Hatsusamides A and B: Two New Metabolites Produced by the Deep-Sea-Derived Fungal Strain Penicillium steckii FKJ-0213
Abstract
1. Introduction
2. Results and Discussion
2.1. Structure Elucidation of 1 and 2
2.2. Anti-Malarial Activity of 1–4
2.3. Cytotoxicity of 1–4 against Five Human Cancer Cell Lines
3. Materials and Methods
3.1. General Experimental Procedures
3.2. Isolation and Identification of Strain FKI-0213
3.3. Fermentation of Strain FKJ-0213 and Isolation of 1–4
3.4. Hydrolysis of 1
3.5. X-ray Diffraction Analysis of 3
3.6. Cytotoxic Activity of 1–4 against Several Tumor Cell Lines
3.7. Anti-Malarial Activity of 1–4
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Nagel, D.W.; Pachler, K.G.R.; Steyn, P.S.; Wessels, P.; Gafner, G.; Kruger, G.J. X-ray structure of oxaline: A novel alkaloid from Penicillium oxalicum. J. Chem. Soc. Chem. Commun. 1974, 24, 1021–1022. [Google Scholar] [CrossRef]
- Hirano, A.; Iwai, Y.; Masuma, R.; Tei, K.; Ōmura, S. Neoxaline a new alkaloid produced by Aspergillus japonicus. Production, isolation and properties. J. Antibiot. 1979, 32, 781–785. [Google Scholar] [CrossRef] [PubMed]
- Konda, Y.; Onda, M.; Hirano, A.; Ōmura, S. Oxaline and neoxaline. Chem. Pharm. Bull. 1980, 28, 2987–2993. [Google Scholar] [CrossRef]
- Koizumi, Y.; Arai, Y.; Tomoda, H.; Ōmura, S. Oxaline, a fungal alkaloid, arrests the cell cycle in the M phase by inhibition of tublin polymerization. Biochim. Biohys. Acta 2004, 1693, 47–55. [Google Scholar] [CrossRef] [PubMed]
