First Record of Juncaceicola as Endophytic Fungi Associated with Deschampsia antarctica Desv.
Abstract
1. Introduction
2. Materials and Methods
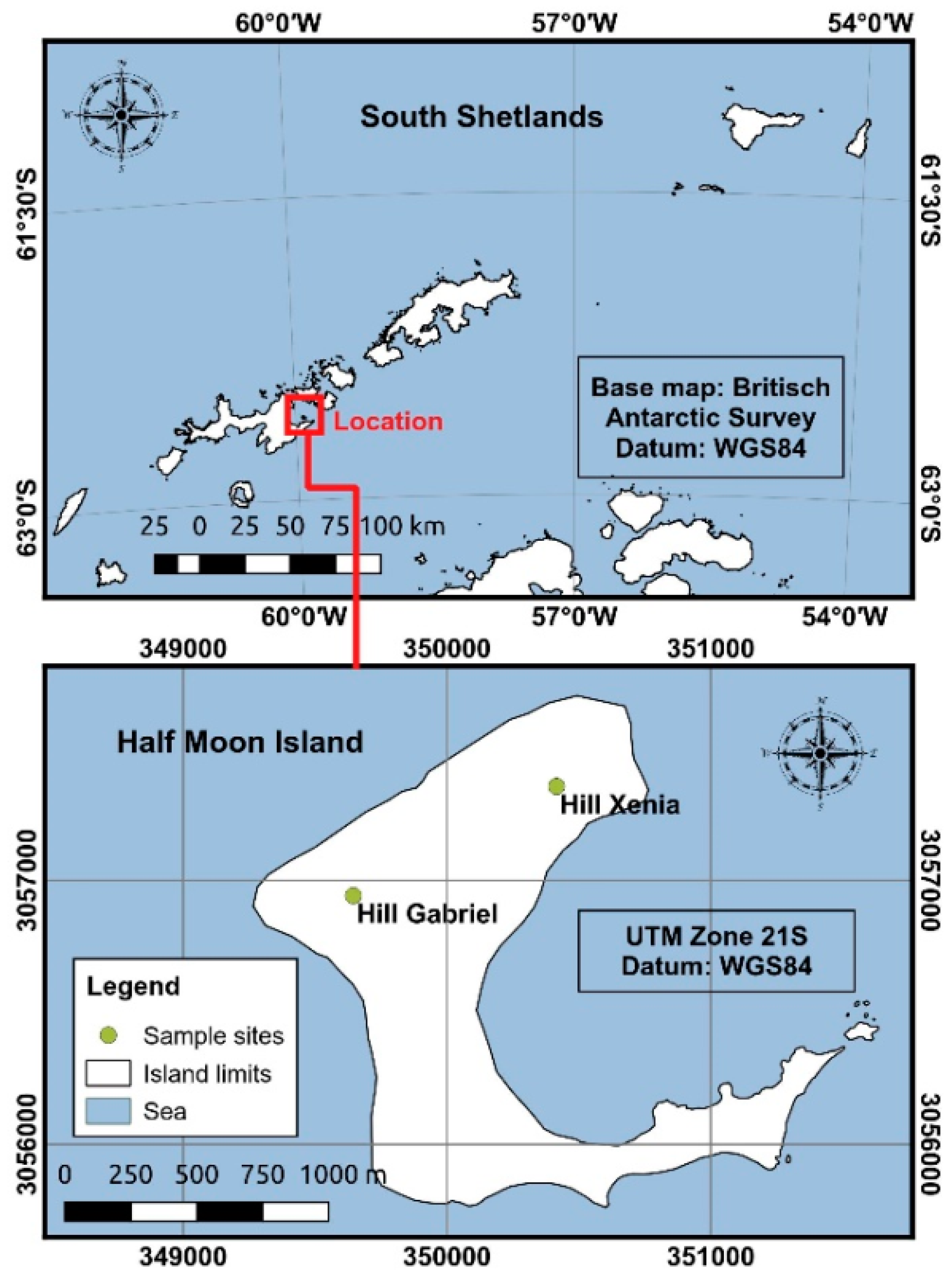
2.1. Study Area and Sample Collection
2.2. Isolation of the Endophytic Fungus
2.3. DNA Extraction, Amplification and Purification
2.4. Phylogenetic Analysis
2.5.Effects of Temperature on Mycelia Growth
2.6. Morphological Description
3. Results
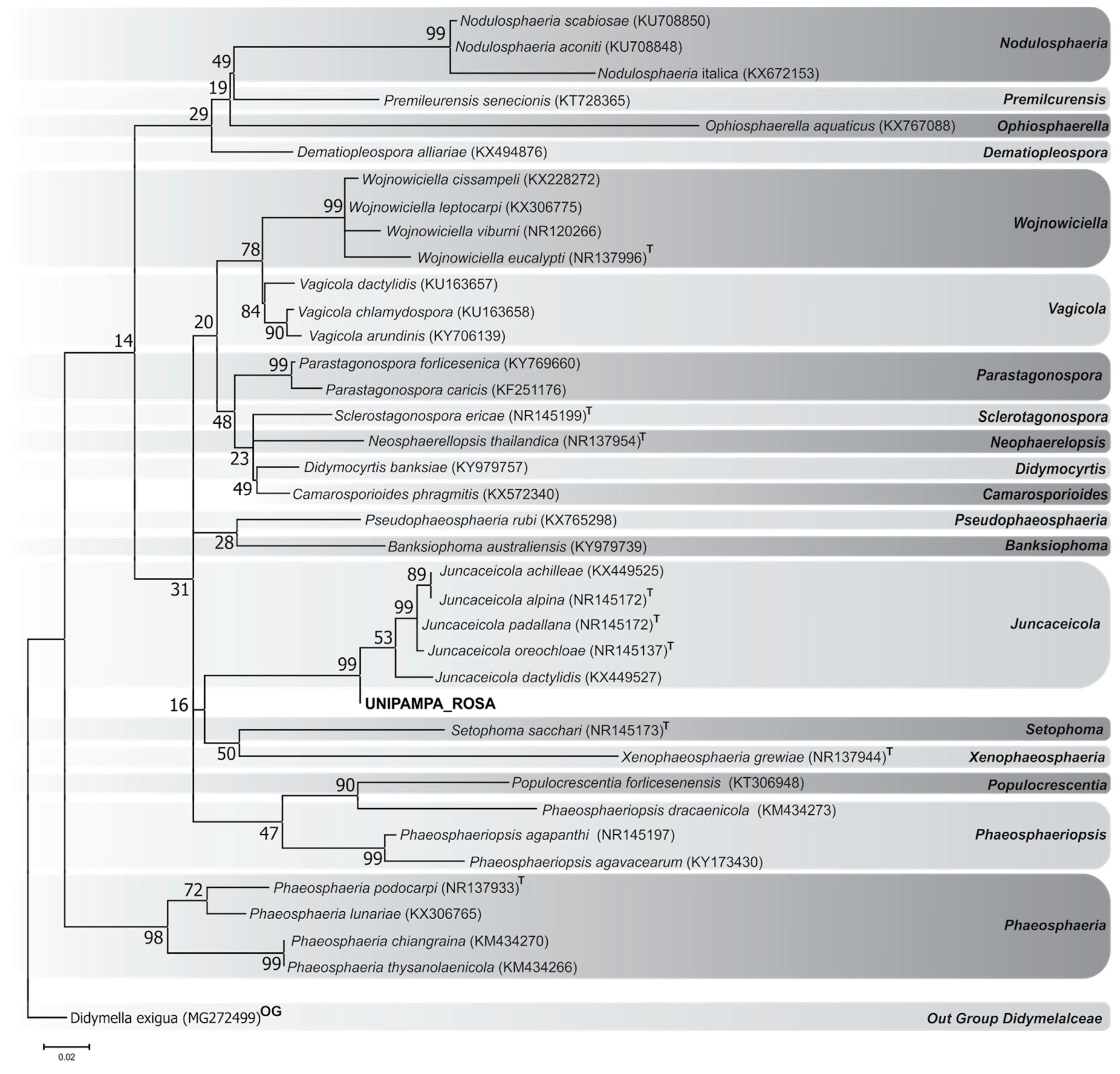
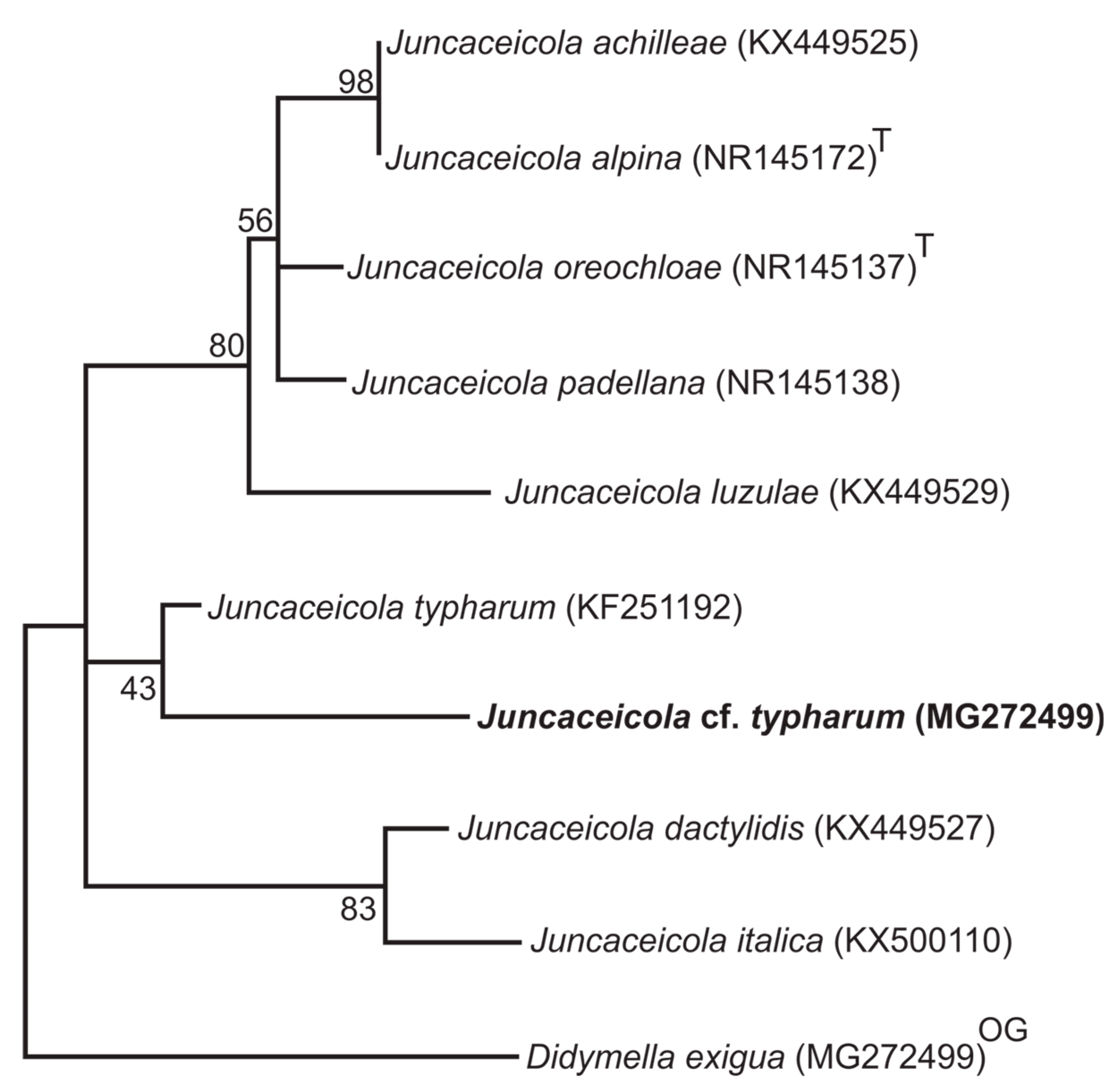
3.1. Phylogenetic Analyzes
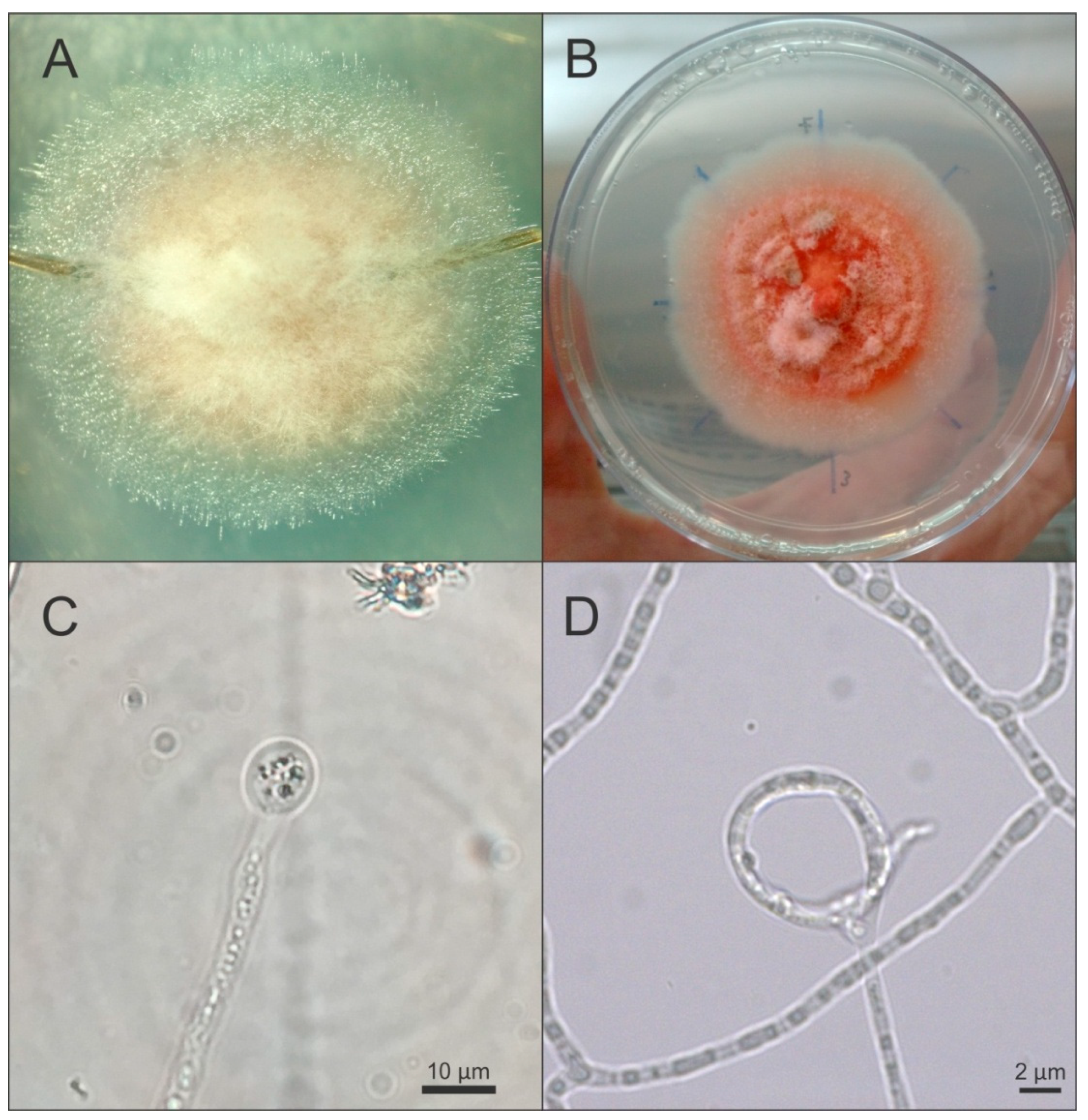
3.2. Morphological Description
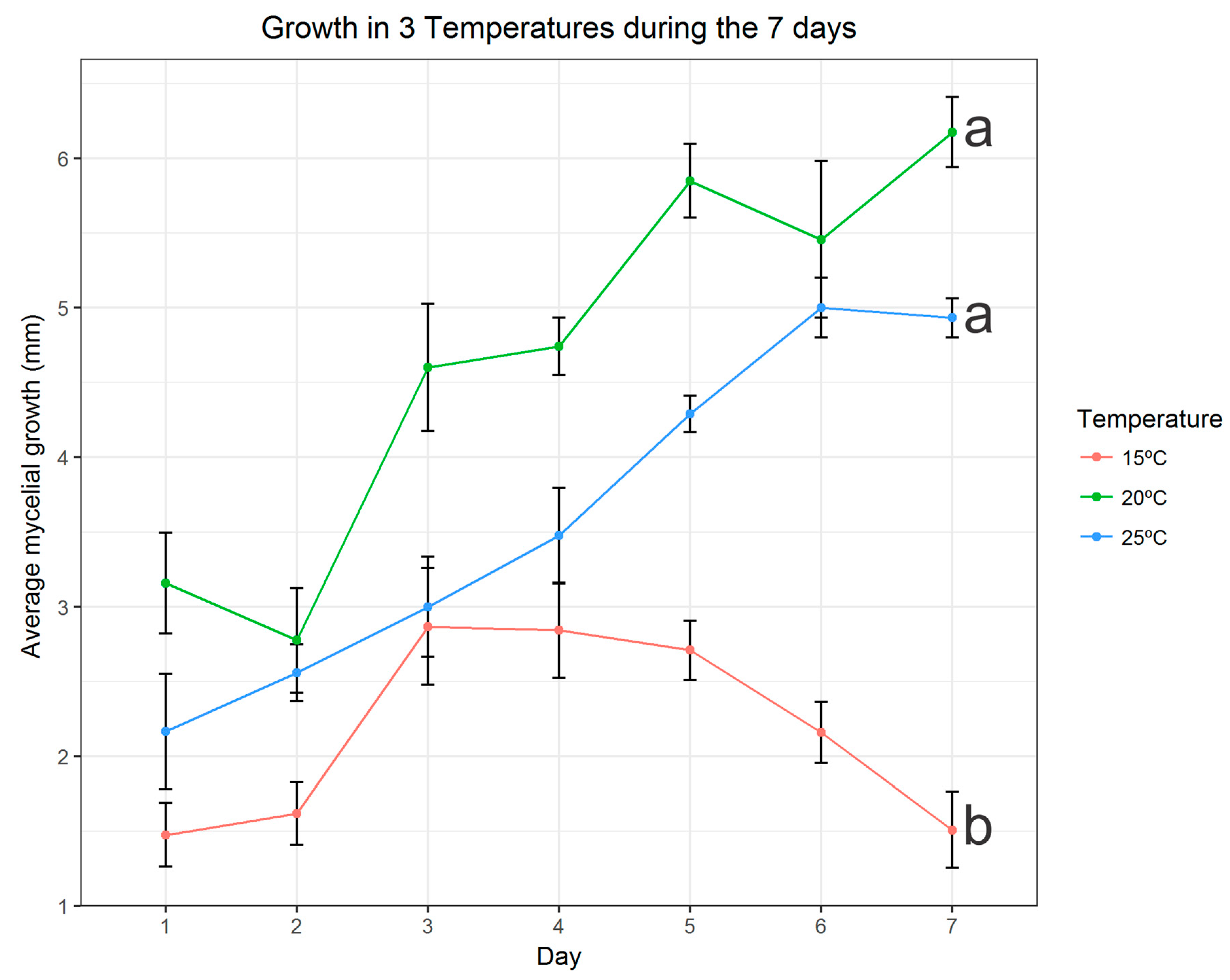
3.3. Effects of Temperature on Mycelia Growth
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Torres-Díaz, C.; Gallardo-Cerda, J.; Lavin, P.; Oses, R.; Carrasco-Urra, F.; Atala, C.; Acuña-Rodríguez, I.S.; Convey, P.; Molina-Montenegro, M.A. Biological Interactions and Simulated Climate Change Modulates the Ecophysiological Performance of Colobanthus quitensis in the Antarctic Ecosystem. PLoS ONE 2016, 11. [Google Scholar] [CrossRef] [PubMed]
- Chénard, C.; Lauro, F.M. (Eds.) Microbial Ecology of Extreme Environments; Springer International Publishing: Cham, Switzerland, 2017. [Google Scholar]
