Effects of Electrospinning on the Viability of Ten Species of Lactic Acid Bacteria in Poly(Ethylene Oxide) Nanofibers
Abstract
1. Introduction
2. Materials and Methods
2.1. Materials
2.2. Bacterial Strains and Culturing Conditions
2.3. Characterization of Bacterial Strains
2.3.1. Determination of Cell Surface Charge of the Bacteria
2.3.2. Determination of Cell Surface Hydrophobicity of the Bacteria
2.3.3. Determination of Mass of the Bacterial Cells
2.3.4. Growth Curves of Lactobacillus spp. and L. lactis
2.4. Preparation of Polymer Solutions with the Bacteria
2.5. Rheological Characterization of Polymer Solutions with the Bacteria
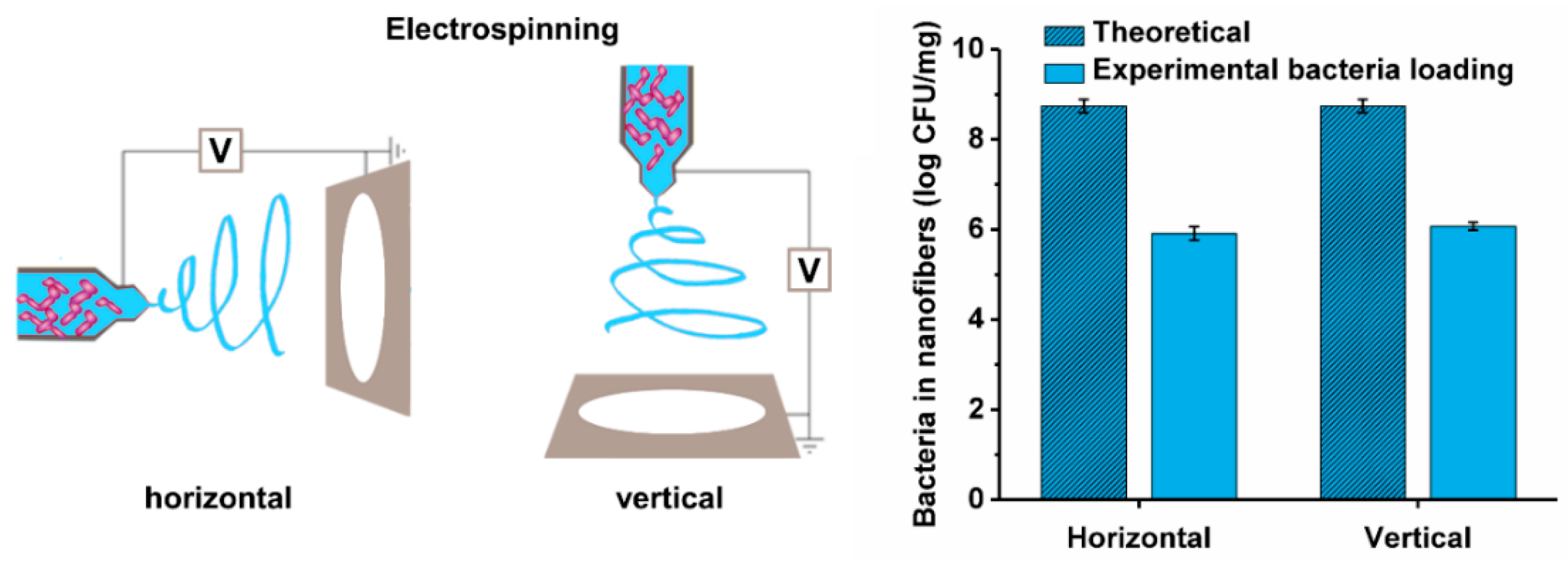
2.6. Preparation of Bacteria-Loaded Nanofibers by Electrospinning
2.7. Characterization of Nanofibers Loaded with the Bacteria
2.7.1. Morphology of the Bacterial Cells and Nanofibers
2.7.2. Viability of the Bacteria
2.8. Statistics
3. Results
3.1. Physical Characteristics of the Lactobacillus spp. and L. lactis
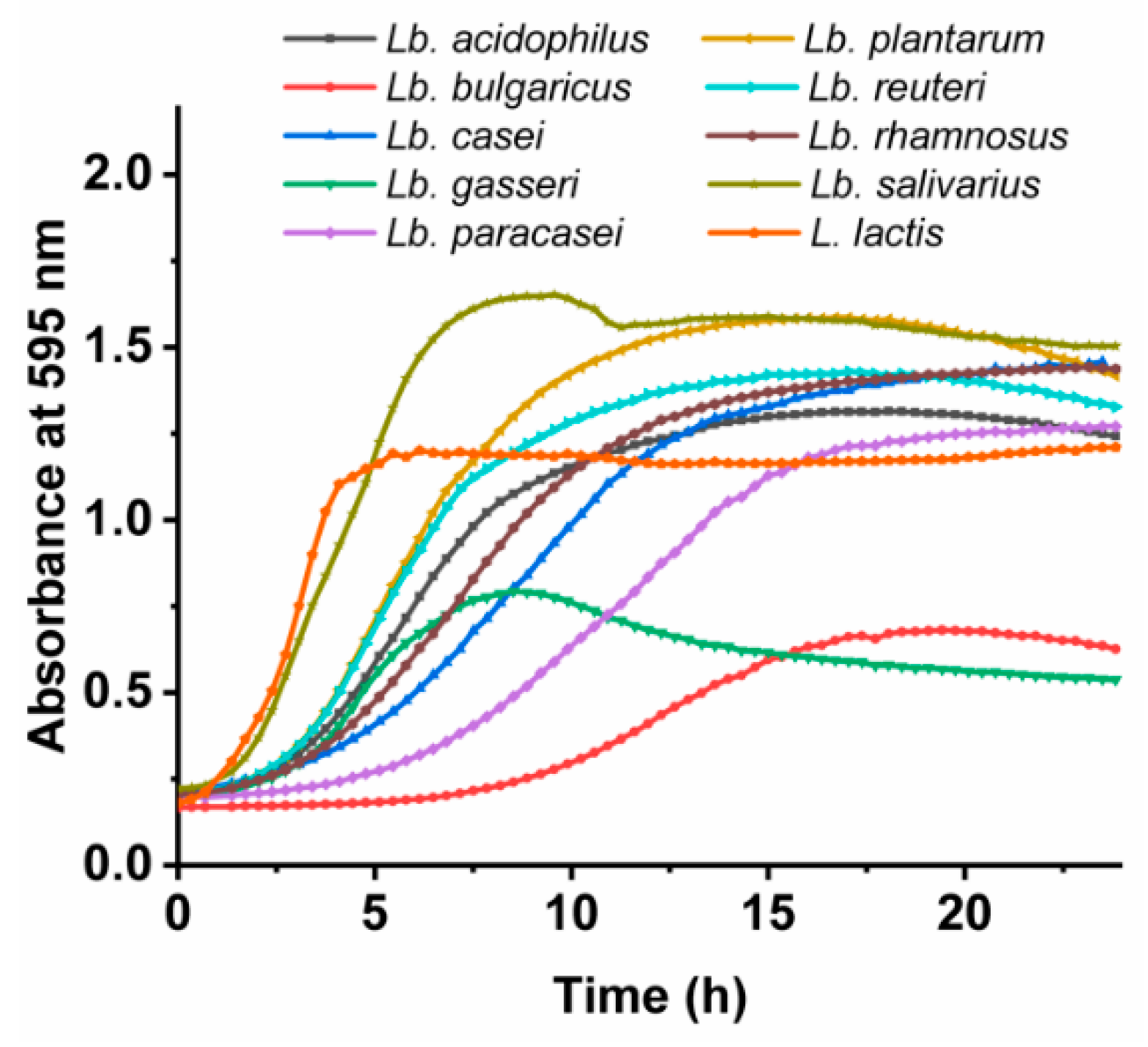
3.2. Growth Characteristics of the Lactobacillus spp. and L. lactis
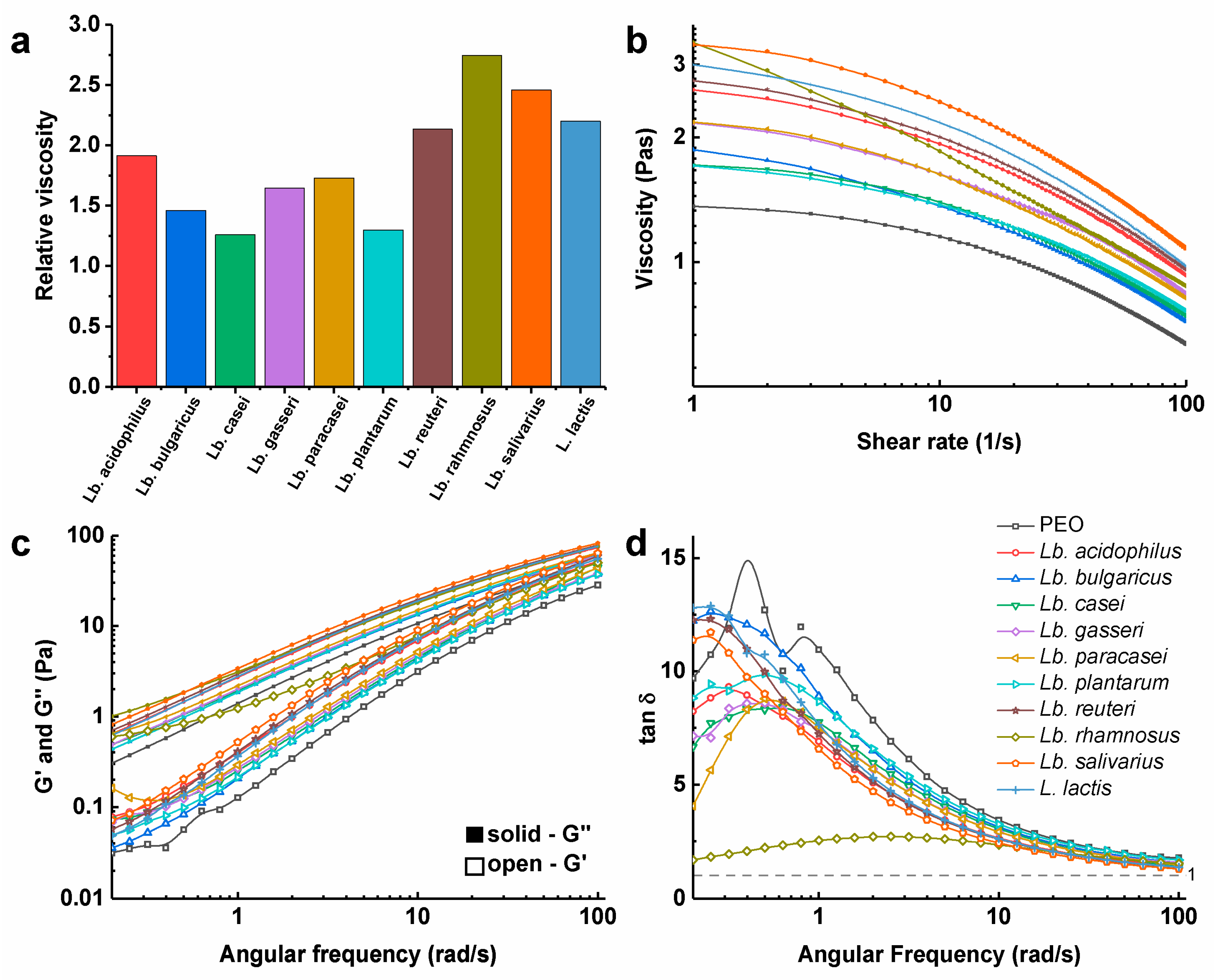
3.3. Viscosity of Polymer Solutions with the Dispersed Bacteria
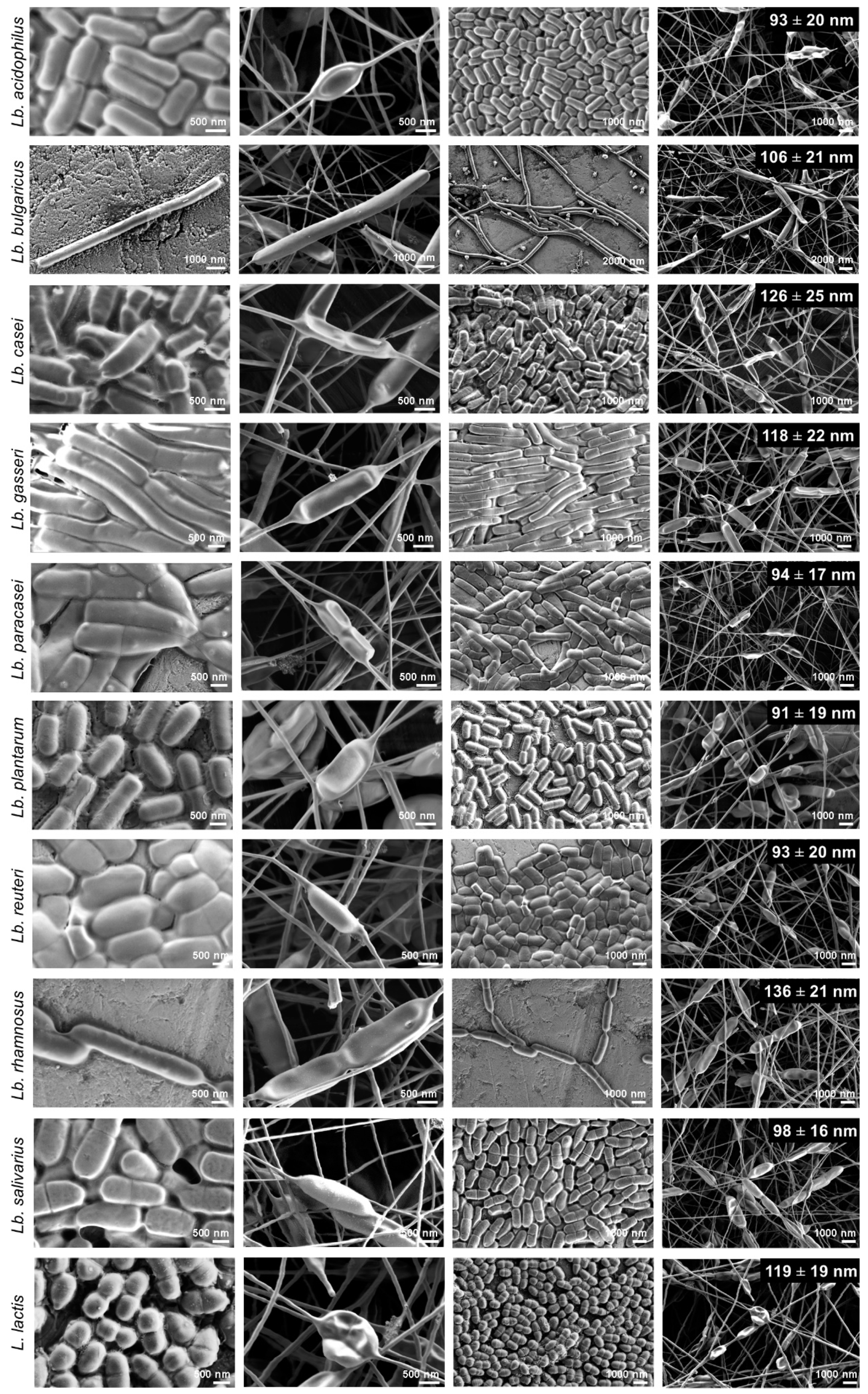
3.4. Morphology of Nanofibers with the Bacteria
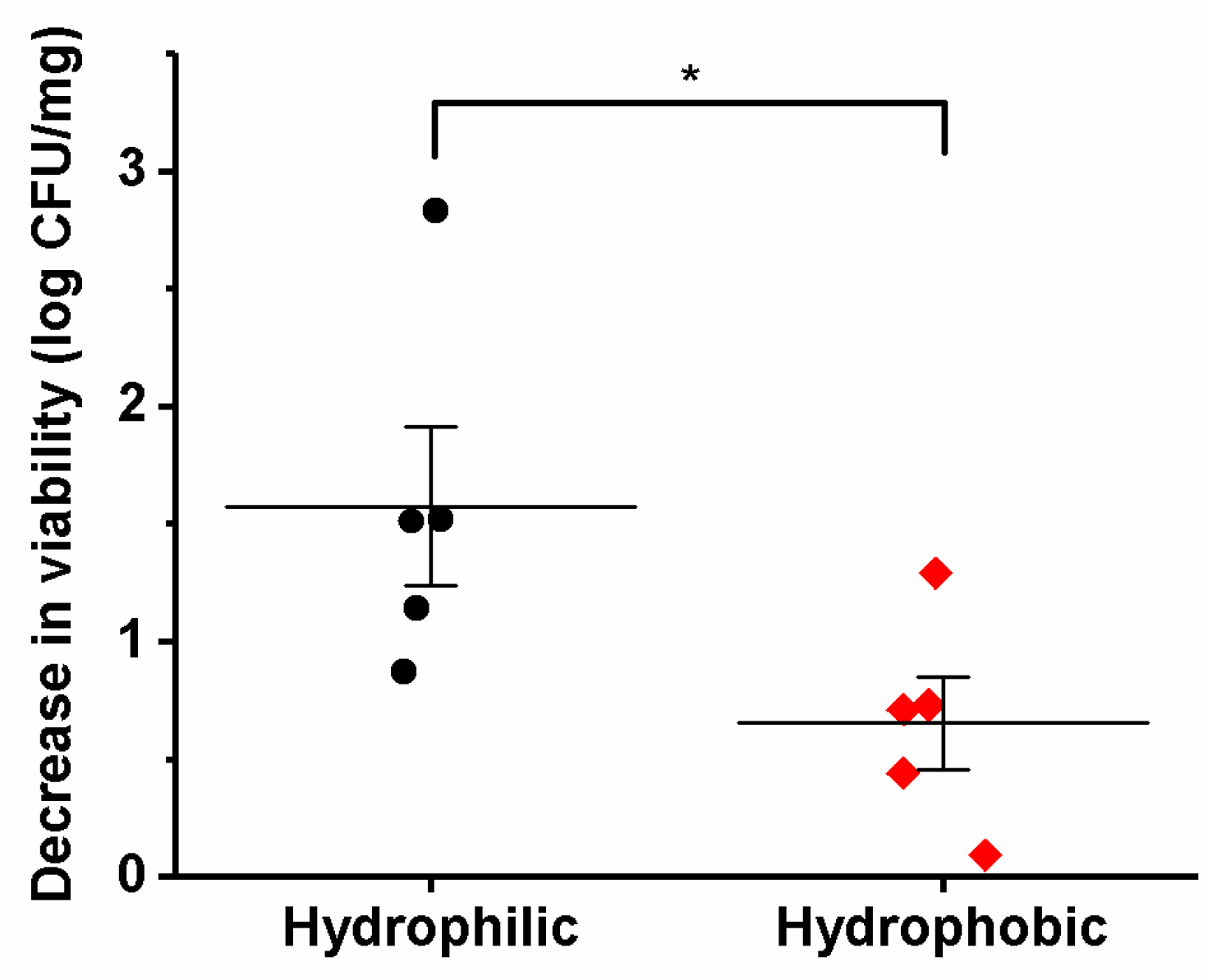
3.5. Viability of the Different Lactic Acid Bacteria after Electrospinning
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Salvetti, E.; Torriani, S.; Felis, G.E. The Genus Lactobacillus: A Taxonomic Update. Probiotics Antimicrob. Proteins 2012, 4, 217–226. [Google Scholar] [CrossRef] [PubMed]
- Sun, Z.; Harris, H.M.; McCann, A.; Guo, C.; Argimon, S.; Zhang, W.; Yang, X.; Jeffery, I.B.; Cooney, J.C.; Kagawa, T.F.; et al. Expanding the biotechnology potential of lactobacilli through comparative genomics of 213 strains and associated genera. Nat. Commun. 2015, 6, 8322. [Google Scholar] [CrossRef] [PubMed]
