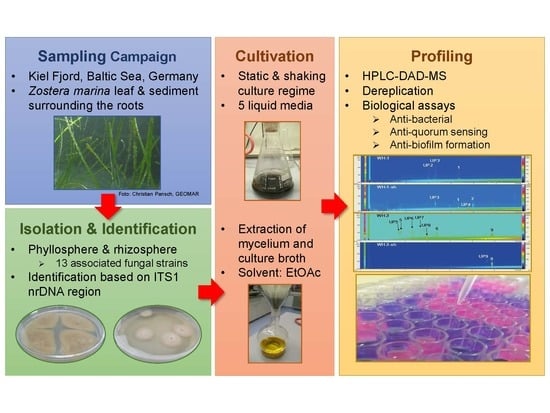
Rapid Metabolome and Bioactivity Profiling of Fungi Associated with the Leaf and Rhizosphere of the Baltic Seagrass Zostera marina
Abstract
:1. Introduction
2. Results
2.1. Identification and Extraction of Marine Fungi
2.2. Extraction, Metabolite, and Bioactivity Profile of Fungi Associated with Zostera marina
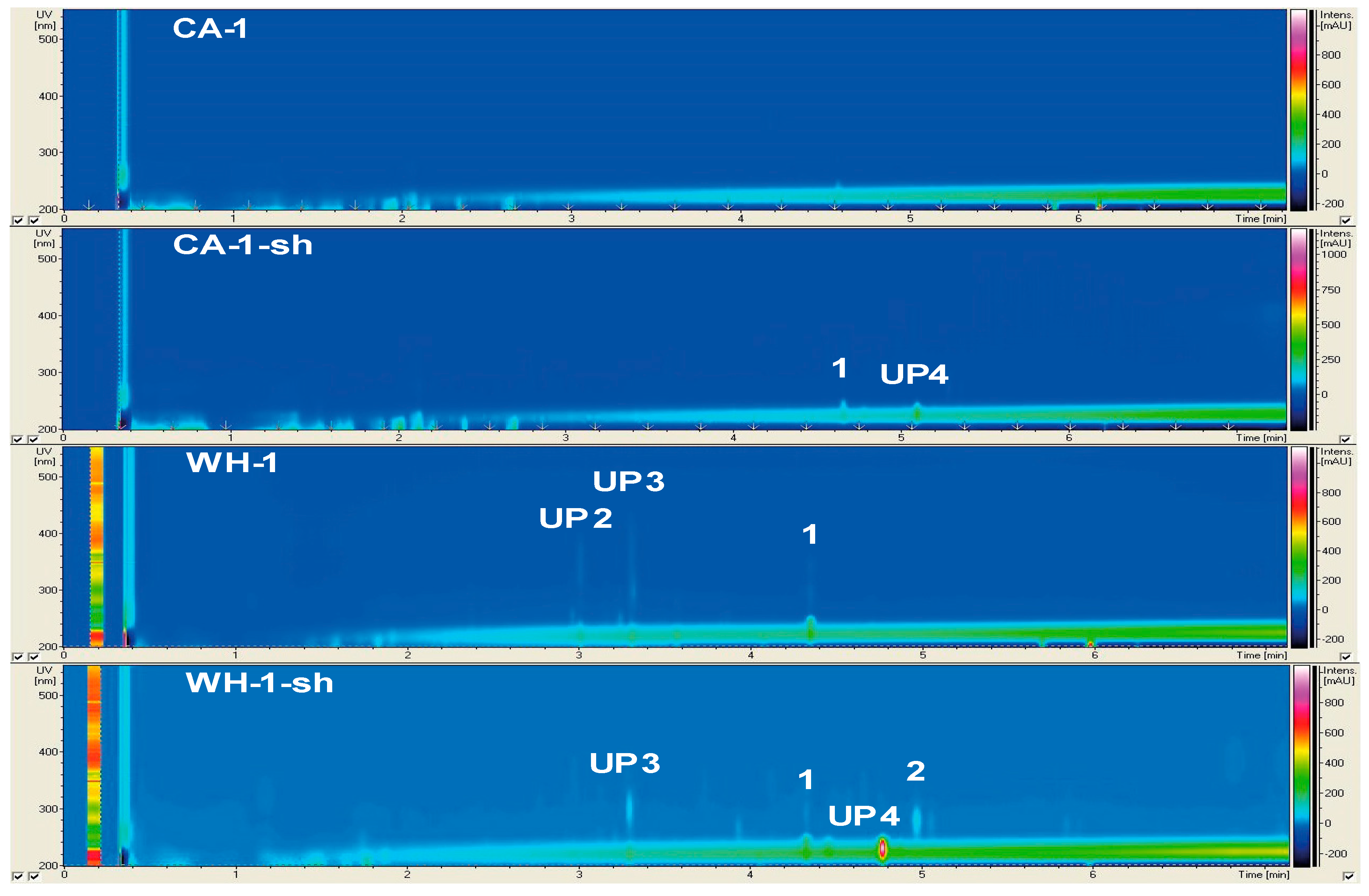
2.2.1. Secondary Metabolite Profile of Phoma macrostoma (Strain 1)
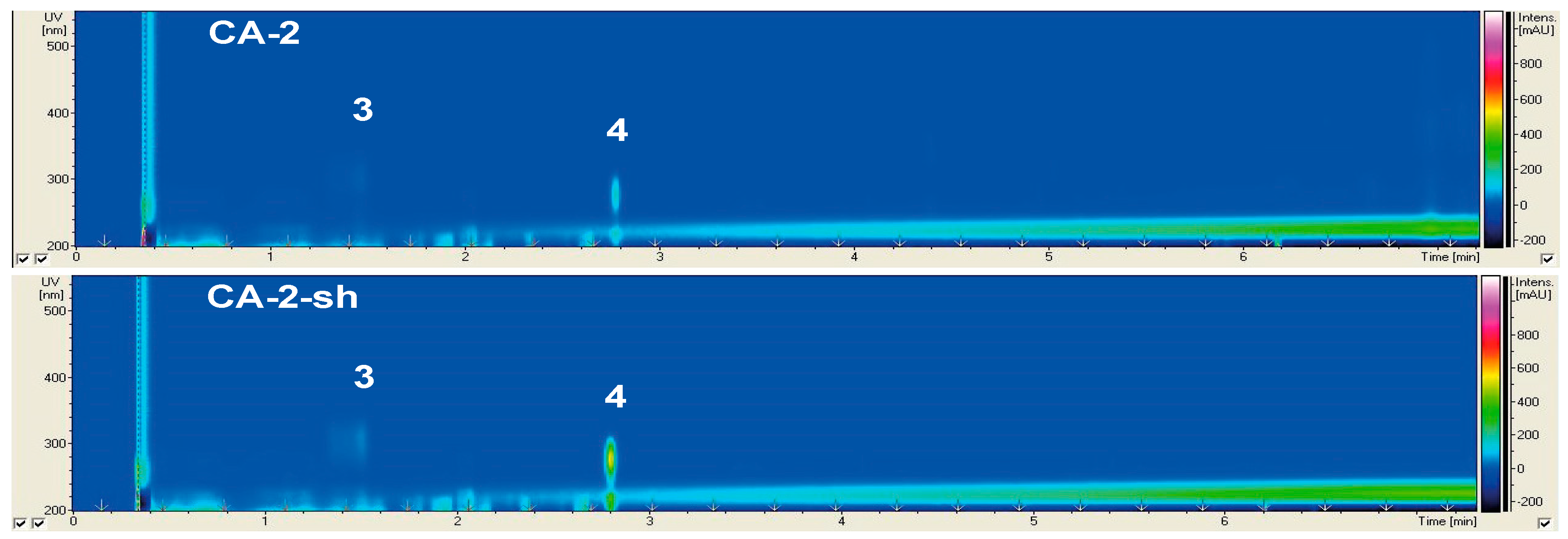
2.2.2. Secondary Metabolite Profile of Cladosporium langeronii (Strain 2)
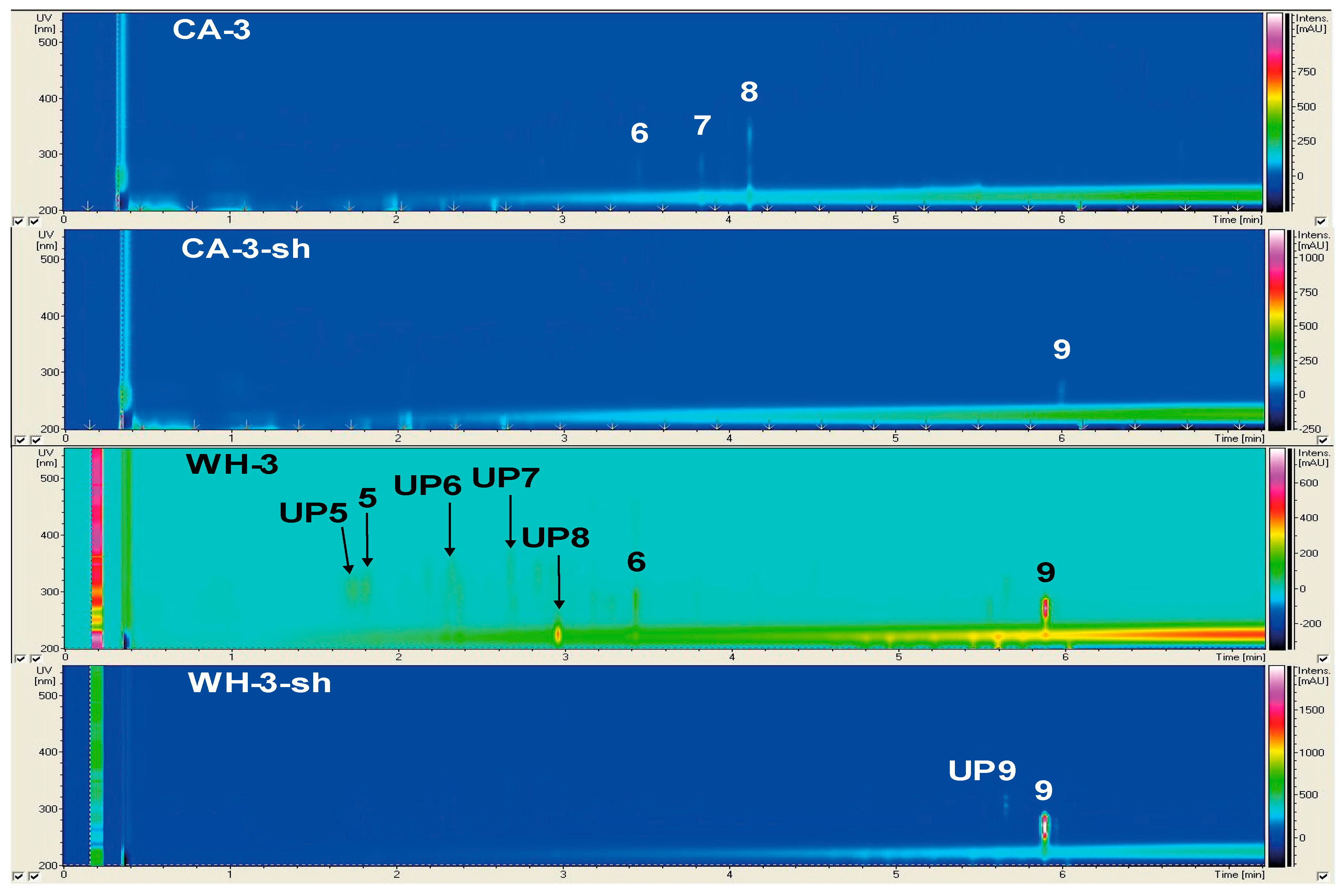
2.2.3. Secondary Metabolite Profile of Trichoderma harzianum (Strain 3)
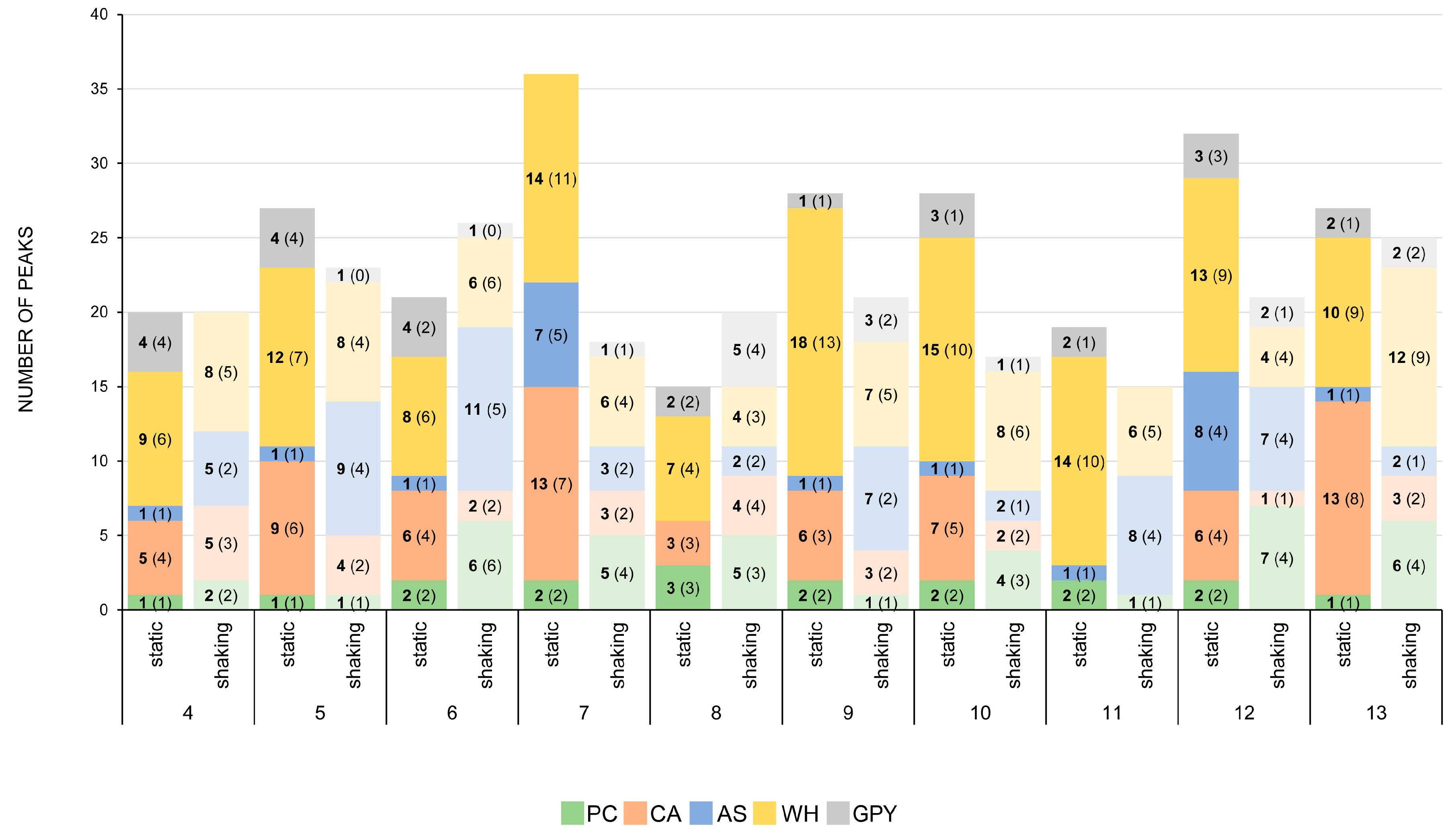
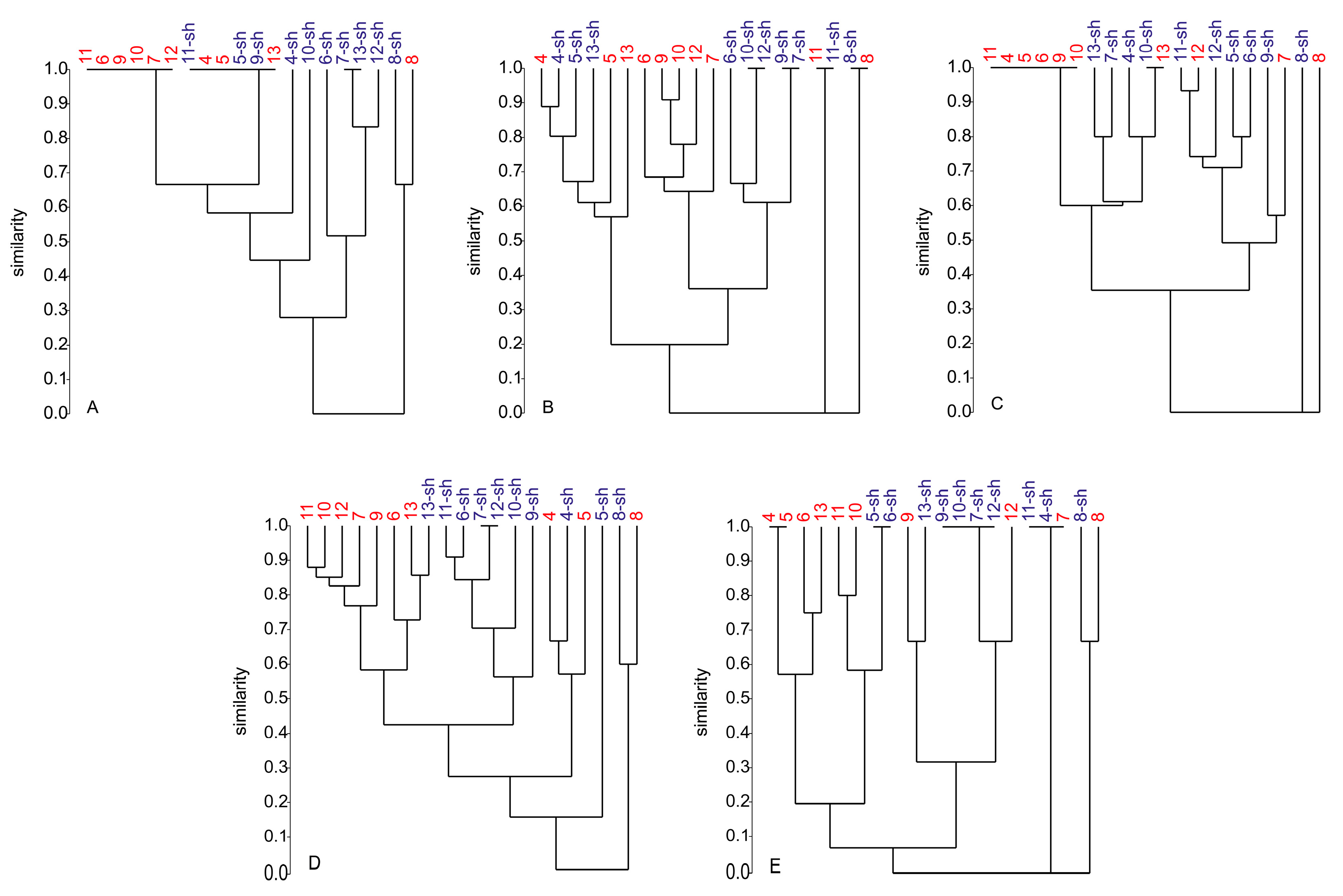
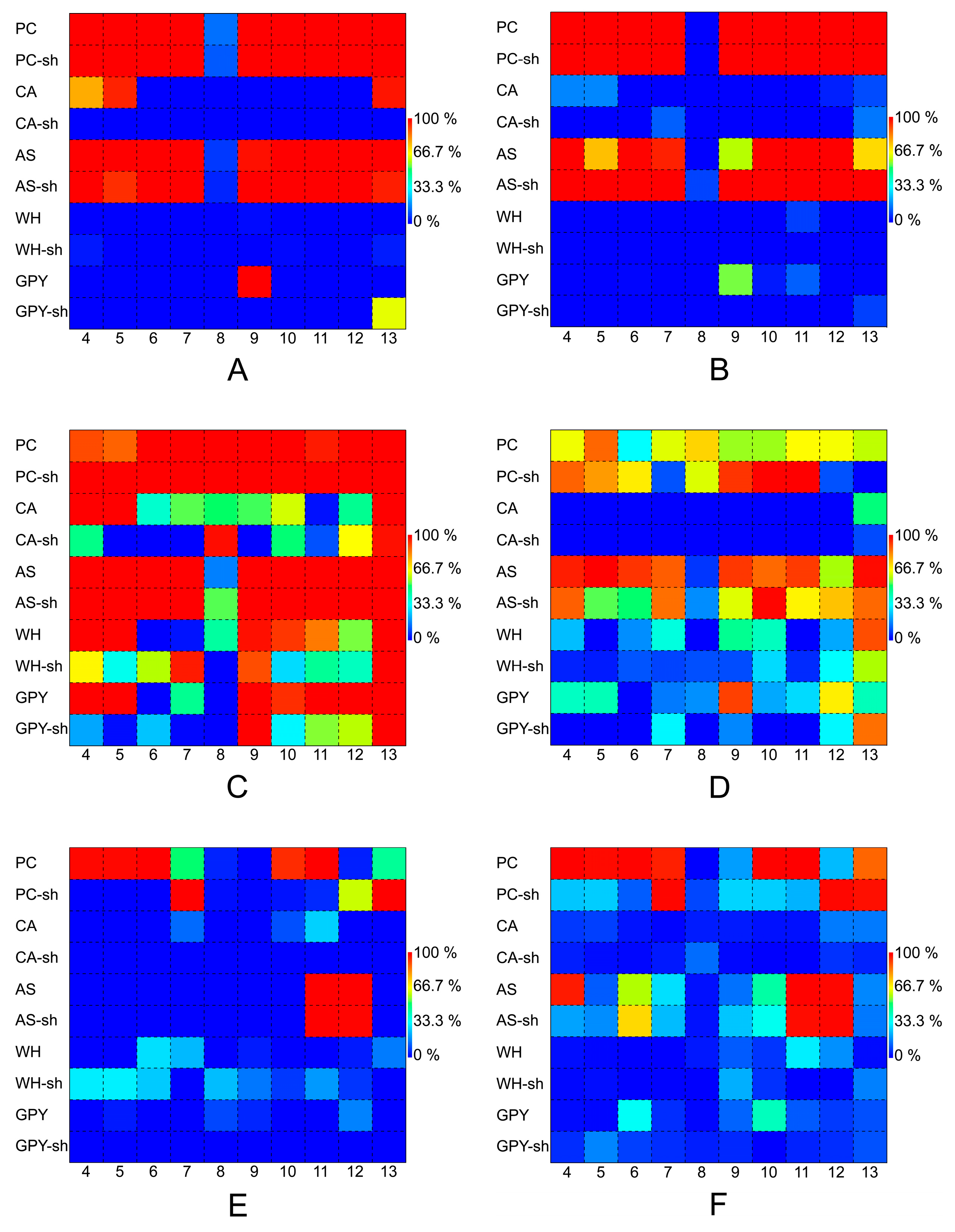
2.2.4. Secondary Metabolite Profile of Penicillium spp. (Strains 4–13)
2.2.5. Bioactivity of Penicillium Species
3. Discussions
3.1. Fungi Associated with the Leaf and Rhizosphere of Baltic Z. marina
3.2. Dereplication of Secondary Metabolite Profiles of Z. marina Associated Fungi
3.3. The Influence of Media and Culture Regime on the Secondary Metabolome of Z. marina Associated Fungi
3.4. Bioactivities of the Extracts of Z. marina Associated Fungi
4. Materials and Methods
4.1. Collection of Zostera marina
4.2. Isolation of Fungi
4.3. Identification of Fungal Strains
4.4. Cultivation and Extraction of Marine Fungi
4.5. LC-DAD-MS Analyses and Dereplication
4.6. Bioassays
4.6.1. Antibacterial Activity
4.6.2. Quorum Sensing Inhibition
4.6.3. Biofilm Inhibition
4.7. Statistics
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Lee, H.; Golicz, A.A.; Bayer, P.E.; Jiao, Y.; Tang, H.; Paterson, A.H.; Sablok, G.; Krishnaraj, R.R.; Chan, C.-K.K.; Batley, J.; et al. The genome of a southern hemisphere seagrass species (Zostera muelleri). Plant Physiol. 2016, 172, 272–283. [Google Scholar] [CrossRef] [PubMed]
- Bengtsson, M.M.; Bühler, A.; Brauer, A.; Dahlke, S.; Schubert, H.; Blindow, I. Eelgrass leaf surface microbiomes are locally variable and highly correlated with epibiotic eukaryotes. Front. Microbiol. 2017, 8, 1312. [Google Scholar] [CrossRef] [PubMed]
- Boström, C.; Baden, S.; Bockelmann, A.-C.; Dromph, K.; Fredriksen, S.; Gustafsson, C.; Krause-Jensen, D.; Möller, T.; Nielsen, S.L.; Olesen, B.; et al. Distribution, structure and function of Nordic eelgrass (Zostera marina) ecosystems: Implications for coastal management and conservation. Aquat. Conserv. Mar. Freshw. Ecosyst. 2014, 24, 410–434. [Google Scholar] [CrossRef] [PubMed]
- Olsen, J.L.; Rouzé, P.; Verhelst, B.; Lin, Y.-C.; Bayer, T.; Collen, J.; Dattolo, E.; De Paoli, E.; Dittami, S.; Maumus, F.; et al. The genome of the seagrass Zostera marina reveals angiosperm adaptation to the sea. Nature 2016, 530, 331–335. [Google Scholar] [CrossRef] [PubMed]
- Fahimipour, A.K.; Kardish, M.R.; Lang, J.M.; Green, J.L.; Eisen, J.A.; Stachowicz, J.J. Global-scale structure of the eelgrass microbiome. Appl. Environ. Microbiol. 2017, 83, e03391-16. [Google Scholar] [CrossRef]
- Guan, C.; Parrot, D.; Wiese, J.; Sönnichsen, F.D.; Saha, M.; Tasdemir, D.; Weinberger, F. Identification of rosmarinic acid and sulfated flavonoids as inhibitors of microfouling on the surface of eelgrass Zostera marina. Biofouling 2017, 33, 867–880. [Google Scholar] [CrossRef] [PubMed]
