Serratia marcescens: A Versatile Opportunistic Pathogen with Emerging Clinical and Biotechnological Significance
Abstract
1. Introduction
2. Taxonomy and General Characteristics
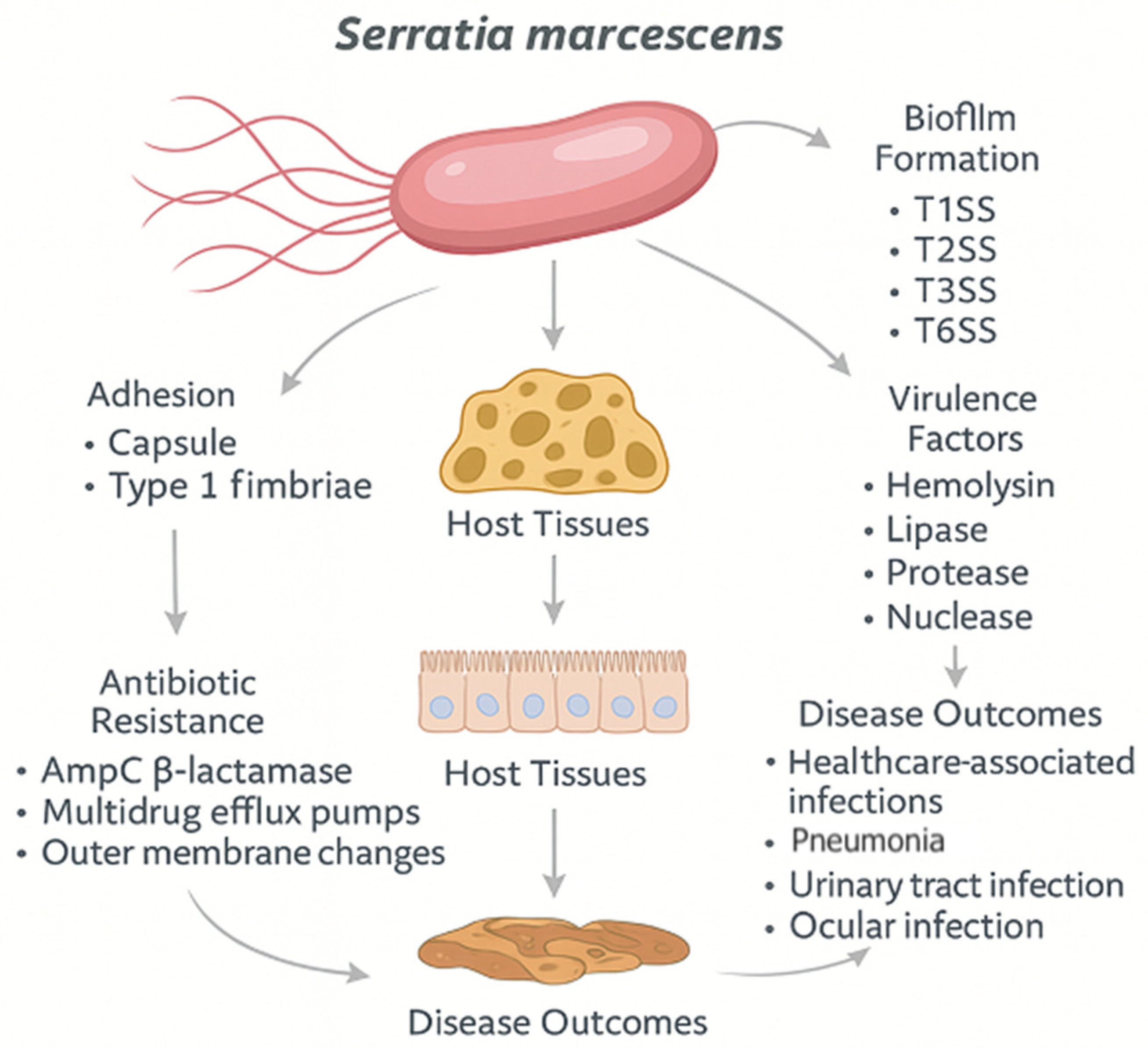
3. Virulence Factors
- Proteases (e.g., serralysin): Degrade host tissue components and immune mediators such as immunoglobulins and complement proteins.
- Lipases and phospholipases: Involved in membrane disruption and nutrient liberation.
- DNases and nucleases: Facilitate evasion of neutrophil extracellular traps (NETs).
- Type I Secretion System (T1SS): Transports hemolysin (ShlA).
- Type III Secretion System (T3SS): Injects effector proteins into host cells (limited distribution in clinical strains).
- Type IV Secretion System (T4SS): Implicated in horizontal gene transfer and virulence gene dissemination.
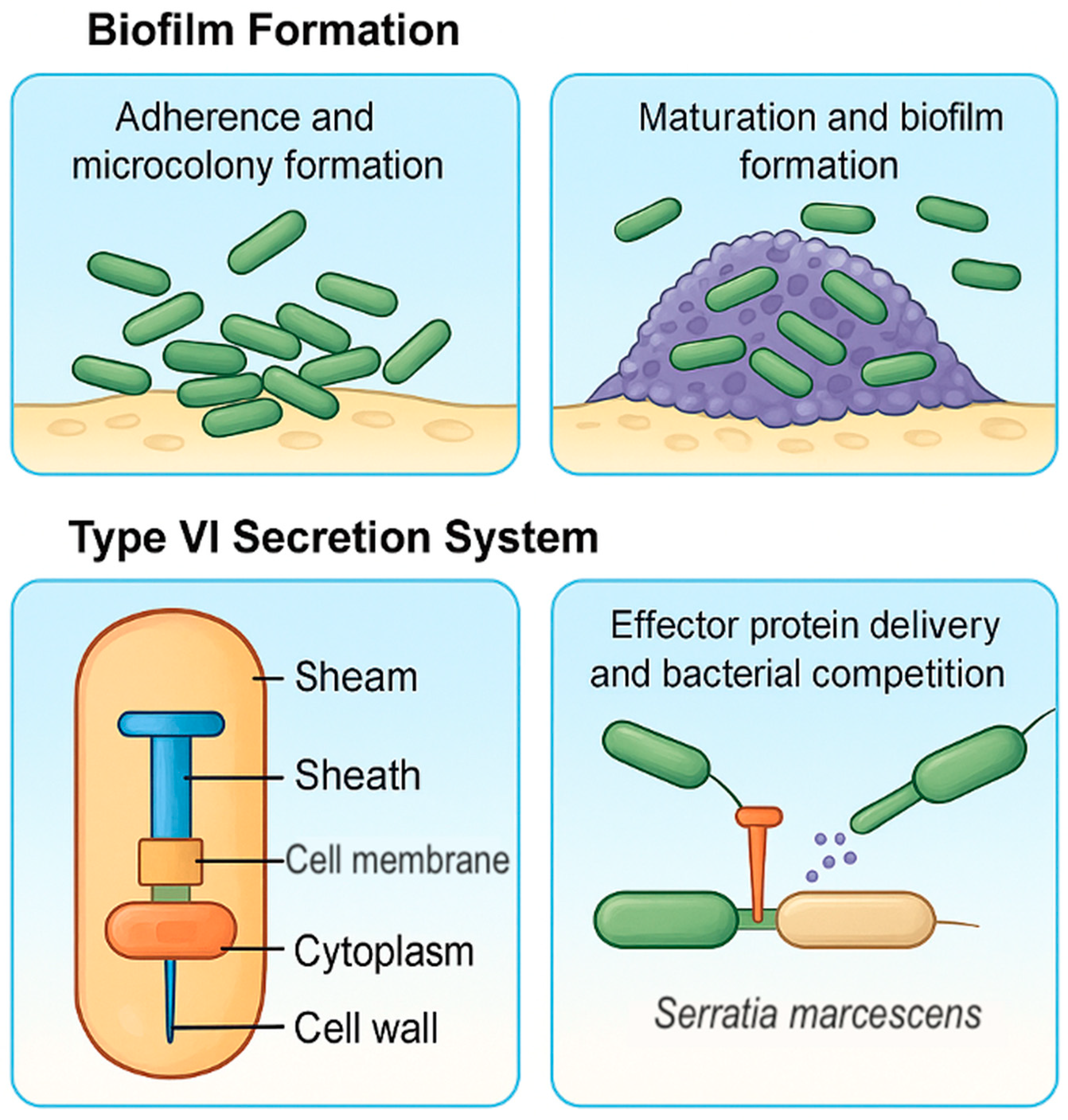
- Type VI Secretion System (T6SS): Functions in bacterial competition and delivery of toxins to eukaryotic cells. The T6SS is particularly active in biofilm and polymicrobial environments. The Type VI Secretion System is especially crucial in interbacterial antagonism, particularly in polymicrobial environments. T6SS in S. marcescens is encoded by hcp/vgrG loci and is closely regulated by quorum sensing and stress responses [48].
