Reducing Response Time in Motor Imagery Using A Headband and Deep Learning †
Abstract
1. Introduction
2. Related Work
3. Materials and Methods
3.1. Experiment Protocols
3.1.1. Exploring Response Time Reduction
- The participant is sitting down on a chair with arms extended in parallel, resting on a table.
- A bottle of water is on the table, approximately 5 cm to the left of the left hand.
- The participant’s head faces forward, while the eyes rotate to the left, looking to the bottle.
- We asked the participant to imagine picking up the bottle with the left hand, but without moving the hand only thinking about it for 20 s.
- Then, we asked the participant to relax and to close the eyes for 20 s.
- We move the bottle on the right side of the table (approximately 5 cm to the right of the right hand).
- The participant repeats steps 3–5, but for right hand motor imagery.
- We repeated steps 2–7 for 20 times for each participant.
3.1.2. Validating the Response Time Reduction
- The participant is sitting down on a chair with arms extended in parallel, resting on a table.
- Two bottles of water are on the table. One of them is approximately 5 cm to the left of the left hand and the other bottle is 5 cm to the right of the right hand.
- The participant’s head face forward, while the eyes rotate to the left, looking to the bottle.
- We asked the participant to imagine picking up the bottle with the right hand, but without moving the hand; only thinking about it for 6 s.
- Then, we asked the participant to imagine picking up the bottle with the left hand, but without moving the hand, only thinking about it for 6 s.
- We repeated steps 4–5 for 5 times for each participant: 1 min in total.
- Then, we repeated steps 1–6 for 20 times.
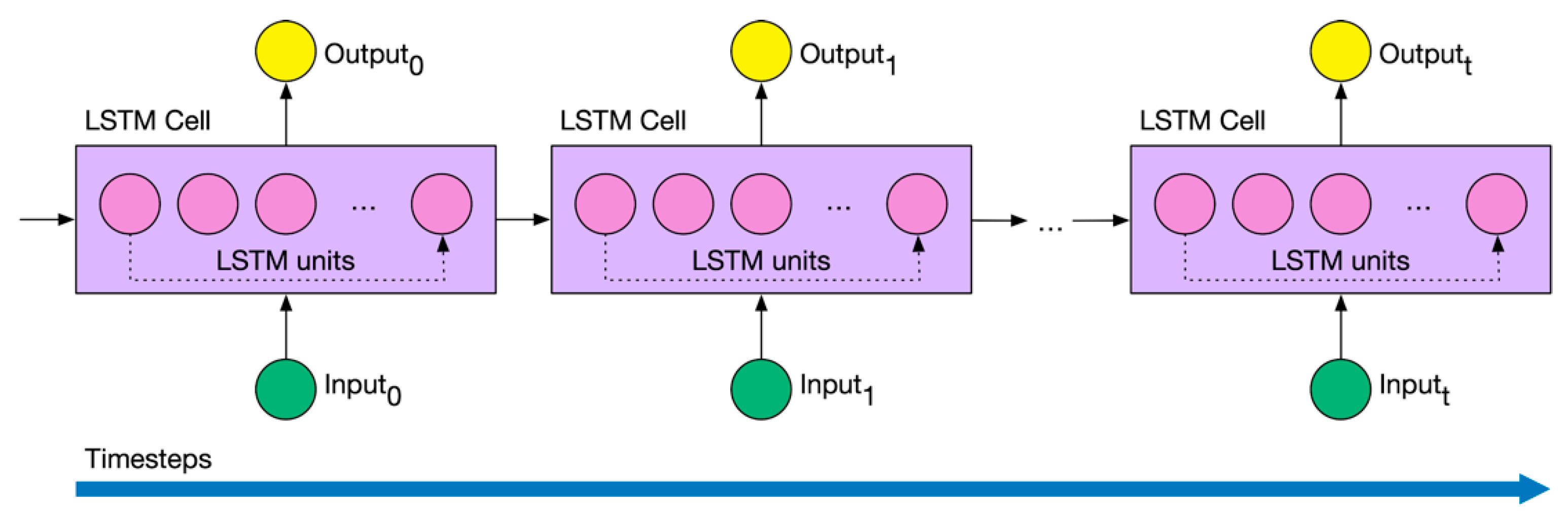
3.2. Deep Learning Foundations
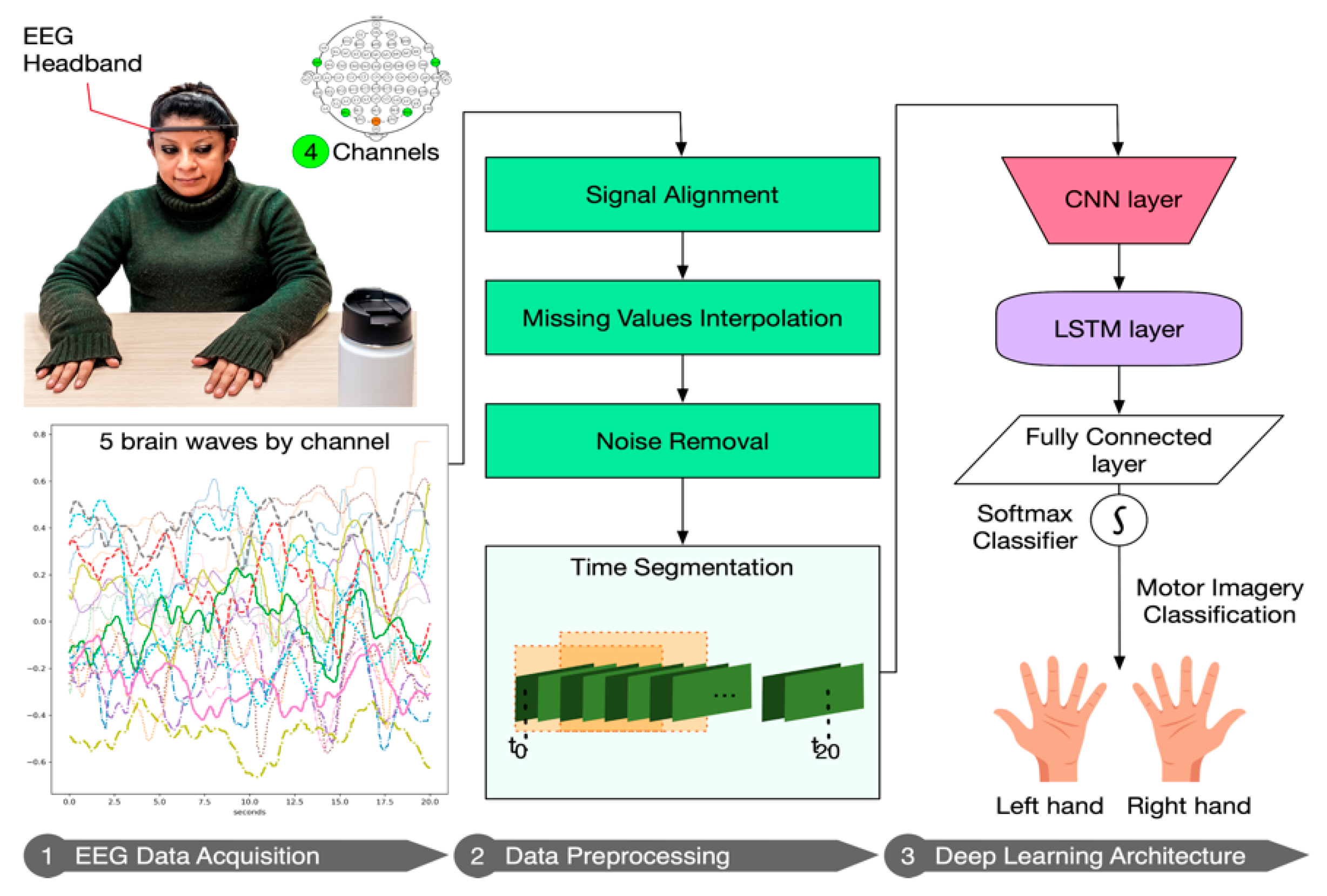
3.3. Deep Learning Pipeline
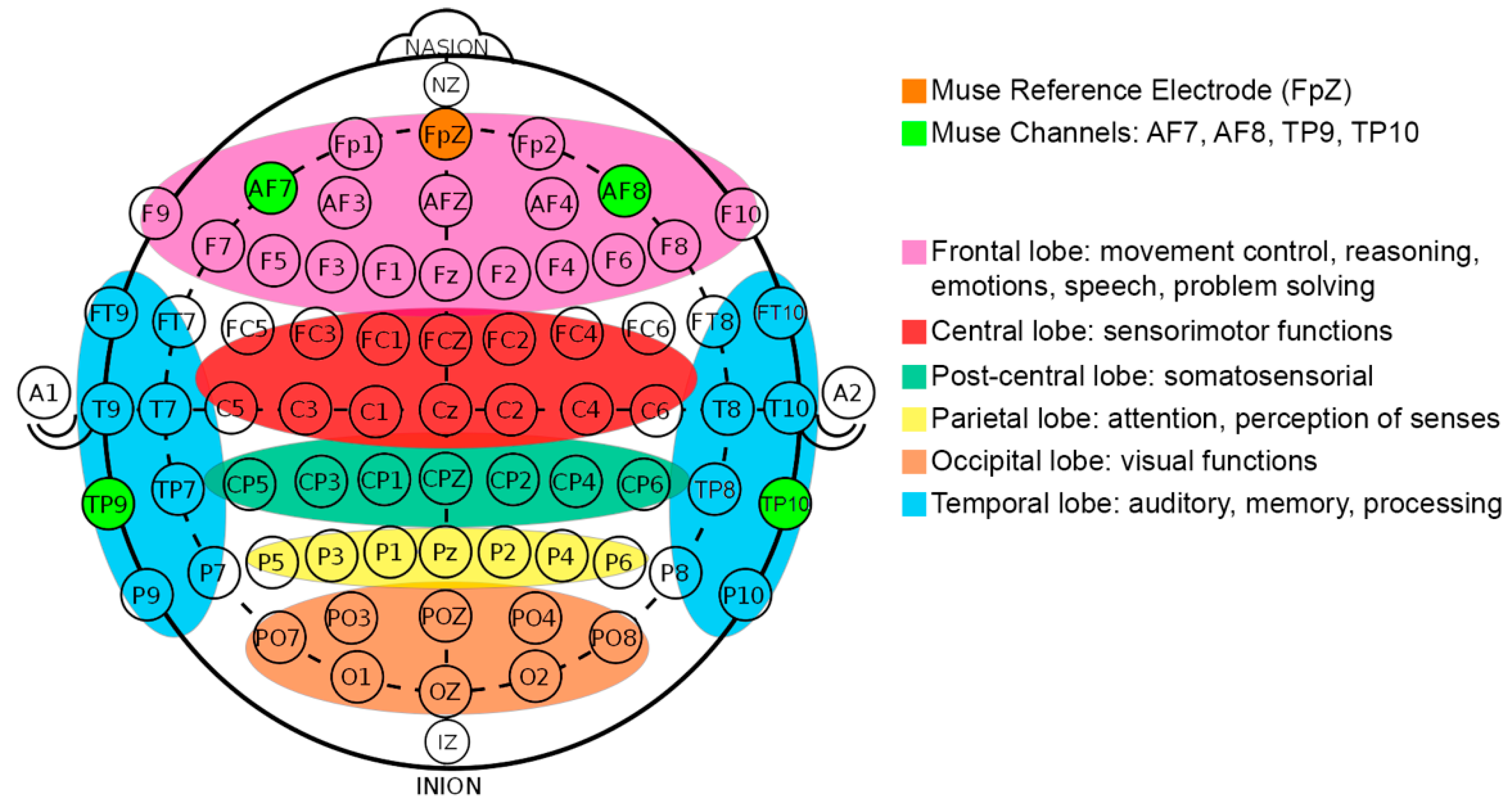
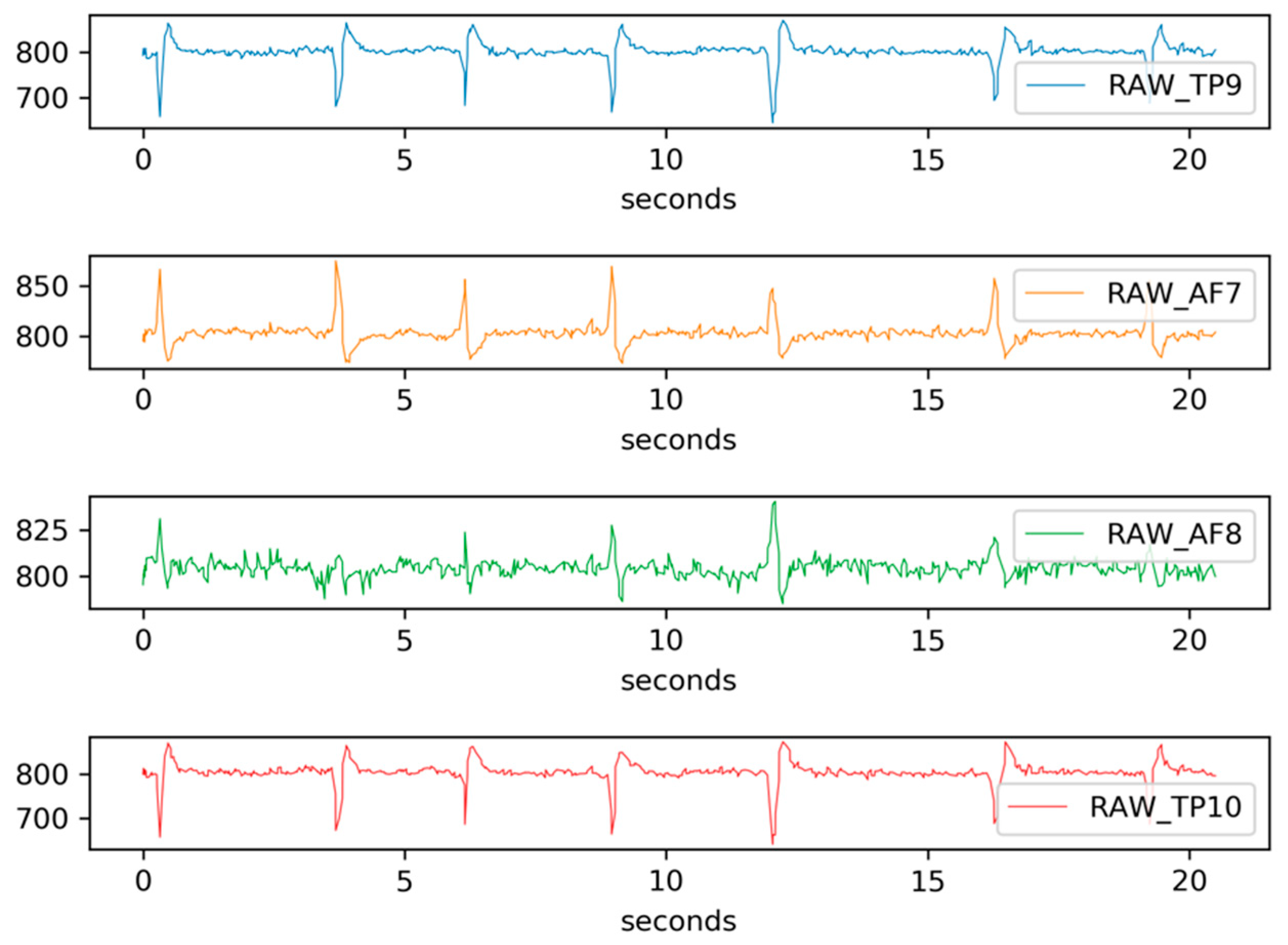
3.3.1. EEG Data Acquisition
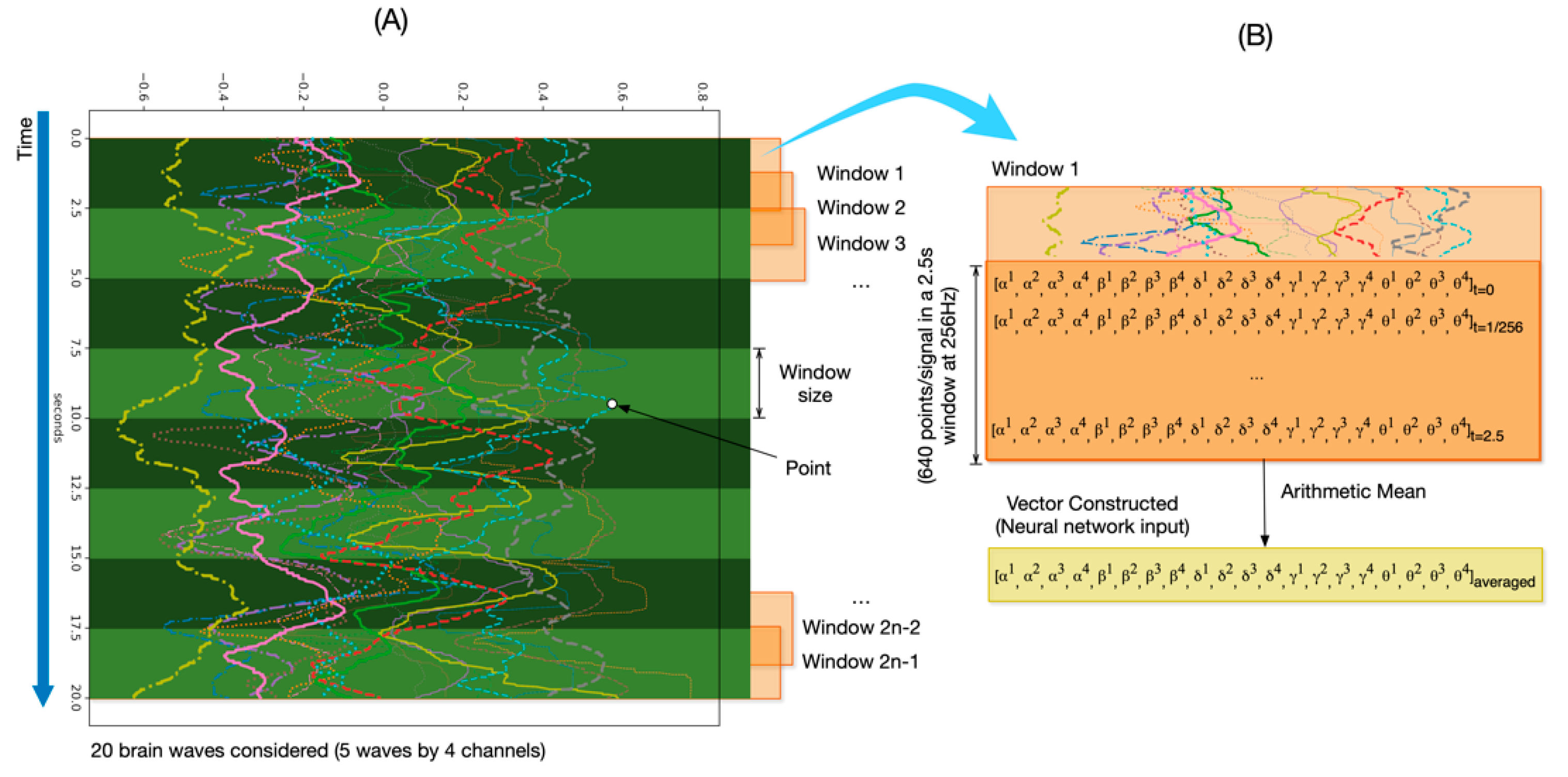
