Tenofovir, Another Inexpensive, Well-Known and Widely Available Old Drug Repurposed for SARS-COV-2 Infection
Abstract
1. Introduction
2. Methodology and Literature Search Strategy
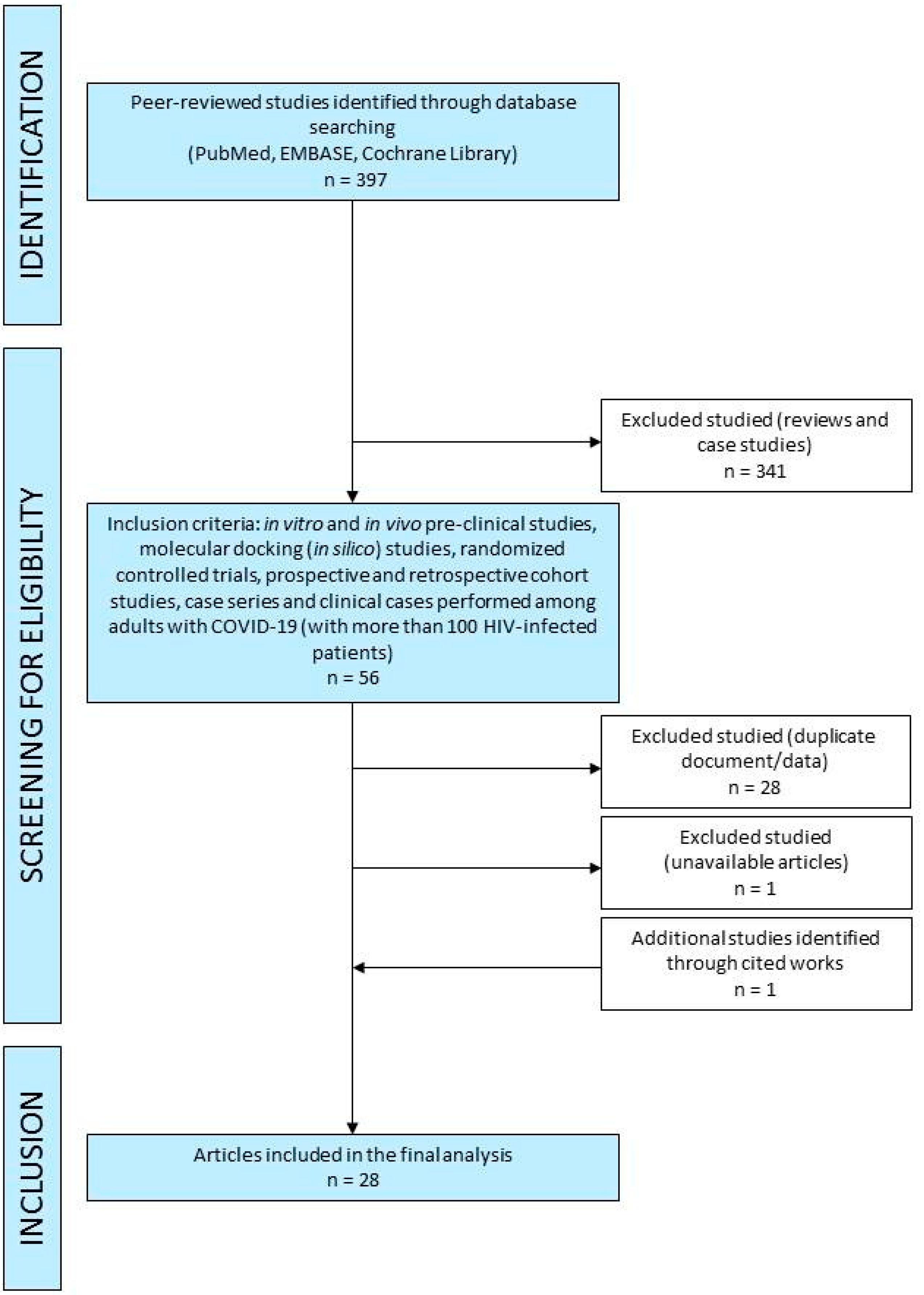
2.1. Literature Search
2.2. Selection of Studies
3. Pre-Clinical Studies
3.1. In Silico Studies
3.2. In Vitro Studies
3.3. In Vivo Studies
4. Clinical Data
4.1. Retrospective Studies
4.2. Clinical Trials
5. Discussion and Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Available online: https://www.who.int/news-room/q-a-detail/coronavirus-disease-(covid-19)-vaccines (accessed on 12 April 2021).
- Kumar, A.; Dowling, W.E.; Román, R.G.; Chaudhari, A.; Gurry, C.; Le, T.T.; Tollefson, S.; Clark, C.E.; Bernasconi, V.; Kristiansen, P.A. Status Report on COVID-19 Vaccines Development. Curr. Infect. Dis. Rep. 2021, 23, 1–12. [Google Scholar] [CrossRef]
- Golob, J.L.; Lugogo, N.; Lauring, A.S.; Lok, A.S. SARS-CoV-2 vaccines: A triumph of science and collaboration. JCI Insight 2021, 2021. [Google Scholar] [CrossRef]
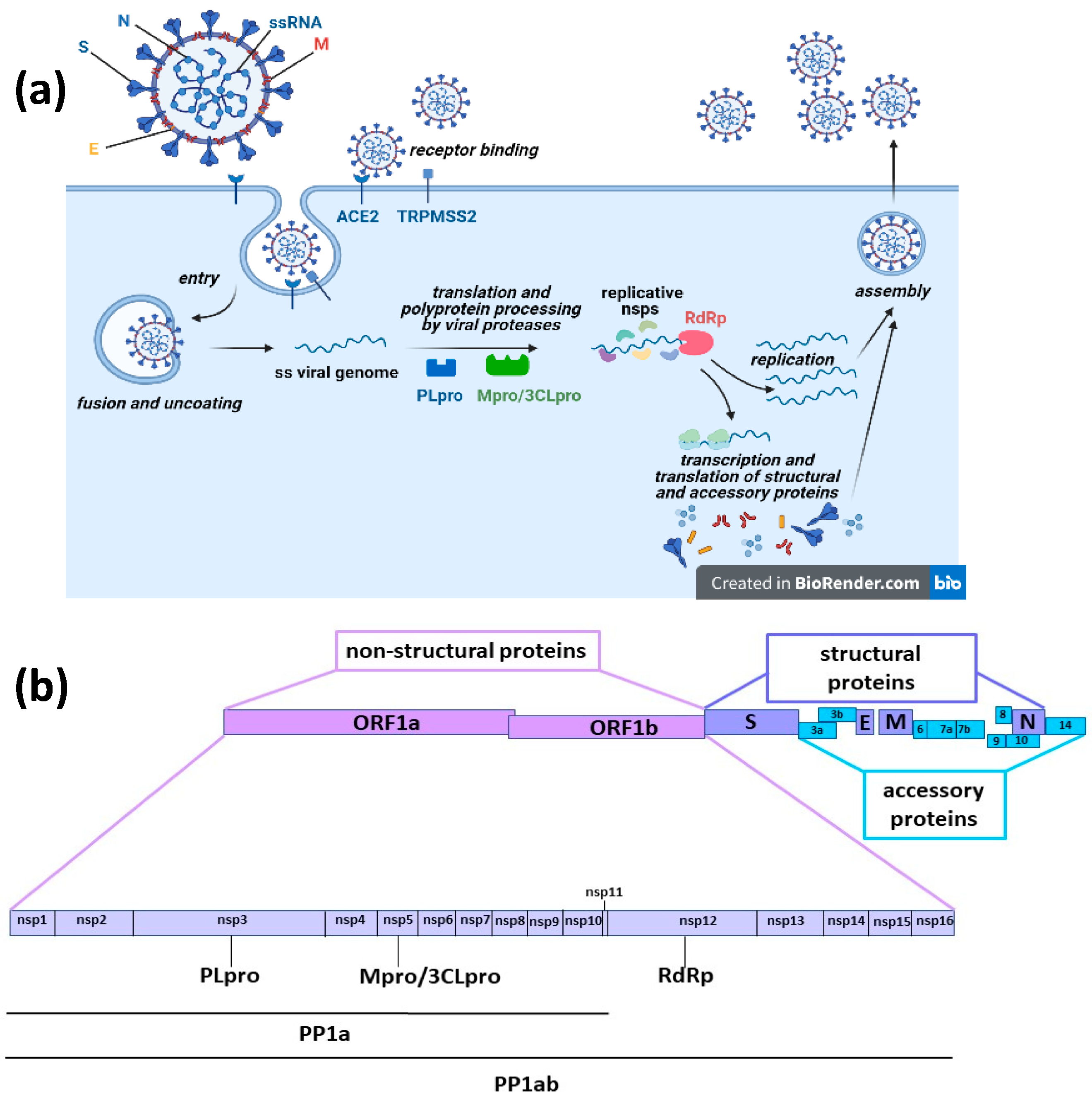
- Jeong, G.U.; Song, H.; Yoon, G.Y.; Kim, D.; Kwon, Y.-C. Therapeutic Strategies against COVID-19 and Structural Characterization of SARS-CoV-2: A Review. Front. Microbiol. 2020, 11, 1723. [Google Scholar] [CrossRef] [PubMed]
- Romano, M.; Ruggiero, A.; Squeglia, F.; Maga, G.; Berisio, R. A Structural View of SARS-CoV-2 RNA Replication Machinery: RNA Synthesis, Proofreading and Final Capping. Cells 2020, 9, 1267. [Google Scholar] [CrossRef]
