Isolation and Full-Length Sequence Analysis of a Pestivirus from Aborted Lamb Fetuses in Italy
Abstract
1. Introduction
2. Materials and Methods
2.1. Clinical Cases and Sampling
2.2. Laboratory Investigation
2.3. Molecular Analyses
3. Results
3.1. Clinical Cases
3.2. Virus Isolation and Identification
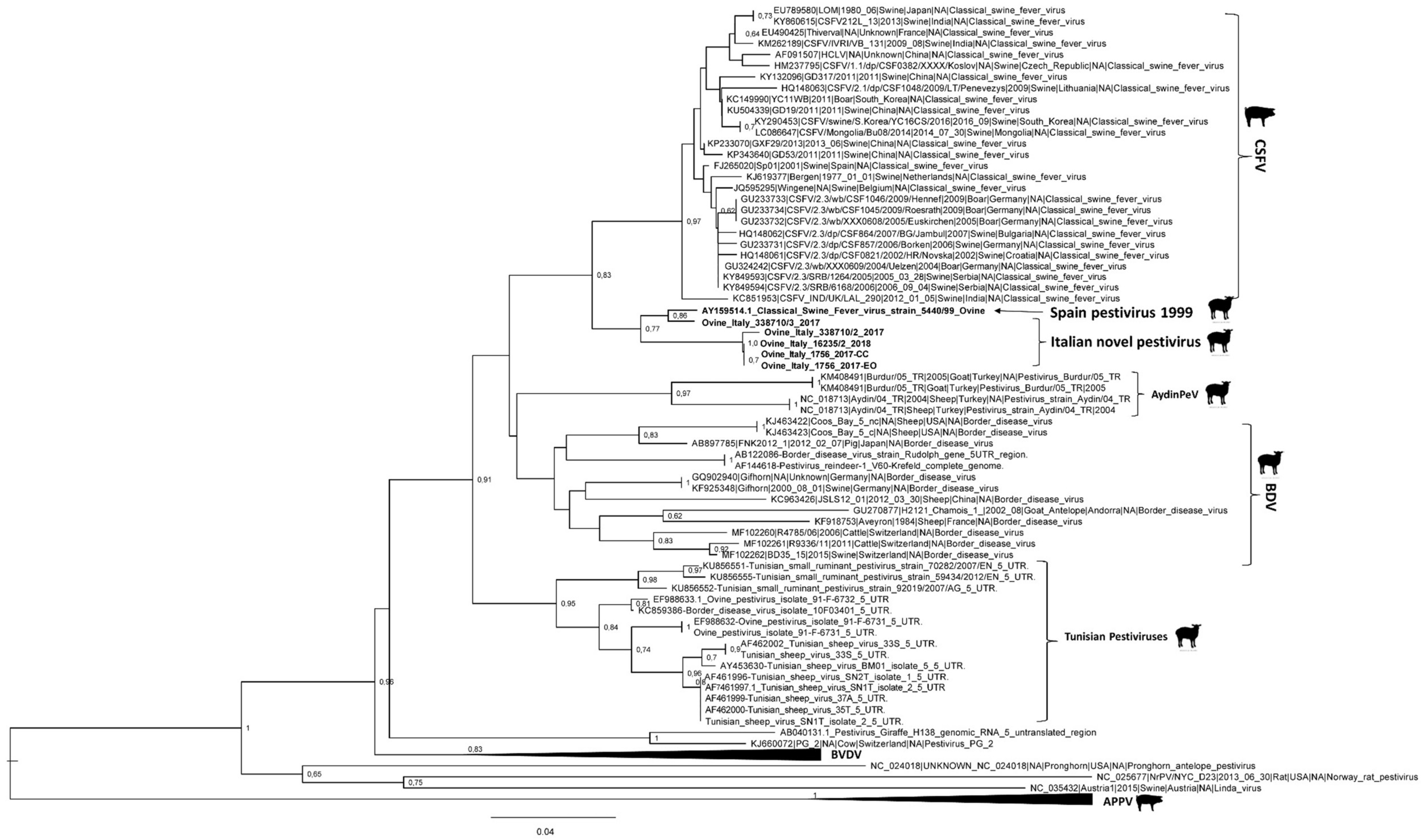
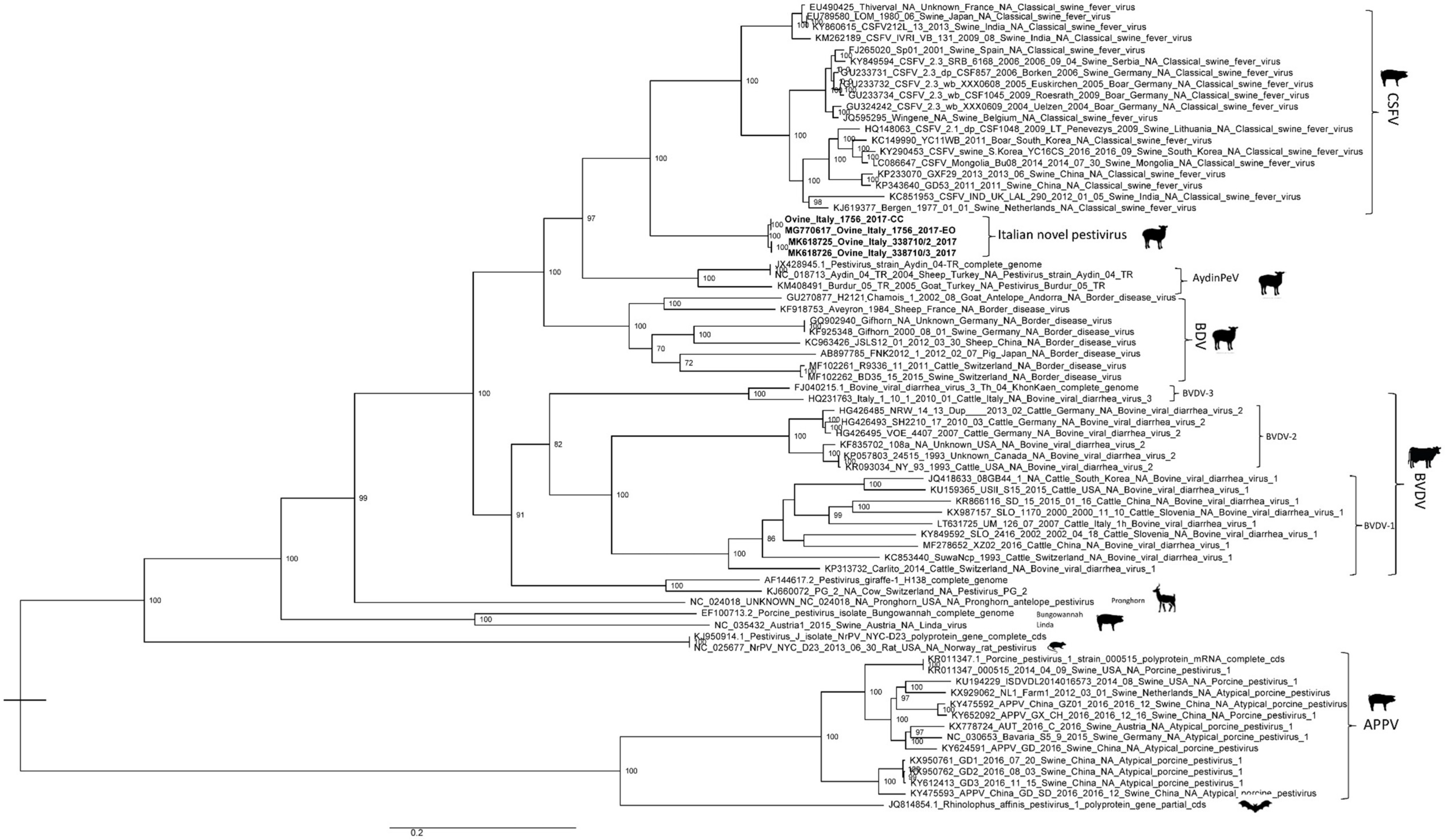
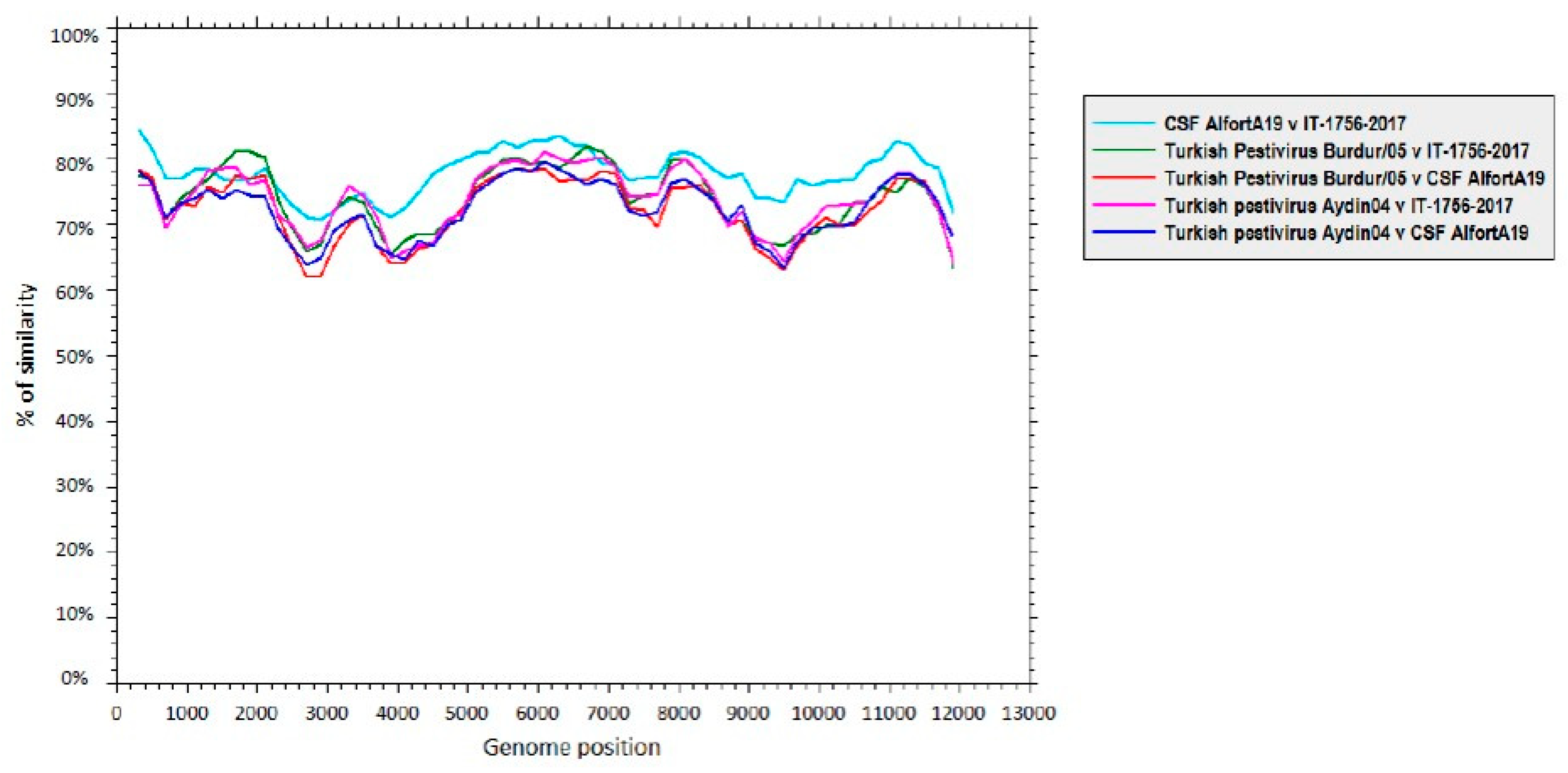
3.3. Genome Characterization
4. Discussion
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Smith, D.B.; Meyers, G.; Bukh, J.; Gould, E.A.; Monath, T.; Scott Muerhoff, A.; Pletnev, A.; Rico-Hesse, R.; Stapleton, J.T.; Simmonds, P.; et