Physiological Roles and Mechanisms of Action of Class I TCP Transcription Factors
Abstract
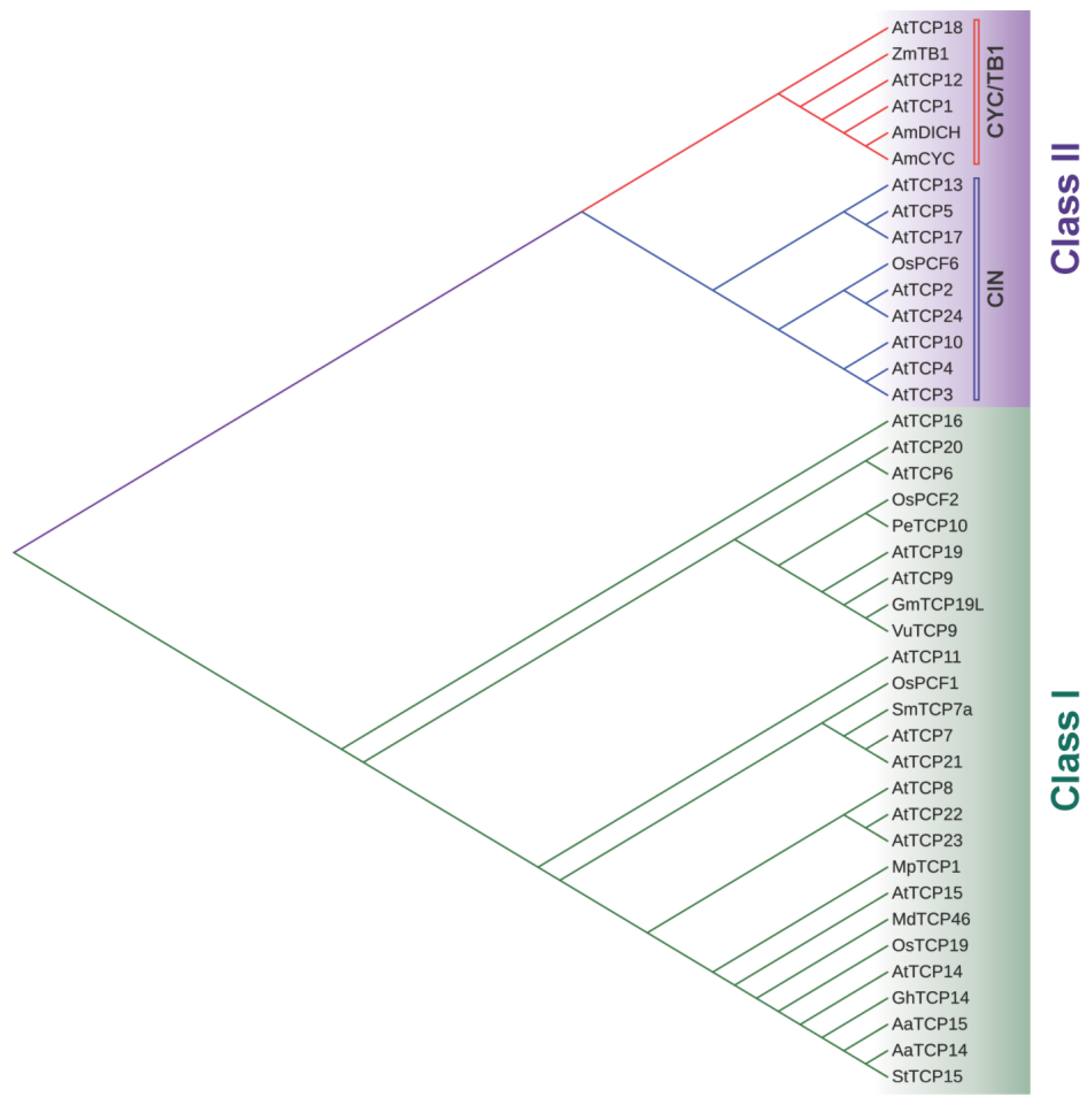
1. Introduction
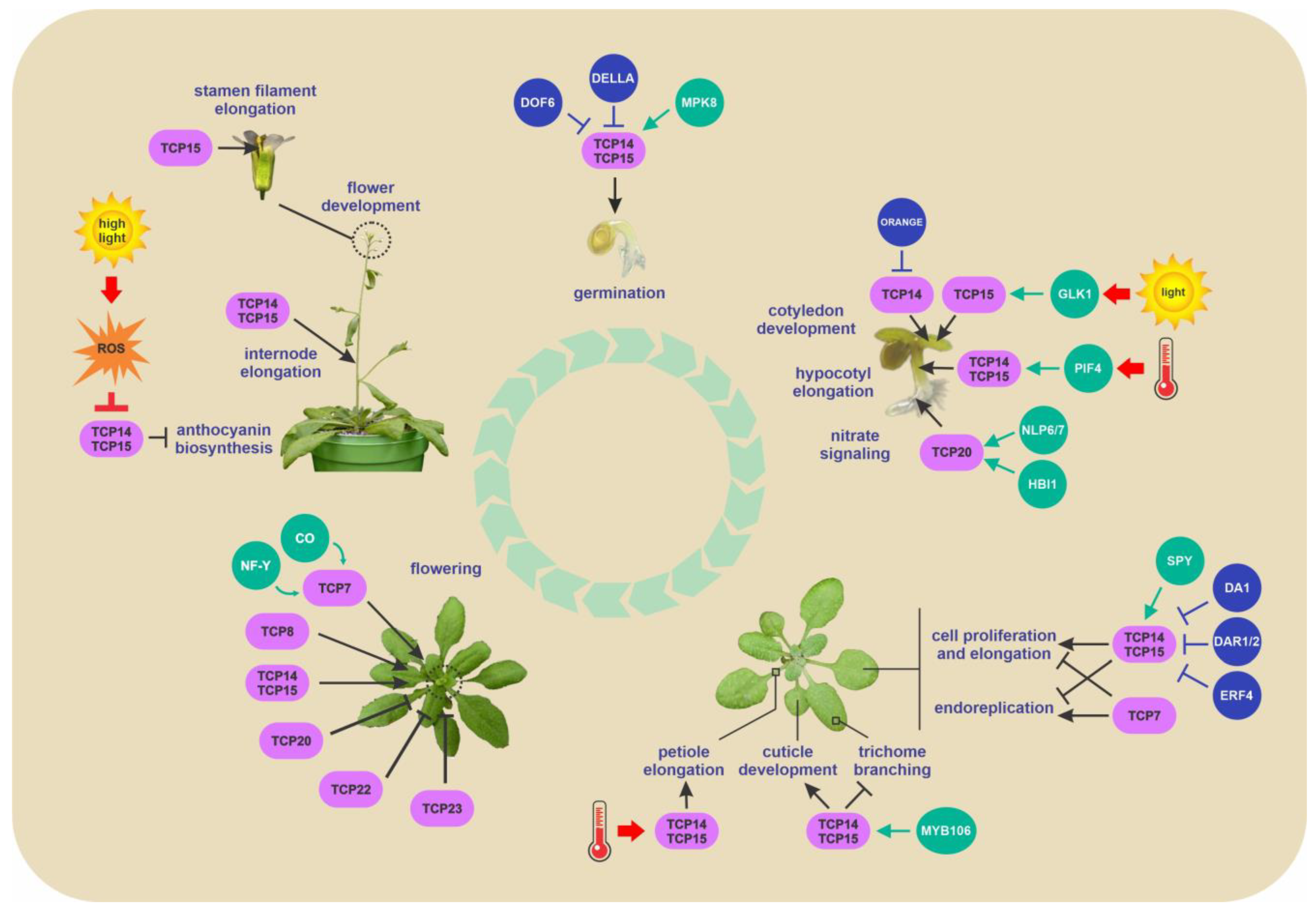
2. Biological Processes Modulated by Class I TCPs
2.1. Role of Class I TCPs in Cell Division, Growth and Expansion
2.2. Role of Class I TCPs in Germination
2.3. Role of Class I TCPs in Epidermis Development
2.4. Role of Class I TCPs in Flowering
2.5. Role of Class I TCPs in Response to Light and Temperature
2.6. Role of Class I TCPs in Nitrate and Copper Homeostasis
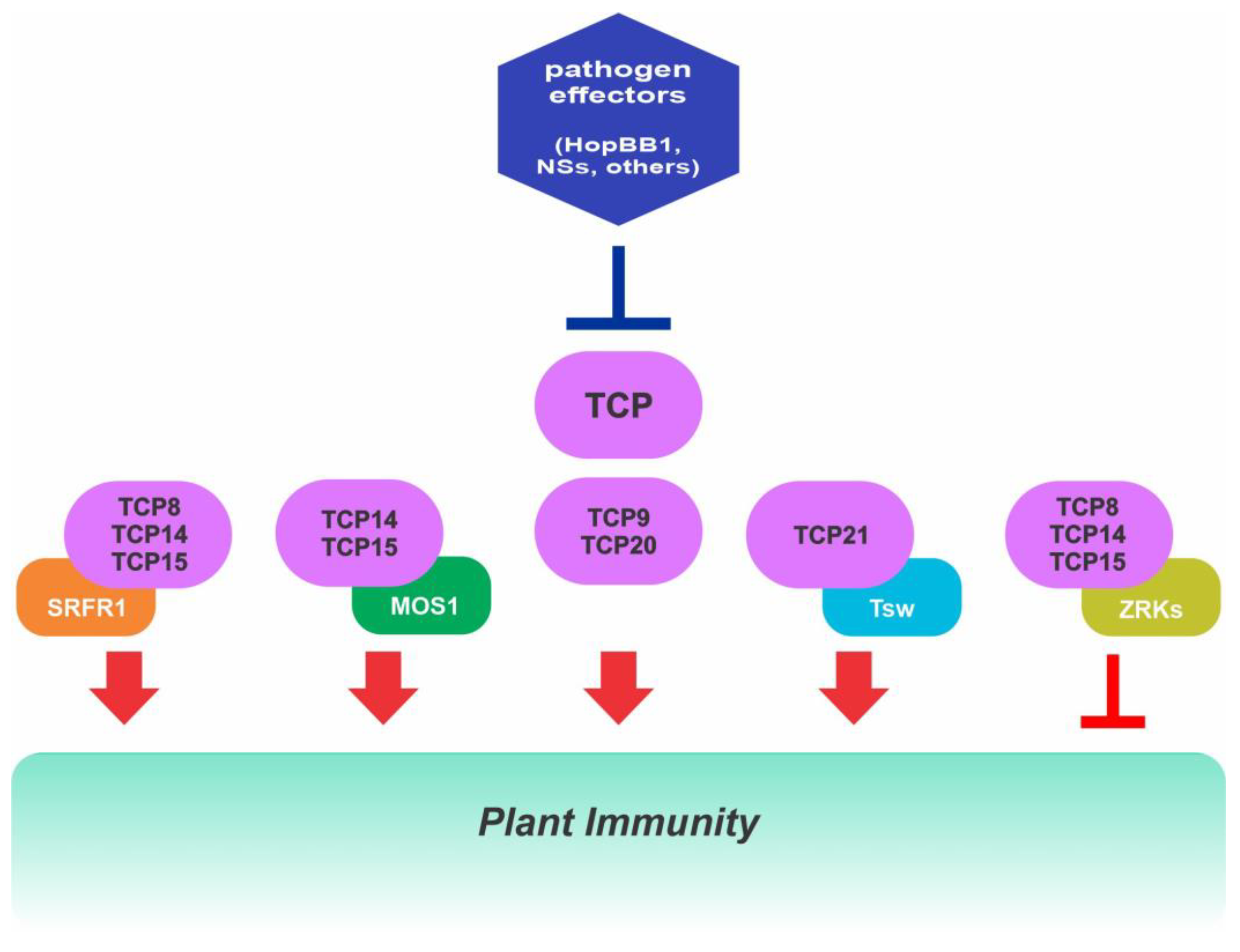
2.7. Role of Class I TCPs in Immunity
2.8. Role of Class I TCPs in Abiotic Stress Responses
3. Modulation of Class I TCP Protein Activity
3.1. Interaction with Non-TCP Transcriptional Regulators
3.2. Class I TCPs in Redox Signaling
3.3. Subcellular Distribution of Class I TCPs
4. Concluding Remarks and Perspectives
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Cubas, P.; Lauter, N.; Doebley, J.; Coen, E. The TCP Domain: A Motif Found in Proteins Regulating Plant Growth and Development. Plant J. 1999, 18, 215–222. [Google Scholar] [CrossRef]
- Doebley, J.; Stec, A.; Hubbard, L. The Evolution of Apical Dominance in Maize. Nature 1997, 386, 485–488. [Google Scholar] [CrossRef]
- Luo, D.; Carpenter, R.; Vincent, C.; Copsey, L.; Coen, E. Origin of Floral Asymmetry in Antirrhinum. Nature 1996, 383, 794–799. [Google Scholar] [CrossRef]
