Metabolism of Cannabidiol in Respiratory-Associated Cells and HepG2-Derived Cells and Molecular Docking of Cannabidiol and Its Metabolites with CYP Enzymes and Cannabinoid Receptors
Abstract
1. Introduction
2. Results
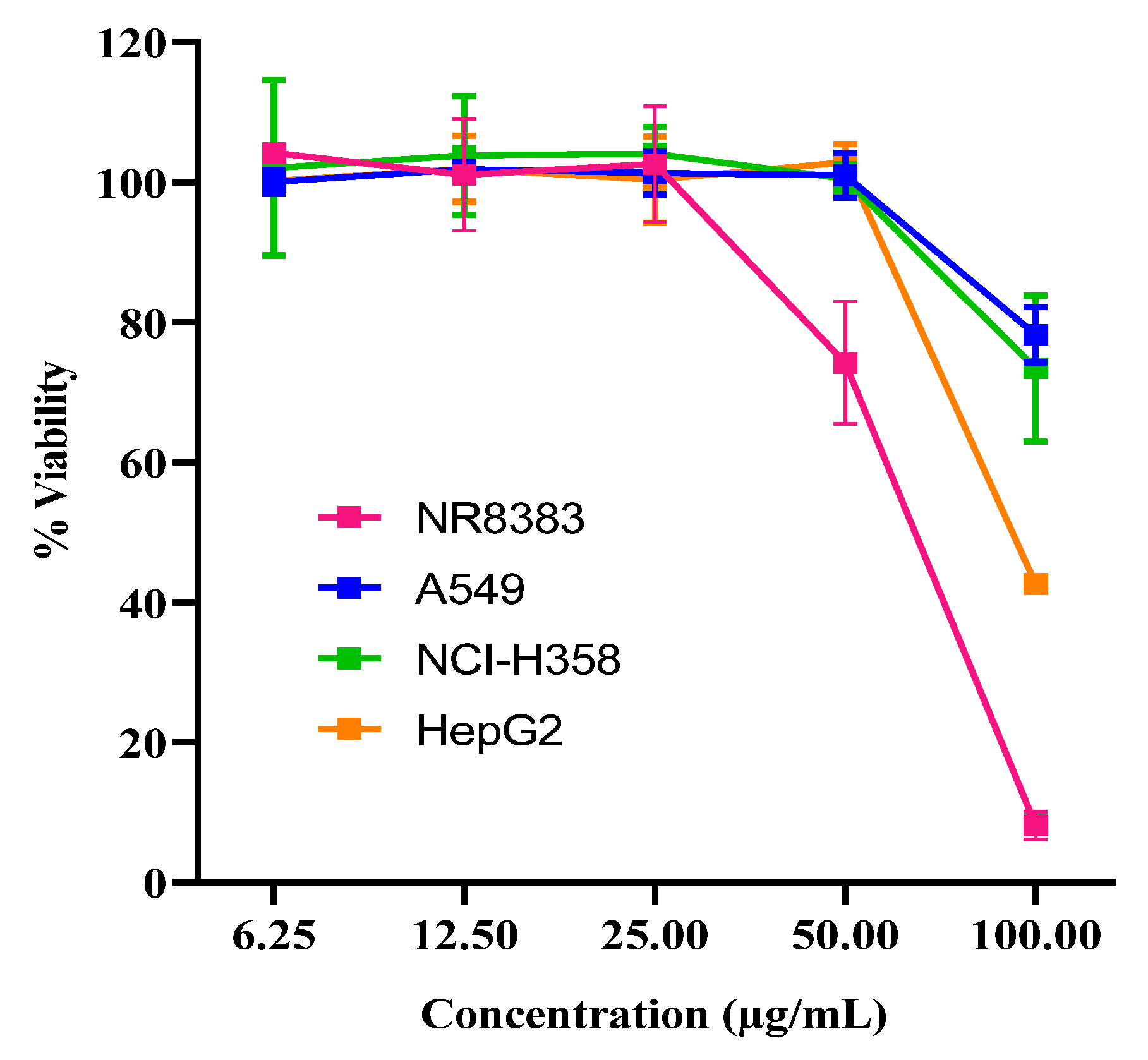
2.1. Cell Viability Assay
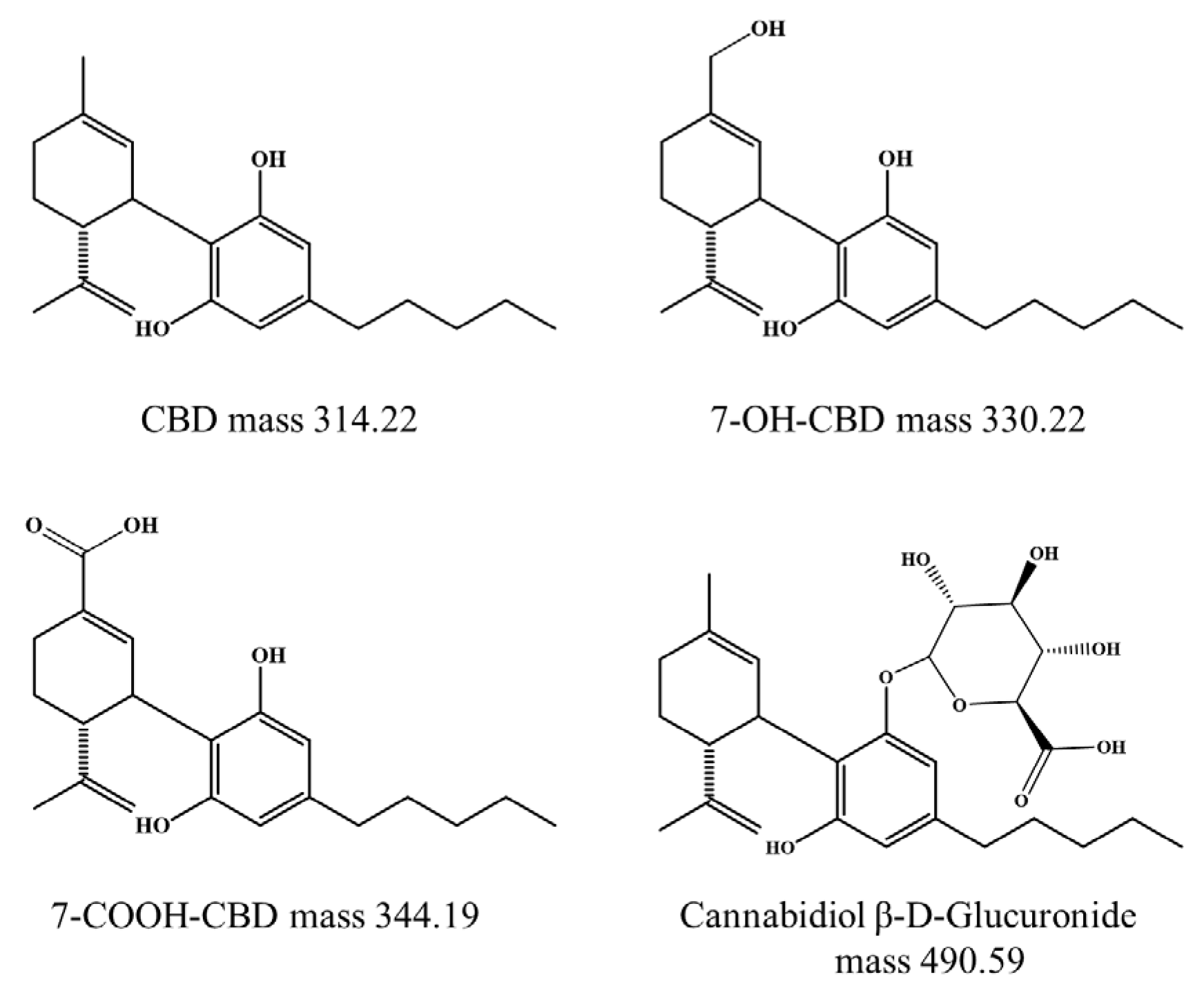
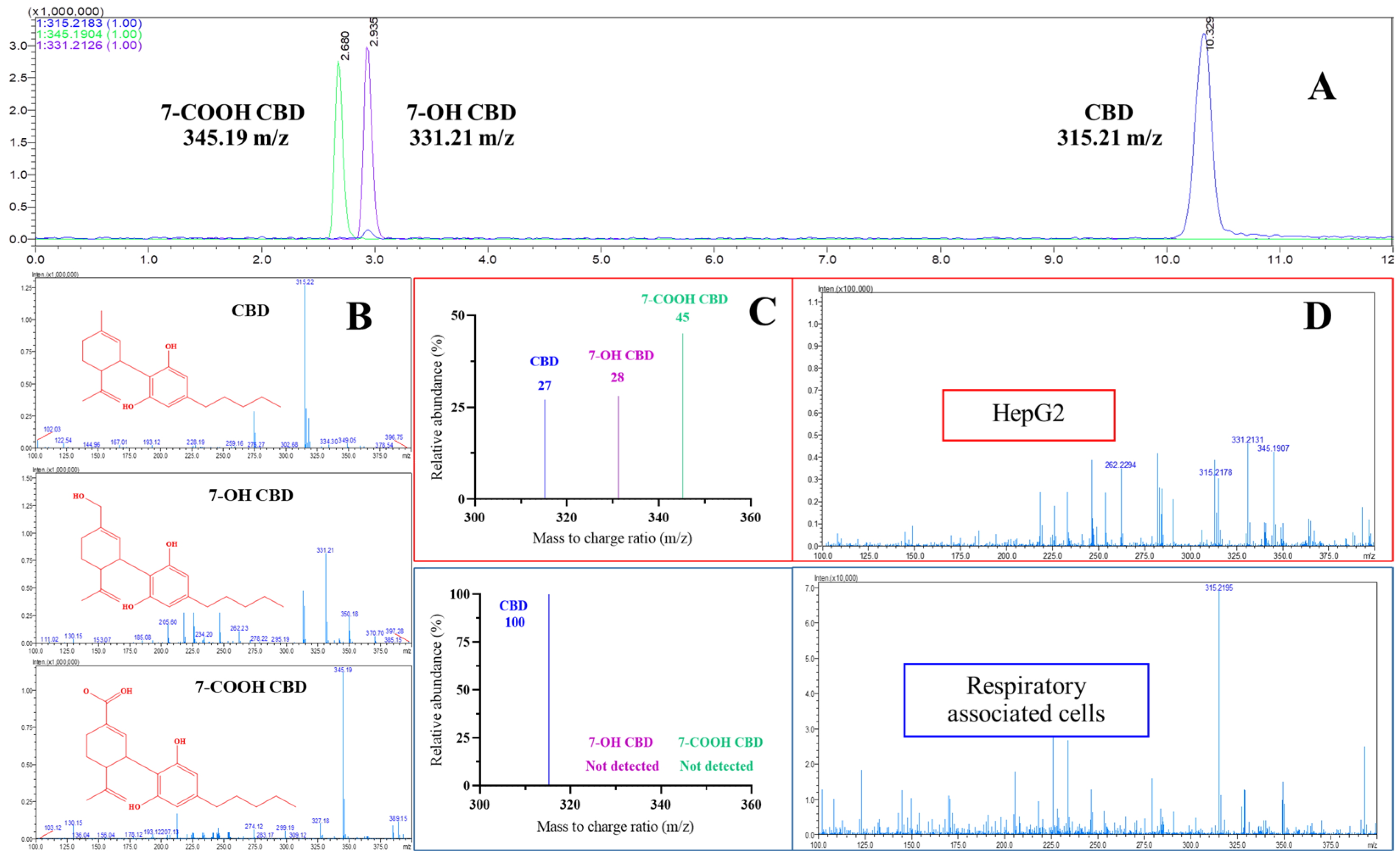
2.2. Analysis of CBD by LC-MS/IT-TOF
2.3. CBD Metabolism in HepG2 Cells and Lung-Associated Cells
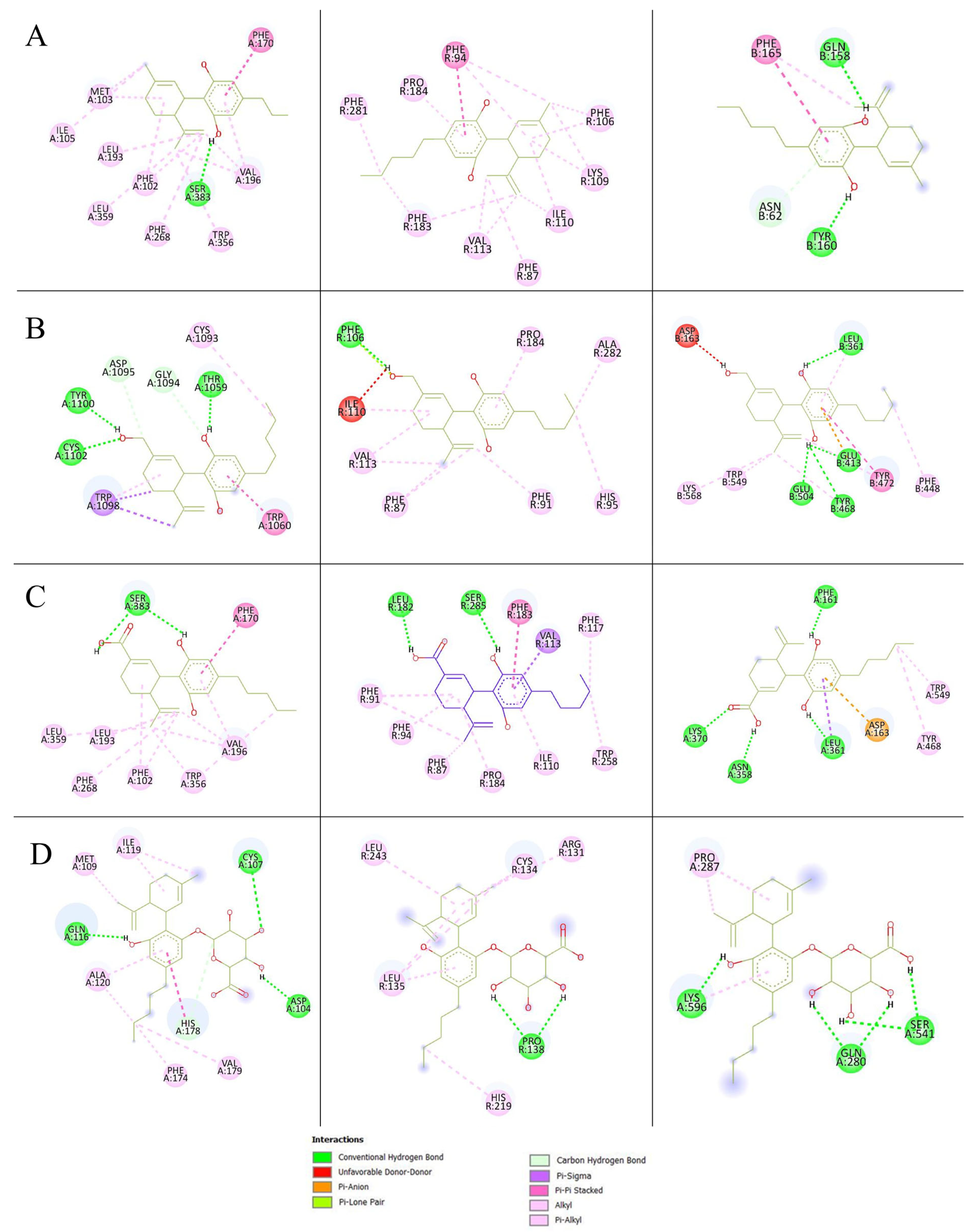
2.4. Binding Interaction by Molecular Docking
3. Discussion
4. Materials and Methods
4.1. Materials
4.2. Chemical Analysis of CBD
4.3. Liquid Chromatography–Tandem Ion Trap Time-of-Flight Mass Spectrometry (LC-MS/IT-TOF) of CBD and CBD Metabolites
4.4. Cell Line Conditions
4.4.1. Alveolar Macrophage Cells
4.4.2. Human Alveolar Cells
4.4.3. Human Bronchioalveolar Carcinoma Cells
4.4.4. Human Hepatocellular Carcinoma Cell Line
4.4.5. Cell Proliferation and Viability Assay
4.5. Drug Metabolism
4.6. Liquid–Liquid Extraction of CBD and Metabolites from the Cell Culture
4.7. Molecular Docking Prediction
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| CBD | Cannabidiol |
| CBD-G | Cannabidiol-glucuronide |
| CB1 | Cannabinoid 1 receptor |
| CB2 | Cannabinoid 2 receptor |
| CYP | Cytochrome |
| LC-MS/IT-TOF | Liquid Chromatography–Tandem Ion Trap Time-of-Flight Mass Spectrometry |
| PHE | Phenylalanine |
| RMSD | Root mean standard deviation |
| TRP | Tryptophan |
| THC | Tetrahydrocannabinol |
References
- Stout, S.M.; Cimino, N.M. Exogenous cannabinoids as substrates, inhibitors, and inducers of human drug metabolizing enzymes: A systematic review. Drug Metab. Rev. 2014, 46, 86–95. [Google Scholar] [CrossRef]
