Is Standardization Necessary for Sharing of a Large Mid-Infrared Soil Spectral Library?
Abstract
1. Introduction
2. Materials and Methods
- Can useful predictions of soil properties be achieved on a secondary instrument without accounting for calibration transfer or having to build new models directly from the secondary spectra? Relatedly, how much of a loss in predictive performance is incurred by using spectra obtained on the secondary instrument?
- Can calibration transfer significantly improve predictions on the secondary instrument? What is the optimal combination of calibration transfer and predictive modeling approach?
2.1. Soil Samples, Measured Properties and Scanning Spectrometers
2.2. Spectral Preprocessing and Calibration Transfer
2.3. Outlier Detection and Predictive Model Development
2.4. Model Performance
3. Results
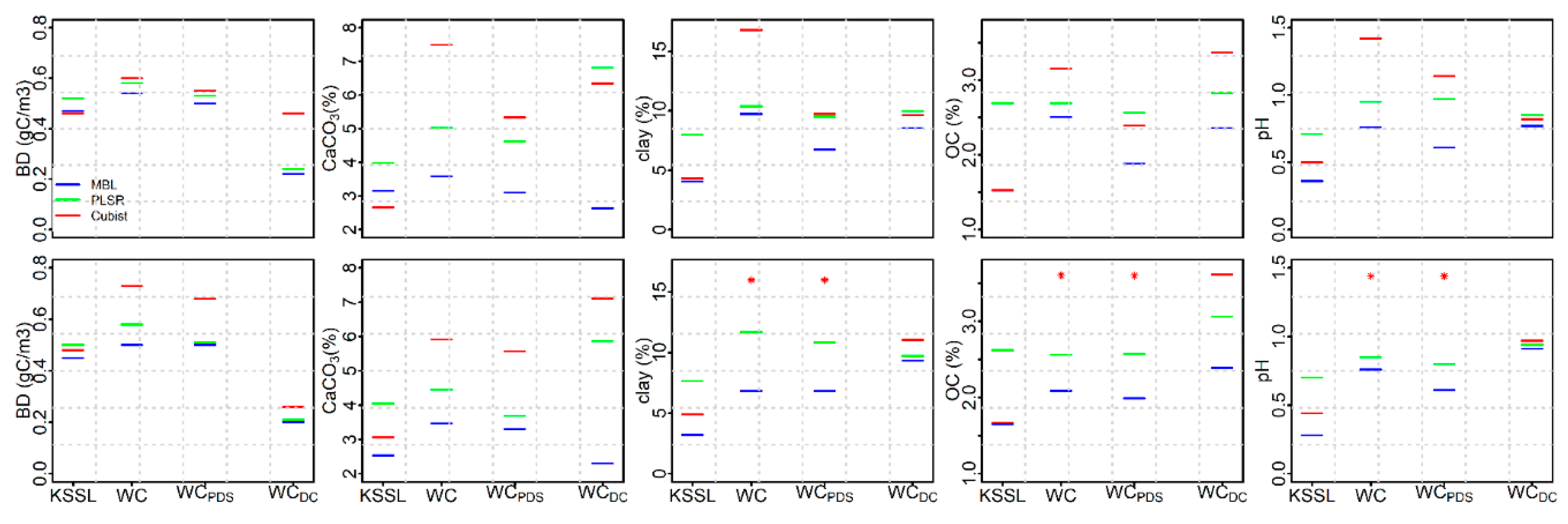
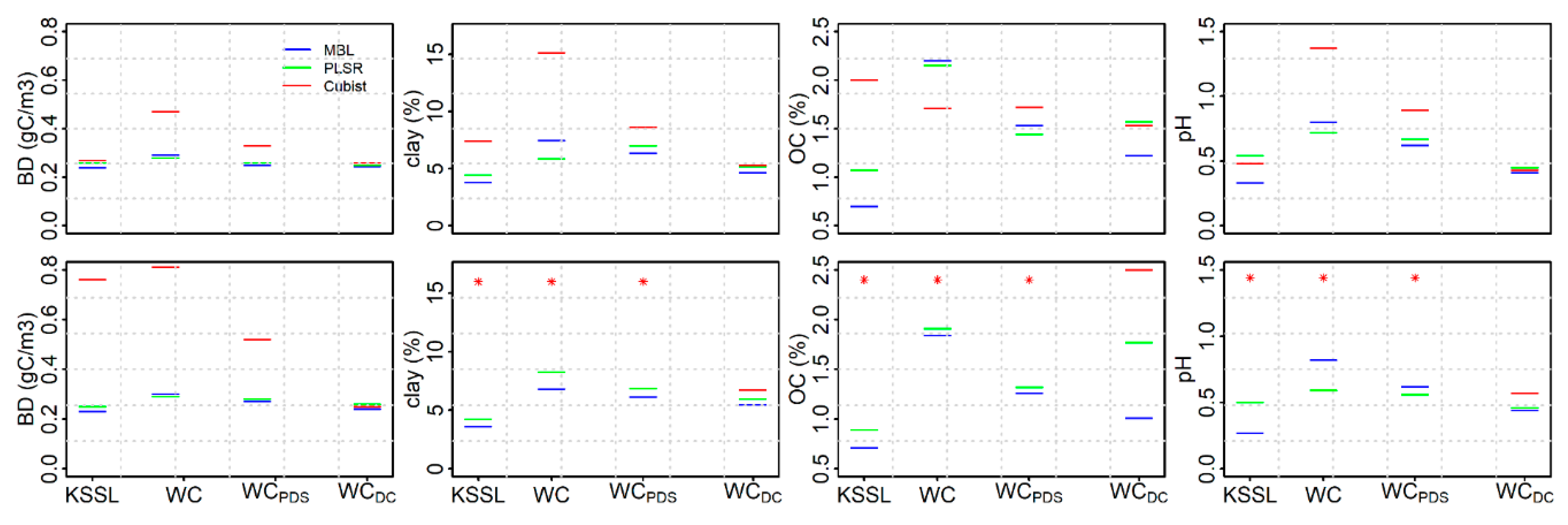
3.1. Performance of KSSL Predictive Models
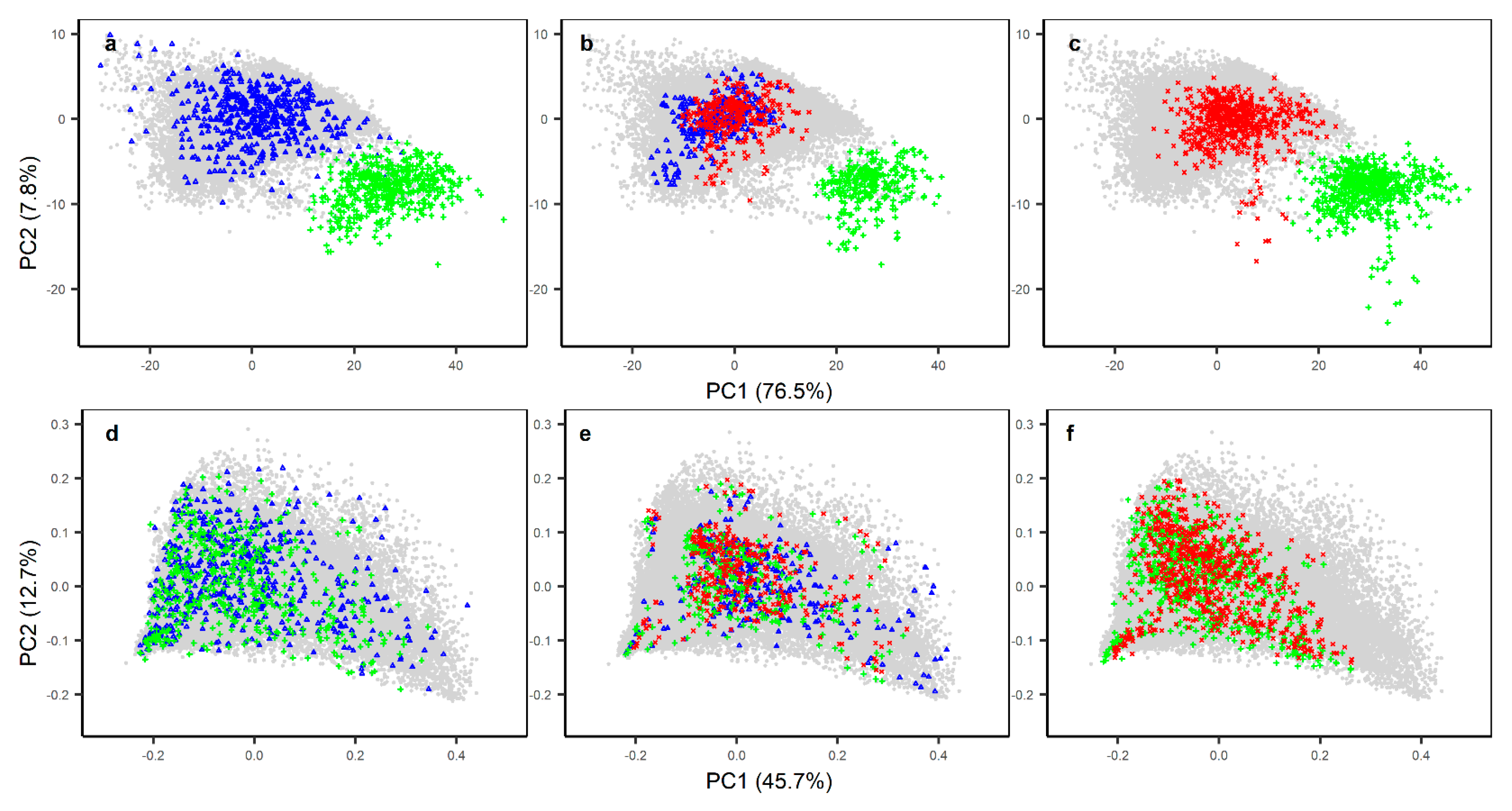
3.2. Performance of KSSL Predictive Models on KSSL and Woodwell Spectra
3.3. The Effect of Calibration Transfer on Secondary (Woodwell) Spectra
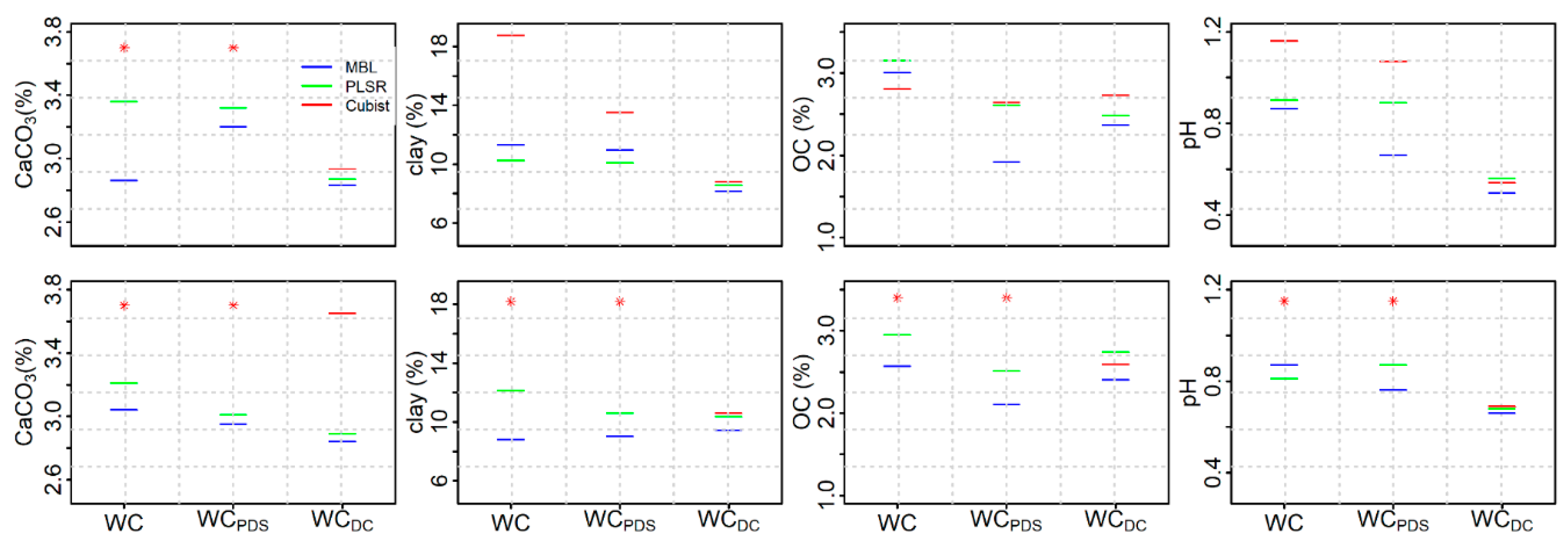
3.4. Performance of KSSL Predictive Models on Calibration Transfer (WoodwellPDS) Spectra
3.5. Performance of Woodwell Direct Calibration Models Using Leave-One-Out Cross Validation
4. Discussion
4.1. Model Performance without Accounting for Calibration Transfer
4.2. Effect of Calibration Transfer on Spectra Acquired Using Different Spectrometers
