Engineering RNA Virus Interference via the CRISPR/Cas13 Machinery in Arabidopsis
Abstract
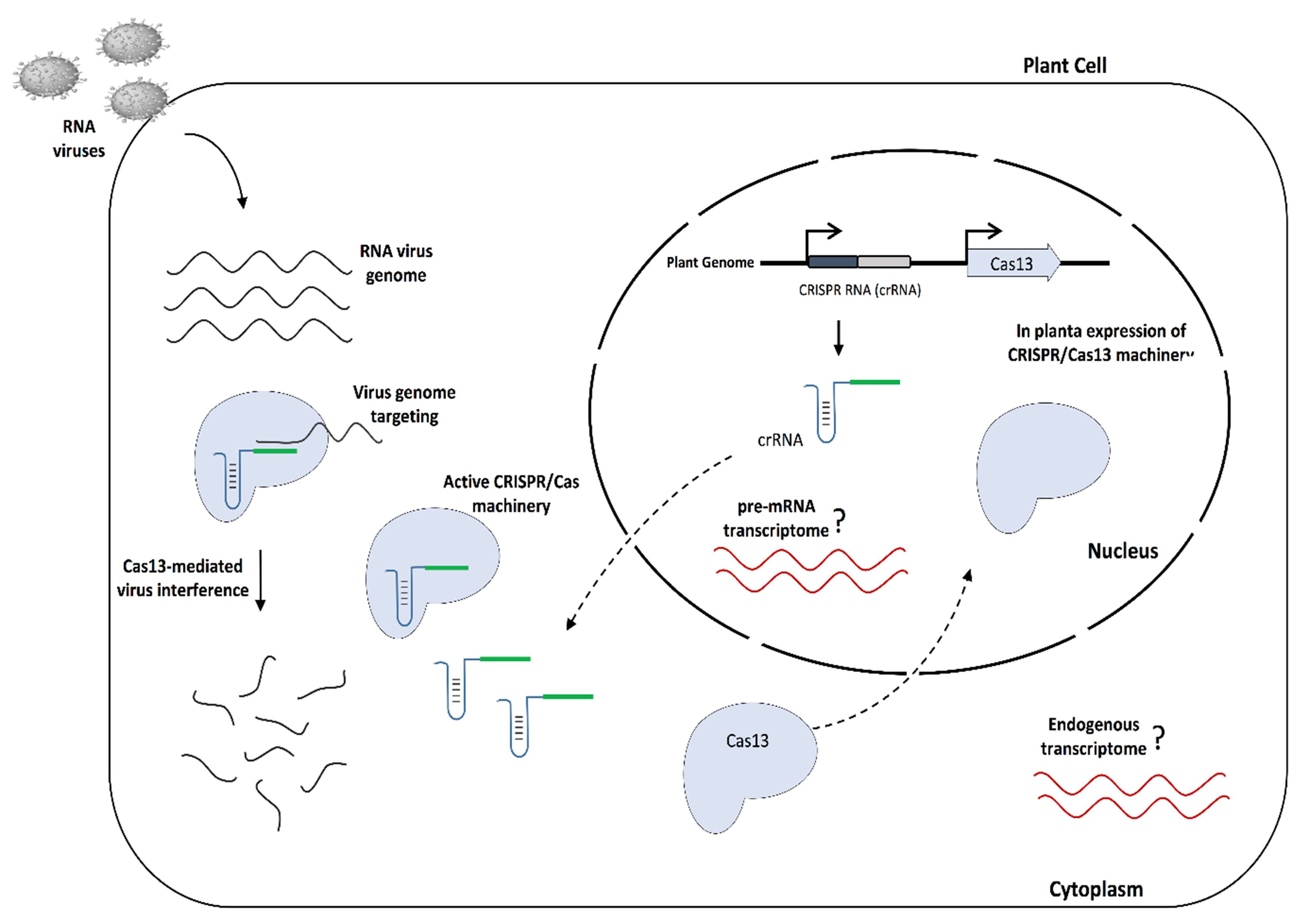
:1. Introduction
2. Material and Methods
2.1. Cloning of crRNAs under AtU6-26 Promoter
2.2. Plant Material
2.3. Arabidopsis Infection by TuMV-GFP via Sap Inoculation and Imaging under Ultraviolet (UV) Light
2.4. Immunoblot Analysis
2.5. RNA Isolation and Northern Blot Analysis
3. Results
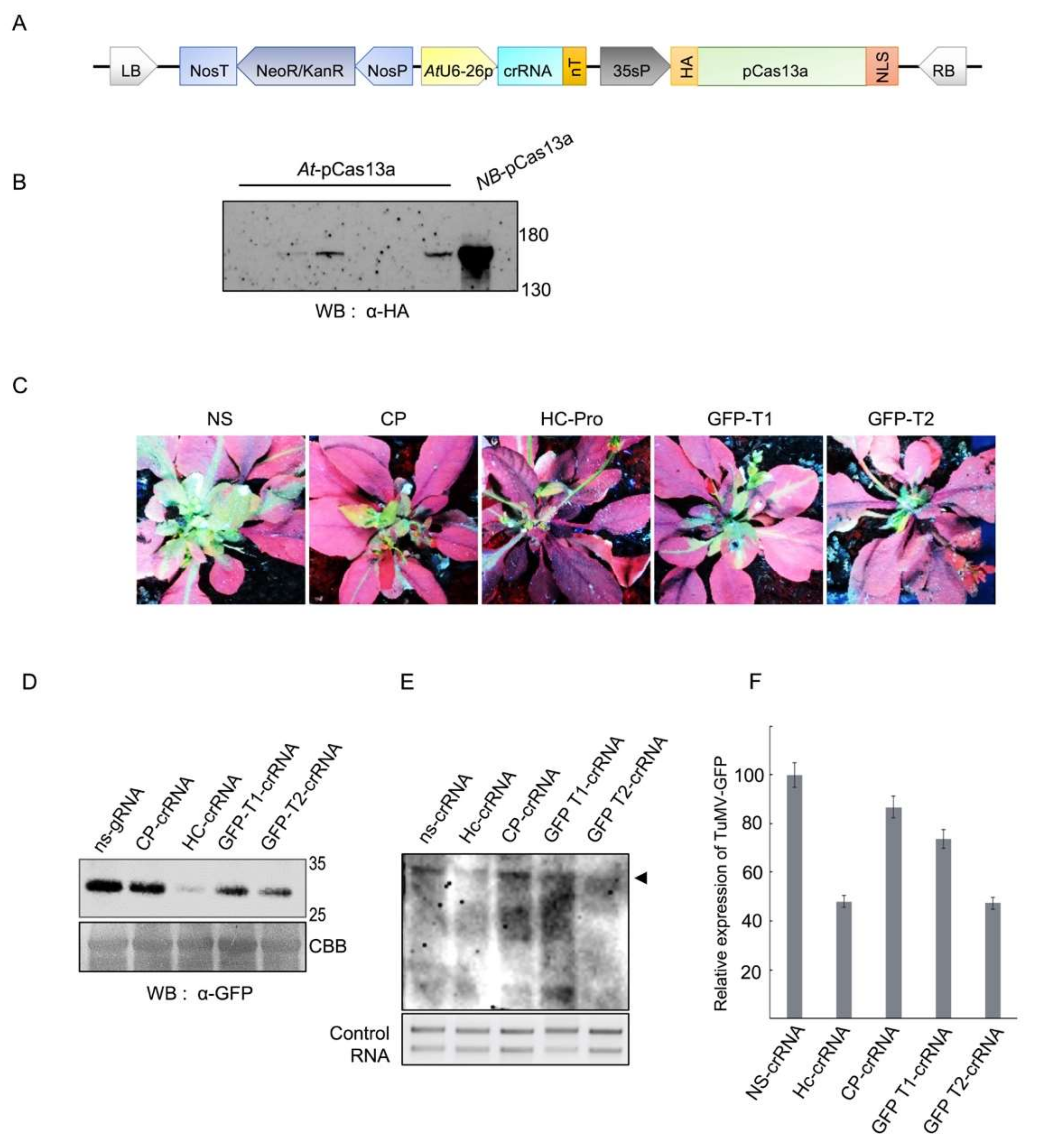
3.1. Construction, Over-Expression and Detection of the Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR)/pCas13a Machinery in A. thaliana Plants
3.2. CRISPR/pCas13a-Mediated TuMV-GFP Virus Interference in A. thaliana Plants
4. Discussion
Author Contributions
Funding
Conflicts of Interest
References
- Anderson, P.K.; Cunningham, A.A.; Patel, N.G.; Morales, F.J.; Epstein, P.R.; Daszak, P. Emerging infectious diseases of plants: Pathogen pollution, climate change and agrotechnology drivers. Trends Ecol. Evol. 2004, 19, 535–544. [Google Scholar] [CrossRef] [PubMed]
- Legg, J.P.; Thresh, J.M. Cassava mosaic virus disease in east africa: A dynamic disease in a changing environment. Virus Res. 2000, 71, 135–149. [Google Scholar] [CrossRef]
- Sasaya, T.; Nakazono-Nagaoka, E.; Saika, H.; Aoki, H.; Hiraguri, A.; Netsu, O.; Uehara-Ichiki, T.; Onuki, M.; Toki, S.; Saito, K.; et al. Transgenic strategies to confer resistance against viruses in rice plants. Front. Microbiol. 2014, 4, 409. [Google Scholar] [CrossRef]
- Sudarshana, M.R.; Roy, G.; Falk, B.W. Methods for engineering resistance to plant viruses. Methods Mol. Biol. 2007, 354, 183–195. [Google Scholar] [PubMed]
- Baulcombe, D.C. Mechanisms of pathogen-derived resistance to viruses in transgenic plants. Plant Cell 1996, 8, 1833–1844. [Google Scholar] [CrossRef] [PubMed]
- Beachy, R.N.; Loeschfries, S.; Tumer, N.E. Coat protein-mediated resistance against virus-infection. Ann. Rev. Phytopathol. 1990, 28, 451–474. [Google Scholar] [CrossRef]
