Laminariales Host Does Impact Lipid Temperature Trajectories of the Fungal Endophyte Paradendryphiella salina (Sutherland.)
Abstract
1. Introduction
2. Results
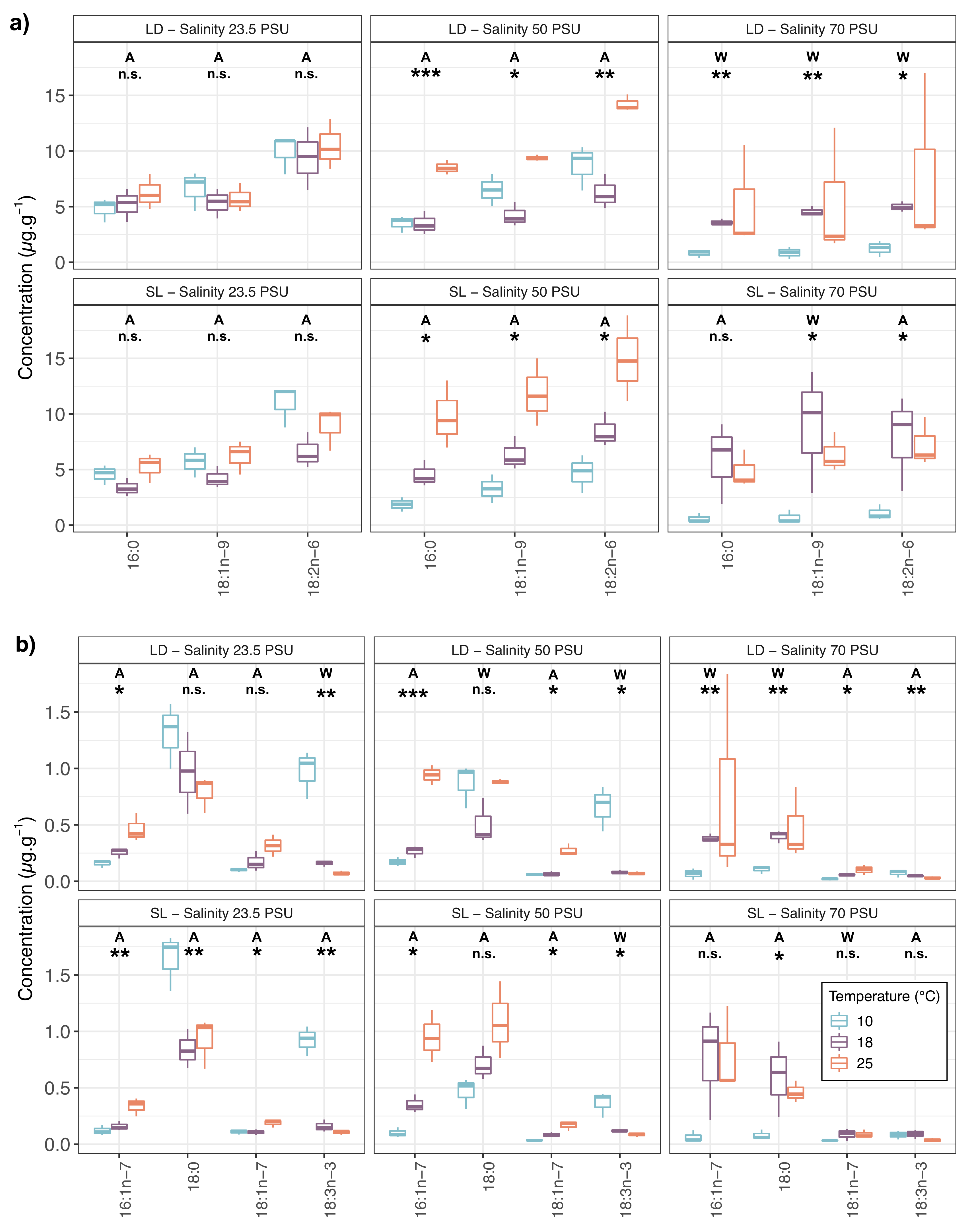
2.1. Fatty Acid Compositions
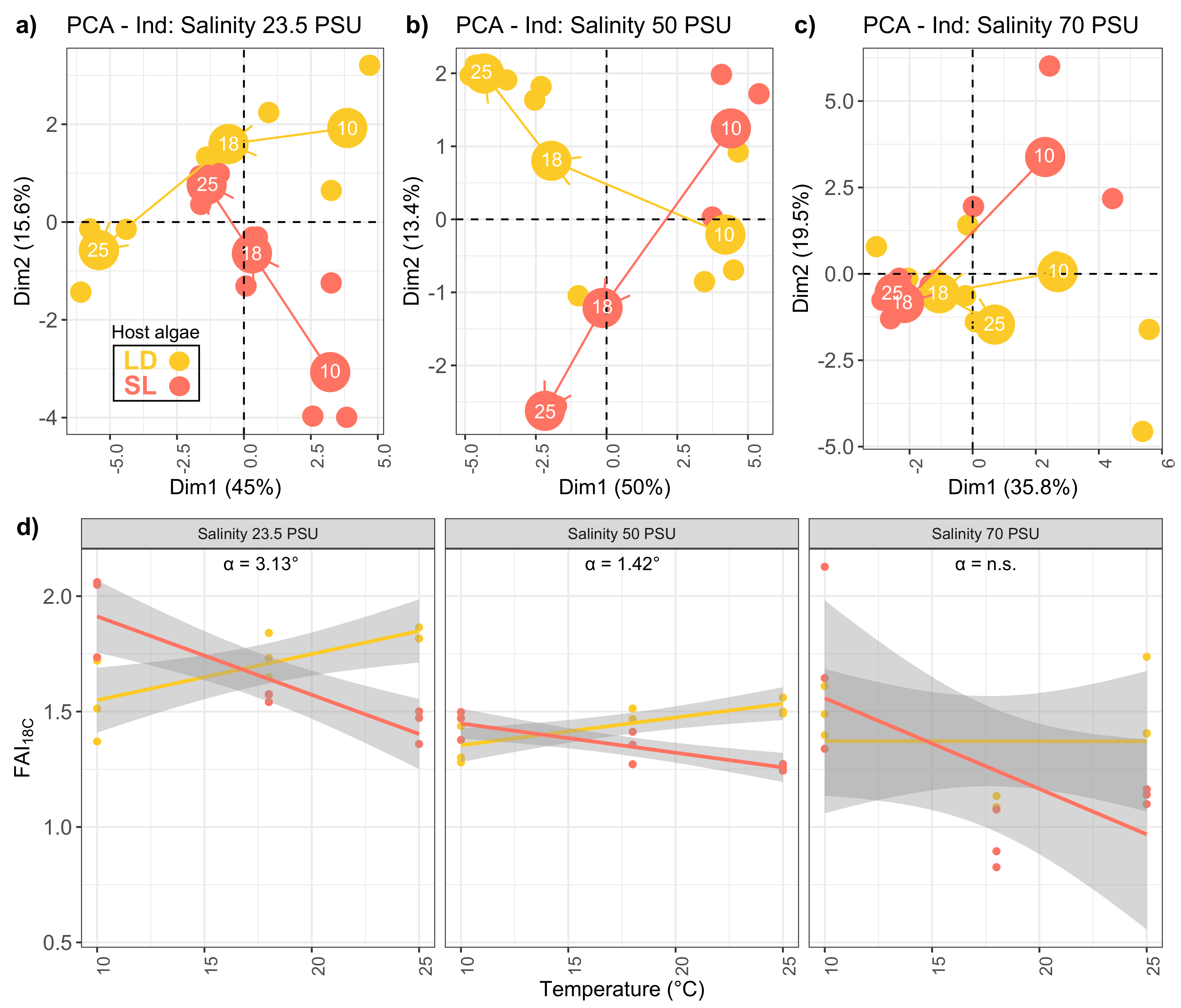
2.2. Lipid Trajectories
3. Discussion
3.1. Fatty Acid Compositions of P. salina in Relation to Experimental Conditions
3.2. Divergent Fatty Acid Trajectories in P. salina Revealed Adaptive Strategies to Temperature Changes
4. Materials and Methods
4.1. Reagents and Chemicals
4.2. Strain Isolation, Cultivation and Identification
4.3. Experimental Design
4.4. Fatty Acid Extraction
4.5. Statistics
4.5.1. Univariate
4.5.2. Multivariate
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| LD | Laminaria digitata |
| MPB | Microphytobenthos |
| MUFA | Monounsaturated fatty acid |
| PUFA | Polyunsaturated fatty acid |
| QS | Quorum Sensing |
| SFA | Saturated fatty acid |
| SL | Saccharina latissima |
| TFA | Total Fatty Acids |
| PSU | practical salinity units |
| ANOVA | analysis of variance |
| PCA | principal component analyses |
| FAME | Fatty acid methyl esters |
| NIST | National Institute of Standards and Technology |
| FID | flame ionization detector |
References
- Bartsch, I.; Wiencke, C.; Bischof, K.; Buchholz, C.M.; Buck, B.H.; Eggert, A.; Feuerpfeil, P.; Hanelt, D.; Jacobsen, S.; Karez, R.; et al. The genus Laminaria sensu lato: Recent insights and developments. Eur. J. Phycol. 2008, 43, 1–86. [Google Scholar] [CrossRef]
- Vallet, M.; Strittmatter, M.; Murúa, P.; Lacoste, S.; Dupont, J.; Hubas, C.; Genta-Jouve, G.; Gachon, C.M.M.; Kim, G.H.; Prado, S. Chemically-Mediated Interactions Between Macroalgae, Their Fungal Endophytes, and Protistan Pathogens. Front. Microbiol. 2018, 9. [Google Scholar] [CrossRef] [PubMed]
- Sutherland, G.K. Marine Fungi Imperfecti. New Phytol. 1916, 15, 35–48. [Google Scholar] [CrossRef]
- Michaelis, K.C.; Gessner, R.V.; Romano, M.A. Population Genetics and Systematics of Marine Species of Dendryphiella. Mycologia 1987, 79, 514–518. [Google Scholar] [CrossRef]