4.3. The Need for Calibration Transfer
4.4. Direct Calibration Using WOODWELL Spectra Are Not Always the Best Model
4.5. Best Model Performance
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- McCarty, G.W.; Reeves, J.B.; Reeves, V.B.; Follett, R.F.; Kimble, J.M. Mid-infrared and near-infrared diffuse reflectance spectroscopy for soil carbon measurement. Soil Sci. Soc. Am. J. 2002, 66, 640–646. [Google Scholar] [CrossRef]
- Shepherd, K.D.; Walsh, M.G. Development of reflectance spectral libraries for characterization of soil properties. Soil Sci. Soc. Am. J. 2002, 66, 988–998. [Google Scholar] [CrossRef]
- Chang, C.-W.; Laird, D.A.; Mausbach, M.J.; Hurburgh, C.R. Near-infrared reflectance spectroscopy–principal components regression analyses of soil properties. Soil Sci. Soc. Am. J. 2001, 65, 480–490. [Google Scholar] [CrossRef]
- Stenberg, B.; Rossel, R.A.V.; Mouazen, A.M.; Wetterlind, J. Visible and near infrared spectroscopy in soil science. In Advances in Agronomy; Elsevier: Amsterdam, The Netherlands, 2010; Volume 107, pp. 163–215. ISBN 0065-2113. [Google Scholar]
- Fernandez-Ahumada, E.; Garrido-Varo, A.; Guerrero, J.E.; Perez-Marin, D.; Fearn, T. Taking NIR calibrations of feed compounds from the laboratory to the process: Calibration transfer between predispersive and postdispersive instruments. J. Agric. Food Chem. 2008, 56, 10135–10141. [Google Scholar] [CrossRef] [PubMed]
- Ge, Y.; Morgan, C.L.; Grunwald, S.; Brown, D.J.; Sarkhot, D.V. Comparison of soil reflectance spectra and calibration models obtained using multiple spectrometers. Geoderma 2011, 161, 202–211. [Google Scholar] [CrossRef]
