Abstract
Viruses have a dual nature: particles are “passive substances” lacking chemical energy transformation, whereas infected cells are “active substances” turning-over energy. How passive viral substances convert to active substances, comprising viral replication and assembly compartments has been of intense interest to virologists, cell and molecular biologists and immunologists. Infection starts with virus entry into a susceptible cell and delivers the viral genome to the replication site. This is a multi-step process, and involves the cytoskeleton and associated motor proteins. Likewise, the egress of progeny virus particles from the replication site to the extracellular space is enhanced by the cytoskeleton and associated motor proteins. This overcomes the limitation of thermal diffusion, and transports virions and virion components, often in association with cellular organelles. This review explores how the analysis of viral trajectories informs about mechanisms of infection. We discuss the methodology enabling researchers to visualize single virions in cells by fluorescence imaging and tracking. Virus visualization and tracking are increasingly enhanced by computational analyses of virus trajectories as well as in silico modeling. Combined approaches reveal previously unrecognized features of virus-infected cells. Using select examples of complementary methodology, we highlight the role of actin filaments and microtubules, and their associated motors in virus infections. In-depth studies of single virion dynamics at high temporal and spatial resolutions thereby provide deep insight into virus infection processes, and are a basis for uncovering underlying mechanisms of how cells function.
Keywords:
Modeling; simulation; computing; quantitative microscopy; fluorescent virions; microscopy; single particle tracking; trajectory segmentation; click chemistry; tracking; trafficking; membrane traffic; fluorescence microscopy; immunofluorescence microscopy; electron microscopy; microtubule; intracellular transport; machine learning; virus infection mechanisms; DNA virus; RNA virus; enveloped virus; nonenveloped virus; cell biology; virus entry; cytoskeleton; infection; receptor; internalization; innate immunity; virion uncoating; endocytosis; gene expression; gene therapy; actin; kinesin; dynein; myosin; nuclear pore complex; adenovirus; herpesvirus; herpes simplex virus; influenza virus; hepatitis B virus; baculovirus; human immunodeficiency virus HIV; parvovirus; adeno-associated virus AAV; simian virus 40 1. Introduction
Viruses affect all forms of life, from bacteria to humans. They are a product of co-evolution with their hosts, and cause disease, or assist in gene and anti-microbial therapies [1,2,3,4]. Virus particles, virions, require the assistance from the host cells to cause an infection, and transfer viral genes into host cells. Infection is a complex subversion process, which gives rise to latent, persistent or lytic outcomes, and cell survival or death [5]. Virions are a container with structural proteins and DNA or RNA genomes inside, sometimes wrapped with a lipid membrane and sugars. Although virions emerge from cells, their water content is several fold lower than that of cells [6]. This implies that they are tightly packaged, and contain entropic pressure [7,8,9]. Virions are considerably smaller than cells, although some of them can reach the size of bacterial cells [10]. Despite their simplicity, virus particles from different families exhibit a large structural diversity, and particles from a single virus type can contain genomes that are variable in sequence but preserve overall function. Viral genomes encode enzymes for virus replication, maturation, genome integration into the host chromosomes, as well as structural and regulatory proteins for building virions and tuning the immune system, proliferation and apoptosis.
Virions deliver their genome into host cells by using receptors, attachment factors and facilitators of the host mediating binding to and activation of cells [11]. Cell signalling, endocytic uptake, endosomal escape and cytoplasmic transport all directly or indirectly depend on the actin or microtubule cytoskeleton [12,13,14,15,16,17,18,19,20,21,22,23,24]. For an overview of virus entry pathways by the cytoskeleton, see Figure 1.
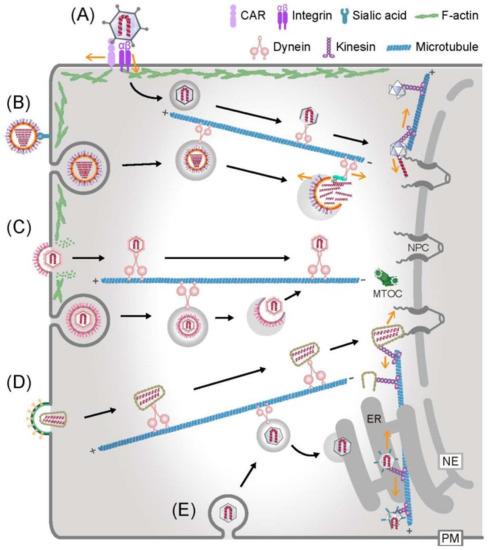
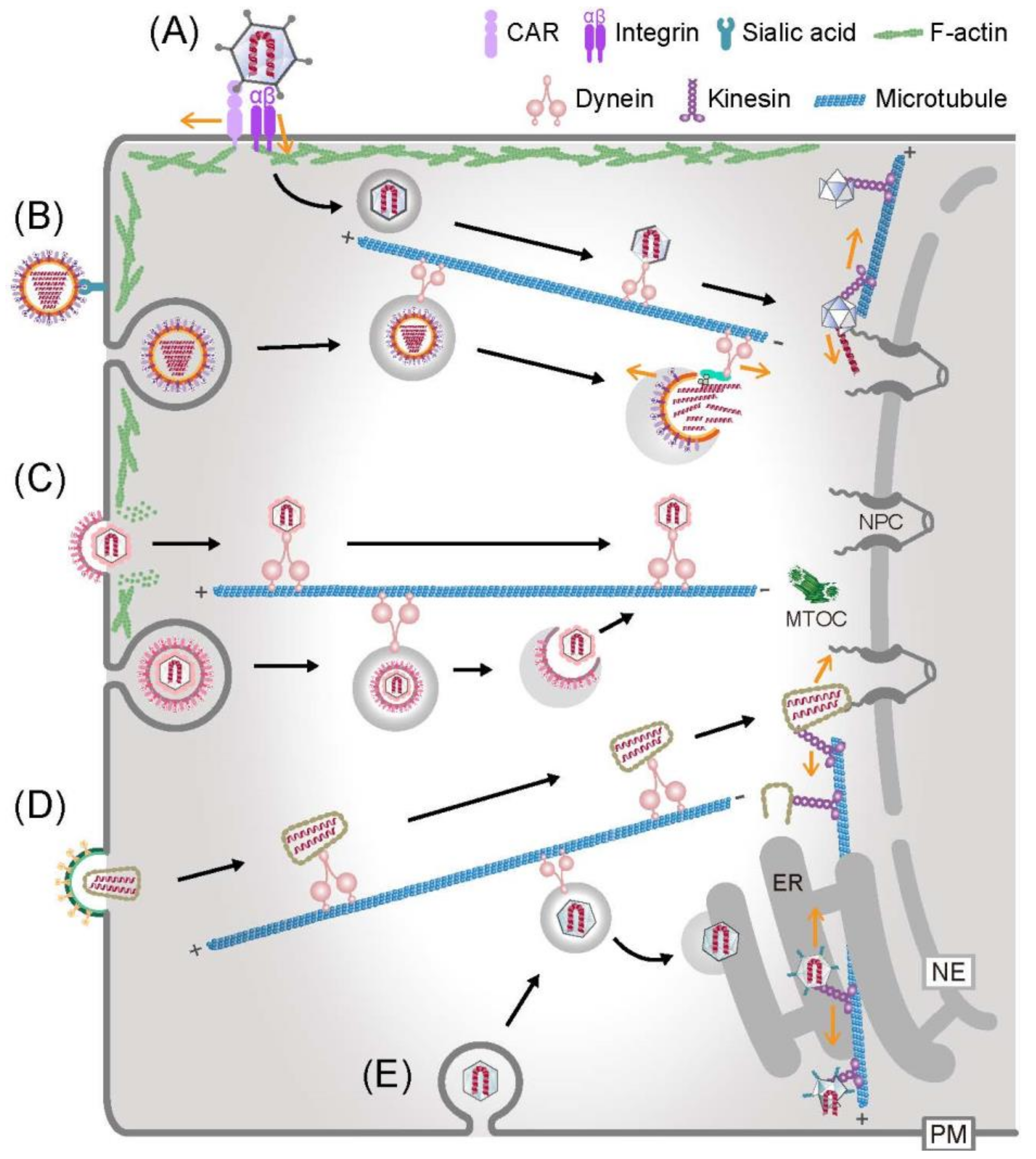
Figure 1.
Examples of virus entry and interactions with the cytoskeleton with a focus on microtubules. Adenovirus (A), influenza virus (B), herpesvirus (C), human immunodeficiency virus (D) and simian virus 40 (E) enter into the cytoplasm either by a direct fusion of viral membrane and host plasma membrane (PM), or by receptor-mediated endocytosis, endosome rupture, or endoplasmic reticulum (ER) membrane penetration. Subsequently, viruses engage with the cytoskeleton and motor proteins to move towards the replication sites. Mechanical forces from the virus–motor protein interactions and opposing forces, such as actin-anchored integrins (A), the nuclear pore complex (NPC) (A), reverse transcription in the viral particle (D) or the site of ER penetration (E) are thought to facilitate virion disruption and release the viral genome (dark yellow arrows).
Before a viral genome is transcribed and replicated, it is at least partially uncoated from the capsid. Genome uncoating requires a series of sequential interactions of the virion with host factors. This concept was initially demonstrated with adenovirus (AdV), a non-enveloped DNA virus, which starts its uncoating program by shedding the fiber proteins at the cell surface, and continues releasing minor virion components in a stepwise manner [25,26,27,28]. For some viruses, such as influenza virus (IV) and AdV, complete genome uncoating requires the acto-myosin and microtubule cytoskeleton [11,29,30]. Other viruses, such as human immunodeficiency virus (HIV) or poxviruses transcribe their genome while located in the cytosol and at least partly wrapped by their capsid [31,32,33,34]. This strategy is thought to provide protection to the viral genome from innate sensors in the cytoplasm [35,36,37,38].
Viruses replicating in the nucleus typically dissociate their genome from the capsid before the genome enters the nucleus, although very small virions, such as adeno-associated viruses are thought to uncoat their genome in the nucleoplasm [39,40,41,42,43,44]. From the site of replication in the cytosol or the nuclear compartment, newly assembled particles then leave the infected cell by mechanisms that unanimously require the assistance of the host cell [45,46,47,48,49]. This is essential because diffusion purely depends on the thermal energy, and does not suffice for effective transport of particles larger than 50 nm across the crowded cytosol [50,51,52,53,54]. In this review we discuss how the imaging of virion dynamics and trafficking in entry and egress can be analyzed by fluorescence microscopy, single particle tracking, and trajectory analyses, and thereby shed new light on cell function in health and disease.
2. In the Extracellular Milieu—Tracking and Modelling of Fluorescent Animal Virus Particles
The classical approach to track the motility of virions has been by fluorescence microscopy of fluorophore-tagged particles [14,52,55,56,57,58,59,60]. A list of fluorescent virus particles from both non-enveloped and enveloped families, and the methods by which the virions were prepared is presented in Table 1. The list also indicates the cytoskeletal elements and the compartments highlighted by virion tracking experiments.

Table 1.
Fluorescent animal viruses.
Imaging studies revealed that before virions attach to cells, the particles diffuse in the extracellular medium. The cell-free medium differs drastically from the intracellular environment in molecular crowdedness [123], ion composition [124], and bulk currents [125,126]. Virus particles in the extracellular medium are subjected to advection due to thermal flow caused by temperature differences, due to the active flow of mucus caused by ciliary motions [127], or the flow of blood or lymph fluids.
Apart from the bulk currents, diffusion represents an important driving force carrying viral particles in the extracellular milieu, and contributes to virus spread in micropopulations of cells [126]. For example, the diffusion constant of particles smaller than 100 nm, such as HPV is largely unaffected by the viscosity in mucus, whereas larger particles, such as herpes simplex virus are strongly affected by mucus [127]. Similar results were reported with label-free virus particles by ultrahigh-speed scattering-based imaging, where vaccinia virus (VV) particles moved on the plasma membrane of host cells with diffusion coefficients in the range of 1 μm2/s [128]. Along the same lines, the diffusion constants (D) of AdV particles (approx. 90 nm in diameter) in extracellular medium are in the range of about 10 µm2/s, as determined by ultrahigh-speed total internal reflection fluorescence (TIRF) microscopy and single-particle tracking of fluorophore-tagged AdV-C2 using spherical fluorescent beads of different diameters as standards [125]. The results are in good agreement with the prediction of the diffusion constant (D) from the Einstein–Stokes equation, where D inversely depends on the diameter of the particle and the viscosity of the medium. Remarkably, numerous approaches have been employed towards biophysical modelling cell-free spread of viruses, including ordinary and partial differential equations (ODE and PDE) [129], agent-based models [130], cellular automata (CA), and not least multi-modal approaches combining two or more modelling methods [125,126,131]. ODE- and PDE-based approaches are among the most simplistic ways to represent dynamic changes of one or multiple parameters in a biophysical system.
3. The Way in and out—Actin-Based Virion Transport
After passing through the extracellular medium, the next step in the infection process is the attachment to a target cell. Eukaryotic cells lacking a cell wall, such as vertebrate cells, are shaped by a layer of actin filaments (F-actin), which also serves to reinforce the plasma membrane and imposes a barrier towards the outside of the cell. The F-actin layer is dynamically regulated by polymerisation and depolymerisation reactions, for example allowing the formation of endocytic pits and vesicles [132]. It provides opportunities for virions to bind to cell extensions, such as filopodia [14,60]. F-actin also provides contractile tracks for myosin motor proteins, and can be arranged into net-like patterns by crosslinking proteins [133]. Upon attachment to the cell surface, virions transmit forward signals into the cell, for example triggering signal transduction pathways akin to growth factors, and prepare the cell for endocytic uptake and infection [13,15,134].
The tracking of fluorophore-tagged single virions in cell culture has revealed that upon initial contacts with attachment factors and receptors on the plasma membrane, virions move outside of the cell in quasi two-dimensional diffusive motions for several seconds until their motion gets confined to small areas of a few hundred nanometers in diameter (reviewed in [14,60]). Virions remain highly mobile by engaging with the cytoskeleton through transmembrane receptors and intracellular adaptors, a process termed “surfing” (reviewed in [14,60]). Notably, fluorophore-tagged retroviruses, such as HIV, murine leukemia virus (MLV) or avian leukosis virus (ALV) [87], murine polyomavirus [107], HPV16 [79] and AdV-C2 [27] were shown to drift along filopodia, a process which is coupled to the myosin-dependent flow of filamentous (F)-actin towards the cell body where endocytic uptake processes occur. Retrograde F-actin flow was originally shown for growth cones of Aplysia neurons [135,136].
Other actin-dependent processes are subverted by viruses during entry, assembly and egress from the infected cell (reviewed in [137,138]). They include actin polymerisation-mediated movement of incoming baculovirus across the cytosol to where the virions eventually reach the cell nucleus [139], the involvement of dynamic F-actin in the entry of Influenza A virus (IAV) into polarized epithelial cells [140], herpes simplex virus (HSV) type 1 entry into cells [141], or a rapid actin-dependent intracellular movement of poliovirus and VV particles [82,116,142,143,144,145]
On the way out, actin-driven viral egress is well described for VV. The cell-associated enveloped particle induces the formation of an actin tail, and moves at an average speed of 2.8 µm/min [143,146]. This propelling force may aid in the transmission of virus particles between cells [144]. Actin-dependent transport of nucleocapsids for virion budding at the plasma membrane has also been reported for baculovirus and filovirus [139,147,148]. At the plasma membrane, F-actin interacts directly or indirectly with the matrix protein of filoviruses to drive the budding of virions, possibly involving myosin motors [149,150].
4. Subcellular Regulation of Microtubule-Dependent Virion Transport on the Way in and out
Microtubules are polarized filaments formed by tubulins and microtubule-associated proteins. In many eukaryotic cell types, the minus-ends are located near the cell center and plus-ends point to the plasma membrane [151]. In addition to maintaining the structure of the cells and providing the framework for cell division, microtubules serve as tracks for the intracellular transport of organelles, proteins, and RNA-protein complexes [152]. Microtubules support long range virion transport at µm/s speed. The role of microtubules in virus entry and egress from infected cells has been extensively reviewed [46,51,52,57,137,138,153,154,155,156,157,158,159]. Microtubules are composed of different isotypes of alpha and beta tubulin, which are subject to a range of post-translational modifications (PTM) [160,161,162]. PTM make up the “tubulin code”. The specific chemistry of the “tubulin code” facilitates the functional diversification, and enables interactions of subsets of microtubules with specific sets of microtubule-associated proteins, including motor proteins.