- Dela Cruz, T.E.; Wagner, S.; Schulz, B. Physiological responses of marine Dendryphiella species from different geographical locations. Mycol. Prog. 2006, 5, 108–119. [Google Scholar] [CrossRef]
- Kireev, Y.V.; Konovalova, O.P.; Myuge, N.S.; Shnyreva, A.V.; Bubnova, E.N. Cultural properties and taxonomic position of Helminthosporium-like fungal isolates from the White Sea. Microbiology 2015, 84, 665–676. [Google Scholar] [CrossRef]
- Tourneroche, A.; Lami, R.; Hubas, C.; Blanchet, E.; Vallet, M.; Escoubeyrou, K.; Paris, A.; Prado, S. Bacterial–Fungal Interactions in the Kelp Endomicrobiota Drive Autoinducer-2 Quorum Sensing. Front. Microbiol. 2019, 10, 1693. [Google Scholar] [CrossRef]
- Vallet, M.; Chong, Y.M.; Tourneroche, A.; Genta-Jouve, G.; Hubas, C.; Lami, R.; Gachon, C.; Klochkova, T.; Chan, K.G.; Prado, S. Novel α-hydroxy γ-butenolides of kelp endophytes disrupt bacterial cell-to-cell signaling. Front. Mar. Sci. 2020. [Google Scholar] [CrossRef]
- Pilgaard, B.; Wilkens, C.; Herbst, F.A.; Vuillemin, M.; Rhein-Knudsen, N.; Meyer, A.S.; Lange, L. Proteomic enzyme analysis of the marine fungus Paradendryphiella salina reveals alginate lyase as a minimal adaptation strategy for brown algae degradation. Sci. Rep. 2019, 9, 1–13. [Google Scholar] [CrossRef]
- Barbosa, M.; Fernandes, F.; Pereira, D.M.; Azevedo, I.C.; Sousa-Pinto, I.; Andrade, P.B.; Valentão, P. Fatty acid patterns of the kelps Saccharina latissima, Saccorhiza polyschides and Laminaria ochroleuca: Influence of changing environmental conditions. Arab. J. Chem. 2020, 13, 45–58. [Google Scholar] [CrossRef]
- Dawczynski, C.; Schubert, R.; Jahreis, G. Amino acids, fatty acids, and dietary fibre in edible seaweed products. Food Chem. 2007, 103, 891–899. [Google Scholar] [CrossRef]
- Weete, J.D. Fatty Acids. In Lipid Biochemistry of Fungi and Other Organisms; Springer: Boston, MA, USA, 1980; pp. 49–95. [Google Scholar] [CrossRef]
- Bärenstrauch, M.; Mann, S.; Jacquemin, C.; Bibi, S.; Sylla, O.K.; Baudouin, E.; Buisson, D.; Prado, S.; Kunz, C. Molecular crosstalk between the endophyte Paraconiothyrium variabile and the phytopathogen Fusarium oxysporum—Modulation of lipoxygenase activity and beauvericin production during the interaction. Fungal Genet. Biol. 2020, 139, 103383. [Google Scholar] [CrossRef] [PubMed]