- Liu, Y.; Jiang, Q.; Fei, T.; Wang, J.; Shi, T.; Guo, K.; Li, X.; Chen, Y. Transferability of a visible and near-infrared model for soil organic matter estimation in riparian landscapes. Remote Sens. 2014, 6, 4305–4322. [Google Scholar] [CrossRef]
- Honorato, F.A.; Galvão, R.K.H.; Pimentel, M.F.; de Barros Neto, B.; Araújo, M.C.U.; de Carvalho, F.R. Robust modeling for multivariate calibration transfer by the successive projections algorithm. Chemom. Intell. Lab. Syst. 2005, 76, 65–72. [Google Scholar] [CrossRef]
- Seybold, C.A.; Ferguson, R.; Wysocki, D.; Bailey, S.; Anderson, J.; Nester, B.; Schoeneberger, P.; Wills, S.; Libohova, Z.; Hoover, D. Application of Mid-Infrared Spectroscopy in Soil Survey. Soil Sci. Soc. Am. J. 2019, 83, 1746–1759. [Google Scholar] [CrossRef]
- Baldock, J.A.; Hawke, B.; Sanderman, J.; Macdonald, L.M. Predicting contents of carbon and its component fractions in Australian soils from diffuse reflectance mid-infrared spectra. Soil Res. 2013, 51, 577–595. [Google Scholar] [CrossRef]
- Brown, D.J.; Shepherd, K.D.; Walsh, M.G.; Mays, M.D.; Reinsch, T.G. Global soil characterization with VNIR diffuse reflectance spectroscopy. Geoderma 2006, 132, 273–290. [Google Scholar] [CrossRef]
- Viscarra Rossel, R.A.; Webster, R.; Bui, E.N.; Baldock, J.A. Baseline map of organic carbon in Australian soil to support national carbon accounting and monitoring under climate change. Glob. Chang. Biol. 2014, 20, 2953–2970. [Google Scholar] [CrossRef] [PubMed]
