Continuous m-Health Data Authentication Using Wavelet Decomposition for Feature Extraction
Abstract
1. Introduction
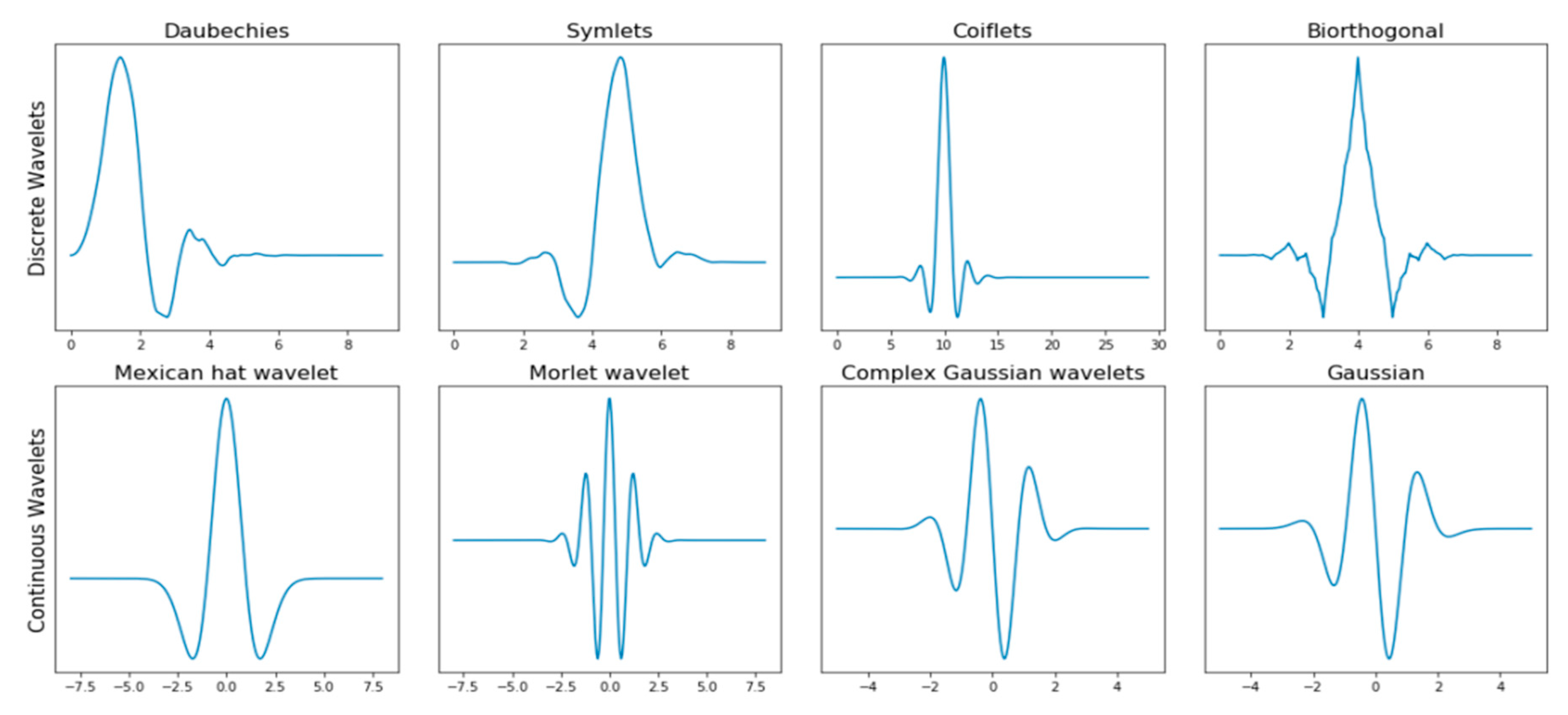
2. State of the Art
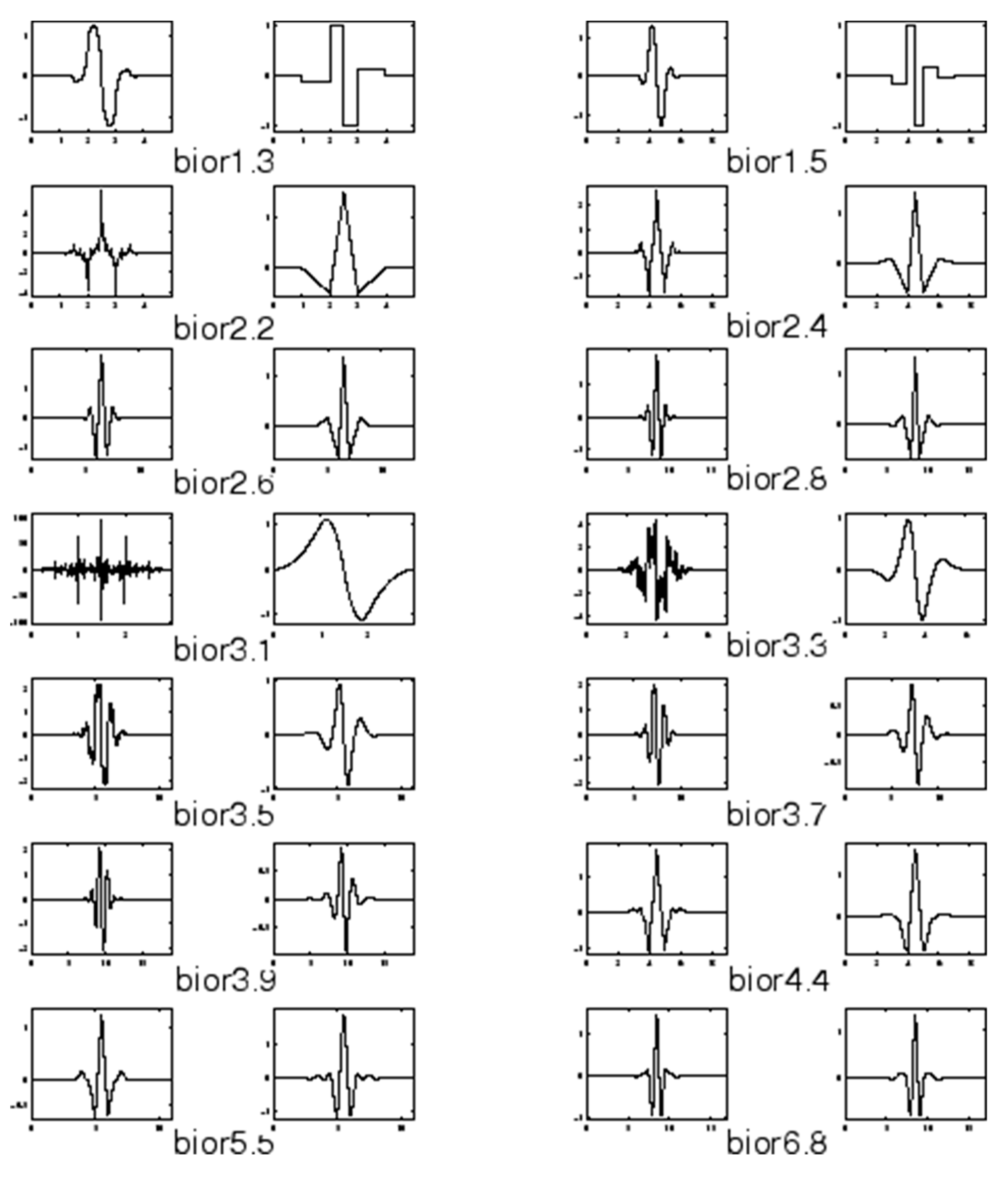
2.1. Biorthogonal Wavelets
2.2. Heart Rate Variability
3. Methodology
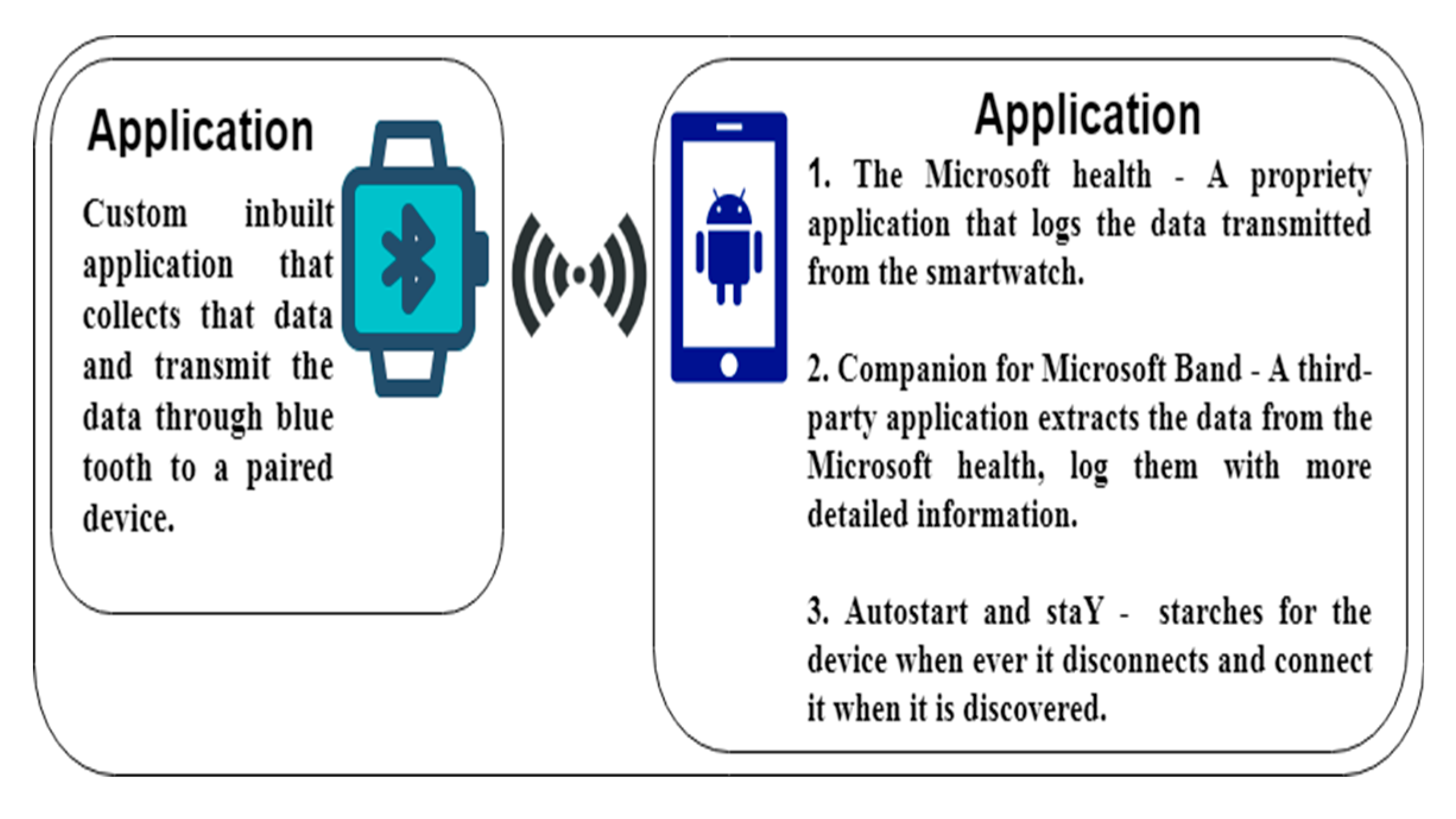
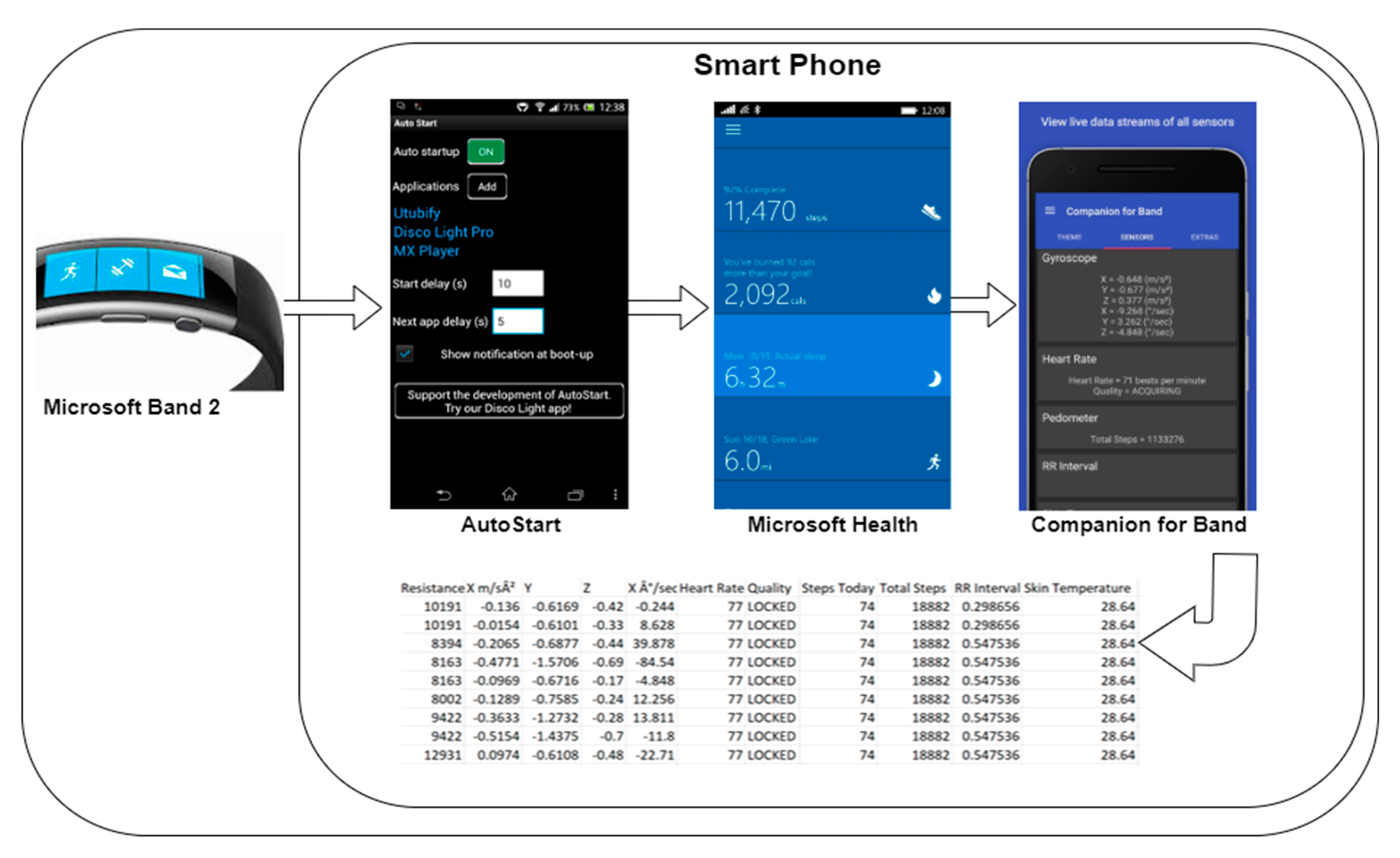
3.1. Transparent Data Extraction
3.2. Data Segmentation for Continuous Authentication
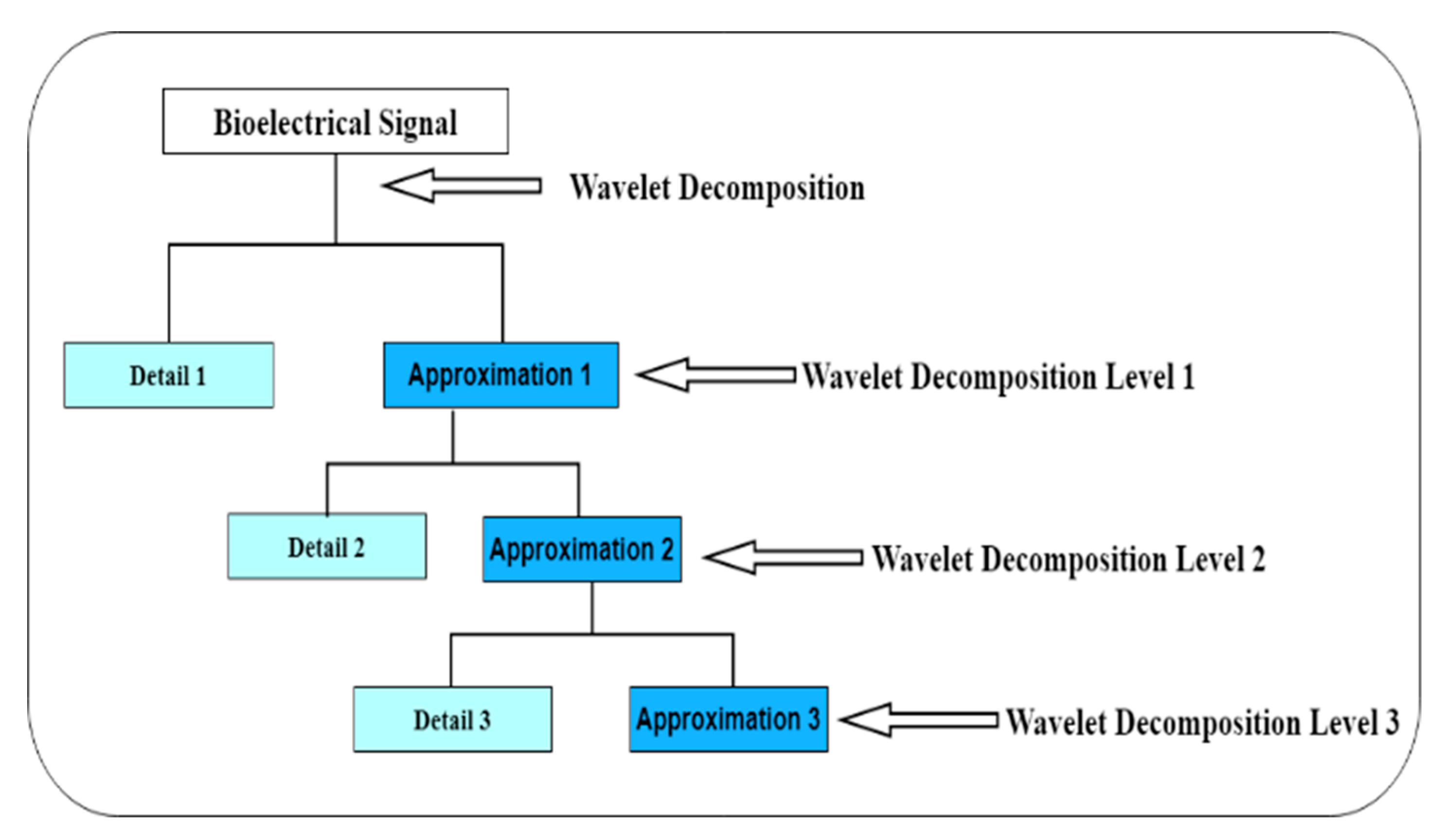
3.3. Biorthogonal Wavelet Decomposition
- Low-pass decomposition filter = Lo_D
- High-pass decomposition filter = Hi_D
- Low-pass reconstruction filter = Lo_R