- Inaba, N.; Trainer, V.L.; Onishi, Y.; Ishii, K.I.; Wyllie-Echeverria, S.; Imai, I. Algicidal and growth-inhibiting bacteria associated with seagrass and macroalgae beds in Puget Sound, WA, USA. Harmful Algae 2017, 62, 136–147. [Google Scholar] [CrossRef] [PubMed]
- Papazian, S.; Parrot, D.; Burýšková, B.; Weinberger, F.; Tasdemir, D. Surface chemical defence of the eelgrass Zostera marina against microbial foulers. Sci. Rep. 2019, 9, 3323. [Google Scholar] [CrossRef]
- Vandenkoornhuyse, P.; Quaiser, A.; Duhamel, M.; Le Van, A.; Dufresne, A. The importance of the microbiome of the plant holobiont. New Phytol. 2015, 206, 1196–1206. [Google Scholar] [CrossRef]
- Ugarelli, K.; Chakrabarti, S.; Laas, P.; Stingl, U. The seagrass holobiont and its microbiome. Microorganisms 2017, 5, 81. [Google Scholar] [CrossRef]
- Ettinger, C.L.; Voerman, S.E.; Lang, J.M.; Stachowicz, J.J.; Eisen, J.A. Microbial communities in sediment from Zostera marina patches, but not the Z. marina leaf or root microbiomes, vary in relation to distance from patch edge. PeerJ 2017, 5, e3246. [Google Scholar] [CrossRef] [PubMed]
- Ettinger, C.L.; Williams, S.L.; Abbott, J.M.; Stachowicz, J.J.; Eisen, J.A. Microbiome succession during ammonification in eelgrass bed sediments. PeerJ 2017, 5, e3674. [Google Scholar] [CrossRef]
- Kirichuk, N.N.; Pivkin, M.V. Filamentous fungi associated with the seagrass Zostera marina Linnaeus, 1753 of Rifovaya Bay (Peter the Great Bay, the Sea of Japan). Russ. J. Mar. Biol. 2015, 41, 351–355. [Google Scholar] [CrossRef]
- Afiyatullov, S.S.; Leshchenko, E.V.; Sobolevskaya, M.P.; Gerasimenko, A.V.; Khudyakova, Y.V.; Kirichuk, N.N.; Mikhailov, V.V. New 3-[2′(R)-hydroxybutyl]-7-hydroxyphtalide from marine isolate of the fungus Penicillium claviforme. Chem. Nat. Compd. 2015, 51, 111–115. [Google Scholar] [CrossRef]
- Afiyatullov, S.S.; Leshchenko, E.V.; Sobolevskaya, M.P.; Denisenko, V.A.; Kirichuk, N.N.; Khudyakova, Y.V.; Hoai, T.P.T.; Dmitrenok, P.S.; Menchinskaya, E.S.; Pislyagin, E.A.; et al. New eudesmane sesquiterpenes from the marine-derived fungus Penicillium thomii. Phytochem. Lett. 2015, 14, 209–214. [Google Scholar] [CrossRef]
- Hyde, K.D.; Sarma, V.V.; Jones, E.B.G. Morphology and taxonomy of higher marine fungi. In Marine Mycology—A practical Approach, 1st ed.; Hyde, K.D., Pointing, S.B., Eds.; Fungal Diversity Press: Hong Kong, China, 2000; pp. 172–204. [Google Scholar]
- Ruby, E.G. Lessons from a cooperative, bacterial-animal association: The Vibrio fischeri—Euprymna scolopes light organ symbiosis. Annu. Rev. Microbiol. 1996, 50, 591–624. [Google Scholar] [CrossRef]
- Ruby, E.G.; Urbanowski, M.; Campbell, J.; Dunn, A.; Faini, M.; Gunsalus, R.; Lostroh, P.; Lupp, C.; McCann, J.; Millikan, D.; et al. Complete genome sequence of Vibrio fischeri: A symbiotic bacterium with pathogenic congeners. Proc. Natl. Acad. Sci. USA 2005, 102, 3004–3009. [Google Scholar] [CrossRef] [PubMed]
- Scheel, L.; Perone, V.B.; Larkin, R.L.; Kupel, R.E. The isolation and characterization of two phototoxic furanocoumarins (psoralens) from diseased celery. Biochemistry 1963, 2, 1127–1131. [Google Scholar] [CrossRef] [PubMed]