- Lu, R.; Martin-Hernandez, A.M.; Peart, J.R.; Malcuit, I.; Baulcombe, D.C. Virus-induced gene silencing in plants. Methods 2003, 30, 296–303. [Google Scholar] [CrossRef]
- Simon-Mateo, C.; Garcia, J.A. Antiviral strategies in plants based on rna silencing. Biochim. Biophys. Acta 2011, 1809, 722–731. [Google Scholar] [CrossRef]
- Piquerez, S.J.; Harvey, S.E.; Beynon, J.L.; Ntoukakis, V. Improving crop disease resistance: Lessons from research on arabidopsis and tomato. Front. Plant Sci. 2014, 5, 671. [Google Scholar] [CrossRef]
- Pumplin, N.; Voinnet, O. Rna silencing suppression by plant pathogens: Defence, counter-defence and counter-counter-defence. Nat. Rev. Microbiol. 2013, 11, 745–760. [Google Scholar] [CrossRef]
- Voinnet, O. Induction and suppression of rna silencing: Insights from viral infections. Nat. Rev. Genet. 2005, 6, 206–220. [Google Scholar] [CrossRef] [PubMed]
- Sorek, R.; Lawrence, C.M.; Wiedenheft, B. Crispr-mediated adaptive immune systems in bacteria and archaea. Annu. Rev. Biochem. 2013, 82, 237–266. [Google Scholar] [CrossRef] [PubMed]
- Adli, M. The crispr tool kit for genome editing and beyond. Nat. Commun. 2018, 9, 1911. [Google Scholar] [CrossRef] [PubMed]
- Komor, A.C.; Badran, A.H.; Liu, D.R. Crispr-based technologies for the manipulation of eukaryotic genomes. Cell 2017, 168, 20–36. [Google Scholar] [CrossRef] [PubMed]
- Rath, D.; Amlinger, L.; Rath, A.; Lundgren, M. The crispr-cas immune system: Biology, mechanisms and applications. Biochimie 2015, 117, 119–128. [Google Scholar] [CrossRef]
- Mahas, A.; Mahfouz, M. Engineering virus resistance via crispr-cas systems. Curr. Opin. Virol. 2018, 32, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Ali, Z.; Abulfaraj, A.; Idris, A.; Ali, S.; Tashkandi, M.; Mahfouz, M.M. Crispr/cas9-mediated viral interference in plants. Genome Biol. 2015, 16, 238. [Google Scholar] [CrossRef]