- Kanehisa, M.; Goto, S.; Furumichi, M.; Tanabe, M.; Hirakawa, M. KEGG for representation and analysis of molecular networks involving diseases and drugs. Nucleic Acids Res. 2010, 38, D355–D360. [Google Scholar] [CrossRef] [PubMed]
- Hong, J.H.; Jang, S.; Heo, Y.M.; Min, M.; Lee, H.; Lee, Y.M.; Lee, H.; Kim, J.J. Investigation of marine-derived fungal diversity and their wxploitable biological activities. Mar. Drugs 2015, 13, 4137–4155. [Google Scholar] [CrossRef] [PubMed]
- Daletos, G.; Ebrahim, W.; Ancheeva, E.; EI-Neketi, M.; Song, W.; Proksch, P. Natural products from deep-sea- derived fungi-A new source of novel bioactive compounds. Curr. Med. Chem. 2018, 25, 186–207. [Google Scholar] [CrossRef] [PubMed]
- Nagano, Y.; Nagahama, T. Fungal diversity in deep-sea extreme environments. Fungal Ecol. 2012, 5, 463–471. [Google Scholar] [CrossRef]
- Matsuo, H.; Nonaka, K.; Nagano, Y.; Yabuki, A.; Fujikura, K.; Takahashi, Y.; Ōmura, S.; Nakashima, T. New metabolites, sarcopodinols A and B, isolated form deep-sea derived fungal strain Sarcopodium sp. FKJ-0025. Biosci. Biotechnol. Biochem. 2018, 82, 1323–1326. [Google Scholar] [CrossRef] [PubMed]
- Matsuo, H.; Mokudai, T.; Higo, M.; Nonaka, K.; Nagano, Y.; Nagahama, T.; Niwano, Y.; Takahashi, Y.; Ōmura, S.; Nakashima, T. Cipralphelin, a new anti-oxidative N-cinnamoyl tripeptide produced by the deep-sea derived fungal strain Penicillium brevicompactum FKJ-0123. J. Antibiot. 2019, 72, 775–778. [Google Scholar] [CrossRef] [PubMed]