- Nippon Kayaku KK. Lipoxygenase Inhibitor, Antitumour NK-A-17-e-233-II—Produced by Cultivation of Phoma sp. NK-A-17-e-233. Japan Patent Office JP63216894-A, 1988. [Google Scholar]
- Zeeck, A. Agistatins, Metabolites from Fusarium sp., Process for Their Preparation and Their Use. Deutsches Patent- und Markenamt EP0492318A2, 13 December 1991. [Google Scholar]
- Schlörke, O.; Zeeck, A. Orsellides A–E: An example for 6-deoxyhexose derivatives produced by fungi. Eur. J. Org. Chem. 2006, 2006, 1043–1049. [Google Scholar] [CrossRef]
- Hirota, A.; Suzuki, A.; Aizawa, K.; Tamura, S. Structure of Cyl-2, a novel cyclotetrapeptide from Cylindrocladium scoparium. Agric. Biol. Chem. 1973, 37, 955–956. [Google Scholar] [CrossRef]
- Hayashi, H.; Nakatani, T.; Inoue, Y.; Nakayama, M.; Nozaki, H. New dihydroquinolinone toxic to Artemia salina produced by Penicillium sp. NTC-47. Biosci. Biotechnol. Biochem. 1997, 61, 914–916. [Google Scholar] [CrossRef]
- Dickinson, J.M.; Hanson, J.R.; Hitchcock, P.B.; Claydon, N. Structure and biosynthesis of harzianopyridone, an antifungal metabolite of Trichoderma harzianum. J. Chem. Soc. Perkin Trans. 1 1989, 11, 1885–1887. [Google Scholar] [CrossRef]
- Robins, M.J.; Robins, R.K. Purine nucleosides. XXIV. A new method for the synthesis guanine nucleosides. The preparation of 2′-deoxy-α and β-guanosines and the corresponding N2-methyl derivatives. J. Phys. Chem. 1969, 34, 2160–2163. [Google Scholar] [CrossRef]
- Newell, S.Y. Fungi and bacteria in or on leaves of eelgrass (Zostera marina L.) from Chesapeake Bay. Appl. Environ. Microbiol. 1981, 41, 1219–1224. [Google Scholar]
- Kossuga, M.H.; Romminger, S.; Xavier, C.; Milanetto, M.C.; do Valle, M.C.; Pimenta, E.F.; Morais, R.P.; de Carvalho, E.; Mizuno, C.M.; Coradello, L.F.C.; et al. Evaluating methods for the isolation of marine-derived fungal strains and production of bioactive secondary metabolites. Braz. J. Pharmacogn. 2012, 22, 257–267. [Google Scholar] [CrossRef] [Green Version]
- Cúcio, C.; Engelen, A.H.; Costa, R.; Muyzer, G. Rhizosphere microbiomes of European seagrasses are selected by the plant, but are not species specific. Front. Microbiol. 2016, 7, 440. [Google Scholar] [CrossRef]
- Pollard, P.C.; Moriarty, D.J.W. Organic carbon decomposition, primary and bacterial productivity, and sulphate reduction, in tropical seagrass beds of the Gulf of Carpentaria, Australia. Mar. Ecol. Prog. Ser. 1991, 69, 149–159. [Google Scholar] [CrossRef]
- Brodersen, K.E.; Nielsen, D.A.; Ralph, P.J.; Kühl, M. Oxic microshield and local pH enhancement protects Zostera muelleri from sediment derived hydrogen sulphide. New Phytol. 2015, 205, 1264–1276. [Google Scholar] [CrossRef]
- Brodersen, K.E.; Siboni, N.; Nielsen, D.A.; Pernice, M.; Ralph, P.J.; Seymour, J.; Kühl, M. Seagrass rhizosphere microenvironment alters plant-associated microbial community composition. Environ. Microbiol. 2018, 20, 2854–2864. [Google Scholar] [CrossRef]
- Bode, H.B.; Bethe, B.; Höfs, R.; Zeeck, A. Big effects from small changes: Possible ways to explore nature’s chemical diversity. Chembiochem 2002, 3, 619–627. [Google Scholar] [CrossRef]
- Rai, M.; Deshmukh, P.; Gade, A.; Ingle, A.; Kövics, G.J.; Irinyi, L. Phoma saccardo: Distribution, secondary metabolite production and biotechnological applications. Crit. Rev. Microbiol. 2009, 35, 182–196. [Google Scholar] [CrossRef]
