African Horse Sickness: A Review of Current Understanding and Vaccine Development
Abstract
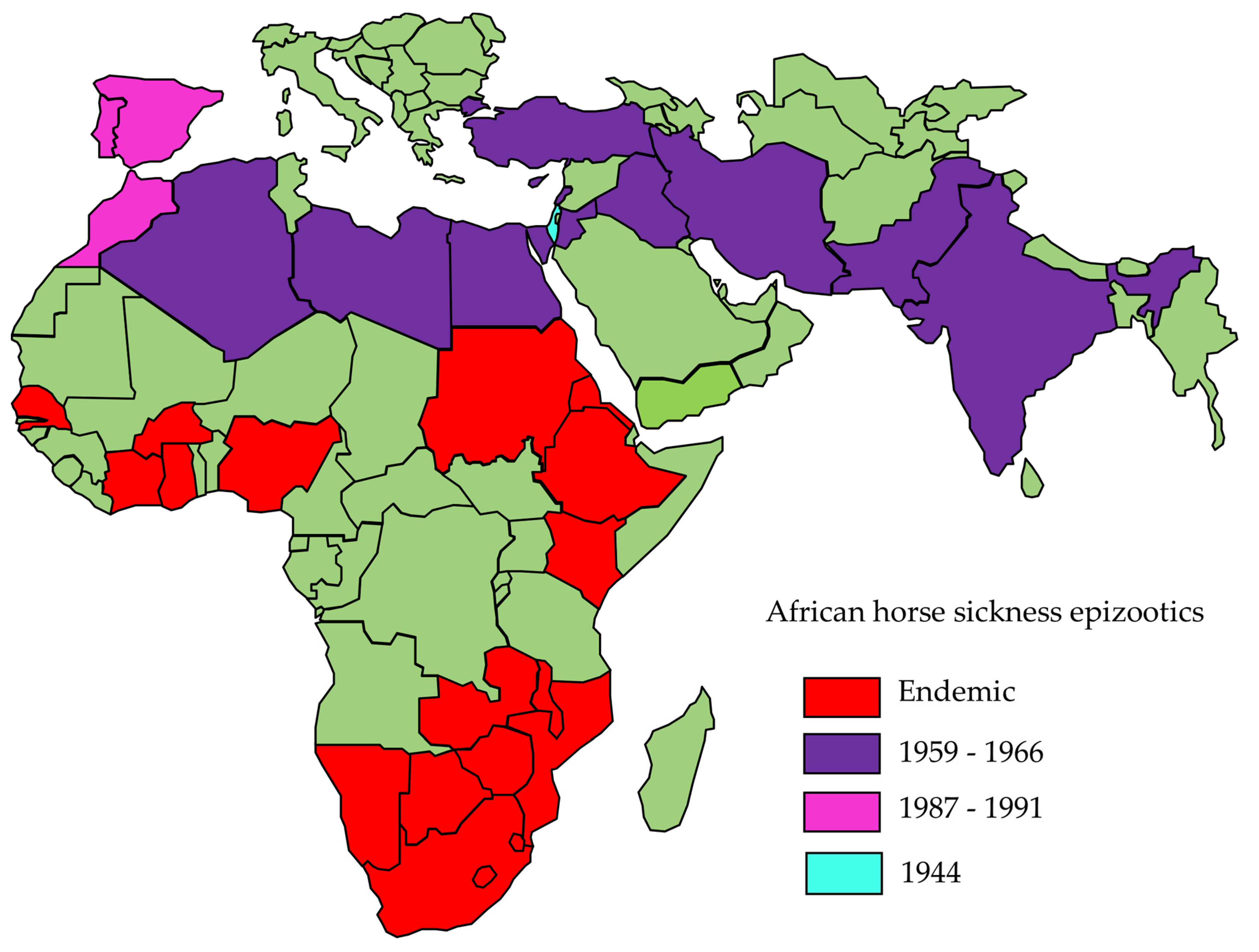
1. Introduction
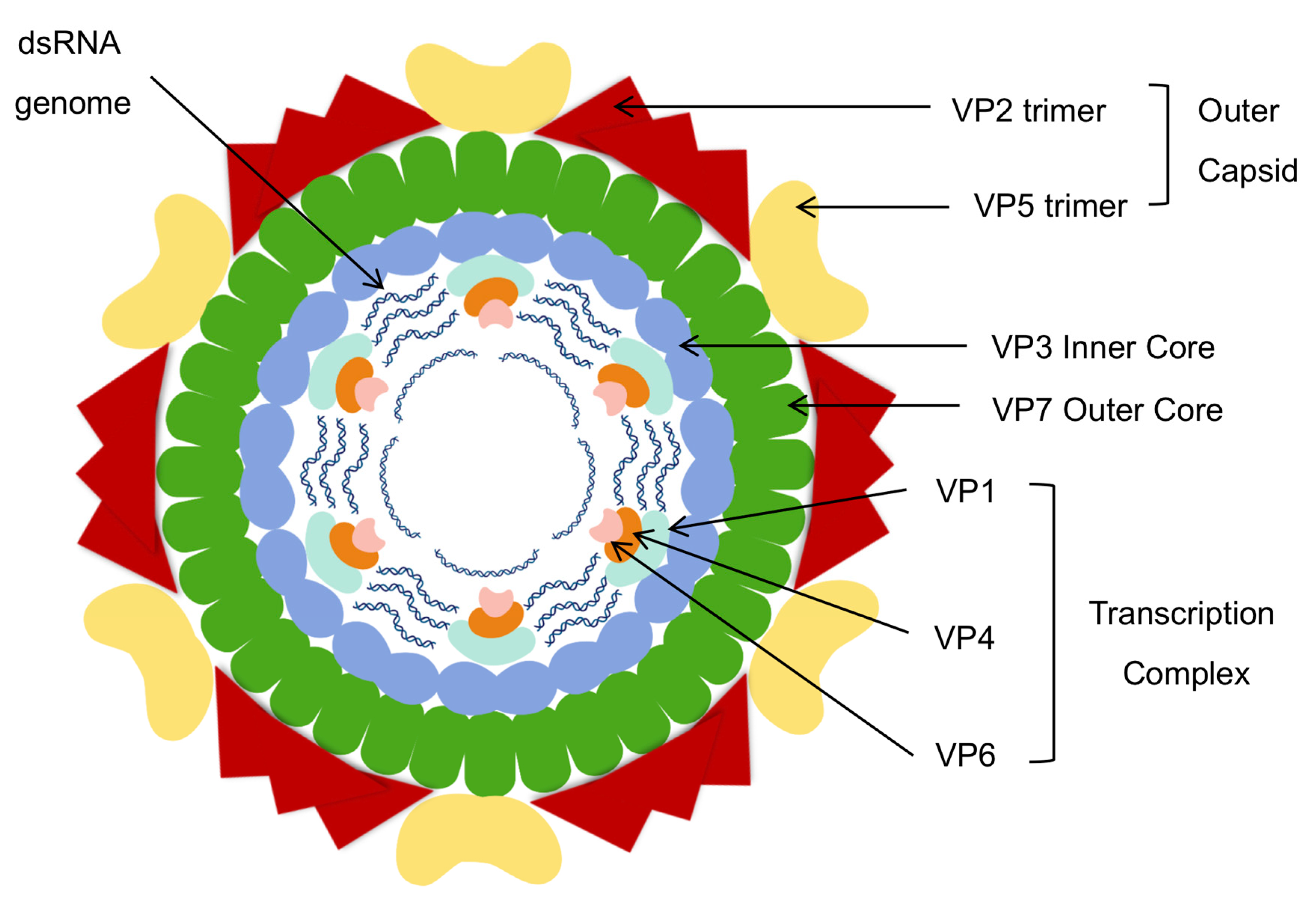
2. African Horse Sickness Virus
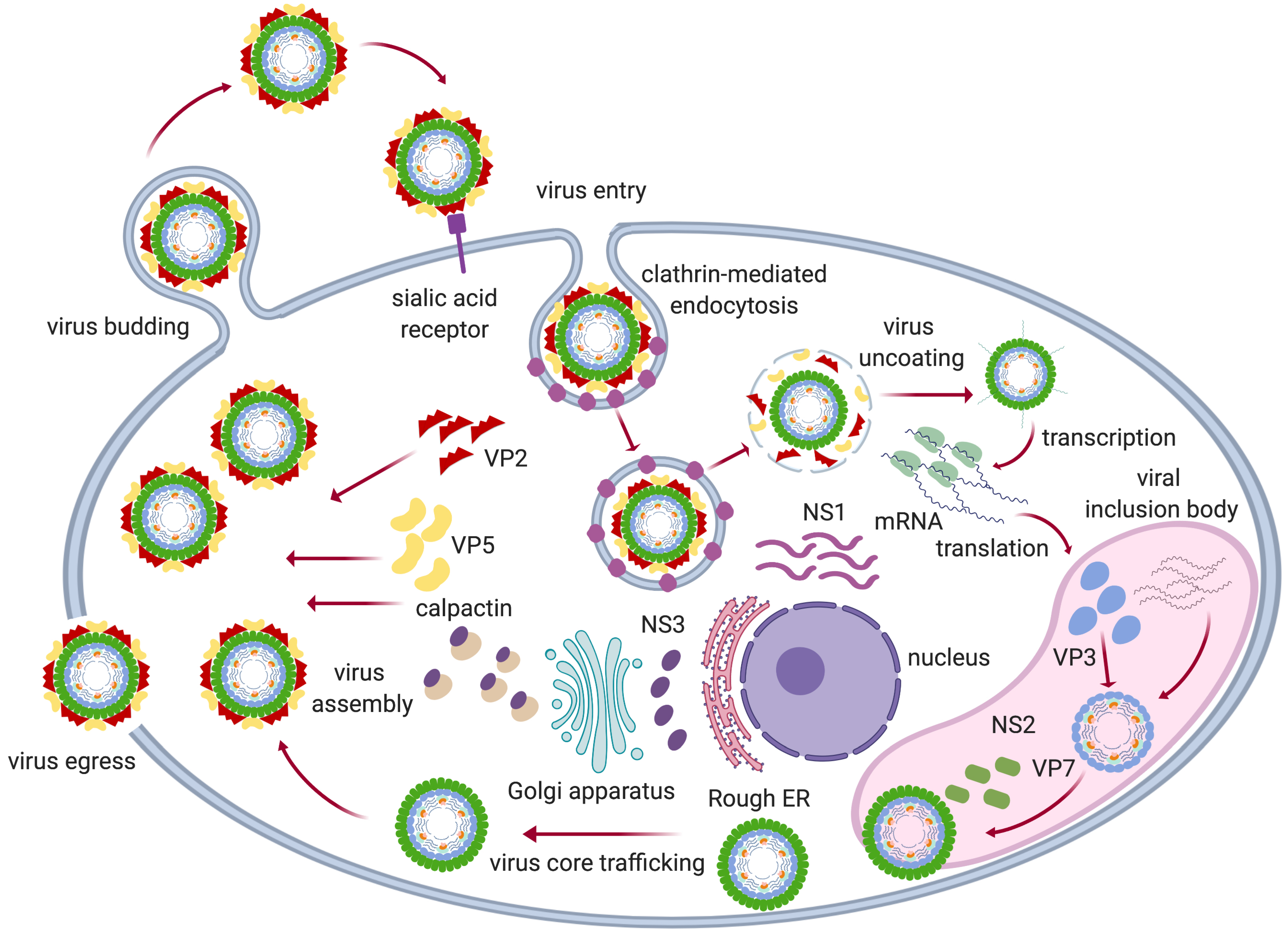
3. Viral Infection and Replication
4. African Horse Sickness Disease
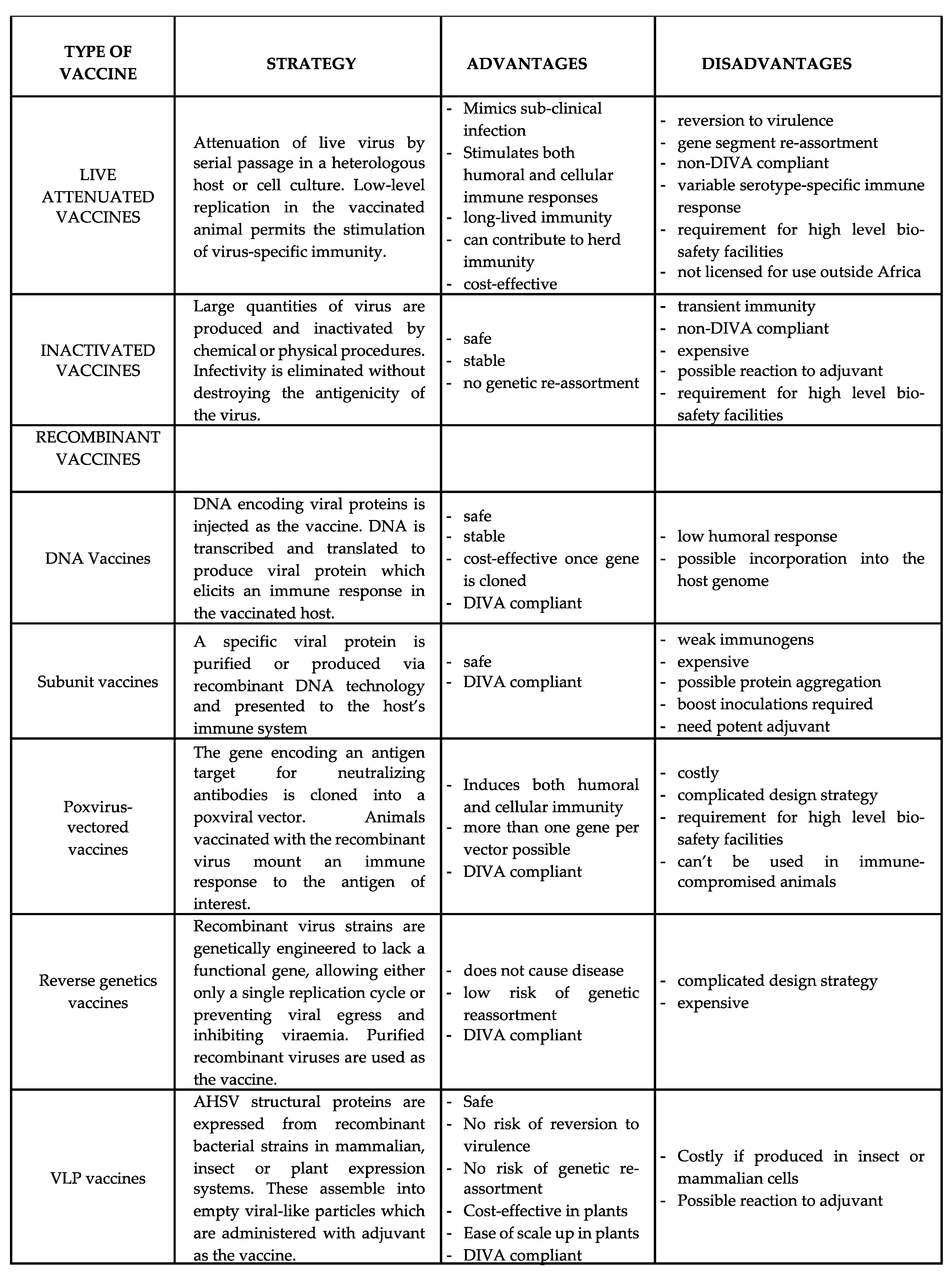
5. Prevention and Control
5.1. Live Attenuated Vaccines
5.2. Inactivated Vaccines
5.3. Recombinant Vaccines
5.3.1. DNA Vaccines
5.3.2. Subunit Vaccines
5.3.3. Poxvirus-Vectored Vaccines
5.3.4. Reverse Genetics Vaccines
5.3.5. Virus-Like Particle Vaccines
6. Conclusions
Patents
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Mellor, P.S.; Hamblin, C. African horse sickness. Vet. Res. 2004, 35, 445–466. [Google Scholar] [CrossRef] [PubMed]
- Henning, M.W. African Horse Sickness, Perdesiekte, Pestis Equorum. In Animal Diseases of South Africa, 3rd ed.; Central News Agency Ltd.: Pretoria, Africa, 1956; pp. 785–808. [Google Scholar]
- Coetzer, J.A.W.; Guthrie, A.J. African horse sickness. In Infectious Diseases of Livestock, 2nd ed.; Coetzer, J.A.W., Tustin, R.C., Eds.; Oxford University Press: Cape Town, Africa, 2004; pp. 1231–1246. [Google Scholar]
- Vandenbergh, S. The story of a disease: African horsesickness and its direct influence on the necessary development of veterinary science in South Africa c. 1890s–1920s. Historia 2010, 55, 243–262. [Google Scholar]
- Lubroth, J. African Horsesickness and the epizootic in Spain, 1987. Equine Pract. 1988, 10, 26–33. [Google Scholar]
- Mirchamsy, H.; Hazrati, A. A review on etiology and pathogeny of African horse sickness. Arch. Razi Inst. 1973, 25, 23–46. [Google Scholar]
- Mellor, P. African horse sickness: Transmission and epidemiology. Vet. Res. 1993, 24, 199–212. [Google Scholar] [PubMed]
- Howell, P.G. The 1960 epizootic of African Horsesickness in the Middle East and SW Asia. J. S. Afr. Vet. Assoc. 1960, 31, 329–334. [Google Scholar]
- De Vos, C.J.; Hoek, C.A.; Nodelijk, G. Risk of introducing African horse sickness virus into the Netherlands by international equine movements. Prev. Vet. Med. 2012, 106, 108–122. [Google Scholar] [CrossRef] [PubMed]
- Herholz, C.; Fussel, A.E.; Timoney, P.; Schwermer, H.; Bruckner, L.; Leadon, D. Equine travellers to the Olympic Games in Hong Kong 2008: A review of worldwide challenges to equine health, with particular reference to vector-borne diseases. Equine Vet. J. 2008, 40, 87–95. [Google Scholar] [CrossRef] [PubMed]
- Hopley, R.; Toth, B. Focus on: African horse sickness. Vet. Rec. 2013, 173, 13–14. [Google Scholar] [CrossRef]
- M’Fadyean, J. African horse-sickness. J. Comp. Pathol. Ther. 1900, 13, 1–20. [Google Scholar] [CrossRef]
- Howell, P.G. The isolation and identification of further antigenic types of African horsesickness virus. Onderstepoort J. Vet. Res. 1962, 29, 139–149. [Google Scholar]
- McIntosh, B.M. Immunological types of horsesickness virus and their significance in immunization. Onderstepoort J. Vet. Res. 1958, 27, 465–536. [Google Scholar]
- Calisher, C.H.; Mertens, P.P. Taxonomy of African horse sickness viruses. In African Horse Sickness; Mellor, P.S., Baylis, M., Hamblin, C., Mertens, P.P.C., Calisher, C.H., Eds.; Springer: Vienna, Austria, 1998. [Google Scholar]
- Du Toit, R.M. The transmission of bluetongue and horse sickness by Culicoides. Onderstepoort J. Vet. Sci. Anim. Ind. 1944, 19, 7–16. [Google Scholar]
- Mellor, P.S.; Boorman, J.; Jennings, M. The multiplication of African horse-sickness virus in two species ofCulicoides (Diptera, Ceratopogonidae). Arch. Virol. 1975, 47, 351–356. [Google Scholar] [CrossRef] [PubMed]
- Boorman, J.; Mellor, P.S.; Penn, M.; Jennings, M. The growth of African horse-sickness virus in embryonated hen eggs and the transmission of virus byCulicoides variipennis Coquillett (Diptera, Ceratopogonidae). Arch. Virol. 1975, 47, 343–349. [Google Scholar] [CrossRef] [PubMed]
- Carpenter, S.; Mellor, P.S.; Fall, A.G.; Garros, C.; Venter, G.J. African Horse Sickness Virus: History, Transmission, and Current Status. Annu. Rev. Entomol. 2017, 62, 343–358. [Google Scholar] [CrossRef]
- Roy, P. Genetically engineered structure-based vaccine for bluetongue disease. Vet. Ital. 2004, 40, 594–600. [Google Scholar]
- Barnard, B.J.H. Epidemiology of African horse sickness and the role of the zebra in South Africa. In African Horse Sickness; Mellor, P.S., Baylis, M., Hamblin, C., Mertens, P.P.C., Calisher, C.H., Eds.; Springer: Vienna, Austria, 1998; pp. 13–19. [Google Scholar]
- Theiler, A. The susceptibility of the dog to african horse-sickness. J. Comp. Pathol. Ther. 1910, 23, 315–325. [Google Scholar] [CrossRef]
- McIntosh, B.M. Horsesickness antibodies in the sera of dogs in enzootic areas. J. S. Afr. Vet. Assoc. 1955, 26, 269–272. [Google Scholar]
- Oellermann, R.; Els, H.; Erasmus, B. Characterization of African horsesiekness virus. Arch. Fur Die Gesamte Virusforsch. 1970, 29, 163–174. [Google Scholar] [CrossRef]