- Howlett, A.C.; Abood, M.E. CB1 and CB2 receptor pharmacology. Adv. Pharmacol. 2017, 80, 169–206. [Google Scholar] [CrossRef] [PubMed]
- Peng, J.; Fan, M.; An, C.; Ni, F.; Huang, W.; Luo, J. A narrative review of molecular mechanism and therapeutic effect of cannabidiol (CBD). Basic Clin. Pharmacol. Toxicol. 2022, 130, 439–456. [Google Scholar] [CrossRef] [PubMed]
- Bansal, S.; Paine, M.F.; Unadkat, J.D. Comprehensive predictions of cytochrome P450 (P450)-mediated in vivo cannabinoid-drug interactions based on reversible and time-dependent P450 inhibition in human liver microsomes. Drug Metab. Dispos. 2022, 50, 351–360. [Google Scholar] [CrossRef] [PubMed]
- Bardhi, K.; Coates, S.; Watson, C.J.; Lazarus, P. Cannabinoids and drug metabolizing enzymes: Potential for drug-drug interactions and implications for drug safety and efficacy. Expert Rev. Clin. Pharmacol. 2022, 15, 1443–1460. [Google Scholar] [CrossRef]
- Jiang, R.; Yamaori, S.; Takeda, S.; Yamamoto, I.; Watanabe, K. Identification of cytochrome P450 enzymes responsible for metabolism of cannabidiol by human liver microsomes. Life Sci. 2011, 89, 165–170. [Google Scholar] [CrossRef]
- Beers, J.L.; Fu, D.; Jackson, K.D. Cytochrome P450–catalyzed metabolism of cannabidiol to the active metabolite 7-hydroxy-cannabidiol. Drug Metab. Dispos. 2021, 49, 882–891. [Google Scholar] [CrossRef]
- Ujváry, I.; Hanuš, L. Human metabolites of cannabidiol: A review on their formation, biological activity, and relevance in therapy. Cannabis Cannabinoid Res. 2016, 1, 90–101. [Google Scholar] [CrossRef]