- Kosugi, S.; Ohashi, Y. PCF1 and PCF2 Specifically Bind to Cis Elements in the Rice Proliferating Cell Nuclear Antigen Gene. Plant Cell 1997, 9, 1607–1619. [Google Scholar] [CrossRef] [PubMed]
- Howarth, D.G.; Donoghue, M.J. Phylogenetic Analysis of the “ECE” (CYC/TB1) Clade Reveals Duplications Predating the Core Eudicots. Proc. Natl. Acad. Sci. USA 2006, 103, 9101–9106. [Google Scholar] [CrossRef]
- Martín-Trillo, M.; Cubas, P. TCP Genes: A Family Snapshot Ten Years Later. Trends Plant Sci. 2010, 15, 31–39. [Google Scholar] [CrossRef] [PubMed]
- Navaud, O.; Dabos, P.; Carnus, E.; Tremousaygue, D.; Hervé, C. TCP Transcription Factors Predate the Emergence of Land Plants. J. Mol. Evol. 2007, 65, 23–33. [Google Scholar] [CrossRef]
- Kosugi, S.; Ohashi, Y. DNA Binding and Dimerization Specificity and Potential Targets for the TCP Protein Family. Plant J. 2002, 30, 337–348. [Google Scholar] [CrossRef] [PubMed]
- Viola, I.L.; Uberti Manassero, N.G.; Ripoll, R.; Gonzalez, D.H. The Arabidopsis Class I TCP transcription factor AtTCP11 is a developmental regulator with distinct DNA-binding properties due to the presence of a Threonine residue at position 15 of the TCP domain. Biochem. J. 2011, 435, 143–155. [Google Scholar] [CrossRef]
- Viola, I.L.; Reinheimer, R.; Ripoll, R.; Manassero, N.G.U.; Gonzalez, D.H. Determinants of the DNA binding specificity of class I and class II TCP transcription factors. J. Biol. Chem. 2012, 287, 347–356. [Google Scholar] [CrossRef] [PubMed]