- Coetzer, J.A.W.; Erasmus, B.J. African Horse Sickness. In Infectious Diseases of Livestock with Special Reference to Southern Africa; Coetzer, J.A.W., Thomson, G.R., Tustin, R.C., Eds.; Oxford Uiversity Press: Oxford, UK, 1994; pp. 460–475. [Google Scholar]
- Grubman, M.J.; Lewis, S.A. Identification and characterization of the structural and nonstructural proteins of African horsesickness virus and determination of the genome coding assignments. Virology 1992, 186, 444–451. [Google Scholar] [CrossRef]
- Bremer, C.W. A gel electrophoretic study of the protein and nucleic acid components of African horsesickness virus. Onderstepoort J. Vet. Res. 1976, 43, 193–199. [Google Scholar] [PubMed]
- Bremer, C.W.; Huismans, H.; Van Dijk, A.A. Characterization and cloning of the African horsesickness virus genome. J. Gen. Virol. 1990, 71, 793–799. [Google Scholar] [CrossRef]
- Roy, P.; Mertens, P.P.; Casal, I. African horse sickness virus structure. Comp. Immunol. Microbiol. Infect. Dis. 1994, 17, 243–273. [Google Scholar] [CrossRef]
- Zhang, X.; Patel, A.; Celma, C.C.; Yu, X.; Roy, P.; Zhou, Z.H. Atomic model of a nonenveloped virus reveals pH sensors for a coordinated process of cell entry. Nat. Struct. Mol. Biol. 2016, 23, 74–80. [Google Scholar] [CrossRef]
- Grimes, J.M.; Burroughs, J.N.; Gouet, P.; Diprose, J.M.; Malby, R.; Zientara, S.; Mertens, P.P.C.; Stuart, D.I. The atomic structure of the bluetongue virus core. Nature 1998, 395, 470–478. [Google Scholar] [CrossRef]
- Zhang, X.; Boyce, M.; Bhattacharya, B.; Zhang, X.; Schein, S.; Roy, P.; Zhou, Z.H. Bluetongue virus coat protein VP2 contains sialic acid-binding domains, and VP5 resembles enveloped virus fusion proteins. Proc. Natl. Acad. Sci. USA 2010, 107, 6292–6297. [Google Scholar] [CrossRef]
- Nason, E.L.; Rothagel, R.; Mukherjee, S.K.; Kar, A.K.; Forzan, M.; Prasad, B.V.; Roy, P. Interactions between the inner and outer capsids of bluetongue virus. J. Virol. 2004, 78, 8059–8067. [Google Scholar] [CrossRef]
- Roy, P. Bluetongue virus structure and assembly. Curr. Opin. Virol. 2017, 24, 115–123. [Google Scholar] [CrossRef]
- Iwata, H.; Yamagaw, M.; Roy, P. Evolutionary relationships among the gnat-transmitted orbiviruses that cause African horse sickness, bluetongue, and epizootic hemorrhagic disease as evidenced by their capsid protein sequences. Virology 1992, 191, 251–261. [Google Scholar] [CrossRef]
- Caspar, D.L.D.; Klug, A. Physical principles in the construction of regular viruses. Cold Spring Harb. Symp. Quant. Biol. 1962, 27, 1–24. [Google Scholar] [CrossRef] [PubMed]
- Stuart, D.I.; Gouet, P.; Grimes, J.; Malby, R.; Diprose, J.; Zientara, S.; Burroughs, J.N.; Mertens, P.P. Structural studies of orbivirus particles. Arch. Virol. Suppl. 1998, 14, 235–250. [Google Scholar] [PubMed]
- Manole, V.; Laurinmaki, P.; Van Wyngaardt, W.; Potgieter, C.A.; Wright, I.M.; Venter, G.J.; van Dijk, A.A.; Sewell, B.T.; Butcher, S.J. Structural insight into African horsesickness virus infection. J. Virol. 2012, 86, 7858–7866. [Google Scholar] [CrossRef] [PubMed]
- Gouet, P.; Diprose, J.M.; Grimes, J.M.; Malby, R.; Burroughs, J.N.; Zientara, S.; Stuart, D.I.; Mertens, P.P. The highly ordered double-stranded RNA genome of bluetongue virus revealed by crystallography. Cell 1999, 97, 481–490. [Google Scholar] [CrossRef]
- Stuart, D.; Grimes, J. Structural Studies on Orbivirus Proteins and Particles. In Reoviruses: Entry, Assembly and Morphogenesis. Current Topics in Microbiology and Immunology; Roy, P., Ed.; Springer: Berlin/Heidelberg, Germany, 2006; pp. 221–244. [Google Scholar]
- Chuma, T.; Le Blois, H.; Sanchez-Vizcaino, J.M.; Diaz-Laviada, M.; Roy, P. Expression of the major core antigen VP7 of African horsesickness virus by a recombinant baculovirus and its use as a group-specific diagnostic reagent. J. Gen. Virol. 1992, 73, 925–931. [Google Scholar] [CrossRef] [PubMed]
- Grimes, J.; Basak, A.K.; Roy, P.; Stuart, D. The crystal structure of bluetongue virus VP7. Nature 1995, 373, 167. [Google Scholar] [CrossRef] [PubMed]
- Basak, A.K.; Gouet, P.; Grimes, J.; Roy, P.; Stuart, D. Crystal structure of the top domain of African horse sickness virus VP7: Comparisons with bluetongue virus VP7. J. Virol. 1996, 70, 3797–3806. [Google Scholar] [PubMed]
- Grimes, J.M.; Jakana, J.; Ghosh, M.; Basak, A.K.; Roy, P.; Chiu, W.; Stuart, D.I.; Prasad, B.V.V. An atomic model of the outer layer of the bluetongue virus core derived from X-ray crystallography and electron cryomicroscopy. Structure 1997, 5, 885–893. [Google Scholar] [CrossRef]
- Roy, P.; Noad, R. Bluetongue Virus Assembly and Morphogenesis. In Reoviruses: Entry, Assembly and Morphogenesis. Current Topics in Microbiology and Immunology; Roy, P., Ed.; Springer: Berlin/Heidelberg, Germany, 2006; pp. 87–116. [Google Scholar]