- Fischer, G.J.; Keller, N.P. Production of cross-kingdom oxylipins by pathogenic fungi: An update on their role in development and pathogenicity. J. Microbiol. 2016, 54, 254–264. [Google Scholar] [CrossRef] [PubMed]
- Savchenko, T.V.; Zastrijnaja, O.M.; Klimov, V.V. Oxylipins and plant abiotic stress resistance. Biochemistry 2014, 79, 362–375. [Google Scholar] [CrossRef] [PubMed]
- Hurd, C.L. Water motion, marine macroalgal physiology, and production. J. Phycol. 2000, 36, 453–472. [Google Scholar] [CrossRef]
- Norton, T.; Mathieson, A.; Neushul, M. A Review of Some Aspects of Form and Function in Seaweeds. Bot. Mar. 1982, 25, 501–510. [Google Scholar] [CrossRef]
- Bekkby, T.; Smit, C.; Gundersen, H.; Rinde, E.; Steen, H.; Tveiten, L.; Gitmark, J.K.; Fredriksen, S.; Albretsen, J.; Christie, H.C. The abundance of kelp is modified by the combined impact of depth, waves and currents. Front. Mar. Sci. 2019, 6. [Google Scholar] [CrossRef]
- Karsten, U. Research note: Salinity tolerance of Arctic kelps from Spitsbergen. Phycol. Res. 2007, 55, 257–262. [Google Scholar] [CrossRef]
- Küpper, F.C.; Gaquerel, E.; Boneberg, E.M.; Morath, S.; Salaün, J.P.; Potin, P. Early events in the perception of lipopolysaccharides in the brown alga Laminaria digitata include an oxidative burst and activation of fatty acid oxidation cascades. J. Exp. Bot. 2006, 57, 1991–1999. [Google Scholar] [CrossRef]
- Ritter, A.; Cabioch, L.; Brillet-Guéguen, L.; Corre, E.; Cosse, A.; Dartevelle, L.; Duruflé, H.; Fasshauer, C.; Goulitquer, S.; Thomas, F.; et al. Herbivore-induced chemical and molecular responses of the kelps Laminaria digitata and Lessonia spicata. PLoS ONE 2017, 12, e0173315. [Google Scholar] [CrossRef]
- Küpper, F.C.; Gaquerel, E.; Cosse, A.; Adas, F.; Peters, A.F.; Müller, D.G.; Kloareg, B.; Salaün, J.P.; Potin, P. Free Fatty Acids and Methyl Jasmonate Trigger Defense Reactions in Laminaria digitata. Plant Cell Physiol. 2009, 50, 789–800. [Google Scholar] [CrossRef] [PubMed]