- Dangal, S.R.; Sanderman, J.; Wills, S.; Ramirez-Lopez, L. Accurate and Precise Prediction of Soil Properties from a Large Mid-Infrared Spectral Library. Soil Syst. 2019, 3, 11. [Google Scholar] [CrossRef]
- Andrew, A.; Fearn, T. Transfer by orthogonal projection: Making near-infrared calibrations robust to between-instrument variation. Chemom. Intell. Lab. Syst. 2004, 72, 51–56. [Google Scholar] [CrossRef]
- Feudale, R.N.; Woody, N.A.; Tan, H.; Myles, A.J.; Brown, S.D.; Ferré, J. Transfer of multivariate calibration models: A review. Chemom. Intell. Lab. Syst. 2002, 64, 181–192. [Google Scholar] [CrossRef]
- Workman, J.J. A review of calibration transfer practices and instrument differences in spectroscopy. Appl. Spectrosc. 2018, 72, 340–365. [Google Scholar] [CrossRef] [PubMed]
- Shi, G.; Han, L.; Yang, Z.; Chen, L.; Liu, X. Near infrared spectroscopy calibration transfer for quantitative analysis of fish meal mixed with soybean meal. J. Near Infrared Spectrosc. 2010, 18, 217–223. [Google Scholar] [CrossRef]
- Ji, W.; Li, S.; Chen, S.; Shi, Z.; Rossel, R.A.V.; Mouazen, A.M. Prediction of soil attributes using the Chinese soil spectral library and standardized spectra recorded at field conditions. Soil Tillage Res. 2016, 155, 492–500. [Google Scholar] [CrossRef]
- Alam, T.M.; Alam, M.K.; McIntyre, S.K.; Volk, D.E.; Neerathilingam, M.; Luxon, B.A. Investigation of chemometric instrumental transfer methods for high-resolution NMR. Anal. Chem. 2009, 81, 4433–4443. [Google Scholar] [CrossRef]