- Bubnova, E.N. Fungal diversity in bottom sediments of the Kara Sea. Bot. Mar. 2010, 53, 595–600. [Google Scholar] [CrossRef]
- Zhang, X.-Y.; Tang, G.-L.; Xu, X.-Y.; Nong, X.-H.; Qi, S.-H. Insights into deep-sea sediment fungal communities from the East Indian Ocean using targeted environmental sequencing combined with traditional cultivation. PLoS ONE 2014, 9, e109118. [Google Scholar] [CrossRef]
- Garg, B.J.; Saraswat, A.; Bhatia, A.; Katare, O.P. Topical treatment in vitiligo and the potential uses of new drug delivery systems. Indian J. Dermatol. Venereol. Leprol. 2010, 76, 231–238. [Google Scholar] [CrossRef]
- Leite, V.C.; Santos, R.F.; Lee, C.C.; Guillo, L.A. Psoralen derivatives and longwave ultraviolet irradiation are active in vitro against human melanoma cell line. J. Photochem. Photobiol. 2004, 76, 49–53. [Google Scholar] [CrossRef]
- Venkatachalam, A.; Govinda Rajulu, M.B.; Thirunavukkarasu, N.; Suryanarayanan, T.S. Endophytic fungi of marine algae and seagrasses: A novel source of chitin modifying enzymes. Mycosphere 2015, 6, 345–355. [Google Scholar] [CrossRef]
- Panno, L.; Voyron, S.; Anastasi, A.; Mussat Sartor, R.; Varese, G.C. Biodiversity of marine fungi associated with the seagrass Posidonia oceanica: An ecological and biotechnological perspective. Biol. Mar. Mediterr. 2011, 18, 85–88. [Google Scholar]
- Panno, L.; Bruno, M.; Voyron, S.; Anastasi, A.; Gnavi, G.; Miserere, L.; Varese, G.C. Diversity, ecological role and potential biotechnological applications of marine fungi associated to the seagrass Posidonia oceanica. New Biotechnol. 2013, 30, 685–694. [Google Scholar] [CrossRef]
- Miao, F.P.; Liang, X.-R.; Yin, X.-L.; Wang, G.; Ji, N.-Y. Absolute configurations of unique harziane diterpenes from Trichoderma species. Org. Lett. 2012, 14, 3815–3817. [Google Scholar] [CrossRef]
- Hermosa, R.; Viterbo, A.; Chet, I.; Monte, E. Plant-beneficial effects of Trichoderma and of its genes. Microbiology 2012, 158, 17–25. [Google Scholar] [CrossRef]
- Hirota, A.; Suzuki, A.; Suzuki, H.; Tamura, S. Isolation and biological activity of cyl-2, a metabolite of Cylindrocladium scoparium. Agric. Biol. Chem. 1973, 37, 643–647. [Google Scholar] [CrossRef]
- Kusano, M.; Koshino, H.; Uzawa, J.; Fujioka, S.; Kawano, T.; Kimura, Y. Nematicidal alkaloids and related compounds produced by the fungus Penicillium cf. simplicissimum. Biosci. Biotechnol. Biochem. 2000, 64, 2559–2568. [Google Scholar] [CrossRef]
- Cutler, H.G.; Jacyno, J.M. Biological activity of (–)-harziano-pyridone isolated from Trichoderma harzianum. Agric. Biol. Chem. 1991, 55, 2629–2631. [Google Scholar] [CrossRef]
- Roslan, H.A.; Ngo, C.S.; Muid, S. Genetic diversity of Penicillium species isolated from various sources in Sarawak, Malaysia. J. Cell Mol. Biol. 2010, 7, 13–23. [Google Scholar]
- Rateb, M.E.; Ebel, R. Secondary metabolites of fungi from marine habitats. Nat. Prod. Rep. 2011, 28, 290–344. [Google Scholar] [CrossRef]
- Yurchenko, A.N.; Smetanina, O.F.; Ivanets, E.V.; Kalinovsky, A.I.; Khudyakova, Y.V.; Kirichuk, N.N.; Popov, R.S.; Bokemeyer, C.; von Amsberg, G.; Chingizova, E.A.; et al. Pretrichodermamides D–F from a marine algicolous fungus Penicillium sp. KMM 4672. Mar. Drugs 2016, 14, 122. [Google Scholar] [CrossRef]