al. Proposed revision to the taxonomy of the genus Pestivirus, family Flaviviridae. J. Gen. Virol. 2017, 98, 2106–2112. [Google Scholar] [CrossRef]
- Becher, P.; Orlich, M.; Shannon, A.D.; Horner, G.; Konig, M.; Thiel, H. –J. Phylogenetic analysis of pestiviruses from domestic and wild ruminants. J. Gen. Virol. 1997, 78, 1357–1366. [Google Scholar] [CrossRef]
- Terpstra, C.; Wensvoort, G. Natural infections of pigs with bovine viral diarrhoea virus associated with signs resembling swine fever. Res. Vet. Sci. 1998, 45, 137–142. [Google Scholar] [CrossRef]
- Vilcek, S.; Belak, S. Genetic identification of pestivirus strain Frijters as a border disease virus from pigs. J. Virol. Methods 1996, 60, 103–108. [Google Scholar] [CrossRef]
- Vilcek, S.; Nettleton, P.F.; Paton, D.J.; Belak, S. Molecular characterization of ovine pestiviruses. J. Gen. Virol. 1997, 78, 725–735. [Google Scholar] [CrossRef]
- Paton, D.J. Pestivirus diversity. J. Comp. Pathol. 1995, 112, 215–236. [Google Scholar] [CrossRef]
- OIE Manual for Terrestrial Animals 2018 cap 3.4.7 par B1.2: Bovine viral diarrhoea – Diagnostic techniques – Detection of the agent – Nucleic acid detection. Available online: https://www.oie.int/fileadmin/Home/eng/Health_standards/tahm/3.04.07_BVD.pdf (accessed on 24 July 2019).