Increasing evidence indicates that viruses use and modify the tubulin code by signalling to microtubules and affecting microtubule stability and motor protein preference. On the way in, herpesvirus and HIV engage with microtubules and promote microtubule stabilization through a series of protein–protein interactions involving end-binding protein 1 (EB1) and other plus-end tracking proteins (+TIP), such as cytoplasmic linker protein 170 (CLIP-170) and kinesin family member 4 (Kif4) [163,164,165]. AdV induces microtubule growth via the Rac1 signalling pathway, and may therefore increase the chance of the formation of stable microtubules and virion engagement with these tracks [166]. Kaposi’s sarcoma-associated herpesvirus (KSHV) and HIV promote the stabilization of microtubules through the ezrin–radixin–moesin family proteins [167,168].
On the way out, IV, AdV, HIV and human herpesvirus 8 enhance the acetylation and the stability of microtubules [166,167,169,170]. In the case of IV, tubulin acetylation appears to directly promote the transport of progeny viral genomes towards the budding site at the apical plasma membrane. Yet other viruses, such as VV or HSV1, produce viral proteins with MAP-like activities to stabilize microtubules [171,172]. The HSV1 protein Us3 mimics Akt to activate cytoplasmic linker-associated proteins, which are plus-end tracking proteins (+TIP) that lead to the formation of stable microtubules [173]. HIV stabilizes microtubules by engaging the host protein suppressor of cytokine signaling 1 (SOCS1) and viral Gag protein [174]. These examples show that both incoming and outgoing virus particles stabilze microtubule tracks by PTM or dedicated viral proteins to enhance their entry or egress processes.
5. Virion Dismantling by the Cytoskeleton on the Way in
Viruses are too large to efficiently diffuse through the crowded cytoplasm [175]. They evolved mechanisms to take advantage of cellular transport processes, which not only affect their subcellular localization but also exert force on the particles. This has been discovered with AdV, which uses mechanical cues from motile coxsackievirus AdV receptors (CAR) and stationary integrin co-receptors on the plasma membrane of epithelial cells to shed the fibers and open-up the capsid to release the membrane lytic protein VI [9,25,26,27,30,176,177]. Virion motility on the plasma membrane depends on CAR, actin turnover and myosin-2 activity, and works against the holding force of either nonmotile CAR attached to other fibers on the virion or integrins, which are stationary. How the AdV particle exposes its membrane lytic protein in macrophages which lack CAR but provide the entry receptor SR-A6 (MARCO) is currently unknown [178]. Upon virion endocytosis and endosomal escape where the membrane lytic viral protein VI is fully exposed from the virion, the partially dismantled capsids are transported to the nuclear envelope in a microtubule-dependent manner [63,179,180].
Another important question has remained unresolved until recently, namely how any virus particle, which traffics on microtubules can detach from microtubules in order to bind to the nuclear pore complex (NPC) in the nuclear envelope. This question is of key importance for herpes viruses, HIV, parvoviruses, IV viral ribonucleo-protein complexes (vRNP), human foamy virus and AdV, which all deliver their genome into the nucleus of postmitotic cells [52]. Mechanisms of cellular cargo unloading from microtubules vary depending on the types of motors and cargos [181,182]. In the case of AdV, it was recently shown that the nuclear export factor CRM1 has a key role in controlling the virus–microtubule interactions at the juxtanuclear region [183]. Upon the inhibition of CRM1 by leptomycin B, the incoming AdV is constrained on the microtubules, and fails to reach the NPC. If the virions are allowed to detach from microtubules in the juxta-nuclear region, they attach to the NPC via a hexon-Nup214 interaction [183,184,185]. At the NPC, the molecular motor kinesin-1, which is tethered to the virion is activated by Nup358, and then exerts a mechanical force, which breaks open the weakened capsid and releases the viral genome for import into the nucleus through the NPC [29].
The mechanical uncoating concepts proposed for AdV have been adapted for IAV entry [186]. In this scenario, the vRNPs are dissociated from the endosomal membrane by dynein and myosin motors, upon exposure to the cytosol and fusion of the viral membrane with the limiting endosomal membrane. Along a similar line, conventional kinesin has been implicated in membrane penetration of SV40 particles from the ER lumen to the cytosol [187]. Membrane penetration depended on acetylated microtubules in agreement with the earlier notion that kinesin-1 preferably moved on acetylated microtubules [188,189]. In addition, the destabilization of the HIV capsid was reported to depend on microtubules and dynein motor activity [190]. In summary, it becomes increasingly clear that actin filaments, microtubules and associated motors serve as cues for scheduled on-site virus capsid disassembly events, and thereby boost viral infection.
6. Tool Box—Virion Imaging in Cells
Fluorescence microscopy has empowered virologists to determine how virions interact with cell surface attachment factors, receptors, and facilitators for cell entry [11,13,14,56,57,71,105,178,191]. Imaging post-entry steps has shown how the cytoskeleton and its motors support virion trafficking and uncoating, leading to gene delivery and infection [52,138,155,156,157,192,193]. Here we provide an overview of imaging modalities by light microscopy with viruses (Table 2).

Table 2.
Techniques for live cell visualization of viruses and pros and cons in virus tracking experiments. Additional examples of techniques used for virion imaging are listed in Table 5.
Diffraction-limited light microscopy has a spatial resolution in the range of several hundred nanometers, as described by Abbé’s law [220]. Most viruses are between 20–400 nm in size, below the diffraction limit, and appear as point sources in micrographs [221]. However, at high fluorescence signal from the particles and low background one can use the point spread function to determine the position of the particle with much higher precision than the resolution of the microscope. Single particle tracking at high signal to noise ratio is thus possible at low tens of nanometer accuracy, and is providing unprecedented information (see also below). For example, particles that have been derivatized with organic fluorophores or genetically fused to a fluorescent protein, such as GFP could be tracked below the diffraction limit of the light microscope [222]. Alternatively, fluorescence can be induced on the virion by the so-called split-GFP technology, where an engineered particle contains a GFP segment, and binds to a protein of interest with another GFP segment in presence of soluble GFP core, which together leads to complementation of fluorescence on the virion [223]. This three-way complementation informs about virion proximity to cytoplasmic proteins of interest.
Fluorescent AdV particles were shown early on to use the microtubule-dependent transport system to traffic to the nucleus and uncoat the genome at the nuclear pore complex [29,63,224]. Fluorophore-tagged SV40 revealed actin-dependent virus entry [225], and fluorophore-tagged human papillomavirus (HPV) 31 pseudovirions were tracked by confocal fluorescence microscopy to reveal the movement of single virus particles via retrograde transport from the filopodium periphery toward the cell body [80,226]. Experiments with quantum-dot labeled IAV particles suggested a motor switch on the endosome from myosin VI to cytoplasmic dynein [227], and GFP-tagged HSV1 capsids were found to traffic on microtubules [228,229,230,231]. In addition, the bis-arsenical fluorescein derivative FlAsH reveal trafficking of HIV particles [232]. For IAV, organic fluorophore labeled isolated vRNPs that were microinjected into cells revealed cytoplasmic trafficking [233], whereas fluorophore-tagged IAV particles that were endocytosed into cells highlighted the impact of the microtubule network on endosomal transport processes [104]. A summary of fluorescent virus particles has been provided in Table 1.
The recent development of superresolution microscopy, including single molecule localisation microscopy (SMLM) [204,205], stimulated emission depletion microscopy (STED) [202], and structured illumination microscopy (SIM) [234], has been breaking the diffraction-limited resolution barrier. However, most superresolution microscopy methods have a low acquisition speed, high phototoxicity and require heavy post-acquisition processing, which makes volumetric time-lapse imaging of live cells impractical [235]. Nonetheless, superresolution fluorescence microscopy has been used to gain subcellular localization information of virions and viral genomes in chemically fixed cells with single particle resolution [14,52,56,57,66,183,236]. This has demonstrated, for example, that cytosolic AdV particles are leaky containers that have undergone limited disassembly, yet still perfectly enclose the viral genome and shield it against cytosolic sensors and innate immunity [28,65,66].
In addition to fluorescence microscopy, label-free microscopy based on interferometry has been used to track the motions of extracellular vesicles and HSV1 particles, as well as SV40 particles on supported membrane bilayers harbouring the SV40 glycolipid receptor [215,237]. Optical interferometry is a technique which records an interference pattern of superimposing electromagnetic waves following interaction with an object. Interferometry is a low-invasive approach to reveal the dynamics of virion trafficking at high temporal and spatial resolution.
Another emerging development to unravel molecular details of virus–host interactions is force spectroscopy applied to single virions, including atomic force microscopy (AFM) and optical tweezers (OT) [238]. This allows for the mapping of physical properties of virions and the impact of particle physics on the entry of HIV, HSV1 and AdV into cells [8,9,239,240,241,242,243,244]. AFM and OT have also been used for determining force–distance curves in virion binding to cells using microscale thermophoresis, and for mapping of the interactions of virion components with potential cell surface adhesion molecules [238,245].
Another label-free imaging modality to track virions was recently introduced, coherent brightfield microscopy (COBRIM). COBRIM detects scattered light by imaging-based interferometry. It can be combined with digital image processing and post-processing to remove background scatter noise from cellular structures, and thereby allowed to track the motions of VV particle at nanometer accuracy and microsecond temporal resolution [128].
7. Tool Box—Single Virus Particle Tracking
Initially, virion motions were manually tracked with limited precision and small numbers of trajectories [62]. Advances in virion labelling, microscopy engineering and image analyses have recently allowed for live-cell observation of thousands of virus particles at high temporal and spatial resolution. Single particle tracking experiments are analyzed in three distinct steps: particle detection and tracking, trajectory classification, and physical modeling [200]. A typical workflow of a virion imaging and tracking experiment is depicted in Figure 2.
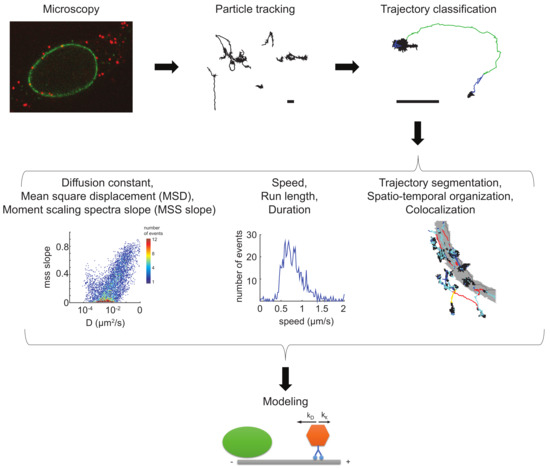
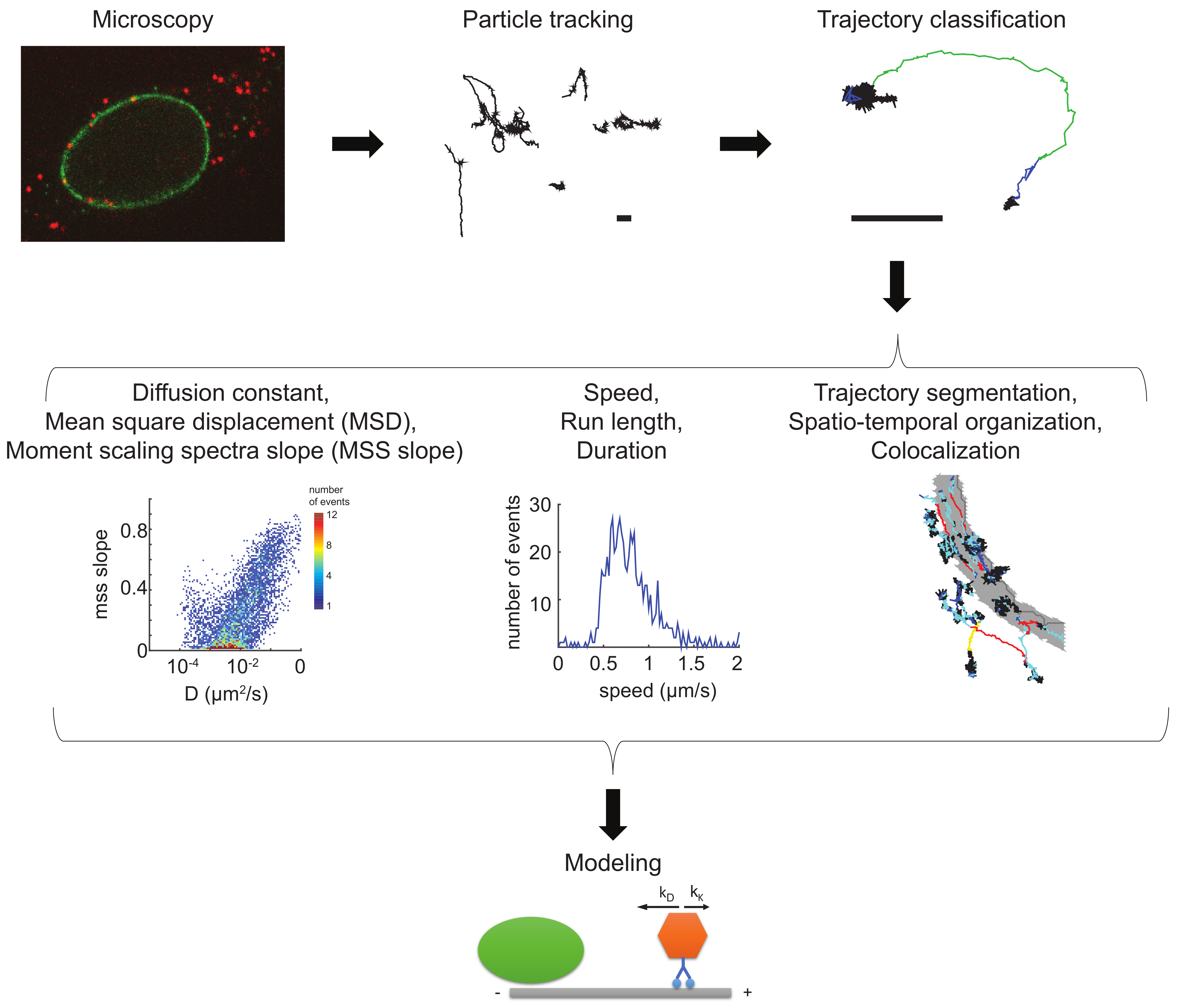
Figure 2.
Workflow of a virus imaging and single particle tracking experiment. Images from fluorescent viral particles infecting host cells are acquired at high temporal and spatial resolution (microscopy). Images are processed and particles detected and tracked over time (particle tracking). The resulting virus trajectories can be analyzed by a plethora of different and orthogonal approaches, including diffusion and moment scaling spectrum slope measurements, and trajectory segmentation (schematic depiction in the center row). Motion properties can be extracted from entire tracks or from segments. Virus particles in motion can contain fluorescent cellular marker proteins or localize with subcellular compartments. Parameters extracted from the motion behavior can be used for various biocomputational modeling approaches to generate predictions that can be tested in follow-up experiments (modeling). Scale bars: 1 µm. The segmented tracks on grey shaded nuclear outlines of HeLa cells have been derived from previously published data, and were adapted with permission from The Company of Biologists Ltd. (Cambridge, UK) [183].
Below, we highlight recent applications in virus imaging and single particle tracking, and discuss advances and challenges. Viral particles are detected in time-lapse image series, time-resolved particle positions are recorded, and the resulting trajectories are analysed and particle motion is classified (reviewed in [196,246,247,248]). Imaging modalities used for virus tracking range from wide field microscopy, total internal reflection microscopy (TIRF) to ring sheet light-sheet microscopy and confocal microscopy. All these techniques allow for the detection of virions labelled with dozens of photostable small organic fluorescent dyes or fluorescent proteins at high signal to noise ratios [210,249]. The strong signals emitted from the diffraction limited viral particles allow for highly accurate particle localization in the nanometer range [249], well below the resolution of the microscopes.
For dim signals from particles labelled with only few copies of GFP-fusion proteins [65], or for signals from rapidly moving particles [180,250], the detection of particles has been challenging. Rapidly moving particles cover a longer distance during the acquisition time of the image and depending on the frame rate in image acquisition, their signal may spread over a large area and lose peak intensity. As the signal degenerates, the particles become harder to detect and localization becomes less accurate. This has been noticed, for example, by advanced wavelet-based transforms, which are mathematical operations to characterize spatial image information over large space scales with greater accuracy than intensity-based segmentation algorithms. More recent developments have replaced the wavelet-based transforms by model-based detection algorithms, which score low signal to noise image data [251], and improve the detection of dim diffraction limited objects, such as clathrin-coated pits [252].
A variety of tracking algorithms are available for diffraction-limited objects, such as virus particles. The performance of trackers was assessed in an unbiased “grand-challenge” on synthetic and real data, including virus motion [253]. The goal for research groups was to independently apply their algorithms on a set of test cases, and to compare the results based on common evaluation criteria. Performance varied depending on the task, and certain trackers performed particularly well on the virus motion data [249,254,255].