- Convey, P. Antarctic terrestrial biodiversity in a changing world. Polar Biol. 2011, 34, 1629–1641. [Google Scholar] [CrossRef]
- Marcías, M.L.; Deregibus, D.; Saravia, L.A.; Campana, G.L.; Quartino, M.L. Life between tides: Spatial and temporal variations of an intertidal macroalgal community at Potter Peninsula, South Shetland Islands, Antarctica. Estuar. Coast. Shelf Sci. 2017, 187, 193–203. [Google Scholar] [CrossRef]
- Ruisi, S.; Barreca, D.; Selbmann, L.; Zucconi, L.; Onofri, S. Fungi in Antarctica. Rev. Environ. Sci. Biotechnol. 2007, 6, 127–141. [Google Scholar] [CrossRef]
- Franco, D.C.; Signori, C.N.; Duarte, R.T.D.; Nakayama, C.R.; Campos, L.S.; Pellizari, V.H. High Prevalence of Gammaproteobacteria in the Sediments of Admiralty Bay and North Bransfield Basin, Northwestern Antarctic Peninsula. Front. Microbiol. 2017, 8. [Google Scholar] [CrossRef] [PubMed]
- Makhalanyane, T.P.; Van Goethem, M.W.; Cowan, D.A. Microbial diversity and functional capacity in polar soils. Curr. Opin. Biotechnol. 2016, 38, 159–166. [Google Scholar] [CrossRef] [PubMed]
- De Menezes, G.C.A.; Godinho, V.M.; Porto, B.A.; Gonçalves, V.N.; Rosa, L.H. Antarctomyces pellizariae sp. nov., a new, endemic, blue, snow resident psychrophilic ascomycete fungus from Antarctica. Extremophiles 2017, 21, 259–269. [Google Scholar] [CrossRef] [PubMed]