- Bosma, E.F.; Forster, J.; Nielsen, A.T. Lactobacilli and pediococci as versatile cell factories - Evaluation of strain properties and genetic tools. Biotechnol. Adv. 2017, 35, 419–442. [Google Scholar] [CrossRef] [PubMed]
- Guarino, A.; Guandalini, S.; Lo Vecchio, A. Probiotics for Prevention and Treatment of Diarrhea. J. Clin. Gastroenterol. 2015, 49 (Suppl. 1), S37–S45. [Google Scholar] [CrossRef]
- AlFaleh, K.; Anabrees, J. Probiotics for prevention of necrotizing enterocolitis in preterm infants. Cochrane Database Syst. Rev. 2014. [Google Scholar] [CrossRef] [PubMed]
- Cai, J.; Zhao, C.; Du, Y.; Zhang, Y.; Zhao, M.; Zhao, Q. Comparative efficacy and tolerability of probiotics for antibiotic-associated diarrhea: Systematic review with network meta-analysis. United Eur. Gastroenterol. J. 2018, 6, 169–180. [Google Scholar] [CrossRef] [PubMed]
- Ganji-Arjenaki, M.; Rafieian-Kopaei, M. Probiotics are a good choice in remission of inflammatory bowel diseases: A meta analysis and systematic review. J. Cell. Physiol. 2018, 233, 2091–2103. [Google Scholar] [CrossRef] [PubMed]
- Rupa, P.; Mine, Y. Recent advances in the role of probiotics in human inflammation and gut health. J. Agric. Food Chem. 2012, 60, 8249–8256. [Google Scholar] [CrossRef] [PubMed]
- Matsubara, V.H.; Bandara, H.M.; Ishikawa, K.H.; Mayer, M.P.; Samaranayake, L.P. The role of probiotic bacteria in managing periodontal disease: A systematic review. Expert Rev. Anti. Infect. Ther. 2016, 14, 643–655. [Google Scholar] [CrossRef]
- Borges, S.; Barbosa, J.; Teixeira, P. Drug Delivery Systems for Vaginal Infections. Front. Clin. Drug Res. 2016, 2, 233. [Google Scholar]
- Petrova, M.I.; Lievens, E.; Malik, S.; Imholz, N.; Lebeer, S. Lactobacillus species as biomarkers and agents that can promote various aspects of vaginal health. Front. Physiol. 2015, 6, 81. [Google Scholar] [CrossRef] [PubMed]
- Belkaid, Y.; Naik, S. Compartmentalized and systemic control of tissue immunity by commensals. Nat. Immunol. 2013, 14, 646. [Google Scholar] [CrossRef] [PubMed]