- Bu, Y.-Y.; Yamazaki, H.; Takahashi, O.; Kirikoshi, R.; Ukai, K.; Namikoshi, M. Penicyrones A and B, an epimeric pair of α-pyrone-type polyketides produced by the marine-derived Penicillium sp. J. Antibiot. 2016, 69, 57–61. [Google Scholar] [CrossRef]
- Guo, W.; Zhang, Z.; Zhu, T.; Gu, Q.; Li, D. Penicyclones A–E, antibacterial polyketides from the deep-sea-derived fungus Penicillium sp. F23-2. J. Nat. Prod. 2015, 78, 2699–2703. [Google Scholar] [CrossRef]
- Puel, O.; Galtier, P.; Oswald, I.P. Biosynthesis and toxicological effects of patulin. Toxins 2010, 2, 613–631. [Google Scholar] [CrossRef]
- Prandini, A.; Tansini, G.; Sigolo, S.; Filippi, L.; Laporta, M.; Piva, G. On the occurrence of aflatoxin M1 in milk and dairy products. Food Chem. Toxicol. 2009, 47, 984–991. [Google Scholar] [CrossRef]
- Brady, S.F.; Wagenaar, M.M.; Singh, M.P.; Janso, J.E.; Clardy, J. The cytosporones, new octaketide antibiotics isolated from an endophytic fungus. Org. Lett. 2000, 2, 4043–4046. [Google Scholar] [CrossRef]
- Roll, D.M.; Manning, J.K.; Carter, G.T. Hongoquercins A and B, new sesquiterpenoid antibiotics: Isolation, structure elucidation, and antibacterial activity. J. Antibiot. 1998, 51, 635–639. [Google Scholar] [CrossRef]
- Zheng, C.-J.; Kim, C.-J.; Bae, K.S.; Kim, Y.-H.; Kim, W.-G. Bionectins A-C, epidithiodioxopiperazines with anti-MRSA activity, from Bionectra byssicola F120. J. Nat. Prod. 2006, 69, 1816–1819. [Google Scholar] [CrossRef]
- Weber, H.A.; Baenziger, N.C.; Gloer, J.B. Podosporin A: A novel antifungal metabolite from the coprophilous fungus Podospora decipiens (Wint.) Niessl. J. Org. Chem. 1988, 53, 4567–4569. [Google Scholar] [CrossRef]
- Suzuki, S.; Hosoe, T.; Nozawa, K.; Kawai, K.I.; Yaguchi, T.; Udagawa, S.I. Antifungal substances against pathogenic fungi, talaroconvolutins, from Talaromyces convolutes. J. Nat. Prod. 2000, 63, 768–772. [Google Scholar] [CrossRef]
- TePaske, M.R.; Gloer, J.B.; Wicklow, D.T.; Dowd, P.F. Tubingensin A: An antiviral carbazole alkaloid from the sclerotia of Aspergillus tubingensis. J. Org. Chem. 1989, 54, 4743–4746. [Google Scholar] [CrossRef]
- TePaske, M.R.; Gloer, J.B. The structure of tubingensin B: A cytotoxic carbazole alkaloid from the sclerotia of Aspergillus tubingensis. Tetrahedron Lett. 1989, 30, 5965–5968. [Google Scholar] [CrossRef]
- Ishikawa, Y.; Morimoto, K.; Iseki, S. Atrovenetin as a potent antioxidant compound from Penicillium species. J. Am. Oil Chem. Soc. 1991, 68, 666–668. [Google Scholar] [CrossRef]
- Ling, K.H.; Liou, H.-H.; Yang, C.-M.; Yang, C.-K. Isolation, chemical structure, acute toxicity, and some physicochemical properties of territrem C from Aspergillus terreus. Appl. Environ. Microbiol. 1984, 47, 98–100. [Google Scholar]
- Kyriakidis, N.; Waight, E.S.; Day, J.B.; Mantle, P.G. Novel metabolites from Penicillium crustosum, including penitrem E, a tremorgenic mycotoxin. Appl. Environ. Microbiol. 1981, 42, 61–62. [Google Scholar]
- Penn, J.; Biddle, J.R.; Mantle, P.G.; Bilton, J.N.; Sheppard, R.N. Pennigritrem, a naturally-occurring penitrem A analogue with novel cyclisation in the diterpenoid moiety. J. Chem. Soc. Perkin Trans. 1 1992, 1, 23–26. [Google Scholar] [CrossRef]