- Arya, R.; Kumari, S.; Pandey, B.; Mistry, H.; Bihani, S.C.; Das, A.; Prashar, V.; Gupta, G.D.; Panicker, L.; Kumar, M. Structural insights into SARS-CoV-2 proteins. J. Mol. Biol. 2021, 433, 166725. [Google Scholar] [CrossRef] [PubMed]
- Available online: https://www.fda.gov/news-events/press-announcements/fda-approves-first-treatment-covid-19 (accessed on 12 April 2021).
- Magro, P.; Zanella, I.; Pescarolo, M.; Castelli, F.; Quiros-Roldan, E. Lopinavir/ritonavir: Repurposing an old drug for HIV infection in COVID-19 treatment. Biomed. J. 2021, 44, 43–53. [Google Scholar] [CrossRef]
- Chan, J.F.-W.; Yao, Y.; Yeung, M.-L.; Deng, W.; Bao, L.; Jia, L.; Li, F.; Xiao, C.; Gao, H.; Yu, P.; et al. Treatment with Lopinavir/Ritonavir or Interferon-β1b Improves Outcome of MERS-CoV Infection in a Nonhuman Primate Model of Common Marmoset. J. Infect. Dis. 2015, 212, 1904–1913. [Google Scholar] [CrossRef] [PubMed]
- Cao, B.; Wang, Y.; Wen, D.; Liu, W.; Wang, J.; Fan, G.; Ruan, L.; Song, B.; Cai, Y.; Wei, M.; et al. A Trial of Lopinavir–Ritonavir in Adults Hospitalized with Severe Covid-19. N. Engl. J. Med. 2020, 382, 1787–1799. [Google Scholar] [CrossRef]
- WHO Solidarity Trial Consortium. Repurposed Antiviral Drugs for Covid-19-Interim WHO Solidarity Trial Results. N. Engl. J. Med. 2021, 384, 497–511. [Google Scholar] [CrossRef]
- RECOVERY Collaborative Group. Lopinavir-ritonavir in patients admitted to hospital with COVID-19 (RECOVERY): A randomised, controlled, open-label, platform trial. Lancet 2020, 396, 1345–1352. [Google Scholar] [CrossRef]
- Available online: https://www.who.int/news-room/fact-sheets/detail/hiv-aids (accessed on 12 April 2021).