- High-pass reconstruction filter = Hi_R
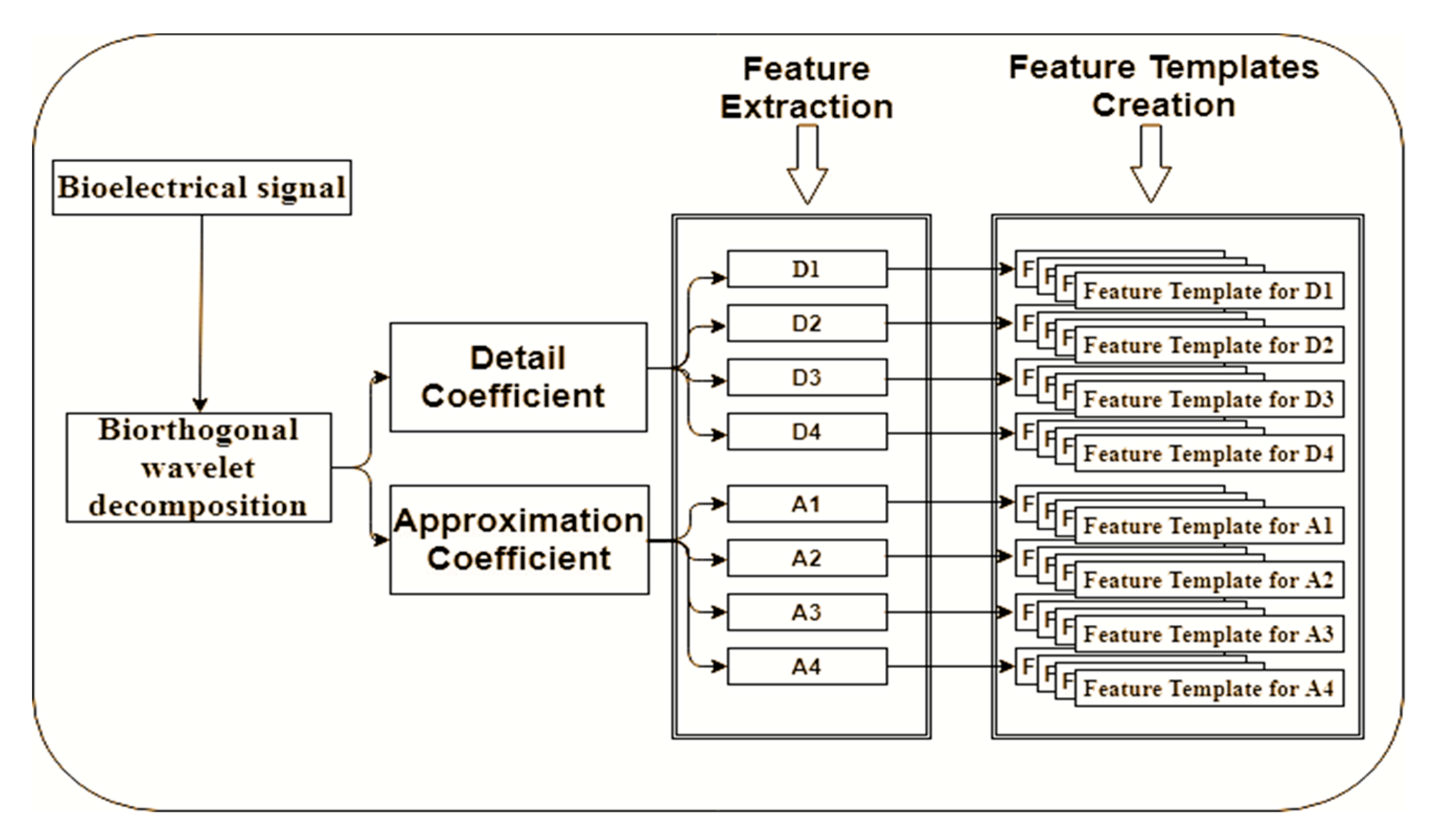
3.4. Feature Extraction
- Variance: This is the sum of square distance of the bioelectrical signal.
- Mean of the energy: The signal mean of the energy is the energy average value of the bioelectrical signal.
- Minimum energy: This is the lowest energy value of the bioelectrical signal
- Maximum energy: This is the highest energy value of the bioelectrical signal.
- Mean: These are the values diversity of the data around the median.
- Minimum amplitude: This is the lowest point from the equilibrium point of the bioelectrical signal.