- Sycuro, L.K.; Fredricks, D.N. Microbiota of the Genitourinary Tract. The Human Microbiota: How Microbial Communities Affect. Health and Disease; John Wiley Sons: Hoboken, NJ, USA, 2013; pp. 167–210. [Google Scholar]
- Ravel, J.; Gajer, P.; Abdo, Z.; Schneider, G.M.; Koenig, S.S.; McCulle, S.L.; Karlebach, S.; Gorle, R.; Russell, J.; Tacket, C.O.; et al. Vaginal microbiome of reproductive-age women. Proc. Natl. Acad. Sci. USA 2011, 108, 4680–4687. [Google Scholar] [CrossRef] [PubMed]
- Gao, S.; Tang, G.; Hua, D.; Xiong, R.; Han, J.; Jiang, S.; Zhang, Q.; Huang, C. Stimuli-responsive bio-based polymeric systems and their applications. J. Mater. Chem. B 2019, 7, 709–729. [Google Scholar] [CrossRef]
- Ding, Q.; Xu, X.; Yue, Y.; Mei, C.; Huang, C.; Jiang, S.; Wu, Q.; Han, J. Nanocellulose-Mediated Electroconductive Self-Healing Hydrogels with High Strength, Plasticity, Viscoelasticity, Stretchability, and Biocompatibility toward Multifunctional Applications. ACS App. Mater. Inter. 2018, 10, 27987–28002. [Google Scholar] [CrossRef] [PubMed]
- Ghorani, B.; Tucker, N. Fundamentals of electrospinning as a novel delivery vehicle for bioactive compounds in food nanotechnology. Food Hydrocolloid 2015, 51, 227–240. [Google Scholar] [CrossRef]
- Mirtič, J.; Rijavec, T.; Zupančič, S.; Zvonar Pobirk, A.; Lapanje, A.; Kristl, J. Development of probiotic-loaded microcapsules for local delivery: Physical properties, cell release and growth. Eur. J. Pharm. Sci. 2018, 121, 178–187. [Google Scholar] [CrossRef] [PubMed]
- Zupancčič, Š.; Rijavec, T.; Lapanje, A.; Petelin, M.; Kristl, J.; Kocbek, P. Nanofibers with Incorporated Autochthonous Bacteria as Potential Probiotics for Local Treatment of Periodontal Disease. Biomacromolecules 2018, 19, 4299–4306. [Google Scholar] [CrossRef]
- Hu, X.; Liu, S.; Zhou, G.; Huang, Y.; Xie, Z.; Jing, X. Electrospinning of polymeric nanofibers for drug delivery applications. J. Control. Release 2014, 185, 12–21. [Google Scholar] [CrossRef]
- Chou, S.F.; Carson, D.; Woodrow, K.A. Current strategies for sustaining drug release from electrospun nanofibers. J. Control. Release 2015, 220, 584–591. [Google Scholar] [CrossRef]