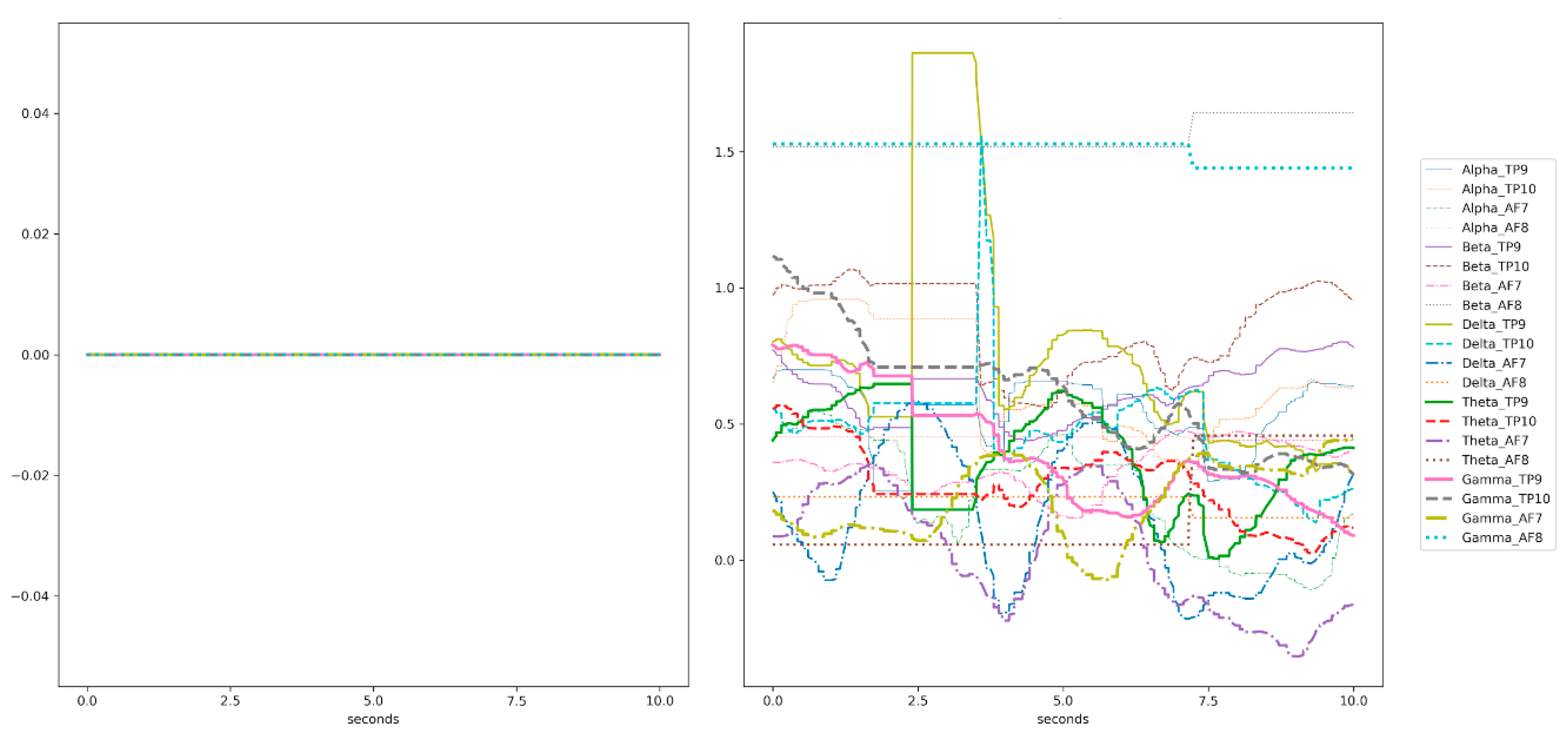
3.3.2. Data Preprocessing
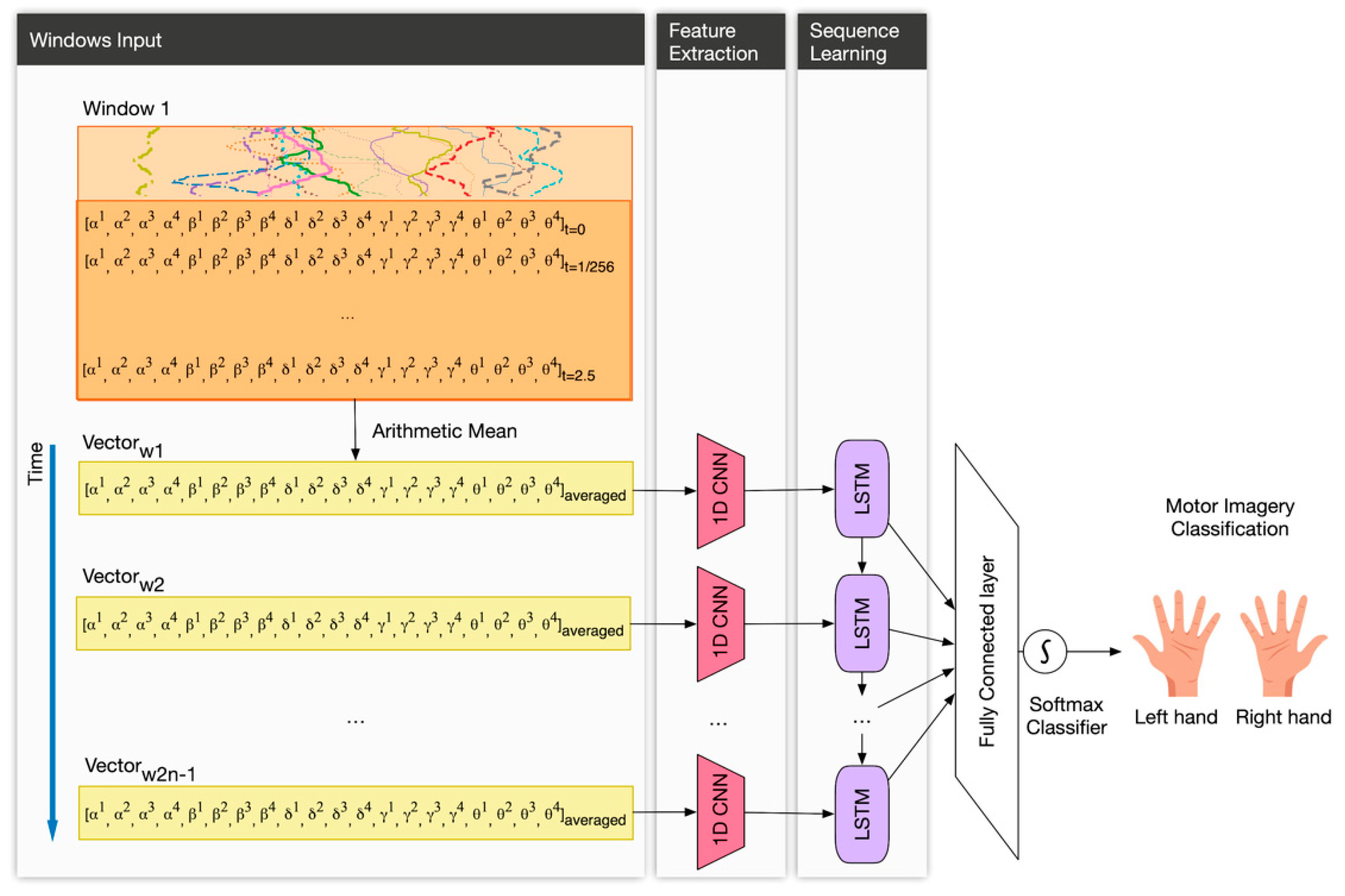
3.3.3. Deep Learning Architecture
- The input layer. This layer is three-dimensional following this triplet (samples: None, timesteps: number of windows, features: 20). In our case, samples are the number of observations, i.e., the total number of recordings, which the DL model does not know a priori and for that reason is “None”. Since we used 50% overlapping of sliding windows, we defined the timesteps as the number of windows covering the full session (in total, 2n − 1 windows). In addition, features, in our case, are the number of signals, i.e., twenty.
- A 1D-CNN layer with 32 filters of size 1; and kernel size equals 1, which means that each output is calculated based on the previous 1 timestep.
- A LSTM layer with 32 neurons. In order to prevent overfitting, we used 0.1 dropout, 0.0000001 regularizer and early stopping [44]. Early stopping ensures that the neural network stops the training at a certain epoch where further training would overfit the model.
- A fully connected layer with a softmax function activation, for performing the classification of the two motor intentions.
4. Results
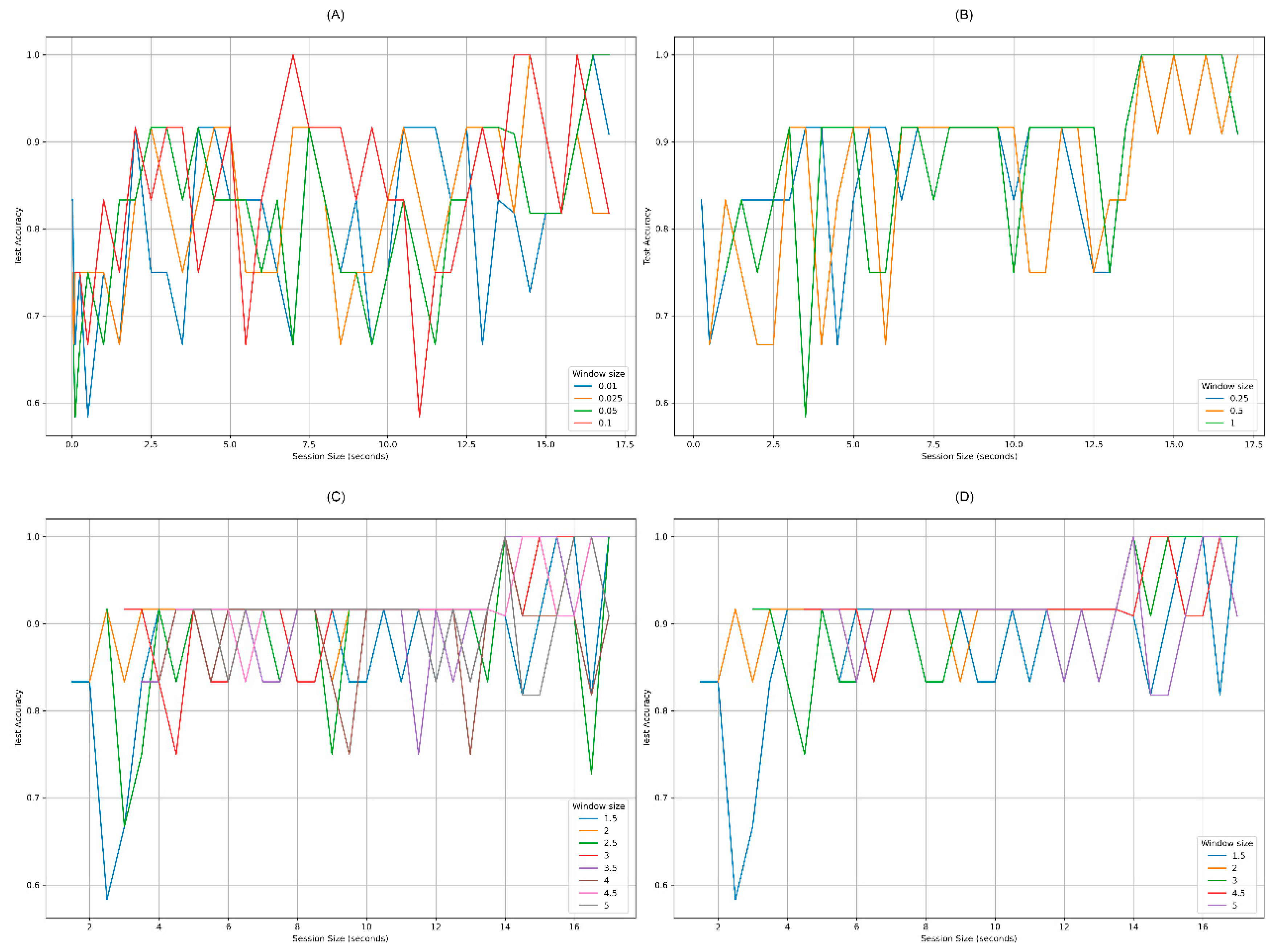
4.1. Set of Experiments 1: Exploring Response Time Reduction
4.2. Experiment 2: Validating the Response Time Reduction
5. Conclusions and Future Work
Author Contributions
Funding

Acknowledgments
Conflicts of Interest
Abbreviation
| AFX | position of an electrode in BCI, with number X |
| BCI | brain computer interfaces |
| CNN | convolutional neural network |
| DL | deep learning |
| EEG | electroencephalography |
| Fpz | reference electrode in BCI |
| LR | logistic regression |
| LSTM | long short-term memory |
| ML | machine learning |
| NN | neural network |
| RNN | recurrent neural network |
| SDK | software development kits |
| SVM | support vector machines |
| TPX | position of an electrode in BCI, with number X |
References