Nonetheless, tracking of heterogeneous and crowded motions of virions on the cell surface or in the cytoplasm remains challenging, especially if the particles appear and disappear from the focal plane of observation. Particle appearance/disappearance can be due to the curvilinear rails of microtubules serving as tracks for high speed virion motions [52]. To computationally connect discontinuous tracks, gap closing algorithms have been successfully applied post-processing [251].
Molecular crowding occurs, for example, when virions are tethered to microtubule or actin filaments at high filament density. Heterogeneous virion motions exhibit rapidly changing directionality, stop-and-go bursts, long periods of spatial confinement or fast transitions between motion types [200]. To improve the tracking in crowded conditions, a piecewise-stationary motion model smoother (PMMS) approach was developed and tested on the cytoplasmic motion of AdV particles [256]. PMMS applies an iterative recursive tracking approach, where particles are tracked in both forward and backward direction over multiple iterations. This helps recovering tracks that undergo spontaneous transitions in motion patters [256], for example between confined and directed motion, as it is frequently observed for cytoplasmic motion of AdV (see Figure 3). Significantly, the advent of advanced light sheet microscopy will enhance further developments in tracking of diffraction limited objects in 3D [257,258,259].
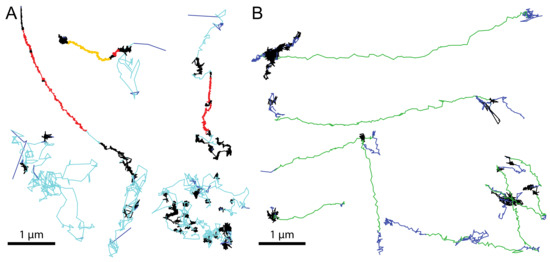
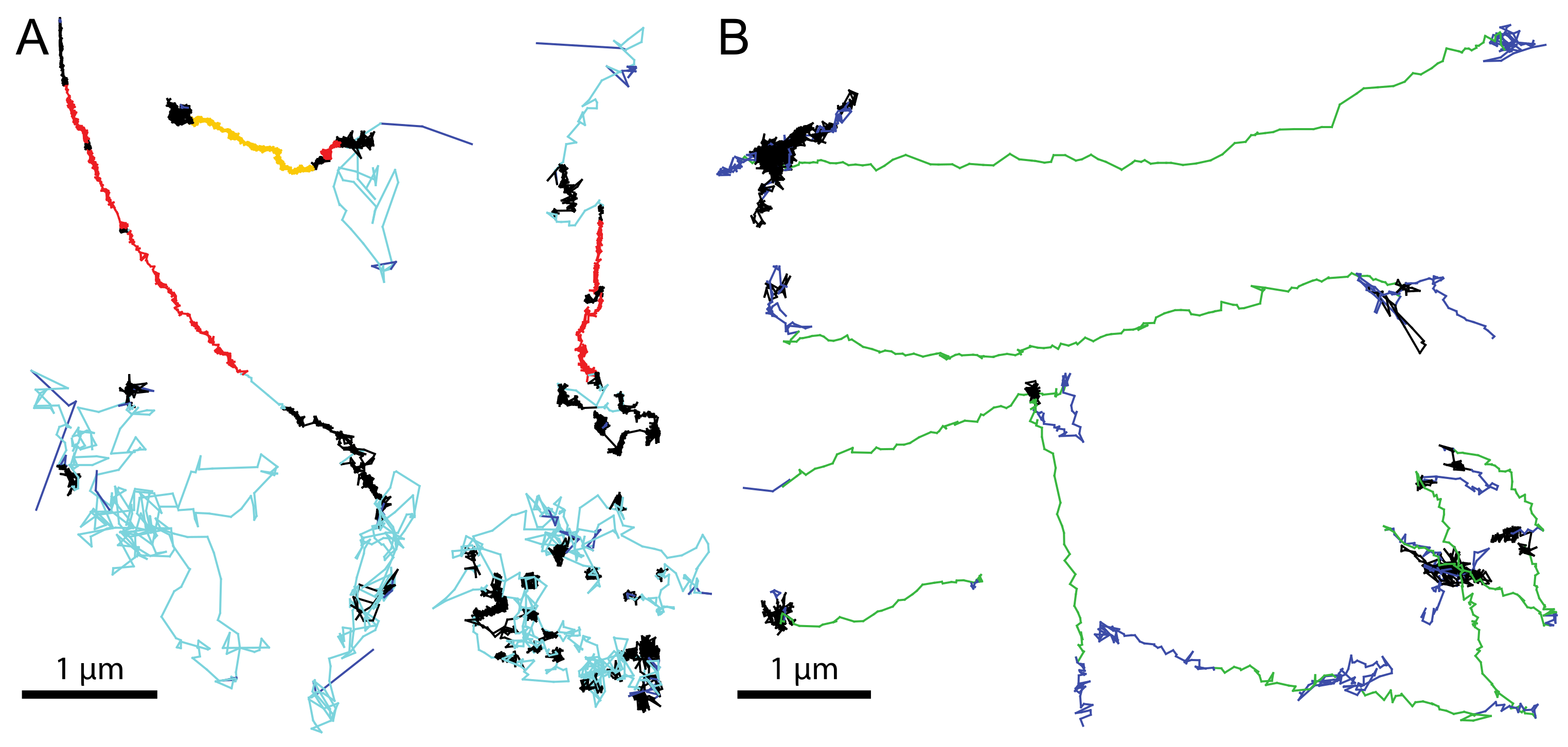
Figure 3.
Examples of virion trajectories on the cell surface and in the cytosol. Segmented trajectories from AdV at the cell surface (A) and in the cytoplasm (B). Cell surface motion was classified in diffusion (cyan), slow drift (red), fast drift (orange), confined motion (black) and not classified steps (blue). Cytoplasmic motion was classified into directed motion (green), fast and slow drifts (orange and red, respectively), confined motion (black) and not classified steps (blue). Scale bars are 1 µm. Technical details of the tracking and segmentation procedures are described in [27]. The segmented tracks in panel A were derived from previously published data, and adapted with permission from Elsevier [27]. The segmented tracks in panel B were adapted from [183], with permission from The Company of Biologists Ltd.
8. Tool Box—Trajectory Analyses
Various classification methods for single particle trajectories have been proposed. Significantly, all the microscopy software packages for object tracking contain a trajectory classifier [247,260,261]. Table 3 provides a compilation of different tracking and trajectory analysis software used for viruses.

Table 3.
Open source tracking and trajectory analysis packages. Adapted from [253,261].
The most commonly used trajectory classification describes the speeds of the entire tracks, or domains within heterogeneous tracks. For example, the reported transport speeds of virions on microtubules covered a wide range of speeds for different viruses, with top speeds in the range of several µm/s (Table 4).

Table 4.
Virion speeds on microtubules.
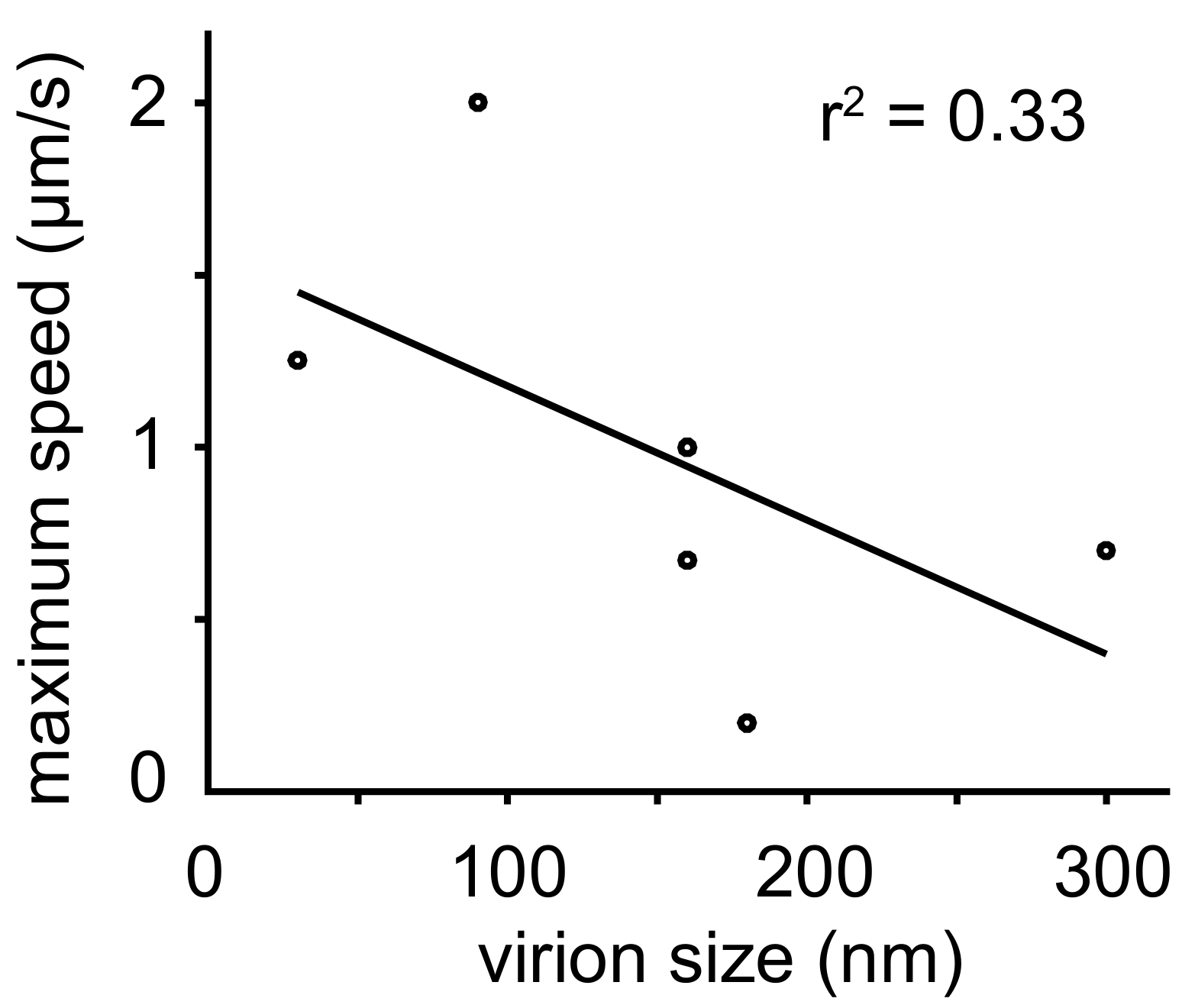
A meta-analysis revealed an inverse correlation between particle size and transport speed (see Figure 4). Small viruses like FMDV (25–30 nm) and AdV (90 nm) moved at maximum speeds of 1–2 µm/s, while much larger rabies virus (180 nm) and VV intracellular enveloped particle (IEV) (300 nm) were transported at speeds around 0.5 µm/s. Speed reduction could be explained by increased drag forces on particles with larger radii in agreement with the Stokes law [268]. Larger particles may also be slowed down in a crowded cytoplasm by entanglements with organelles and filaments. Alternatively, larger viruses may offer more binding sites for motor proteins of opposite directionality and hence may get slowed down by tug-of-war [250].
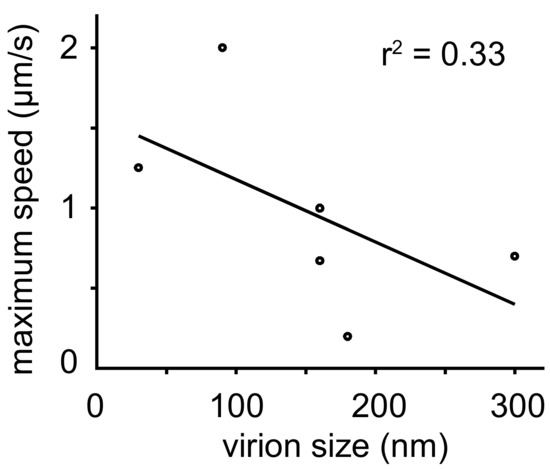
Figure 4.
Plot of virion speed on microtubules versus particle size. Microtubule-based speed of viral particles inversely correlates with particle size. Reported maximum speeds and particle sizes are shown. For details and references, see Table 4.
In addition to the overall particle velocity, the peak speeds and continuous drifting speeds, the mean square displacement (MSD), diffusion coefficients, the moment scaling spectra slopes (SMSS), and segmentations have been widely used for characterization and classification of viral trajectories [269,270]. A compilation of motion features that have been extracted by virus trajectory analyses is provided in Table 5.

Table 5.
Trajectory analyses extract a range of virus motion features. Total internal reflection (TIRF) microscopy.
In fact, some of the first virus tracking experiments were reported with fluorescent IV particles on the surface of human fibroblast in cold medium [103,271]. MSD analyses indicated that the particles underwent random diffusion and continuous drifts, albeit at very low frequency. The slope of the momentum scaling spectrum classifies the motion diffusivity into confined, diffusive and superdiffusive motions [249,272]. SMSS analysis is useful and powerful for moving object analysis [273], and especially for viruses, since virus trajectories are heterogeneous and rarely highly processive due to virion interactions with a range of host factors, as amply demonstrated by cell surface motion analyses of AdV, for example [27]. In addition, SMSS analysis identified filopodial drifting motions of murine poliomavirus virus-like particles and HPV16 pseudoviruses, which are assembled from recombinant capsid proteins and lack a viral genome [79,196].
Heterogeneous motion behavior is readily revealed by imaging at high spatial and temporal resolution [274]. For example, transient confinement zones have been investigated for a variety of molecules [275]. Algorithms have been developed to detect jumps between adjacent confinement corrals and sudden changes in diffusion coefficients [276,277,278] or SMSS for intracellular Mason–Pfizer Monkey retroviral (M-PMV) particles [261].
Differential motion behavior was also detected by moving window approaches, where motion parameters are extracted from sub-segments and changes in motion behavior are defined by thresholding methods [276,279] or segment classification [280]. In addition, confinement zones were detected using Bayesian methods [281,282], Hidden Markov models [283] and particle filtering methods [284]. Likewise, Hidden Markov models were used to extract dynamic colocalization events from multi-channel image data in the case of HIV membrane fusion events [285].
Importantly, machine learning approaches were introduced for trajectory analysis including support vector machines (SVM) for AdV motion analyses [200]. Virion trajectories were segmented into different motion modes including confined motion, directed motion and drifts with supervised SVM classification [200] (Figure 4). Subsequently, neural networks were developed for the analysis of membrane receptor motion [286]. While single particle tracking algorithms converged to more robust solutions in the past decade [253], novel motion analysis procedures are being developed. This will likely enhance the analyses of complex and heterogeneous motion behaviors in 2D and 3D biological samples, and multi-channel data sets. Interference approaches and analyses of complex virus motions are expected to give insight into biological mechanisms of host cell and tissue infections in the course of disease.
9. Tool Box—Physical Models Describing the Movements of Incoming Virus Particles
Computational modeling and simulations can provide in silico experimental data to enhance mechanisms and give insight into interaction processes. Increasingly, aspects of infection dynamics are addressed by mathematical and computational modeling (for reviews, see [287,288,289]). Parameters extracted from virus tracking experiments can be used in data-driven models which are also known as “top-down” models. They explore patterns and make correlations from statistical or machine learning methods. Examples include correlation analyses, reconstruction of molecular interaction networks from high-content screening datasets, and classification of viral motion types using machine learning [200,290,291]. Data-driven models suggest mechanisms, and can indicate necessity of a process or a molecule in a perturbation experiment [289]. This is notably different, yet complimentary to so called “bottom-up” modelling, where physical models are constructed based on mechanistic hypothesis derived from prior data or theoretical reasoning [289]. In contrast, the “top-down” approach allows for discovery of new patterns concealed in the data.
With regard to cytoskeletal transport, bidirectional active transport along microtubules, propelled by dynein and kinesin motors has been an attractive subject for modelling studies. A general mathematical model for stop-and-go virus motion on microtubules was proposed [287,292,293]. In addition, a stochastic computational model of AdV cytoplasmic transport was built using parameters extracted from single particle tracking and trajectory segmentation experiments [250]. This model allowed the possibility of a tug-of-war between motors of opposite migration on microtubules, such as cytoplasmic dynein and conventional kinesin [294]. The model also allowed predictions of the number of active motors attached to single virions during fast directed motions, namely two to three, and the number of the virion binding sites for the microtubule motors, which implied the major virion capsid protein hexon [250]. Subsequently, empirical evidence confirmed that hexon directly binds to the microtubule minus-end trafficking motor protein, the dynein complex [64].