- Nasrin, S.; Watson, C.J.; Perez-Paramo, Y.X.; Lazarus, P. Cannabinoid metabolites as inhibitors of major hepatic CYP450 enzymes, with implications for cannabis-drug interactions. Drug Metab. Dispos. 2021, 49, 1070–1080. [Google Scholar] [CrossRef]
- Chayasirisobhon, S. Mechanisms of action and pharmacokinetics of cannabis. Perm. J. 2020, 25, 1–3. [Google Scholar] [CrossRef]
- Taylor, L.; Gidal, B.; Blakey, G.; Tayo, B.; Morrison, G. A phase I, randomized, double-blind, placebo-controlled, single ascending dose, multiple dose, and food effect trial of the safety, tolerability and pharmacokinetics of highly purified cannabidiol in healthy subjects. CNS Drugs 2018, 32, 1053–1067. [Google Scholar] [CrossRef]
- Devinsky, O.; Kraft, K.; Rusch, L.; Fein, M.; Leone-Bay, A. Improved bioavailability with dry powder cannabidiol inhalation: A phase 1 clinical study. J. Pharm. Sci. 2021, 110, 3946–3952. [Google Scholar] [CrossRef]
- Ashton, J.C.; Glass, M. The cannabinoid CB2 receptor as a target for inflammation-dependent neurodegeneration. Curr. Neuropharmacol. 2007, 5, 73–80. [Google Scholar] [CrossRef]
- Spindle, T.R.; Cone, E.J.; Goffi, E.; Weerts, E.M.; Mitchell, J.M.; Winecker, R.E.; Bigelow, G.E.; Flegel, R.R.; Vandrey, R. Pharmacodynamic effects of vaporized and oral cannabidiol (CBD) and vaporized CBD-dominant cannabis in infrequent cannabis users. Drug Alcohol Depend. 2020, 211, 107937. [Google Scholar] [CrossRef]
- Burstein, S. Cannabidiol (CBD) and its analogs: A review of their effects on inflammation. Bioorg. Med. Chem. 2015, 23, 1377–1385. [Google Scholar] [CrossRef]