- Zupančič, Š. Core-shell nanofibers as drug delivery systems. Acta Pharm. 2019, 69, 131–153. [Google Scholar] [CrossRef] [PubMed]
- Fung, W.Y.; Yuen, K.H.; Liong, M.T. Agrowaste-based nanofibers as a probiotic encapsulant: Fabrication and characterization. J. Agric. Food Chem. 2011, 59, 8140–8147. [Google Scholar] [CrossRef] [PubMed]
- Lopez-Rubio, A.; Sanchez, E.; Wilkanowicz, S.; Sanz, Y.; Lagaron, J.M. Electrospinning as a useful technique for the encapsulation of living bifidobacteria in food hydrocolloids. Food Hydrocolloid 2012, 28, 159–167. [Google Scholar] [CrossRef]
- Nagy, Z.K.; Wagner, I.; Suhajda, A.; Tobak, T.; Harasztos, A.H.; Vigh, T.; Soti, P.L.; Pataki, H.; Molnar, K.; Marosi, G. Nanofibrous solid dosage form of living bacteria prepared by electrospinning. Express Polym. Lett. 2014, 8, 352–361. [Google Scholar] [CrossRef]
- Skrlec, K.; Zupancic, S.; Prpar Mihevc, S.; Kocbek, P.; Kristl, J.; Berlec, A. Development of electrospun nanofibers that enable high loading and long-term viability of probiotics. Eur. J. Pharm. Biopharm. 2019, 136, 108–119. [Google Scholar] [CrossRef] [PubMed]
- Ceylan, Z.; Meral, R.; Karakaş, C.Y.; Dertli, E.; Yilmaz, M.T. A novel strategy for probiotic bacteria: Ensuring microbial stability of fish fillets using characterized probiotic bacteria-loaded nanofibers. Innov. Food Sci. Emerg. Technol. 2018, 48, 212–218. [Google Scholar] [CrossRef]
- Amna, T.; Hassan, M.S.; Pandeya, D.R.; Khil, M.S.; Hwang, I.H. Classy non-wovens based on animate L. gasseri-inanimate poly(vinyl alcohol): Upstream application in food engineering. Appl. Microbiol. Biotechnol. 2013, 97, 4523–4531. [Google Scholar] [CrossRef] [PubMed]
- Palomino, M.M.; Allievi, M.C.; Fina Martin, J.; Waehner, P.M.; Prado Acosta, M.; Sanchez Rivas, C.; Ruzal, S.M. Draft Genome Sequence of the Probiotic Strain Lactobacillus acidophilus ATCC 4356. Genome Announc. 2015, 3. [Google Scholar] [CrossRef] [PubMed]