4. Biofilm Formation
- Initial attachment—mediated by flagella, fimbriae, and surface adhesins.
- Irreversible attachment—involving enhanced EPS production and expression of specific biofilm genes.
- Maturation I and II—3D architecture develops, with water channels and nutrient gradients.
- Dispersion—cells detach and colonize new surfaces.
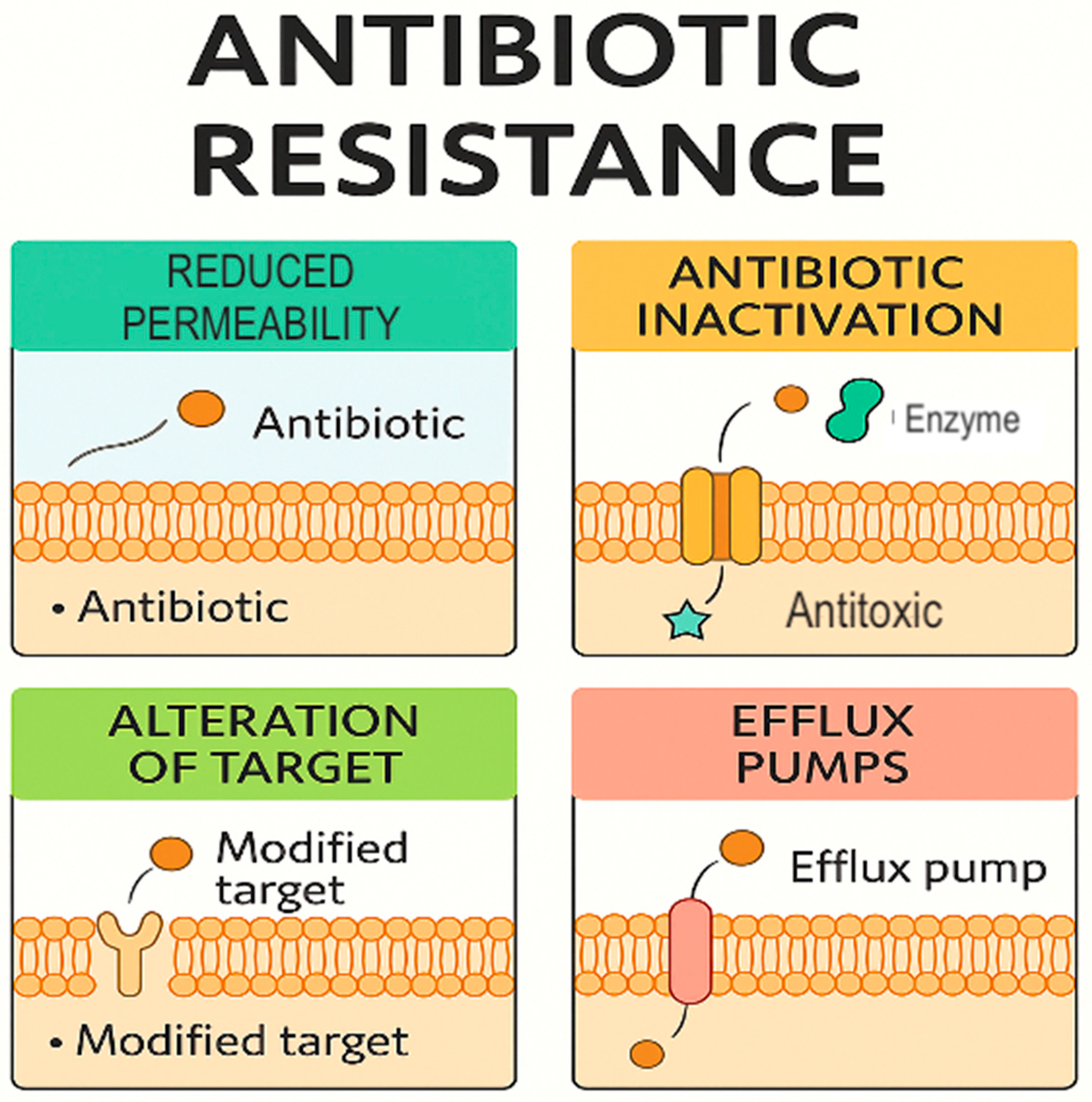
5. Antibiotic Resistance
- Extended-spectrum β-lactamases (ESBLs)
- Carbapenemases, e.g., KPC, NDM, VIM;
- Aminoglycoside-modifying enzymes;
- Fluoroquinolone resistance.