- O’Sullivan, S.E.; Jensen, S.S.; Kolli, A.R.; Nikolajsen, G.N.; Bruun, H.Z.; Hoeng, J. Strategies to improve cannabidiol bioavailability and drug delivery. Pharmaceuticals 2024, 17, 244. [Google Scholar] [CrossRef]
- Hložek, T.; Uttl, L.; Kadeřábek, L.; Balíková, M.; Lhotková, E.; Horsley, R.R.; Nováková, P.; Šíchová, K.; Štefková, K.; Tylš, F.; et al. Pharmacokinetic and behavioural profile of THC, CBD, and THC+CBD combination after pulmonary, oral, and subcutaneous administration in rats and confirmation of conversion in vivo of CBD to THC. Eur. Neuropsychopharmacol. 2017, 27, 1223–1237. [Google Scholar] [CrossRef] [PubMed]
- Tai, W.; Arnold, J.C.; Chan, H.-K.; Kwok, P.C.L. Spray freeze dried cannabidiol with dipalmitoylphosphatidylcholine (DPPC) for inhalation and solubility enhancement. Int. J. Pharm. 2024, 659, 124235. [Google Scholar] [CrossRef] [PubMed]
- Enlo-Scott, Z.; Bäckström, E.; Mudway, I.; Forbes, B. Drug metabolism in the lungs: Opportunities for optimising inhaled medicines. Expert Opin. Drug Metab. Toxicol. 2021, 17, 611–625. [Google Scholar] [CrossRef] [PubMed]
- Kicman, A.; Toczek, M. The effects of cannabidiol, a non-intoxicating compound of cannabis, on the cardiovascular system in health and disease. Int. J. Mol. Sci. 2020, 21, 6740. [Google Scholar] [CrossRef]
- Pigliasco, F.; Malaca, S.; Faro, A.F.L.; Tini, A.; Cangemi, G.; Cafaro, A.; Barco, S.; Riva, A.; Pisati, A.; Amadori, E.; et al. Cannabidiol, ∆9-tetrahydrocannabinol, and metabolites in human blood by volumetric absorptive microsampling and LCMS/MS following controlled administration in epilepsy patients. Front. Pharmacol. 2022, 13, 1038754. [Google Scholar] [CrossRef]