- Dunn, J.; Runge, R.; Snyder, M. Wearables and the medical revolution. Per. Med. 2018, 15, 429–448. [Google Scholar] [CrossRef] [PubMed]
- Aggravi, M.; Salvietti, G.; Prattichizzo, D. Haptic assistive bracelets for blind skier guidance. In Proceedings of the 7th Augmented Human International Conference 2016 on–AH ’16, Geneva, Switzerland, 25–27 February 2016; pp. 1–4. [Google Scholar]
- Majumder, S.; Mondal, T.; Deen, M. Wearable sensors for remote health monitoring. Sensors 2017, 17, 130. [Google Scholar] [CrossRef] [PubMed]
- Elsaleh, T.; Enshaeifar, S.; Rezvani, R.; Acton, S.T.; Janeiko, V.; Bermudez-Edo, M. IoT-Stream: A lightweight ontology for internet of Things data streams and Its USE with data analytics and event Detection SERVICES. Sensors 2020, 20, 953. [Google Scholar] [CrossRef]
- Enshaeifar, S.; Zoha, A.; Markides, A.; Skillman, S.; Acton, S.T.; Elsaleh, T.; Hassanpour, M.; Ahrabian, A.; Kenny, M.; Klein, S.; et al. Health management and pattern analysis of daily living activities of people with dementia using in-home sensors and machine learning techniques. PLoS ONE 2018, 13, e0195605. [Google Scholar] [CrossRef] [PubMed]
- Fico, G.; Montalva, J.-B.; Medrano, A.; Liappas, N.; Mata-Díaz, A.; Cea, G.; Arredondo, M.T. Co-creating with consumers and stakeholders to understand the benefit of internet of things in smart living environments for ageing well: The approach adopted in the Madrid Deployment Site of the ACTIVAGE Large Scale Pilot. In IFMBE Proceedings; Springer: Singapore, 2018; pp. 1089–1092. [Google Scholar]
- Greene, B.R.; McManus, K.; Redmond, S.J.; Caulfield, B.; Quinn, C.C. Digital assessment of falls risk, frailty, and mobility impairment using wearable sensors. Npj Digit. Med. 2019, 2, 1–7. [Google Scholar] [CrossRef]
- Zhang, X.; Yao, L.; Huang, C.; Sheng, Q.Z.; Wang, X. Intent recognition in smart living Through DEEP recurrent neural networks. In Proceedings of the International Conference on Neural Information Processing, Guangzhou, China, 14–18 November 2017; pp. 748–758. [Google Scholar]