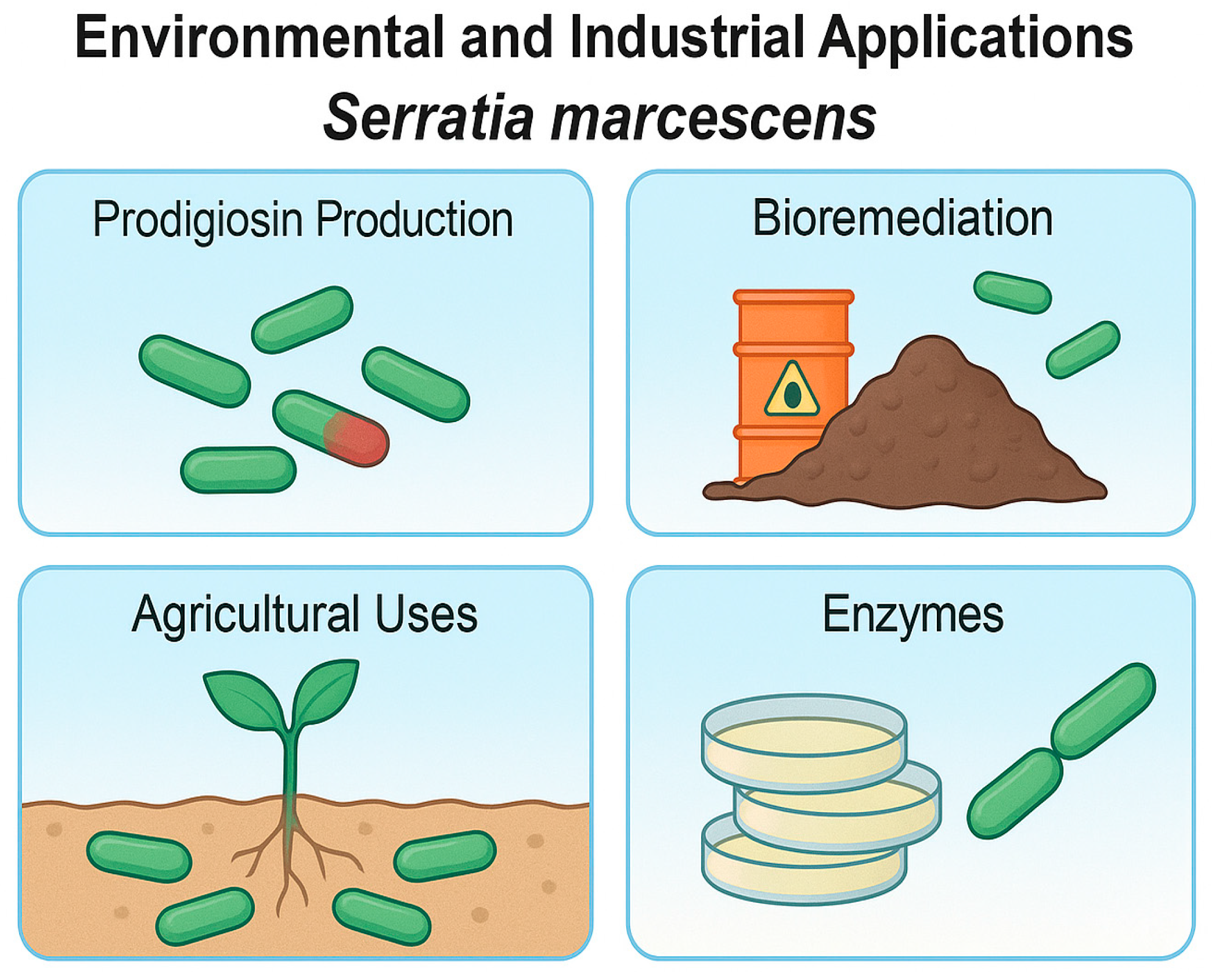
6. Environmental and Industrial Aspects. Industrially Relevant Enzymes from S. marcescens
7. Clinical Applications and Innovative Therapies
- Bacteriophage Therapy: Specific lytic phages targeting S. marcescens have shown effectiveness in disrupting biofilms and working together with antibiotics. Experimental and compassionate-use cases demonstrate clinical benefits in treating difficult-to-treat device-related infections and chronic wounds [97].
- Phage-Antibiotic Synergy (PAS): Using phages with sub-inhibitory amounts of antibiotics (such as ciprofloxacin, cefepime) can improve bacterial elimination and slow resistance development. This synergy is especially useful in treating biofilm-related infections where monotherapy often fails [98].
- CRISPR (Clustered Regularly Interspaced Short Palindromic Repeats) and Cas (CRISPR-associated proteins): Synthetic CRISPR constructs delivered through conjugative plasmids or engineered phages are being tested to selectively eliminate resistance genes in Enterobacterales. Although not yet approved for clinical use, preclinical studies have shown success in making MDR strains of S. marcescens more sensitive to standard antibiotics [99].
- Nanoparticle-Based Delivery: Nanocarriers, such as liposomes, dendrimers, and silver nanoparticles, can be utilized to deliver antibiotics or phages directly to biofilm-embedded S. marcescens, thereby increasing local drug concentration and overcoming matrix barriers [101].
8. Current Challenges and Future Directions
9. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| HAIs | healthcare-associated infections |
| ICUs | intensive care units |
| MDR | multidrug-resistant |
| XDR | extensively drug-resistant |
| NETs | neutrophil extracellular traps |
| T1SS | Type I Secretion System |
| T3SS | Type III Secretion System |
| T4SS | Type IV Secretion System |
| T6SS | Type VI Secretion System |
| OMPs | outer membrane proteins |
| AHLs | N-acyl homoserine lactones |
| EPS | extracellular polymeric substance |
| QS | quorum sensing |
| eDNA | extracellular DNA |
| ESBLs | Extended-spectrum β-lactamases |
| CRISPR | Clustered Regularly Interspaced Short Palindromic Repeats |
| CDC | Disease Control and Prevention |
| WHO | World Health Organization |
References
- Hejazi, A.; Falkiner, F.R. Serratia marcescens. J. Med. Microbiol. 1997, 46, 903–912. [Google Scholar] [CrossRef] [PubMed]
- Khanna, A.; Khanna, M.; Aggarwal, A. Serratia marcescens- a rare opportunistic nosocomial pathogen and measures to limit its spread in hospitalized patients. J. Clin. Diagn. Res. 2013, 7, 243–246. [Google Scholar] [CrossRef]
- Serratia marcescens. Available online: https://en.wikipedia.org/wiki/Serratia_marcescens (accessed on 15 September 2025).