- Steinhoff, F.S.; Graeve, M.; Wiencke, C.; Wulff, A.; Bischof, K. Lipid content and fatty acid consumption in zoospores/developing gametophytes of Saccharina latissima (Laminariales, Phaeophyceae) as potential precursors for secondary metabolites as phlorotannins. Polar Biol. 2011, 34, 1011–1018. [Google Scholar] [CrossRef]
- Neidleman, S.L. Effects of temperature on lipid unsaturation. Biotechnol. Genet. Eng. Rev. 1987, 5, 245–268. [Google Scholar] [CrossRef] [PubMed]
- Kjer, J.; Debbab, A.; Aly, A.H.; Proksch, P. Methods for isolation of marine-derived endophytic fungi and their bioactive secondary products. Nat. Protoc. 2010, 5, 479–490. [Google Scholar] [CrossRef]
- Kientz, B.; Thabard, M.; Cragg, S.M.; Pope, J.; Hellio, C. A new method for removing microflora from macroalgal surfaces: An important step for natural product discovery. Bot. Mar. 2011, 54, 457–469. [Google Scholar] [CrossRef]
- Bligh, E.G.; Dyer, W.J. A Rapid Method of Total Lipid Extraction and Purification. Can. J. Biochem. Physiol. 1959, 37, 911–917. [Google Scholar] [CrossRef]
- Meziane, T.; Tsuchiya, M. Fatty acids as tracers of organic matter in the sediment and food web of a mangrove/intertidal flat ecosystem, Okinawa, Japan. Mar. Ecol. Prog. Ser. 2000, 200, 49–57. [Google Scholar] [CrossRef]
- Passarelli, C.; Meziane, T.; Thiney, N.; Boeuf, D.; Jesus, B.; Ruivo, M.; Jeanthon, C.; Hubas, C. Seasonal variations of the composition of microbial biofilms in sandy tidal flats: Focus of fatty acids, pigments and exopolymers. Estuarine Coast. Shelf Sci. 2015, 153, 29–37. [Google Scholar] [CrossRef]
- Fahy, E.; Subramaniam, S.; Brown, H.A.; Glass, C.K.; Merrill, A.H.; Murphy, R.C.; Raetz, C.R.; Russell, D.W.; Seyama, Y.; Shaw, W.; et al. A comprehensive classification system for lipids. J. Lipid Res. 2005, 46, 839–861. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2019. [Google Scholar]
- Wickham, H. ggplot2: Elegant Graphics for Data Analysis; Springer: New York, NY, USA, 2016. [Google Scholar]
- Kassambara, A. rstatix: Pipe-Friendly Framework for Basic Statistical Tests. 2020. Available online: https://CRAN.R-project.org/package=rstatix (accessed on 21 May 2020).