- Ali, Z.; Ali, S.; Tashkandi, M.; Zaidi, S.S.; Mahfouz, M.M. Crispr/cas9-mediated immunity to geminiviruses: Differential interference and evasion. Sci. Rep. 2016, 6, 26912. [Google Scholar] [CrossRef]
- Tashkandi, M.; Ali, Z.; Aljedaani, F.; Shami, A.; Mahfouz, M.M. Engineering resistance against tomato yellow leaf curl virus via the crispr/cas9 system in tomato. Plant Signal. Behav. 2018, 13, e1525996. [Google Scholar]
- Baltes, N.J. Conferring resistance to geminiviruses with the crispr–cas prokaryotic immune system. Nat. Plants 2015, 1, 15145. [Google Scholar] [CrossRef]
- Abudayyeh, O.O.; Gootenberg, J.S.; Konermann, S.; Joung, J.; Slaymaker, I.M.; Cox, D.B.; Shmakov, S.; Makarova, K.S.; Semenova, E.; Minakhin, L.; et al. C2c2 is a single-component programmable rna-guided rna-targeting crispr effector. Science 2016, 353, aaf5573. [Google Scholar] [CrossRef] [PubMed]
- East-Seletsky, A.; O’Connell, M.R.; Knight, S.C.; Burstein, D.; Cate, J.H.; Tjian, R.; Doudna, J.A. Two distinct rnase activities of crispr-c2c2 enable guide-rna processing and rna detection. Nature 2016, 538, 270–273. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Li, X.; Wang, J.; Wang, M.; Chen, P.; Yin, M.; Li, J.; Sheng, G.; Wang, Y. Two distant catalytic sites are responsible for c2c2 rnase activities. Cell 2017, 168, 121–134e12. [Google Scholar] [CrossRef] [PubMed]
- Mahas, A.; Neal Stewart, C., Jr.; Mahfouz, M.M. Harnessing crispr/cas systems for programmable transcriptional and post-transcriptional regulation. Biotechnol. Adv. 2017, 36, 295–310. [Google Scholar] [CrossRef] [PubMed]
- Abudayyeh, O.O.; Gootenberg, J.S.; Essletzbichler, P.; Han, S.; Joung, J.; Belanto, J.J.; Verdine, V.; Cox, D.B.T.; Kellner, M.J.; Regev, A.; et al. Rna targeting with crispr-cas13. Nature 2017, 550, 280–284. [Google Scholar] [CrossRef]