- Grant, J.K.; Vincent, L.; Ebner, B.; Hurwitz, B.E.; Alcaide, M.L.; Martinez, C. Early Insights into COVID-19 in Persons Living with HIV and Cardiovascular Manifestations. J. AIDS HIV Treat. 2020, 2, 68–74. [Google Scholar] [CrossRef] [PubMed]
- Zhu, F.; Cao, Y.; Xu, S.; Zhou, M. Co-infection of SARS-CoV-2 and HIV in a patient in Wuhan city, China. J. Med. Virol. 2020, 92, 529–530. [Google Scholar] [CrossRef]
- Riva, A.; Conti, F.; Bernacchia, D.; Pezzati, L.; Sollima, S.; Merli, S.; Siano, M.; Lupo, A.; Rusconi, S.; Cattaneo, D.; et al. Darunavir does not prevent SARS-CoV-2 infection in HIV patients. Pharmacol. Res. 2020, 157, 104826. [Google Scholar] [CrossRef] [PubMed]
- Blanco, J.L.; Ambrosioni, J.; Garcia, F.; Martínez, E.; Soriano, A.; Mallolas, J.; Miro, J.M. COVID-19 in patients with HIV: Clinical case series. Lancet HIV 2020, 7, e314–e316. [Google Scholar] [CrossRef]
- Singh, A. In severe COVID-19, adding lopinavir–ritonavir to usual care did not improve mortality at 28 days. Ann. Intern. Med. 2021, 174, JC3. [Google Scholar] [CrossRef]
- Mellor, M.M.; Bast, A.C.; Jones, N.R.; Roberts, N.W.; Ordóñez-Mena, J.M.; Reith, A.J.; Butler, C.C.; Matthews, P.C.; Dorward, J. Risk of adverse COVID-19 outcomes for people living with HIV. AIDS 2021, F1–F10. [Google Scholar] [CrossRef]
- Patel, R.H.; Acharya, A.; Chand, H.S.; Mohan, M.; Byrareddy, S.N. HIV and SARS-CoV-2 Co-infection: A Systematic Review of the Literature and Challenges. AIDS Res. Hum. Retrovir. 2021. [Google Scholar] [CrossRef]
- Gesesew, H.A.; Koye, D.N.; Fetene, D.M.; Woldegiorgis, M.; Kinfu, Y.; Geleto, A.B.; Melaku, Y.A.; Mohammed, H.; Alene, K.A.; Awoke, M.A.; et al. Risk factors for COVID-19 infection, disease severity and related deaths in Africa: A systematic review. BMJ Open 2021, 11, e044618. [Google Scholar] [CrossRef]
- Brown, L.B.; Spinelli, M.A.; Gandhi, M. The interplay between HIV and COVID-19: Summary of the data and responses to date. Curr. Opin. HIV AIDS 2021, 16, 63–73. [Google Scholar] [CrossRef]
- Quiros-Roldan, E.; Biasiotto, G.; Zanella, I. Letter to the Editor on “Bonafè M, Prattichizzo F, Giuliani A, Storci G, Sabbatinelli J, Olivieri, F. Inflamm-aging: Why older men are the most susceptible to SARS-CoV-2 complicated outcomes. Cytokine Growth Factor Rev”. Cytokine Growth Factor Rev. 2020, 54, 1–2. [Google Scholar] [CrossRef]
- Monforte, A.D.; Bonnet, F.; Bucher, H.; Pourcher, V.; Pantazis, N.; Pelchen-Matthews, A.; Touloumi, G.; Wolf, E. What do the changing patterns of comorbidity burden in people living with HIV mean for long-term management? Perspectives from European HIV cohorts. HIV Med. 2020, 21, 3–16. [Google Scholar] [CrossRef]
- Guaraldi, G.; Milic, J.; Mussini, C. Aging with HIV. Curr. HIV/AIDS Rep. 2019, 16, 475–481. [Google Scholar] [CrossRef]
- Jones, R.; Nelson, M.; Bracchi, M.; Asboe, D.; Boffito, M. COVID-19 in patients with HIV. Lancet HIV 2020, 7, e383. [Google Scholar] [CrossRef]
- Byrd, K.M.; Beckwith, C.G.; Garland, J.M.; Johnson, J.E.; Aung, S.; Cu-Uvin, S.; Farmakiotis, D.; Flanigan, T.; Gillani, F.S.; Macias-Gil, R.; et al. SARS-CoV-2 and HIV coinfection: Clinical experience from Rhode Island, United States. J. Int. AIDS Soc. 2020, 23. [Google Scholar] [CrossRef]
- Xu, Z.; Zhang, C.; Wang, F.-S. COVID-19 in people with HIV. Lancet HIV 2020, 7, e524–e526. [Google Scholar] [CrossRef]
- Wu, Q.; Chen, T.; Zhang, H. Recovery from the coronavirus disease-2019 (COVID-19) in two patients with coexisted (HIV) infection. J. Med. Virol. 2020, 92, 2325–2327. [Google Scholar] [CrossRef]
- Vizcarra, P.; Pérez-Elías, M.J.; Quereda, C.; Moreno, A.; Vivancos, M.J.; Dronda, F.; Casado, J.L.; Moreno, S.; Fortún, J.; Navas, E.; et al. Description of COVID-19 in HIV-infected individuals: A single-centre, prospective cohort. Lancet HIV 2020, 7, e554–e564. [Google Scholar] [CrossRef]
- Härter, G.; Spinner, C.D.; Roider, J.; Bickel, M.; Krznaric, I.; Grunwald, S.; Schabaz, F.; Gillor, D.; Postel, N.; Mueller, M.C.; et al. COVID-19 in people living with human immunodeficiency virus: A case series of 33 patients. Infection 2020, 48, 681–686. [Google Scholar] [CrossRef]
- Okoh, A.K.; Bishburg, E.; Grinberg, S.; Nagarakanti, S. COVID-19 Pneumonia in Patients with HIV: A Case Series. JAIDS J. Acquir. Immune Defic. Syndr. 2020, 85, e4–e5. [Google Scholar] [CrossRef]