- Dray, S.; Dufour, A.B. The ade4 Package: Implementing the Duality Diagram for Ecologists. J. Stat. Softw. 2007, 22, 1–20. [Google Scholar] [CrossRef]
- Kassambara, A.; Mundt, F. Factoextra: Extract and Visualize the Results of Multivariate Data Analyses. 2019. Available online: https://CRAN.R-project.org/package=factoextra (accessed on 21 May 2020).
- Wilke, C.O. cowplot: Streamlined Plot Theme and Plot Annotations for ggplot2. 2019. Available online: https://CRAN.R-project.org/package=cowplot (accessed on 21 May 2020).
| 12:0 | 14:0 | 14:1n-5 | 15:0 | 16:0 | 16:1n-7 | 16:1n-9 | 16:2n-4 | |||||||||||
| Host algae | Temperature | Salinity | mean | sd | mean | sd | mean | sd | mean | sd | mean | sd | mean | sd | mean | sd | mean | sd |
| LD | T10 | S1 | 0.02 | 0.022 | 0.04 | 0.007 | 0.01 | 0.001 | <dl | <dl | 4.79 | 1.069 | 0.16 | 0.035 | 0.03 | 0.005 | 0.01 | 0.003 |
| SL | T10 | S1 | <dl | <dl | 0.03 | 0.005 | <dl | <dl | 0.01 | 0.001 | 4.55 | 0.899 | 0.12 | 0.045 | 0.03 | 0.008 | 0.01 | 0.002 |
| LD | T18 | S1 | 0.01 | 0.010 | 0.03 | 0.011 | 0.01 | 0.000 | 0.01 | 0.002 | 5.2 | 1.472 | 0.25 | 0.044 | 0.04 | 0.008 | 0.02 | 0.004 |
| SL | T18 | S1 | <dl | <dl | 0.02 | 0.003 | <dl | <dl | 0.01 | 0.002 | 3.36 | 0.810 | 0.16 | 0.040 | 0.03 | 0.008 | 0.01 | 0.001 |
| LD | T25 | S1 | 0.01 | 0.005 | 0.03 | 0.004 | 0.02 | 0.015 | 0.02 | 0.005 | 6.24 | 1.586 | 0.46 | 0.126 | 0.06 | 0.016 | 0.03 | 0.009 |
| SL | T25 | S1 | <dl | <dl | 0.03 | 0.005 | 0.01 | 0.003 | 0.01 | 0.003 | 5.26 | 1.311 | 0.34 | 0.081 | 0.04 | 0.008 | 0.02 | 0.004 |
| LD | T10 | S2 | 0.01 | 0.006 | 0.03 | 0.007 | <dl | <dl | 0.01 | 0.001 | 3.49 | 0.741 | 0.17 | 0.040 | 0.02 | 0.005 | 0.01 | 0.002 |
| SL | T10 | S2 | 0.01 | 0.011 | 0.02 | 0.001 | <dl | <dl | <dl | <dl | 1.87 | 0.640 | 0.1 | 0.044 | 0.01 | 0.005 | <dl | <dl |
| LD | T18 | S2 | 0.01 | 0.017 | 0.03 | 0.004 | 0.01 | 0.003 | 0.01 | 0.003 | 3.47 | 1.061 | 0.27 | 0.053 | 0.04 | 0.008 | 0.01 | 0.004 |
| SL | T18 | S2 | 0.01 | 0.001 | 0.04 | 0.009 | 0.01 | 0.001 | 0.02 | 0.004 | 4.55 | 1.194 | 0.35 | 0.081 | 0.04 | 0.008 | 0.02 | 0.005 |
| LD | T25 | S2 | 0.01 | 0.002 | 0.08 | 0.030 | 0.02 | 0.010 | 0.02 | 0.005 | 8.5 | 0.643 | 0.94 | 0.088 | 0.09 | 0.008 | 0.07 | 0.004 |
| SL | T25 | S2 | <dl | <dl | 0.09 | 0.031 | 0.02 | 0.003 | 0.03 | 0.007 | 9.8 | 3.034 | 0.95 | 0.230 | 0.08 | 0.014 | 0.05 | 0.014 |