- Zhang, X.; Yao, L.; Wang, X.; Monaghan, J.; Mcalpine, D.; Zhang, Y. A Survey on deep learning based brain Computer INTERFACE: Recent advances and New frontiers. arXiv 2019, arXiv:physics/0402096. [Google Scholar]
- Nicolas-Alonso, L.F.; Gomez-Gil, J. Brain computer interfaces–A review. Sensors 2012, 12, 1211–1279. [Google Scholar] [CrossRef]
- InteraXon Muse 2: Brain Sensing Headband–Technology Enhanced Meditatiomn. Available online: https://choosemuse.com/muse-2/ (accessed on 27 January 2020).
- Jurcak, V.; Tsuzuki, D.; Dan, I. 10/20, 10/10, and 10/5 systems revisited: Their validity as relative head-surface-based positioning systems. Neuroimage 2007, 34, 1600–1611. [Google Scholar] [CrossRef]
- Garcia-Moreno, F.M.; Bermudez-Edo, M.; Rodriguez-Fortiz, M.J.; Garrido, J.L. A CNN-LSTM deep Learning classifier for motor imagery EEG detection using a low-invasive and low-Cost BCI headband. In Proceedings of the 16th International Conference on Intelligent Environments (IE), Madrid, Spain, 20–23 July 2020; pp. 84–91. [Google Scholar]
- Bermudez-Edo, M.; Barnaghi, P. Spatio-temporal analysis for smart city data. In Proceedings of the Companion of the The Web Conference 2018 on The Web Conference 2018–WWW ’18, Lyon, France, 23–27 April 2018. [Google Scholar]
- LeCun, Y.; Bengio, Y.; Hinton, G. Deep learning. Nature 2015, 521, 436–444. [Google Scholar] [CrossRef]
- Ngiam, J.; Khosla, A.; Kim, M.; Nam, J.; Lee, H.; Ng, A.Y. Multimodal deep learning. In Proceedings of the 28th International Conference on Machine Learning, ICML 2011, Bellevue, WA, USA, 28 June–2 July 2011. [Google Scholar]
- Ravi, D.; Wong, C.; Deligianni, F.; Berthelot, M.; Andreu-Perez, J.; Lo, B.; Yang, G.-Z. Deep learning for health informatics. IEEE J. Biomed. Health Inf. 2017, 21, 4–21. [Google Scholar] [CrossRef]
- Roy, Y.; Banville, H.; Albuquerque, I.; Gramfort, A.; Falk, T.H.; Faubert, J. Deep learning-based electroencephalography analysis: A systematic review. J. Neural Eng. 2019, 16, 051001. [Google Scholar] [CrossRef] [PubMed]