- Limn, C.-K.; Staeuber, N.; Monastyrskaya, K.; Gouet, P.; Roy, P. Functional dissection of the major structural protein of bluetongue virus: Identification of key residues within VP7 essential for capsid assembly. J. Virol. 2000, 74, 8658–8669. [Google Scholar] [CrossRef] [PubMed]
- Roy, P.; Hirasawa, T.; Fernandez, M.; Blinov, V.M.; Rodrique, S.-V.J.M. The complete sequence of the group-specific antigen, VP7, of African horsesickness disease virus serotype 4 reveals a close relationship to bluetongue virus. J. Gen. Virol. 1991, 72, 1237–1241. [Google Scholar] [CrossRef]
- Burroughs, J.N.; O’Hara, R.S.; Smale, C.J.; Hamblin, C.; Walton, A.; Armstrong, R.; Mertens, P.P. Purification and properties of virus particles, infectious subviral particles, cores and VP7 crystals of African horsesickness virus serotype 9. J. Gen. Virol. 1994, 75, 1849–1857. [Google Scholar] [CrossRef] [PubMed]
- Burrage, T.G.; Trevejo, R.; Stone-Marschat, M.; Laegreid, W.W. Neutralizing epitopes of African horsesickness virus serotype 4 are located on VP2. Virology 1993, 196, 799–803. [Google Scholar] [CrossRef] [PubMed]
- Hewat, E.A.; Booth, T.F.; Roy, P. Structure of bluetongue virus particles by cryoelectron microscopy. J. Struct. Biol. 1992, 109, 61–69. [Google Scholar] [CrossRef]
- Boyce, M.; Celma, C.P.; Roy, P. Bluetongue virus non-structural protein 1 is a positive regulator of viral protein synthesis. Virol. J. 2012, 9, 178. [Google Scholar] [CrossRef] [PubMed]
- Van Staden, V.; Theron, J.; Greyling, B.; Huismans, H.; Nel, L. A comparison of the nucleotide sequences of cognate NS2 genes of three different orbiviruses. Virology 1991, 185, 500–504. [Google Scholar] [CrossRef]
- Patel, A.; Roy, P. The molecular biology of Bluetongue virus replication. Virus Res. 2014, 182, 5–20. [Google Scholar] [CrossRef] [PubMed]
- Quan, M.; Van Vuuren, M.; Howell, P.G.; Groenewald, D.; Guthrie, A.J. Molecular epidemiology of the African horse sickness virus S10 gene. J. Gen. Virol. 2008, 89, 1159–1168. [Google Scholar] [CrossRef] [PubMed]
- Celma, C.C.P.; Roy, P. Interaction of calpactin light chain (S100A10/p11) and a viral NS protein is essential for intracellular trafficking of non-enveloped Bluetongue virus. J. Virol. 2011, 85, 4783–4791. [Google Scholar] [CrossRef] [PubMed]
- Zwart, L.; Potgieter, C.A.; Clift, S.J.; van Staden, V. Characterising Non-Structural Protein NS4 of African Horse Sickness Virus. PLoS ONE 2015, 10, e0124281. [Google Scholar] [CrossRef] [PubMed]
- Ratinier, M.; Caporale, M.; Golder, M.; Franzoni, G.; Allan, K.; Nunes, S.F.; Armezzani, A.; Bayoumy, A.; Rixon, F.; Shaw, A. Identification and characterization of a novel non-structural protein of bluetongue virus. PLoS Pathog. 2011, 7, e1002477. [Google Scholar] [CrossRef] [PubMed]
- Hassan, S.H.; Wirblich, C.; Forzan, M.; Roy, P. Expression and functional characterization of bluetongue virus VP5 protein: Role in cellular permeabilization. J. Virol. 2001, 75, 8356–8367. [Google Scholar] [CrossRef] [PubMed]
- Mecham, J.O.; McHolland, L.E. Measurement of bluetongue virus binding to a mammalian cell surface receptor by an in situ immune fluorescent staining technique. J. Virol. Methods 2010, 165, 112–115. [Google Scholar] [CrossRef] [PubMed]
- Stevens, L.M.; Moffat, K.; Cooke, L.; Nomikou, K.; Mertens, P.P.C.; Jackson, T.; Darpel, K.E. A low-passage insect-cell isolate of bluetongue virus uses a macropinocytosis-like entry pathway to infect natural target cells derived from the bovine host. J. Gen. Virol. 2019, 100, 568–582. [Google Scholar] [CrossRef] [PubMed]
- Eaton, B.T.; Crameri, G.S. The Site of Bluetongue Virus Attachment to Glycophorins from a Number of Animal Erythrocytes. J. Gen. Virol. 1989, 70, 3347–3353. [Google Scholar] [CrossRef] [PubMed]
- Hassan, S.S.; Roy, P. Expression and Functional Characterization of Bluetongue Virus VP2 Protein: Role in Cell Entry. J. Virol. 1999, 73, 9832. [Google Scholar] [PubMed]
- Marchi, P.R.; Rawlings, P.; Burroughs, J.N.; Wellby, M.; Mertens, P.P.C.; Mellor, P.S.; Wade-Evans, A.M. Proteolytic cleavage of VP2, an outer capsid protein of African horse sickness virus, by species-specific serum proteases enhances infectivity in Culicoides. J. Gen. Virol. 1995, 76, 2607–2611. [Google Scholar] [CrossRef] [PubMed]
- Forzan, M.; Marsh, M.; Roy, P. Bluetongue virus entry into cells. J. Virol. 2007, 81, 4819–4827. [Google Scholar] [CrossRef]
- Forzan, M.; Wirblich, C.; Roy, P. A capsid protein of nonenveloped Bluetongue virus exhibits membrane fusion activity. Proc. Natl. Acad. Sci. USA 2004, 101, 2100–2105. [Google Scholar] [CrossRef]
- Boyce, M.; Wehrfritz, J.; Noad, R.; Roy, P. Purified recombinant bluetongue virus VP1 exhibits RNA replicase activity. J. Virol. 2004, 78, 3994–4002. [Google Scholar] [CrossRef]
- Ramadevi, N.; Burroughs, N.J.; Mertens, P.P.C.; Jones, I.M.; Roy, P. Capping and methylation of mRNA by purified recombinant VP4 protein of bluetongue virus. Proc. Natl. Acad. Sci. USA 1998, 95, 13537–13542. [Google Scholar] [CrossRef]