| LD | T10 | S3 | 0.01 | 0.002 | 0.01 | 0.004 | <dl | <dl | <dl | <dl | 0.82 | 0.373 | 0.07 | 0.049 | <dl | <dl | <dl | <dl |
| SL | T10 | S3 | <dl | <dl | 0.01 | 0.004 | <dl | <dl | <dl | <dl | 0.62 | 0.418 | 0.06 | 0.051 | <dl | <dl | <dl | <dl |
| LD | T18 | S3 | <dl | <dl | 0.04 | 0.004 | <dl | <dl | 0.01 | 0.001 | 3.6 | 0.277 | 0.38 | 0.037 | 0.01 | 0.003 | 0.02 | 0.002 |
| SL | T18 | S3 | <dl | <dl | 0.07 | 0.044 | <dl | <dl | 0.02 | 0.013 | 5.91 | 3.655 | 0.76 | 0.493 | 0.03 | 0.020 | 0.03 | 0.017 |
| LD | T25 | S3 | 0.03 | 0.037 | 0.07 | 0.038 | <dl | <dl | 0.01 | 0.005 | 5.19 | 4.629 | 0.76 | 0.937 | 0.02 | 0.021 | 0.06 | 0.081 |
| SL | T25 | S3 | <dl | <dl | 0.07 | 0.036 | <dl | <dl | 0.02 | 0.008 | 4.87 | 1.677 | 0.78 | 0.383 | 0.03 | 0.008 | 0.04 | 0.022 |
| 16:2n-6 | 17:0 | 17:1n-7 | 17:1n-9 | 17:2n-5 | 18:0 | 18:1n-7 | 18:1n-9 | |||||||||||
| Host algae | Temperature | Salinity | mean | sd | mean | sd | mean | sd | mean | sd | mean | sd | mean | sd | mean | sd | mean | sd |
| LD | T10 | S1 | 0.01 | 0.002 | 0.03 | 0.006 | 0.01 | 0.003 | <dl | <dl | <dl | <dl | 1.31 | 0.291 | 0.1 | 0.015 | 6.6 | 1.774 |
| SL | T10 | S1 | 0.01 | 0.010 | 0.04 | 0.002 | 0.01 | 0.001 | 0.01 | 0.008 | <dl | <dl | 1.64 | 0.251 | 0.11 | 0.022 | 5.71 | 1.359 |
| LD | T18 | S1 | 0.02 | 0.003 | 0.03 | 0.006 | <dl | <dl | <dl | <dl | <dl | <dl | 0.97 | 0.363 | 0.17 | 0.090 | 5.34 | 1.333 |
| SL | T18 | S1 | 0.01 | 0.004 | 0.03 | 0.008 | 0.01 | 0.001 | <dl | <dl | <dl | <dl | 0.84 | 0.174 | 0.11 | 0.023 | 4.21 | 0.980 |
| LD | T25 | S1 | 0.04 | 0.014 | 0.04 | 0.016 | 0.01 | 0.002 | <dl | <dl | 0.01 | 0.002 | 0.79 | 0.161 | 0.32 | 0.098 | 5.72 | 1.262 |
| SL | T25 | S1 | 0.02 | 0.004 | 0.03 | 0.007 | 0.01 | 0.001 | <dl | <dl | <dl | <dl | 0.93 | 0.224 | 0.19 | 0.034 | 6.23 | 1.515 |
| LD | T10 | S2 | <dl | <dl | 0.01 | 0.001 | <dl | <dl | <dl | <dl | <dl | <dl | 0.87 | 0.195 | 0.06 | 0.009 | 6.5 | 1.456 |
| SL | T10 | S2 | <dl | <dl | 0.01 | 0.007 | <dl | <dl | <dl | <dl | <dl | <dl | 0.47 | 0.135 | 0.04 | 0.007 | 3.27 | 1.289 |
| SL | T18 | S2 | 0.01 | 0.004 | 0.02 | 0.005 | 0.01 | 0.001 | <dl | <dl | <dl | <dl | 0.71 | 0.150 | 0.08 | 0.019 | 6.33 | 1.516 |
| LD | T25 | S2 | 0.05 | 0.008 | 0.03 | 0.008 | 0.01 | 0.001 | 0.01 | 0.001 | 0.01 | 0.000 | 0.88 | 0.021 | 0.27 | 0.054 | 9.38 | 0.268 |