- Gervasoni, C.; Meraviglia, P.; Riva, A.; Giacomelli, A.; Oreni, L.; Minisci, D.; Atzori, C.; Ridolfo, A.; Cattaneo, D. Clinical Features and Outcomes of Patients with Human Immunodeficiency Virus With COVID-19. Clin. Infect. Dis. 2020, 71, 2276–2278. [Google Scholar] [CrossRef]
- Lesko, C.R.; Bengtson, A.M. HIV and COVID-19: Intersecting Epidemics with Many Unknowns. Am. J. Epidemiol. 2021, 190, 10–16. [Google Scholar] [CrossRef]
- Karmen-Tuohy, S.; Carlucci, P.M.; Zervou, F.N.; Zacharioudakis, I.M.; Rebick, G.; Klein, E.; Reich, J.; Jones, S.; Rahimian, J. Outcomes among HIV-Positive Patients Hospitalized with COVID-19. JAIDS J. Acquir. Immune Defic. Syndr. 2020, 85, 6–10. [Google Scholar] [CrossRef]
- Mirzaei, H.; McFarland, W.; Karamouzian, M.; Sharifi, H. COVID-19 among People Living with HIV: A Systematic Review. AIDS Behav. 2021, 25, 85–92. [Google Scholar] [CrossRef] [PubMed]
- Yang, R.; Gui, X.; Zhang, Y.; Xiong, Y.; Gao, S.; Ke, H. Clinical characteristics of COVID-19 patients with HIV coinfection in Wuhan, China. Expert Rev. Respir. Med. 2021, 15, 403–409. [Google Scholar] [CrossRef] [PubMed]
- Lee, K.; Yap, S.; Ngeow, Y.; Lye, M. COVID-19 in People Living with HIV: A Systematic Review and Meta-Analysis. Int. J. Environ. Res. Public Health 2021, 18, 3554. [Google Scholar] [CrossRef] [PubMed]
- Mohindra, R.; Kanta, P.; Porchezhian, P.; Goyal, K.; Suri, V.; Singh, M.P.; Pvm, L.; Bhalla, A. COVID-19 infection in a HIV positive health care worker: First case report from a tertiary care hospital of North India. VirusDisease 2021, 2021, 1–5. [Google Scholar] [CrossRef]
- Calza, L.; Bon, I.; Tadolini, M.; Borderi, M.; Colangeli, V.; Badia, L.; Verucchi, G.; Rossini, G.; Vocale, C.; Gaibani, P.; et al. COVID-19 in patients with HIV-1 infection: A single-centre experience in northern Italy. Infection 2021, 49, 333–337. [Google Scholar] [CrossRef] [PubMed]
- Hoffmann, C.; Casado, J.L.; Härter, G.; Vizcarra, P.; Moreno, A.; Cattaneo, D.; Meraviglia, P.; Spinner, C.D.; Schabaz, F.; Grunwald, S.; et al. Immune deficiency is a risk factor for severe COVID-19 in people living with HIV. HIV Med. 2021, 22, 372–378. [Google Scholar] [CrossRef] [PubMed]
- Johnston, R. The first 6 months of HIV-SARS-CoV-2 coinfection: Outcomes for 6947 individuals. Curr. Opin. HIV AIDS 2021, 16, 54–62. [Google Scholar] [CrossRef]
- Joob, B.; Wiwanitkit, V. SARS-CoV-2 and HIV. J. Med. Virol. 2020, 92, 1415. [Google Scholar] [CrossRef]
- Hu, Y.; Ma, J.; Huang, H.; Vermund, S.H. Coinfection with HIV and SARS-CoV-2 in Wuhan, China: A 12-Person Case Series. JAIDS J. Acquir. Immune Defic. Syndr. 2020, 85, 1–5. [Google Scholar] [CrossRef] [PubMed]
- Ridgway, J.P.; Farley, B.; Benoit, J.-L.; Frohne, C.; Hazra, A.; Pettit, N.; Pho, M.; Pursell, K.; Saltzman, J.; Schmitt, J.; et al. A Case Series of Five People Living with HIV Hospitalized with COVID-19 in Chicago, Illinois. AIDS Patient Care STDs 2020, 34, 331–335. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Cheng, X.; Wang, R.; Zeng, X. Computed tomography imaging of an HIV-infected patient with coronavirus disease 2019. J. Med. Virol. 2020, 92, 1774–1776. [Google Scholar] [CrossRef] [PubMed]
- Laurence, J. Why Aren’t People Living with HIV at Higher Risk for Developing Severe Coronavirus Disease 2019 (COVID-19)? AIDS Patient Care STDs 2020, 34, 247–248. [Google Scholar] [CrossRef] [PubMed]
- Balzarini, J.; Holy, A.; Jindrich, J.; Naesens, L.; Snoeck, R.; Schols, D.; De Clercq, E. Differential antiherpesvirus and antiretrovirus effects of the (S) and (R) enantiomers of acyclic nucleoside phosphonates: Potent and selective in vitro and in vivo antiretrovirus activities of (R)-9-(2-phosphonomethoxypropyl)-2,6-diaminopurine. Antimicrob. Agents Chemother. 1993, 37, 332–338. [Google Scholar] [CrossRef]
- Edited Tsai, C.-C.; Follis, K.E.; Sabo, A.; Beck, T.W.; Grant, R.F.; Bischofberger, N.; Benveniste, R.E.; Black, R. Prevention of SIV Infection in Macaques by (R)-9-(2-Phosphonylmethoxypropyl)adenine. Science 1995, 270, 1197–1199. [Google Scholar] [CrossRef] [PubMed]
- Elfiky, A.A. Ribavirin, Remdesivir, Sofosbuvir, Galidesivir, and Tenofovir against SARS-CoV-2 RNA dependent RNA polymerase (RdRp): A molecular docking study. Life Sci. 2020, 253, 117592. [Google Scholar] [CrossRef]