- Nakashima, T.; Takahashi, Y.; Ōmura, S. Search for new compounds from Kitasato microbial library by physicochemical screening. Biochem. Pharmacol. 2017, 134, 42–55. [Google Scholar] [CrossRef] [PubMed]
- Ando, K.; Azuma, T.; Matsuda, Y.; Morishita, Y.; Saito, Y. Compound gs-1302. Japan patent JPH07179391A, 22 December 1993. [Google Scholar]
- Davis, R.A.; Longden, J.; Avery, V.M.; Healy, P.C. The isolation, structure determination and cytotoxicity of the new fungal metabolite, trichodermamide C. Bioorg. Med. Chem. Lett. 2008, 18, 2836–2839. [Google Scholar] [PubMed]
- Kuramoto, M.; Yamada, K.; Shikano, M.; Yazawa, K.; Arimoto, H.; Okamura, T.; Uemura, D. Tanzawaic acids A, B, C, and D: Inhibitors of superoxide anion production from Penicillium citrinum. Chem. Lett. 1997, 9, 885–886. [Google Scholar]
- Capon, R.J.; Ratnayake, R.; Stewart, M.; Lacey, E.; Tennant, S.; Gill, J.H. Aspergillazines A-E: Novel heterocyclic dipeptides from an Australian strain of Aspergillus unilateralis. Org. Biomol. Chem. 2005, 3, 123–129. [Google Scholar] [CrossRef] [PubMed]
- Lin, Y.; Shao, Z.; Jiang, G.; Zhou, S.; Cai, J.; Vrijmoed, L.L.P.; Jones, E.B.G. Penicillazine, a unique quinone derivative with 4H-5,6-dihydro-1,2-oxazine ring system from the marine fungus Penicillium sp. (strain #386) from the South China Sea. Tetrahedron 2000, 56, 9607–9609. [Google Scholar]
- Iwatsuki, M.; Takada, S.; Mori, M.; Ishiyama, A.; Namatame, M.; Nishimura-Tsukashima, A.; Nonaka, K.; Masuma, R.; Otoguro, K.; Ōmura, S. In vitro and In vivo antimalarial activity of puberulic acid and its new analogs, viticolins A-C, produced by Penicillium sp. FKI-4410. J. Antibiot. 2011, 64, 183–188. [Google Scholar] [CrossRef] [PubMed]