- Standard deviation (STD): This is the square root of the variance of a random variation.
- Maximum amplitude: This is the highest point from the equilibrium point of the bioelectrical signal.
- Range: This is the difference between the highest signal value and the lowest signal value.
- Peak2peak: This is the difference between the maximum and minimum values of the bioelectrical signal.
- Root mean square (RMS): The RMS is the measurement of the magnitude of a set values within the signal.
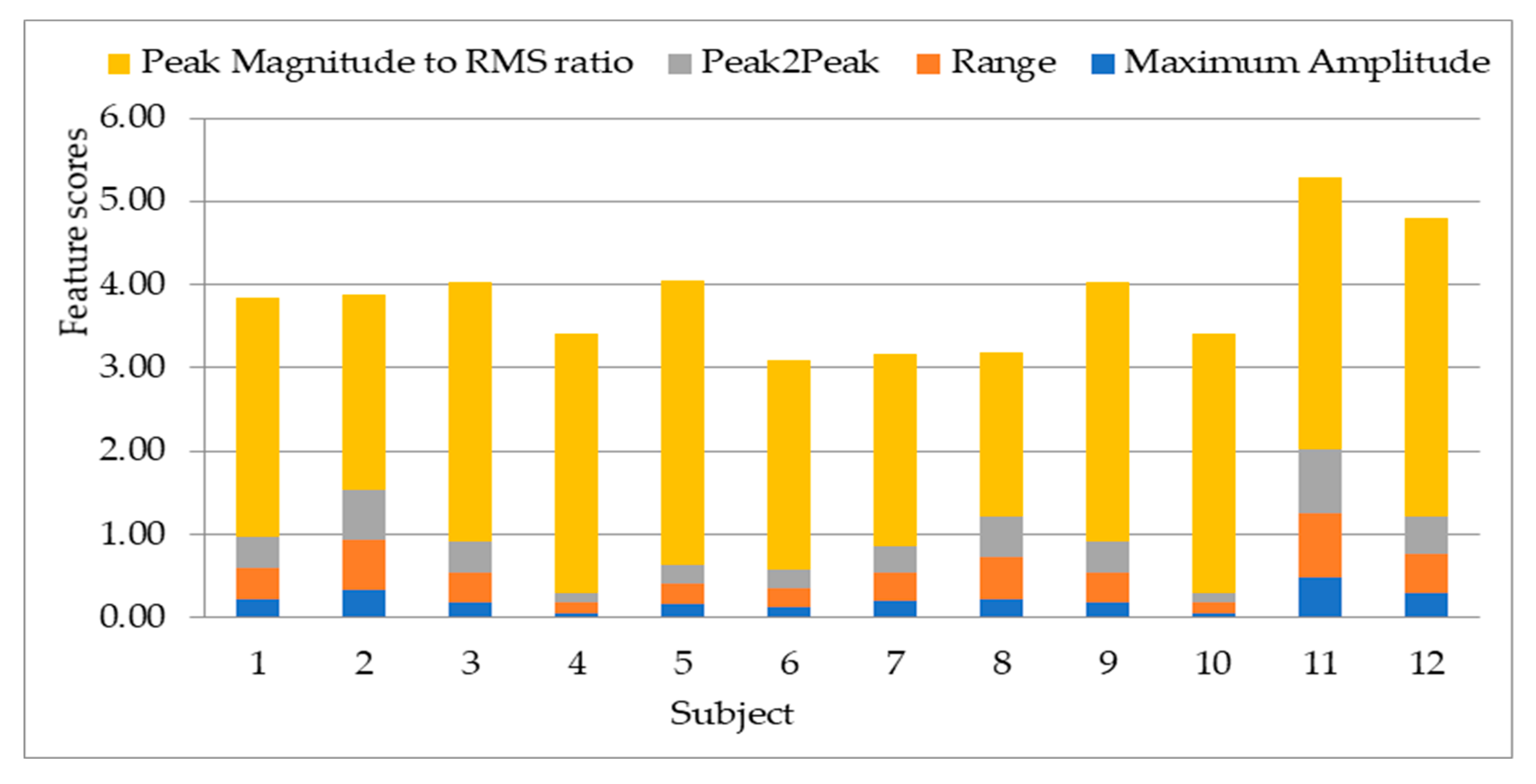
- Peak magnitude to RMS ratio: This is the ratio of the largest absolute value of a signal to the root mean square (RMS) value of that signal.