- Kar, A.K.; Bhattacharya, B.; Roy, P. Bluetongue virus RNA binding protein NS2 is a modulator of viral replication and assembly. BMC Mol. Biol. 2007, 8, 4. [Google Scholar] [CrossRef] [PubMed]
- Modrof, J.; Lymperopoulos, K.; Roy, P. Phosphorylation of bluetongue virus nonstructural protein 2 is essential for formation of viral inclusion bodies. J. Virol. 2005, 79, 10023–10031. [Google Scholar] [CrossRef] [PubMed]
- Mohl, B.-P.; Roy, P. Cellular casein kinase 2 and protein phosphatase 2A modulate replication site assembly of bluetongue virus. J. Biol. Chem. 2016, 291, 14566–14574. [Google Scholar] [CrossRef] [PubMed]
- Beaton, A.R.; Rodriguez, J.; Reddy, Y.K.; Roy, P. The membrane trafficking protein calpactin forms a complex with bluetongue virus protein NS3 and mediates virus release. Proc. Natl. Acad. Sci. USA 2002, 99, 13154–13159. [Google Scholar] [CrossRef] [PubMed]
- Celma, C.C.P.; Roy, P. A viral nonstructural protein regulates bluetongue virus trafficking and release. J. Virol. 2009, 83, 6806–6816. [Google Scholar] [CrossRef] [PubMed]
- Venter, E.; Van der Merwe, C.F.; Buys, A.V.; Huismans, H.; Van Staden, V. Comparative ultrastructural characterization of African horse sickness virus-infected mammalian and insect cells reveals a novel potential virus release mechanism from insect cells. J. Gen. Virol. 2014, 95, 642–651. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Mertens, P.P.C.; Burroughs, J.N.; Walton, A.; Wellby, M.P.; Fu, H.; O’Hara, R.S.; Brookes, S.M.; Mellor, P.S. Enhanced infectivity of modified bluetongue virus particles for two insect cell lines and for TwoCulicoidesVector species. Virology 1996, 217, 582–593. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Ruoslahti, E. Rgd and Other Recognition Sequences for Integrins. Annu. Rev. Cell Dev. Biol. 1996, 12, 697–715. [Google Scholar] [CrossRef] [PubMed]
- Burrage, T.G.; Laegreid, W.W. African horsesickness: Pathogenesis and immunity. Comp. Immunol. Microbiol. Infect. Dis. 1994, 17, 275–285. [Google Scholar] [CrossRef]
- Wohlsein, P.; Pohlenz, J.F.; Davidson, F.L.; Salt, J.S.; Hamblin, C. Immunohistochemical demonstration of African horse sickness viral antigen in formalin-fixed equine tissues. Vet. Pathol. 1997, 34, 568–574. [Google Scholar] [CrossRef] [PubMed]
- Erasmus, B. A new Approach to Polyvalent Immunization Against African Horsesickness. In Proceedings of the 4th International Conference on Equine Infectious Diseases, Lyon, France, 24–27 September 1976; Bryans, J.T., Gerber, H., Eds.; Veterinary Publications: Princeton, NJ, USA, 1978; pp. 401–403. [Google Scholar]
- Erasmus, B.J. The Pathogenesis of African Horsesickness. In Equine Infectious Diseases; Karger Publishers: Basel, Switzerland, 1974; pp. 1–11. [Google Scholar]
- Alexander, R.A. Studies on the neurotropic virus of horsesickness III: The intracerebral protection test and its application to the study of immunity. Onderstepoort J. Vet. Sci. Anim. Ind. 1935, 4, 349–377. [Google Scholar]
- Bentley, L.; Fehrsen, J.; Jordaan, F.; Huismans, H.; Du Plessis, D.H. Identification of antigenic regions on VP2 of African horsesickness virus serotype 3 by using phage-displayed epitope libraries. J. Gen. Virol. 2000, 81, 993–1000. [Google Scholar] [CrossRef] [PubMed]
- Mathebula, E.M.; Faber, F.E.; van Wyngaardt, W.; van Schalkwyk, A.; Pretorius, A.; Fehrsen, J. B-cell epitopes of African horse sickness virus serotype 4 recognised by immune horse sera. Onderstepoort J. Vet. Res. 2017, 84, 1–12. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Martinez-Torrecuadrada, J.L.; Langeveld, J.P.; Meloen, R.H.; Casal, J.I. Definition of neutralizing sites on African horse sickness virus serotype 4 VP2 at the level of peptides. J. Gen. Virol. 2001, 82, 2415–2424. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Venter, M.; Napier, G.; Huismans, H. Cloning, sequencing and expression of the gene that encodes the major neutralisation-specific antigen of African horsesickness virus serotype 9. J. Virol. Methods 2000, 86, 41–53. [Google Scholar] [CrossRef]
- De la Poza, F.; Marin-Lopez, A.; Castillo-Olivares, J.; Calvo-Pinilla, E.; Ortego, J. Identification of CD8 T cell epitopes in VP2 and NS1 proteins of African horse sickness virus in IFNAR(-/-) mice. Virus Res. 2015, 210, 149–153. [Google Scholar] [CrossRef] [PubMed]
- El Garch, H.; Crafford, J.E.; Amouyal, P.; Durand, P.Y.; Edlund Toulemonde, C.; Lemaitre, L.; Cozette, V.; Guthrie, A.; Minke, J.M. An African horse sickness virus serotype 4 recombinant canarypox virus vaccine elicits specific cell-mediated immune responses in horses. Vet. Immunol. Immunopathol. 2012, 149, 76–85. [Google Scholar] [CrossRef] [PubMed]
- Pretorius, A.; Van Kleef, M.; Van Wyngaardt, W.; Heath, J. Virus-specific CD8+ T-cells detected in PBMC from horses vaccinated against African horse sickness virus. Vet. Immunol. Immunopathol. 2012, 146, 81–86. [Google Scholar] [CrossRef]