- Santiago, I.F.; Rosa, C.A.; Rosa, L.H. Endophytic symbiont yeasts associated with the Antarctic angiosperms Deschampsia antarctica and Colobanthus quitensis. Polar Biol. 2017, 40, 177–183. [Google Scholar] [CrossRef]
- Ding, Z.; Li, L.; Che, Q.; Li, D.; Gu, Q.; Zhu, T. Richness and bioactivity of culturable soil fungi from the Fildes Peninsula, Antarctica. Extremophiles 2016, 20, 425–435. [Google Scholar] [CrossRef] [PubMed]
- Gonçalves, V.N.; Vitoreli, G.A.; de Menezes, G.C.A.; Mendes, C.R.B.; Secchi, E.R.; Rosa, C.A.; Rosa, L.H. Taxonomy, phylogeny and ecology of cultivable fungi present in seawater gradients across the Northern Antarctica Peninsula. Extremophiles 2017. [Google Scholar] [CrossRef] [PubMed]
- Rosa, L.H.; Vaz, A.B.M.; Caligiorne, R.B.; Campolina, S.; Rosa, C.A. Endophytic fungi associated with the Antarctic grass Deschampsia antarctica Desv. (Poaceae). Polar Biol. 2009, 32, 161–167. [Google Scholar] [CrossRef]
- Rosa, L.H.; de Almeida Vieira, M.L.; Santiago, I.F.; Rosa, C.A. Endophytic fungi community associated with the dicotyledonous plant Colobanthus quitensis (Kunth) Bartl. (Caryophyllaceae) in Antarctica: Endophytic fungi associated with Colobanthus quitensis. FEMS Microbiol. Ecol. 2010. [Google Scholar] [CrossRef] [PubMed]