10. Conclusions and Outlook
An increasingly refined picture is emerging of how virus particles traffic on the cytoskeleton during entry and egress from cells. The knowledge emerges from analyses of virion trajectories by computational methods, and in silico simulation experiments. Advances in virion imaging and motion analyses provide a basis to inform about biological mechanisms, and foster the development of anti-viral therapeutics. They critically enhance the conceptual understanding of cell functions, based on the notion that viruses interact with thousands of proteins in an infected cell, and are major drivers of host adaption in evolution, and immune regulation.
Open questions and challenges comprise virus trafficking studies in cultured cells, primary cells and tissues, and the complex communication processes of the host to and from the pathogen. For example, the deciphering of how cytokines, chemokines and other signalling molecules affect the way viruses use the cytoskeleton and tune cytoskeletal dynamics will inform passive and active pro- and anti-viral mechanisms. Deep analyses of virion motions in cells will further inform on underlying mechanisms of cell-to-cell heterogeneity of infection, a phenomenon which is largely unexplored but well known to every virologist. Finally, time-controlled virus infection analyses will better prepare the field to reach out to even more complex settings, for example the elucidation of the interactions between eukaryotes, microbes and viruses in the course of human health and disease.
Acknowledgments
The work was funded by a grants from the Swiss National Science Foundation no. 310030B_160316 and Sinergia CRSII5_170929 / 1. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Lederberg, J. Infectious history. Science 2000, 288, 287–293. [Google Scholar] [CrossRef] [PubMed]
- Waehler, R.; Russell, S.J.; Curiel, D.T. Engineering targeted viral vectors for gene therapy. Nat. Rev. Genet. 2007, 8, 573–587. [Google Scholar] [CrossRef] [PubMed]
- Ludmir, E.B.; Enquist, L.W. Viral genomes are part of the phylogenetic tree of life. Nat. Rev. Microbiol. 2009, 7, 615. [Google Scholar] [CrossRef] [PubMed]
- Greber, U.F.; Arnberg, N.; Wadell, G.; Benko, M.; Kremer, E.J. Adenoviruses—From pathogens to therapeutics: A report on the 10th international adenovirus meeting. Cell. Microbiol. 2013, 15, 16–23. [Google Scholar] [CrossRef] [PubMed]
- Virgin, H.W.; Wherry, E.J.; Ahmed, R. Redefining chronic viral infection. Cell 2009, 138, 30–50. [Google Scholar] [CrossRef] [PubMed]
- Dobos, P.; Hallett, R.; Kells, D.T.; Sorensen, O.; Rowe, D. Biophysical studies of infectious pancreatic necrosis virus. J. Virol. 1977, 22, 150–159. [Google Scholar] [PubMed]
- Evilevitch, A. Physical evolution of pressure-driven viral infection. Biophys. J. 2013, 104, 2113–2114. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Greber, U.F. How cells tune viral mechanics—Insights from biophysical measurements of influenza virus. Biophys. J. 2014, 106, 2317–2321. [Google Scholar] [CrossRef] [PubMed]
- Greber, U.F. Virus and host mechanics support membrane penetration and cell entry. J. Virol. 2016, 90, 3802–3805. [Google Scholar] [CrossRef] [PubMed]
- Abergel, C.; Legendre, M.; Claverie, J.M. The rapidly expanding universe of giant viruses: Mimivirus, pandoravirus, pithovirus and mollivirus. FEMS Microbiol. Rev. 2015, 39, 779–796. [Google Scholar] [CrossRef] [PubMed]
- Yamauchi, Y.; Greber, U.F. Principles of virus uncoating: Cues and the snooker ball. Traffic 2016, 17, 569–592. [Google Scholar] [CrossRef] [PubMed]
- Mercer, J.; Schelhaas, M.; Helenius, A. Virus entry by endocytosis. Annu. Rev. Biochem. 2010, 79, 803–833. [Google Scholar] [CrossRef] [PubMed]
- Wolfrum, N.; Greber, U.F. Adenovirus signalling in entry. Cell. Microbiol. 2013, 15, 53–62. [Google Scholar] [CrossRef] [PubMed]
- Burckhardt, C.J.; Greber, U.F. Virus movements on the plasma membrane support infection and transmission between cells. PLoS. Pathog. 2009, 5, e1000621. [Google Scholar] [CrossRef] [PubMed]
- Greber, U.F.; Carafoli, E. Signalling takes control of nucleo-cytoplasmic trafficking. Workshop on signal-regulated nuclear transport. EMBO Rep. 2002, 3, 410–414. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Blaas, D. Viral entry pathways: The example of common cold viruses. Wien Med. Wochenschr. 2016, 166, 211–226. [Google Scholar] [CrossRef] [PubMed]
- Albornoz, A.; Hoffmann, A.B.; Lozach, P.Y.; Tischler, N.D. Early bunyavirus-host cell interactions. Viruses 2016, 8, 143. [Google Scholar] [CrossRef] [PubMed]
- Boulant, S.; Stanifer, M.; Lozach, P.Y. Dynamics of virus-receptor interactions in virus binding, signaling, and endocytosis. Viruses 2015, 7, 2794–2815. [Google Scholar] [CrossRef] [PubMed]
- Grove, J.; Marsh, M. The cell biology of receptor-mediated virus entry. J. Cell Biol. 2011, 195, 1071–1082. [Google Scholar] [CrossRef] [PubMed]
- Cotmore, S.F.; Tattersall, P. Parvoviral host range and cell entry mechanisms. Adv. Virus Res. 2007, 70, 183–232. [Google Scholar] [PubMed]
- Gruenberg, J.; van der Goot, F.G. Mechanisms of pathogen entry through the endosomal compartments. Nat. Rev. Mol. Cell Biol. 2006, 7, 495–504. [Google Scholar] [CrossRef] [PubMed]
- Sieczkarski, S.B.; Whittaker, G.R. Viral entry. Curr. Top. Microbiol. Immunol. 2005, 285, 1–23. [Google Scholar] [PubMed]
- Lyman, M.G.; Enquist, L.W. Herpesvirus interactions with the host cytoskeleton. J. Virol. 2009, 83, 2058–2066. [Google Scholar] [CrossRef] [PubMed]
- Hogue, I.B.; Bosse, J.B.; Engel, E.A.; Scherer, J.; Hu, J.R.; Del Rio, T.; Enquist, L.W. Fluorescent protein approaches in alpha herpesvirus research. Viruses 2015, 7, 5933–5961. [Google Scholar] [CrossRef] [PubMed]
- Greber, U.F.; Willetts, M.; Webster, P.; Helenius, A. Stepwise dismantling of adenovirus 2 during entry into cells. Cell 1993, 75, 477–486. [Google Scholar] [CrossRef]
- Nakano, M.Y.; Boucke, K.; Suomalainen, M.; Stidwill, R.P.; Greber, U.F. The first step of adenovirus type 2 disassembly occurs at the cell surface, independently of endocytosis and escape to the cytosol. J. Virol. 2000, 74, 7085–7095. [Google Scholar] [CrossRef] [PubMed]
- Burckhardt, C.J.; Suomalainen, M.; Schoenenberger, P.; Boucke, K.; Hemmi, S.; Greber, U.F. Drifting motions of the adenovirus receptor car and immobile integrins initiate virus uncoating and membrane lytic protein exposure. Cell Host Microbe 2011, 10, 105–117. [Google Scholar] [CrossRef] [PubMed]
- Suomalainen, M.; Luisoni, S.; Boucke, K.; Bianchi, S.; Engel, D.A.; Greber, U.F. A direct and versatile assay measuring membrane penetration of adenovirus in single cells. J. Virol. 2013, 87, 12367–12379. [Google Scholar] [CrossRef] [PubMed]
- Strunze, S.; Engelke, M.F.; Wang, I.H.; Puntener, D.; Boucke, K.; Schleich, S.; Way, M.; Schoenenberger, P.; Burckhardt, C.J.; Greber, U.F. Kinesin-1-mediated capsid disassembly and disruption of the nuclear pore complex promote virus infection. Cell Host Microbe 2011, 10, 210–223. [Google Scholar] [CrossRef] [PubMed]
- Luisoni, S.; Greber, U.F. Biology of adenovirus cell entry—Receptors, pathways, mechanisms. In Adenoviral Vectors for Gene Therapy, 2nd ed.; Curiel, D., Ed.; Academic Press, Elsevier: London, UK, 2016; pp. 27–58. [Google Scholar]
- Moss, B. Poxviridae: The viruses and their replication. In Fields Virology, 5th ed.; Knipe, D.M., Howley, P.M., Griffin, D.E., Lamb, R.A., Martin, M.A., Roizman, B., Straus, S.E., Eds.; Lippincott Williams & Wilkins: Philadelphia, PA, USA, 2007; Volume 2, pp. 2905–2945. [Google Scholar]
- Yakimovich, A.; Huttunen, M.; Zehnder, B.; Coulter, L.J.; Gould, V.; Schneider, C.; Kopf, M.; McInnes, C.J.; Greber, U.F.; Mercer, J. Inhibition of poxvirus gene expression and genome replication by bisbenzimide derivatives. J. Virol. 2017, 91. [Google Scholar] [CrossRef] [PubMed]
- Peng, K.; Muranyi, W.; Glass, B.; Laketa, V.; Yant, S.R.; Tsai, L.; Cihlar, T.; Muller, B.; Krausslich, H.G. Quantitative microscopy of functional hiv post-entry complexes reveals association of replication with the viral capsid. eLife 2014, 3, e04114. [Google Scholar] [CrossRef] [PubMed]
- Jacques, D.A.; McEwan, W.A.; Hilditch, L.; Price, A.J.; Towers, G.J.; James, L.C. HIV-1 uses dynamic capsid pores to import nucleotides and fuel encapsidated DNA synthesis. Nature 2016, 536, 349–353. [Google Scholar] [CrossRef] [PubMed]
- Hendrickx, R.; Stichling, N.; Koelen, J.; Kuryk, L.; Lipiec, A.; Greber, U.F. Innate immunity to adenovirus. Hum. Gene. Ther. 2014, 25, 265–284. [Google Scholar] [CrossRef] [PubMed]
- Xiao, T.S.; Fitzgerald, K.A. The cGAS-STING pathway for DNA sensing. Mol. Cell 2013, 51, 135–139. [Google Scholar] [CrossRef] [PubMed]
- Schoggins, J.W.; Randall, G. Lipids in innate antiviral defense. Cell Host Microbe 2013, 14, 379–385. [Google Scholar] [CrossRef] [PubMed]
- Kobiler, O.; Drayman, N.; Butin-Israeli, V.; Oppenheim, A. Virus strategies for passing the nuclear envelope barrier. Nucleus 2012, 3, 526–539. [Google Scholar] [CrossRef] [PubMed]
- Flatt, J.W.; Greber, U.F. Viral mechanisms for docking and delivering at nuclear pore complexes. Semin. Cell Dev. Biol. 2017, 68, 59–71. [Google Scholar] [CrossRef] [PubMed]
- Mettenleiter, T.C. Breaching the barrier-the nuclear envelope in virus infection. J. Mol. Biol. 2016, 428, 1949–1961. [Google Scholar] [CrossRef] [PubMed]
- Flatt, J.W.; Greber, U.F. Misdelivery at the nuclear pore complex-stopping a virus dead in its tracks. Cells 2015, 4, 277–296. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, Y.; Craigie, R. The road to chromatin—Nuclear entry of retroviruses. Nat. Rev. Microbiol. 2007, 5, 187–196. [Google Scholar] [CrossRef] [PubMed]
- Aydin, I.; Schelhaas, M. Viral genome tethering to host cell chromatin: Cause and consequences. Traffic 2016, 17, 327–340. [Google Scholar] [CrossRef] [PubMed]
- Nicolson, S.C.; Samulski, R.J. Recombinant adeno-associated virus utilizes host cell nuclear import machinery to enter the nucleus. J. Virol. 2014, 88, 4132–4144. [Google Scholar] [CrossRef] [PubMed]
- Basyuk, E.; Galli, T.; Mougel, M.; Blanchard, J.M.; Sitbon, M.; Bertrand, E. Retroviral genomic RNAs are transported to the plasma membrane by endosomal vesicles. Dev. Cell 2003, 5, 161–174. [Google Scholar] [CrossRef]
- Taylor, M.P.; Enquist, L.W. Axonal spread of neuroinvasive viral infections. Trends Microbiol. 2015, 23, 283–288. [Google Scholar] [CrossRef] [PubMed]
- Bird, S.W.; Kirkegaard, K. Nonlytic spread of naked viruses. Autophagy 2015, 11, 430–431. [Google Scholar] [CrossRef] [PubMed]
- Bird, S.W.; Kirkegaard, K. Escape of non-enveloped virus from intact cells. Virology 2015, 479–480, 444–449. [Google Scholar] [CrossRef] [PubMed]
- Ravindran, M.S.; Bagchi, P.; Cunningham, C.N.; Tsai, B. Opportunistic intruders: How viruses orchestrate er functions to infect cells. Nat. Rev. Microbiol. 2016, 14, 407–420. [Google Scholar] [CrossRef] [PubMed]
- Luby-Phelps, K.; Castle, P.E.; Taylor, D.L.; Lanni, F. Hindered diffusion of inert tracer particles in the cytoplasm of mouse 3T3 cells. Proc. Natl. Acad. Sci. USA 1987, 84, 4910–4913. [Google Scholar] [CrossRef] [PubMed]
- Dohner, K.; Sodeik, B. The role of the cytoskeleton during viral infection. Curr. Top. Microbiol. Immunol. 2005, 285, 67–108. [Google Scholar] [PubMed]
- Greber, U.F.; Way, M. A superhighway to virus infection. Cell 2006, 124, 741–754. [Google Scholar] [CrossRef] [PubMed]
- Leopold, P.L.; Crystal, R.G. Intracellular trafficking of adenovirus: Many means to many ends. Adv. Drug Deliv. Rev. 2007, 59, 810–821. [Google Scholar] [CrossRef] [PubMed]
- Seksek, O.; Biwersi, J.; Verkman, A.S. Translational diffusion of macromolecule-sized solutes in cytoplasm and nucleus. J. Cell Biol. 1997, 138, 131–142. [Google Scholar] [CrossRef] [PubMed]
- Nakano, M.Y.; Greber, U.F. Quantitative microscopy of fluorescent adenovirus entry. J. Struct. Biol. 2000, 129, 57–68. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Sun, E.; He, J.; Zhuang, X. Live cell imaging of viral entry. Curr. Opin. Virol. 2013, 3, 34–43. [Google Scholar] [CrossRef] [PubMed]
- Brandenburg, B.; Zhuang, X. Virus trafficking—Learning from single-virus tracking. Nat. Rev. Microbiol. 2007, 5, 197–208. [Google Scholar] [CrossRef] [PubMed]
- Greber, U.F. Viral trafficking violations in axons: The herpesvirus case. Proc. Natl. Acad. Sci. USA 2005, 102, 5639–5640. [Google Scholar] [CrossRef] [PubMed]
- Meier, O.; Greber, U.F. Adenovirus endocytosis. J. Gene Med. 2003, 5, 451–462. [Google Scholar] [CrossRef] [PubMed]
- Mothes, W.; Sherer, N.M.; Jin, J.; Zhong, P. Virus cell-to-cell transmission. J. Virol. 2010, 84, 8360–8368. [Google Scholar] [CrossRef] [PubMed]
- Lux, K.; Goerlitz, N.; Schlemminger, S.; Perabo, L.; Goldnau, D.; Endell, J.; Leike, K.; Kofler, D.M.; Finke, S.; Hallek, M.; et al. Green fluorescent protein-tagged adeno-associated virus particles allow the study of cytosolic and nuclear trafficking. J. Virol. 2005, 79, 11776–11787. [Google Scholar] [CrossRef] [PubMed]
- Leopold, P.L.; Ferris, B.; Grinberg, I.; Worgall, S.; Hackett, N.R.; Crystal, R.G. Fluorescent virions: Dynamic tracking of the pathway of adenoviral gene transfer vectors in living cells. Hum. Gene Ther. 1998, 9, 367–378. [Google Scholar] [CrossRef] [PubMed]