- Weyer, C.T.; Grewar, J.D.; Burger, P.; Rossouw, E.; Lourens, C.; Joone, C.; le Grange, M.; Coetzee, P.; Venter, E.; Martin, D.P.; et al. African Horse Sickness Caused by Genome Reassortment and Reversion to Virulence of Live, Attenuated Vaccine Viruses, South Africa, 2004–2014. Emerg. Infect. Dis. 2016, 22, 2087–2096. [Google Scholar] [CrossRef]
- Weyer, C.T. African Horse Sickness Outbreak Investigation and Disease Surveillance Using Molecular Techniques. Ph.D. Thesis, University of Pretoria, Pretoria, South Africa, 2017. [Google Scholar]
- Sailleau, C.; Hamblin, C.; Paweska, J.; Zientara, S. Identification and differentiation of the nine African horse sickness virus serotypes by RT–PCR amplification of the serotype-specific genome segment 2. J. Gen. Virol. 2000, 81, 831–837. [Google Scholar] [CrossRef]
- Maree, S.; Paweska, J.T. Preparation of recombinant African horse sickness virus VP7 antigen via a simple method and validation of a VP7-based indirect ELISA for the detection of group-specific IgG antibodies in horse sera. J. Virol. Methods 2005, 125, 55–65. [Google Scholar] [CrossRef] [PubMed]
- Weyer, C.T.; Joone, C.; Lourens, C.W.; Monyai, M.S.; Koekemoer, O.; Grewar, J.D.; van Schalkwyk, A.; Majiwa, P.O.; MacLachlan, N.J.; Guthrie, A.J. Development of three triplex real-time reverse transcription PCR assays for the qualitative molecular typing of the nine serotypes of African horse sickness virus. J. Virol. Methods 2015, 223, 69–74. [Google Scholar] [CrossRef] [PubMed]
- Agüero, M.; Gomez-Tejedor, C.; Cubillo, Á.M.; Rubio, C.; Romero, E.; Jiménez-Clavero, M.A. Real-time fluorogenic reverse transcription polymerase chain reaction assay for detection of African horse sickness virus. J. Vet. Diagn. Investig. 2008, 20, 325–328. [Google Scholar] [CrossRef] [PubMed]
- Guthrie, A.J.; MacLachlan, N.J.; Joone, C.; Lourens, C.W.; Weyer, C.T.; Quan, M.; Monyai, M.S.; Gardner, I.A. Diagnostic accuracy of a duplex real-time reverse transcription quantitative PCR assay for detection of African horse sickness virus. J. Virol. Methods 2013, 189, 30–35. [Google Scholar] [CrossRef] [PubMed]
- Quan, M.; Lourens, C.W.; MacLachlan, N.J.; Gardner, I.A.; Guthrie, A.J. Development and optimisation of a duplex real-time reverse transcription quantitative PCR assay targeting the VP7 and NS2 genes of African horse sickness virus. J. Virol. Methods 2010, 167, 45–52. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez-Sanchez, B.; Fernandez-Pinero, J.; Sailleau, C.; Zientara, S.; Belak, S.; Arias, M.; Sanchez-Vizcaino, J.M. Novel gel-based and real-time PCR assays for the improved detection of African horse sickness virus. J. Virol. Methods 2008, 151, 87–94. [Google Scholar] [CrossRef] [PubMed]
- OIE; Word Organisation for Animal Health. Manual of Diagnostic Tests and Vaccines for Terrestrial Animals, 8th ed.; World Organisation for Animal Health: Paris, France, 2018. [Google Scholar]
- Koekemoer, J.J.O. Serotype-specific detection of African horsesickness virus by real-time PCR and the influence of genetic variations. J. Virol. Methods 2008, 154, 104–110. [Google Scholar] [CrossRef] [PubMed]
- Bachanek-Bankowska, K.; Maan, S.; Castillo-Olivares, J.; Manning, N.M.; Maan, N.S.; Potgieter, A.C.; Di Nardo, A.; Sutton, G.; Batten, C.; Mertens, P.P. Real time RT-PCR assays for detection and typing of African horse sickness virus. PLoS ONE 2014, 9, e93758. [Google Scholar] [CrossRef] [PubMed]
- Meiswinkel, R.; Baylis, M.; Labuschagne, K. Stabling and the protection of horses from Culicoides bolitinos (Diptera: Ceratopogonidae), a recently identified vector of African horse sickness. Bull. Entomol. Res. 2000, 90, 509–515. [Google Scholar] [CrossRef]
- Alexander, R.A. The immunization of horses and mules against Horse Sickness by means of the neurotropic virus of mice and guinea-pigs. Onderstepoort J. Vet. Sci Anim Ind 1934, 2, 375–391. [Google Scholar]
- Erasmus, B.J. Cultivation of horsesickness virus in tissue culture. Nature 1963, 200, 716. [Google Scholar] [CrossRef] [PubMed]
- Von Teichman, B.F.; Smit, T.K. Evaluation of the pathogenicity of African Horsesickness (AHS) isolates in vaccinated animals. Vaccine 2008, 26, 5014–5021. [Google Scholar] [CrossRef] [PubMed]
- Von Teichman, B.F.; Dungu, B.; Smit, T.K. In vivo cross-protection to African horse sickness Serotypes 5 and 9 after vaccination with Serotypes 8 and 6. Vaccine 2010, 28, 6505–6517. [Google Scholar] [CrossRef] [PubMed]
- Molini, U.; Marucchella, G.; Maseke, A.; Ronchi, G.F.; Di Ventura, M.; Salini, R.; Scacchia, M.; Pini, A. Immunization of horses with a polyvalent live-attenuated African horse sickness vaccine: Serological response and disease occurrence under field conditions. Trials Vaccinol. 2015, 4, 24–28. [Google Scholar] [CrossRef]
- Weyer, C.T.; Grewar, J.D.; Burger, P.; Joone, C.; Lourens, C.; MacLachlan, N.J.; Guthrie, A.J. Dynamics of African horse sickness virus nucleic acid and antibody in horses following immunization with a commercial polyvalent live attenuated vaccine. Vaccine 2017, 35, 2504–2510. [Google Scholar] [CrossRef] [PubMed]