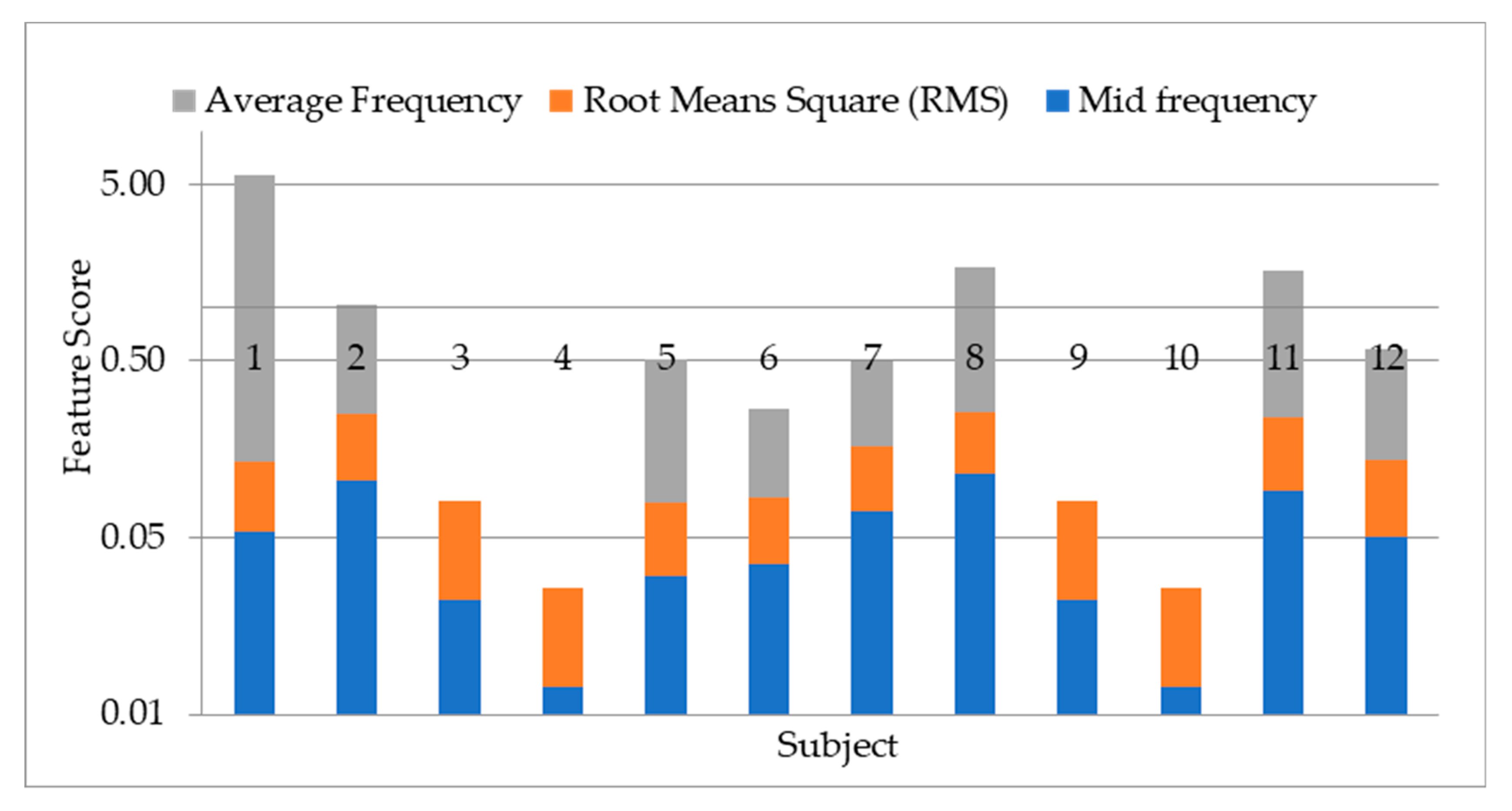
- Average frequency: this the arithmetic mean of the signal frequency.
3.5. Classification
3.6. Experimentation
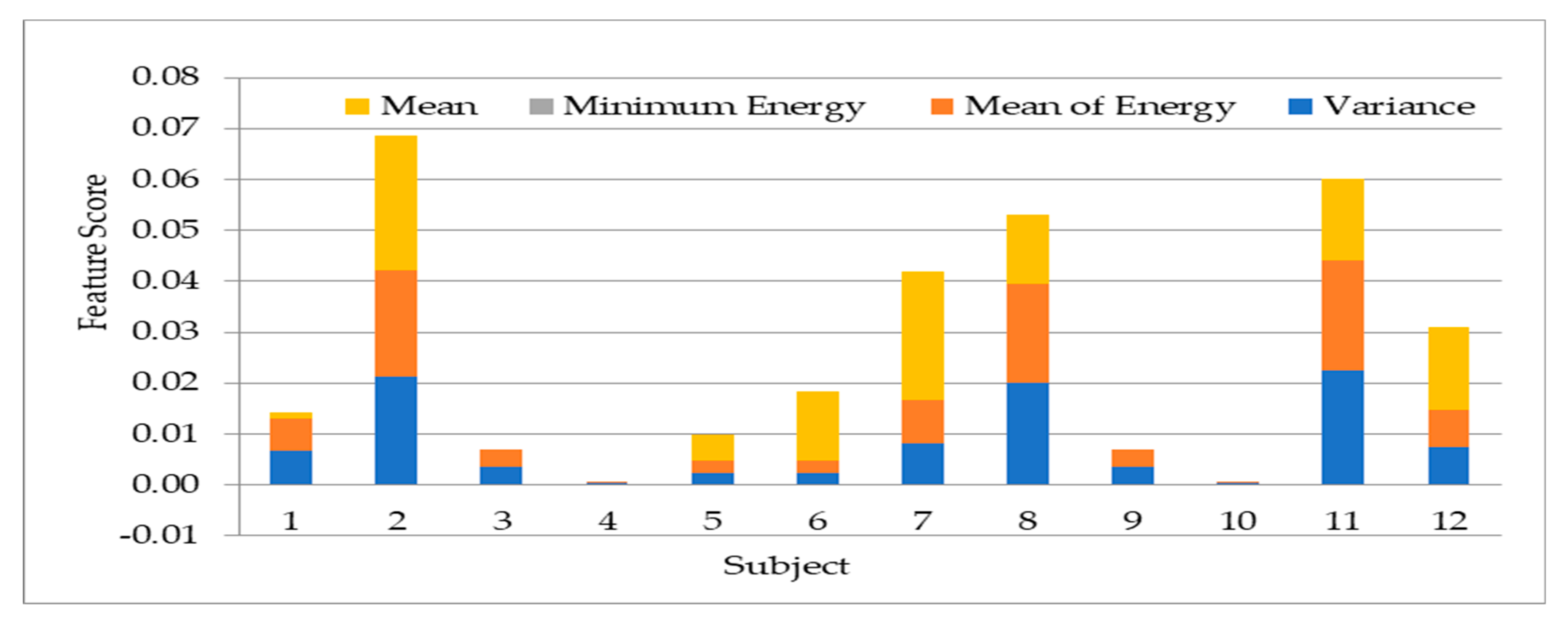
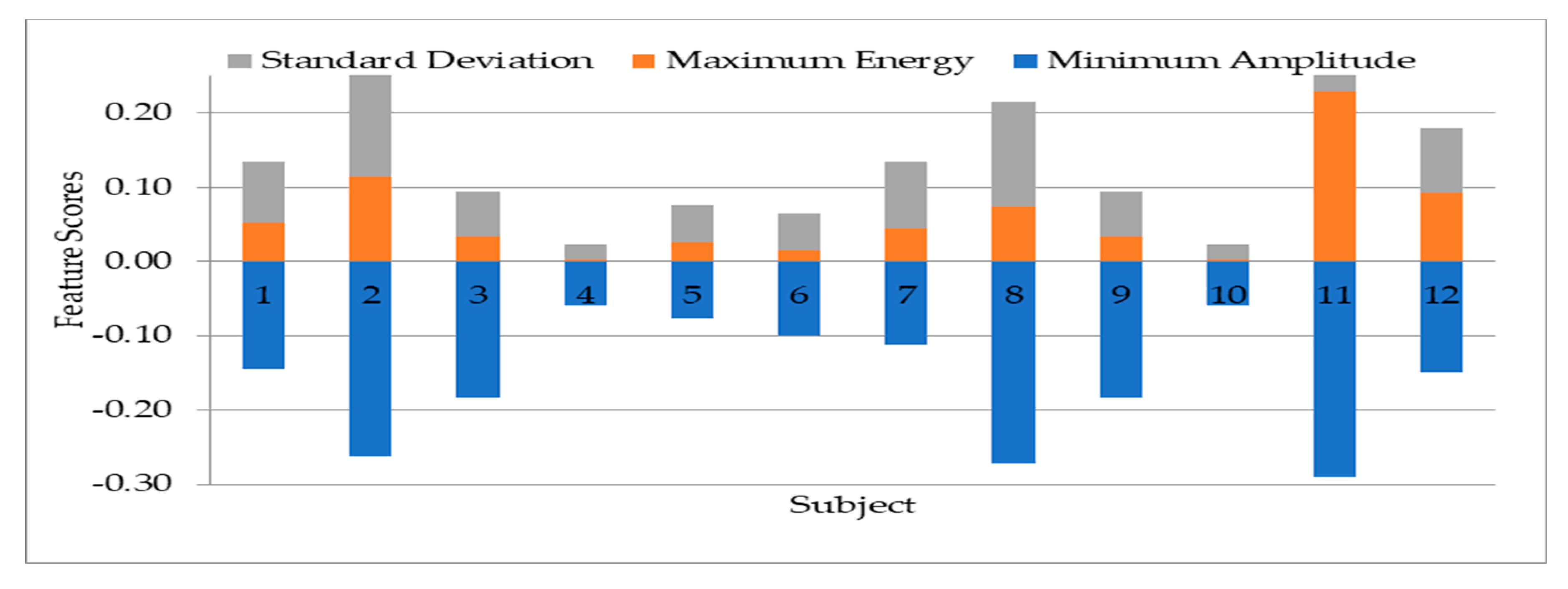
- Experiment 1: the experimentation to select the most suitable features for use in the other experiments. The experiment used 12 subjects’ data for the 13 statistical features extraction experimentation.
- Experiment 2: the second experimentation is the classification of the sub-bands after extracting features. The 30 subjects’ data are used with 12 selected features for each sub-band for both the approximation of coefficient and detail coefficient for fifteen biorthogonal wavelet family.
- Experiment 3: this experiment is done on the fusion of the sub-bands. This experiment also used 30 subjects’ data with the 12 selected features for fusion of all the sub-band for both the approximation of coefficient and detail coefficient classification.
- Used case experiment: this is the used case data authentication using 30 subjects’ data.
4. Results