- Suomalainen, M.; Nakano, M.Y.; Keller, S.; Boucke, K.; Stidwill, R.P.; Greber, U.F. Microtubule-dependent plus- and minus end-directed motilities are competing processes for nuclear targeting of adenovirus. J. Cell Biol. 1999, 144, 657–672. [Google Scholar] [CrossRef] [PubMed]
- Bremner, K.H.; Scherer, J.; Yi, J.; Vershinin, M.; Gross, S.P.; Vallee, R.B. Adenovirus transport via direct interaction of cytoplasmic dynein with the viral capsid hexon subunit. Cell Host Microbe 2009, 6, 523–535. [Google Scholar] [CrossRef] [PubMed]
- Puntener, D.; Engelke, M.F.; Ruzsics, Z.; Strunze, S.; Wilhelm, C.; Greber, U.F. Stepwise loss of fluorescent core protein V from human adenovirus during entry into cells. J. Virol. 2011, 85, 481–496. [Google Scholar] [CrossRef] [PubMed]
- Wang, I.H.; Suomalainen, M.; Andriasyan, V.; Kilcher, S.; Mercer, J.; Neef, A.; Luedtke, N.W.; Greber, U.F. Tracking viral genomes in host cells at single-molecule resolution. Cell Host Microbe 2013, 14, 468–480. [Google Scholar] [CrossRef] [PubMed]
- Meier, O.; Boucke, K.; Hammer, S.V.; Keller, S.; Stidwill, R.P.; Hemmi, S.; Greber, U.F. Adenovirus triggers macropinocytosis and endosomal leakage together with its clathrin-mediated uptake. J. Cell Biol. 2002, 158, 1119–1131. [Google Scholar] [CrossRef] [PubMed]
- Imelli, N.; Meier, O.; Boucke, K.; Hemmi, S.; Greber, U.F. Cholesterol is required for endocytosis and endosomal escape of adenovirus type 2. J. Virol. 2004, 78, 3089–3098. [Google Scholar] [CrossRef] [PubMed]
- Meier, O.; Gastaldelli, M.; Boucke, K.; Hemmi, S.; Greber, U.F. Early steps of clathrin-mediated endocytosis involved in phagosomal escape of Fcgamma receptor-targeted adenovirus. J. Virol. 2005, 79, 2604–2613. [Google Scholar] [CrossRef] [PubMed]
- Gastaldelli, M.; Imelli, N.; Boucke, K.; Amstutz, B.; Meier, O.; Greber, U.F. Infectious adenovirus type 2 transport through early but not late endosomes. Traffic 2008, 9, 2265–2278. [Google Scholar] [CrossRef] [PubMed]
- Sirena, D.; Lilienfeld, B.; Eisenhut, M.; Kalin, S.; Boucke, K.; Beerli, R.R.; Vogt, L.; Ruedl, C.; Bachmann, M.F.; Greber, U.F.; et al. The human membrane cofactor CD46 is a receptor for species B adenovirus serotype 3. J. Virol. 2004, 78, 4454–4462. [Google Scholar] [CrossRef] [PubMed]
- Amstutz, B.; Gastaldelli, M.; Kalin, S.; Imelli, N.; Boucke, K.; Wandeler, E.; Mercer, J.; Hemmi, S.; Greber, U.F. Subversion of CtBP1-controlled macropinocytosis by human adenovirus serotype 3. EMBO J. 2008, 27, 956–969. [Google Scholar] [CrossRef] [PubMed]
- Kalin, S.; Amstutz, B.; Gastaldelli, M.; Wolfrum, N.; Boucke, K.; Havenga, M.; DiGennaro, F.; Liska, N.; Hemmi, S.; Greber, U.F. Macropinocytotic uptake and infection of human epithelial cells with species B2 adenovirus type 35. J. Virol. 2010, 84, 5336–5350. [Google Scholar] [CrossRef] [PubMed]
- Hueffer, K.; Parker, J.S.; Weichert, W.S.; Geisel, R.E.; Sgro, J.Y.; Parrish, C.R. The natural host range shift and subsequent evolution of canine parvovirus resulted from virus-specific binding to the canine transferrin receptor. J. Virol. 2003, 77, 1718–1726. [Google Scholar] [CrossRef] [PubMed]
- Cureton, D.K.; Harbison, C.E.; Cocucci, E.; Parrish, C.R.; Kirchhausen, T. Limited transferrin receptor clustering allows rapid diffusion of canine parvovirus into clathrin endocytic structures. J. Virol. 2012, 86, 5330–5340. [Google Scholar] [CrossRef] [PubMed]
- Pietiainen, V.; Marjomaki, V.; Upla, P.; Pelkmans, L.; Helenius, A.; Hyypia, T. Echovirus 1 endocytosis into caveosomes requires lipid rafts, dynamin II, and signaling events. Mol. Biol. Cell 2004, 15, 4911–4925. [Google Scholar] [CrossRef] [PubMed]
- Martin-Acebes, M.A.; Vazquez-Calvo, A.; Gonzalez-Magaldi, M.; Sobrino, F. Foot-and-mouth disease virus particles inactivated with binary ethylenimine are efficiently internalized into cultured cells. Vaccine 2011, 29, 9655–9662. [Google Scholar] [CrossRef] [PubMed]
- Smith, J.L.; Campos, S.K.; Ozbun, M.A. Human papillomavirus type 31 uses a caveolin 1- and dynamin 2-mediated entry pathway for infection of human keratinocytes. J. Virol. 2007, 81, 9922–9931. [Google Scholar] [CrossRef] [PubMed]
- Schelhaas, M.; Ewers, H.; Rajamaki, M.L.; Day, P.M.; Schiller, J.T.; Helenius, A. Human papillomavirus type 16 entry: Retrograde cell surface transport along actin-rich protrusions. PLoS. Pathog. 2008, 4, e1000148. [Google Scholar] [CrossRef] [PubMed]
- Smith, J.L.; Lidke, D.S.; Ozbun, M.A. Virus activated filopodia promote human papillomavirus type 31 uptake from the extracellular matrix. Virology 2008, 381, 16–21. [Google Scholar] [CrossRef] [PubMed]
- Brandenburg, B.; Lee, L.Y.; Lakadamyali, M.; Rust, M.J.; Zhuang, X.; Hogle, J.M. Imaging poliovirus entry in live cells. PLoS Biol. 2007, 5, e183. [Google Scholar] [CrossRef] [PubMed]
- Vaughan, J.C.; Brandenburg, B.; Hogle, J.M.; Zhuang, X. Rapid actin-dependent viral motility in live cells. Biophys. J. 2009, 97, 1647–1656. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Ehrlich, M.; Boll, W.; Van Oijen, A.; Hariharan, R.; Chandran, K.; Nibert, M.L.; Kirchhausen, T. Endocytosis by random initiation and stabilization of clathrin-coated pits. Cell 2004, 118, 591–605. [Google Scholar] [CrossRef] [PubMed]
- Kremser, L.; Konecsni, T.; Blaas, D.; Kenndler, E. Fluorescence labeling of human rhinovirus capsid and analysis by capillary electrophoresis. Anal. Chem. 2004, 76, 4175–4181. [Google Scholar] [CrossRef] [PubMed]
- Pelkmans, L.; Puntener, D.; Helenius, A. Local actin polymerization and dynamin recruitment in sv40-induced internalization of caveolae. Science 2002, 296, 535–539. [Google Scholar] [CrossRef] [PubMed]
- Akita, H.; Enoto, K.; Tanaka, H.; Harashima, H. Particle tracking analysis for the intracellular trafficking of nanoparticles modified with african swine fever virus protein p54-derived peptide. Mol. Ther. 2013, 21, 309–317. [Google Scholar] [CrossRef] [PubMed]
- Lehmann, M.J.; Sherer, N.M.; Marks, C.B.; Pypaert, M.; Mothes, W. Actin- and myosin-driven movement of viruses along filopodia precedes their entry into cells. J. Cell Biol. 2005, 170, 317–325. [Google Scholar] [CrossRef] [PubMed]
- Hoornweg, T.E.; van Duijl-Richter, M.K.; Ayala Nunez, N.V.; Albulescu, I.C.; van Hemert, M.J.; Smit, J.M. Dynamics of chikungunya virus cell entry unraveled by single-virus tracking in living cells. J. Virol. 2016, 90, 4745–4756. [Google Scholar] [CrossRef] [PubMed]
- Van der Schaar, H.M.; Rust, M.J.; Chen, C.; van der Ende-Metselaar, H.; Wilschut, J.; Zhuang, X.; Smit, J.M. Dissecting the cell entry pathway of dengue virus by single-particle tracking in living cells. PLoS. Pathog. 2008, 4, e1000244. [Google Scholar] [CrossRef] [PubMed]
- Nanbo, A.; Imai, M.; Watanabe, S.; Noda, T.; Takahashi, K.; Neumann, G.; Halfmann, P.; Kawaoka, Y. Ebolavirus is internalized into host cells via macropinocytosis in a viral glycoprotein-dependent manner. PLoS. Pathog. 2010, 6, e1001121. [Google Scholar] [CrossRef] [PubMed]
- Carette, J.E.; Raaben, M.; Wong, A.C.; Herbert, A.S.; Obernosterer, G.; Mulherkar, N.; Kuehne, A.I.; Kranzusch, P.J.; Griffin, A.M.; Ruthel, G.; et al. Ebola virus entry requires the cholesterol transporter Niemann-Pick C1. Nature 2011, 477, 340–343. [Google Scholar] [CrossRef] [PubMed]
- Costello, D.A.; Millet, J.K.; Hsia, C.Y.; Whittaker, G.R.; Daniel, S. Single particle assay of coronavirus membrane fusion with proteinaceous receptor-embedded supported bilayers. Biomaterials 2013, 34, 7895–7904. [Google Scholar] [CrossRef] [PubMed]
- Costello, D.A.; Daniel, S. Single particle tracking assay to study coronavirus membrane fusion. Methods Mol. Biol. 2015, 1282, 183–194. [Google Scholar] [PubMed]
- Elliott, G.; O’Hare, P. Live-cell analysis of a green fluorescent protein-tagged herpes simplex virus infection. J. Virol. 1999, 73, 4110–4119. [Google Scholar] [PubMed]
- Donnelly, M.; Elliott, G. Fluorescent tagging of herpes simplex virus tegument protein VP13/14 in virus infection. J. Virol. 2001, 75, 2575–2583. [Google Scholar] [CrossRef] [PubMed]
- Sugimoto, K.; Uema, M.; Sagara, H.; Tanaka, M.; Sata, T.; Hashimoto, Y.; Kawaguchi, Y. Simultaneous tracking of capsid, tegument, and envelope protein localization in living cells infected with triply fluorescent herpes simplex virus 1. J. Virol. 2008, 82, 5198–5211. [Google Scholar] [CrossRef] [PubMed]
- De Oliveira, A.P.; Glauser, D.L.; Laimbacher, A.S.; Strasser, R.; Schraner, E.M.; Wild, P.; Ziegler, U.; Breakefield, X.O.; Ackermann, M.; Fraefel, C. Live visualization of herpes simplex virus type 1 compartment dynamics. J. Virol. 2008, 82, 4974–4990. [Google Scholar] [CrossRef] [PubMed]
- Hao, X.; Shang, X.; Wu, J.; Shan, Y.; Cai, M.; Jiang, J.; Huang, Z.; Tang, Z.; Wang, H. Single-particle tracking of hepatitis B virus-like vesicle entry into cells. Small 2011, 7, 1212–1218. [Google Scholar] [CrossRef] [PubMed]
- Coller, K.E.; Berger, K.L.; Heaton, N.S.; Cooper, J.D.; Yoon, R.; Randall, G. RNA interference and single particle tracking analysis of hepatitis C virus endocytosis. PLoS. Pathog. 2009, 5, e1000702. [Google Scholar] [CrossRef] [PubMed]
- McDonald, D.; Vodicka, M.A.; Lucero, G.; Svitkina, T.M.; Borisy, G.G.; Emerman, M.; Hope, T.J. Visualization of the intracellular behavior of HIV in living cells. J. Cell Biol. 2002, 159, 441–452. [Google Scholar] [CrossRef] [PubMed]
- Sherer, N.M.; Lehmann, M.J.; Jimenez-Soto, L.F.; Ingmundson, A.; Horner, S.M.; Cicchetti, G.; Allen, P.G.; Pypaert, M.; Cunningham, J.M.; Mothes, W. Visualization of retroviral replication in living cells reveals budding into multivesicular bodies. Traffic 2003, 4, 785–801. [Google Scholar] [CrossRef] [PubMed]
- Sherer, N.M.; Lehmann, M.J.; Jimenez-Soto, L.F.; Horensavitz, C.; Pypaert, M.; Mothes, W. Retroviruses can establish filopodial bridges for efficient cell-to-cell transmission. Nat. Cell Biol. 2007, 9, 310–315. [Google Scholar] [CrossRef] [PubMed]
- Anderson, C.M.; Georgiou, G.N.; Morrison, I.E.; Stevenson, G.V.; Cherry, R.J. Tracking of cell surface receptors by fluorescence digital imaging microscopy using a charge-coupled device camera. Low-density lipoprotein and influenza virus receptor mobility at 4 degrees C. J. Cell Sci. 1992, 101 Pt 2, 415–425. [Google Scholar] [PubMed]
- Lakadamyali, M.; Rust, M.J.; Babcock, H.P.; Zhuang, X. Visualizing infection of individual influenza viruses. Proc. Natl. Acad. Sci. USA 2003, 100, 9280–9285. [Google Scholar] [CrossRef] [PubMed]
- Rust, M.J.; Lakadamyali, M.; Zhang, F.; Zhuang, X. Assembly of endocytic machinery around individual influenza viruses during viral entry. Nat. Struct. Mol. Biol. 2004, 11, 567–573. [Google Scholar] [CrossRef] [PubMed]
- Smith, A.E.; Lilie, H.; Helenius, A. Ganglioside-dependent cell attachment and endocytosis of murine polyomavirus-like particles. FEBS Lett. 2003, 555, 199–203. [Google Scholar] [CrossRef]
- Ewers, H.; Smith, A.E.; Sbalzarini, I.F.; Lilie, H.; Koumoutsakos, P.; Helenius, A. Single-particle tracking of murine polyoma virus-like particles on live cells and artificial membranes. Proc. Natl. Acad. Sci. USA 2005, 102, 15110–15115. [Google Scholar] [CrossRef] [PubMed]
- Piccinotti, S.; Kirchhausen, T.; Whelan, S.P. Uptake of rabies virus into epithelial cells by clathrin-mediated endocytosis depends upon actin. J. Virol. 2013, 87, 11637–11647. [Google Scholar] [CrossRef] [PubMed]
- Xu, H.; Hao, X.; Wang, S.; Wang, Z.; Cai, M.; Jiang, J.; Qin, Q.; Zhang, M.; Wang, H. Real-time imaging of rabies virus entry into living vero cells. Sci. Rep. 2015, 5, 11753. [Google Scholar] [CrossRef] [PubMed]
- Finke, S.; Brzozka, K.; Conzelmann, K.K. Tracking fluorescence-labeled rabies virus: Enhanced green fluorescent protein-tagged phosphoprotein p supports virus gene expression and formation of infectious particles. J. Virol. 2004, 78, 12333–12343. [Google Scholar] [CrossRef] [PubMed]
- Wan, X.Y.; Zheng, L.L.; Gao, P.F.; Yang, X.X.; Li, C.M.; Li, Y.F.; Huang, C.Z. Real-time light scattering tracking of gold nanoparticles-bioconjugated respiratory syncytial virus infecting HEp-2 cells. Sci. Rep. 2014, 4, 4529. [Google Scholar] [CrossRef] [PubMed]
- Helenius, A.; Kartenbeck, J.; Simons, K.; Fries, E. On the entry of semliki forest virus into BHK-21 cells. J. Cell Biol. 1980, 84, 404–420. [Google Scholar] [CrossRef] [PubMed]
- Vonderheit, A.; Helenius, A. Rab7 associates with early endosomes to mediate sorting and transport of semliki forest virus to late endosomes. PLoS Biol. 2005, 3, e233. [Google Scholar] [CrossRef] [PubMed]
- Jose, J.; Tang, J.; Taylor, A.B.; Baker, T.S.; Kuhn, R.J. Fluorescent protein-tagged sindbis virus E2 glycoprotein allows single particle analysis of virus budding from live cells. Viruses 2015, 7, 6182–6199. [Google Scholar] [CrossRef] [PubMed]
- Lozach, P.Y.; Mancini, R.; Bitto, D.; Meier, R.; Oestereich, L.; Overby, A.K.; Pettersson, R.F.; Helenius, A. Entry of bunyaviruses into mammalian cells. Cell Host Microbe 2010, 7, 488–499. [Google Scholar] [CrossRef] [PubMed]