- Bashyal, B.P.; Kithsiri Wijeratne, E.M.; Faeth, S.H.; Leslie Gunatilaka, A.A. Globosumones A-C, cytotoxic orsellinic acid esters from the Sonoran Desert endophytic fungus Chaetomium globosum. J. Nat. Prod. 2005, 68, 724–728. [Google Scholar] [CrossRef]
- Lin, Z.-J.; Lu, Z.-Y.; Zhu, T.-J.; Fang, Y.-C.; Gu, Q.-Q.; Zhu, W.-M. Penicillenols from Penicillium sp. GQ-7, an endophytic fungus associated with Aegiceras corniculatum. Chem. Pharm. Bull. 2008, 56, 217–221. [Google Scholar] [CrossRef]
- Fill, T.P.; Pallinib, H.F.; da Silva Amaral, L.; da Silva, J.V.; Bidóia, D.L.; Peron, F.; Garcia, F.P.; Nakamura, C.V.; Rodrigues-Filho, E. Copper and manganese cations alter secondary metabolism in the fungus Penicillium brasilianum. J. Braz. Chem. Soc. 2016, 27, 1444–1451. [Google Scholar] [CrossRef]
- Gao, S.-S.; Shang, Z.; Li, X.-M.; Li, C.-S.; Cui, C.-M.; Wang, B.-G. Secondary metabolites produced by solid fermentation of the marine-derived fungus Penicillium commune QSD-17. Biosci. Biotechnol. Biochem. 2012, 76, 358–360. [Google Scholar] [CrossRef]
- Rodríguez-Ortiz, R.; Mehta, B.J.; Avalos, J.; Limón, M.C. Stimulation of bikaverin production by sucrose and by salt starvation in Fusarium fujikuroi. Appl. Microbiol. Biotechnol. 2010, 85, 1991–2000. [Google Scholar] [CrossRef]
- Fan, B.; Parrot, D.; Blümel, M.; Labes, A.; Tasdemir, D. Influence of OSMAC-based cultivation in metabolome and anticancer activity of fungi associated with the brown alga Fucus vesiculosus. Mar. Drugs 2019, 17, 67. [Google Scholar] [CrossRef]
- Davis, N.D.; Diener, U.L.; Eldridge, D.W. Production of aflatoxins B1 and G1 by Aspergillus flavus in a semisynthetic medium. Appl. Environ. Microbiol. 1966, 14, 378–380. [Google Scholar]
- Chang, S.-C.; Wei, Y.-H.; Wei, D.-L.; Chen, Y.-Y.; Jong, S.-C. Factors affecting the production of eremofortin C and PR toxin in Penicillium roqueforti. Appl. Environ. Microbiol. 1991, 57, 2581–2585. [Google Scholar]
- Wang, L.; Ridgway, D.; Gu, T.; Moo-Young, M. Bioprocessing strategies to improve heterologous protein production in filamentous fungal fermentations. Biotechnol. Adv. 2005, 23, 115–129. [Google Scholar] [CrossRef]
- Bigelis, R.; He, H.; Yang, H.Y.; Chang, L.-P.; Greenstein, M. Production of fungal antibiotics using polymeric solid supports in solid-state and liquid fermentation. J. Ind. Microbiol. Biotechnol. 2006, 33, 815–826. [Google Scholar] [CrossRef]
- Guo, W.; Peng, J.; Zhu, T.; Gu, Q.; Keyzers, R.A.; Li, D. Sorbicillamines A-E, nitrogen-containing sorbicillinoids from the deep-sea-derived fungus Penicillium sp. F23−2. J. Nat. Prod. 2013, 76, 2106–2112. [Google Scholar] [CrossRef]
- Austin, B.; Austin, D.; Sutherland, R.; Thompson, F.; Swings, J. Pathogenicity of Vibrios to rainbow trout (Oncorhynchus mykiss, Walbaum) and Artemia nauplii. Environ. Microbiol. 2005, 7, 1488–1495. [Google Scholar] [CrossRef]
- Mikkelsen, H.; Lund, V.; Martinsen, L.-C.; Gravningen, K.; Schrøder, M.B. Variability among Vibrio anguillarum O2 isolates from Atlantic cod (Gadus morhua L.): Characterisation and vaccination studies. Aquaculture 2007, 266, 16–25. [Google Scholar] [CrossRef]
- Hickey, M.E.; Lee, J.-L. A comprehensive review of Vibrio (Listonella) anguillarum: Ecology, pathology and prevention. Rev. Aquac. 2018, 10, 585–610. [Google Scholar] [CrossRef]