- Salguero-Chaparro, L.; Palagos, B.; Peña-Rodríguez, F.; Roger, J.M. Calibration transfer of intact olive NIR spectra between a pre-dispersive instrument and a portable spectrometer. Comput. Electron. Agric. 2013, 96, 202–208. [Google Scholar] [CrossRef]
- Yu, B.; Ji, H.; Kang, Y. Standardization of near infrared spectra based on multi-task learning. Spectrosc. Lett. 2016, 49, 23–29. [Google Scholar] [CrossRef]
- Gemperline, P.J.; Cho, J.; Aldridge, P.K.; Sekulic, S.S. Appearance of discontinuities in spectra transformed by the piecewise direct instrument standardization procedure. Anal. Chem. 1996, 68, 2913–2915. [Google Scholar] [CrossRef] [PubMed]
- De Noord, O.E. Multivariate calibration standardization. Chemom. Intell. Lab. Syst. 1994, 25, 85–97. [Google Scholar] [CrossRef]
- Lin, Z.; Wu, B.; Wang, H.; Wang, Q. Study on near infrared spectroscopy model transfer by using piecewise direct standardization at different temperature. J. Instrum. Anal. 2008, 27, 1330–1333. [Google Scholar]
- Kennard, R.W.; Stone, L.A. Computer aided design of experiments. Technometrics 1969, 11, 137–148. [Google Scholar] [CrossRef]
- Soil Survey Staff. Kellogg Soil Survey Laboratory Methods Manual; Soil Survey Investigations Report; Report No. 42; United States Department of Agriculture (USDA); Natural Resources Conservation Service (NRCS): Washington, DC, USA, 2014.
- Tóth, G.; Jones, A.; Montanarella, L. LUCAS Topsoil Survey—Methodology, Data and Results, JRC Technical Reports EUR 26102; Publication Office of the European Union: Luxembourg, 2013. [Google Scholar]