- OIE Manual for Terrestrial Animals 2016 cap 2.1.4 par B1, B1.1, B1.2, B1.3: Brucellosis [Brucella abortus, B. melitensis, B. suis (Infection with B. abortus, B. melitensis, B. suis)] – Diagnostic techniques – Identification of the agent – Staining methods – Collection of samples and culture – Identification and typing. Available online: https://www.oie.int/fileadmin/Home/eng/Health_standards/tahm/3.01.04_BRUCELLOSIS.pdf (accessed on 24 July 2019).
- Berri, M.; Arricau-Bouvery, N.; Rodolakis, A. PCR-based detection of Coxiella burnetii from clinical samples. Methods Mol. Biol. 2003, 216, 53–161. [Google Scholar]
- Ehricht, R.; Slickers, P.; Goellner, S.; Hotzel, H.; Sachse, K. Optimized DNA microarray assay allows detection and genotyping of single PCR-amplifiable target copies. Mol. Cell Probes 2006, 20, 60–63. [Google Scholar] [CrossRef]
- Vicari, N.; Santoni, R.; Vigo, P.G.; Magnino, S. A PCR-RFLP assay targeting the 16S ribosomal gene for the diagnosis of animal chlamydioses. In Proceedings of the 5th Meeting of the European Society for Chlamydia Research (Ed.: Judith Deak), Budapest, Hungary, 1–4 September 2004; 2004. [Google Scholar]
- Van Kuppeveld, F.J.; van der Logt, J.T.; Angulo, A.F.; van Zoest, M.J.; Quint, W.G.; Niesters, H.G.; Galama, J.M.; Melchers, W.J. Genus- and species-specific identification of mycoplasmas by 16S rRNA amplification. Appl. Environ. Microbiol. 1992, 58, 2606–2615, Erratum in: Appl. Environ. Microbiol. 1993, 59, 655. [Google Scholar]
- Müller, N.; Zimmermann, V.; Hentrich, B.; Gottstein, B. Diagnosis of Neospora caninum and Toxoplasma gondii infection by PCR and DNA hybridization immunoassay. J. Clin. Microbiol. 1996, 34, 2850–2852. [Google Scholar]
- Menotti, J.; Garin, Y.J.; Thulliez, P.; Sérugue, M.C.; Stanislawiak, J.; Ribaud, P.; de Castro, N.; Houzé, S.; Derouin, F. Evaluation of a new 5′-nuclease real-time PCR assay targeting the Toxoplasma gondii AF146527 genomic repeat. Clin. Microbiol. Infect. 2010, 16, 363–368. [Google Scholar] [CrossRef]
- Schulz, C.; van der Poel, W.H.M.; Ponsart, C.; Cay, A.B.; Steinbach, F.; Zientara, S.; Beer, M.; Hoffmann, B. European interlaboratory comparison of Schmallenberg virus (SBV) real-time RT-PCR detection in experimental and field samples: the method of extraction is critical for SBV RNA detection in semen. J. Vet. Diagn. Invest. 2015, 27, 422–430. [Google Scholar] [CrossRef]
- Letellier, C.; Kerkhofs, P.; Wellemans, G.; Vanopdenbosch, E. Detection and genotyping of bovine diarrhea virus by reverse transcription-polymerase chain amplification of the 5′ untranslated region. Vet. Microbiol. 1999, 64, 155–167. [Google Scholar] [CrossRef]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6. Molecular Evolutionary Genetics Analysis Version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef]
