Coconut Milk-Derived Bioactive Peptides as Multifunctional Agents Against Hyperglycemia, Oxidative Stress, and Glycation: An Integrated Experimental and Computational Study
Abstract
1. Introduction
2. Results and Discussion
2.1. Extraction, Estimation, and Profiling of Coconut Milk Proteins
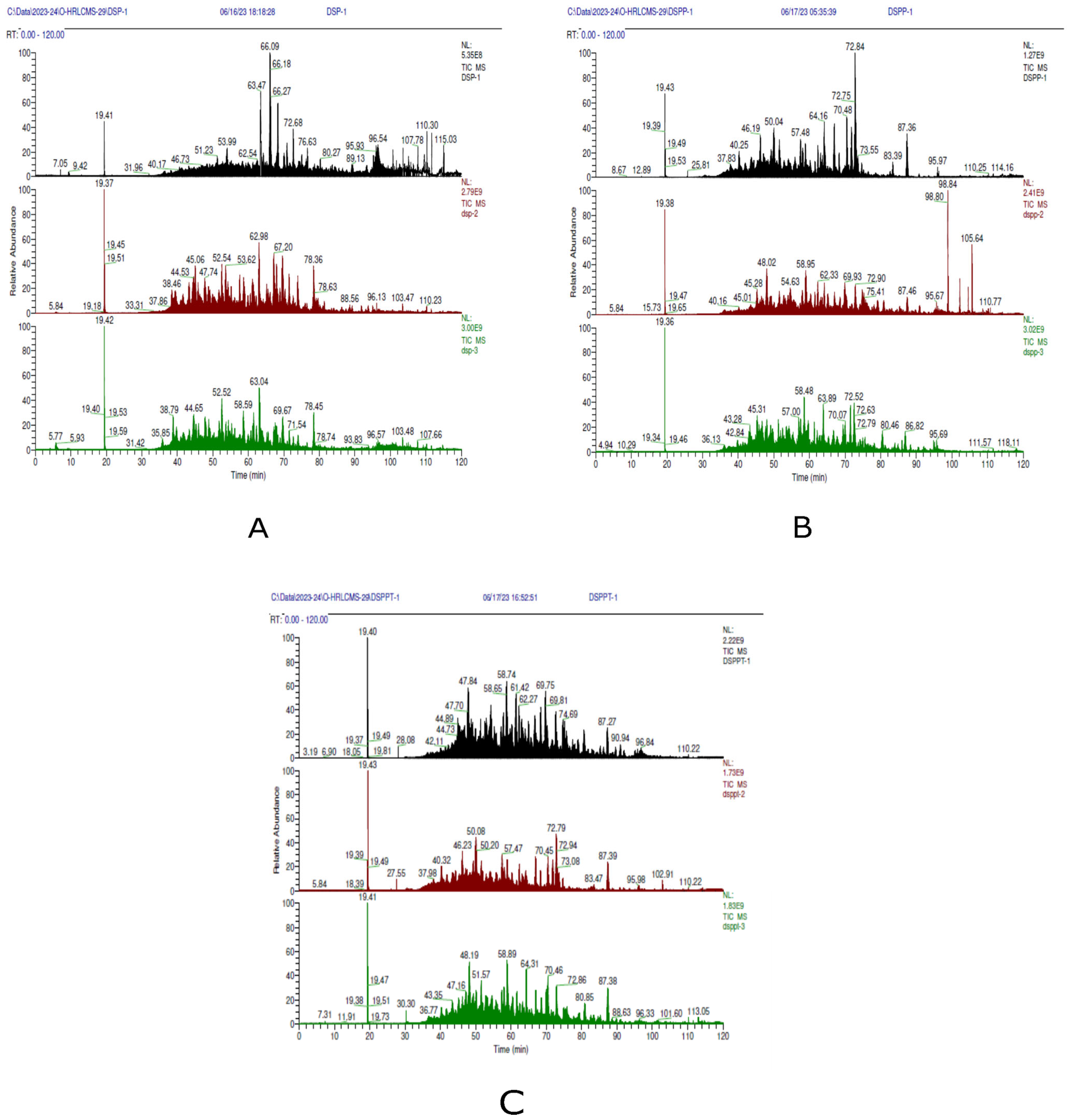
2.2. Identification of Peptides by Nano-ESI-Orbitrap-LC-MS/MS
2.3. In Silico Screening and Identification of Lead Peptides
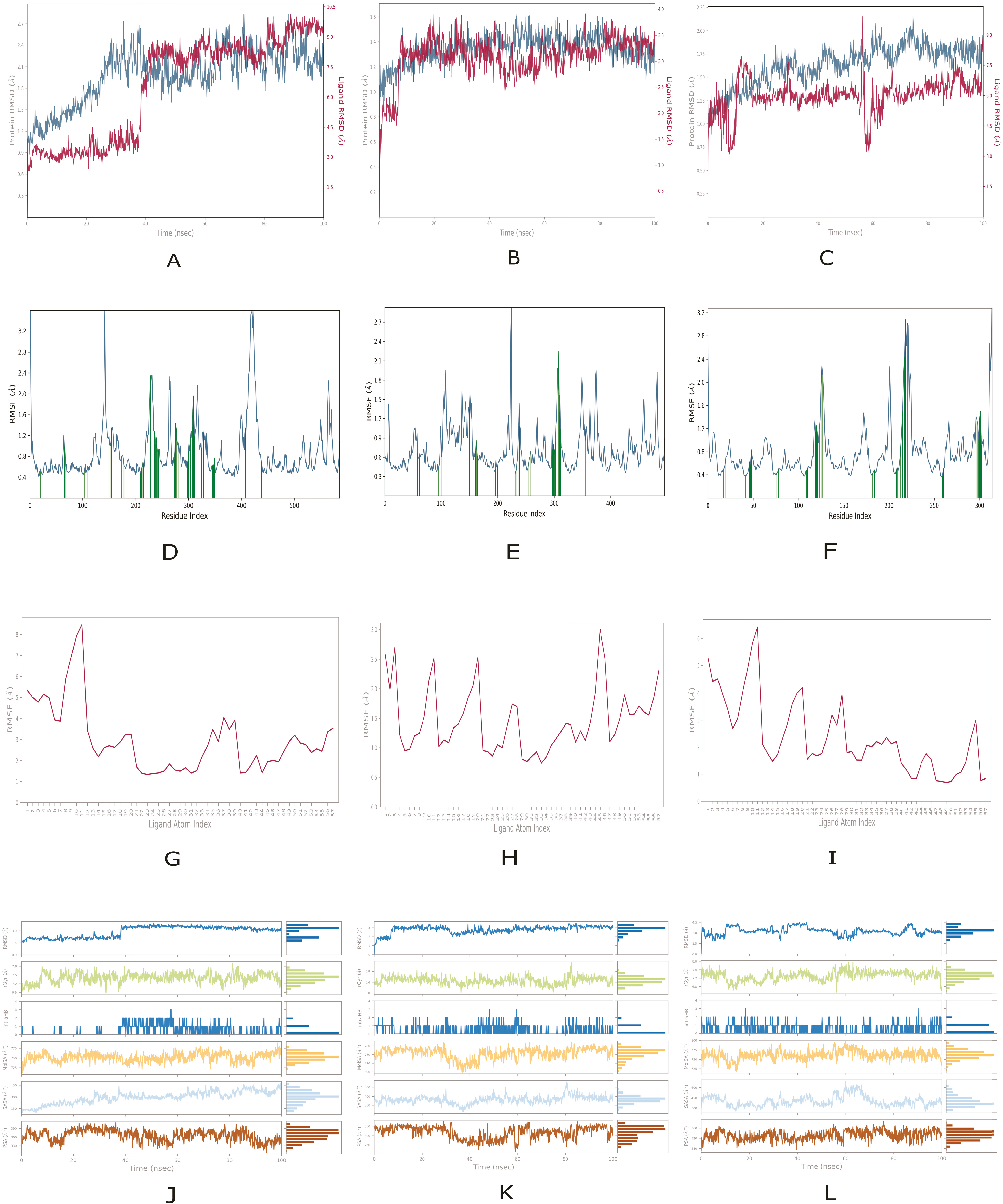
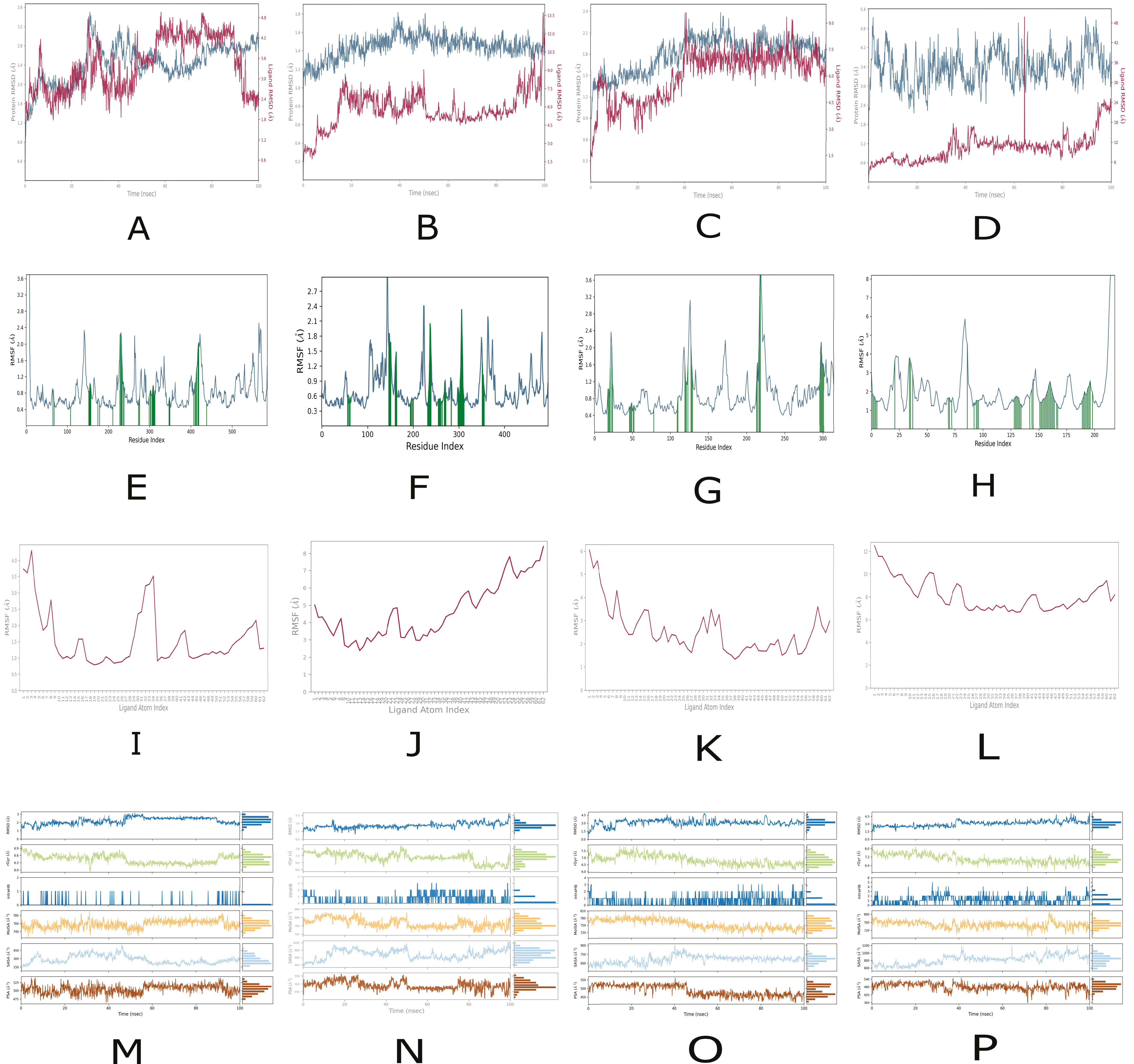
2.4. Molecular Dynamics Analysis of Peptide and Reference Drug Complexes
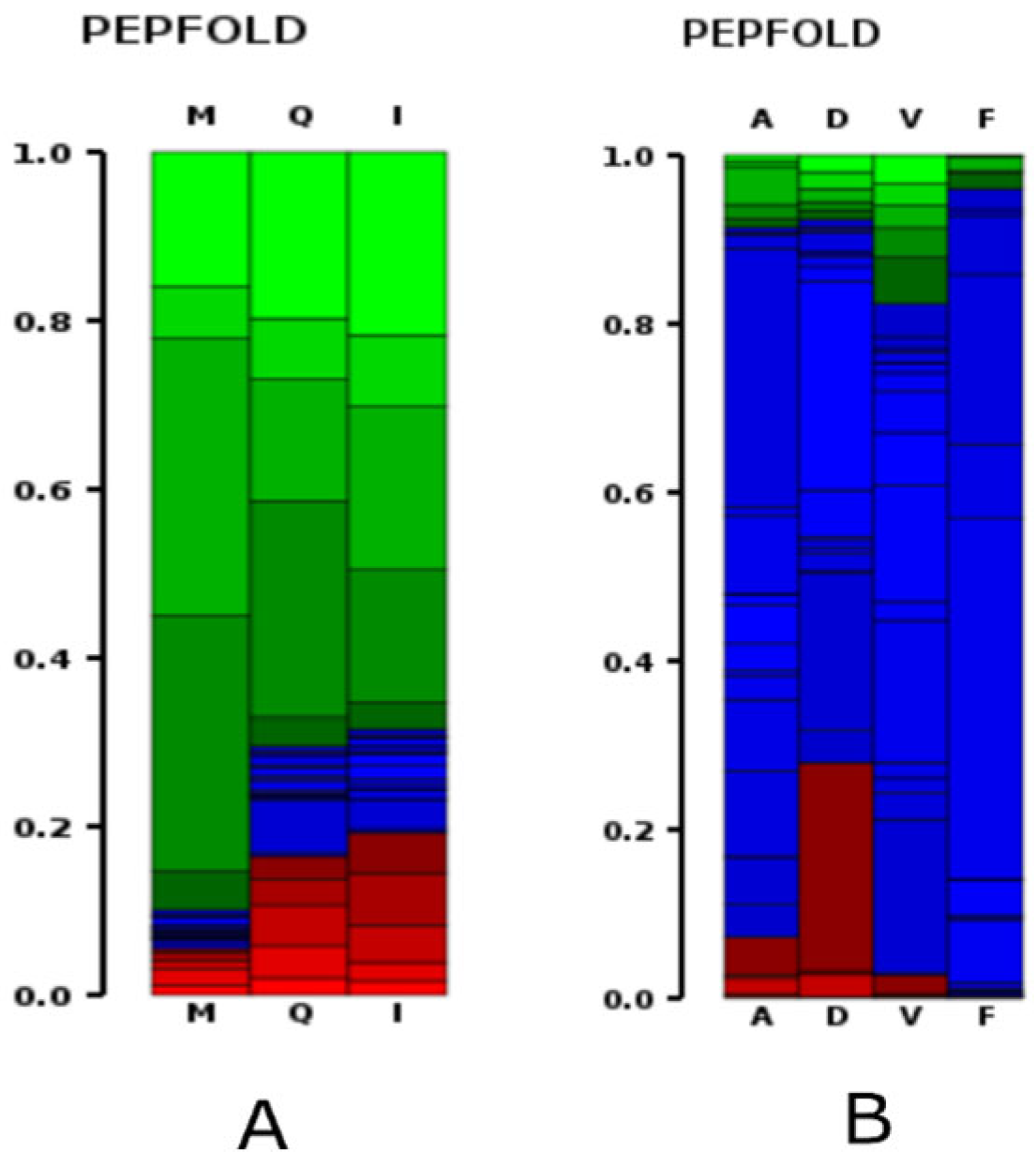
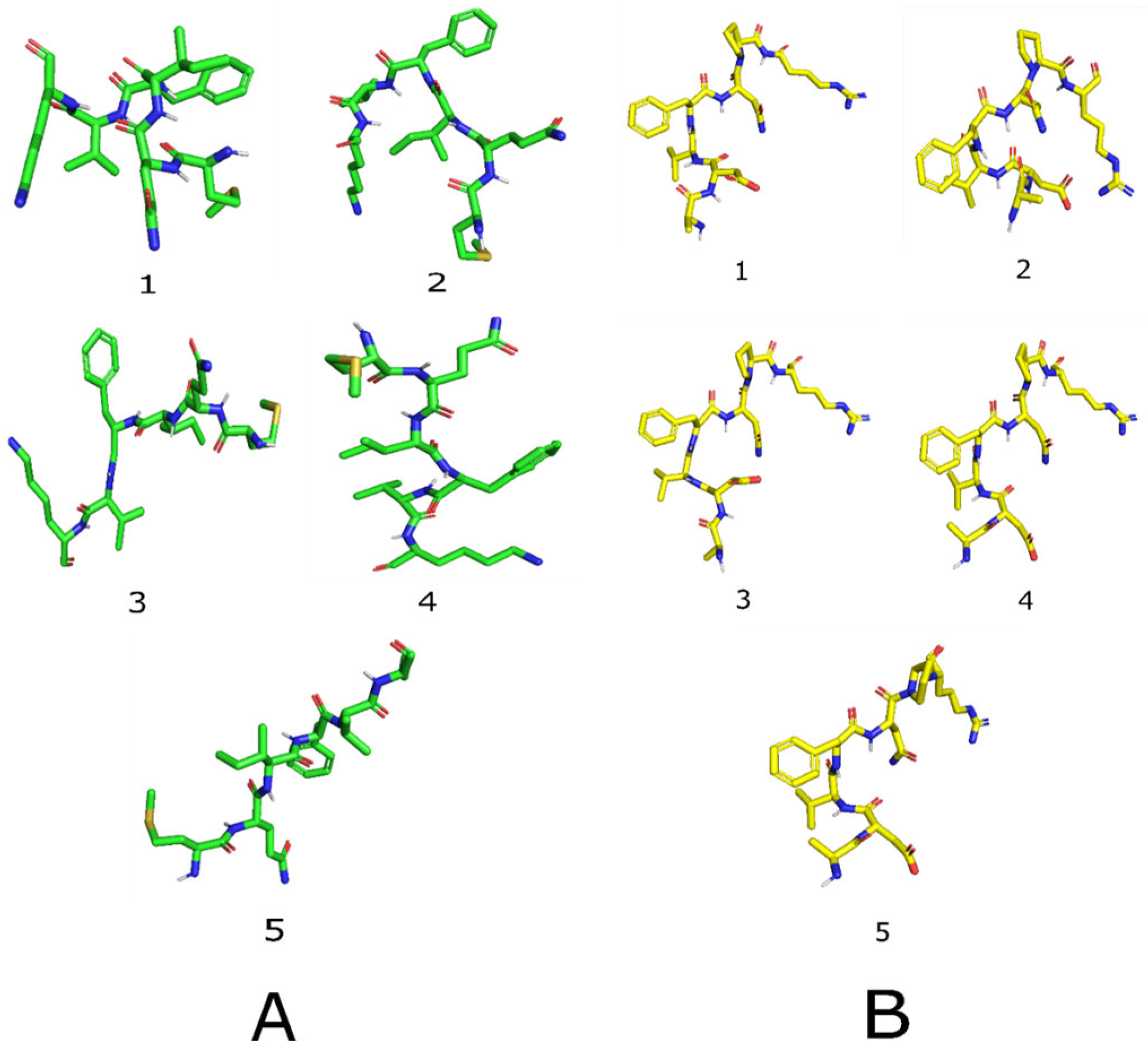
2.5. Peptide Structure Prediction
2.6. Safety Assessment of Coconut Milk Peptides by Hemolytic Assay
| Enzymes | Treatment | Mode of inhibition x | Km (mM) | Vmax 103(µM/min)−1 | Ki(µg) y,z |
|---|---|---|---|---|---|
| α—Glucosidase | Control | Competitive | 2.26 | 15.15 | 0.95 ± 0.09 |
| IC20—13.13 µg | 1.78 | 14.87 | |||
| IC40—26.26 µg | 0.60 | 14.68 | |||
| IC60—39.39 µg | 0.18 | 14.50 | |||
| α—Amylase | Control | Competitive | 3.15 | 28.28 | 1.34 ± 0.12 |
| IC20—33.55 µg | 2.00 | 27.75 | |||
| IC40—67.10 µg | 0.79 | 26.26 | |||
| IC60—100.60 µg | 0.22 | 27.50 | |||
| Aldose reductase | Control | Competitive | 6.30 | 47.35 | 0.78 ± 0.05 |
| IC20—8.12 µg | 4.86 | 46.64 | |||
| IC40—16.23 µg | 2.02 | 46.02 | |||
| IC60—24.35 µg | 0.66 | 45.55 |
2.7. Evaluation of In Vitro Antihyperglycemic, Anti-Glycation Activities of Peptides and Enzyme Inhibition Kinetics of the Lead Peptide
2.8. Evaluation of Methyl Glyoxal Scavenging Activity of Peptides
| Samples | IC50 (µg/mL/µM) / EC50 (µg/mL/µM) | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| Anti-Hyperglycemic Assays | Anti-Glycation Assay (Methylglyoxal Scavenging) | Safety Assessment of Peptides: Hemolytic Assay (HC50) (µg/mL/µM) | Anti-Oxidant Assays | ||||||
| α-Glucosidase Inhibition | α-Amylase Inhibition | Aldose Reductase Inhibition | EC50 (µg/mL/µM) at RT | EC50 (µg/mL/µM) at 37 °C, 2 h | DPPH Radical Scavenging Assay | ABTS Radical Scavenging Assay | Superoxide Radical Scavenging Assay | ||
| Peptide 1 | 38.98 ± 0.23 b (50.96) | 88.25 ± 0.34 b (115.36) | 22.51 ± 0.04 b (29.43) | 215.17 ± 0.44 b (281.28) | 190.13 ± 0.50 b (248.54) | 220.85 ± 0.20 b (288.70) | 26.14 ± 0.12 c (34.17) | 27.89 ± 0.17 c (36.46) | 52.32 ± 0.30 b (68.39) |
| Peptide 2 | 32.82 ± 0.17 a (40.13) | 83.88 ± 0.22 a (102.56) | 20.29 ± 0.02 a (24.81) | 208.10 ± 0.65 a (254.44) | 182.17 ± 0.23 a (222.73) | 231.53 ± 0.14 c (283.08) | 22.14 ± 0.12 a (27.07) | 17.89 ± 0.20 a (21.87) | 42.45 ± 0.30 a (51.90) |
| * Acarbose | 46.16 ± 0.23 c (71.50) | 89.62 ± 0.46 c (138.82) | - | - | - | - | - | - | - |
| ** Quercetin | - | - | 25.22 ± 0.07 c (83.44) | - | - | - | - | - | - |
| *** Creatine | - | - | - | 230.05 ± 0.34 c (1754.37) | - | - | - | - | |
| **** Triton-X-100 | - | - | - | - | - | 105.25 ± 0.17 a (162.67) | - | - | - |
| ***** Ascorbic acid | - | - | - | - | - | - | 28.65 ± 0.17 b (162.67) | 22.74 ± 0.27 b (129.12) | 57.71 ± 0.16 c (327.67) |
2.9. Evaluation of the Antioxidant Potential of Coconut Milk Peptides
3. Materials and Methods
3.1. Materials
3.2. Extraction, Estimation, and Profiling of Coconut Milk Proteins
3.3. In Vitro Gastro-Intestinal Protein Digestion
3.4. Identification of Peptides by Nano-LC-MS/MS Orbitrap Analysis
3.5. Physico-Chemical Properties of Peptides
3.6. Molecular Docking
3.7. Molecular Dynamics Simulations
3.8. Peptide Structure Prediction
3.9. Synthesis of Lead Peptides