- Wang, Y.; Lysaght, M.J.; Kowalski, B.R. Improvement of multivariate calibration through instrument standardization. Anal. Chem. 1992, 64, 562–564. [Google Scholar] [CrossRef]
- Ertlen, D.; Schwartz, D.; Trautmann, M.; Webster, R.; Brunet, D. Discriminating between organic matter in soil from grass and forest by near-infrared spectroscopy. Eur. J. Soil Sci. 2010, 61, 207–216. [Google Scholar] [CrossRef]
- Waruru, B.K.; Shepherd, K.D.; Ndegwa, G.M.; Sila, A.; Kamoni, P.T. Application of mid-infrared spectroscopy for rapid characterization of key soil properties for engineering land use. Soils Found. 2015, 55, 1181–1195. [Google Scholar] [CrossRef]
- Swierenga, H.; Haanstra, W.G.; De Weijer, A.P.; Buydens, L.M.C. Comparison of two different approaches toward model transferability in NIR spectroscopy. Appl. Spectrosc. 1998, 52, 7–16. [Google Scholar] [CrossRef]
- Bergman, E.-L.; Brage, H.; Josefson, M.; Svensson, O.; Sparén, A. Transfer of NIR calibrations for pharmaceutical formulations between different instruments. J. Pharm. Biomed. Anal. 2006, 41, 89–98. [Google Scholar] [CrossRef]
- Abdi, H. Partial least squares regression and projection on latent structure regression (PLS Regression). Wiley Interdiscip. Rev. Comput. Stat. 2010, 2, 97–106. [Google Scholar] [CrossRef]
- Kuhn, M.; Weston, S.; Keefer, C.; Coulter, N. Cubist Models for Regression. R Package Vignette R Package Version 0.0. 2012. Volume 18, pp. 223–244. Available online: https://cran.r-project.org/web/packages/Cubist/Cubist.pdf (accessed on 20 November 2020).