- Nguyen, L.T.; Schmidt, H.A.; von Haeseler, A.; Minh, B.Q. IQ-TREE: A fast and effective stochastic algorithm for estimating maximum likelihood phylogenies. Mol. Biol. Evol. 2015, 32, 268–274. [Google Scholar] [CrossRef]
- Pickett, B.E.; Sadat, E.L.; Zhang, Y.; Noronha, J.M.; Squires, R.B.; Hunt, V.; Liu, M.; Kumar, S.; Zaremba, S.; Gu, Z.; et al. ViPR: an open bioinformatics database and analysis resource for virology research. Nucleic Acids Res. 2012, 40, D593–D598. [Google Scholar] [CrossRef]
- Kalyaanamoorthy, S.; Minh, B.Q.; Wong, T.K.F.; von Haeseler, A.; Jermiin, L.S. ModelFinder: Fast Model Selection for Accurate Phylogenetic Estimates. Nat. Methods 2017, 14, 587–589. [Google Scholar] [CrossRef]
- Giammarioli, M.; Rossi, E.; Casciari, C.; Bazzucchi, M.; Torresi, C.; De Mia, G.M. Genetic characterization of border disease virus (BDV) isolates from small ruminants in Italy. Virus Genes 2015, 50, 321–324. [Google Scholar] [CrossRef]
- Ciulli, S.; Purpari, G.; Agnello, S.; Di Marco, P.; Di Bella, S.; Volpe, E.; Mira, F.; de AguiarSaldanhaPinheiro, A.C.; Vullo, S.; Guercio, A. Evidence for Tunisian-Like Pestiviruses Presence in Small Ruminants in Italy Since 2007. Transbound Emerg. Dis. 2017, 64, 1243–1253. [Google Scholar] [CrossRef]
- Hurtado, A.; García-Pérez, A.L.; Aduriz, G.; Juste, R.A. Genetic diversity of ruminant pestiviruses from Spain. Virus Res. 2003, 92, 67–73. [Google Scholar] [CrossRef]
- Oguzoglu, T.C. A Review of Border disease virus (BDV)infections in Ruminants: Molecular Characterization, Pathogenesis and Control. Anim. Health Prod. Hyg. 2012, 1, 1–9. [Google Scholar]
- Rios, L.; Coronado, L.; Naranjo-Feliciano, D.; Martínez-Pérez, O.; Perera, C.L.; Hernandez-Alvarez, L.; Díaz de Arce, H.; Núñez, J.I.; Ganges, L.; Pérez, L.J. Deciphering the emergence, genetic diversity and evolution of classical swine fever virus. Sci. Rep. 2017, 7, 17887. [Google Scholar] [CrossRef]
| Sample Id. | Lesions | Laboratory Outcome | ||
|---|---|---|---|---|
| Pestivirus | Chlamydophila abortus | |||
| EO | CC | |||
| 1756/2017 (1+2) pool | (1) mummified (2) hairy fleece with abnormal yellow pigmentation | + | + | negative |
| 338710-1/2017 | hemorrhagic subcutaneous edema, serum hemorrhagic fluid in cavities | − | − | positive |
| 338710-2/2017 | + | + | positive | |
| 338710-3/2017 | + | + | positive | |
| 350462/2017 | no lesions | − | − | negative |
| 350530/2017 | no lesions | − | − | negative |