- Courcot, E.; Leclerc, J.; Lafitte, J.-J.; Mensier, E.; Jaillard, S.; Gosset, P.; Shirali, P.; Pottier, N.; Broly, F.; Lo-Guidice, J.-M. Xenobiotic metabolism and disposition in human lung cell models: Comparison with In Vivo expression profiles. Drug Metab. Dispos. 2012, 40, 1953–1965. [Google Scholar] [CrossRef]
- Hukkanen, J.; Pelkonen, O.; Hakkola, J.; Raunio, H. Expression and regulation of xenobiotic-metabolizing cytochrome P450 (CYP) enzymes in human lung. Crit. Rev. Toxicol. 2002, 32, 391–411. [Google Scholar] [CrossRef] [PubMed]
- Falzon, M.; McMahin, J.B.; Schuller, H.M. Xenobiotic-metabolizing enzyme activity in human non-small-cell derived lung cancer cell lines. Biochem. Pharmacol. 1986, 35, 563–568. [Google Scholar] [CrossRef]
- Hukkanen, J.; Lassila, A.; Paivarinta, K.; Valanne, S.; Sarpo, S.; Hakkola, J.; Pelkonen, O.; Raunio, H. Induction and regulation of xenobiotic-metabolizing cytochrome P450s in the human A549 lung adenocarcinoma cell line. Am. J. Respir. Cell Mol. Biol. 2000, 22, 360–366. [Google Scholar] [CrossRef] [PubMed]
- Yang, G.; Ge, S.; Singh, R.; Basu, S.; Shatzer, K.; Zen, M.; Liu, J.; Tu, Y.; Zhang, C.; Wei, J.; et al. Glucuronidation: Driving factors and their impact on glucuronide disposition. Drug Metab. Rev. 2017, 49, 105–138. [Google Scholar] [CrossRef] [PubMed]
- Nelson, K.M.; Bisson, J.; Singh, G.; Graham, J.G.; Chen, S.-N.; Friesen, J.B.; Dahlin, J.L.; Niemitz, M.; Walters, M.A.; Pauli, G.F. The essential medicinal chemistry of cannabidiol (CBD). J. Med. Chem. 2020, 63, 12137–12155. [Google Scholar] [CrossRef]