3.10. Safety Assessment of Coconut Milk Peptides by Hemolytic Assay
3.11. α-Glucosidase Inhibition Assay
3.12. α-Amylase Inhibition Assay
3.13. Aldose-Reductase Inhibition Assay
3.14. Kinetics of Enzyme Inhibition
3.15. Maillard Reaction Models and Evaluation of Methyl Glyoxal Scavenging Activity
3.16. Antioxidant Assays
3.17. Statistical Analysis
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ramu, R.; Shirahatti, P.S.; Zameer, F.; Ranganatha, L.V.; Nagendra Prasad, M.N. Inhibitory effect of banana (Musa sp. var. Nanjangud rasa bale) flower extract and its constituents Umbelliferone and Lupeol on α-glucosidase, aldose reductase and glycation at multiple stages. S. Afr. J. Bot. 2014, 95, 54–63. [Google Scholar] [CrossRef]
- Rabbani, N.; Thornalley, P.J. Dicarbonyl stress in cell and tissue dysfunction contributing to ageing and disease. Biochem. Biophys. Res. Commun. 2015, 458, 221–226. [Google Scholar] [CrossRef]
- Taguchi, K.; Fukami, K. RAGE signaling regulates the progression of diabetic complications. Front. Pharmacol. 2023, 14, 1128872. [Google Scholar] [CrossRef] [PubMed]
- Su, J.; Lucchesi, P.A.; Gonzalez-Villalobos, R.A.; Palen, D.I.; Rezk, B.M.; Suzuki, Y.; Boulares, H.A.; Matrougui, K. Role of advanced glycation end products with oxidative stress in resistance artery dysfunction in type 2 diabetic mice. Arterioscler. Thromb. Vasc. Biol. 2008, 28, 1432–1438. [Google Scholar] [CrossRef]
- Kerkeni, M.; Saïdi, A.; Bouzidi, H.; Letaief, A.; Ben Yahia, S.; Hammami, M. Pentosidine as a biomarker for microvascular complications in type 2 diabetic patients. Diabetes Vasc. Dis. Res. 2013, 10, 239–245. [Google Scholar] [CrossRef]
- Rochín-Medina, J.J.; Ramírez-Serrano, E.S.; Ramírez, K. Inhibition of α-glucosidase activity by potential peptides derived from fermented spent coffee grounds. Food Chem. 2024, 454, 139791. [Google Scholar] [CrossRef]
- Fadimu, G.J.; Gill, H.; Farahnaky, A.; Truong, T. Improving the enzymolysis efficiency of lupin protein by ultrasound pretreatment: Effect on antihypertensive, antidiabetic and antioxidant activities of the hydrolysates. Food Chem. 2022, 383, 132457. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Wang, J.; Liu, Y.; Sun, B. Peptides derived from oats improve insulin sensitivity. J. Biomed. Sci. 2015, 4, 1. [Google Scholar] [CrossRef]
- Gengan, G.; Zaini, N.S.M.; Saari, N.; Hussin, A.S.M.; Jaafar, A.H.; Hasan, H.; Lim, E.J.; Abd Al Qadr Imad Wan-Mohtar, W.; Abd Rahim, M.H. Nutritional and therapeutic benefits of coconut milk and its potential as a plant-based functional yogurt alternative: A review. Food Sci. Hum. Wellness 2025, 14, 9250004. [Google Scholar] [CrossRef]
- Alatawi, K.A.; Alshubaily, F.A. Coconut products alleviate hyperglycaemic, hyperlipidimic and nephropathy indices in streptozotocin-induced diabetic wistar rats. Saudi J. Biol. Sci. 2021, 28, 4224–4231. [Google Scholar] [CrossRef]
- Salil, G.; Nevin, K.G.; Rajamohan, T. Arginine rich coconut kernel protein modulates diabetes in alloxan treated rats. Chem. Biol. Interact. 2011, 189, 107–111. [Google Scholar] [CrossRef] [PubMed]
- Karoui, R.; Bouaicha, I. A review on nutritional quality of animal and plant-based milk alternatives: A focus on protein. Front. Nutr. 2024, 11, 1378556. [Google Scholar] [CrossRef]
- Kwon, K.S.; Bae, D.; Park, K.H.; Rhee, K.C. Aqueous extraction and membrane techniques improve coconut protein concentrate functionality. J. Food Sci. 1996, 61, 753–756. [Google Scholar] [CrossRef]
- Senphan, T.; Benjakul, S. Comparative Study on Virgin Coconut Oil Extraction Using Protease from Hepatopancreas of Pacific White Shrimp and Alcalase. J. Food Process. Preserv. 2017, 41, e12771. [Google Scholar] [CrossRef]