- Ramirez-Lopez, L.; Behrens, T.; Schmidt, K.; Rossel, R.; Dematte, J.; Scholten, T. Distance and similarity-search metrics for use with soil vis-NIR spectra. Geoderma 2013, 199, 43–53. [Google Scholar] [CrossRef]
- Ramirez-Lopez, L.; Stevens, A. Resemble: Regression and Similarity Evaluation for Memory-Based Learning in Spectral Chemometrics. R Package Version 122. Available online: https://cran.r-project.org/web/packages/resemble/resemble.pdf (accessed on 20 November 2020).
- Bellon-Maurel, V.; McBratney, A. Near-infrared (NIR) and mid-infrared (MIR) spectroscopic techniques for assessing the amount of carbon stock in soils–Critical review and research perspectives. Soil Biol. Biochem. 2011, 43, 1398–1410. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2018; Available online: https://www.r-project.org/ (accessed on 20 November 2020).
- Terhoeven-Urselmans, T.; Vagen, T.-G.; Spaargaren, O.; Shepherd, K.D. Prediction of soil fertility properties from a globally distributed soil mid-infrared spectral library. Soil Sci. Soc. Am. J. 2010, 74, 1792–1799. [Google Scholar] [CrossRef]
- Rossel, R.V.; Behrens, T. Using data mining to model and interpret soil diffuse reflectance spectra. Geoderma 2010, 158, 46–54. [Google Scholar] [CrossRef]
- Friedman, J.H. Greedy function approximation: A gradient boosting machine. Ann. Stat. 2001, 1189–1232. [Google Scholar] [CrossRef]
- Ramirez-Lopez, L.; Behrens, T.; Schmidt, K.; Stevens, A.; Demattê, J.A.M.; Scholten, T. The spectrum-based learner: A new local approach for modeling soil vis–NIR spectra of complex datasets. Geoderma 2013, 195, 268–279. [Google Scholar] [CrossRef]
- Minasny, B.; McBratney, A.B.; Stockmann, U.; Hong, S.Y. Cubist, a Regression Rule Approach for use in Calibration of NIR Spectra. In Proceedings of the NIR 2013—16th International Conference on Near Infrared Spectroscopy, la Grande-Motte, France, 2–7 June 2013; Volume 630. [Google Scholar]
- Coûteaux, M.-M.; Berg, B.; Rovira, P. Near infrared reflectance spectroscopy for determination of organic matter fractions including microbial biomass in coniferous forest soils. Soil Biol. Biochem. 2003, 35, 1587–1600. [Google Scholar] [CrossRef]
- Minasny, B.; McBratney, A.B. Regression rules as a tool for predicting soil properties from infrared reflectance spectroscopy. Chemom. Intell. Lab. Syst. 2008, 94, 72–79. [Google Scholar] [CrossRef]
- Wang, Z.; Dean, T.; Kowalski, B. Additive background correction in multivariate instrument standardization. Anal. Chem. 1995, 67, 2379–2385. [Google Scholar] [CrossRef]
- Luo, X.; Ikehata, A.; Sashida, K.; Piao, S.; Okura, T.; Terada, Y. Calibration transfer across near infrared spectrometers for measuring hematocrit in the blood of grazing cattle. J. Near Infrared Spectrosc. 2017, 25, 15–25. [Google Scholar] [CrossRef]
- Rossel, R.V.; Walvoort, D.J.J.; McBratney, A.B.; Janik, L.J.; Skjemstad, J.O. Visible, near infrared, mid infrared or combined diffuse reflectance spectroscopy for simultaneous assessment of various soil properties. Geoderma 2006, 131, 59–75. [Google Scholar] [CrossRef]