- Upson, R.; Newsham, K.; Read, D. Root-fungal associations of Colobanthus quitensis and Deschampsia antarctica in the maritime and subantarctic. Arct. Antarct. Alp. Res. 2008, 40, 592–599. [Google Scholar] [CrossRef]
- Khiralla, A.; Spina, R.; Yagi, S.; Mohamed, I.; Laurain-Mattar, D. Endophytic Fungi: Occurrence, Classification, Function and Natural Products. Endophytic Fungi Diversity Characterization and Biocontrol; Hughes, E., Ed.; Nova Science Publishers: New York, NY, USA, 2017; pp. 14–39. ISBN 978-1-53610-341-0. [Google Scholar]
- Chowdhary, K.; Kaushik, N. Fungal endophyte diversity and bioactivity in the Indian medicinal plant Ocimum sanctum Linn. PLoS ONE 2015, 10, e0141444. [Google Scholar] [CrossRef] [PubMed]
- Arnold, A.E. Endophytic Fungi: Hidden Components of Tropical Community Ecology. Tropical Forest Community Ecology; Carson, W., Schnitzer, S., Eds.; Wiley-Blackwell: Hoboken, NJ, USA, 2008; pp. 178–188. ISBN 978-1-405-11897-2. [Google Scholar]
- Hokama, Y.; Savi, D.; Assad, B.; Aluizio, R.; Gomes-Figueiredo, J.; Adamoski, D. Endophytic Fungi Isolated from Vochysia divergens in the Pantanal, Matogrosso do sul: Diversity, Phylogeny and Biocontrol of Phyllosticta citricarpa. In Endophytic Fungi: Diversity Characterization and Biocontrol; Hughes, E., Ed.; Nova Science Publishers: New York, NY, USA, 2017; pp. 1–25. ISBN 978-1-53610-341-0. [Google Scholar]
- Martin, F.; Cullen, D.; Hibbett, D.; Pisabarro, A.; Spatafora, J.W.; Baker, S.E.; Grigoriev, I.V. Sequencing the fungal tree of life. New Phytol. 2011, 190, 818–821. [Google Scholar] [CrossRef] [PubMed]
- Araújo, W.L.; Lacava, P.T.; Marco, J.; Lima, A.O.S.; Sobral, J.K.; Azevedo, J.L.; Pizzirani-Kleiner, A.A. Isolamento de Microrganismos Endofíticos. Guiaprático: Isolamento e Caracterização de Microrganismos Endofíticos. Guia Prático: Isolamento e Caracterização de Microrganismos Endofíticos; Luiz de Queiroz: Piracicaba, Brasil, 2010; p. 167. [Google Scholar]
- Strobel, G.; Daisy, B. Bioprospecting for microbial endophytes and their natural products. Microbiol. Mol. Biol. Rev. 2003, 67, 491–502. [Google Scholar] [CrossRef] [PubMed]
- Castellani, A. Maintenance and cultivation of common pathogenic fungi of man in sterile distilled water. Further researches. J. Trop. Med. Hyg. 1967, 70, 181. [Google Scholar]