- van de Guchte, M.; Penaud, S.; Grimaldi, C.; Barbe, V.; Bryson, K.; Nicolas, P.; Robert, C.; Oztas, S.; Mangenot, S.; Couloux, A.; et al. The complete genome sequence of Lactobacillus bulgaricus reveals extensive and ongoing reductive evolution. Proc. Natl. Acad. Sci. USA 2006, 103, 9274–9279. [Google Scholar] [CrossRef] [PubMed]
- Toh, H.; Oshima, K.; Nakano, A.; Takahata, M.; Murakami, M.; Takaki, T.; Nishiyama, H.; Igimi, S.; Hattori, M.; Morita, H. Genomic adaptation of the Lactobacillus casei group. PLoS ONE 2013, 8, e75073. [Google Scholar] [CrossRef] [PubMed]
- Azcarate-Peril, M.A.; Altermann, E.; Goh, Y.J.; Tallon, R.; Sanozky-Dawes, R.B.; Pfeiler, E.A.; O’Flaherty, S.; Buck, B.L.; Dobson, A.; Duong, T.; et al. Analysis of the genome sequence of Lactobacillus gasseri ATCC 33323 reveals the molecular basis of an autochthonous intestinal organism. Appl. Environ. Microbiol. 2008, 74, 4610–4625. [Google Scholar] [CrossRef] [PubMed]
- Saulnier, D.M.; Santos, F.; Roos, S.; Mistretta, T.A.; Spinler, J.K.; Molenaar, D.; Teusink, B.; Versalovic, J. Exploring metabolic pathway reconstruction and genome-wide expression profiling in Lactobacillus reuteri to define functional probiotic features. PLoS ONE 2011, 6, e18783. [Google Scholar] [CrossRef] [PubMed]
- Morita, H.; Toh, H.; Oshima, K.; Murakami, M.; Taylor, T.D.; Igimi, S.; Hattori, M. Complete genome sequence of the probiotic Lactobacillus rhamnosus ATCC 53103. J. Bacteriol. 2009, 191, 7630–7631. [Google Scholar] [CrossRef] [PubMed]
- Linares, D.M.; Kok, J.; Poolman, B. Genome sequences of Lactococcus lactis MG1363 (revised) and NZ9000 and comparative physiological studies. J. Bacteriol. 2010, 192, 5806–5812. [Google Scholar] [CrossRef] [PubMed]
- Perez, P.F.; Minnaard, Y.; Disalvo, E.A.; De Antoni, G.L. Surface properties of bifidobacterial strains of human origin. Appl. Environ. Microbiol. 1998, 64, 21–26. [Google Scholar] [PubMed]
- Baranyi, J.; Roberts, T.A. A dynamic approach to predicting bacterial growth in food. Int. J. Food Microbiol. 1994, 23, 277–294. [Google Scholar] [CrossRef]