- Altschul, S.F.; Medden, T.L.; Schaffer, A.A.; Zhang, J.; Zhang, Z.; Miller, W.; Lipman, D.J. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res. 1997, 25, 3389–3402. [Google Scholar] [CrossRef] [PubMed]
- Otoguro, K.; Kohana, A.; Manabe, C.; Ishiyama, A.; Ui, H.; Shiomi, K.; Yamada, H.; Ōmura, S. Potent antimalarial activities of polyether antibiotic, X-206. J. Antibiot. 2001, 54, 658–663. [Google Scholar] [CrossRef] [PubMed]
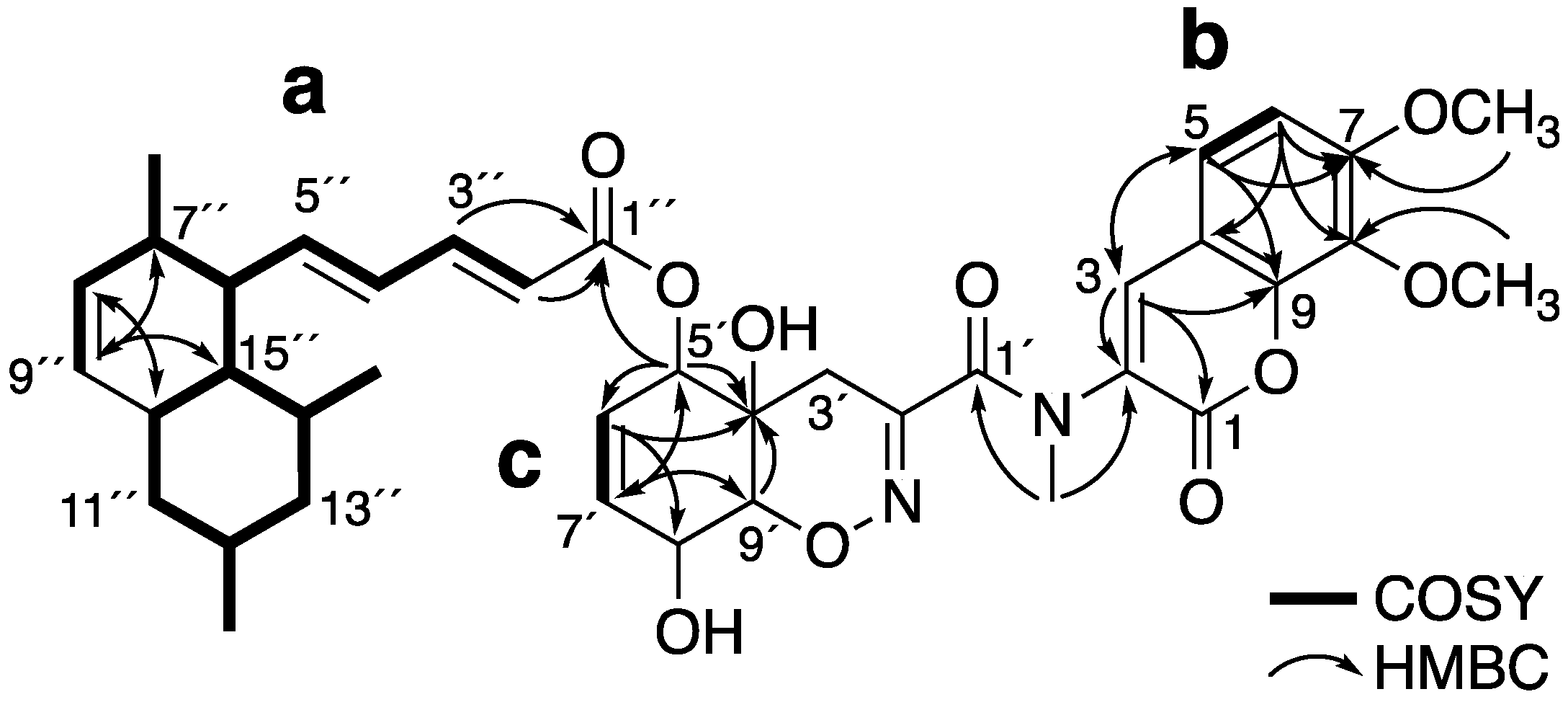
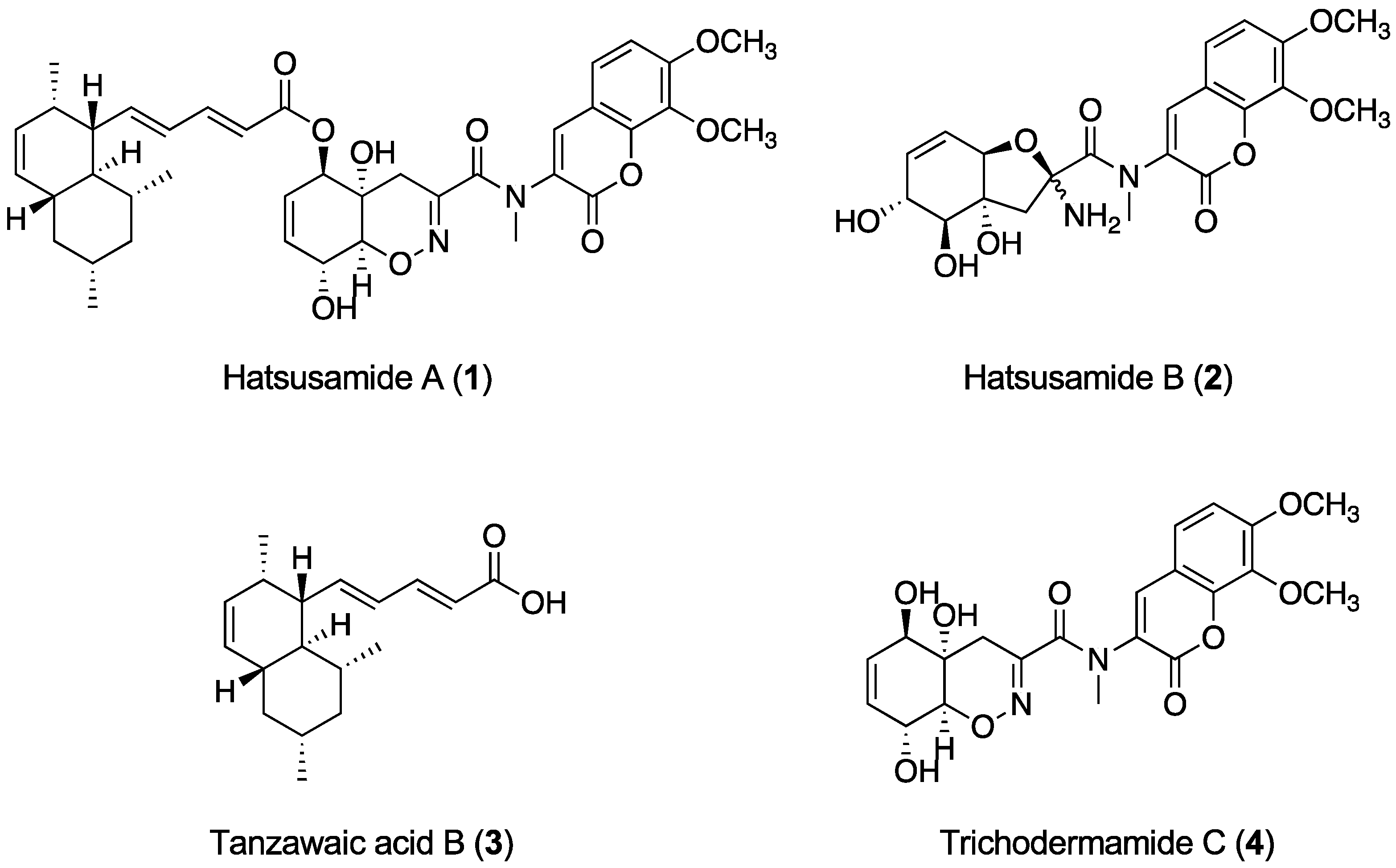
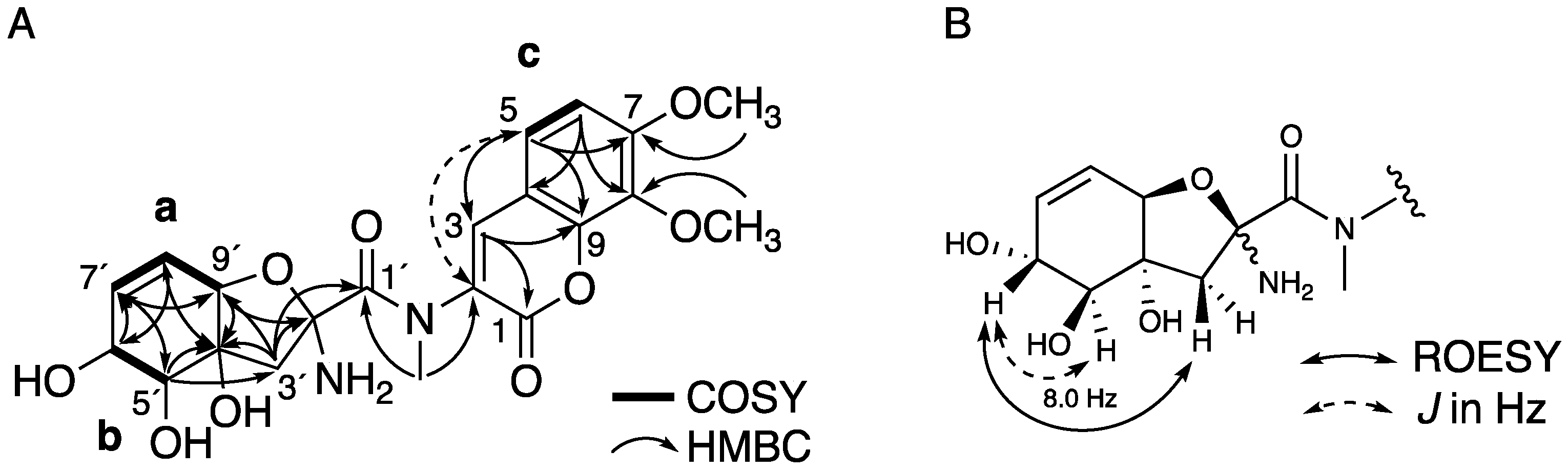
| Hatsusamide A (1) a | Hatsusamide B (2) b | |||
|---|---|---|---|---|
| Position | δH | δC | δH | δC |
| 1 | 159.9 | 166.8 | ||
| 2 | 129.6 | 131.2 | ||
| 3 | 7.91 (s, 1H) | 137.6 | 7.25 (s, 1H) | 119.9 |
| 4 | 114.8 | 115.5 | ||
| 5 | 7.39 (d, J = 9.2 Hz, 1H) | 124.2 | 6.86 (d, J = 8.9 Hz, 1H) | 126.4 |
| 6 | 7.13 (d, J = 9.2 Hz, 1H) | 110.5 | 6.59 (d, J = 8.9 Hz, 1H) | 104.5 |
| 7 | 156.4 | 155.5 | ||
| 8 | 136.9 | 137.5 | ||
| 9 | 147.4 | 150.9 | ||
| 7-OCH3 | 3.97 (s, 3H) | 56.8 | 3.87 (s, 3H) | 56.4 |
| 8-OCH3 | 3.90 (s, 3H) | 61.3 | 3.79 (s, 3H) | 61.1 |
| N-CH3 | 3.32 (s, 3H) | 37.0 | 2.90 (s, 3H) | 35.2 |
| 1′ | 166.3 | 166.0 | ||
| 2′ | 152.5 | 90.7 | ||
| 3′ | N.D. | N.D. | 2.09 (d, J = 14.0 Hz, 1H) | 40.6 |
| N.D. | 3.04 (d, J = 14.0 Hz, 1H) | |||
| 4′ | 67.4 | 83.2 | ||
| 5′ | 5.61 (m, 1H) | 76.9 | 3.70 (d, 8.0 Hz, 1H) | 78.8 |