- Ward, B.M.; Moss, B. Visualization of intracellular movement of vaccinia virus virions containing a green fluorescent protein-B5R membrane protein chimera. J. Virol. 2001, 75, 4802–4813. [Google Scholar] [CrossRef] [PubMed]
- Ward, B.M.; Moss, B. Vaccinia virus intracellular movement is associated with microtubules and independent of actin tails. J. Virol. 2001, 75, 11651–11663. [Google Scholar] [CrossRef] [PubMed]
- Rietdorf, J.; Ploubidou, A.; Reckmann, I.; Holmstrom, A.; Frischknecht, F.; Zettl, M.; Zimmermann, T.; Way, M. Kinesin-dependent movement on microtubules precedes actin-based motility of vaccinia virus. Nat. Cell Biol. 2001, 3, 992–1000. [Google Scholar] [CrossRef] [PubMed]
- Mercer, J.; Helenius, A. Vaccinia virus uses macropinocytosis and apoptotic mimicry to enter host cells. Science 2008, 320, 531–535. [Google Scholar] [CrossRef] [PubMed]
- Cureton, D.K.; Massol, R.H.; Saffarian, S.; Kirchhausen, T.L.; Whelan, S.P. Vesicular stomatitis virus enters cells through vesicles incompletely coated with clathrin that depend upon actin for internalization. PLoS. Pathog. 2009, 5, e1000394. [Google Scholar] [CrossRef] [PubMed]
- Cureton, D.K.; Massol, R.H.; Whelan, S.P.; Kirchhausen, T. The length of vesicular stomatitis virus particles dictates a need for actin assembly during clathrin-dependent endocytosis. PLoS. Pathog. 2010, 6, e1001127. [Google Scholar] [CrossRef] [PubMed]
- Makino, Y.; Suzuki, T.; Hasebe, R.; Kimura, T.; Maeda, A.; Takahashi, H.; Sawa, H. Establishment of tracking system for west nile virus entry and evidence of microtubule involvement in particle transport. J. Virol. Methods 2014, 195, 250–257. [Google Scholar] [CrossRef] [PubMed]
- Goodsell, D.S. Inside a living cell. Trends Biochem. Sci. 1991, 16, 203–206. [Google Scholar] [CrossRef]
- Lang, F. Mechanisms and significance of cell volume regulation. J. Am. Coll. Nutr. 2007, 26, 613S–623S. [Google Scholar] [CrossRef] [PubMed]
- Yakimovich, A.; Gumpert, H.; Burckhardt, C.J.; Lutschg, V.A.; Jurgeit, A.; Sbalzarini, I.F.; Greber, U.F. Cell-free transmission of human adenovirus by passive mass transfer in cell culture simulated in a computer model. J. Virol. 2012, 86, 10123–10137. [Google Scholar] [CrossRef] [PubMed]
- Yakimovich, A.; Yakimovich, Y.; Schmid, M.; Mercer, J.; Sbalzarini, I.F.; Greber, U.F. Infectio: A generic framework for computational simulation of virus transmission between cells. mSphere 2016, 1. [Google Scholar] [CrossRef] [PubMed]
- Olmsted, S.S.; Padgett, J.L.; Yudin, A.I.; Whaley, K.J.; Moench, T.R.; Cone, R.A. Diffusion of macromolecules and virus-like particles in human cervical mucus. Biophys. J. 2001, 81, 1930–1937. [Google Scholar] [CrossRef]
- Huang, Y.F.; Zhuo, G.Y.; Chou, C.Y.; Lin, C.H.; Chang, W.; Hsieh, C.L. Coherent brightfield microscopy provides the spatiotemporal resolution to study early stage viral infection in live cells. ACS Nano 2017, 11, 2575–2585. [Google Scholar] [CrossRef] [PubMed]
- Komarova, N.L.; Wodarz, D. ODE models for oncolytic virus dynamics. J. Theor. Biol. 2010, 263, 530–543. [Google Scholar] [CrossRef] [PubMed]
- Wodarz, D.; Komarova, N. Towards predictive computational models of oncolytic virus therapy: Basis for experimental validation and model selection. PLoS ONE 2009, 4, e4271. [Google Scholar] [CrossRef] [PubMed]
- Paiva, L.R.; Binny, C.; Ferreira, S.C., Jr.; Martins, M.L. A multiscale mathematical model for oncolytic virotherapy. Cancer Res. 2009, 69, 1205–1211. [Google Scholar] [CrossRef] [PubMed]
- Mercer, J.; Helenius, A. Virus entry by macropinocytosis. Nat. Cell Biol. 2009, 11, 510–520. [Google Scholar] [CrossRef] [PubMed]
- Pollard, T.D.; Cooper, J.A. Actin, a central player in cell shape and movement. Science 2009, 326, 1208–1212. [Google Scholar] [CrossRef] [PubMed]
- Greber, U.F. Signalling in viral entry. Cell. Mol. Life Sci. 2002, 59, 608–626. [Google Scholar] [CrossRef] [PubMed]
- Lin, C.H.; Espreafico, E.M.; Mooseker, M.S.; Forscher, P. Myosin drives retrograde F-actin flow in neuronal growth cones. Biol. Bull. 1997, 192, 183–185. [Google Scholar] [CrossRef] [PubMed]
- Medeiros, N.A.; Burnette, D.T.; Forscher, P. Myosin II functions in actin-bundle turnover in neuronal growth cones. Nat. Cell Biol. 2006, 8, 215–226. [Google Scholar] [CrossRef] [PubMed]
- Taylor, M.P.; Koyuncu, O.O.; Enquist, L.W. Subversion of the actin cytoskeleton during viral infection. Nat. Rev. Microbiol. 2011, 9, 427–439. [Google Scholar] [CrossRef] [PubMed]
- Welch, M.D.; Way, M. Arp2/3-mediated actin-based motility: A tail of pathogen abuse. Cell Host Microbe 2013, 14, 242–255. [Google Scholar] [CrossRef] [PubMed]
- Ohkawa, T.; Volkman, L.E.; Welch, M.D. Actin-based motility drives baculovirus transit to the nucleus and cell surface. J. Cell Biol. 2010, 190, 187–195. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.; Whittaker, G.R. Role of the actin cytoskeleton during influenza virus internalization into polarized epithelial cells. Cell. Microbiol. 2007, 9, 1672–1682. [Google Scholar] [CrossRef] [PubMed]
- Devadas, D.; Koithan, T.; Diestel, R.; Prank, U.; Sodeik, B.; Dohner, K. Herpes simplex virus internalization into epithelial cells requires Na+/H+ exchangers and p21-activated kinases but neither clathrin- nor caveolin-mediated endocytosis. J. Virol. 2014, 88, 13378–13395. [Google Scholar] [CrossRef] [PubMed]
- Newsome, T.P.; Scaplehorn, N.; Way, M. SRC mediates a switch from microtubule- to actin-based motility of vaccinia virus. Science 2004, 306, 124–129. [Google Scholar] [CrossRef] [PubMed]
- Leite, F.; Way, M. The role of signalling and the cytoskeleton during vaccinia virus egress. Virus Res. 2015, 209, 87–99. [Google Scholar] [CrossRef] [PubMed]
- Doceul, V.; Hollinshead, M.; van der Linden, L.; Smith, G.L. Repulsion of superinfecting virions: A mechanism for rapid virus spread. Science 2010, 327, 873–876. [Google Scholar] [CrossRef] [PubMed]
- Smith, G.L.; Law, M. The exit of vaccinia virus from infected cells. Virus Res. 2004, 106, 189–197. [Google Scholar] [CrossRef] [PubMed]
- Cudmore, S.; Cossart, P.; Griffiths, G.; Way, M. Actin-based motility of vaccinia virus. Nature 1995, 378, 636–638. [Google Scholar] [CrossRef] [PubMed]
- Schudt, G.; Kolesnikova, L.; Dolnik, O.; Sodeik, B.; Becker, S. Live-cell imaging of marburg virus-infected cells uncovers actin-dependent transport of nucleocapsids over long distances. Proc. Natl. Acad. Sci. USA 2013, 110, 14402–14407. [Google Scholar] [CrossRef] [PubMed]
- Schudt, G.; Dolnik, O.; Kolesnikova, L.; Biedenkopf, N.; Herwig, A.; Becker, S. Transport of ebolavirus nucleocapsids is dependent on actin polymerization: Live-cell imaging analysis of ebolavirus-infected cells. J. Infect. Dis. 2015, 212 (Suppl. S2), S160–S166. [Google Scholar] [CrossRef] [PubMed]
- Adu-Gyamfi, E.; Digman, M.A.; Gratton, E.; Stahelin, R.V. Single-particle tracking demonstrates that actin coordinates the movement of the ebola virus matrix protein. Biophys. J. 2012, 103, L41–L43. [Google Scholar] [CrossRef] [PubMed]
- Kolesnikova, L.; Bohil, A.B.; Cheney, R.E.; Becker, S. Budding of marburgvirus is associated with filopodia. Cell. Microbiol. 2007, 9, 939–951. [Google Scholar] [CrossRef] [PubMed]
- Kirschner, M.; Mitchison, T. Beyond self-assembly: From microtubules to morphogenesis. Cell 1986, 45, 329–342. [Google Scholar] [CrossRef]
- Welte, M.A. Bidirectional transport along microtubules. Curr. Biol. 2004, 14, R525–R537. [Google Scholar] [CrossRef] [PubMed]
- Dodding, M.P.; Way, M. Coupling viruses to dynein and kinesin-1. EMBO J. 2011, 30, 3527–3539. [Google Scholar] [CrossRef] [PubMed]
- Frischknecht, F.; Way, M. Surfing pathogens and the lessons learned for actin polymerization. Trends Cell Biol. 2001, 11, 30–38. [Google Scholar] [CrossRef]
- Ploubidou, A.; Way, M. Viral transport and the cytoskeleton. Curr. Opin. Cell Biol. 2001, 13, 97–105. [Google Scholar] [CrossRef]
- Radtke, K.; Dohner, K.; Sodeik, B. Viral interactions with the cytoskeleton: A hitchhiker’s guide to the cell. Cell. Microbiol. 2006, 8, 387–400. [Google Scholar] [CrossRef] [PubMed]
- Smith, G.A.; Enquist, L.W. Break ins and break outs: Viral interactions with the cytoskeleton of mammalian cells. Annu. Rev. Cell Dev. Biol. 2002, 18, 135–161. [Google Scholar] [CrossRef] [PubMed]
- Ward, B.M. The taking of the cytoskeleton one two three: How viruses utilize the cytoskeleton during egress. Virology 2011, 411, 244–250. [Google Scholar] [CrossRef] [PubMed]
- Naghavi, M.H.; Walsh, D. Microtubule regulation and function during virus infection. J. Virol. 2017. [Google Scholar] [CrossRef] [PubMed]
- Roll-Mecak, A. Intrinsically disordered tubulin tails: Complex tuners of microtubule functions? Semin. Cell Dev. Biol. 2015, 37, 11–19. [Google Scholar] [CrossRef] [PubMed]
- Janke, C. The tubulin code: Molecular components, readout mechanisms, and functions. J. Cell Biol. 2014, 206, 461–472. [Google Scholar] [CrossRef] [PubMed]
- Verhey, K.J.; Gaertig, J. The tubulin code. Cell Cycle 2007, 6, 2152–2160. [Google Scholar] [CrossRef] [PubMed]
- Malikov, V.; da Silva, E.S.; Jovasevic, V.; Bennett, G.; de Souza Aranha Vieira, D.A.; Schulte, B.; Diaz-Griffero, F.; Walsh, D.; Naghavi, M.H. HIV-1 capsids bind and exploit the kinesin-1 adaptor FEZ1 for inward movement to the nucleus. Nat. Commun. 2015, 6, 6660. [Google Scholar] [CrossRef] [PubMed]
- Jovasevic, V.; Naghavi, M.H.; Walsh, D. Microtubule plus end-associated CLIP-170 initiates HSV-1 retrograde transport in primary human cells. J. Cell Biol. 2015, 211, 323–337. [Google Scholar] [CrossRef] [PubMed]
- Sabo, Y.; Walsh, D.; Barry, D.S.; Tinaztepe, S.; de Los Santos, K.; Goff, S.P.; Gundersen, G.G.; Naghavi, M.H. HIV-1 induces the formation of stable microtubules to enhance early infection. Cell Host Microbe 2013, 14, 535–546. [Google Scholar] [CrossRef] [PubMed]
- Warren, J.C.; Rutkowski, A.; Cassimeris, L. Infection with replication-deficient adenovirus induces changes in the dynamic instability of host cell microtubules. Mol. Biol. Cell 2006, 17, 3557–3568. [Google Scholar] [CrossRef] [PubMed]
- Naranatt, P.P.; Krishnan, H.H.; Smith, M.S.; Chandran, B. Kaposi’s sarcoma-associated herpesvirus modulates microtubule dynamics via rhoa-gtp-diaphanous 2 signaling and utilizes the dynein motors to deliver its DNA to the nucleus. J. Virol. 2005, 79, 1191–1206. [Google Scholar] [CrossRef] [PubMed]
- Naghavi, M.H.; Valente, S.; Hatziioannou, T.; de Los Santos, K.; Wen, Y.; Mott, C.; Gundersen, G.G.; Goff, S.P. Moesin regulates stable microtubule formation and limits retroviral infection in cultured cells. EMBO J. 2007, 26, 41–52. [Google Scholar] [CrossRef] [PubMed]
- Husain, M.; Harrod, K.S. Enhanced acetylation of alpha-tubulin in influenza a virus infected epithelial cells. FEBS Lett. 2011, 585, 128–132. [Google Scholar] [CrossRef] [PubMed]
- Valenzuela-Fernandez, A.; Alvarez, S.; Gordon-Alonso, M.; Barrero, M.; Ursa, A.; Cabrero, J.R.; Fernandez, G.; Naranjo-Suarez, S.; Yanez-Mo, M.; Serrador, J.M.; et al. Histone deacetylase 6 regulates human immunodeficiency virus type 1 infection. Mol. Biol. Cell 2005, 16, 5445–5454. [Google Scholar] [CrossRef] [PubMed]
- Elliott, G.; O’Hare, P. Herpes simplex virus type 1 tegument protein VP22 induces the stabilization and hyperacetylation of microtubules. J. Virol. 1998, 72, 6448–6455. [Google Scholar] [PubMed]
- Ploubidou, A.; Moreau, V.; Ashman, K.; Reckmann, I.; Gonzalez, C.; Way, M. Vaccinia virus infection disrupts microtubule organization and centrosome function. EMBO J. 2000, 19, 3932–3944. [Google Scholar] [CrossRef] [PubMed]
- Naghavi, M.H.; Gundersen, G.G.; Walsh, D. Plus-end tracking proteins, CLASPs, and a viral Akt mimic regulate herpesvirus-induced stable microtubule formation and virus spread. Proc. Natl. Acad. Sci. USA 2013, 110, 18268–18273. [Google Scholar] [CrossRef] [PubMed]
- Nishi, M.; Ryo, A.; Tsurutani, N.; Ohba, K.; Sawasaki, T.; Morishita, R.; Perrem, K.; Aoki, I.; Morikawa, Y.; Yamamoto, N. Requirement for microtubule integrity in the SOCS1-mediated intracellular dynamics of HIV-1 Gag. FEBS Lett. 2009, 583, 1243–1250. [Google Scholar] [CrossRef] [PubMed]
- Luby-Phelps, K. Cytoarchitecture and physical properties of cytoplasm: Volume, viscosity, diffusion, intracellular surface area. Int. Rev. Cytol. 2000, 192, 189–221. [Google Scholar] [PubMed]
- Luisoni, S.; Suomalainen, M.; Boucke, K.; Tanner, L.B.; Wenk, M.R.; Guan, X.L.; Grzybek, M.; Coskun, U.; Greber, U.F. Co-option of membrane wounding enables virus penetration into cells. Cell Host Microbe 2015, 18, 75–85. [Google Scholar] [CrossRef] [PubMed]
- Luisoni, S.; Bauer, M.; Prasad, V.; Boucke, K.; Papadopoulos, C.; Meyer, H.; Hemmi, S.; Suomalainen, M.; Greber, U.F. Endosomophagy clears disrupted early endosomes but not virus particles during virus entry into cells. Matters 2016. [Google Scholar] [CrossRef]
- Stichling, N.; Suomalainen, M.; Flatt, J.W.; Schmid, M.; Pacesa, M.; Hemmi, S.; Jungraithmayr, W.; Maler, M.D.; Freudenberg, M.A.; Pluckthun, A.; et al. Lung macrophage scavenger receptor SR-A6 (MARCO) is an adenovirus type-specific virus entry receptor. PLoS. Pathog. 2018, 14, e1006914. [Google Scholar] [CrossRef] [PubMed]