- Edited Li, G.; Yue, T.; Zhang, P.; Gu, W.; Gao, L.-J.; Tan, L. Drug Discovery of Nucleos(t)ide Antiviral Agents: Dedicated to Prof. Dr. Erik De Clercq on Occasion of His 80th Birthday. Molecules 2021, 26, 923. [Google Scholar] [CrossRef]
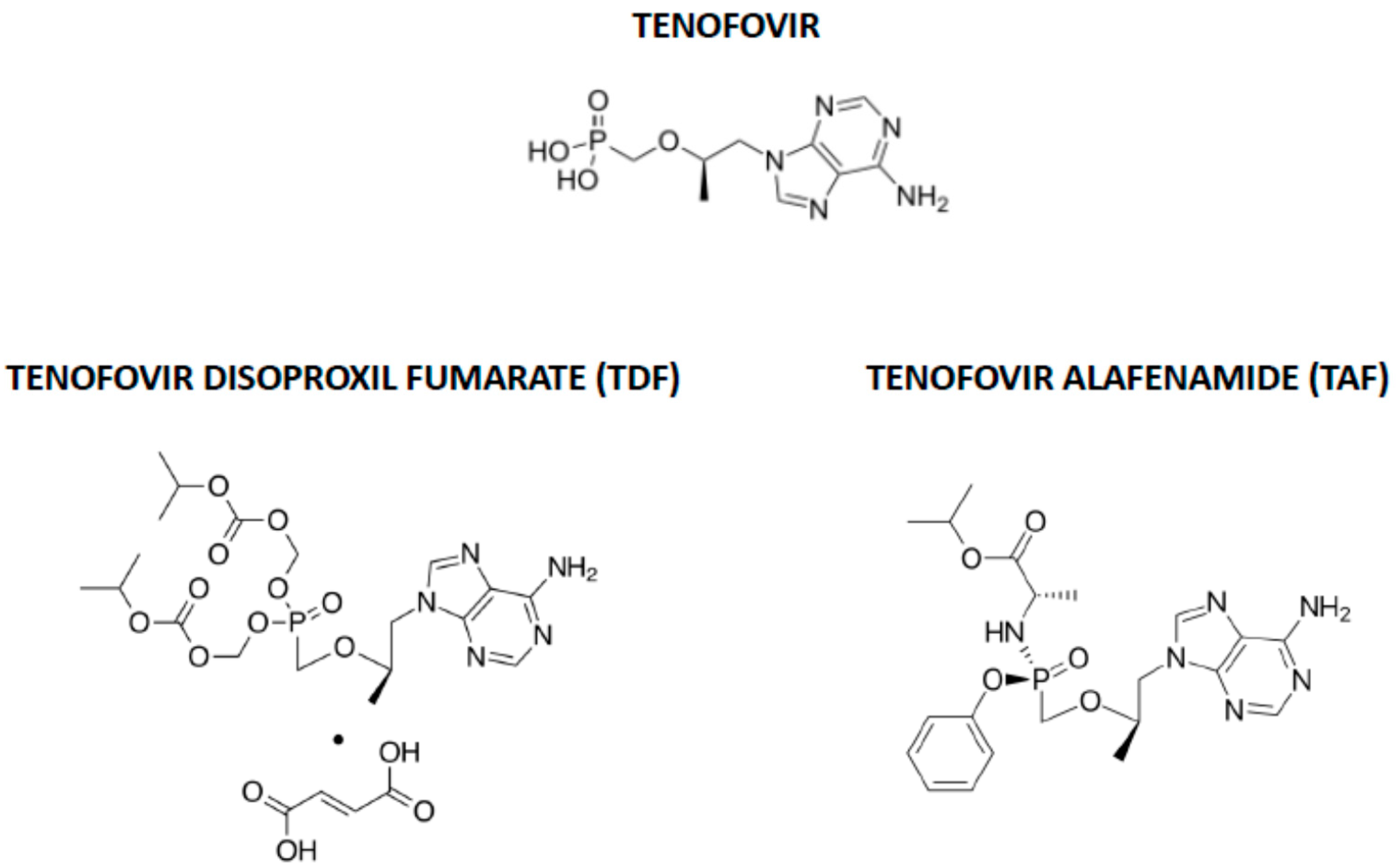
- Wassner, C.; Bradley, N.; Lee, Y. A Review and Clinical Understanding of Tenofovir: Tenofovir Disoproxil Fumarate versus Tenofovir Alafenamide. J. Int. Assoc. Provid. AIDS Care (JIAPAC) 2020, 19. [Google Scholar] [CrossRef]
- Mu, Y.; Pham, M.; Podany, A.T.; Cory, T.J. Evaluating emtricitabine + rilpivirine + tenofovir alafenamide in combination for the treatment of HIV-infection. Expert Opin. Pharmacother. 2020, 21, 389–397. [Google Scholar] [CrossRef]
- Waters, L.; Mehta, V.; Gogtay, J.; Boffito, M. The evidence for using tenofovir disoproxil fumarate plus lamivudine as a nucleoside analogue backbone for the treatment of HIV. J. Virus Erad. 2021, 7, 100028. [Google Scholar] [CrossRef]
- Santevecchi, B.A.; Miller, S.; Childs-Kean, L.M. Doing More with Less: Review of Dolutegravir-Lamivudine, a Novel Single-Tablet Regimen for Antiretroviral-Naïve Adults with HIV-1 Infection. Ann. Pharmacother. 2020, 54, 1252–1259. [Google Scholar] [CrossRef]
- Buchbinder, S.P.; Liu, A.Y. CROI 2019: Advances in HIV prevention and plans to end the epidemic. Top. Antivir. Med. 2019, 27, 8–25. [Google Scholar] [PubMed]
- Biello, K.B.; Mimiaga, M.J.; Valente, P.K.; Saxena, N.; Bazzi, A.R. The Past, Present, and Future of PrEP implementation among People Who Use Drugs. Curr. HIV/AIDS Rep. 2021. [Google Scholar] [CrossRef]
- Özdener-Poyraz, A.E.; Slugocki, M.; Kalabalik-Hoganson, J.; Han, J. Pre-Exposure Prophylaxis (PrEP) in the Prevention of HIV: Strategies, Target Populations and Upcoming Treatments. HIV/AIDS Res. Palliat. Care 2020, 12, 283–293. [Google Scholar] [CrossRef] [PubMed]
- Mayer, K.H.; Allan-Blitz, L.-T. PrEP 1.0 and Beyond: Optimizing a Biobehavioral Intervention. JAIDS J. Acquir. Immune Defic. Syndr. 2019, 82, S113–S117. [Google Scholar] [CrossRef]
- Riddell, J.; Amico, K.R.; Mayer, K.H. HIV Preexposure Prophylaxis. JAMA 2018, 319, 1261–1268. [Google Scholar] [CrossRef]
- Fonner, V.A.; Dalglish, S.L.; Kennedy, C.E.; Baggaley, R.; O’Reilly, K.R.; Koechlin, F.M.; Rodolph, M.; Hodges-Mameletzis, I.; Grant, R.M. Effectiveness and safety of oral HIV preexposure prophylaxis for all populations. AIDS 2016, 30, 1973–1983. [Google Scholar] [CrossRef]
- Sun, S.; Yang, Q.; Sheng, Y.; Fu, Y.; Sun, C.; Deng, C. Investigational drugs with dual activity against HBV and HIV (Review). Exp. Ther. Med. 2020, 21. [Google Scholar] [CrossRef]
- Roade, L.; Riveiro-Barciela, M.; Esteban, R.; Buti, M. Long-term efficacy and safety of nucleos(t)ides analogues in patients with chronic hepatitis B. Ther. Adv. Infect. Dis. 2021, 8. [Google Scholar] [CrossRef]
- Hall, S.; Howell, J.; Visvanathan, K.; Thompson, A. The Yin and the Yang of Treatment for Chronic Hepatitis B—When to Start, When to Stop Nucleos(t)ide Analogue Therapy. Viruses 2020, 12, 934. [Google Scholar] [CrossRef]
- Marrazzo, J.M.; Rabe, L.; Kelly, C.; Richardson, B.; Deal, C.; Schwartz, J.L.; Chirenje, Z.M.; Piper, J.; Morrow, R.A.; Hendrix, C.W.; et al. Tenofovir Gel for Prevention of Herpes Simplex Virus Type 2 Acquisition: Findings from the VOICE Trial. J. Infect. Dis. 2019, 219, 1940–1947. [Google Scholar] [CrossRef]
- Chaix, M.-L.; Charreau, I.; Pintado, C.; Delaugerre, C.; Mahjoub, N.; Cotte, L.; Capitant, C.; Raffi, F.; Cua, E.; Pialoux, G.; et al. Effect of On-Demand Oral Pre-exposure Prophylaxis with Tenofovir/Emtricitabine on Herpes Simplex Virus-1/2 Incidence among Men Who Have Sex with Men: A Substudy of the ANRS IPERGAY Trial. Open Forum. Infect. Dis. 2018, 5. [Google Scholar] [CrossRef] [PubMed]
- Andrei, G.; Gillemot, S.; Topalis, D.; Snoeck, R. The Anti–Human Immunodeficiency Virus Drug Tenofovir, a Reverse Transcriptase Inhibitor, Also Targets the Herpes Simplex Virus DNA Polymerase. J. Infect. Dis. 2017, 217, 790–801. [Google Scholar] [CrossRef] [PubMed]
- Celum, C.; Hong, T.; Cent, A.; Donnell, D.; Morrow, R.; Baeten, J.M.; Firnhaber, C.; Grinsztejn, B.; Hosseinipour, M.C.; Lalloo, U.; et al. Herpes Simplex Virus Type 2 Acquisition among HIV-1–Infected Adults Treated with Tenofovir Disoproxyl Fumarate as Part of Combination Antiretroviral Therapy: Results from the ACTG A5175 PEARLS Study. J. Infect. Dis. 2017, 215, 907–910. [Google Scholar] [CrossRef]