- Herigstad, B.; Hamilton, M.; Heersink, J. How to optimize the drop plate method for enumerating bacteria. J. Microbiol. Methods 2001, 44, 121–129. [Google Scholar] [CrossRef]
- Lebeer, S.; Verhoeven, T.L.; Francius, G.; Schoofs, G.; Lambrichts, I.; Dufrene, Y.; Vanderleyden, J.; De Keersmaecker, S.C. Identification of a Gene Cluster for the Biosynthesis of a Long, Galactose-Rich Exopolysaccharide in Lactobacillus rhamnosus GG and Functional Analysis of the Priming Glycosyltransferase. Appl. Environ. Microbiol. 2009, 75, 3554–3563. [Google Scholar] [CrossRef]
- Zeidan, A.A.; Poulsen, V.K.; Janzen, T.; Buldo, P.; Derkx, P.M.F.; Oregaard, G.; Neves, A.R. Polysaccharide production by lactic acid bacteria: From genes to industrial applications. FEMS Microbiol. Rev. 2017, 41, S168–S200. [Google Scholar] [CrossRef]
- Reznik, S.N.; Yarin, A.L.; Zussman, E.; Bercovici, L. Evolution of a compound droplet attached to a core-shell nozzle under the action of a strong electric field. Phys. Fluids 2006, 18, 062101. [Google Scholar] [CrossRef]
- Kajdič, S.; Vrečer, F.; Kocbek, P. Preparation of poloxamer-based nanofibers for enhanced dissolution of carvedilol. Eur. J. Pharm. Sci. 2018, 117, 331–340. [Google Scholar] [CrossRef] [PubMed]
- Krasowska, A.; Sigler, K. How microorganisms use hydrophobicity and what does this mean for human needs? Front. Cell Infect. Microbiol. 2014, 4, 112. [Google Scholar] [CrossRef] [PubMed]
- Zadravec, P.; Štrukelj, B.; Berlec, A. Improvement of LysM-mediated surface display of designed ankyrin repeat proteins (DARPins) in recombinant and nonrecombinant strains of Lactococcus lactis and Lactobacillus Species. Appl. Environ. Microbiol. 2015, 81, 2098–2106. [Google Scholar] [CrossRef] [PubMed][Green Version]
| Strain | Source | Fermentation Type | Genome Size [bp] | Reference |
|---|---|---|---|---|
| Lactobacillus sp. | ||||
| Lb. acidophilus ATCC 4356 | Infant feces | Homofermentative | 1956.699 | [29] |
| Lb. delbrueckii ssp. bulgaricus ATCC 11842 | Yoghurt | Homofermentative | 1864.998 | [30] |