- Li, X.; Dierckens, K.; Bossier, P.; Defoirdt, T. The impact of quorum sensing on the virulence of Vibrio anguillarum towards gnotobiotic sea bass (Dicentrarchus labrax) larvae. Aquac. Res. 2018, 49, 3686–3689. [Google Scholar] [CrossRef]
- Sawabe, T.; Tanaka, R.; Iqbal, M.M.; Tajima, K.; Ezura, Y.; Ivanova, E.P.; Christen, R. Assignment of Alteromonas elyakovii KMM 162T and five strains isolated from spotwounded fronds of Laminaria japonica to Pseudoalteromonas elyakovii comb. nov. and the extended description of the species. Int. J. Syst. Evol. Microbiol. 2000, 50, 256–271. [Google Scholar] [CrossRef]
- Shaaban, M.; Elgam, A.; Habib, E.E. Biotechnological applications of quorum sensing inhibition as novel therapeutic strategies for multidrug resistant pathogens. Microb. Pathog. 2019, 127, 138–143. [Google Scholar] [CrossRef]
- Dunlap, P.V. Quorum regulation of luminescence in Vibrio fischeri. J. Molec. Microbiol. Biotechnol. 1999, 1, 5–12. [Google Scholar]
- Kalia, V.C. Quorum sensing inhibitors: An overview. Biotechnol. Adv. 2013, 31, 224–245. [Google Scholar] [CrossRef]
- Gellatly, S.L.; Hancock, R.E.W. Pseudomonas aeruginosa: New insights into pathogenesis and host defenses. Pathog. Dis. 2013, 67, 159–173. [Google Scholar] [CrossRef]
- Rasmussen, T.B.; Skindersoe, M.E.; Bjarnsholt, T.; Phipps, R.K.; Christensen, K.B.; Jensen, P.O.; Andersen, J.B.; Koch, B.; Larsen, T.O.; Hentzer, M.; et al. Identity and effects of quorum-sensing inhibitors produced by Penicillium species. Microbiology 2005, 151, 1325–1340. [Google Scholar] [CrossRef]
- Ianiri, G.; Pinedo, C.; Fratianni, A.; Panfili, G.; Castoria, R. Patulin degradation by the biocontrol yeast Sporobolomyces sp. is an inducible process. Toxins 2017, 9, 61. [Google Scholar] [CrossRef]
- Kavanagh, F. Activities of twenty-two antibacterial substances against nine species of bacteria. J. Bacteriol. 1947, 54, 761–766. [Google Scholar]
- White, T.J.; Bruns, T.; Lee, S.; Taylor, J. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In PCR Protocols: A Guide to Methods and Applications, 1st ed.; Innis, M.A., Gelfand, D.H., Sninsky, J.J., White, T.J., Eds.; Academic Press Inc.: New York, NY, USA, 1990; pp. 315–322. ISBN 978-0123721815. [Google Scholar]
- Ramette, A. Multivariate analyses inmicrobial ecology. FEMS Microbiol. Ecol. 2007, 62, 142–160. [Google Scholar] [CrossRef]
| Strain Number | Origin | Morphology | Next Related Strain | Similarity (%) | GenBank Accession Number |
|---|---|---|---|---|---|
| 1 | L | Phoma sp. | Phoma macrostoma | 100 | MN166393 |
| 2 | S | Cladosporium sp. | Cladosporium langeronii | 99 | MN166394 |
| 3 | S | Trichoderma sp. | Trichoderma harzianum | 100 | MN166395 |
| 4 | S | Penicillium sp. | Penicillium antarcticum | 99 | MN172369 |
| 5 | S | Penicillium sp. | Penicillium antarcticum | 100 | MN166396 |
| 6 | S | Penicillium sp. | Penicillium antarcticum | 100 | MN166397 |
| 7 | S | Penicillium sp. | Penicillium antarcticum | 100 | MN166398 |
| 8 | S | Penicillium sp. | Penicillium atramentosum | 100 | MN166399 |
| 9 | S | Penicillium sp. | Penicillium atrovenetum | 100 | MN166400 |
| 10 | S | Penicillium sp. | Penicillium atrovenetum | 100 | MN166401 |
| 11 | S | Penicillium sp. | Penicillium atrovenetum | 100 | MN166402 |