- Watanabe, K.; Usami, N.; Osada, S.; Narimatsu, S.; Yamamoto, I.; Yoshimura, H. Cannabidiol metabolism revisited: Tentative identification of novel decarbonylated metabolites of cannabidiol formed by human liver microsomes and recombinant cytochrome P450 3A4. Forensic Toxicol. 2019, 37, 449–455. [Google Scholar] [CrossRef]
- Raunio, H.; Hakkola, J.; Pelkonen, O. Regulation of CYP3A genes in the human respiratory tract. Chem. Biol. Interact. 2005, 151, 53–62. [Google Scholar] [CrossRef]
- Rossi, F.; Tortora, C.; Argenziano, M.; Di Paola, A.; Punzo, F. Cannabinoid Receptor Type 2: A Possible Target in SARS-CoV-2 (CoV-19) Infection. Int. J. Mol. Sci. 2020, 21, 3809. [Google Scholar] [CrossRef]
- Jakowiecki, J.; Abel, R.; Orzeł, U.; Pasznik, P.; Preissner, R.; Filipek, S. Allosteric modulation of the CB1 cannabinoid receptor by cannabidiol—A molecular modeling study of the N-terminal domain and the allosteric-orthosteric coupling. Molecules 2021, 26, 2456. [Google Scholar] [CrossRef]
- Yuan, J.; Jiang, C.; Wang, J.; Chen, C.-J.; Hao, Y.; Zhao, G.; Feng, Z.; Xie, X.-Q. In silico prediction and validation of CB2 allosteric binding sites to aid the design of allosteric modulators. Molecules 2022, 27, 453. [Google Scholar] [CrossRef]
- Thomas, A.; Baillie, G.; Phillips, A.; Razdan, R.; Ross, R.A.; Pertwee, R. Cannabidiol displays unexpectedly high potency as an antagonist of CB1 and CB2 receptor agonists in vitro. Br. J. Pharmacol. 2007, 150, 613–623. [Google Scholar] [CrossRef] [PubMed]
- Rao, Q.; Zhang, T.; Dai, M.; Li, B.; Pu, Q.; Zhao, M.; Cheng, Y.; Yan, D.; Zhao, Q.; Wu, Z.E.; et al. Comparative metabolomic profiling of the metabolic differences of Δ9-tetrahydrocannabinol and cannabidiol. Molecules 2022, 27, 7573. [Google Scholar] [CrossRef]