- Sun, L.; Zou, X.; Jiang, M.; Wu, X.; Chen, Y.; Wang, Q.; Wang, Q.; Chen, L.; Wu, Y. The Crystal Structure of the TCP Domain of PCF6 in Oryza sativa L. Reveals an RHH-like Fold. FEBS Lett. 2020, 594, 1296–1306. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Xu, Y.; Nie, J.; Chen, H.; Qin, G.; Wang, B.; Su, X.-D. DNA–TCP Complex Structures Reveal a Unique Recognition Mechanism for TCP Transcription Factor Families. Nucleic Acids Res. 2022, 51, 434–448. [Google Scholar] [CrossRef] [PubMed]
- Valsecchi, I.; Guittard-Crilat, E.; Maldiney, R.; Habricot, Y.; Lignon, S.; Lebrun, R.; Miginiac, E.; Ruelland, E.; Jeannette, E.; Lebreton, S. The Intrinsically Disordered C-Terminal Region of Arabidopsis Thaliana TCP8 Transcription Factor Acts Both as a Transactivation and Self-Assembly Domain. Mol. Biosyst. 2013, 9, 2282. [Google Scholar] [CrossRef]
- Thieulin-Pardo, G.; Avilan, L.; Kojadinovic, M.; Gontero, B. Fairy “tails”: Flexibility and function of intrinsically disordered extensions in the photosynthetic world. Front. Mol. Biosci. 2015, 2, 23. [Google Scholar] [CrossRef] [PubMed]
- Danisman, S.; Van Dijk, A.D.J.; Bimbo, A.; Van Der Wal, F.; Hennig, L.; De Folter, S.; Angenent, G.C.; Immink, R.G.H. Analysis of Functional Redundancies within the Arabidopsis TCP Transcription Factor Family. J. Exp. Bot. 2013, 64, 5673–5685. [Google Scholar] [CrossRef]