- Suomalainen, M.; Nakano, M.Y.; Boucke, K.; Keller, S.; Greber, U.F. Adenovirus-activated PKA and p38/MAPK pathways boost microtubule-mediated nuclear targeting of virus. EMBO J. 2001, 20, 1310–1319. [Google Scholar] [CrossRef] [PubMed]
- Engelke, M.F.; Burckhardt, C.J.; Morf, M.K.; Greber, U.F. The dynactin complex enhances the speed of microtubule-dependent motions of adenovirus both towards and away from the nucleus. Viruses 2011, 3, 233–253. [Google Scholar] [CrossRef] [PubMed]
- Hirokawa, N.; Niwa, S.; Tanaka, Y. Molecular motors in neurons: Transport mechanisms and roles in brain function, development, and disease. Neuron 2010, 68, 610–638. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.J. Regulation of dynein-dynactin-driven vesicular transport. Traffic 2017, 18, 336–347. [Google Scholar] [CrossRef] [PubMed]
- Wang, I.H.; Burckhardt, C.J.; Yakimovich, A.; Morf, M.K.; Greber, U.F. The nuclear export factor CRM1 controls juxta-nuclear microtubule-dependent virus transport. J. Cell Sci. 2017, 130, 2185–2195. [Google Scholar] [CrossRef] [PubMed]
- Trotman, L.C.; Mosberger, N.; Fornerod, M.; Stidwill, R.P.; Greber, U.F. Import of adenovirus DNA involves the nuclear pore complex receptor CAN/Nup214 and histone H1. Nat. Cell Biol. 2001, 3, 1092–1100. [Google Scholar] [CrossRef] [PubMed]
- Strunze, S.; Trotman, L.C.; Boucke, K.; Greber, U.F. Nuclear targeting of adenovirus type 2 requires CRM1-mediated nuclear export. Mol. Biol. Cell 2005, 16, 2999–3009. [Google Scholar] [CrossRef] [PubMed]
- Banerjee, I.; Miyake, Y.; Nobs, S.P.; Schneider, C.; Horvath, P.; Kopf, M.; Matthias, P.; Helenius, A.; Yamauchi, Y. Influenza a virus uses the aggresome processing machinery for host cell entry. Science 2014, 346, 473–477. [Google Scholar] [CrossRef] [PubMed]
- Ravindran, M.S.; Engelke, M.F.; Verhey, K.J.; Tsai, B. Exploiting the kinesin-1 molecular motor to generate a virus membrane penetration site. Nat. Commun. 2017, 8, 15496. [Google Scholar] [CrossRef] [PubMed]
- Reed, N.A.; Cai, D.; Blasius, T.L.; Jih, G.T.; Meyhofer, E.; Gaertig, J.; Verhey, K.J. Microtubule acetylation promotes kinesin-1 binding and transport. Curr. Biol. 2006, 16, 2166–2172. [Google Scholar] [CrossRef] [PubMed]
- Konishi, Y.; Setou, M. Tubulin tyrosination navigates the kinesin-1 motor domain to axons. Nat Neurosci. 2009, 12, 559–567. [Google Scholar] [CrossRef] [PubMed]
- Pawlica, P.; Le Sage, V.; Poccardi, N.; Tremblay, M.J.; Mouland, A.J.; Berthoux, L. Functional evidence for the involvement of microtubules and dynein motor complexes in TRIM5alpha-mediated restriction of retroviruses. J. Virol. 2014, 88, 5661–5676. [Google Scholar] [CrossRef] [PubMed]
- Suomalainen, M.; Greber, U.F. Uncoating of non-enveloped viruses. Curr. Opin. Virol. 2013, 3, 27–33. [Google Scholar] [CrossRef] [PubMed]
- Smith, G.L.; Murphy, B.J.; Law, M. Vaccinia virus motility. Annu. Rev. Microbiol. 2003, 57, 323–342. [Google Scholar] [CrossRef] [PubMed]
- Zaichick, S.V.; Bohannon, K.P.; Smith, G.A. Alphaherpesviruses and the cytoskeleton in neuronal infections. Viruses 2011, 3, 941–981. [Google Scholar] [CrossRef] [PubMed]
- Lichtman, J.W.; Conchello, J.A. Fluorescence microscopy. Nat. Methods 2005, 2, 910–919. [Google Scholar] [CrossRef] [PubMed]
- Axelrod, D. Chapter 7: Total internal reflection fluorescence microscopy. Methods Cell Biol. 2008, 89, 169–221. [Google Scholar] [PubMed]
- Ewers, H.; Schelhaas, M. Analysis of virus entry and cellular membrane dynamics by single particle tracking. Methods Enzymol. 2012, 506, 63–80. [Google Scholar] [PubMed]
- Inoue, S. Foundations of confocal scanned imaging in light microscopy. In Handbook of Biological Confocal Microscopy, 3rd ed.; Pawley, J.B., Ed.; Springer Science & Business Media, LLC: New York, NY, USA, 2006. [Google Scholar]
- Graf, R.; Rietdorf, J.; Zimmermann, T. Live cell spinning disk microscopy. Adv. Biochem. Eng. Biotechnol. 2005, 95, 57–75. [Google Scholar] [PubMed]
- Oreopoulos, J.; Berman, R.; Browne, M. Spinning-disk confocal microscopy: Present technology and future trends. Methods Cell Biol. 2014, 123, 153–175. [Google Scholar] [PubMed]
- Helmuth, J.A.; Burckhardt, C.J.; Koumoutsakos, P.; Greber, U.F.; Sbalzarini, I.F. A novel supervised trajectory segmentation algorithm identifies distinct types of human adenovirus motion in host cells. J. Struct. Biol. 2007, 159, 347–358. [Google Scholar] [CrossRef] [PubMed]
- Huang, B.; Babcock, H.; Zhuang, X. Breaking the diffraction barrier: Super-resolution imaging of cells. Cell 2010, 143, 1047–1058. [Google Scholar] [CrossRef] [PubMed]
- Hell, S.W.; Wichmann, J. Breaking the diffraction resolution limit by stimulated emission: Stimulated-emission-depletion fluorescence microscopy. Opt. Lett. 1994, 19, 780–782. [Google Scholar] [CrossRef] [PubMed]
- Gustafsson, M.G. Nonlinear structured-illumination microscopy: Wide-field fluorescence imaging with theoretically unlimited resolution. Proc. Natl. Acad. Sci. USA 2005, 102, 13081–13086. [Google Scholar] [CrossRef] [PubMed]
- Rust, M.J.; Bates, M.; Zhuang, X. Sub-diffraction-limit imaging by stochastic optical reconstruction microscopy (storm). Nat. Methods 2006, 3, 793–795. [Google Scholar] [CrossRef] [PubMed]
- Betzig, E.; Patterson, G.H.; Sougrat, R.; Lindwasser, O.W.; Olenych, S.; Bonifacino, J.S.; Davidson, M.W.; Lippincott-Schwartz, J.; Hess, H.F. Imaging intracellular fluorescent proteins at nanometer resolution. Science 2006, 313, 1642–1645. [Google Scholar] [CrossRef] [PubMed]
- Willig, K.I.; Kellner, R.R.; Medda, R.; Hein, B.; Jakobs, S.; Hell, S.W. Nanoscale resolution in GFP-based microscopy. Nat. Methods 2006, 3, 721–723. [Google Scholar] [CrossRef] [PubMed]
- Manley, S.; Gillette, J.M.; Patterson, G.H.; Shroff, H.; Hess, H.F.; Betzig, E.; Lippincott-Schwartz, J. High-density mapping of single-molecule trajectories with photoactivated localization microscopy. Nat. Methods 2008, 5, 155–157. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.; Deerinck, T.J.; Sigal, Y.M.; Babcock, H.P.; Ellisman, M.H.; Zhuang, X. Correlative stochastic optical reconstruction microscopy and electron microscopy. PLoS ONE 2015, 10, e0124581. [Google Scholar] [CrossRef] [PubMed]
- Stelzer, E.H. Light-sheet fluorescence microscopy for quantitative biology. Nat. Methods 2015, 12, 23–26. [Google Scholar] [CrossRef] [PubMed]
- Bosse, J.B.; Hogue, I.B.; Feric, M.; Thiberge, S.Y.; Sodeik, B.; Brangwynne, C.P.; Enquist, L.W. Remodeling nuclear architecture allows efficient transport of herpesvirus capsids by diffusion. Proc. Natl. Acad. Sci. USA 2015, 112, E5725–5733. [Google Scholar] [CrossRef] [PubMed]
- Hoyer, P.; de Medeiros, G.; Balazs, B.; Norlin, N.; Besir, C.; Hanne, J.; Krausslich, H.G.; Engelhardt, J.; Sahl, S.J.; Hell, S.W.; et al. Breaking the diffraction limit of light-sheet fluorescence microscopy by resolft. Proc. Natl. Acad. Sci. USA 2016, 113, 3442–3446. [Google Scholar] [CrossRef] [PubMed]
- Gorelik, J.; Shevchuk, A.; Ramalho, M.; Elliott, M.; Lei, C.; Higgins, C.F.; Lab, M.J.; Klenerman, D.; Krauzewicz, N.; Korchev, Y. Scanning surface confocal microscopy for simultaneous topographical and fluorescence imaging: Application to single virus-like particle entry into a cell. Proc. Natl. Acad. Sci. USA 2002, 99, 16018–16023. [Google Scholar] [CrossRef] [PubMed]
- Binnig, G.; Quate, C.F.; Gerber, C. Atomic force microscope. Phys. Rev. Lett. 1986, 56, 930–933. [Google Scholar] [CrossRef] [PubMed]
- Pan, Y.; Wang, S.; Shan, Y.; Zhang, D.; Gao, J.; Zhang, M.; Liu, S.; Cai, M.; Xu, H.; Li, G.; et al. Ultrafast tracking of a single live virion during the invagination of a cell membrane. Small 2015, 11, 2782–2788. [Google Scholar] [CrossRef] [PubMed]
- Ewers, H.; Jacobsen, V.; Klotzsch, E.; Smith, A.E.; Helenius, A.; Sandoghdar, V. Label-free optical detection and tracking of single virions bound to their receptors in supported membrane bilayers. Nano Lett. 2007, 7, 2263–2266. [Google Scholar] [CrossRef] [PubMed]
- Larson, D.R.; Johnson, M.C.; Webb, W.W.; Vogt, V.M. Visualization of retrovirus budding with correlated light and electron microscopy. Proc. Natl. Acad. Sci. USA 2005, 102, 15453–15458. [Google Scholar] [CrossRef] [PubMed]
- Morgan, C.; Godman, G.C.; Breitenfeld, P.M.; Rose, H.M. A correlative study by electron and light microscopy of the development of type 5 adenovirus. I. Electron microscopy. J. Exp. Med. 1960, 112, 373–382. [Google Scholar] [CrossRef] [PubMed]
- Godman, G.C.; Morgan, C.; Breitenfeld, P.M.; Rose, H.M. A correlative study by electron and light microscopy of the development of type 5 adenovirus. II. Light microscopy. J. Exp. Med. 1960, 112, 383–402. [Google Scholar] [CrossRef] [PubMed]
- Schellenberger, P.; Kaufmann, R.; Siebert, C.A.; Hagen, C.; Wodrich, H.; Grunewald, K. High-precision correlative fluorescence and electron cryo microscopy using two independent alignment markers. Ultramicroscopy 2014, 143, 41–51. [Google Scholar] [CrossRef] [PubMed]
- Hell, S.W. Far-field optical nanoscopy. Science 2007, 316, 1153–1158. [Google Scholar] [CrossRef] [PubMed]
- Matthews, R.E. A classification of virus groups based on the size of the particle in relation to genome size. J. Gen. Virol. 1975, 27, 135–149. [Google Scholar] [CrossRef] [PubMed]
- Tsien, R.Y. The green fluorescent protein. Annu. Rev. Biochem. 1998, 67, 509–544. [Google Scholar] [CrossRef] [PubMed]
- Kamiyama, D.; Sekine, S.; Barsi-Rhyne, B.; Hu, J.; Chen, B.; Gilbert, L.A.; Ishikawa, H.; Leonetti, M.D.; Marshall, W.F.; Weissman, J.S.; et al. Versatile protein tagging in cells with split fluorescent protein. Nat. Commun. 2016, 7, 11046. [Google Scholar] [CrossRef] [PubMed]
- Kelkar, S.A.; Pfister, K.K.; Crystal, R.G.; Leopold, P.L. Cytoplasmic dynein mediates adenovirus binding to microtubules. J. Virol. 2004, 78, 10122–10132. [Google Scholar] [CrossRef] [PubMed]
- Schelhaas, M.; Shah, B.; Holzer, M.; Blattmann, P.; Kuhling, L.; Day, P.M.; Schiller, J.T.; Helenius, A. Entry of human papillomavirus type 16 by actin-dependent, clathrin- and lipid raft-independent endocytosis. PLoS. Pathog. 2012, 8, e1002657. [Google Scholar] [CrossRef] [PubMed]
- Bienkowska-Haba, M.; Sapp, M. The cytoskeleton in papillomavirus infection. Viruses 2011, 3, 260–271. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.J.; Xia, L.; Liu, S.L.; Sun, E.Z.; Wu, Q.M.; Wen, L.; Zhang, Z.L.; Pang, D.W. A “driver switchover” mechanism of influenza virus transport from microfilaments to microtubules. ACS Nano 2017, 12, 474–484. [Google Scholar] [CrossRef] [PubMed]
- Nagel, C.H.; Dohner, K.; Fathollahy, M.; Strive, T.; Borst, E.M.; Messerle, M.; Sodeik, B. Nuclear egress and envelopment of herpes simplex virus capsids analyzed with dual-color fluorescence HSV1(17+). J. Virol. 2008, 82, 3109–3124. [Google Scholar] [CrossRef] [PubMed]
- Dohner, K.; Radtke, K.; Schmidt, S.; Sodeik, B. Eclipse phase of herpes simplex virus type 1 infection: Efficient dynein-mediated capsid transport without the small capsid protein vp26. J. Virol. 2006, 80, 8211–8224. [Google Scholar] [CrossRef] [PubMed]
- Douglas, M.W.; Diefenbach, R.J.; Homa, F.L.; Miranda-Saksena, M.; Rixon, F.J.; Vittone, V.; Byth, K.; Cunningham, A.L. Herpes simplex virus type 1 capsid protein VP26 interacts with dynein light chains RP3 and Tctex1 and plays a role in retrograde cellular transport. J. Biol. Chem. 2004, 279, 28522–28530. [Google Scholar] [CrossRef] [PubMed]
- Diefenbach, R.J.; Miranda-Saksena, M.; Diefenbach, E.; Holland, D.J.; Boadle, R.A.; Armati, P.J.; Cunningham, A.L. Herpes simplex virus tegument protein us11 interacts with conventional kinesin heavy chain. J. Virol. 2002, 76, 3282–3291. [Google Scholar] [CrossRef] [PubMed]
- Arhel, N.; Genovesio, A.; Kim, K.A.; Miko, S.; Perret, E.; Olivo-Marin, J.C.; Shorte, S.; Charneau, P. Quantitative four-dimensional tracking of cytoplasmic and nuclear HIV-1 complexes. Nat. Methods 2006, 3, 817–824. [Google Scholar] [CrossRef] [PubMed]
- Babcock, H.P.; Chen, C.; Zhuang, X. Using single-particle tracking to study nuclear trafficking of viral genes. Biophys. J. 2004, 87, 2749–2758. [Google Scholar] [CrossRef] [PubMed]
- Bailey, B.; Farkas, D.L.; Taylor, D.L.; Lanni, F. Enhancement of axial resolution in fluorescence microscopy by standing-wave excitation. Nature 1993, 366, 44–48. [Google Scholar] [CrossRef] [PubMed]
- York, A.G.; Chandris, P.; Nogare, D.D.; Head, J.; Wawrzusin, P.; Fischer, R.S.; Chitnis, A.; Shroff, H. Instant super-resolution imaging in live cells and embryos via analog image processing. Nat. Methods 2013, 10, 1122–1126. [Google Scholar] [CrossRef] [PubMed]
- Gray, R.D.; Beerli, C.; Pereira, P.M.; Scherer, K.M.; Samolej, J.; Bleck, C.K.; Mercer, J.; Henriques, R. Virusmapper: Open-source nanoscale mapping of viral architecture through super-resolution microscopy. Sci. Rep. 2016, 6, 29132. [Google Scholar] [CrossRef] [PubMed]
- Ymeti, A.; Greve, J.; Lambeck, P.V.; Wink, T.; van Hovell, S.W.; Beumer, T.A.; Wijn, R.R.; Heideman, R.G.; Subramaniam, V.; Kanger, J.S. Fast, ultrasensitive virus detection using a young interferometer sensor. Nano Lett. 2007, 7, 394–397. [Google Scholar] [CrossRef] [PubMed]
- Herrmann, A.; Sieben, C. Single-virus force spectroscopy unravels molecular details of virus infection. Integr. Biol. Quant. Biosci. Nano Macro 2015, 7, 620–632. [Google Scholar] [CrossRef] [PubMed]