- Ramadan, R.A.; Vasilakos, A.V. Brain computer interface: Control signals review. Neurocomputing 2017, 223, 26–44. [Google Scholar] [CrossRef]
- Bird, J.J.; Faria, D.R.; Manso, L.J.; Ekárt, A.; Buckingham, C.D. A deep evolutionary approach to bioinspired classifier optimisation for brain-machine interaction. Complexity 2019, 2019, 4316548. [Google Scholar] [CrossRef]
- Lotte, F. Signal processing approaches to minimize or suppress calibration time in oscillatory activity-based brain–computer interfaces. Proc. IEEE 2015, 103, 871–890. [Google Scholar] [CrossRef]
- Wiechert, G.; Triff, M.; Liu, Z.; Yin, Z.; Zhao, S.; Zhong, Z.; Zhaou, R.; Lingras, P. Identifying users and activities with cognitive signal processing from a wearable headband. In Proceedings of the IEEE 15th International Conference on Cognitive Informatics & Cognitive Computing (ICCI*CC), Palo Alto, CA, USA, 8 August 2016; pp. 129–136. [Google Scholar]
- Salehzadeh, A.; Calitz, A.P.; Greyling, J. Human activity recognition using deep electroencephalography learning. Biomed. Signal Process. Control 2020, 62, 102094. [Google Scholar] [CrossRef]
- Zhao, D.; MacDonald, S.; Gaudi, T.; Uribe-Quevedo, A.; Martin, M.V.; Kapralos, B. Facial expression detection employing a brain computer interface. In Proceedings of the 9th International Conference on Information, Intelligence, Systems and Applications (IISA), Zakynthos, Greece, 23–25 July 2018; pp. 1–2. [Google Scholar]
- Rodriguez, P.I.; Mejia, J.; Mederos, B.; Moreno, N.E.; Mendoza, V.M. Acquisition, analysis and classification of EEG signals for control design. arXiv 2018, arXiv:physics/0402096. [Google Scholar]
- Li, Z.; Xu, J.; Zhu, T. Recognition of Brain Waves of Left and Right Hand Movement Imagery with Portable Electroencephalographs. arXiv 2015, arXiv:physics/0402096. [Google Scholar]
- Zhang, D.; Yao, L.; Zhang, X.; Wang, S.; Chen, W.; Boots, R. EEG-based intention recognition from spatio-temporal representations via Cascade and parallel convolutional Recurrent neural networks. arXiv 2017, arXiv:physics/0402096. [Google Scholar]
