholar] [CrossRef]
- Kamilaris, A.; Prenafeta-Boldú, F.X. Deep learning in agriculture: A survey. Comput. Electron. Agric. 2018, 147, 70–90. [Google Scholar] [CrossRef]
- Maruyama, A.; Kuwagata, T.; Ohba, K. Measurement Error of Plant Area Index using Plant Canopy Analyzer and its Dependence on Mean Tilt Angle of The Foliage. J. Agric. Meteorol. 2005, 61, 229–233. [Google Scholar] [CrossRef][Green Version]
- Fang, H.; Li, W.; Wei, S.; Jiang, C. Seasonal variation of leaf area index (LAI) over paddy rice fields in NE China: Intercomparison of destructive sampling, LAI-2200, digital hemispherical photography (DHP), and AccuPAR methods. Agric. For. Meteorol. 2014, 198, 126–141. [Google Scholar] [CrossRef]
- Chen, J.M.; Rich, P.M.; Gower, S.T.; Norman, J.M.; Plummer, S. Leaf area index of boreal forests: Theory, techniques, and measurements. J. Geophys. Res. Atmos. 1997, 102, 29429–29443. [Google Scholar] [CrossRef]





| Camera | Spectral Band (nm) | Resolution (Pixels) |
|---|---|---|
| Rededge-MX (multispectral camera) | 475 (Blue), 560 (Green), 668 (Red), 717 (Rededge), 840 (NIR) | 1280 × 960 |
| Zenmuse X4S (RGB camera) | R, G, B | 5472 × 3648 |
| Index | Formula | Reference | |
|---|---|---|---|
| VIs | NDVI (λ1, λ2) | (Rλ1 − Rλ2)/(Rλ1 + Rλ2) | Jordan [34] |
| SR (λ1, λ2) | Rλ1/Rλ2 | Jordan [34] | |
| MSR (λ1, λ2) | ((Rλ1/Rλ2) − 1)/((Rλ1/Rλ2) + 1)0.5 | Chen [35] | |
| SAVI (λ1, λ2) | 1.5(Rλ1 − Rλ2)/(Rλ1 + Rλ2 + 0.5) | Huete [36] | |
| CIs | VARI | (g − r)/(g + r − b) | Gitelson et al. [37] |
| E × G | 2g − r – b | Woebbecke et al. [38] | |
| E × R | 1.4r – g | Meyer & Neto [39] | |
| E × B | 1.4b – g | Mao et al. [40] | |
| NGRDI | (g − r)/(g + r) | Tucker [41] | |
| MGRVI | (g2 − r2)/(g2 + r2) | Tucker [41] | |
| GLA | (2g − r − b)/(2g + r + b) | Louhaichi et al. [42] | |
| RGBVI | (g2 − b × r)/(g2 + b × r) | Bendig et al. [43] | |
| VEG | g/(rab(1 − a)), a = 0.667 | Hague et al. [44] | |
| Input Dataset | Epoch | Batch Size | Optimizer | Weight Decay |
|---|---|---|---|---|
| CIs | 100 | 16 | Adam | 0 |
| Images | 100 | 16 | Adam | 0 |
| CIs + Images | 100 | 8 | Adam | 0.01 |
| Index | Model | Regression Equation | |
|---|---|---|---|
| VI | NDVI (NIR, Red) | Exponential | y = 0.0809 × exp(4.41 × x) |
| NDVI (NIR, Rededge) | Linear | y = 8.17 × x − 0.363 | |
| NDVI (Rededge, Red) | Exponential | y = 0.112 × exp(5.09 × x) | |
| SR (NIR, Red) | Logarithmic | y = 1.58 × In(x) − 0.707 | |
| SR (NIR, Rededge) | Logarithmic | y = 3.10 × In(x) + 0.0131 | |
| SR (Rededge, Red) | Linear | y = 0.794 × x − 0.781 | |
| MSR (NIR, Red) | Linear | y = 0.859 × x − 0.154 | |
| MSR (NIR, Rededge) | Linear | y = 2.77 × x − 0.644 | |