| 16235-1/2018 | hemorrhagic subcutaneous edema, serum hemorrhagic fluid in cavities | − | − | negative |
| 16235-2/2018 | hairy fleece with abnormal yellow pigmentation | + | − | negative |
| 30564-1/2018 | mummified | − | − | positive |
| 30564-2/2018 | mummified | − | − | positive |
| 37146/2018 | hemorrhagic subcutaneous edema, serum hemorrhagic fluid in cavities liver degeneration | − | − | positive |
| PestivirusSpecies/Strain | 1- | 2- | 3- | 4- | 5- | 6- | 7- | 8- | 9- | 10- | 11- | 12- | 13- | 14- | 15- | 16- | 17- | 18- | 19- | 20- | 21- | 22- | 23- | 24- | 25- | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1. Ovine/Italy/1756/2017CC | XXX | |||||||||||||||||||||||||
| 2. Ovine/Italy/1756/2017EO | 100 | XXX | ||||||||||||||||||||||||
| 3. Ovine/Italy/16235-2/2018 | 99.5 | 99.5 | XXX | |||||||||||||||||||||||
| 4. Ovine/Italy/338710-2/2017 | 99.5 | 99.5 | 99.1 | XXX | ||||||||||||||||||||||
| 5. Ovine/Italy/338710-3/2017 | 94.8 | 94.8 | 95.3 | 94.3 | XXX | |||||||||||||||||||||
| C | 6. AF091661_CSFV_strain_Brescia | 87.6 | 87.6 | 88.1 | 87.0 | 91.2 | XXX | |||||||||||||||||||
| 7. U90951_CSFV_strain_Alfort_A19 | 87.5 | 87.5 | 88.1 | 87.0 | 91.2 | 98.1 | XXX | |||||||||||||||||||
| 8. X87939_CSFV_strain_Alfort/187 | 87.5 | 87.5 | 88.1 | 87.0 | 91.2 | 98.1 | 100 | XXX | ||||||||||||||||||
| 9. FJ265020_CSFV_Sp01_Spain | 88.1 | 88.1 | 88.6 | 87.5 | 90.7 | 95.8 | 95.7 | 95.7 | XXX | |||||||||||||||||
| nc | 10. AY159514.1_Pestivirus_strain_5440/99 | 95.3 | 95.3 | 95.8 | 94.8 | 98.6 | 90.7 | 90.7 | 90.7 | 90.2 | XXX | |||||||||||||||
| nc | 11. AF462000-Tunisian_sheep_virus_35T | 86.4 | 86.4 | 87.0 | 85.8 | 85.9 | 82.0 | 82.5 | 82.5 | 81.5 | 86.4 | XXX | ||||||||||||||
| G | 12. AB040131.1_Pestivirus_Giraffe_H138 | 72.0 | 72.0 | 72.6 | 71.2 | 71.4 | 68.5 | 71.2 | 71.2 | 69.1 | 72.7 | 79.4 | XXX | |||||||||||||
| I | 13. KM408491_Pestivirus_Burdur/05_TR | 83.6 | 83.6 | 84.2 | 84.2 | 80.8 | 80.1 | 80.0 | 80.0 | 82.4 | 81.3 | 80.1 | 71.2 | XXX | ||||||||||||
| 14. NC_018713_Pestivirus_Aydin/04_TR | 81.9 | 81.9 | 82.5 | 82.5 | 80.2 | 81.8 | 80.6 | 80.6 | 82.9 | 82.0 | 81.9 | 76.4 | 92.9 | XXX | ||||||||||||
| A | 15. CS114709_BVDV1_NADL | 71.8 | 71.8 | 71.1 | 71.0 | 71.1 | 71.9 | 71.7 | 71.7 | 69.6 | 70.4 | 69.0 | 63.9 | 74.1 | 73.5 | XXX | ||||||||||
| 16. AF091605_BVDV1_strain_Oregon_C24V | 70.5 | 70.5 | 71.2 | 69.7 | 72.5 | 71.9 | 71.8 | 71.8 | 70.3 | 73.1 | 69.0 | 63.8 | 72.9 | 71.0 | 93.3 | XXX | ||||||||||
| B | 17. L32886_BVDV2_strain_890 | 70.3 | 70.3 | 70.9 | 69.5 | 66.1 | 64.5 | 65.7 | 65.7 | 65.6 | 66.1 | 68.5 | 68.0 | 65.9 | 62.5 | 72.4 | 69.8 | XXX | ||||||||