- Chen, W.; Wang, S.; Zhang, X.; Yao, L.; Yue, L.; Qian, B.; Li, X. EEG-based motion intention recognition via multi-task RNNs. In Proceedings of the 2018 SIAM International Conference on Data Mining, San Diego, CA, USA, 3–5 May 2018; pp. 279–287. [Google Scholar]
- Kaya, M.; Binli, M.K.; Ozbay, E.; Yanar, H.; Mishchenko, Y. A large electroencephalographic motor imagery dataset for electroencephalographic brain computer interfaces. Sci. Data 2018, 5, 180211. [Google Scholar] [CrossRef]
- Brunner, C.; Leeb, R.; Müller-Putz, G.R.; Schlögl, A.; Pfurtscheller, G. BCI Competition IV 2008–Graz Data Set A. Available online: http://www.bbci.de/competition/iv/desc_2a.pdf (accessed on 24 November 2020).
- Brunner, C.; Leeb, R.; Müller-Putz, G.R.; Schlögl, A.; Pfurtscheller, G. BCI Competition 2008–Graz Data Set B. Available online: http://www.bbci.de/competition/iv/desc_2b.pdf (accessed on 24 November 2020).
- Bhattacharyya, S.; Khasnobish, A.; Konar, A.; Tibarewala, D.N.; Nagar, A.K. Performance analysis of left/right hand movement classification from EEG signal by intelligent algorithms. In Proceedings of the IEEE Symposium on Computational Intelligence, Cognitive Algorithms, Mind, and Brain (CCMB), Paris, France, 11–15 April 2011; pp. 1–8. [Google Scholar]
- Tang, Z.; Li, C.; Sun, S. Single-trial EEG classification of motor imagery using deep convolutional neural networks. Optik 2017, 130, 11–18. [Google Scholar] [CrossRef]
- Tabar, Y.R.; Halici, U. A novel deep learning approach for classification of EEG motor imagery signals. J. Neural Eng. 2017, 14, 016003. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Jiang, A.; Liu, X.; Shang, J.; Zhang, L. LSTM-based EEG classification in motor imagery tasks. IEEE Trans. Neural Syst. Rehabil. Eng. 2018, 26, 2086–2095. [Google Scholar] [CrossRef]