| MSR (Rededge, Red) | Linear | y = 2.54 × x − 1.61 | |
| SAVI (NIR, Red) | Exponential | y = 0.0810 × exp(2.94 × x) | |
| SAVI (NIR, Rededge) | Linear | y = 5.45 × x − 0.363 | |
| SAVI (Rededge, Red) | Exponential | y = 0.113 × exp(3.39 × x) | |
| CI | VARI | Exponential | y = 0.252 × exp(5.74 × x) |
| E × G | Linear | y = 12.3 × x − 0.175 | |
| E × R | Exponential | y = 1.36 × exp(−11.7 × x) | |
| E×B | Linear | y = -24.7 × x + 3.25 | |
| NGRDI | Exponential | y = 0.275 × exp(9.72 × x) | |
| MGRVI | Exponential | y = 0.258 × exp(5.40 × x) | |
| GLA | Linear | y = 18.1 × x − 0.238 | |
| RGBVI | Exponential | y = 0.261 × exp(6.13 × x) | |
| VEG | Linear | y = 5.99 × x − 6.01 | |
| Algorithm | Input Dataset | Equation | R2 | RMSE |
|---|---|---|---|---|
| ANN | CIs | y = 1.00x | 0.940 | 0.401 |
| Images | y = 1.02x | 0.906 | 0.568 | |
| CIs + Images | y = 1.01x | 0.828 | 0.659 | |
| PLSR | CIs | y = 1.01x | 0.939 | 0.422 |
| Images | y = 0.957x | 0.252 | 1.697 | |
| CIs + Images | y = 0.982x | 0.715 | 0.940 | |
| RF | CIs | y = 1.02x | 0.939 | 0.436 |
| Images | y = 0.996x | 0.851 | 0.585 | |
| CIs + Images | y = 0.993x | 0.957 | 0.342 | |
| SVR | CIs | y = 0.932x | 0.945 | 0.399 |
| Images | y = 0.967x | 0.882 | 0.549 | |
| CIs + Images | y = 0.967x | 0.883 | 0.549 |
| Input Dataset | Equation | R2 | RMSE |
|---|---|---|---|
| CIs | y = 0.994x | 0.900 | 0.605 |
| Images | y = 0.991x | 0.979 | 0.280 |
| CIs + Images | y = 1.01x | 0.989 | 0.203 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yamaguchi, T.; Tanaka, Y.; Imachi, Y.; Yamashita, M.; Katsura, K. Feasibility of Combining Deep Learning and RGB Images Obtained by Unmanned Aerial Vehicle for Leaf Area Index Estimation in Rice. Remote Sens. 2021, 13, 84. https://doi.org/10.3390/rs13010084
Yamaguchi T, Tanaka Y, Imachi Y, Yamashita M, Katsura K. Feasibility of Combining Deep Learning and RGB Images Obtained by Unmanned Aerial Vehicle for Leaf Area Index Estimation in Rice. Remote Sensing. 2021; 13(1):84. https://doi.org/10.3390/rs13010084
Chicago/Turabian StyleYamaguchi, Tomoaki, Yukie Tanaka, Yuto Imachi, Megumi Yamashita, and Keisuke Katsura. 2021. "Feasibility of Combining Deep Learning and RGB Images Obtained by Unmanned Aerial Vehicle for Leaf Area Index Estimation in Rice" Remote Sensing 13, no. 1: 84. https://doi.org/10.3390/rs13010084
APA StyleYamaguchi, T., Tanaka, Y., Imachi, Y., Yamashita, M., & Katsura, K. (2021). Feasibility of Combining Deep Learning and RGB Images Obtained by Unmanned Aerial Vehicle for Leaf Area Index Estimation in Rice. Remote Sensing, 13(1), 84. https://doi.org/10.3390/rs13010084