- Tangsuphoom, N.; Coupland, J.N. Effect of surface-active stabilizers on the surface properties of coconut milk emulsions. Food Hydrocoll. 2009, 23, 1801–1809. [Google Scholar] [CrossRef]
- Garcia, R.N.; Arocena, R.V.; Laurena, A.C.; Tecson-Mendoza, E.M. 11S and 7S globulins of coconut (Cocos nucifera L.): Purification and characterization. J. Agric. Food Chem. 2005, 53, 1734–1739. [Google Scholar] [CrossRef]
- Fadimu, G.J.; Farahnaky, A.; Gill, H.; Olalere, O.A.; Gan, C.Y.; Truong, T. In-Silico Analysis and Antidiabetic Effect of α-Amylase and α-Glucosidase Inhibitory Peptides from Lupin Protein Hydrolysate: Enzyme-Peptide Interaction Study Using Molecular Docking Approach. Foods 2022, 11, 3375. [Google Scholar] [CrossRef]
- Mohd Rodhi, A.; Yap, P.G.; Olalere, O.A.; Gan, C.Y. Exploring α-Glucosidase Inhibitory Peptides: Structure-Activity Relationship Analysis and Perspectives for Designing Potential Anti-Diabetic Agents. Jundishapur J. Nat. Pharm. Prod. 2023, 18, e139988. [Google Scholar] [CrossRef]
- Karami, Z.; Akbari-adergani, B. Bioactive food derived peptides: A review on correlation between structure of bioactive peptides and their functional properties. J. Food Sci. Technol. 2019, 56, 535–547. [Google Scholar] [CrossRef] [PubMed]
- Majura, J.J.; Cao, W.; Chen, Z.; Htwe, K.K.; Li, W.; Du, R.; Zhang, P.; Zheng, H.; Gao, J. The current research status and strategies employed to modify food-derived bioactive peptides. Front. Nutr. 2022, 9, 950823. [Google Scholar] [CrossRef]
- Remanan, M.K.; Wu, J. Antioxidant activity in cooked and simulated digested eggs. Food Funct. 2014, 5, 1464–1474. [Google Scholar] [CrossRef]
- Wang, W.; Liu, X.; Li, Y.; You, H.; Yu, Z.; Wang, L.; Liu, X.; Ding, L. Identification and Characterization of Dipeptidyl Peptidase-IV Inhibitory Peptides from Oat Proteins. Foods 2022, 11, 1406. [Google Scholar] [CrossRef]
- Darewicz, M.; Pliszka, M.; Borawska-Dziadkiewicz, J.; Minkiewicz, P.; Iwaniak, A. Multi-Bioactivity of Protein Digests and Peptides from Oat (Avena sativa L.) Kernels in the Prevention of the Cardiometabolic Syndrome. Molecules 2022, 27, 7907. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Zhang, Y.; Yu, T.; He, J.; Cui, J.; Wang, J.; Cheng, X.; Fan, J. Oat globulin peptides regulate antidiabetic drug targets and glucose transporters in Caco-2 cells. J. Funct. Foods 2018, 42, 12–20. [Google Scholar] [CrossRef]
- Sánchez, A.; Vázquez, A. Bioactive peptides: A review. Food Qual. Saf. 2017, 1, 29–46. [Google Scholar] [CrossRef]
- Oyebamiji, A.K.; Akintayo, E.T.; Akintayo, C.O.; Aworinde, H.O.; Adekunle, O.D.; Akintelu, S.A. Biochemical Study on Bioactive Peptides: Potential Anti-Aldose Reductase Targeting Diabetes Mellitus. Lett. Appl. NanoBioSci. 2024, 13, 10–33263. [Google Scholar]
- Kashung, P.; Karuthapandian, D. Milk-derived bioactive peptides. Food Prod. Process. Nutr. 2025, 7, 6. [Google Scholar] [CrossRef]
- Lin, H.; Yang, Y.; Wang, X.; Chung, M.; Zhang, L.; Cai, S.; Pan, X.; Pan, Y. Targeting the AGEs-RAGE axis: Pathogenic mechanisms and therapeutic interventions in diabetic wound healing. Front. Med. 2025, 12, 1667620. [Google Scholar] [CrossRef]
- Patil, S.P.; Goswami, A.; Kalia, K.; Kate, A.S. Plant-Derived Bioactive Peptides: A Treatment to Cure Diabetes. Int. J. Pept. Res. Ther. 2020, 26, 955–968. [Google Scholar] [CrossRef]
- Park, H.; Boyington, J.C. The 1.5 Å Crystal Structure of Human Receptor for Advanced Glycation Endproducts (RAGE) Ectodomains Reveals Unique Features Determining Ligand Binding. J. Biol. Chem. 2010, 285, 40762–40770. [Google Scholar] [CrossRef] [PubMed]