- Tavares-Carreon, F.; De Anda-Mora, K.; Rojas-Barrera, I.C.; Andrade, A. Serratia marcescens antibiotic resistance mechanisms of an opportunistic pathogen: A literature review. PeerJ 2023, 11, e14399. [Google Scholar] [CrossRef]
- Traub, W.H. Antibiotic Susceptibility of Serratia marcescens and Serratia liquefaciens. Chemotherapy 2000, 46, 315–321. [Google Scholar] [CrossRef]
- Cristina, M.L.; Sartini, M.; Spagnolo, A.M. Serratia marcescens Infections in Neonatal Intensive Care Units (NICUs). Int. J. Environ. Res. Public Health 2019, 16, 610. [Google Scholar] [CrossRef]
- Zhu, H.; Li, F.; Cao, X.; Zhang, Y.; Liu, C.; Chen, Y.; Shen, H. Epidemiology, resistance profiles, and risk factors of multidrug- and carbapenem-resistant Serratia marcescens infections: A retrospective study of 242 cases. BMC Infect. Dis. 2025, 25, 1105. [Google Scholar] [CrossRef]
- Darshan, N.; Manonmani, H.K. Prodigiosin and its potential applications. J. Food Sci. Technol. 2015, 52, 5393–5407. [Google Scholar] [CrossRef]
- Islan, G.A.; Rodenak-Kladniew, B.; Noacco, N.; Duran, N.; Castro, G.R. Prodigiosin: A promising biomolecule with many potential biomedical applications. Bioengineered 2022, 13, 14227–14258. [Google Scholar] [CrossRef] [PubMed]
- Cosimato, I.; Santella, B.; Rufolo, S.; Sabatini, P.; Galdiero, M.; Capunzo, M.; Boccia, G.; Folliero, V.; Franci, G. Current Epidemiological Status and Antibiotic Resistance Profile of Serratia marcescens. Antibiotics 2024, 13, 323. [Google Scholar] [CrossRef]
- Bartolomeo Bizio. Available online: https://en.wikipedia.org/wiki/Bartolomeo_Bizio (accessed on 15 September 2025).
- Nazzaro, G. Etymologia: Serratia marcescens. Emerg. Infect. Dis. 2019, 25, 2012. [Google Scholar] [CrossRef]
- THE TAXONOMIC STATUS OF SERRATIA MARCESCENS BIZIO. Available online: https://scispace.com/pdf/the-taxonomic-status-of-serratia-marcescens-bizio-3et58oputa.pdf (accessed on 15 September 2025).
- Species Serratia marcescens. Available online: https://lpsn.dsmz.de/species/serratia-marcescens (accessed on 15 September 2025).
- Family Yersiniaceae. Available online: https://lpsn.dsmz.de/family/yersiniaceae (accessed on 15 September 2025).
- Janda, J.M.; Abbott, S.L. The Changing Face of the Family Enterobacteriaceae (Order: “Enterobacterales”): New Members, Taxonomic Issues, Geographic Expansion, and New Diseases and Disease Syndromes. Clin. Microbiol. Rev. 2021, 34, e00174-20. [Google Scholar] [CrossRef] [PubMed]
- Serratia marcescens. Available online: https://micropspbgmu.ru/downloads/Serratia%20marcescens.pdf (accessed on 15 September 2025).
- Gupta, V.; Sharma, S.; Pal, K.; Goyal, P.; Agarwal, D.; Chander, J. Serratia, No Longer an Uncommon Opportunistic Pathogen—Case Series & Review of Literature. Infect. Disord. Drug Targets 2021, 21, e300821191666. [Google Scholar] [CrossRef]
- Grimont, F.; Grimont, P.A.D. The Genus Serratia. In The Prokaryotes: A Handbook on the Biology of Bacteria Volume 6: Proteobacteria: Gamma Subclass; Dworkin, M., Falkow, S., Rosenberg, E., Schleifer, K.-H., Stackebrandt, E., Eds.; Springer New York: New York, NY, USA, 2006; pp. 219–244. [Google Scholar]
- Romanowski, E.G.; Lehner, K.M.; Martin, N.C.; Patel, K.R.; Callaghan, J.D.; Stella, N.A.; Shanks, R.M.Q. Thermoregulation of Prodigiosin Biosynthesis by Serratia marcescens is Controlled at the Transcriptional Level and Requires HexS. Pol. J. Microbiol. 2019, 68, 43–50. [Google Scholar] [CrossRef]
- Shimuta, K.; Ohnishi, M.; Iyoda, S.; Gotoh, N.; Koizumi, N.; Watanabe, H. The hemolytic and cytolytic activities of Serratia marcescens phospholipase A (PhlA) depend on lysophospholipid production by PhlA. BMC Microbiol. 2009, 9, 261. [Google Scholar] [CrossRef]
- Roy, P.; Ahmed, N.H.; Grover, R.K. Non-pigmented strain of serratia marcescens: An unusual pathogen causing pulmonary infection in a patient with malignancy. J. Clin. Diagn. Res. 2014, 8, Dd05–Dd06. [Google Scholar] [CrossRef]
- Mahlen, S.D. Serratia Infections: From Military Experiments to Current Practice. Clin. Microbiol. Rev. 2011, 24, 755–791. [Google Scholar] [CrossRef]
- Pérez-Viso, B.; Aracil-Gisbert, S.; Coque, T.M.; Del Campo, R.; Ruiz-Garbajosa, P.; Cantón, R. Evaluation of CHROMagar™-Serratia agar, a new chromogenic medium for the detection and isolation of Serratia marcescens. Eur. J. Clin. Microbiol. Infect. Dis. 2021, 40, 2593–2596. [Google Scholar] [CrossRef]
- Pramanik, A.; Könninger, U.; Selvam, A.; Braun, V. Secretion and activation of the Serratia marcescens hemolysin by structurally defined ShlB mutants. Int. J. Med. Microbiol. 2014, 304, 351–359. [Google Scholar] [CrossRef] [PubMed]
- Walker, G.; Hertle, R.; Braun, V. Activation of Serratia marcescens hemolysin through a conformational change. Infect. Immun. 2004, 72, 611–614. [Google Scholar] [CrossRef]
- Guérin, J.; Bigot, S.; Schneider, R.; Buchanan, S.K.; Jacob-Dubuisson, F. Two-Partner Secretion: Combining Efficiency and Simplicity in the Secretion of Large Proteins for Bacteria-Host and Bacteria-Bacteria Interactions. Front. Cell. Infect. Microbiol. 2017, 7, 148. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Morrison, A.J.; Thibodeau, P.H. Interdomain Contacts and the Stability of Serralysin Protease from Serratia marcescens. PLoS ONE 2015, 10, e0138419. [Google Scholar] [CrossRef]
- Kida, Y.; Inoue, H.; Shimizu, T.; Kuwano, K. Serratia marcescens serralysin induces inflammatory responses through protease-activated receptor 2. Infect. Immun. 2007, 75, 164–174. [Google Scholar] [CrossRef]