- Aman, R.; Ali, Z.; Butt, H.; Mahas, A.; Aljedaani, F.; Khan, M.Z.; Ding, S.; Mahfouz, M. Rna virus interference via crispr/cas13a system in plants. Genome Biol. 2018, 19, 1. [Google Scholar] [CrossRef]
- Jing, X.; Xie, B.; Chen, L.; Zhang, N.; Jiang, Y.; Qin, H.; Wang, H.; Hao, P.; Yang, S.; Li, X. Implementation of the crispr-cas13a system in fission yeast and its repurposing for precise rna editing. Nucleic Acids Res. 2018, 46, e90. [Google Scholar] [CrossRef] [PubMed]
- Ali, Z.; Abul-faraj, A.; Li, L.; Ghosh, N.; Piatek, M.; Mahjoub, A.; Aouida, M.; Piatek, A.; Baltes, N.J.; Voytas, D.F.; et al. Efficient virus-mediated genome editing in plants using the crispr/cas9 system. Mol. Plant 2015, 8, 1288–1291. [Google Scholar] [CrossRef]
- Gibbs, A.; Ohshima, K. Potyviruses and the digital revolution. Annu. Rev. Phytopathol. 2010, 48, 205–223. [Google Scholar] [CrossRef]
- Brian Mahy, M.v.R. Desk Encyclopedia of Plant and Fungal Virology; Academic Press: Cambridge, MA, USA, 2009; p. 632. [Google Scholar]
- Ma, X.; Zhu, Q.; Chen, Y.; Liu, Y.G. Crispr/cas9 platforms for genome editing in plants: Developments and applications. Mol. Plant 2016, 9, 961–974. [Google Scholar] [CrossRef]
- Svitashev, S.; Schwartz, C.; Lenderts, B.; Young, J.K.; Mark Cigan, A. Genome editing in maize directed by crispr-cas9 ribonucleoprotein complexes. Nat. Commun. 2016, 7, 13274. [Google Scholar] [CrossRef]
- Ali, Z.; Mahas, A.; Mahfouz, M. Crispr/cas13 as a tool for rna interference. Trends Plant Sci. 2018, 23, 374–378. [Google Scholar] [CrossRef] [PubMed]
- Clough, S.J.; Bent, A.F. Floral dip: A simplified method for agrobacterium-mediated transformation of arabidopsis thaliana. Plant J. 1998, 16, 735–743. [Google Scholar] [CrossRef] [PubMed]
| Oligo Name | Sequence (5′-3′) | Purpose |
|---|---|---|
| GFP-T1-U6-Top strand | gattgccaccccaatatcgaaggggactaaaacaacaggtagttttccagtagtgcaaatatttttttttc | TuMV-GFP-T1 crRNA |