- Bongarzone, S.; Savickas, V.; Luzi, F.; Gee, A.D. Targeting the Receptor for Advanced Glycation Endproducts (RAGE): A Medicinal Chemistry Perspective. J. Med. Chem. 2017, 60, 7213–7232. [Google Scholar] [CrossRef]
- Deane, R.; Singh, I.; Sagare, A.P.; Bell, R.D.; Ross, N.T.; LaRue, B.; Love, R.; Perry, S.; Paquette, N.; Deane, R.J.; et al. A Multimodal RAGE-Specific Inhibitor Reduces Amyloid β–Mediated Brain Disorder in a Mouse Model of Alzheimer Disease. J. Clin. Investig. 2012, 122, 1377–1392. [Google Scholar] [CrossRef]
- Xue, J.; Ray, R.; Singer, D.; Böhme, D.; Burz, D.S.; Rai, V.; Hoffmann, R.; Shekhtman, A. The Receptor for Advanced Glycation End Products (RAGE) Specifically Recognizes Methylglyoxal-Derived AGEs. Biochemistry 2014, 53, 3327–3335. [Google Scholar] [CrossRef]
- Fritz, G. RAGE: A Single Receptor Fits Multiple Ligands. Trends Biochem. Sci. 2011, 36, 625–632. [Google Scholar] [CrossRef]
- Hollingsworth, S.A.; Dror, R.O. Molecular Dynamics Simulation for All. Neuron 2018, 99, 1129–1143. [Google Scholar] [CrossRef] [PubMed]
- De Vivo, M.; Masetti, M.; Bottegoni, G.; Cavalli, A. Role of Molecular Dynamics in Drug Discovery. J. Med. Chem. 2016, 59, 4035–4061. [Google Scholar] [CrossRef]
- Gautier, R.; Douguet, D.; Antonny, B.; Drin, G. HELIQUEST: A web server to screen sequences with specific α-helical properties. Bioinformatics 2008, 24, 2101–2102. [Google Scholar] [CrossRef] [PubMed]
- Wright, P.E.; Dyson, H.J. Intrinsically Unstructured Proteins: Re-Assessing the Protein Structure–Function Paradigm. J. Mol. Biol. 1999, 293, 321–331. [Google Scholar] [CrossRef] [PubMed]
- Nussinov, R.; Liu, Y.; Zhang, W.; Jang, H. Protein conformational ensembles in function: Roles and mechanisms. RSC Chem. Biol. 2023, 4, 850–864. [Google Scholar] [CrossRef]
- Robles-Loaiza, A.A.; Pinos-Tamayo, E.A.; Mendes, B.; Ortega-Pila, J.A.; Proaño-Bolaños, C.; Plisson, F.; Teixeira, C.; Gomes, P.; Almeida, J.R. Traditional and Computational Screening of Non-Toxic Peptides and Approaches to Improving Selectivity. Pharmaceuticals 2022, 15, 323. [Google Scholar] [CrossRef]
- Greco, I.; Molchanova, N.; Holmedal, E.; Jenssen, H.; Hummel, B.D.; Watts, J.L.; Håkansson, J.; Hansen, P.R.; Svenson, J. Correlation between hemolytic activity, cytotoxicity and systemic in vivo toxicity of synthetic antimicrobial peptides. Sci. Rep. 2020, 10, 13206. [Google Scholar] [CrossRef]
- Wang, R.; Zhao, H.; Pan, X.; Orfila, C.; Lu, W.; Ma, Y. Preparation of bioactive peptides with antidiabetic, antihypertensive, and antioxidant activities and identification of α-glucosidase inhibitory peptides from soy protein. Food Sci. Nutr. 2019, 7, 1848–1856. [Google Scholar] [CrossRef]
- Rafique, H.; Dong, R.; Wang, X.; Alim, A.; Aadil, R.M.; Li, L.; Zou, L.; Hu, X. Dietary-Nutraceutical Properties of Oat Protein and Peptides. Front. Nutr. 2022, 9, 950400. [Google Scholar] [CrossRef]
- Patil, S.M.; Jayaram, L.; Ramu, R. Systems biology-based proteo-peptidomic profiling, bioactivity-guided antidiabetic and antioxidant potential of bovine colostrum proteome hydrolysates. Food Biosci. 2025, 73, 107775. [Google Scholar] [CrossRef]
- González-Montoya, M.; Hernández-Ledesma, B.; Mora-Escobedo, R.; Martínez-Villaluenga, C. Bioactive Peptides from Germinated Soybean with Anti-Diabetic Potential by Inhibition of Dipeptidyl Peptidase-IV, α-Amylase, and α-Glucosidase Enzymes. Int. J. Mol. Sci. 2018, 19, 2883. [Google Scholar] [CrossRef] [PubMed]