| SL | T25 | S2 | 0.02 | 0.003 | 0.02 | 0.003 | 0.01 | 0.001 | <dl | <dl | <dl | <dl | 1.09 | 0.341 | 0.17 | 0.045 | 11.85 | 3.027 |
| LD | T10 | S3 | <dl | <dl | <dl | <dl | <dl | <dl | <dl | <dl | <dl | <dl | 0.11 | 0.035 | 0.02 | 0.011 | 0.85 | 0.551 |
| SL | T10 | S3 | <dl | <dl | <dl | <dl | <dl | <dl | <dl | <dl | <dl | <dl | 0.08 | 0.043 | 0.03 | 0.004 | 0.7 | 0.599 |
| LD | T18 | S3 | <dl | <dl | 0.01 | 0.002 | <dl | <dl | <dl | <dl | <dl | <dl | 0.4 | 0.056 | 0.06 | 0.007 | 4.53 | 0.443 |
| SL | T18 | S3 | <dl | <dl | 0.01 | 0.008 | 0.01 | 0.005 | <dl | <dl | <dl | <dl | 0.6 | 0.336 | 0.09 | 0.054 | 8.92 | 5.547 |
| LD | T25 | S3 | <dl | <dl | 0.01 | 0.002 | 0.01 | 0.005 | <dl | <dl | <dl | <dl | 0.47 | 0.318 | 0.1 | 0.046 | 5.38 | 5.823 |
| SL | T25 | S3 | <dl | <dl | 0.01 | 0.002 | 0.01 | 0.002 | <dl | <dl | <dl | <dl | 0.46 | 0.096 | 0.09 | 0.037 | 6.37 | 1.769 |
| 18:2n-6 | 18:3n-3 | 20:0 | 20:1n-9 | 20:2n-9 | 22:0 | |||||||||||||
| Host algae | Temperature | Salinity | mean | sd | mean | sd | mean | sd | mean | sd | mean | sd | mean | sd | ||||
| LD | T10 | S1 | 9.92 | 1.743 | 0.97 | 0.214 | 0.04 | 0.009 | 0.01 | 0.005 | 0.03 | 0.005 | 0.02 | 0.003 | ||||
| SL | T10 | S1 | 10.98 | 1.901 | 0.92 | 0.133 | 0.03 | 0.006 | 0.01 | 0.001 | 0.04 | 0.005 | 0.02 | 0.002 | ||||
| LD | T18 | S1 | 9.38 | 2.822 | 0.16 | 0.027 | 0.03 | 0.015 | 0.01 | 0.002 | 0.04 | 0.015 | 0.02 | 0.005 | ||||
| SL | T18 | S1 | 6.59 | 1.594 | 0.16 | 0.054 | 0.02 | 0.003 | 0.01 | 0.002 | 0.02 | 0.004 | 0.01 | 0.001 | ||||
| LD | T25 | S1 | 10.48 | 2.265 | 0.07 | 0.018 | 0.02 | 0.004 | 0.01 | 0.005 | 0.07 | 0.017 | 0.02 | 0.002 | ||||
| SL | T25 | S1 | 8.95 | 1.945 | 0.11 | 0.019 | 0.02 | 0.005 | 0.01 | 0.001 | 0.04 | 0.022 | 0.02 | 0.004 | ||||
| LD | T10 | S2 | 8.71 | 2.021 | 0.66 | 0.199 | 0.04 | 0.008 | 0.01 | 0.003 | 0.05 | 0.023 | 0.01 | 0.005 | ||||
| SL | T10 | S2 | 4.7 | 1.691 | 0.37 | 0.113 | 0.02 | 0.007 | 0.01 | 0.002 | 0.03 | 0.019 | 0.01 | 0.008 | ||||
| LD | T18 | S2 | 6.23 | 1.565 | 0.08 | 0.015 | 0.02 | 0.008 | <dl | 0.002 | 0.05 | 0.026 | 0.01 | 0.003 | ||||
| SL | T18 | S2 | 8.46 | 1.561 | 0.12 | 0.011 | 0.03 | 0.007 | 0.01 | 0.003 | 0.03 | 0.012 | 0.02 | 0.003 | ||||
| LD | T25 | S2 | 14.24 | 0.745 | 0.07 | 0.017 | 0.04 | 0.005 | 0.01 | 0.003 | 0.05 | 0.019 | 0.02 | 0.002 | ||||
| SL | T25 | S2 | 14.92 | 3.861 | 0.09 | 0.019 | 0.05 | 0.017 | 0.01 | 0.004 | 0.05 | 0.017 | 0.04 | 0.011 | ||||