- Mirchamsy, H.; Taslimi, H. Inactivated African horse sickness virus cell culture vaccine. Immunology 1968, 14, 81. [Google Scholar] [PubMed]
- Weyer, C.T.; Quan, M.; Joone, C.; Lourens, C.W.; MacLachlan, N.J.; Guthrie, A.J. African horse sickness in naturally infected, immunised horses. Equine Vet. J. 2013, 45, 117–119. [Google Scholar] [CrossRef]
- Lelli, R.; Molini, U.; Ronchi, G.F.; Rossi, E.; Franchi, P.; Ulisse, S.; Armillotta, G.; Capista, S.; Khaiseb, S.; Di Ventura, M. Inactivated and adjuvanted vaccine for the control of the African horse sickness virus serotype 9 infection: Evaluation of efficacy in horses and guinea-pig model. Vet. Ital. 2013, 49, 89–98. [Google Scholar]
- Erasmus, B.J. Preliminary observations on the value of the guinea-pig in determining the innocuity and antigenicity of neurotropic attenuated horsesickness strains. Onderstepoort J. Vet. Res. 1963, 30, 11–22. [Google Scholar]
- Romito, M.; Du Plessis, D.H.; Viljoen, G.J. Immune responses in a horse inoculated with the VP2 gene of African horsesickness virus. Onderstepoort J. Vet. Res. 1999, 66, 139–144. [Google Scholar]
- Roy, P.; Bishop, D.H.; Howard, S.; Aitchison, H.; Erasmus, B. Recombinant baculovirus-synthesized African horsesickness virus (AHSV) outer-capsid protein VP2 provides protection against virulent AHSV challenge. J. Gen. Virol. 1996, 77, 2053–2057. [Google Scholar] [CrossRef] [PubMed]
- Martinez-Torrecuadrada, J.L.; Diaz-Laviada, M.; Roy, P.; Sanchez, C.; Vela, C.; Sanchez-Vizcaino, J.M.; Casal, J.I. Full protection against African horsesickness (AHS) in horses induced by baculovirus-derived AHS virus serotype 4 VP2, VP5 and VP7. J. Gen. Virol. 1996, 77, 1211–1221. [Google Scholar] [CrossRef] [PubMed]
- Kanai, Y.; van Rijn, P.A.; Maris-Veldhuis, M.; Kaname, Y.; Athmaram, T.N.; Roy, P. Immunogenicity of recombinant VP2 proteins of all nine serotypes of African horse sickness virus. Vaccine 2014, 32, 4932–4937. [Google Scholar] [CrossRef] [PubMed]
- Du Plessis, M.; Cloete, M.; Aitchison, H.; Van Dijk, A.A. Protein aggregation complicates the development of baculovirus-expressed African horsesickness virus serotype 5 VP2 subunit vaccines. Onderstepoort J. Vet. Res. 1998, 65, 321–329. [Google Scholar] [PubMed]
- Scanlen, M.; Paweska, J.T.; Verschoor, J.A.; van Dijk, A.A. The protective efficacy of a recombinant VP2-based African horsesickness subunit vaccine candidate is determined by adjuvant. Vaccine 2002, 20, 1079–1088. [Google Scholar] [CrossRef]
- Guthrie, A.J.; Quan, M.; Lourens, C.W.; Audonnet, J.C.; Minke, J.M.; Yao, J.; He, L.; Nordgren, R.; Gardner, I.A.; Maclachlan, N.J. Protective immunization of horses with a recombinant canarypox virus vectored vaccine co-expressing genes encoding the outer capsid proteins of African horse sickness virus. Vaccine 2009, 27, 4434–4438. [Google Scholar] [CrossRef] [PubMed]
- Calvo-Pinilla, E.; Casanova, I.; Bachanek-Bankowska, K.; Chiam, R.; Maan, S.; Nieto, J.M.; Ortego, J.; Mertens, P.P.C. A modified vaccinia Ankara virus (MVA) vaccine expressing African horse sickness virus (AHSV) VP2 protects against AHSV challenge in an IFNAR -/- mouse model. PLoS ONE 2011, 6, e16503. [Google Scholar]
- Manning, N.M.; Bachanek-Bankowska, K.; Mertens, P.P.C.; Castillo-Olivares, J. Vaccination with recombinant Modified Vaccinia Ankara (MVA) viruses expressing single African horse sickness virus VP2 antigens induced cross-reactive virus neutralising antibodies (VNAb) in horses when administered in combination. Vaccine 2017, 35, 6024–6029. [Google Scholar] [CrossRef]
- Alberca, B.; Bachanek-Bankowska, K.; Cabana, M.; Calvo-Pinilla, E.; Viaplana, E.; Frost, L.; Gubbins, S.; Urniza, A.; Mertens, P.; Castillo-Olivares, J. Vaccination of horses with a recombinant modified vaccinia Ankara virus (MVA) expressing African horse sickness (AHS) virus major capsid protein VP2 provides complete clinical protection against challenge. Vaccine 2014, 32, 3670–3674. [Google Scholar] [CrossRef]
- Chiam, R.; Sharp, E.; Maan, S.; Rao, S.; Mertens, P.; Blacklaws, B.; Davis-Poynter, N.; Wood, J.; Castillo-Olivares, J. Induction of antibody responses to African horse sickness virus (AHSV) in ponies after vaccination with recombinant modified vaccinia Ankara (MVA). PLoS ONE 2009, 4, e5997. [Google Scholar] [CrossRef]
- Calvo-Pinilla, E.; Gubbins, S.; Mertens, P.; Ortego, J.; Castillo-Olivares, J. The immunogenicity of recombinant vaccines based on modified Vaccinia Ankara (MVA) viruses expressing African horse sickness virus VP2 antigens depends on the levels of expressed VP2 protein delivered to the host. Antivir. Res. 2018, 154, 132–139. [Google Scholar] [CrossRef] [PubMed]