| 18. AY379547_BVDV2_strain_Giessen_6 | 71.6 | 71.6 | 72.3 | 70.8 | 68.9 | 66.6 | 67.8 | 67.8 | 67.6 | 68.9 | 68.5 | 70.0 | 71.3 | 69.3 | 74.9 | 72.4 | 90.9 | XXX | ||||||||
| D | 19. AB122086-BDV_strain_Rudolph | 79.0 | 79.0 | 79.6 | 78.3 | 79.6 | 82.0 | 81.9 | 81.9 | 81.9 | 79.6 | 78.7 | 70.1 | 83.0 | 81.3 | 67.7 | 65.6 | 61.8 | 64.0 | XXX | ||||||
| 20. U65022_BDV_strain_Moredun_cp | 82.5 | 82.5 | 82.0 | 82.5 | 80.7 | 79.6 | 79.5 | 79.5 | 80.8 | 80.7 | 78.7 | 65.7 | 82.6 | 80.9 | 66.6 | 63.7 | 60.7 | 61.4 | 89.6 | XXX | ||||||
| 21. EF693989_BDV_isolate_90-F-6227 | 77.7 | 77.7 | 77.1 | 77.6 | 75.8 | 80.9 | 80.8 | 80.8 | 81.5 | 75.8 | 74.5 | 68.7 | 81.0 | 79.8 | 68.9 | 66.1 | 65.5 | 63.9 | 88.1 | 91.7 | XXX | |||||
| 22. AY738080_BDV_Chamois | 79.5 | 79.5 | 78.9 | 79.4 | 77.1 | 78.4 | 78.3 | 78.3 | 77.1 | 77.7 | 75.0 | 68.7 | 77.4 | 78.0 | 66.2 | 68.3 | 63.9 | 63.9 | 82.3 | 84.7 | 84.7 | XXX | ||||
| 23. EF693998_BDV_isolate_96-F-7624 | 79.5 | 79.5 | 78.9 | 79.4 | 78.3 | 76.5 | 76.4 | 76.4 | 78.3 | 77.7 | 74.3 | 65.6 | 77.0 | 73.9 | 68.3 | 67.0 | 63.3 | 61.7 | 86.8 | 87.9 | 87.4 | 88.5 | XXX | |||
| 24. EF693996_BDV_isolate_94-F-7446/1 | 80.9 | 80.9 | 80.3 | 80.8 | 81.5 | 80.9 | 80.8 | 80.8 | 80.9 | 80.3 | 80.1 | 70.0 | 78.0 | 78.6 | 70.3 | 68.9 | 65.3 | 63.9 | 89.2 | 91.8 | 90.3 | 89.6 | 90.1 | XXX | ||
| 25. KT072634_BDV_isolate_Italy-103761 | 78.2 | 78.2 | 78.8 | 77.4 | 77.5 | 80.0 | 79.9 | 79.9 | 81.8 | 77.5 | 74.1 | 66.3 | 78.9 | 76.5 | 68.9 | 68.9 | 64.4 | 63.6 | 89.6 | 88.0 | 86.4 | 84.0 | 88.5 | 88.1 | XXX | |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sozzi, E.; Lavazza, A.; Gaffuri, A.; Bencetti, F.C.; Prosperi, A.; Lelli, D.; Chiapponi, C.; Moreno, A. Isolation and Full-Length Sequence Analysis of a Pestivirus from Aborted Lamb Fetuses in Italy. Viruses 2019, 11, 744. https://doi.org/10.3390/v11080744
Sozzi E, Lavazza A, Gaffuri A, Bencetti FC, Prosperi A, Lelli D, Chiapponi C, Moreno A. Isolation and Full-Length Sequence Analysis of a Pestivirus from Aborted Lamb Fetuses in Italy. Viruses. 2019; 11(8):744. https://doi.org/10.3390/v11080744
Chicago/Turabian StyleSozzi, Enrica, Antonio Lavazza, Alessandra Gaffuri, Fabio Carlo Bencetti, Alice Prosperi, Davide Lelli, Chiara Chiapponi, and Ana Moreno. 2019. "Isolation and Full-Length Sequence Analysis of a Pestivirus from Aborted Lamb Fetuses in Italy" Viruses 11, no. 8: 744. https://doi.org/10.3390/v11080744
APA StyleSozzi, E., Lavazza, A., Gaffuri, A., Bencetti, F. C., Prosperi, A., Lelli, D., Chiapponi, C., & Moreno, A. (2019). Isolation and Full-Length Sequence Analysis of a Pestivirus from Aborted Lamb Fetuses in Italy. Viruses, 11(8), 744. https://doi.org/10.3390/v11080744