- Danisman, S. TCP Transcription Factors at the Interface between Environmental Challenges and the Plant’s Growth Responses. Front. Plant Sci. 2016, 7, 1930. [Google Scholar] [CrossRef]
- Uberti Manassero, N.G.; Viola, I.L.; Welchen, E.; Gonzalez, D.H. TCP Transcription Factors: Architectures of Plant Form. Biomol. Concepts 2013, 4, 111–127. [Google Scholar] [CrossRef]
- Busch, A.; Deckena, M.; Almeida-Trapp, M.; Kopischke, S.; Kock, C.; Sch€ Ussler, E.; Tsiantis, M.; Mith€ Ofer, A.; Zachgo, S. MpTCP1 Controls Cell Proliferation and Redox Processes in Marchantia Polymorpha. New Phytol. 2019, 224, 1627–1641. [Google Scholar] [CrossRef]
- Mukhopadhyay, P.; Tyagi, A.K. OsTCP19 Influences Developmental and Abiotic Stress Signaling by Modulating ABI4-Mediated Pathways. Sci. Rep. 2015, 5, 1–11. [Google Scholar] [CrossRef]
- Feng, Z.-J.; Xu, S.-C.; Liu, N.; Zhang, G.-W.; Hu, Q.-Z.; Gong, Y.-M. Soybean TCP Transcription Factors: Evolution, Classification, Protein Interaction and Stress and Hormone Responsiveness. Plant Physiol. Biochem. 2018, 127, 129–142. [Google Scholar] [CrossRef]
- Koyama, T.; Sato, F.; Ohme-Takagi, M. Roles of MiR319 and TCP Transcription Factors in Leaf Development. Plant Physiol. 2017, 175, 874–885. [Google Scholar] [CrossRef] [PubMed]