- Gibas, K.M.; Berg, P.V.D.; Powell, V.E.; Krakower, D.S. Drug Resistance during HIV Pre-Exposure Prophylaxis. Drugs 2019, 79, 609–619. [Google Scholar] [CrossRef] [PubMed]
- Tang, L.S.Y.; Covert, E.; Wilson, E.; Kottilil, S. Chronic Hepatitis B Infection. JAMA 2018, 319, 1802–1813. [Google Scholar] [CrossRef]
- Park, E.-S.; Lee, A.R.; Kim, D.H.; Lee, J.-H.; Yoo, J.-J.; Ahn, S.H.; Sim, H.; Park, S.; Kang, H.S.; Won, J.; et al. Identification of a quadruple mutation that confers tenofovir resistance in chronic hepatitis B patients. J. Hepatol. 2019, 70, 1093–1102. [Google Scholar] [CrossRef]
- DeJong, C.; Spinelli, M.A.; Okochi, H.; Gandhi, M. Tenofovir-based PrEP for COVID-19. AIDS 2021. [Google Scholar] [CrossRef]
- Edited Udofia, I.A.; Gbayo, K.O.; Oloba-Whenu, O.A.; Ogunbayo, T.B.; Isanbor, C. In silico studies of selected multi-drug targeting against 3CLpro and nsp12 RNA-dependent RNA-polymerase proteins of SARS-CoV-2 and SARS-CoV. Netw. Model. Anal. Health Inform. Bioinform. 2021, 10, 1–12. [Google Scholar] [CrossRef]
- Poustforoosh, A.; Hashemipour, H.; Tüzün, B.; Pardakhty, A.; Mehrabani, M.; Nematollahi, M.H. Evaluation of potential anti-RNA-dependent RNA polymerase (RdRP) drugs against the newly emerged model of COVID-19 RdRP using computational methods. Biophys. Chem. 2021, 272, 106564. [Google Scholar] [CrossRef]
- Tiwari, V. Denovo designing, retro-combinatorial synthesis, and molecular dynamics analysis identify novel antiviral VTRM1.1 against RNA-dependent RNA polymerase of SARS CoV2 virus. Int. J. Biol. Macromol. 2021, 171, 358–365. [Google Scholar] [CrossRef]
- ElFiky, A.A. SARS-CoV-2 RNA dependent RNA polymerase (RdRp) targeting: An in silico perspective. J. Biomol. Struct. Dyn. 2020, 2020, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Edited Grahl, M.V.; Alcará, A.M.; de Souza, O.N.; Ligabue-Braun, R.; Perin, A.P.A.; Moro, C.F.; Pinto, É.S.; Feltes, B.C.; Ghilardi, I.M.; Rodrigues, F.V.; et al. Evaluation of drug repositioning by molecular docking of pharmaceutical resources available in the Brazilian healthcare system against SARS-CoV-2. Inform. Med. Unlocked 2021, 23, 100539. [Google Scholar] [CrossRef]
- Copertino, D.C., Jr.; Lima, B.C.C.; Duarte, R.R.R.; Powell, T.R.; Ormsby, C.E.; Wilkin, T.; Gulick, R.M.; Rougvie, M.D.M.; Nixon, D.F. Antiretroviral drug activity and potential for pre-exposure prophylaxis against COVID-19 and HIV infection. J. Biomol. Struct. Dyn. 2021, 2021, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Hasan, K.; Kamruzzaman, M.; Bin Manjur, O.H.; Mahmud, A.; Hussain, N.; Alam Mondal, M.S.; Hosen, I.; Bello, M.; Rahman, A. Structural analogues of existing anti-viral drugs inhibit SARS-CoV-2 RNA dependent RNA polymerase: A computational hierarchical investigation. Heliyon 2021, 7, e06435. [Google Scholar] [CrossRef]
- Dallocchio, R.N.; Dessì, A.; De Vito, A.; Delogu, G.; Serra, P.A.; Madeddu, G. Early combination treatment with existing HIV antivirals: An effective treatment for COVID-19? Eur. Rev. Med. Pharmacol. Sci. 2021, 25, 2435–2448. [Google Scholar] [PubMed]
- Salpini, R.; Alkhatib, M.; Costa, G.; Piermatteo, L.; Ambrosio, F.A.; Di Maio, V.C.; Scutari, R.; Duca, L.; Berno, G.; Fabeni, L.; et al. Key genetic elements, single and in clusters, underlying geographically dependent SARS-CoV-2 genetic adaptation and their impact on binding affinity for drugs and immune control. J. Antimicrob. Chemother. 2021, 76, 396–412. [Google Scholar] [CrossRef]
- Yun, Y.; Song, H.; Ji, Y.; Huo, D.; Han, F.; Li, F.; Jiang, N. Identification of therapeutic drugs against COVID-19 through computational investigation on drug repurposing and structural modification. J. Biomed. Res. 2020, 34, 458–469. [Google Scholar] [CrossRef] [PubMed]
- Toor, H.G.; Banerjee, D.I.; Rath, S.L.; Darji, S.A. Computational drug re-purposing targeting the spike glycoprotein of SARS-CoV-2 as an effective strategy to neutralize COVID-19. Eur. J. Pharmacol. 2021, 890, 173720. [Google Scholar] [CrossRef]
- Shah, B.; Modi, P.; Sagar, S.R. In silico studies on therapeutic agents for COVID-19: Drug repurposing approach. Life Sci. 2020, 252, 117652. [Google Scholar] [CrossRef]
- Edited Rahman, M.R.; Banik, A.; Chowdhury, I.M.; Sajib, E.H.; Sarkar, S. Identification of potential antivirals against SARS-CoV-2 using virtual screening method. Inform. Med. Unlocked 2021, 23, 100531. [Google Scholar] [CrossRef] [PubMed]