- Schirrmeister, R.T.; Springenberg, J.T.; Fiederer, L.D.J.; Glasstetter, M.; Eggensperger, K.; Tangermann, M.; Hutter, F.; Burgard, W.; Ball, T. Deep learning with convolutional neural networks for EEG decoding and visualization. Hum. Brain Mapp. 2017, 38, 5391–5420. [Google Scholar] [CrossRef] [PubMed]
- Ha, K.-W.; Jeong, J.-W. Motor imagery EEG classification using capsule networks. Sensors 2019, 19, 2854. [Google Scholar] [CrossRef] [PubMed]
- Lawhern, V.J.; Solon, A.J.; Waytowich, N.R.; Gordon, S.M.; Hung, C.P.; Lance, B.J. EEGNet: A compact convolutional neural network for EEG-based brain–computer interfaces. J. Neural Eng. 2018, 15, 056013. [Google Scholar] [CrossRef]
- Blankertz, B.; Dornhege, G.; Krauledat, M.; Muller, K.-R.; Kunzmann, V.; Losch, F.; Curio, G. The Berlin brain–computer interface: EEG-based communication without subject training. IEEE Trans. Neural Syst. Rehabil. Eng. 2006, 14, 147–152. [Google Scholar] [CrossRef]
- Tomioka, R.; Aihara, K.; Müller, K.-R. Logistic Regression for Single Trial EEG Classification. In Proceedings of the Twentieth Annual Conference on Neural Information Processing Systems, Vancouver, BC, Canada, 4–7 December 2006; pp. 1377–1384. [Google Scholar]
- Schmidt, R.F.; Thews, G. Human Phys.; Springer: Berlin/Heidelberg, Germany, 1989. [Google Scholar]
- Garcia-Moreno, F.M.; Bermudez-Edo, M.; Garrido, J.L.; Rodríguez-García, E.; Pérez-Mármol, J.M.; Rodríguez-Fórtiz, M.J. A microservices e-Health system for ecological frailty assessment using wearables. Sensors 2020, 20, 3427. [Google Scholar] [CrossRef]
- Bermudez-Edo, M.; Barnaghi, P.; Moessner, K. Analysing real world data streams with spatio-temporal correlations: Entropy vs. Pearson correlation. Autom. Constr. 2018, 88, 87–100. [Google Scholar] [CrossRef]
- Pereyra, G.; Tucker, G.; Chorowski, J.; Kaiser, Ł.; Hinton, G. Regularizing neural networks by penalizing confident output distributions. In Proceedings of the 5th International Conference on Learning Representations (ICLR 2017), Toulon, France, 24–26 April 2017. [Google Scholar]
- Chollet, F. Others Keras. Available online: https://keras.io (accessed on 24 November 2020).