4.1. Result on Experiment 1: Selection of Statistical Features
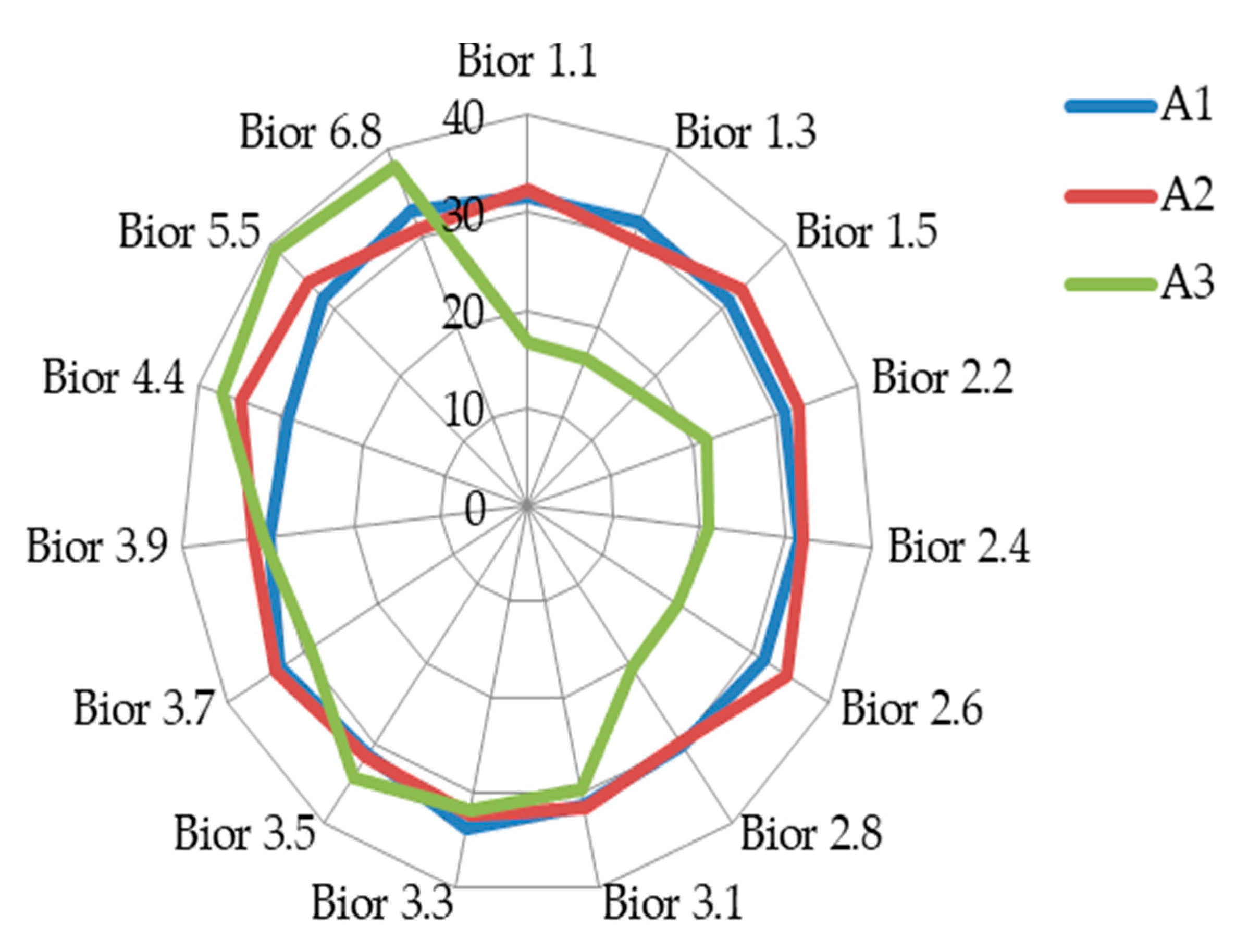
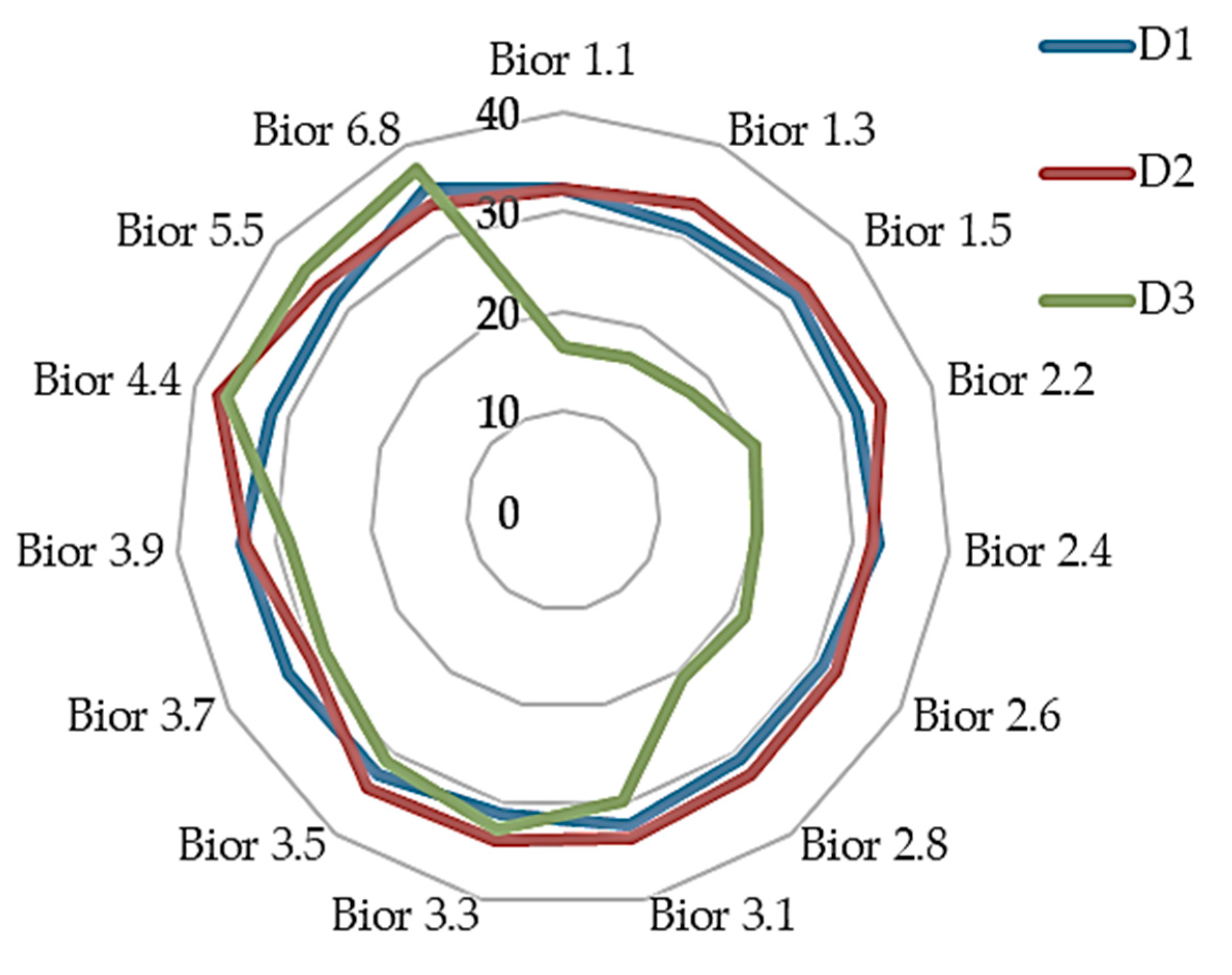
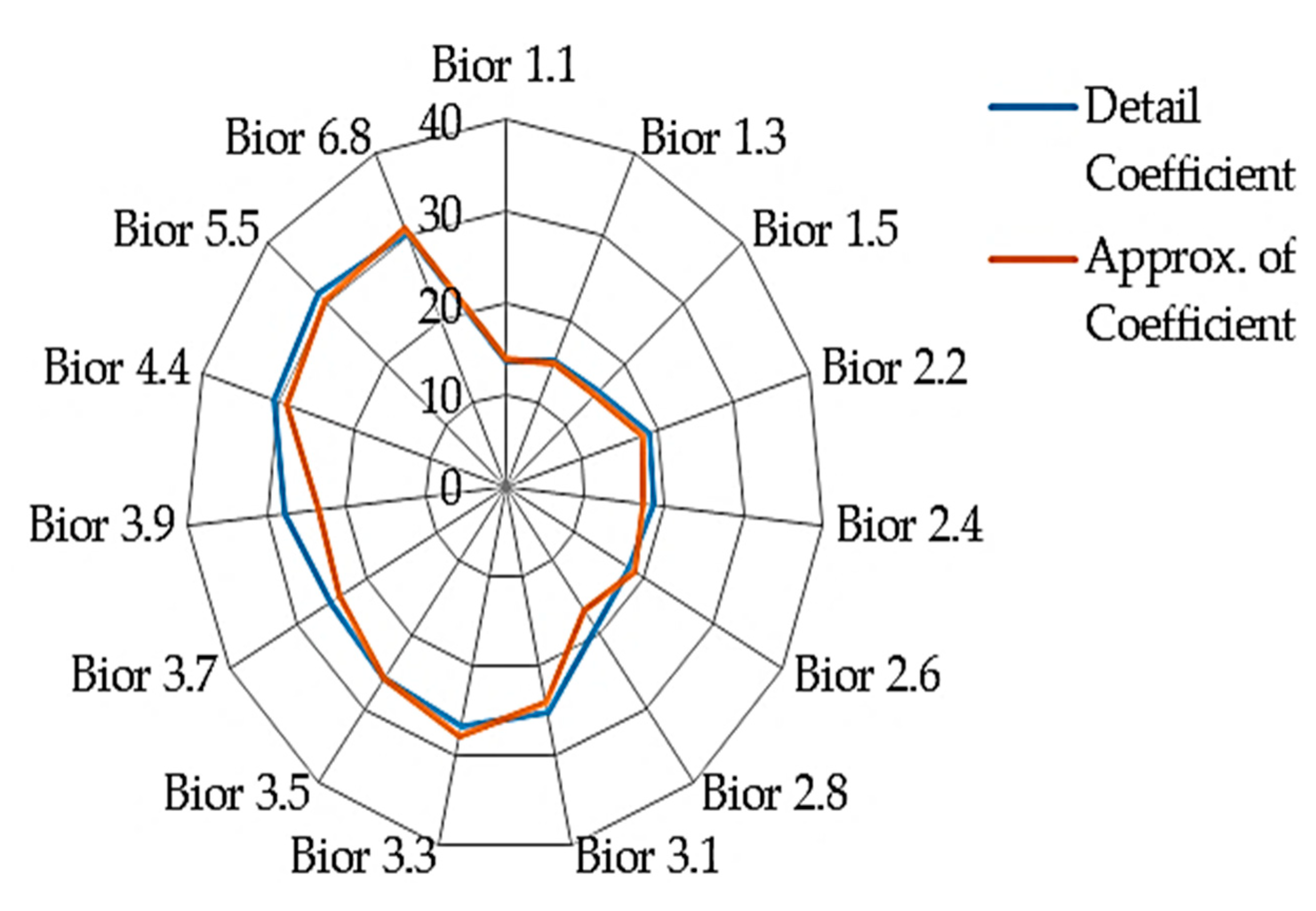
4.2. Result on Experiment 2: Classification Sub-Band Features
4.3. Result on Experiment 3: Fusion of Sub-Band Features
5. Use Case and Evaluation
5.1. Use Case Architecture
5.1.1. The Smartwatch
5.1.2. The Smartphone
5.1.3. The Smartphone
5.1.4. Healthcare Data Management Facility
5.2. Data Authentication
5.3. Use Case Evaluation
5.4. Discussion
6. Conclusions and Future Work
Author Contributions
Funding
Conflicts of Interest
References
- Available online: https://www.statista.com/statistics/330695/number-of-smartphone-users-worldwide/ (accessed on 22 April 2020).
- Khodadadi, T.; Islam, A.M.; Baharun, S.; Komaki, S. Evaluation of recognition-based graphical password schemes in terms of usability and security attributes. Int. J. Electr. Comput. Eng. 2016, 6, 2939. [Google Scholar]
- Saevanee, H.; Clarke, N.; Furnell, S. SMS linguistic profiling authentication on mobile device. In Proceedings of the 2011 5th IEEE International Conference on Network and System Security, Institute of Electrical and Electronics Engineers, Milan, Italy, 6–8 September 2011; pp. 224–228. [Google Scholar]
- Enamamu, T.S.; Clarke, N.; Dowland, P.S.; Li, F. Smartwatch based body-temperature authentication. In Proceedings of the IEEE International Conference on Computing Networking and Informatics (ICCNI), Institute of Electrical and Electronics Engineers, Lagos, Nigeria, 29–31 October 2017; pp. 1–7. [Google Scholar]
- Peng, Z.; Chu, F. Application of the wavelet transform in machine condition monitoring and fault diagnostics: A review with bibliography. Mech. Syst. Signal Process. 2004, 18, 199–221. [Google Scholar] [CrossRef]
- Enamamu, T.S.; Clarke, N.; Dowland, P.S.; Li, F. Transparent authentication: Utilising heart rate for user authentication. In Proceedings of the 12th IEEE International Conference for Internet Technology and Secured Transactions (ICITST), Institute of Electrical and Electronics Engineers, Cambridge, UK, 11–14 December 2017. [Google Scholar]
- Available online: https://www.who.int/goe/publications/goe_mhealth_web.pdf (accessed on 18 December 2019).