| 6′ | 5.53 (ddd, J = 10.4, 2.1, 2.1 Hz, 1H) | 126.3 | 4.14 (m, 1H) | 72.1 |
| 7′ | 5.69 (d, J = 10.4 Hz, 1H) | 131.3 | 5.62 (ddd, J = 10.3, 2.4, 1.4 Hz, 1H) | 131.4 |
| 8′ | 4.07 (br s, 1H) | 67.9 | 5.50 (ddd, J = 10.3, 3.0, 2.6 Hz, 1H) | 127.8 |
| 9′ | 4.01 (br d, 7.4 Hz, 1H) | 83.5 | 4.66 (br dd, 3.0, 2.4 1H) | 86.9 |
| 1″ | 167.0 | |||
| 2″ | 5.96 (d, J = 15.2 Hz, 1H) | 119.5 | ||
| 3″ | 7.41 (dd, J = 15.2, 10.9 Hz, 1H) | 146.7 | ||
| 4″ | 6.30 (dd, J = 14.9, 10.9 Hz, 1H) | 127.5 | ||
| 5″ | 6.42 (dd, J = 14.9, 10.1 Hz, 1H) | 151.7 | ||
| 6″ | 2.49 (ddd, J = 10.1, 9.9, 5.5 Hz, 1H) | 50.1 | ||
| 7″ | 2.20 (m, 1H) | 37.7 | ||
| 8″ | 5.61 (m, 1H) | 133.0 | ||
| 9″ | 5.46 (ddd, J = 9.4, 1.9, 1.9 Hz, 1H) | 132.9 | ||
| 10″ | 1.81–1.88 (m, 1H) | 43.4 | ||
| 11″ | 0.74–0.83 (overlap, 1H) | 42.5 | ||
| 1.71–1.77 (m, 1H) | ||||
| 12″ | 1.51–1.59 (m, 1H) | 33.2 | ||
| 13″ | 0.74–0.83 (overlap, 1H) | 47.4 | ||
| 1.63–1.68 (m, 1H) | ||||
| 14″ | 1.37–1.45 (m, 1H) | 37.3 | ||
| 15″ | 0.97 (overlap, 1H) | 47.7 | ||
| 7″-CH3 | 0.98 (d, 7.4 Hz, 3H) | 16.8 | ||
| 12″-CH3 | 0.88 (d, 6.9 Hz, 3H) | 22.7 | ||
| 14″-CH3 | 0.96 (d, 6.3 Hz, 3H) | 23.1 | ||
| Compounds | IC50 (μM) Values Against Two Strains | |
|---|---|---|
| K1 | FCR3 | |
| 1 | 27.2 | 27.9 |
| 2 | >50 | >50 |
| 3 | 78.5 | 79.2 |
| 4 | >50 | >50 |
| Chloroquine a | 0.17 | 0.038 |
| Compounds | IC50 (μM) Values Against Five Human Tumor Cell Lines | ||||
|---|---|---|---|---|---|
| HeLa S3 | HT29 | A549 | H1299 | Panc1 | |
| 1 | 15.0 | 6.8 | 13.7 | 18.7 | 12.9 |