- Pang, H.B.; Hevroni, L.; Kol, N.; Eckert, D.M.; Tsvitov, M.; Kay, M.S.; Rousso, I. Virion stiffness regulates immature HIV-1 entry. Retrovirology 2013, 10, 4. [Google Scholar] [CrossRef] [PubMed]
- Ortega-Esteban, A.; Condezo, G.N.; Perez-Berna, A.J.; Chillon, M.; Flint, S.J.; Reguera, D.; San Martin, C.; de Pablo, P.J. Mechanics of viral chromatin reveals the pressurization of human adenovirus. ACS Nano 2015, 9, 10826–10833. [Google Scholar] [CrossRef] [PubMed]
- Ortega-Esteban, A.; Bodensiek, K.; San Martin, C.; Suomalainen, M.; Greber, U.F.; de Pablo, P.J.; Schaap, I.A. Fluorescence tracking of genome release during mechanical unpacking of single viruses. ACS Nano 2015, 9, 10571–10579. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Sieben, C.; Ludwig, K.; Hofer, C.T.; Chiantia, S.; Herrmann, A.; Eghiaian, F.; Schaap, I.A. pH-controlled two-step uncoating of influenza virus. Biophys. J. 2014, 106, 1447–1456. [Google Scholar] [CrossRef] [PubMed]
- Snijder, J.; Radtke, K.; Anderson, F.; Scholtes, L.; Corradini, E.; Baines, J.; Heck, A.J.R.; Wuite, G.J.L.; Sodeik, B.; Roos, W.H. Vertex-specific proteins pUL7 and pUL125 mechanically reinforce herpes simplex virus capsids. J. Virol. 2017, 91, e00123-17. [Google Scholar] [CrossRef] [PubMed]
- Snijder, J.; Reddy, V.S.; May, E.R.; Roos, W.H.; Nemerow, G.R.; Wuite, G.J. Integrin and defensin modulate the mechanical properties of adenovirus. J. Virol. 2013, 87, 2756–2766. [Google Scholar] [CrossRef] [PubMed]
- Alsteens, D.; Newton, R.; Schubert, R.; Martinez-Martin, D.; Delguste, M.; Roska, B.; Muller, D.J. Nanomechanical mapping of first binding steps of a virus to animal cells. Nat. Nanotechnol. 2017, 12, 177–183. [Google Scholar] [CrossRef] [PubMed]
- Saxton, M.J.; Jacobson, K. Single-particle tracking: Applications to membrane dynamics. Annu. Rev. Biophys. Biomol. Struct. 1997, 26, 373–399. [Google Scholar] [CrossRef] [PubMed]
- Manzo, C.; Garcia-Parajo, M.F. A review of progress in single particle tracking: From methods to biophysical insights. Rep. Prog. Phys. 2015, 78, 124601. [Google Scholar] [CrossRef] [PubMed]
- Chenouard, N.; Dufour, A.; Olivo-Marin, J.C. Tracking algorithms chase down pathogens. Biotechnol. J. 2009, 4, 838–845. [Google Scholar] [CrossRef] [PubMed]
- Sbalzarini, I.F.; Koumoutsakos, P. Feature point tracking and trajectory analysis for video imaging in cell biology. J. Struct. Biol. 2005, 151, 182–195. [Google Scholar] [CrossRef] [PubMed]
- Gazzola, M.; Burckhardt, C.J.; Bayati, B.; Engelke, M.; Greber, U.F.; Koumoutsakos, P. A stochastic model for microtubule motors describes the in vivo cytoplasmic transport of human adenovirus. PLoS Comput. Biol. 2009, 5, e1000623. [Google Scholar] [CrossRef] [PubMed]
- Jaqaman, K.; Loerke, D.; Mettlen, M.; Kuwata, H.; Grinstein, S.; Schmid, S.L.; Danuser, G. Robust single-particle tracking in live-cell time-lapse sequences. Nat. Methods 2008, 5, 695–702. [Google Scholar] [CrossRef] [PubMed]
- Aguet, F.; Antonescu, C.N.; Mettlen, M.; Schmid, S.L.; Danuser, G. Advances in analysis of low signal-to-noise images link dynamin and AP2 to the functions of an endocytic checkpoint. Dev. Cell 2013, 26, 279–291. [Google Scholar] [CrossRef] [PubMed]
- Chenouard, N.; Smal, I.; de Chaumont, F.; Maska, M.; Sbalzarini, I.F.; Gong, Y.; Cardinale, J.; Carthel, C.; Coraluppi, S.; Winter, M.; et al. Objective comparison of particle tracking methods. Nat. Methods 2014, 11, 281–289. [Google Scholar] [CrossRef] [PubMed]
- Godinez, W.J.; Lampe, M.; Worz, S.; Muller, B.; Eils, R.; Rohr, K. Deterministic and probabilistic approaches for tracking virus particles in time-lapse fluorescence microscopy image sequences. Med. Image Anal. 2009, 13, 325–342. [Google Scholar] [CrossRef] [PubMed]
- Coraluppi, S.; Carthel, C. Multi-stage multiple-hypothesis tracking. J. Adv. Inf. Fusion 2011, 6, 57–67. [Google Scholar]
- Roudot, P.; Ding, L.; Jaqaman, K.; Kervrann, C.; Danuser, G. Piecewise-stationary motion modeling and iterative smoothing to track heterogeneous particle motions in dense environments. IEEE Trans. Image Process. 2017, 26, 5395–5410. [Google Scholar] [CrossRef] [PubMed]
- Welf, E.S.; Driscoll, M.K.; Dean, K.M.; Schafer, C.; Chu, J.; Davidson, M.W.; Lin, M.Z.; Danuser, G.; Fiolka, R. Quantitative multiscale cell imaging in controlled 3D microenvironments. Dev. Cell 2016, 36, 462–475. [Google Scholar] [CrossRef] [PubMed]
- Keller, P.J. Imaging morphogenesis: Technological advances and biological insights. Science 2013, 340, 1234168. [Google Scholar] [CrossRef] [PubMed]
- Dean, K.M.; Roudot, P.; Welf, E.S.; Danuser, G.; Fiolka, R. Deconvolution-free subcellular imaging with axially swept light sheet microscopy. Biophys. J. 2015, 108, 2807–2815. [Google Scholar] [CrossRef] [PubMed]
- Lee, D.W.; Hsu, H.L.; Bacon, K.B.; Daniel, S. Image restoration and analysis of influenza virions binding to membrane receptors reveal adhesion-strengthening kinetics. PLoS ONE 2016, 11, e0163437. [Google Scholar] [CrossRef] [PubMed]
- Rigano, A.; Galli, V.; Clark, J.M.; Pereira, L.E.; Grossi, L.; Luban, J.; Giulietti, R.; Leidi, T.; Hunter, E.; Valle, M.; et al. Omega: A software tool for the management, analysis, and dissemination of intracellular trafficking data that incorporates motion type classification and quality control. bioRxiv 2018. [Google Scholar] [CrossRef]
- Applegate, K.T.; Besson, S.; Matov, A.; Bagonis, M.H.; Jaqaman, K.; Danuser, G. Plustiptracker: Quantitative image analysis software for the measurement of microtubule dynamics. J. Struct. Biol. 2011, 176, 168–184. [Google Scholar] [CrossRef] [PubMed]
- Ng, M.R.; Besser, A.; Danuser, G.; Brugge, J.S. Substrate stiffness regulates cadherin-dependent collective migration through myosin-II contractility. J. Cell Biol. 2012, 199, 545–563. [Google Scholar] [CrossRef] [PubMed]
- Rink, J.; Ghigo, E.; Kalaidzidis, Y.; Zerial, M. Rab conversion as a mechanism of progression from early to late endosomes. Cell 2005, 122, 735–749. [Google Scholar] [CrossRef] [PubMed]
- Tinevez, J.Y.; Perry, N.; Schindelin, J.; Hoopes, G.M.; Reynolds, G.D.; Laplantine, E.; Bednarek, S.Y.; Shorte, S.L.; Eliceiri, K.W. Trackmate: An open and extensible platform for single-particle tracking. Methods 2017, 115, 80–90. [Google Scholar] [CrossRef] [PubMed]
- De Chaumont, F.; Dallongeville, S.; Chenouard, N.; Herve, N.; Pop, S.; Provoost, T.; Meas-Yedid, V.; Pankajakshan, P.; Lecomte, T.; Le Montagner, Y.; et al. Icy: An open bioimage informatics platform for extended reproducible research. Nat. Methods 2012, 9, 690–696. [Google Scholar] [CrossRef] [PubMed]
- Vallotton, P.; van Oijen, A.M.; Whitchurch, C.B.; Gelfand, V.; Yeo, L.; Tsiavaliaris, G.; Heinrich, S.; Dultz, E.; Weis, K.; Grunwald, D. Diatrack particle tracking software: Review of applications and performance evaluation. Traffic 2017, 18, 840–852. [Google Scholar] [CrossRef] [PubMed]
- Holzwarth, G.; Bonin, K.; Hill, D.B. Forces required of kinesin during processive transport through cytoplasm. Biophys. J. 2002, 82, 1784–1790. [Google Scholar] [CrossRef]
- Michalet, X. Mean square displacement analysis of single-particle trajectories with localization error: Brownian motion in an isotropic medium. Phys. Rev. E Stat. Nonlinear Soft Matter Phys. 2010, 82, 41914. [Google Scholar] [CrossRef] [PubMed]
- Qian, H.; Sheetz, M.P.; Elson, E.L. Single particle tracking. Analysis of diffusion and flow in two-dimensional systems. Biophys. J. 1991, 60, 910–921. [Google Scholar] [CrossRef]
- Morrison, I.E.; Anderson, C.M.; Georgiou, G.N.; Cherry, R.J. Measuring diffusion coefficients of labelled particles on cell surfaces by digital fluorescence microscopy. Biochem. Soc. Trans. 1990, 18, 938. [Google Scholar] [CrossRef] [PubMed]
- Ferrari, R.; Manfroi, A.J.; Young, W.R. Strongly and weakly self-similar diffusion. Physica D 2001, 154, 111137. [Google Scholar] [CrossRef]
- Siebrasse, J.P.; Djuric, I.; Schulze, U.; Schluter, M.A.; Pavenstadt, H.; Weide, T.; Kubitscheck, U. Trajectories and single-particle tracking data of intracellular vesicles loaded with either SNAP-Crb3A or SNAP-Crb3B. Data Brief 2016, 7, 1665–1669. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Saxton, M.J. Lateral diffusion in an archipelago. Single-particle diffusion. Biophys. J. 1993, 64, 1766–1780. [Google Scholar] [CrossRef]
- Kusumi, A.; Nakada, C.; Ritchie, K.; Murase, K.; Suzuki, K.; Murakoshi, H.; Kasai, R.S.; Kondo, J.; Fujiwara, T. Paradigm shift of the plasma membrane concept from the two-dimensional continuum fluid to the partitioned fluid: High-speed single-molecule tracking of membrane molecules. Annu. Rev. Biophys. Biomol. Struct. 2005, 34, 351–378. [Google Scholar] [CrossRef] [PubMed]
- Simson, R.; Sheets, E.D.; Jacobson, K. Detection of temporary lateral confinement of membrane proteins using single-particle tracking analysis. Biophys. J. 1995, 69, 989–993. [Google Scholar] [CrossRef]
- Meilhac, N.; Le Guyader, L.; Salome, L.; Destainville, N. Detection of confinement and jumps in single-molecule membrane trajectories. Phys. Rev. E Stat. Nonlinear Soft Matter Phys. 2006, 73, 11915. [Google Scholar] [CrossRef] [PubMed]
- Montiel, D.; Cang, H.; Yang, H. Quantitative characterization of changes in dynamical behavior for single-particle tracking studies. J. Phys. Chem. B 2006, 110, 19763–19770. [Google Scholar] [CrossRef] [PubMed]
- Huet, S.; Karatekin, E.; Tran, V.S.; Fanget, I.; Cribier, S.; Henry, J.P. Analysis of transient behavior in complex trajectories: Application to secretory vesicle dynamics. Biophys. J. 2006, 91, 3542–3559. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.L.; Perillo, E.P.; Liu, C.; Yu, P.; Chou, C.K.; Hung, M.C.; Dunn, A.K.; Yeh, H.C. Segmentation of 3D trajectories acquired by TSUNAMI microscope: An application to EGFR trafficking. Biophys. J. 2016, 111, 2214–2227. [Google Scholar] [CrossRef] [PubMed]
- Bosch, P.J.; Kanger, J.S.; Subramaniam, V. Classification of dynamical diffusion states in single molecule tracking microscopy. Biophys. J. 2014, 107, 588–598. [Google Scholar] [CrossRef] [PubMed]
- Turkcan, S.; Masson, J.B. Bayesian decision tree for the classification of the mode of motion in single-molecule trajectories. PLoS ONE 2013, 8, e82799. [Google Scholar] [CrossRef] [PubMed]
- Slator, P.J.; Cairo, C.W.; Burroughs, N.J. Detection of diffusion heterogeneity in single particle tracking trajectories using a hidden markov model with measurement noise propagation. PLoS ONE 2015, 10, e0140759. [Google Scholar] [CrossRef] [PubMed]
- Bernstein, J.; Fricks, J. Analysis of single particle diffusion with transient binding using particle filtering. J. Theor. Biol. 2016, 401, 109–121. [Google Scholar] [CrossRef] [PubMed]
- Godinez, W.J.; Lampe, M.; Koch, P.; Eils, R.; Muller, B.; Rohr, K. Identifying virus-cell fusion in two-channel fluorescence microscopy image sequences based on a layered probabilistic approach. IEEE Trans. Med. Imaging 2012, 31, 1786–1808. [Google Scholar] [CrossRef] [PubMed]
- Dosset, P.; Rassam, P.; Fernandez, L.; Espenel, C.; Rubinstein, E.; Margeat, E.; Milhiet, P.E. Automatic detection of diffusion modes within biological membranes using back-propagation neural network. BMC Bioinform. 2016, 17, 197. [Google Scholar] [CrossRef] [PubMed]
- Lagache, T.; Dauty, E.; Holcman, D. Physical principles and models describing intracellular virus particle dynamics. Curr. Opin. Microbiol. 2009, 12, 439–445. [Google Scholar] [CrossRef] [PubMed]
- Boianelli, A.; Nguyen, V.K.; Ebensen, T.; Schulze, K.; Wilk, E.; Sharma, N.; Stegemann-Koniszewski, S.; Bruder, D.; Toapanta, F.R.; Guzman, C.A.; et al. Modeling influenza virus infection: A roadmap for influenza research. Viruses 2015, 7, 5274–5304. [Google Scholar] [CrossRef] [PubMed]
- Sbalzarini, I.F.; Greber, U.F. How computational models enable mechanistic insights into virus infection. In Influenza Virus: Methods & Protocols; Yamauchi, Y., Ed.; Springer: New York, NY, USA, 2018; in press. [Google Scholar]
- Siebourg-Polster, J.; Mudrak, D.; Emmenlauer, M.; Ramo, P.; Dehio, C.; Greber, U.; Frohlich, H.; Beerenwinkel, N. Nemix: Single-cell nested effects models for probabilistic pathway stimulation. PLoS Comput. Biol. 2015, 11, e1004078. [Google Scholar] [CrossRef] [PubMed]
- Ramo, P.; Drewek, A.; Arrieumerlou, C.; Beerenwinkel, N.; Ben-Tekaya, H.; Cardel, B.; Casanova, A.; Conde-Alvarez, R.; Cossart, P.; Csucs, G.; et al. Simultaneous analysis of large-scale rnai screens for pathogen entry. BMC Genom. 2014, 15, 1162. [Google Scholar] [CrossRef] [PubMed]
- Hancock, W.O. Bidirectional cargo transport: Moving beyond tug of war. Nat. Rev. Mol. Cell Biol. 2014, 15, 615–628. [Google Scholar] [CrossRef] [PubMed]
- Lagache, T.; Holcman, D. Quantifying intermittent transport in cell cytoplasm. Phys. Rev. E Stat. Nonlinear Soft Matter Phys. 2008, 77, 30901. [Google Scholar] [CrossRef] [PubMed]
- Muller, M.J.; Klumpp, S.; Lipowsky, R. Tug-of-war as a cooperative mechanism for bidirectional cargo transport by molecular motors. Proc. Natl. Acad. Sci. USA 2008, 105, 4609–4614. [Google Scholar] [CrossRef] [PubMed]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).