- Admassu, H.; Gasmalla, M.A.A.; Yang, R.; Zhao, W. Identification of Bioactive Peptides with α-Amylase Inhibitory Potential from Enzymatic Protein Hydrolysates of Red Seaweed (Porphyra spp). J. Agric. Food Chem. 2018, 66, 4872–4882. [Google Scholar] [CrossRef] [PubMed]
- Maradesha, T.; Patil, S.M.; Phanindra, B.; Achar, R.R.; Silina, E.; Stupin, V.; Ramu, R. Multiprotein Inhibitory Effect of Dietary Polyphenol Rutin from Whole Green Jackfruit Flour Targeting Different Stages of Diabetes Mellitus: Defining a Bio-Computational Stratagem. Separations 2022, 9, 262. [Google Scholar] [CrossRef]
- Balestri, F.; Moschini, R.; Mura, U.; Cappiello, M.; Del Corso, A. In Search of Differential Inhibitors of Aldose Reductase. Biomolecules 2022, 12, 485. [Google Scholar] [CrossRef]
- Patil, S.M.; Martiz, R.M.; Satish, A.M.; Shbeer, A.M.; Ageel, M.; Al-Ghorbani, M.; Ranganatha, L.V.; Parameswaran, S.; Ramu, R. Discovery of Novel Coumarin Derivatives as Potential Dual Inhibitors against α-Glucosidase and α-Amylase for the Management of Post-Prandial Hyperglycemia via Molecular Modelling Approaches. Molecules 2022, 27, 3888. [Google Scholar] [CrossRef]
- Risum, A.B.; Bevilacqua, M.; Li, C.; Engholm-Keller, K.; Poojary, M.M.; Rinnan, Å.; Lund, M.N. Resolving fluorescence spectra of Maillard reaction products formed on bovine serum albumin using parallel factor analysis. Food Res. Int. 2024, 178, 113950. [Google Scholar] [CrossRef]
- Zhu, H.; Poojary, M.M.; Andersen, M.L.; Lund, M.N. The effect of molecular structure of polyphenols on the kinetics of the trapping reactions with methylglyoxal. Food Chem. 2020, 319, 126500. [Google Scholar] [CrossRef]
- Deng, Y.; Wang, X.; Zhang, Y.; Zhang, C.; Xie, P.; Huang, L. Inhibitory effect of Ginkgo biloba seeds peptides on methylglyoxal-induced glycations. Food Chem. Toxicol. 2023, 172, 113587. [Google Scholar] [CrossRef]
- Schalkwijk, C.G.; Stehouwer, C.D.A. Methylglyoxal, a highly reactive dicarbonyl compound, in diabetes, its vascular complications, and other age-related diseases. Physiol. Rev. 2020, 100, 407–461. [Google Scholar] [CrossRef]
- Brings, S.; Fleming, T.; De Buhr, S.; Beijer, B.; Lindner, T.; Wischnjow, A.; Kender, Z.; Peters, V.; Kopf, S.; Haberkorn, U.; et al. A scavenger peptide prevents methylglyoxal induced pain in mice. Biochim. Biophys. Acta-Mol. Basis Dis. 2017, 1863, 654–662. [Google Scholar] [CrossRef]
- Wang, L.; Ma, M.; Yu, Z.; Du, S.-K. Preparation and identification of antioxidant peptides from cottonseed proteins. Food Chem. 2021, 352, 129399. [Google Scholar] [CrossRef]
- Chen, M.-L.; Ning, P.; Jiao, Y.; Xu, Z.; Cheng, Y.-H. Extraction of antioxidant peptides from rice dreg protein hydrolysate via an angling method. Food Chem. 2021, 337, 128069. [Google Scholar] [CrossRef] [PubMed]
- Suetsuna, K.; Ukeda, H.; Ochi, H. Isolation and characterization of free radical scavenging activities peptides derived from casein. J. Nutr. Biochem. 2000, 11, 128–131. [Google Scholar] [CrossRef] [PubMed]
- Chambal, B.; Bergenståhl, B.; Dejmek, P. Edible proteins from coconut milk press cake; one step alkaline extraction and characterization by electrophoresis and mass spectrometry. Food Res. Int. 2012, 47, 146–151. [Google Scholar] [CrossRef]
- Naik, A.; Venu, G.V.; Prakash, M.; Raghavarao, K.S.M.S. Dehydration of coconut skim milk and evaluation of functional properties. CYTA-J. Food 2014, 12, 227–234. [Google Scholar] [CrossRef]
- Patil, S.M.; Ramu, R.; Goh, K.W. Protocol for isolation, fractionation, and system biology-based profiling of gastrointestinal digested dairy colostrum and milk proteome. STAR Protoc. 2025, 6, 103657. [Google Scholar] [CrossRef]