| 12 | S | Penicillium sp. | Penicillium atrovenetum | 100 | MN166403 |
| 13 | S | Penicillium sp. | Penicillium atrovenetum | 100 | MN166404 |
| Peak/Compound | Medium | UVmax (nm) | m/z [M + H]+ | Identity | Reference |
|---|---|---|---|---|---|
| 1 | PC, PC-sh, CA-sh, WH, WH-sh | 220, 245, 292, 342 | 229.1 | Trioxsalen | [19] |
| 2 | WH-sh | 222, 279 | 277.2 | NK-A 17E-233II | [20] |
| 3 | CA, CA-sh | 221, 289, 310 | 211.1 | Agistatin D | [21] |
| 4 | CA, CA-sh | 205, 216, 222, 277 | 295.1 | Orsellide D | [22] |
| 5 | WH | 215, 306 | 311.3 | Orsellide C | [22] |
| 6 | CA, WH | 223, 275, 288 | 599.3 | Cyl-2 | [23] |
| 7 | CA | 223, 280 | 316.1 | Quinolinone B | [24] |
| 8 | CA | 227, 269, 334 | 282.1 | Harzianopyridone | [25] |
| 9 | PC, CA-sh, WH, WH-sh | 223, 261, 271, 281 | 282.2 | 2′-Deoxyribofuranosylguanine; 2N-Me | [26] |
| UP1 | PC-sh | 221, 292 | 325.2 | n.k. | |
| UP2 | PC, PC-sh, WH | 220, 275, 325, 360 | 273.1 | n.k. | |
| UP3 | PC, PC-sh, WH, WH-sh | 218, 296, 364 | 289.1 | n.k. | |
| UP4 | CA-sh, WH-sh | 232 | 279.3 | n.k. | |
| UP5 | WH | 215, 303 | 327.3 | n.k. | |
| UP6 | WH | 218, 297 | 312.2 | n.k. | |
| UP7 | WH | 220, 266, 276, 287 | 393.1 | n.k. | |
| UP8 | WH | 226 | 245.2 | n.k. | |
| UP9 | WH-sh | 222, 280, 292, 305, 319 | 626.2 | n.k. |
| Medium | Inhibition (%) of A. fischeri QS/Growth Inhibition | Inhibition (%) of Ps. aeruginosa Biofilm/Growth Inhibition | ||||
|---|---|---|---|---|---|---|
| Phoma macrostoma | Cladosporium langeronii | Trichoderma harzianum | Phoma macrostoma | Cladosporium langeronii | Trichoderma harzianum | |
| PC | 100/0 | 100/31 | 94/70 | 5 /24 | 0/0 | 0/3 |
| PC-sh | 97/0 | 93/37 | 47/0 | 0/6 | 0/7 | 17/3 |
| CA | 99/0 | 95/0 | 90/0 | 0/16 | 0/5 | 0/3 |
| CA-sh | 0/0 | 30/0 | 81/0 | 0/2 | 0/1 | 0/2 |
| AS | 73/0 | 0/0 | 47/5 | 0/8 | 0/7 | 0/9 |
| AS-sh | 53/0 | 77/0 | 76/0 | 12/1 | 0/3 | 11/4 |
| WH | 100/17 | 98/0 | 97/46 | 0/21 | 0/3 | 0/1 |
| WH-sh | 80/0 | 95/0 | 70/0 | 10/5 | 0/1 | 0/20 |
| GPY | 34 /0 | 1/13 | 20/0 | 0/5 | 0/0 | 0/2 |
| GPY-sh | 7/29 | 40/0 | 0/35 | 4/3 | 0/1 | 0/9 |
| Standard | 100A/100B | 100A/100B | 100A/100B | 100C/100C | 100C/100C | 100C/100C |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Petersen, L.-E.; Marner, M.; Labes, A.; Tasdemir, D. Rapid Metabolome and Bioactivity Profiling of Fungi Associated with the Leaf and Rhizosphere of the Baltic Seagrass Zostera marina. Mar. Drugs 2019, 17, 419. https://doi.org/10.3390/md17070419
Petersen L-E, Marner M, Labes A, Tasdemir D. Rapid Metabolome and Bioactivity Profiling of Fungi Associated with the Leaf and Rhizosphere of the Baltic Seagrass Zostera marina. Marine Drugs. 2019; 17(7):419. https://doi.org/10.3390/md17070419
Chicago/Turabian StylePetersen, Lars-Erik, Michael Marner, Antje Labes, and Deniz Tasdemir. 2019. "Rapid Metabolome and Bioactivity Profiling of Fungi Associated with the Leaf and Rhizosphere of the Baltic Seagrass Zostera marina" Marine Drugs 17, no. 7: 419. https://doi.org/10.3390/md17070419
APA StylePetersen, L.-E., Marner, M., Labes, A., & Tasdemir, D. (2019). Rapid Metabolome and Bioactivity Profiling of Fungi Associated with the Leaf and Rhizosphere of the Baltic Seagrass Zostera marina. Marine Drugs, 17(7), 419. https://doi.org/10.3390/md17070419