| GFP-T1-U6-Bottom strand | catggaaaaaaaaatatttgcactactggaaaactacctgttgttttagtccccttcgatattggggtggc | |
| GFP-T2-U6-Top strand | gattgccaccccaatatcgaaggggactaaaacccgtcctccttgaaatcgattcccttaatttttttttc | TuMV-GFP-T2 crRNA |
| GFP-T2-U6- Bottom strand | catggaaaaaaaaattaagggaatcgatttcaaggaggacgggttttagtccccttcgatattggggtggc | |
| HC-Pro-U6-Top strand | gattgccaccccaatatcgaaggggactaaaacccgcttgcttgtccttgggatagctcactttttttttc | TuMV-HC-pro crRNA |
| HC-Pro-U6- Bottom strand | catggaaaaaaaaagtgagctatcccaaggacaagcaagcgggttttagtccccttcgatattggggtggc | |
| CP-U6-Top strand | gattgccaccccaatatcgaaggggactaaaacacactgaaagttccagaggttccagcgttttttttttc | TuMV-CP crRNA |
| CP-U6-Bottom strand | catggaaaaaaaaaacgctggaacctctggaactttcagtgtgttttagtccccttcgatattggggtggc | |
| AtU6-F | atgagctcaagcttcgacttgccttccg | For PCR amplification of AtU6 promoter-crRNA cassette out of the intermediate pUC19 vector |
| AtU6-R | aggtaccacgcgtcactagtataaccatgg |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Aman, R.; Mahas, A.; Butt, H.; Ali, Z.; Aljedaani, F.; Mahfouz, M. Engineering RNA Virus Interference via the CRISPR/Cas13 Machinery in Arabidopsis. Viruses 2018, 10, 732. https://doi.org/10.3390/v10120732
Aman R, Mahas A, Butt H, Ali Z, Aljedaani F, Mahfouz M. Engineering RNA Virus Interference via the CRISPR/Cas13 Machinery in Arabidopsis. Viruses. 2018; 10(12):732. https://doi.org/10.3390/v10120732
Chicago/Turabian StyleAman, Rashid, Ahmed Mahas, Haroon Butt, Zahir Ali, Fatimah Aljedaani, and Magdy Mahfouz. 2018. "Engineering RNA Virus Interference via the CRISPR/Cas13 Machinery in Arabidopsis" Viruses 10, no. 12: 732. https://doi.org/10.3390/v10120732
APA StyleAman, R., Mahas, A., Butt, H., Ali, Z., Aljedaani, F., & Mahfouz, M. (2018). Engineering RNA Virus Interference via the CRISPR/Cas13 Machinery in Arabidopsis. Viruses, 10(12), 732. https://doi.org/10.3390/v10120732