- Smith, P.K.; Krohn, R.I.; Hermanson, G.T.; Mallia, A.K.; Gartner, F.H.; Provenzano, M.D.; Fujimoto, E.K.; Goeke, N.M.; Olson, B.J.; Klenk, D.C. Measurement of protein using bicinchoninic acid. Anal. Biochem. 1985, 150, 76–85. [Google Scholar] [CrossRef]
- Laemmli, U.K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 1970, 227, 680–685. [Google Scholar] [CrossRef]
- Martiz, R.M.; Ramu, R.; Nambisan, H.; Suhail, A.; Raish, M.; Patil, S.M.; Ashwini, P.; Mahesh, B.; Przybyłek, M.; Bełdowski, P.; et al. Identification and 3D modeling of bioactive peptides from Lactobacillus brevis RAMULAB49 protein hydrolysate with in silico ERK1 phosphorylation inhibition activity targeting diabetic nephropathy. PLoS ONE 2025, 20, e0331192. [Google Scholar] [CrossRef]
- Gasteiger, E.; Hoogland, C.; Gattiker, A.; Duvaud, S.; Wilkins, M.R.; Appel, R.D.; Bairoch, A. Protein Identification and Analysis Tools on the ExPASy Server. In The Proteomics Protocols Handbook; Humana Press: Totowa, NJ, USA, 2005; pp. 571–607. [Google Scholar]
- Lear, S.; Cobb, S.L. Pep-Calc.com: A set of web utilities for the calculation of peptide and peptoid properties and automatic mass spectral peak assignment. J. Comput. Aided. Mol. Des. 2016, 30, 271–277. [Google Scholar] [CrossRef] [PubMed]
- Saha, S.; Raghava, G.P.S. AlgPred: Prediction of allergenic proteins and mapping of IgE epitopes. Nucleic Acids Res. 2006, 34, W202–W209. [Google Scholar] [CrossRef] [PubMed]
- Gupta, S.; Kapoor, P.; Chaudhary, K.; Gautam, A.; Kumar, R.; Raghava, G.P.S. In Silico Approach for Predicting Toxicity of Peptides and Proteins. PLoS ONE 2013, 8, e73957. [Google Scholar] [CrossRef]
- Yamamoto, K.; Miyake, H.; Kusunoki, M.; Osaki, S. Crystal structures of isomaltase from Saccharomyces cerevisiae and in complex with its competitive inhibitor maltose. FEBS J. 2010, 277, 4205–4214. [Google Scholar] [CrossRef] [PubMed]
- Bompard-Gilles, C.; Rousseau, P.; Rouge, P.; Payan, F. Substrate mimicry in the active center of a mammalian α-amylase: Structural analysis of an enzyme-inhibitor complex. Structure 1996, 4, 1441–1452. [Google Scholar] [CrossRef]
- Kinoshita, T.; Miyake, H.; Fujii, T.; Takakura, S.; Goto, T. The structure of human recombinant aldose reductase complexed with the potent inhibitor zenarestat. Acta Crystallogr. Sect. D Biol. Crystallogr. 2002, 58, 622–626. [Google Scholar] [CrossRef]
- Koch, M.; Chitayat, S.; Dattilo, B.M.; Schiefner, A.; Diez, J.; Chazin, W.J.; Fritz, G. Structural Basis for Ligand Recognition and Activation of RAGE. Structure 2010, 18, 1342–1352. [Google Scholar] [CrossRef]
- Friesner, R.A.; Banks, J.L.; Murphy, R.B.; Halgren, T.A.; Klicic, J.J.; Mainz, D.T.; Repasky, M.P.; Knoll, E.H.; Shelley, M.; Perry, J.K.; et al. Glide: A New Approach for Rapid, Accurate Docking and Scoring. 1. Method and Assessment of Docking Accuracy. J. Med. Chem. 2004, 47, 1739–1749. [Google Scholar] [CrossRef] [PubMed]
- Bowers, K.J.; Chow, D.E.; Xu, H.; Dror, R.O.; Eastwood, M.P.; Gregersen, B.A.; Klepeis, J.L.; Kolossvary, I.; Moraes, M.A.; Sacerdoti, F.D.; et al. Scalable Algorithms for Molecular Dynamics Simulations on Commodity Clusters. In Proceedings of the ACM/IEEE SC 2006 Conference (SC’06), Tampa, FL, USA, 11–17 November 2006; IEEE: New York, NY, USA, 2007; p. 43. [Google Scholar]
- Toukmaji, A.Y.; Board, J.A. Ewald summation techniques in perspective: A survey. Comput. Phys. Commun. 1996, 95, 73–92. [Google Scholar] [CrossRef]