- AlMotiri, S.H.; Khan, M.A.; Alghamdi, M.A. Mobile Health (m-Health) System in the Context of IoT. In Proceedings of the IEEE 4th International Conference on Future Internet of Things and Cloud Workshops (FiCloudW), Vienna, Austria, 22–24 August 2016; pp. 39–42. [Google Scholar]
- Mallat, S.G.; Heil, C.; Walnut, D.F. A Theory for Multiresolution Signal Decomposition: The Wavelet Representation. IEEE Trans. Pattern Anal. Mach. Intell. 2009, 11, 674–693. [Google Scholar] [CrossRef]
- Addison, P.S.; Walker, J.; Guido, R.C. Time–frequency analysis of biosignals. IEEE Eng. Med. Biol. Mag. 2009, 28, 14–29. [Google Scholar] [CrossRef]
- Subasi, A.; Ercelebi, E. Classification of EEG signals using neural network and logistic regression. Comput. Methods Programs Biomed. 2005, 78, 87–99. [Google Scholar] [CrossRef]
- Subasi, A. EEG signal classification using wavelet feature extraction and a mixture of expert model. Expert Syst. Appl. 2007, 32, 1084–1093. [Google Scholar] [CrossRef]
- Jahankhani, P.; Kodogiannis, V.; Revett, K. EEG Signal Classification Using Wavelet Feature Extraction and Neural Networks. In Proceedings of the IEEE John Vincent Atanasoff 2006 International Symposium on Modern Computing (JVA’06), Sofia, Bulgaria, 3–6 October 2006; pp. 120–124. [Google Scholar]
- Gokhale, M.Y.; Khanduja, D.K. Time Domain Signal Analysis Using Wavelet Packet Decomposition Approach. Int. J. Commun. Netw. Syst. Sci. 2010, 3, 321–329. [Google Scholar] [CrossRef]
- Prabhakar, S.; Mohanty, A.; Sekhar, A. Application of discrete wavelet transform for detection of ball bearing race faults. Tribol. Int. 2002, 35, 793–800. [Google Scholar] [CrossRef]
- Ovanesova, A.; Suarez, L. Applications of wavelet transforms to damage detection in frame structures. Eng. Struct. 2004, 26, 39–49. [Google Scholar] [CrossRef]
- Tsou, C.; Hsieh, C.-H.; Liang, M.-C.; Huang, P.-W.; Lee, S.-Y. ECG acquisition system with heart rate detection and energy harvesting for drivers. In Proceedings of the IEEE International Symposium on Bioelectronics and Bioinformatics (ISBB), Beijing, China, 14–17 October 2015; pp. 31–34. [Google Scholar]
- Laine, A.; Fan, J. Texture classification by wavelet packet signatures. IEEE Trans. Pattern Anal. Mach. Intell. 1993, 15, 1186–1191. [Google Scholar] [CrossRef]
- Büssow, R. An algorithm for the continuous Morlet wavelet transform. Mech. Syst. Signal Process. 2007, 21, 2970–2979. [Google Scholar] [CrossRef]
- Li, L.; Liu, P.; Xing, Y.; Guo, H. Time-frequency analysis of acoustic signals from a high-lift configuration with two wavelet functions. Appl. Acoust. 2018, 129, 155–160. [Google Scholar] [CrossRef]
- Engelbrektsson, J.; Abrahamsson, K.; Breitholtz, J.; Nicholas, M.; Svensson, O.; Wikström, H.; Josefson, M. The impact of Mexican hat and dual-tree complex wavelet transforms on multivariate evaluation of image texture properties. J. Chemom. 2010, 24, 454–463. [Google Scholar] [CrossRef]
- Semwogerere, D. The Use of the Two-Dimensional Continuous Wavelet Transform for Classification of Targets in Flir Imagery. Master’s Thesis, Clark Atlanta University, Atlanta, GA, USA, 1998. [Google Scholar]
- Sharif, I.; Khare, S. Comparative Analysis of Haar and Daubechies Wavelet for Hyper Spectral Image Classification. ISPRS-Int. Arch. Photogramm. Remote. Sens. Spat. Inf. Sci. 2014, 40, 937–941. [Google Scholar] [CrossRef]
- Mahmoodabadi, S.Z.; Ahmadian, A.; Abolhasani, M.D. ECG feature extraction using Daubechies wavelets. In Proceedings of the 5th IASTED International conference on Visualization, Imaging and Image Processing, Benidorm, Spain, 7–9 September 2005. [Google Scholar]
- Clonda, D.; Lina, J.-M.; Goulard, B. Complex Daubechies wavelets: Properties and statistical image modelling. Signal Process. 2004, 84, 1–23. [Google Scholar] [CrossRef]
- Misiti, Y.; Misiti, M.; Oppenheim, G.; Poggi, J.-M. Wavelet Toolbox; The MathWorks Inc.: Natick, MA, USA, 1996; p. 21. [Google Scholar]
- Černá, D.; Finek, V.; Najzar, K. On the exact values of coefficients of coiflets. Cent. Eur. J. Math. 2008, 6, 159–169. [Google Scholar] [CrossRef]
- Stolojescu, C.; Railean, I.; Moga, S.; Isar, A. Comparison of wavelet families with application to WiMAX traffic forecasting. In Proceedings of the 12th IEEE International Conference on Optimization of Electrical and Electronic Equipment, Brasov, Romania, 20–22 May 2010; pp. 932–937. [Google Scholar]
- Sindhura, S.K.; Reddy, S.N.; Kamaraj, P. Comparison of SNR Improvement for Lower Atmospheric Signals Using Wavelets. Int. J. Adv. Res. Electr. Electron. Instrum. Eng. 2015, 4, 8202–8209. [Google Scholar]
- Abhyankar, A.; Schuckers, S. Novel Biorthogonal Wavelet based Iris Recognition for Robust Biometric System. Int. J. Comput. Theory Eng. 2010, 2, 233–237. [Google Scholar] [CrossRef]
- Hariprasath, S.; Venkatasubramaniam, S. Iris pattern recognition using biorthogonal Wavelet Packet Analysis. In Proceedings of the IEEE International Conference on Communication Control and Computing Technologies, Institute of Electrical and Electronics Engineers, Ramanathapuram, India, 7–9 October 2010; pp. 830–834. [Google Scholar]
- Szewczyk, R.; Grabowski, K.; Napieralska, M.; Sankowski, W.; Zubert, M.; Napieralski, A. A reliable iris recognition algorithm based on reverse biorthogonal wavelet transform. Pattern Recognit. Lett. 2012, 33, 1019–1026. [Google Scholar] [CrossRef]
- Prashar, D.; Mnupreet, K. Human Eye Iris Recognition Using Discrete 2d Reverse Biorthogonal Wavelet 6.8. Int. J. Sci. Technol. Res. 2014, 3, 266–270. [Google Scholar]
- Isnanto, R.R. Iris recognition analysis using biorthogonal wavelets tranform for feature extraction. In Proceedings of the 1st IEEE International Conference on Information Technology, Computer, and Electrical Engineering, Semarang, Indonesia, 7–8 November 2014; pp. 183–187. [Google Scholar]
- Kapogiannopoulos, G.S.; Papadakis, M. Character recognition using a biorthogonal discrete wavelet transform. SPIE’s Int. Symp. Optical Sci. Eng. Instrum. 1996, 2825, 384–394. [Google Scholar]
- Terwiesch, P.; Mercorelli, P. A local feature extraction using biorthogonal bases for classification of embedded classes of signals. In Proceedings of the 14th International Symposium on Mathematical Theory of Networks and Systems, Perpignan, France, 19–23 June 2000. [Google Scholar]
- Hema, C.; Paulraj, M.; Kaur, H. Brain signatures: A modality for biometric authentication. In Proceedings of the IEEE International Conference on Electronic Design, Penang, Malaysia, 1–3 December 2008; pp. 1–4. [Google Scholar]