- Kunz, M.R.; Ottaway, J.; Kalivas, J.H.; Andries, E. Impact of standardization sample design on Tikhonov regularization variants for spectroscopic calibration maintenance and transfer. J. Chemom. 2010, 24, 218–229. [Google Scholar] [CrossRef]
- Zhao, Y.; Zhao, Z.; Shan, P.; Peng, S.; Yu, J.; Gao, S. Calibration Transfer Based on Affine Invariance for NIR without Transfer Standards. Molecules 2019, 24, 1802. [Google Scholar] [CrossRef] [PubMed]
- Malli, B.; Birlutiu, A.; Natschläger, T. Standard-free calibration transfer-An evaluation of different techniques. Chemom. Intell. Lab. Syst. 2017, 161, 49–60. [Google Scholar] [CrossRef]
- Dor, E.B.; Ong, C.; Lau, I.C. Reflectance measurements of soils in the laboratory: Standards and protocols. Geoderma 2015, 245, 112–124. [Google Scholar]
- Sagi, O.; Rokach, L. Ensemble learning: A survey. Wiley Interdiscip. Rev. Data Min. Knowl. Discov. 2018, 8, e1249. [Google Scholar] [CrossRef]
| Samples | Variables | N | Mean | Median | SD | Q25 | Q75 |
|---|---|---|---|---|---|---|---|
| Training Set | BD (g/cm3) | 13,667 | 1.15 | 1.26 | 0.44 | 0.95 | 1.46 |
| CaCO3 (wt %) | 9365 | 10.01 | 4.92 | 12.88 | 1.04 | 14.72 | |
| clay (wt %) | 37,187 | 22.71 | 20.93 | 15.79 | 3.15 | 5.70 | |
| OC (wt %) | 55,598 | 8.30 | 1.33 | 14.86 | 0.42 | 5.39 | |
| pH | 39,347 | 6.41 | 6.26 | 1.29 | 5.41 | 7.54 | |
| Set I | BD (g/cm3) | 110 | 1.76 | 1.78 | 0.27 | 1.61 | 1.92 |
| CaCO3 (wt %) | 216 | 12.39 | 3.33 | 18.53 | 0.32 | 15.53 | |
| clay (wt %) | 321 | 24.58 | 20.67 | 19.54 | 7.41 | 37.22 | |
| OC (wt %) | 420 | 6.68 | 0.81 | 12.85 | 0.24 | 5.60 | |
| pH | 341 | 6.58 | 6.50 | 1.48 | 5.34 | 7.93 | |
| Set II | BD (g/cm3) | 290 | 1.23 | 1.31 | 0.42 | 1.04 | 1.52 |
| (NEON) | clay (wt %) | 286 | 17.21 | 13.00 | 14.22 | 5.63 | 25.98 |
| OC (%) | 296 | 3.64 | 0.48 | 9.34 | 0.13 | 2.04 | |
| pH | 296 | 6.40 | 5.84 | 1.41 | 5.34 | 7.90 | |
| Set III | CaCO3 (wt %) | 605 | 4.66 | 0.10 | 10.45 | 0.00 | 2.40 |
| (LUCAS) | clay (wt %) | 605 | 21.68 | 17.00 | 17.75 | 7.00 | 34.00 |
| OC (wt %) | 605 | 5.14 | 1.99 | 9.66 | 1.14 | 3.45 | |
| pH | 605 | 6.32 | 6.32 | 1.31 | 5.21 | 7.53 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dangal, S.R.S.; Sanderman, J. Is Standardization Necessary for Sharing of a Large Mid-Infrared Soil Spectral Library? Sensors 2020, 20, 6729. https://doi.org/10.3390/s20236729
Dangal SRS, Sanderman J. Is Standardization Necessary for Sharing of a Large Mid-Infrared Soil Spectral Library? Sensors. 2020; 20(23):6729. https://doi.org/10.3390/s20236729
Chicago/Turabian StyleDangal, Shree R. S., and Jonathan Sanderman. 2020. "Is Standardization Necessary for Sharing of a Large Mid-Infrared Soil Spectral Library?" Sensors 20, no. 23: 6729. https://doi.org/10.3390/s20236729
APA StyleDangal, S. R. S., & Sanderman, J. (2020). Is Standardization Necessary for Sharing of a Large Mid-Infrared Soil Spectral Library? Sensors, 20(23), 6729. https://doi.org/10.3390/s20236729