- Kadaoluwa Pathirannahalage, S.P.; Meftahi, N.; Elbourne, A.; Weiss, A.C.G.; McConville, C.F.; Padua, A.A.H.; Winkler, D.A.; Costa Gomes, M.F.; Greaves, T.L.; Le, T.C.; et al. Systematic Comparison of the Structural and Dynamic Properties of Commonly Used Water Models for Molecular Dynamics Simulations. J. Chem. Inf. Model. 2021, 61, 4521–4536. [Google Scholar] [CrossRef] [PubMed]
- Rey, J.; Murail, S.; de Vries, S.; Derreumaux, P.; Tuffery, P. PEP-FOLD4: A pH-dependent force field for peptide structure prediction in aqueous solution. Nucleic Acids Res. 2023, 51, W432–W437. [Google Scholar] [CrossRef] [PubMed]
- Selvam, R.; Subashchandran, K.P. Synthesis of Biologically Active Hydrophobic Peptide by Using Novel Polymer Support: Improved Fmoc Solid Phase Methodology. Int. J. Pept. Res. Ther. 2015, 21, 91–97. [Google Scholar] [CrossRef]
- Luna-Vázquez-Gómez, R.; Arellano-García, M.E.; García-Ramos, J.C.; Radilla-Chávez, P.; Salas-Vargas, D.S.; Casillas-Figueroa, F.; Ruiz-Ruiz, B.; Bogdanchikova, N.; Pestryakov, A. Hemolysis of human erythrocytes by argovitTM agnps from healthy and diabetic donors: An in vitro study. Materials 2021, 14, 2792. [Google Scholar] [CrossRef]
- Huligere, S.S.; Kumari, V.B.C.; Desai, S.M.; Wong, L.S.; Firdose, N.; Ramu, R. Investigating the antidiabetic efficacy of dairy-derived Lacticaseibacillus paracasei probiotic strains: Modulating α-amylase and α-glucosidase enzyme functions. Front. Microbiol. 2023, 14, 1288487. [Google Scholar] [CrossRef]
- Chandana Kumari, V.B.; Huligere, S.S.; Shbeer, A.M.; Ageel, M.; Jayanthi, M.K.; Jagadeep Chandra, S.; Ramu, R. Probiotic Potential Lacticaseibacillus casei and Limosilactobacillus fermentum Strains Isolated from Dosa Batter Inhibit α-Glucosidase and α-Amylase Enzymes. Microorganisms 2022, 10, 1195. [Google Scholar] [CrossRef]
- Pravin, N.J.; Kavalapure, R.S.; Gharge, S.; Alegaon, S.G.; Ranade, S.D.; Wong, L.S.; Ramu, R.; Ramya, C.M. Synthesis and In Vitro Biological Evaluation of Quinolone-Based Hydrazones as Potential Antidiabetic Agents Targeting Key Metabolic Enzymes. ACS Omega 2025, 10, 33712–33730. [Google Scholar] [CrossRef]
- Kokila, N.R.; Mahesh, B.; Ramu, R.; Divakara, S.G.; Mruthunjaya, K.; Raghav, N.; Boreddy Shivanandappa, T. Combined In Vitro and In Silico Approach to Define Alangimarckine from Thunbergia mysorensis Leaves as a Potential Inhibitor of α-Glucosidase. J. Biomol. Struct. Dyn. 2025, 43, 1–20. [Google Scholar] [CrossRef]
- Alharbi, O.; Alsaedi, W.H.; Alsehli, M.; Althagafi, S.H.; Alharbi, H.Y.; Asiri, Y.M.; Ramu, R.; Al-Ghorbani, M. Novel Quinoline- and Naphthalene-Incorporated Hydrazineylidene–Propenamide Analogues as Antidiabetic Agents: Design, Synthesis, and Computational Studies. Pharmaceuticals 2024, 17, 1692. [Google Scholar] [CrossRef]
- Lineweaver, H.; Burk, D. The Determination of Enzyme Dissociation Constants. J. Am. Chem. Soc. 1934, 56, 658–666. [Google Scholar] [CrossRef]
- Maradesha, T.; Patil, S.M.; Al-Mutairi, K.A.; Ramu, R.; Madhunapantula, S.V.; Alqadi, T. Inhibitory Effect of Polyphenols from Whole Green Jackfruit Flour against α-Glucosidase, α-Amylase, Aldose Reductase and Glycation at Multiple Stages and Their Interaction: Inhibition Kinetics and Molecular Simulations. Molecules 2022, 27, 1888. [Google Scholar] [CrossRef]
- Kokila, N.R.; Mahesh, B.; Ramu, R.; Roopashree, B.; Mruthunjaya, K. α-Amylase Inhibitory Potential of Thunbergia mysorensis Leaves Extract and Bioactive Compounds by In Vitro and Computational Approach. J. Biomol. Struct. Dyn. 2023, 41, 14887–14903. [Google Scholar] [CrossRef]
- Kwok, T.; Kirkpatrick, G.; Yusof, H.; Portokalakis, I.; Nigam, P.; Owusu-Apenten, R. Rapid Colorimetric Determination of Methylglyoxal Equivalents for Manuka Honey. J. Adv. Biol. Biotechnol. 2016, 7, 1–6. [Google Scholar] [CrossRef]