- Kousarrizi, M.R.N.; Ghanbari, A.A.; Teshnehlab, M.; Shoorehdeli, M.A.; Gharaviri, A. Feature Extraction and Classification of EEG Signals Using Wavelet Transform, SVM and Artificial Neural Networks for Brain Computer Interfaces. In Proceedings of the IEEE International Joint Conference on Bioinformatics, Systems Biology and Intelligent Computing, Shanghai, China, 3–5 August 2009; pp. 352–355. [Google Scholar]
- Guennoun, M.; Abbad, N.; Talom, J.; Rahman, S.M.M.; El-Khatib, K. Continuous authentication by electrocardiogram data. In Proceedings of the IEEE Toronto International Conference Science and Technology for Humanity (TIC-STH), Toronto, ON, Canada, 26–27 September 2009; pp. 40–42. [Google Scholar] [CrossRef]
- Pirbhulal, S.; Zhang, H.; Mukhopadhyay, S.C.; Li, C.; Wang, Y.; Li, G.; Wu, W.; Zhang, Y.-T. An Efficient Biometric-Based Algorithm Using Heart Rate Variability for Securing Body Sensor Networks. Sensors 2015, 15, 15067–15089. [Google Scholar] [CrossRef] [PubMed]
- Tawfik, M.M.; Kamal, H.S.T. Human Identification Using QT Signal and QRS Complex of the ECG. OJEEE 2011, 3, 383–387. [Google Scholar]
- Pourbabaee, B.; Howe-Patterson, M.; Reiher, E.; Benard, F. Deep Convolutional Neural Network for ECG-Based Human Identification; CMBES: Ottawa, ON, Canada, 2018; Volume 41. [Google Scholar]
- Labati, R.D.; Munoz, E.; Piuri, V.; Sassi, R.; Scotti, F. Deep-ECG: Convolutional Neural Networks for ECG biometric recognition. Pattern Recognit. Lett. 2019, 126, 78–85. [Google Scholar] [CrossRef]
- Enamamu; Timibloudi, S. Bioelectrical User Authentication. Ph.D. Thesis, University of Plymouth, Plymouth, UK, 2019. [Google Scholar]
- Tay, D.B.H.; Marusic, S.; Deng, G.; Palaniswami, M. Wavelet theoretic properties of the family of 6/10 biorthogonal filters. IEEE Int. Symp. Commun. Inf. Technol. 2004, 2, 1051–1054. [Google Scholar]
- Reaz, M.B.I.; Hussain, M.S.; Mohd-Yasin, F.; Raez, M. Techniques of EMG signal analysis: Detection, processing, classification and applications (Correction). Biol. Proced. Online 2006, 8, 163. [Google Scholar] [CrossRef] [PubMed]
- Faris, H.; Aljarah, I.; Mirjalili, S. Training feedforward neural networks using multi-verse optimizer for binary classification problems. Appl. Intell. 2016, 45, 322–332. [Google Scholar] [CrossRef]
- Guez, A.; Selinsky, J. Neurocontroller design via supervised and unsupervised learning. J. Intell. Robot. Syst. 1989, 2, 307–335. [Google Scholar] [CrossRef]
- Fernández-Alemán, J.L.; Señor, I.C.; Lozoya, P.; Ángel, O.; Toval, A. Security and privacy in electronic health records: A systematic literature review. J. Biomed. Inform. 2013, 46, 541–562. [Google Scholar] [CrossRef]




| Heart Rate Variability Segmentation Using 3 Sec. | |||
|---|---|---|---|
| Types | Exp. 1 | Exp. 2–3 | Used Case Exp |
| Number of subjects used | 12 | 30 | 30 |
| Sampling rate | 8 | 8 | 8 |
| Data points per segment | 24 | 24 | 24 |
| Num. of Feature Segments of each subject | 150 | 300 | 600 |
| Number recorded time per subject in Seconds | 3600 | 7200 | 14,400 |
| Total number recorded time for all subjects in Seconds | 108,000 | 216,000 | 432,000 |
| No | Wavelet Family | EER of Approximation of Coefficient and Detail Coefficient Classification Comparison in EER (%) | |||||
|---|---|---|---|---|---|---|---|
| D1 | A1 | D2 | A2 | D3 | A3 | ||
| 1 | Bior1.1 | 32.27 | 31.54 | 32.21 | 32.31 | 16.41 | 16.66 |
| 2 | Bior1.3 | 31.02 | 31.77 | 33.51 | 29.80 | 16.79 | 16.56 |
| 3 | Bior1.5 | 32.33 | 31.42 | 33.32 | 33.21 | 17.56 | 17.20 |
| 4 | Bior2.2 | 31.79 | 31.12 | 34.50 | 32.99 | 20.69 | 21.69 |
| 5 | Bior2.4 | 32.59 | 31.55 | 32.06 | 31.87 | 20.02 | 20.95 |
| 6 | Bior2.6 | 30.87 | 31.44 | 32.32 | 34.55 | 21.47 | 19.94 |
| 7 | Bior2.8 | 30.83 | 30.39 | 32.66 | 29.81 | 20.79 | 20.44 |
| 8 | Bior3.1 | 32.16 | 31.40 | 33.67 | 31.58 | 29.82 | 29.70 |
| 9 | Bior3.3 | 31.10 | 33.64 | 33.74 | 32.18 | 32.82 | 31.90 |
| 10 | Bior3.5 | 32.65 | 31.40 | 34.44 | 31.63 | 31.11 | 34.32 |
| 11 | Bior3.7 | 32.73 | 32.90 | 30.08 | 33.67 | 28.52 | 28.95 |
| 12 | Bior3.9 | 33.41 | 29.99 | 33.10 | 31.81 | 28.50 | 30.67 |
| 13 | Bior4.4 | 31.85 | 29.03 | 37.57 | 34.86 | 36.54 | 37.16 |
| 14 | Bior5.5 | 31.61 | 31.87 | 33.67 | 34.05 | 35.93 | 39.01 |
| 15 | Bior6.8 | 35.22 | 33.07 | 33.59 | 30.94 | 37.40 | 37.90 |
| Approximation and Detail Coefficient Classification Comparison in EER (%) | ||
|---|---|---|
| Feature | Bior1.1 | Bior1.3 |
| Approximation of Coefficient Sub-band 3 | 16.66 | 16.56 |
| Detail Coefficient Sub-band 3 | 16.41 | 16.56 |
| Approximation of Coefficient Fusion | 14.00 | 14.83 |
| Detail Coefficient Fusion | 13.80 | 14.89 |
| Sub. | AF | DF | Sub. | AF | DF |
|---|---|---|---|---|---|
| 1 | 0.00 | 0.00 | 16 | 22.84 | 20.76 |
| 2 | 19.04 | 17.46 | 17 | 13.22 | 15.73 |
| 3 | 12.07 | 9.70 | 18 | 14.66 | 10.13 |
| 4 | 19.83 | 20.04 | 19 | 7.11 | 4.74 |
| 5 | 18.89 | 13.00 | 20 | 16.38 | 9.55 |
| 6 | 4.60 | 4.17 | 21 | 13.58 | 11.35 |
| 7 | 13.36 | 14.08 | 22 | 10.20 | 12.72 |
| 8 | 24.21 | 19.11 | 23 | 2.08 | 2.08 |
| 9 | 14.80 | 14.08 | 24 | 19.97 | 9.27 |
| 10 | 18.10 | 17.03 | 25 | 10.13 | 9.48 |
| 11 | 10.06 | 4.45 | 26 | 0.00 | 0.00 |
| 12 | 15.52 | 22.49 | 27 | 8.91 | 8.76 |
| 13 | 12.72 | 13.99 | 28 | 27.23 | 28.81 |
| 14 | 22.56 | 24.71 | 29 | 15.54 | 14.83 |
| 15 | 20.55 | 17.10 | 30 | 3.46 | 3.05 |
| EER Features Fusion Result | 13.17 | 12.42 | |||
| Data Authentication Performance | ||||||
|---|---|---|---|---|---|---|
| Threshold (EER) | Patient | Non-Patient | ||||
| Acceptance | Rejection | Success Rate | Acceptance | Rejection | Success Rate | |
| 9% | 54 | 46 | 54% | 0 | 2000 | 0% |
| 10% | 56 | 44 | 56% | 0 | 2000 | 0% |
| 11% | 81 | 19 | 81% | 6 | 1994 | 0.3% |
| 12% | 84 | 16 | 84% | 8 | 1992 | 0.4% |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Enamamu, T.; Otebolaku, A.; Marchang, J.; Dany, J. Continuous m-Health Data Authentication Using Wavelet Decomposition for Feature Extraction. Sensors 2020, 20, 5690. https://doi.org/10.3390/s20195690
Enamamu T, Otebolaku A, Marchang J, Dany J. Continuous m-Health Data Authentication Using Wavelet Decomposition for Feature Extraction. Sensors. 2020; 20(19):5690. https://doi.org/10.3390/s20195690
Chicago/Turabian StyleEnamamu, Timibloudi, Abayomi Otebolaku, Jims Marchang, and Joy Dany. 2020. "Continuous m-Health Data Authentication Using Wavelet Decomposition for Feature Extraction" Sensors 20, no. 19: 5690. https://doi.org/10.3390/s20195690
APA StyleEnamamu, T., Otebolaku, A., Marchang, J., & Dany, J. (2020). Continuous m-Health Data Authentication Using Wavelet Decomposition for Feature Extraction. Sensors, 20(19), 5690. https://doi.org/10.3390/s20195690