- Calvo-Pinilla, E.; de la Poza, F.; Gubbins, S.; Mertens, P.P.; Ortego, J.; Castillo-Olivares, J. Vaccination of mice with a modified Vaccinia Ankara (MVA) virus expressing the African horse sickness virus (AHSV) capsid protein VP2 induces virus neutralising antibodies that confer protection against AHSV upon passive immunisation. Virus Res. 2014, 180, 23–30. [Google Scholar] [CrossRef] [PubMed]
- Gilbert, S.C. Clinical development of Modified Vaccinia virus Ankara vaccines. Vaccine 2013, 31, 4241–4246. [Google Scholar] [CrossRef] [PubMed]
- Cottingham, M.G.; Carroll, M.W. Recombinant MVA vaccines: Dispelling the myths. Vaccine 2013, 31, 4247–4251. [Google Scholar] [CrossRef] [PubMed]
- Calvo-Pinilla, E.; de la Poza, F.; Gubbins, S.; Mertens, P.P.; Ortego, J.; Castillo-Olivares, J. Antiserum from mice vaccinated with modified vaccinia Ankara virus expressing African horse sickness virus (AHSV) VP2 provides protection when it is administered 48h before, or 48h after challenge. Antivir. Res. 2015, 116, 27–33. [Google Scholar] [CrossRef] [PubMed]
- Lulla, V.; Lulla, A.; Wernike, K.; Aebischer, A.; Beer, M.; Roy, P. Assembly of Replication-Incompetent African Horse Sickness Virus Particles: Rational Design of Vaccines for All Serotypes. J. Virol. 2016, 90, 7405–7414. [Google Scholar] [CrossRef] [PubMed]
- Vermaak, E.; Paterson, D.J.; Conradie, A.; Theron, J. Directed genetic modification of African horse sickness virus by reverse genetics. S. Afr. J. Sci. 2015, 111, 1–8. [Google Scholar] [CrossRef][Green Version]
- Van de Water, S.G.; van Gennip, R.G.; Potgieter, C.A.; Wright, I.M.; van Rijn, P.A. VP2 Exchange and NS3/NS3a Deletion in African Horse Sickness Virus (AHSV) in Development of Disabled Infectious Single Animal Vaccine Candidates for AHSV. J. Virol. 2015, 89, 8764–8772. [Google Scholar] [CrossRef]
- Kaname, Y.; Celma, C.C.; Kanai, Y.; Roy, P. Recovery of African horse sickness virus from synthetic RNA. J. Gen. Virol. 2013, 94, 2259–2265. [Google Scholar] [CrossRef]
- Lulla, V.; Losada, A.; Lecollinet, S.; Kerviel, A.; Lilin, T.; Sailleau, C.; Beck, C.; Zientara, S.; Roy, P. Protective efficacy of multivalent replication-abortive vaccine strains in horses against African horse sickness virus challenge. Vaccine 2017, 35, 4262–4269. [Google Scholar] [CrossRef]
- Van Rijn, P.A.; Maris-Veldhuis, M.A.; Potgieter, C.A.; van Gennip, R.G.P. African horse sickness virus (AHSV) with a deletion of 77 amino acids in NS3/NS3a protein is not virulent and a safe promising AHS Disabled Infectious Single Animal (DISA) vaccine platform. Vaccine 2018, 36, 1925–1933. [Google Scholar] [CrossRef] [PubMed]
- Boyce, M.; Celma, C.C.P.; Roy, P. Development of reverse genetics systems for bluetongue virus: Recovery of infectious virus from synthetic RNA transcripts. J. Virol. 2008, 82, 8339–8348. [Google Scholar] [CrossRef] [PubMed]
- Van Rijn, P.A.; van de Water, S.G.; Feenstra, F.; van Gennip, R.G. Requirements and comparative analysis of reverse genetics for bluetongue virus (BTV) and African horse sickness virus (AHSV). Virol. J. 2016, 13, 119. [Google Scholar] [CrossRef] [PubMed]
- Feenstra, F.; Pap, J.S.; van Rijn, P.A. Application of bluetongue Disabled Infectious Single Animal (DISA) vaccine for different serotypes by VP2 exchange or incorporation of chimeric VP2. Vaccine 2015, 33, 812–818. [Google Scholar] [CrossRef] [PubMed]
- Conradie, A.M.; Stassen, L.; Huismans, H.; Potgieter, C.A.; Theron, J. Establishment of different plasmid only-based reverse genetics systems for the recovery of African horse sickness virus. Virology 2016, 499, 144–155. [Google Scholar] [CrossRef] [PubMed]
- Matsuo, E.; Celma, C.C.; Roy, P. A reverse genetics system of African horse sickness virus reveals existence of primary replication. FEBS Lett. 2010, 584, 3386–3391. [Google Scholar] [CrossRef] [PubMed]
- Celma, C.C.; Stewart, M.; Wernike, K.; Eschbaumer, M.; Gonzalez-Molleda, L.; Breard, E.; Schulz, C.; Hoffmann, B.; Haegeman, A.; De Clercq, K. Replication-deficient particles: New insights into the next generation of bluetongue virus vaccines. J. Virol. 2016, 91, e01892-16. [Google Scholar] [CrossRef]
- Noad, R.; Roy, P. Virus-like particles as immunogens. Trends Microbiol. 2003, 11, 438–444. [Google Scholar] [CrossRef]
- Bachmann, M.F.; Rohrer, U.H.; Kundig, T.M.; Burki, K.; Hengartner, H.; Zinkernagel, R.M. The influence of antigen organization on B cell responsiveness. Science 1993, 262, 1448. [Google Scholar] [CrossRef]
- Lechner, F.; Jegerlehner, A.; Tissot, A.C.; Maurer, P.; Sebbel, P.; Renner, W.A.; Jennings, G.T.; Bachmann, M.F. Virus-Like Particles as a Modular System for Novel Vaccines. Intervirology 2002, 45, 212–217. [Google Scholar] [CrossRef]
- Lua, L.H.L.; Connors, N.K.; Sainsbury, F.; Chuan, Y.P.; Wibowo, N.; Middelberg, A.P.J. Bioengineering virus-like particles as vaccines. Biotechnol. Bioeng. 2014, 111, 425–440. [Google Scholar] [CrossRef] [PubMed]
- Grgacic, E.V.; Anderson, D.A. Virus-like particles: Passport to immune recognition. Methods 2006, 40, 60–65. [Google Scholar] [CrossRef] [PubMed]