- Hussen, E.M.; Endalew, S.A. In vitro antioxidant and free-radical scavenging activities of polar leaf extracts of Vernonia amygdalina. BMC Complement. Med. Ther. 2023, 23, 146. [Google Scholar] [CrossRef] [PubMed]
- Kumar, M.S.; Raghavendra, T.O.; Shashank, M.; Ramu, R.; Mallikarjunaswamy, C.; Harini, R.; Alarifi, S.; Nagaraju, G.; Nirmala, B. Limonia acidissima Fruit Juice–Mediated Eco-Friendly Synthesis of Perovskite ZnSnO3 Nanoparticles: Applications to Photocatalytic, Electrochemical, Antioxidant and Antibacterial Activities. Inorg. Chem. Commun. 2024, 165, 112476. [Google Scholar] [CrossRef]
- Mallikarjunaswamy, C.; Lakshmi Ranganatha, V.; Ramu, R.; Udayabhanu; Nagaraju, G. Facile Microwave-Assisted Green Synthesis of ZnO Nanoparticles: Application to Photodegradation, Antibacterial and Antioxidant. J. Mater. Sci. Mater. Electron. 2020, 31, 1004–1021. [Google Scholar] [CrossRef]
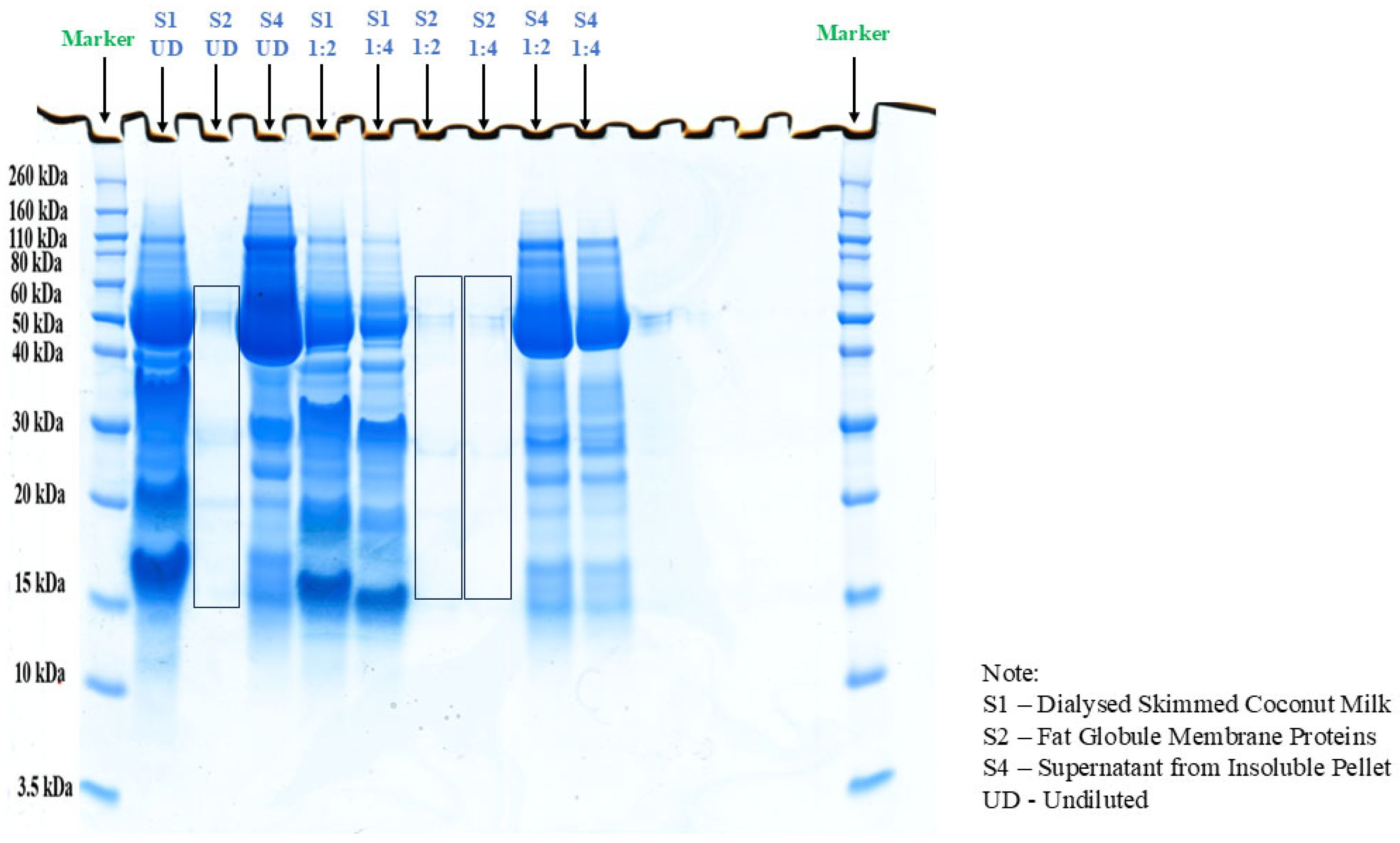
| Samples | Concentration of Protein (µg/mL) | Concentration of Protein (g/100 g) |
|---|---|---|
| Skimmed coconut milk after dialysis | 369 | 0.0365 |
| Crude fat globule membrane (FGM) suspension | 109 | 0.0108 |
| Supernatant from insoluble pellet | 1574 | 0.1558 |
| Peptide/ Standard Drug | Anti-Diabetic Targets | Receptor for Advanced Glycation End-Products (RAGE) | ||||||
|---|---|---|---|---|---|---|---|---|
| α-Glucosidase | α-Amylase | Aldose Reductase | ||||||
| Glide Score (kcal/mol) | THB | Glide Score (kcal/mol) | THB | Glide Score (kcal/mol) | THB | Glide Score (kcal/mol) | THB | |
| Peptide 1 | −9.34 | 3 | −9.64 | 4 | −11.42 | 4 | - | - |
| Peptide 2 | −9.87 | 3 | −9.87 | 6 | −9.54 | 3 | −8.75 | 6 |
| * Acarbose | −12.33 | 7 | −11.40 | 6 | - | - | - | - |
| # Quercetin | - | - | - | - | −10.767 | 3 | - | - |
| & Papaverine | - | - | - | - | - | - | −3.74 | 1 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Naganarasimha, A.S.; Patil, S.M.; Ramu, R.; Przybyłek, M.; Bełdowski, P.; Małolepsza, O.; Bujanowski, S.; Shahid, M. Coconut Milk-Derived Bioactive Peptides as Multifunctional Agents Against Hyperglycemia, Oxidative Stress, and Glycation: An Integrated Experimental and Computational Study. Int. J. Mol. Sci. 2026, 27, 360. https://doi.org/10.3390/ijms27010360
Naganarasimha AS, Patil SM, Ramu R, Przybyłek M, Bełdowski P, Małolepsza O, Bujanowski S, Shahid M. Coconut Milk-Derived Bioactive Peptides as Multifunctional Agents Against Hyperglycemia, Oxidative Stress, and Glycation: An Integrated Experimental and Computational Study. International Journal of Molecular Sciences. 2026; 27(1):360. https://doi.org/10.3390/ijms27010360
Chicago/Turabian StyleNaganarasimha, Akshaya Simha, Shashank M. Patil, Ramith Ramu, Maciej Przybyłek, Piotr Bełdowski, Olga Małolepsza, Sławomir Bujanowski, and Mudassar Shahid. 2026. "Coconut Milk-Derived Bioactive Peptides as Multifunctional Agents Against Hyperglycemia, Oxidative Stress, and Glycation: An Integrated Experimental and Computational Study" International Journal of Molecular Sciences 27, no. 1: 360. https://doi.org/10.3390/ijms27010360
APA StyleNaganarasimha, A. S., Patil, S. M., Ramu, R., Przybyłek, M., Bełdowski, P., Małolepsza, O., Bujanowski, S., & Shahid, M. (2026). Coconut Milk-Derived Bioactive Peptides as Multifunctional Agents Against Hyperglycemia, Oxidative Stress, and Glycation: An Integrated Experimental and Computational Study. International Journal of Molecular Sciences, 27(1), 360. https://doi.org/10.3390/ijms27010360