- White, T.J.; Bruns, T.D.; Lee, S.B. Amplification and Direct Sequencing of Fungal Ribosomal RNA Genes for Phylogenetics. In PCR Protocols: A Guide to Methods and Applications; Innis, M.A., Gelfand, D.H., Sninsky, J.J., White, T.J., Eds.; Academic Press: San Diego, CA, USA, 1990; pp. 315–322. ISBN 0123721814. [Google Scholar]
- Hall, T.A. BioEdit: A User-Friendly Biological Sequence Alignment Editor and Analysis Program for Windows 95/98/NT. In Nucleic acids Symposium Series; Information Retrieval Ltd.: London, UK, 1999; pp. 95–98. [Google Scholar]
- Goodstadt, L.; Ponting, C.P. CHROMA: Consensus-based colouring of multiple alignments for publication. Bioinformatics 2001, 17, 845–846. [Google Scholar] [CrossRef] [PubMed]
- Burland, T.G. DNASTAR’s Lasergene sequence analysis software. Bioinform. Methods Protoc. 2000, 71–91. [Google Scholar] [CrossRef]
- Altschul, S.F.; Madden, T.L.; Schäffer, A.A.; Zhang, J.; Zhang, Z.; Miller, W.; Lipman, D.J. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res. 1997, 25, 3389–3402. [Google Scholar] [CrossRef] [PubMed]
- Nei, M.; Kumar, S. Molecular Evolution and Phylogenetics; Oxford University Press: Oxford, UK, 2000; ISBN 9780195135855. [Google Scholar]