| 2 | >100 | >100 | >100 | >100 | >100 |
| 3 | >100 | >100 | >100 | >100 | >100 |
| 4 | >100 | >100 | >100 | >100 | >100 |
| Staurosporine | <2.0 | <2.0 | <2.0 | <2.0 | <2.0 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Matsuo, H.; Hokari, R.; Ishiyama, A.; Iwatsuki, M.; Higo, M.; Nonaka, K.; Nagano, Y.; Takahashi, Y.; Ōmura, S.; Nakashima, T. Hatsusamides A and B: Two New Metabolites Produced by the Deep-Sea-Derived Fungal Strain Penicillium steckii FKJ-0213. Mar. Drugs 2020, 18, 513. https://doi.org/10.3390/md18100513
Matsuo H, Hokari R, Ishiyama A, Iwatsuki M, Higo M, Nonaka K, Nagano Y, Takahashi Y, Ōmura S, Nakashima T. Hatsusamides A and B: Two New Metabolites Produced by the Deep-Sea-Derived Fungal Strain Penicillium steckii FKJ-0213. Marine Drugs. 2020; 18(10):513. https://doi.org/10.3390/md18100513
Chicago/Turabian StyleMatsuo, Hirotaka, Rei Hokari, Aki Ishiyama, Masato Iwatsuki, Mayuka Higo, Kenichi Nonaka, Yuriko Nagano, Yōko Takahashi, Satoshi Ōmura, and Takuji Nakashima. 2020. "Hatsusamides A and B: Two New Metabolites Produced by the Deep-Sea-Derived Fungal Strain Penicillium steckii FKJ-0213" Marine Drugs 18, no. 10: 513. https://doi.org/10.3390/md18100513
APA StyleMatsuo, H., Hokari, R., Ishiyama, A., Iwatsuki, M., Higo, M., Nonaka, K., Nagano, Y., Takahashi, Y., Ōmura, S., & Nakashima, T. (2020). Hatsusamides A and B: Two New Metabolites Produced by the Deep-Sea-Derived Fungal Strain Penicillium steckii FKJ-0213. Marine Drugs, 18(10), 513. https://doi.org/10.3390/md18100513