- Weakland, D.R.; Smith, S.N.; Bell, B.; Tripathi, A.; Mobley, H.L.T. The Serratia marcescens Siderophore Serratiochelin Is Necessary for Full Virulence during Bloodstream Infection. Infect. Immun. 2020, 88, e00117-20. [Google Scholar] [CrossRef]
- Kalivoda, E.J.; Stella, N.A.; O’Dee, D.M.; Nau, G.J.; Shanks, R.M. The cyclic AMP-dependent catabolite repression system of Serratia marcescens mediates biofilm formation through regulation of type 1 fimbriae. Appl. Environ. Microbiol. 2008, 74, 3461–3470. [Google Scholar] [CrossRef]
- Kalivoda, E.J.; Brothers, K.M.; Stella, N.A.; Schmitt, M.J.; Shanks, R.M. Bacterial cyclic AMP-phosphodiesterase activity coordinates biofilm formation. PLoS ONE 2013, 8, e71267. [Google Scholar] [CrossRef]
- Shanks, R.; Lahr, R.; Stella, N.; Arena, K.; Brothers, K.; Kwak, D.; Liu, X.; Kalivoda, E. A Serratia marcescens PigP Homolog Controls Prodigiosin Biosynthesis, Swarming Motility and Hemolysis and Is Regulated by cAMP-CRP and HexS. PLoS ONE 2013, 8, e57634. [Google Scholar] [CrossRef] [PubMed]
- Shanks, R.M.; Stella, N.A.; Lahr, R.M.; Wang, S.; Veverka, T.I.; Kowalski, R.P.; Liu, X. Serratamolide is a hemolytic factor produced by Serratia marcescens. PLoS ONE 2012, 7, e36398. [Google Scholar] [CrossRef] [PubMed]
- Mariano, G.; Trunk, K.; Williams, D.J.; Monlezun, L.; Strahl, H.; Pitt, S.J.; Coulthurst, S.J. A family of Type VI secretion system effector proteins that form ion-selective pores. Nat. Commun. 2019, 10, 5484. [Google Scholar] [CrossRef]
- Reglinski, M.; Hurst, Q.W.; Williams, D.J.; Gierlinski, M.; Şahin, A.T.; Mathers, K.; Ostrowski, A.; Bergkessel, M.; Zachariae, U.; Pitt, S.J.; et al. A widely-occurring family of pore-forming effectors broadens the impact of the Serratia Type VI secretion system. Embo. J. 2025; advance online publication. [Google Scholar] [CrossRef] [PubMed]
- Labbate, M.; Zhu, H.; Thung, L.; Bandara, R.; Larsen, M.R.; Willcox, M.D.; Givskov, M.; Rice, S.A.; Kjelleberg, S. Quorum-sensing regulation of adhesion in Serratia marcescens MG1 is surface dependent. J. Bacteriol. 2007, 189, 2702–2711. [Google Scholar] [CrossRef]
- Molla, A.; Matsumoto, K.; Oyamada, I.; Katsuki, T.; Maeda, H. Degradation of protease inhibitors, immunoglobulins, and other serum proteins by Serratia protease and its toxicity to fibroblast in culture. Infect. Immun. 1986, 53, 522–529. [Google Scholar] [CrossRef]
- Molla, A.; Akaike, T.; Maeda, H. Inactivation of various proteinase inhibitors and the complement system in human plasma by the 56-kilodalton proteinase from Serratia marcescens. Infect. Immun. 1989, 57, 1868–1871. [Google Scholar] [CrossRef] [PubMed]
- Sharma, P.; Garg, N.; Sharma, A.; Capalash, N.; Singh, R. Nucleases of bacterial pathogens as virulence factors, therapeutic targets and diagnostic markers. Int. J. Med. Microbiol. 2019, 309, 151354. [Google Scholar] [CrossRef] [PubMed]
- Benedik, M.J.; Strych, U. Serratia marcescens and its extracellular nuclease. FEMS Microbiol. Lett. 1998, 165, 1–13. [Google Scholar] [CrossRef]
- Nestle, M.; Roberts, W.K. An extracellular nuclease from Serratia marcescens. II. Specificity of the enzyme. J. Biol. Chem. 1969, 244, 5219–5225. [Google Scholar] [CrossRef] [PubMed]
- Rice, S.A.; Koh, K.S.; Queck, S.Y.; Labbate, M.; Lam, K.W.; Kjelleberg, S. Biofilm Formation and Sloughing in Serratia marcescens Are Controlled by Quorum Sensing and Nutrient Cues. J. Bacteriol. 2005, 187, 3477–3485. [Google Scholar] [CrossRef]
- Li, P.; Kwok, A.H.Y.; Jiang, J.; Ran, T.; Xu, D.; Wang, W.; Leung, F.C.C. Comparative genome analyses of Serratia marcescens FS14 reveal its high antagonistic potential. PLoS ONE 2015, 10, e0123061. [Google Scholar] [CrossRef]
- Murdoch, S.L.; Trunk, K.; English, G.; Fritsch, M.J.; Pourkarimi, E.; Coulthurst, S.J. The opportunistic pathogen Serratia marcescens utilizes type VI secretion to target bacterial competitors. J. Bacteriol. 2011, 193, 6057–6069. [Google Scholar] [CrossRef]
- Alvarez-Martinez, C.E.; Christie, P.J. Biological diversity of prokaryotic type IV secretion systems. Microbiol. Mol. Biol. Rev. 2009, 73, 775–808. [Google Scholar] [CrossRef]
- English, G.; Trunk, K.; Rao, V.A.; Srikannathasan, V.; Hunter, W.N.; Coulthurst, S.J. New secreted toxins and immunity proteins encoded within the Type VI secretion system gene cluster of Serratia marcescens. Mol. Microbiol. 2012, 86, 921–936. [Google Scholar] [CrossRef]
- Hespanhol, J.T.; Nóbrega-Silva, L.; Bayer-Santos, E. Regulation of type VI secretion systems at the transcriptional, posttranscriptional and posttranslational level. Microbiology 2023, 169, 001376. [Google Scholar] [CrossRef]
- Shanks, R.M.Q.; Stella, N.A.; Kalivoda, E.J.; Doe, M.R.; O’Dee, D.M.; Lathrop, K.L.; Guo, F.L.; Nau, G.J. A Serratia marcescens OxyR Homolog Mediates Surface Attachment and Biofilm Formation. J. Bacteriol. 2007, 189, 7262–7272. [Google Scholar] [CrossRef]
- Khilyas, I.V.; Shirshikova, T.V.; Matrosova, L.E.; Sorokina, A.V.; Sharipova, M.R.; Bogomolnaya, L.M. Production of Siderophores by Serratia marcescens and the Role of MacAB Efflux Pump in Siderophores Secretion. BioNanoScience 2016, 6, 480–482. [Google Scholar] [CrossRef]
- Fekrirad, Z.; Gattali, B.; Kashef, N. Quorum sensing-regulated functions of Serratia marcescens are reduced by eugenol. Iran. J. Microbiol. 2020, 12, 451–459. [Google Scholar] [CrossRef] [PubMed]
- Donlan, R.M.; Costerton, J.W. Biofilms: Survival Mechanisms of Clinically Relevant Microorganisms. Clin. Microbiol. Rev. 2002, 15, 167–193. [Google Scholar] [CrossRef]
- Trautner, B.W.; Darouiche, R.O. Role of biofilm in catheter-associated urinary tract infection. Am. J. Infect. Control 2004, 32, 177–183. [Google Scholar] [CrossRef]