- Lebedev, A.V.; Kaelen, M.; Lövdén, M.; Nilsson, J.; Feilding, A.; Nutt, D.J.; Carhart-Harris, R.L. LSD-induced entropic brain activity predicts subsequent personality change. Hum. Brain Mapp. 2016, 37, 3203–3213. [Google Scholar] [CrossRef]
- García-Moreno, F.M.; Rodríguez-García, E.; Rodríguez-Fórtizm, M.; Garrido, J.L.; Bermúdez-Edo, M.; Villaverde-Gutiérrez, C.; Pérez-Mármol, J.M. Designing a smart mobile health system for ecological frailty assessment in elderly. Proceedings 2019, 31, 41. [Google Scholar] [CrossRef]



| Work | Method | Channels | Intrusive | Own Dataset | Subjects | Classes | Session Size | Validation Split | Accuracy |
|---|---|---|---|---|---|---|---|---|---|
| [27] | CNN+LSTM | EEG: 64 | Yes | No | Cross-Subject: 108 | Five | 120 s | 75–25% | 98.3% |
| [28] | LSTM | EEG: 64 | Yes | No | Intra-Subject: 109 | Five | 120 s | 5 × 5-fold | 97.8% |
| [25] | CNN+LSTM | Muse: 4 | Low | Yes | Intra-Subject: 1 | Four | 30 s | 90–10% | 80.13% |
| [13] | CNN+LSTM | Muse: 4 | Low | Yes | Cross-Subject: 4 | Binary | 20 s | 90–10% | 98.9% |
| [26] | SVM | Muse: 4 | Low | Yes | Intra-Subject: 8 | Binary | 10 s | 4-fold | 95.1% |
| [32] | SVM | EEG: 2 | Yes | No | Intra-Subject: 2 | Binary | 9 s | 50–50% | 82.14% |
| [33] | CNN | EEG: 28 | Yes | Yes | Intra-Subject: 2 | Binary | 5 s | 80–20% | 86.41% |
| [36] | RLDA CNN | EEG: 22 EEG: 44 | Yes Yes | No Yes | Intra-Subject: 9 Intra-Subject: 20 | Four Four | 4 s 4 s | ICV | 73.7% 93.9% |
| [35] | LSTM | EEG: 6 | Yes | No | Intra-Subject: 9 | Binary | 4 s | 5 × 5-fold | 79.6% |
| [37] | CNN | EEG: 3 | Yes | No | Intra-Subject: 9 | Binary | 4 s | 60%–40% | 78.44% |
| [34] | CNN+SAE | EEG: 3 | Yes | No | Intra-Subject: 9 | Binary | 4 s | 10 × 10-fold | 77.6% |
| [39,40] | LR | EEG: 128 | Yes | Yes | Intra-Subject: 29 | Three | 3 s | 50%–50% | 90.5% |
| [38] | CNN | EEG: 22 | Yes | No | Intra-Subject: 9 Cross-Subject: 9 | Four | 2 s | 4-fold | ~70% ~40% |
| WS | TrAcc1 | VaAcc1 | TeAcc1 | TrAcc2 | VaAcc2 | TeAcc2 | TrAcc3 | VaAcc3 | TeAcc3 | TrAvg | VaAvg | TeAvg |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 2 s | 0.773846 | 0.760000 | 0.700000 | 0.795385 | 0.746667 | 0.705000 | 0.829231 | 0.786667 | 0.820000 | 0.799487 | 0.764444 | 0.741667 |
| 1.5 s | 0.747692 | 0.753333 | 0.720000 | 0.807692 | 0.806667 | 0.725000 | 0.847692 | 0.880000 | 0.850000 | 0.801025 | 0.813333 | 0.765000 |
| 1 s | 0.912308 | 0.840000 | 0.795000 | 0.829231 | 0.826667 | 0.740000 | 0.949231 | 0.920000 | 0.905000 | 0.896923 | 0.862222 | 0.813333 |
| 0.5 s | 0.920000 | 0.833333 | 0.805000 | 0.906154 | 0.860000 | 0.780000 | 0.940000 | 0.913334 | 0.930000 | 0.922051 | 0.868889 | 0.838333 |
| 0.25 s | 0.896923 | 0.873333 | 0.765000 | 0.832308 | 0.840000 | 0.745000 | 0.916923 | 0.873333 | 0.890000 | 0.882051 | 0.862222 | 0.800000 |
| 0.1 s | 0.841538 | 0.833333 | 0.760000 | 0.803077 | 0.773333 | 0.700000 | 0.840000 | 0.853333 | 0.855000 | 0.828205 | 0.820000 | 0.771667 |
| 0.05 s | 0.835384 | 0.866667 | 0.730000 | 0.692308 | 0.706667 | 0.720000 | 0.818462 | 0.853333 | 0.845000 | 0.782051 | 0.808889 | 0.765000 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Garcia-Moreno, F.M.; Bermudez-Edo, M.; Garrido, J.L.; Rodríguez-Fórtiz, M.J. Reducing Response Time in Motor Imagery Using A Headband and Deep Learning. Sensors 2020, 20, 6730. https://doi.org/10.3390/s20236730
Garcia-Moreno FM, Bermudez-Edo M, Garrido JL, Rodríguez-Fórtiz MJ. Reducing Response Time in Motor Imagery Using A Headband and Deep Learning. Sensors. 2020; 20(23):6730. https://doi.org/10.3390/s20236730
Chicago/Turabian StyleGarcia-Moreno, Francisco M., Maria Bermudez-Edo, José Luis Garrido, and María José Rodríguez-Fórtiz. 2020. "Reducing Response Time in Motor Imagery Using A Headband and Deep Learning" Sensors 20, no. 23: 6730. https://doi.org/10.3390/s20236730
APA StyleGarcia-Moreno, F. M., Bermudez-Edo, M., Garrido, J. L., & Rodríguez-Fórtiz, M. J. (2020). Reducing Response Time in Motor Imagery Using A Headband and Deep Learning. Sensors, 20(23), 6730. https://doi.org/10.3390/s20236730