- Feng, Z.; Chen, M.; Xue, Y.; Liang, T.; Chen, H.; Zhou, Y.; Nolin, T.D.; Smith, R.B.; Xie, X.-Q. MCCS: A novel recognition pattern-based method for fast track discovery of anti-SARS-CoV-2 drugs. Brief. Bioinform. 2021, 22, 946–962. [Google Scholar] [CrossRef] [PubMed]
- Choy, K.-T.; Wong, A.Y.-L.; Kaewpreedee, P.; Sia, S.F.; Chen, D.; Hui, K.P.Y.; Chu, D.K.W.; Chan, M.C.W.; Cheung, P.P.-H.; Huang, X.; et al. Remdesivir, lopinavir, emetine, and homoharringtonine inhibit SARS-CoV-2 replication in vitro. Antivir. Res. 2020, 178, 104786. [Google Scholar] [CrossRef]
- Zandi, K.; Amblard, F.; Musall, K.; Downs-Bowen, J.; Kleinbard, R.; Oo, A.; Cao, D.; Liang, B.; Russell, O.O.; McBrayer, T.; et al. Repurposing Nucleoside Analogs for Human Coronaviruses. Antimicrob. Agents Chemother. 2020, 65, e01652-20. [Google Scholar] [CrossRef] [PubMed]
- Clososki, G.C.; Soldi, R.A.; da Silva, R.M.; Guaratini, T.; Lopes, J.N.C.; Pereira, P.R.R.; Lopes, J.L.C.; dos Santos, T.; Martins, R.B.; Costa, C.S.; et al. Tenofovir Disoproxil Fumarate: New Chemical Developments and Encouraging in vitro Biological Results for SARS-CoV-2. J. Braz. Chem. Soc. 2020, 31, 1552–1556. [Google Scholar] [CrossRef]
- Xie, X.; Muruato, A.E.; Zhang, X.; Lokugamage, K.G.; Fontes-Garfias, C.R.; Zou, J.; Liu, J.; Ren, P.; Balakrishnan, M.; Cihlar, T.; et al. A nanoluciferase SARS-CoV-2 for rapid neutralization testing and screening of anti-infective drugs for COVID-19. Nat. Commun. 2020, 11, 5214. [Google Scholar] [CrossRef]
- Veras, F.P.; Pontelli, M.C.; Silva, C.M.; Toller-Kawahisa, J.E.; De Lima, M.; Nascimento, D.C.; Schneider, A.H.; Caetité, D.; Tavares, L.A.; Paiva, I.M.; et al. SARS-CoV-2–triggered neutrophil extracellular traps mediate COVID-19 pathology. J. Exp. Med. 2020, 217. [Google Scholar] [CrossRef]
- Chien, M.; Anderson, T.K.; Jockusch, S.; Tao, C.; Li, X.; Kumar, S.; Russo, J.J.; Kirchdoerfer, R.N.; Ju, J. Nucleotide Analogues as Inhibitors of SARS-CoV-2 Polymerase, a Key Drug Target for COVID-19. J. Proteome Res. 2020, 19, 4690–4697. [Google Scholar] [CrossRef]
- Park, S.-J.; Yu, K.-M.; Kim, Y.-I.; Kim, S.-M.; Kim, E.-H.; Kim, S.-G.; Casel, M.A.B.; Rollon, R.; Jang, S.-G.; Lee, M.-H.; et al. Antiviral Efficacies of FDA-Approved Drugs against SARS-CoV-2 Infection in Ferrets. mBio 2020, 11. [Google Scholar] [CrossRef]
- Boulle, A.; Davies, M.-A.; Hussey, H.; Ismail, M.; Morden, E.; Vundle, Z.; Zweigenthal, V.; Mahomed, H.; Paleker, M.; Pienaar, D.; et al. Risk Factors for Coronavirus Disease 2019 (COVID-19) Death in a Population Cohort Study from the Western Cape Province, South Africa. Clin. Infect. Dis. 2020. [Google Scholar] [CrossRef]
- Del Amo, J.; Polo, R.; Moreno, S.; Díaz, A.; Martínez, E.; Arribas, J.R.; Jarrín, I.; Hernán, M.A. Incidence and Severity of COVID-19 in HIV-Positive Persons Receiving Antiretroviral Therapy. Ann. Intern. Med. 2020, 173, 536–541. [Google Scholar] [CrossRef] [PubMed]
- del Amo, J.; Polo, R.; Moreno, S.; Díaz, A.; Martínez, E.; Arribas, J.R.; Jarrín, I.; Hernán, M.A. Antiretrovirals and Risk of COVID-19 Diagnosis and Hospitalization in HIV-Positive Persons. Epidemiology 2020, 31, e49–e51. [Google Scholar] [CrossRef] [PubMed]
- Melchjorsen, J.; Risør, M.W.; Søgaard, O.S.; O’Loughlin, K.L.; Chow, S.; Paludan, S.R.; Ellermann-Eriksen, S.; Hedley, D.W.; Minderman, H.; Østergaard, L.; et al. Tenofovir Selectively Regulates Production of Inflammatory Cytokines and Shifts the IL-12/IL-10 Balance in Human Primary Cells. JAIDS J. Acquir. Immune Defic. Syndr. 2011, 57, 265–275. [Google Scholar] [CrossRef]
- Castillo-Mancilla, J.R.; Meditz, A.; Wilson, C.; Zheng, J.-H.; Palmer, B.E.; Lee, E.J.; Gardner, E.M.; Seifert, S.; Kerr, B.; Bushman, L.R.; et al. Reduced Immune Activation during Tenofovir–Emtricitabine Therapy in HIV-Negative Individuals. JAIDS J. Acquir. Immune Defic. Syndr. 2015, 68, 495–501. [Google Scholar] [CrossRef]
- Ayerdi, O.; Puerta, T.; Clavo, P.; Vera, M.; Ballesteros, J.; Fuentes, M.E.; Estrada, V.; Rodríguez, C.; Del Romero, J.; Lejarraga, C.; et al. Preventive Efficacy of Tenofovir/Emtricitabine against Severe Acute Respiratory Syndrome Coronavirus 2 among Pre-Exposure Prophylaxis Users. Open Forum Infect. Dis. 2020, 7, ofaa455. [Google Scholar] [CrossRef]
- Available online: https://clinicaltrials.gov/ct2/show/NCT04519125 (accessed on 12 April 2021).