- Srichana, T.; Chunhachaichana, C.; Suedee, R.; Sawatdee, S.; Changsan, N. Oral inhalation of cannabidiol delivered from a metered dose inhaler to alleviate cytokine production induced by SARS-CoV-2 and pollutants. J. Drug Deliv. Sci. Technol. 2022, 76, 103805. [Google Scholar] [CrossRef]
- Morris, G.M.; Huey, R.; Lindstrom, W.; Sanner, M.F.; Belew, R.K.; Goodsell, D.S.; Olson, A.J. AutoDock4 and AutoDockTools4: Automated docking with selective receptor flexibility. J. Comput. Chem. 2009, 30, 2785–2791. [Google Scholar] [CrossRef]
| Target Protein | PDB ID | RMSD (Å) |
|---|---|---|
| CB1 | 5TGZ | 1.936 |
| CB2 | 6KPF | 0.952 |
| β-Glucuronidase | 3LPG | 1.794 |
| CYP1A2 | 2HI4 | 0.571 |
| CYP2C9 | 1R9O | 1.421 |
| CYP2C19 | 4GQS | 1.462 |
| CYP2D6 | 5TFT | 0.882 |
| CYP3A4 | 1TQN | 0.986 |
| CYP3A5 | 7SV2 | 0.650 |
| Ligand | Macromolecule | ||
|---|---|---|---|
| CB1 (kcal/mol) | CB2 (kcal/mol) | β-Glucuronidase (kcal/mol) | |
| CBD | −9.38 | −9.60 | −8.54 |
| 7-OH-CBD | −9.69 | −9.11 | −8.53 |
| 7-COOH-CBD | −9.01 | −9.16 | −7.89 |
| CBD-G | −6.07 | −6.07 | −6.48 |
| Ligand–Enzyme Interaction | Binding Energy (kcal·mol−1) |
|---|---|
| CBD-CYP1A2 | −9.38 ± 0.05 |
| CBD-CYP2D6 | −9.31 ± 0.10 |
| CBD-CYP3A5 | −8.85 ± 0.07 |
| CBD-CYP2C19 | −8.64 ± 0.09 |
| Cell Lines | Observed m/z | Predicted Formula | Identification |
|---|---|---|---|
| HepG2 | 315.21 | C21H30O2 | CBD |
| 331.21 | C21H30O3 | 7-OH-CBD (hydroxylation) | |
| 345.19 | C21H28O4 | 7-COOH-CBD (oxidation) | |
| NCI-H358 | 315.21 | C21H30O2 | CBD |
| A549 | 315.21 | C21H30O2 | CBD |