- Felsenstein, J. Confidence limits on phylogenies: An approach using the bootstrap. Evolution 1985, 39, 783–791. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed]
- Posada, D. jModelTest: Phylogenetic model averaging. Mol Biolevol. 2008, 25, 1253–1256. [Google Scholar] [CrossRef] [PubMed]
- MycoBank. Available online: http://www.mycobank.org (accessed on 10 November 2017).
- Index Fungorum. Available online: http://www.indexfungorum.org (accessed on 10 November 2017).
- Royston, P. Remark AS R94: A remark on Algorithm AS 181: The W test for normality. Appl. Stat. 1995, 44, 547–551. [Google Scholar] [CrossRef]
- Hollander, M.; Douglas, A. Wolfe, Nonparametric Statistical Methods; John Wiley & Sons: New York, NY, USA, 1999; pp. 68–75. [Google Scholar]
- Chambers, J.M.; Freeny, A.; Heiberger, R.M. Analysis of Variance; Designed Experiments; Chapter 5 of Statistical Models; Chambers, J.M., Hastie, T.J., Eds.; Wadsworth & Brooks/Cole: New York, NY, USA, 1992. [Google Scholar]
- Nobles, M.K. Identification of cultures of Wood in habiting hymenomycetes. Can. J. Bot. 1965, 43, 1097–1139. [Google Scholar] [CrossRef]
- OACC: The Online Auction Colour Chart: The New Language of Colour for Buyers and Sellers; OACC: Palo Alto, CA, USA, 2004; p. 12.