| Lb. casei ATCC 393 | Cheese | Facultative heterofermentative | 2924.929 | [31] |
| Lb. gasseri ATCC 33323 | Human isolate | Homofermentative | 1894.360 | [32] |
| Lb. paracasei ATCC 25302 | Milk product | Facultative heterofermentative | 2991.737 | NCBI |
| Lb. plantarum ATCC 8014 | n/a | Facultative heterofermentative | 3254.764 | NCBI |
| Lb. reuteri ATCC 55730 | Breast milk | Heterofermentative | 2036.000 | [33] |
| Lb. rhamnosus ATCC 53103 | Human intestine | Homofermentative | 3005.051 | [34] |
| Lb. salivarius ATCC 11741 | Infant feces | Homofermentative | 1956.699 | NCBI |
| Lactococcus sp. | ||||
| Lactococcus lactis ssp. cremoris MG1363 | Cheese | Homofermentative | 2529.478 | [35] |
| Bacteria Species | Average Cell Width [µm] | Average Cell Length [µm] | Zeta Potential [mV] | Hydrophobicity [%] | Mass of 1 × 1010 Bacterial Cells [mg] |
|---|---|---|---|---|---|
| Lb. acidophilus | 0.56 ± 0.04 | 1.28 ± 0.26 | −9.1 ± 5.0 | 78.6 ± 1.0 | 0.65 |
| Lb. delbrueckii ssp. bulgaricus | 0.51 ± 0.07 | 10.82 ± 3.31 | −9.4 ± 5.5 | 28.5 ± 3.9 | 9.14 |
| Lb. casei | 0.58 ± 0.06 | 1.54 ± 0.36 | −0.4 ± 5.5 | 11.3 ± 0.5 | 4.22 |
| Lb. gasseri | 0.65 ± 0.07 | 4.68 ± 1.43 | −7.9 ± 4.6 | 92.5 ± 2.1 | 5.59 |
| Lb. paracasei | 0.68 ± 0.09 | 2.05 ± 0.49 | −23.9 ± 4.4 | 36.4 ± 4.8 | 2.13 |
| Lb. plantarum | 0.52 ± 0.04 | 1.33 ± 0.29 | −12.7 ± 4.0 | 74.2 ± 2.3 | 0.54 |
| Lb. reuteri | 0.72 ± 0.08 | 1.43 ± 0.37 | −13.7 ± 5.7 | 71.9 ± 5.8 | 1.04 |
| Lb. rhamnosus | 0.64 ± 0.08 | 2.16 ± 0.51 | −3.9 ± 4.4 | 31.5 ± 7.7 | 16.31 |
| Lb. salivarius | 0.70 ± 0.07 | 1.39 ± 0.31 | −11.7 ± 4.6 | 90.1 ± 1.3 | 1.92 |
| L. lactis | 0.53 ± 0.05 | 0.74 ± 0.18 | −12.8 ± 5.5 | 24.2 ± 6.3 | 0.48 |
| Bacterial Species | Growth Rate [/h] | Lag Time [h] |
|---|---|---|
| Lb. acidophilus | 0.158 ± 0.005 | 2.31 ± 0.09 |
| Lb. bulgaricus | 0.066 ± 0.001 | 8.14 ± 0.15 |
| Lb. casei | 0.116 ± 0.002 | 3.41 ± 0.07 |
| Lb. gasseri | 0.118 ± 0.004 | 1.60 ± 0.15 |
| Lb. paracasei | 0.102 ± 0.010 | 5.50 ± 0.23 |