| NR8383 | 315.21 | C21H30O2 | CBD |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tongkanarak, K.; Paul, P.K.; Bintang, M.A.K.M.; Suedee, R.; Sawatdee, S.; Srichana, T. Metabolism of Cannabidiol in Respiratory-Associated Cells and HepG2-Derived Cells and Molecular Docking of Cannabidiol and Its Metabolites with CYP Enzymes and Cannabinoid Receptors. Int. J. Mol. Sci. 2025, 26, 8384. https://doi.org/10.3390/ijms26178384
Tongkanarak K, Paul PK, Bintang MAKM, Suedee R, Sawatdee S, Srichana T. Metabolism of Cannabidiol in Respiratory-Associated Cells and HepG2-Derived Cells and Molecular Docking of Cannabidiol and Its Metabolites with CYP Enzymes and Cannabinoid Receptors. International Journal of Molecular Sciences. 2025; 26(17):8384. https://doi.org/10.3390/ijms26178384
Chicago/Turabian StyleTongkanarak, Krittawan, Pijush Kumar Paul, Muhammad A. Khumaini Mudhar Bintang, Roongnapa Suedee, Somchai Sawatdee, and Teerapol Srichana. 2025. "Metabolism of Cannabidiol in Respiratory-Associated Cells and HepG2-Derived Cells and Molecular Docking of Cannabidiol and Its Metabolites with CYP Enzymes and Cannabinoid Receptors" International Journal of Molecular Sciences 26, no. 17: 8384. https://doi.org/10.3390/ijms26178384
APA StyleTongkanarak, K., Paul, P. K., Bintang, M. A. K. M., Suedee, R., Sawatdee, S., & Srichana, T. (2025). Metabolism of Cannabidiol in Respiratory-Associated Cells and HepG2-Derived Cells and Molecular Docking of Cannabidiol and Its Metabolites with CYP Enzymes and Cannabinoid Receptors. International Journal of Molecular Sciences, 26(17), 8384. https://doi.org/10.3390/ijms26178384