| LD | T10 | S3 | 1.24 | 0.743 | 0.07 | 0.036 | <dl | <dl | <dl | <dl | 0.01 | 0.004 | <dl | <dl | ||||
| SL | T10 | S3 | 1.08 | 0.697 | 0.08 | 0.038 | <dl | <dl | <dl | <dl | <dl | <dl | <dl | <dl | ||||
| LD | T18 | S3 | 4.99 | 0.463 | 0.05 | 0.004 | 0.02 | 0.003 | 0.01 | 0.002 | 0.02 | 0.005 | 0.01 | 0.002 | ||||
| SL | T18 | S3 | 7.84 | 4.272 | 0.09 | 0.041 | 0.03 | 0.015 | 0.01 | 0.005 | 0.02 | 0.006 | 0.01 | 0.005 | ||||
| LD | T25 | S3 | 7.75 | 8.020 | 0.03 | 0.011 | 0.02 | 0.020 | 0.01 | 0.006 | 0.03 | 0.020 | 0.02 | 0.009 | ||||
| SL | T25 | S3 | 7.25 | 2.175 | 0.04 | 0.017 | 0.02 | 0.004 | 0.01 | 0.003 | 0.02 | 0.004 | 0.01 | 0.002 | ||||
| df | Sum of Square | Mean Squares | F | p-Value | ||
|---|---|---|---|---|---|---|
| Host algae (host) | 1 | 0.1020 | 0.1020 | 5.661 | 0.022774 | * |
| Temperature (temp) | 2 | 0.4321 | 0.2160 | 11.985 | 0.000102 | *** |
| Salinity (sal) | 2 | 1.2981 | 0.6491 | 36.009 | 2.57 × | *** |
| host:temp interaction | 2 | 0.8079 | 0.4040 | 22.411 | 4.76 × | *** |
| host:sal interaction | 2 | 0.0098 | 0.0049 | 0.272 | 0.763277 | n.s. |
| temp:sal interaction | 4 | 0.6356 | 0.1589 | 8.815 | 4.55 × | *** |
| host:temp:sal interaction | 4 | 0.0804 | 0.0201 | 1.115 | 0.364467 | n.s. |
| Residuals | 36 | 0.6489 | 0.0180 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Vallet, M.; Meziane, T.; Thiney, N.; Prado, S.; Hubas, C. Laminariales Host Does Impact Lipid Temperature Trajectories of the Fungal Endophyte Paradendryphiella salina (Sutherland.). Mar. Drugs 2020, 18, 379. https://doi.org/10.3390/md18080379
Vallet M, Meziane T, Thiney N, Prado S, Hubas C. Laminariales Host Does Impact Lipid Temperature Trajectories of the Fungal Endophyte Paradendryphiella salina (Sutherland.). Marine Drugs. 2020; 18(8):379. https://doi.org/10.3390/md18080379
Chicago/Turabian StyleVallet, Marine, Tarik Meziane, Najet Thiney, Soizic Prado, and Cédric Hubas. 2020. "Laminariales Host Does Impact Lipid Temperature Trajectories of the Fungal Endophyte Paradendryphiella salina (Sutherland.)" Marine Drugs 18, no. 8: 379. https://doi.org/10.3390/md18080379
APA StyleVallet, M., Meziane, T., Thiney, N., Prado, S., & Hubas, C. (2020). Laminariales Host Does Impact Lipid Temperature Trajectories of the Fungal Endophyte Paradendryphiella salina (Sutherland.). Marine Drugs, 18(8), 379. https://doi.org/10.3390/md18080379