- Liu, C.; Sun, D.; Zhu, J.; Liu, J.; Liu, W. The Regulation of Bacterial Biofilm Formation by cAMP-CRP: A Mini-Review. Front. Microbiol. 2020, 11, 802. [Google Scholar] [CrossRef] [PubMed]
- Tang, L.; Schramm, A.; Neu, T.R.; Revsbech, N.P.; Meyer, R.L. Extracellular DNA in adhesion and biofilm formation of four environmental isolates: A quantitative study. FEMS Microbiol. Ecol. 2013, 86, 394–403. [Google Scholar] [CrossRef]
- Stella, N.A.; Romanowski, E.G.; Brothers, K.M.; Calvario, R.C.; Shanks, R.M.Q. IgaA Protein, GumB, Has a Global Impact on the Transcriptome and Surface Proteome of Serratia marcescens. Infect. Immun. 2022, 90, e0039922. [Google Scholar] [CrossRef] [PubMed]
- Tuttobene, M.R.; Bruna, R.E.; Molino, M.V.; García Véscovi, E. PrtA-mediated flagellar turnover is essential for robust biofilm development in Serratia marcescens. Appl. Environ. Microbiol. 2025, 91, e0126125. [Google Scholar] [CrossRef]
- Silva, U.C.M.; da Silva, D.R.C.; Cuadros-Orellana, S.; Moreira, L.M.; Leite, L.R.; Medeiros, J.D.; Felestrino, E.B.; Caneschi, W.L.; Almeida, N.F.; Silva, R.S.; et al. Genomic and phenotypic insights into Serratia interaction with plants from an ecological perspective. Braz. J. Microbiol. 2025, 56, 1219–1239. [Google Scholar] [CrossRef] [PubMed]
- Karaguler, T.; Kahraman, H.; Tuter, M. Analyzing effects of ELF electromagnetic fields on removing bacterial biofilm. Biocybern. Biomed. Eng. 2017, 37, 336–340. [Google Scholar] [CrossRef]
- Olsen, I. Biofilm-specific antibiotic tolerance and resistance. Eur. J. Clin. Microbiol. Infect. Dis. 2015, 34, 877–886. [Google Scholar] [CrossRef] [PubMed]
- Ringel, P.D.; Hu, D.; Basler, M. The Role of Type VI Secretion System Effectors in Target Cell Lysis and Subsequent Horizontal Gene Transfer. Cell Rep. 2017, 21, 3927–3940. [Google Scholar] [CrossRef]
- Coulthurst, S. The Type VI secretion system: A versatile bacterial weapon. Microbiology 2019, 165, 503–515. [Google Scholar] [CrossRef]
- Alcoforado Diniz, J.; Coulthurst, S.J. Intraspecies Competition in Serratia marcescens Is Mediated by Type VI-Secreted Rhs Effectors and a Conserved Effector-Associated Accessory Protein. J. Bacteriol. 2015, 197, 2350–2360. [Google Scholar] [CrossRef]
- Jones, R.N.; Baquero, F.; Privitera, G.; Inoue, M.; Wiedemann, B. Inducible β-lactamase-mediated resistance to third-generation cephalosporins. Clin. Microbiol. Infect. 1997, 3, S7–S20. [Google Scholar] [CrossRef][Green Version]
- Tamma, P.D.; Doi, Y.; Bonomo, R.A.; Johnson, J.K.; Simner, P.J. A Primer on AmpC β-Lactamases: Necessary Knowledge for an Increasingly Multidrug-resistant World. Clin. Infect. Dis. 2019, 69, 1446–1455. [Google Scholar] [CrossRef]
- Gogry, F.A.; Siddiqui, M.T.; Sultan, I.; Haq, Q.M.R. Current Update on Intrinsic and Acquired Colistin Resistance Mechanisms in Bacteria. Front. Med. 2021, 8, 677720. [Google Scholar] [CrossRef]
- Maseda, H.; Hashida, Y.; Konaka, R.; Shirai, A.; Kourai, H. Mutational upregulation of a resistance-nodulation-cell division-type multidrug efflux pump, SdeAB, upon exposure to a biocide, cetylpyridinium chloride, and antibiotic resistance in Serratia marcescens. Antimicrob. Agents Chemother. 2009, 53, 5230–5235. [Google Scholar] [CrossRef]
- Toba, S.; Minato, Y.; Kondo, Y.; Hoshikawa, K.; Minagawa, S.; Komaki, S.; Kumagai, T.; Matoba, Y.; Morita, D.; Ogawa, W.; et al. Comprehensive analysis of resistance-nodulation-cell division superfamily (RND) efflux pumps from Serratia marcescens, Db10. Sci. Rep. 2019, 9, 4854. [Google Scholar] [CrossRef] [PubMed]
- Shirshikova, T.V.; Sierra-Bakhshi, C.G.; Kamaletdinova, L.K.; Matrosova, L.E.; Khabipova, N.N.; Evtugyn, V.G.; Khilyas, I.V.; Danilova, I.V.; Mardanova, A.M.; Sharipova, M.R.; et al. The ABC-Type Efflux Pump MacAB Is Involved in Protection of Serratia marcescens against Aminoglycoside Antibiotics, Polymyxins, and Oxidative Stress. mSphere 2021, 6, e00033-21. [Google Scholar] [CrossRef]
- Zhou, G.; Wang, Q.; Wang, Y.; Wen, X.; Peng, H.; Peng, R.; Shi, Q.; Xie, X.; Li, L. Outer Membrane Porins Contribute to Antimicrobial Resistance in Gram-Negative Bacteria. Microorganisms 2023, 11, 1690. [Google Scholar] [CrossRef]
- Drawz, S.M.; Bonomo, R.A. Three Decades of β-Lactamase Inhibitors. Clin. Microbiol. Rev. 2010, 23, 160–201. [Google Scholar] [CrossRef]
- Reygaert, W.C. An overview of the antimicrobial resistance mechanisms of bacteria. AIMS Microbiol. 2018, 4, 482–501. [Google Scholar] [CrossRef]
- Galgano, M.; Pellegrini, F.; Catalano, E.; Capozzi, L.; Del Sambro, L.; Sposato, A.; Lucente, M.S.; Vasinioti, V.I.; Catella, C.; Odigie, A.E.; et al. Acquired Bacterial Resistance to Antibiotics and Resistance Genes: From Past to Future. Antibiotics 2025, 14, 222. [Google Scholar] [CrossRef] [PubMed]
- Cruz-López, F.; Villarreal-Treviño, L.; Camacho-Ortiz, A.; Morfín-Otero, R.; Flores-Treviño, S.; Garza-González, E. Acquired Genetic Elements that Contribute to Antimicrobial Resistance in Frequent Gram-Negative Causative Agents of Healthcare-Associated Infections. Am. J. Med. Sci. 2020, 360, 631–640. [Google Scholar] [CrossRef]
- Partridge, S.R.; Kwong, S.M.; Firth, N.; Jensen, S.O. Mobile Genetic Elements Associated with Antimicrobial Resistance. Clin. Microbiol. Rev. 2018, 31, e00088-17. [Google Scholar] [CrossRef]
- Prioritization of Pathogens to Guide Discovery, Research and Development of New Antibiotics for Drug-Resistant Bacterial Infections, Including Tuberculosis. Available online: https://www.who.int/publications/i/item/WHO-EMP-IAU-2017.12 (accessed on 15 September 2025).