- Tennakoon, D.S.; Hyde, K.D.; Phookamsak, R.; Wanasinghe, D.N.; Camporesi, E.; Promputtha, I. Taxonomy and Phylogeny of Juncaceicola gen. nov. (Phaeosphaeriaceae, Pleosporinae, Pleosporales). Cryptogam. Mycol. 2016, 37, 135–156. [Google Scholar] [CrossRef]
- Hyde, K.D.; Hongsanan, S.; Jeewon, R.; Bhat, D.J.; McKenzie, E.H.C.; Jones, E.B.G.; Phookamsak, R.; Ariyawansa, H.A.; Boonmee, S.; Zhao, Q.; et al. Fungal diversity notes 367–490: Taxonomic and phylogenetic contributions to fungal taxa. Fungal Divers. 2016, 80, 1–270. [Google Scholar] [CrossRef]
- Hughes, E. (Ed.) Endophytic Fungi: Diversity, Characterization and Biocontrol; Nova Publishers: New York, NY, USA, 2017; pp. 14–39. ISBN 978-1-53610-341-0. [Google Scholar]
- Higgins, K.L.; Arnold, A.E.; Miadlikowska, J.; Sarvate, S.D.; Lutzoni, F. Phylogenetic relationships, host affinity, and geographic structure of boreal and arctic endophytes from three major plant lineages. Mol. Phylogenet. Evol. 2007, 42, 543–555. [Google Scholar] [CrossRef] [PubMed]
- Debbab, A.; Aly, A.H.; Edrada-Ebel, R.; Wray, V.; Müller, W.E.; Totzke, F.; Zirrgiebel, U.; Schächtele, C.; Kubbutat, M.H.; Lin, W.H. Bioactive metabolites from the endophytic fungus Stemphylium globuliferum isolated from Mentha pulegium. J. Nat. Prod. 2009, 72, 626–631. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Ebada, S.S.; Proksch, P. Pestalotiopsis a highly creative genus: Chemistry and bioactivity of secondary metabolites. Fungal Divers. 2010, 44, 15–31. [Google Scholar] [CrossRef]
- Poczai, P.; Hyvönen, J. Nuclear ribosomal spacer region in plant phylogenetics: Problems and prospects. Mol. Biol. Rep. 2010, 37, 1897–1912. [Google Scholar] [CrossRef] [PubMed]
- Brader, G.; Compant, S.; Vescio, K.; Mitter, B.; Trognitz, F.; Ma, LJ.; Sessitsch, A. Ecology and Genomic Insights into Plant-Pathogenic and Plant-Nonpathogenic Endophytes. Ann. Rev. Phytopathol. 2017, 55, 61–83. [Google Scholar] [CrossRef] [PubMed]
- Barbosa, M.V.; Pereira, E.A.; Cury, J.C.; Carneiro, M.A.C. Occurrence of arbuscular mycorrhizal fungi on King George Island, South Shetland Island, Antarctica. Anais Acad. Bras. Ciênc. 2017, 8, 1737–1743. [Google Scholar] [CrossRef] [PubMed]
- Putzke, J.; Pereira, A.B. Phaeosphaeria deschampsii (Ascomycota): A new parasite species of Deschampsia antarctica (Poaceae) described to Antarctica. Ann. Braz. Acad. Sci. 2016, 88, 1967–1969. [Google Scholar] [CrossRef] [PubMed]
- Zucconi, L.; Pagano, S.; Fenice, M.; Selbmann, L.; Tosi, S.; Onofri, S. Growth temperature preferences of fungal strains from Victoria Land, Antarctica. Polar Biol. 1996, 16, 53–61. [Google Scholar] [CrossRef]
- Tosi, S.; Casado, B.; Gerdol, R.; Caretta, G. Fungi isolated from Antarctic mosses. Polar Biol. 2002, 25, 262–268. [Google Scholar]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
De Andrade, G.A.K.; Cañón, E.R.P.; Alves, R.P.; Schmitz, D.; Schünemann, A.L.; De Albuquerque, M.P.; Putzke, J.; Pereira, A.B.; De Carvalho Victoria, F. First Record of Juncaceicola as Endophytic Fungi Associated with Deschampsia antarctica Desv. Diversity 2018, 10, 107. https://doi.org/10.3390/d10040107
De Andrade GAK, Cañón ERP, Alves RP, Schmitz D, Schünemann AL, De Albuquerque MP, Putzke J, Pereira AB, De Carvalho Victoria F. First Record of Juncaceicola as Endophytic Fungi Associated with Deschampsia antarctica Desv. Diversity. 2018; 10(4):107. https://doi.org/10.3390/d10040107
Chicago/Turabian StyleDe Andrade, Guilherme Afonso Kessler, Ehidy Rocio Peña Cañón, Rodrigo Paidano Alves, Daniela Schmitz, Adriano Luis Schünemann, Margéli Pereira De Albuquerque, Jair Putzke, Antônio Batista Pereira, and Filipe De Carvalho Victoria. 2018. "First Record of Juncaceicola as Endophytic Fungi Associated with Deschampsia antarctica Desv." Diversity 10, no. 4: 107. https://doi.org/10.3390/d10040107
APA StyleDe Andrade, G. A. K., Cañón, E. R. P., Alves, R. P., Schmitz, D., Schünemann, A. L., De Albuquerque, M. P., Putzke, J., Pereira, A. B., & De Carvalho Victoria, F. (2018). First Record of Juncaceicola as Endophytic Fungi Associated with Deschampsia antarctica Desv. Diversity, 10(4), 107. https://doi.org/10.3390/d10040107