- Pattenden, L.K.; Middelberg, A.P.J.; Niebert, M.; Lipin, D.I. Towards the preparative and large-scale precision manufacture of virus-like particles. Trends Biotechnol. 2005, 23, 523–529. [Google Scholar] [CrossRef] [PubMed]
- Roy, P.; French, T.; Erasmus, B. Protective efficacy of virus-like particles for bluetongue disease. Vaccine 1992, 10, 28–32. [Google Scholar] [CrossRef]
- Roy, P.; Bishop, D.H.L.; LeBlois, H.; Erasmus, B.J. Long-lasting protection of sheep against bluetongue challenge after vaccination with virus-like particles: Evidence for homologous and partial heterologous protection. Vaccine 1994, 12, 805–811. [Google Scholar] [CrossRef]
- Stewart, M.; Bhatia, Y.; Athmaran, T.N.; Noad, R.; Gastaldi, C.; Dubois, E.; Russo, P.; Thiéry, R.; Sailleau, C.; Bréard, E.; et al. Validation of a novel approach for the rapid production of immunogenic virus-like particles for bluetongue virus. Vaccine 2010, 28, 3047–3054. [Google Scholar] [CrossRef]
- Lomonossoff, G.P.; D’Aoust, M.-A. Plant-produced biopharmaceuticals: A case of technical developments driving clinical deployment. Science 2016, 353, 1237–1240. [Google Scholar] [CrossRef]
- Rybicki, E.P. Plant-made vaccines for humans and animals. Plant. Biotechnol. J. 2010, 8, 620–637. [Google Scholar] [CrossRef]
- Fischer, R.; Schillberg, S.; Buyel, J.F.; Twyman, R.M. Commercial aspects of pharmaceutical protein production in plants. Curr. Pharm. Des. 2013, 19, 5471–5477. [Google Scholar] [CrossRef]
- Ma, J.K.C.; Drake, P.M.W.; Christou, P. Genetic modification: The production of recombinant pharmaceutical proteins in plants. Nat. Rev. Genet. 2003, 4, 794. [Google Scholar] [CrossRef]
- Rybicki, E. History and Promise of Plant-Made Vaccines for Animals. In Prospects of Plant-Based Vaccines in Veterinary Medicine; MacDonald, J., Ed.; Springer International Publishing: Cham, Germany, 2018; pp. 1–22. [Google Scholar]
- Rybicki, E.P. Plant-based vaccines against viruses. Virol. J. 2014, 11, 205. [Google Scholar] [CrossRef] [PubMed]
- Thuenemann, E.C.; Meyers, A.E.; Verwey, J.; Rybicki, E.P.; Lomonossoff, G.P. A method for rapid production of heteromultimeric protein complexes in plants: Assembly of protective bluetongue virus-like particles. Plant Biotechnol. J. 2013, 11, 839–846. [Google Scholar] [CrossRef] [PubMed]
- Van Zyl, A.R.; Meyers, A.E.; Rybicki, E.P. Transient Bluetongue virus serotype 8 capsid protein expression in Nicotiana benthamiana. Biotechnol. Rep. 2016, 9, 15–24. [Google Scholar] [CrossRef] [PubMed]
- Maree, S.; Durbach, S.; Huismans, H. Intracellular production of African horsesickness virus core-like particles by expression of the two major core proteins, VP3 and VP7, in insect cells. J. Gen. Virol. 1998, 79, 333–337. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Maree, S.; Maree, F.F.; Putterill, J.F.; de Beer, T.A.; Huismans, H.; Theron, J. Synthesis of empty african horse sickness virus particles. Virus Res. 2016, 213, 184–194. [Google Scholar] [CrossRef] [PubMed]
- Dennis, S.J.; Meyers, A.E.; Guthrie, A.J.; Hitzeroth, I.I.; Rybicki, E.P. Immunogenicity of plant-produced African horse sickness virus-like particles: Implications for a novel vaccine. Plant Biotechnol. J. 2018, 16, 442–450. [Google Scholar] [CrossRef] [PubMed]
- Dennis, S.J.; O’Kennedy, M.M.; Rutkowska, D.; Tsekoa, T.; Lourens, C.W.; Hitzeroth, I.I.; Meyers, A.E.; Rybicki, E.P. Safety and immunogenicity of plant-produced African horse sickness virus-like particles in horses. Vet. Res. 2018, 49, 105. [Google Scholar] [CrossRef]
- MacLachlan, N.J.; Balasuriya, U.B.; Davis, N.L.; Collier, M.; Johnston, R.E.; Ferraro, G.L.; Guthrie, A.J. Experiences with new generation vaccines against equine viral arteritis, West Nile disease and African horse sickness. Vaccine 2007, 25, 5577–5582. [Google Scholar] [CrossRef]
- House, J.A. Recommendations for African horse sickness vaccines for use in nonendemic areas. Rev. D’élevage Et De Médecine Vétérinaire Des. Pays Trop. 1993, 46, 77–81. [Google Scholar]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dennis, S.J.; Meyers, A.E.; Hitzeroth, I.I.; Rybicki, E.P. African Horse Sickness: A Review of Current Understanding and Vaccine Development. Viruses 2019, 11, 844. https://doi.org/10.3390/v11090844
Dennis SJ, Meyers AE, Hitzeroth II, Rybicki EP. African Horse Sickness: A Review of Current Understanding and Vaccine Development. Viruses. 2019; 11(9):844. https://doi.org/10.3390/v11090844
Chicago/Turabian StyleDennis, Susan J, Ann E Meyers, Inga I Hitzeroth, and Edward P Rybicki. 2019. "African Horse Sickness: A Review of Current Understanding and Vaccine Development" Viruses 11, no. 9: 844. https://doi.org/10.3390/v11090844
APA StyleDennis, S. J., Meyers, A. E., Hitzeroth, I. I., & Rybicki, E. P. (2019). African Horse Sickness: A Review of Current Understanding and Vaccine Development. Viruses, 11(9), 844. https://doi.org/10.3390/v11090844