- Olaitan, A.O.; Morand, S.; Rolain, J.-M. Mechanisms of polymyxin resistance: Acquired and intrinsic resistance in bacteria. Front. Microbiol. 2014, 5, 643. [Google Scholar] [CrossRef]
- Kumar, S.; Bhattacharya, S. Optimization of Serratiopeptidase Production from Serratia marcescens SP6 Using Sequential Strategy of Experimental Designs. Curr. Microbiol. 2025, 82, 371. [Google Scholar] [CrossRef] [PubMed]
- Guryanov, I.; Naumenko, E. Bacterial Pigment Prodigiosin as Multifaceted Compound for Medical and Industrial Application. Appl. Microbiol. 2024, 4, 1702–1728. [Google Scholar] [CrossRef]
- Kulkova, I.; Wróbel, B.; Dobrzyński, J. Serratia spp. as plant growth-promoting bacteria alleviating salinity, drought, and nutrient imbalance stresses. Front. Microbiol. 2024, 15, 1342331. [Google Scholar] [CrossRef]
- Peekate, L.; Ogolo, J. The potential of Serratia marcescens in Bioremediation of Crude-oil Polluted Soil. UMYU J. Microbiol. Res. (UJMR) 2024, 9, 75–83. [Google Scholar] [CrossRef]
- The Use of Crude Shrimp Shell Powder for Chitinase Production by Serratia marcescens WF. Available online: https://www.ftb.com.hr/images/pdfarticles/2006/January-March/44-95.pdf (accessed on 15 September 2025).
- Suzuki, K.; Taiyoji, M.; Sugawara, N.; Nikaidou, N.; Henrissat, B.; Watanabe, T. The third chitinase gene (chiC) of Serratia marcescens 2170 and the relationship of its product to other bacterial chitinases. Biochem J. 1999, 343 Pt 3, 587–596. [Google Scholar] [CrossRef]
- Wang, K.; Yan, P.S.; Cao, L.X. Chitinase from a novel strain of Serratia marcescens JPP1 for biocontrol of aflatoxin: Molecular characterization and production optimization using response surface methodology. Biomed. Res. Int. 2014, 2014, 482623. [Google Scholar] [CrossRef]
- El-Sayed, G.M.; Emam, M.T.H.; Hammad, M.A.; Mahmoud, S.H. Gene Cloning, Heterologous Expression, and In Silico Analysis of Chitinase B from Serratia marcescens for Biocontrol of Spodoptera frugiperda Larvae Infesting Maize Crops. Molecules 2024, 29, 1466. [Google Scholar] [CrossRef] [PubMed]
- Ren, X.-B.; Dang, Y.-R.; Liu, S.-S.; Huang, K.-X.; Qin, Q.-L.; Chen, X.-L.; Zhang, Y.-Z.; Wang, Y.-J.; Li, P.-Y. Identification and Characterization of Three Chitinases with Potential in Direct Conversion of Crystalline Chitin into N,N′-diacetylchitobiose. Mar. Drugs 2022, 20, 165. [Google Scholar] [CrossRef]
- Nair, S.R.; C, S.D. Serratiopeptidase: An integrated View of Multifaceted Therapeutic Enzyme. Biomolecules 2022, 12, 1468. [Google Scholar] [CrossRef] [PubMed]
- Jadhav, S.B.; Shah, N.; Rathi, A.; Rathi, V.; Rathi, A. Serratiopeptidase: Insights into the therapeutic applications. Biotechnol. Rep. 2020, 28, e00544. [Google Scholar] [CrossRef] [PubMed]
- Srivastava, V.; Mishra, S.; Chaudhuri, T.K. Enhanced production of recombinant serratiopeptidase in Escherichia coli and its characterization as a potential biosimilar to native biotherapeutic counterpart. Microb. Cell Factories 2019, 18, 215. [Google Scholar] [CrossRef]
- Sabotič, J.; Kos, J. Microbial and fungal protease inhibitors—Current and potential applications. Appl. Microbiol. Biotechnol. 2012, 93, 1351–1375. [Google Scholar] [CrossRef]
- Priyanka, P.; Tan, Y.; Kinsella, G.K.; Henehan, G.T.; Ryan, B.J. Solvent stable microbial lipases: Current understanding and biotechnological applications. Biotechnol. Lett. 2019, 41, 203–220. [Google Scholar] [CrossRef]
- Lu, Y.; Liu, D.; Jiang, R.; Li, Z.; Gao, X. Prodigiosin: Unveiling the crimson wonder—A comprehensive journey from diverse bioactivity to synthesis and yield enhancement. Front. Microbiol. 2024, 15, 1412776. [Google Scholar] [CrossRef]
- Hamada, M.A.; Mohamed, E.T. Characterization of Serratia marcescens (OK482790)’ prodigiosin along with in vitro and in silico validation for its medicinal bioactivities. BMC Microbiol. 2024, 24, 495. [Google Scholar] [CrossRef] [PubMed]
- WHO Bacterial Priority Pathogens List, 2024: Bacterial Pathogens of Public Health Importance to Guide Research, Development and Strategies to Prevent and Control Antimicrobial Resistance. Available online: https://www.who.int/publications/i/item/9789240093461 (accessed on 15 September 2025).
- WHO Updates List of Drug-Resistant Bacteria Most Threatening to Human Health. Available online: https://www.who.int/news/item/17-05-2024-who-updates-list-of-drug-resistant-bacteria-most-threatening-to-human-health (accessed on 15 September 2025).
- AWaRe Classification of Antibiotics for Evaluation and Monitoring of Use, 2023. Available online: https://www.who.int/publications/i/item/WHO-MHP-HPS-EML-2023.04 (accessed on 15 September 2025).