- Available online: https://clinicaltrials.gov/ct2/show/NCT04405271 (accessed on 12 April 2021).
- Available online: https://clinicaltrials.gov/ct2/show/NCT04575545 (accessed on 12 April 2021).
- Available online: https://clinicaltrials.gov/ct2/show/NCT04712357 (accessed on 12 April 2021).
- Available online: https://clinicaltrials.gov/ct2/show/NCT04685512 (accessed on 12 April 2021).
- Available online: https://clinicaltrials.gov/ct2/show/NCT04359095 (accessed on 12 April 2021).
- Available online: https://clinicaltrials.gov/ct2/show/NCT04334928 (accessed on 12 April 2021).
- Available online: https://clinicaltrials.gov/ct2/show/NCT04812496 (accessed on 12 April 2021).
- Roldan, E.Q.; Biasiotto, G.; Magro, P.; Zanella, I. The possible mechanisms of action of 4-aminoquinolines (chloroquine/hydroxychloroquine) against Sars-Cov-2 infection (COVID-19): A role for iron homeostasis? Pharmacol. Res. 2020, 158, 104904. [Google Scholar] [CrossRef]
- Eze, P.; Mezue, K.N.; Nduka, C.U.; Obianyo, I.; Egbuche, O. Efficacy and safety of chloroquine and hydroxychloroquine for treatment of COVID-19 patients-a systematic review and meta-analysis of randomized controlled trials. Am. J. Cardiovasc. Dis. 2021, 11, 93–107. [Google Scholar]
- Kearney, B.P.; Flaherty, J.F.; Shah, J. Tenofovir Disoproxil Fumarate. Clin. Pharmacokinet. 2004, 43, 595–612. [Google Scholar] [CrossRef]
- Geraghty, R.; Aliota, M.; Bonnac, L. Broad-Spectrum Antiviral Strategies and Nucleoside Analogues. Viruses 2021, 13, 667. [Google Scholar] [CrossRef] [PubMed]
- Dash, P.; Mohapatra, S.; Ghosh, S.; Nayak, B. A Scoping Insight on Potential Prophylactics, Vaccines and Therapeutic Weaponry for the Ongoing Novel Coronavirus (COVID-19) Pandemic- A Comprehensive Review. Front. Pharmacol. 2021, 11, 590154. [Google Scholar] [CrossRef] [PubMed]
- Alavian, G.; Kolahdouzan, K.; Mortezazadeh, M.; Torabi, Z.S. Antiretrovirals for Prophylaxis against COVID-19: A Comprehensive Literature Review. J. Clin. Pharmacol. 2020, 61, 581–590. [Google Scholar] [CrossRef] [PubMed]
| NCT Number [Ref] | Title | Status |
|---|---|---|
| NCT04519125 [100] | Daily Regimen of Tenofovir/Emtricitabine as Prevention for COVID-19 in Health Care Personnel in Colombia | not yet recruiting |
| NCT04405271 [101] | TAF/FTC for Pre-exposure Prophylaxis of COVID-19 in Healthcare Workers (CoviPrep Study) | not yet recruiting |
| NCT04575545 [102] | Prevalence of COVID-19 Infection in a Cohort of Patients Infected by the HIV and Patients Taking PrEP. | not yet recruiting |
| NCT04712357 [103] | Clinical Experimentation with Tenofovir Disoproxyl Fumarate and Emtricitabine for COVID-19 (ARTAN-C19) | recruiting |
| NCT04685512 [104] | Effect of Tenofovir/Emtricitabine in Patients Recently Infected With SARS-COV2 (Covid-19) Discharged Home (AR0-CORONA) | recruiting |
| NCT04359095 [105] | Effectiveness and Safety of Medical Treatment for SARS-CoV-2 (COVID-19) in Colombia | recruiting |
| NCT04334928 [106] | Randomized Clinical Trial for the Prevention of SARS-CoV-2 Infection (COVID-19) in Healthcare Personnel | recruiting |
| NCT04812496 [107] | Tenofovir-DF Versus Hydroxychloroquine in the Treatment of Hospitalized Patients With COVID-19 (TEDHICOV) | completed but no results posted |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zanella, I.; Zizioli, D.; Castelli, F.; Quiros-Roldan, E. Tenofovir, Another Inexpensive, Well-Known and Widely Available Old Drug Repurposed for SARS-COV-2 Infection. Pharmaceuticals 2021, 14, 454. https://doi.org/10.3390/ph14050454
Zanella I, Zizioli D, Castelli F, Quiros-Roldan E. Tenofovir, Another Inexpensive, Well-Known and Widely Available Old Drug Repurposed for SARS-COV-2 Infection. Pharmaceuticals. 2021; 14(5):454. https://doi.org/10.3390/ph14050454
Chicago/Turabian StyleZanella, Isabella, Daniela Zizioli, Francesco Castelli, and Eugenia Quiros-Roldan. 2021. "Tenofovir, Another Inexpensive, Well-Known and Widely Available Old Drug Repurposed for SARS-COV-2 Infection" Pharmaceuticals 14, no. 5: 454. https://doi.org/10.3390/ph14050454
APA StyleZanella, I., Zizioli, D., Castelli, F., & Quiros-Roldan, E. (2021). Tenofovir, Another Inexpensive, Well-Known and Widely Available Old Drug Repurposed for SARS-COV-2 Infection. Pharmaceuticals, 14(5), 454. https://doi.org/10.3390/ph14050454