| Lb. plantarum | 0.199 ± 0.002 | 2.52 ± 0.21 |
| Lb. reuteri | 0.193 ± 0.009 | 1.82 ± 0.51 |
| Lb. rhamnosus | 0.125 ± 0.001 | 1.92 ± 0.34 |
| Lb. salivarius | 0.297 ± 0.028 | 1.11 ± 0.08 |
| L. lactis | 0.326 ± 0.013 | 0.84 ± 0.07 |
| Bacterial Species | Theoretical Bacteria Loading (log CFU/mg) | Experimental Bacteria Loading (log CFU/mg) | Decrease in Viability (log CFU/mg) |
|---|---|---|---|
| Lb. acidophilus | 9.89 ± 0.18 | 9.18 ± 0.21 | 0.71 |
| Lb. bulgaricus | 8.74 ± 0.15 | 5.91 ± 0.15 | 2.83 |
| Lb. casei | 8.85 ± 0.21 | 7.33 ± 0.19 | 1.52 |
| Lb. gasseri | 9.30 ± 0.18 | 9.20 ± 0.11 | 0.09 |
| Lb. paracasei | 9.53 ± 0.12 | 8.02 ± 0.18 | 1.51 |
| Lb. plantarum | 9.99 ± 0.06 | 8.70 ± 0.38 | 1.29 |
| Lb. reuteri | 9.76 ± 0.13 | 9.32 ± 0.04 | 0.44 |
| Lb. rhamnosus | 8.55 ± 0.13 | 7.41 ± 0.16 | 1.14 |
| Lb. salivarius | 9.21 ± 0.06 | 6.75 ± 0.07 | 0.73 |
| L. lactis | 9.06 ± 0.14 | 8.19 ± 0.33 | 0.87 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zupančič, Š.; Škrlec, K.; Kocbek, P.; Kristl, J.; Berlec, A. Effects of Electrospinning on the Viability of Ten Species of Lactic Acid Bacteria in Poly(Ethylene Oxide) Nanofibers. Pharmaceutics 2019, 11, 483. https://doi.org/10.3390/pharmaceutics11090483
Zupančič Š, Škrlec K, Kocbek P, Kristl J, Berlec A. Effects of Electrospinning on the Viability of Ten Species of Lactic Acid Bacteria in Poly(Ethylene Oxide) Nanofibers. Pharmaceutics. 2019; 11(9):483. https://doi.org/10.3390/pharmaceutics11090483
Chicago/Turabian StyleZupančič, Špela, Katja Škrlec, Petra Kocbek, Julijana Kristl, and Aleš Berlec. 2019. "Effects of Electrospinning on the Viability of Ten Species of Lactic Acid Bacteria in Poly(Ethylene Oxide) Nanofibers" Pharmaceutics 11, no. 9: 483. https://doi.org/10.3390/pharmaceutics11090483
APA StyleZupančič, Š., Škrlec, K., Kocbek, P., Kristl, J., & Berlec, A. (2019). Effects of Electrospinning on the Viability of Ten Species of Lactic Acid Bacteria in Poly(Ethylene Oxide) Nanofibers. Pharmaceutics, 11(9), 483. https://doi.org/10.3390/pharmaceutics11090483