- Aranaga, C.; Pantoja, L.D.; Martínez, E.A.; Falco, A. Phage Therapy in the Era of Multidrug Resistance in Bacteria: A Systematic Review. Int. J. Mol. Sci. 2022, 23, 4577. [Google Scholar] [CrossRef] [PubMed]
- Supina, B.S.I.; Dennis, J.J. The Current Landscape of Phage–Antibiotic Synergistic (PAS) Interactions. Antibiotics 2025, 14, 545. [Google Scholar] [CrossRef]
- Bikard, D.; Barrangou, R. Using CRISPR-Cas systems as antimicrobials. Curr. Opin. Microbiol. 2017, 37, 155–160. [Google Scholar] [CrossRef]
- Cherrak, Y.; Filella-Merce, I.; Schmidt, V.; Byrne, D.; Sgoluppi, V.; Chaiaheloudjou, R.; Betzi, S.; Morelli, X.; Nilges, M.; Pellarin, R.; et al. Inhibiting Type VI Secretion System Activity with a Biomimetic Peptide Designed To Target the Baseplate Wedge Complex. mBio 2021, 12, e0134821. [Google Scholar] [CrossRef] [PubMed]
- Alshawwa, S.Z.; Kassem, A.A.; Farid, R.M.; Mostafa, S.K.; Labib, G.S. Nanocarrier Drug Delivery Systems: Characterization, Limitations, Future Perspectives and Implementation of Artificial Intelligence. Pharmaceutics 2022, 14, 883. [Google Scholar] [CrossRef]
- Papp, J.; Iacob, M. Inhibitory potential of some selected essential oils and their main components on the growth and quorum-sensing based pigment production of Serratia marcescens. Stud. Univ. Babeş-Bolyai Biol. 2022, 67, 35–49. [Google Scholar] [CrossRef]
- Zivkovic Zaric, R.; Zaric, M.; Sekulic, M.; Zornic, N.; Nesic, J.; Rosic, V.; Vulovic, T.; Spasic, M.; Vuleta, M.; Jovanovic, J.; et al. Antimicrobial Treatment of Serratia marcescens Invasive Infections: Systematic Review. Antibiotics 2023, 12, 367. [Google Scholar] [CrossRef]
- Veggalam, S.; Kandi, V. Serratia marcescens as an Uncommon Cause of Infection Following Craniectomy: A Case Report and Literature Review. Cureus 2025, 17, e86673. [Google Scholar] [CrossRef]
- Bourdin, T.; Benoit, M.; Monnier, A.; Bédard, E.; Prévost, M.; Charron, D.; Audy, N.; Gravel, S.; Sicard, M.; Quach, C.; et al. Serratia marcescens Colonization in a Neonatal Intensive Care Unit Has Multiple Sources, with Sink Drains as a Major Reservoir. Appl. Environ. Microbiol. 2023, 89, e0010523. [Google Scholar] [CrossRef]
- Guo, Q.; Zhao, X.; Ma, J.; Zhou, Y.; Gao, F.; Huang, W.; Sun, L.; Zhu, S.; Li, L.; Sun, H.; et al. Serratia marcescens outbreak in a neonatal intensive care unit associated with contaminated handwashing sinks. Indian. J. Med. Microbiol. 2024, 52, 100741. [Google Scholar] [CrossRef]
- Alqudah, N.; Tashtoush, M. Contact Lens-Associated Serratia marcescens Keratitis: A Case Report. Clin. Optom. 2025, 17, 243–247. [Google Scholar] [CrossRef]
- Pifer, R.; Harris, V.; Sanders, D.; Crary, M.; Shannon, P. Evaluation of Serratia marcescens Adherence to Contact Lens Materials. Microorganisms 2023, 11, 217. [Google Scholar] [CrossRef] [PubMed]
- Karakonstantis, S.; Rousaki, M.; Kritsotakis, E.I. Cefiderocol: Systematic Review of Mechanisms of Resistance, Heteroresistance and In Vivo Emergence of Resistance. Antibiotics 2022, 11, 723. [Google Scholar] [CrossRef] [PubMed]
- Domingues, S.; Lima, T.; Saavedra, M.J.; Da Silva, G.J. An Overview of Cefiderocol’s Therapeutic Potential and Underlying Resistance Mechanisms. Life 2023, 13, 1427. [Google Scholar] [CrossRef]
- Lin, J.; Xu, L.; Yang, J.; Wang, Z.; Shen, X. Beyond dueling: Roles of the type VI secretion system in microbiome modulation, pathogenesis and stress resistance. Stress. Biol. 2021, 1, 11. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, M.M.; Kayode, H.H.; Okesanya, O.J.; Ukoaka, B.M.; Eshun, G.; Mourid, M.R.; Adigun, O.A.; Ogaya, J.B.; Mohamed, Z.O.; Lucero-Prisno, D.E. CRISPR-Cas Systems in the Fight Against Antimicrobial Resistance: Current Status, Potentials, and Future Directions. Infect. Drug Resist. 2024, 17, 5229–5245. [Google Scholar] [CrossRef]
- Pacia, D.M.; Brown, B.L.; Minssen, T.; Darrow, J.J. CRISPR-phage antibacterials to address the antibiotic resistance crisis: Scientific, economic, and regulatory considerations. J. Law Biosci. 2024, 11, lsad030. [Google Scholar] [CrossRef] [PubMed]
- Fahy, S.; O’Connor, J.A.; Lucey, B.; Sleator, R.D. Hospital Reservoirs of Multidrug Resistant Acinetobacter Species—The Elephant in the Room! Br. J. Biomed. Sci. 2023, 80, 11098. [Google Scholar] [CrossRef] [PubMed]
- Sundermann, A.; Griffith, M.; Srinivasa, V.R.; Waggle, K.; Marsh, J.; Tyne, D.V.; Ayres, A.; Pless, L.; Snyder, G.; Harrison, L. Real-time whole-genome sequencing surveillance for outbreak detection and intervention. Antimicrob. Steward. Healthc. Epidemiol. 2023, 3, s21. [Google Scholar] [CrossRef]
- Ferreira, R.L.; Parente Rocha, J.A.; Leite, V.; Moraes, D.; Graziani, D.; Pranchevicius, M.D.S.; Soares, C.M.A. Proteomic profile of multidrug-resistant Serratia marcescens under meropenem challenge. Microb. Pathog. 2025, 204, 107570. [Google Scholar] [CrossRef]
- Gangadharappa, B.S.; Rajashekarappa, S.; Sathe, G. Proteomic profiling of Serratia marcescens by high-resolution mass spectrometry. Bioimpacts 2020, 10, 123–135. [Google Scholar] [CrossRef]
- D’Souza, S.E.; Khan, K.; Uddin, R. Proteogenomic analysis of Serratia marcescens using computational subtractive genomics approach. PLoS ONE 2023, 18, e0283993. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Boldeanu, L.; Boldeanu, M.V.; Novac, M.B.; Assani, M.-Z.; Radu, L. Serratia marcescens: A Versatile Opportunistic Pathogen with Emerging Clinical and Biotechnological Significance. Int. J. Mol. Sci. 2025, 26, 11479. https://doi.org/10.3390/ijms262311479
Boldeanu L, Boldeanu MV, Novac MB, Assani M-Z, Radu L. Serratia marcescens: A Versatile Opportunistic Pathogen with Emerging Clinical and Biotechnological Significance. International Journal of Molecular Sciences. 2025; 26(23):11479. https://doi.org/10.3390/ijms262311479
Chicago/Turabian StyleBoldeanu, Lidia, Mihail Virgil Boldeanu, Marius Bogdan Novac, Mohamed-Zakaria Assani, and Lucrețiu Radu. 2025. "Serratia marcescens: A Versatile Opportunistic Pathogen with Emerging Clinical and Biotechnological Significance" International Journal of Molecular Sciences 26, no. 23: 11479. https://doi.org/10.3390/ijms262311479
APA StyleBoldeanu, L., Boldeanu, M. V., Novac, M. B., Assani, M.-Z., & Radu, L. (2025). Serratia marcescens: A Versatile Opportunistic Pathogen with Emerging Clinical and Biotechnological Significance. International Journal of Molecular Sciences, 26(23), 11479. https://doi.org/10.3390/ijms262311479