- Fan, S.I.; Zhang, Z.; Song, Y.; Zhang, J.; Wang, P.I. CRISPR/Cas9-Mediated Targeted Mutagenesis of GmTCP19L Increasing Susceptibility to Phytophthora Sojae in Soybean. PLoS ONE 2022, 17, e0267502. [Google Scholar] [CrossRef]
- Hao, J.; Tu, L.; Hu, H.; Tan, J.; Deng, F.; Tang, W.; Nie, Y.; Zhang, X. GbTCP, a Cotton TCP Transcription Factor, Confers Fibre Elongation and Root Hair Development by a Complex Regulating System. J. Exp. Bot. 2012, 63, 6267–6281. [Google Scholar] [CrossRef]
- Liu, Y.J.; An, J.P.; Gao, N.; Wang, X.; Chen, X.X.; Wang, X.F.; Zhang, S.; You, C.X. MdTCP46 interacts with MdABI5 to negatively regulate ABA signalling and drought response in apple. Plant Cell Environ. 2022, 45, 3233–3248. [Google Scholar] [CrossRef]
- Zhang, T.; Qu, Y.; Wang, H.; Wang, J.; Song, A.; Hu, Y.; Chen, S.; Jiang, J.; Chen, F. The Heterologous Expression of a Chrysanthemum TCP-P Transcription Factor CmTCP14 Suppresses Organ Size and Delays Senescence in Arabidopsis thaliana. Plant Physiol. Biochem. 2017, 115, 239–248. [Google Scholar] [CrossRef]
- Wang, Q.; Xu, G.; Zhao, X.; Zhang, Z.; Wang, X.; Liu, X.; Xiao, W.; Fu, X.; Chen, X.; Gao, D.; et al. Transcription Factor TCP20 Regulates Peach Bud Endodormancy by Inhibiting DAM5/DAM6 and Interacting with ABF2. J. Exp. Bot. 2020, 71, 1585–1597. [Google Scholar] [CrossRef] [PubMed]
- Xiong, W.; Zhao, Y.; Gao, H.; Li, Y.; Tang, W.; Ma, L.; Yang, G.; Sun, J. Genomic characterization and expression analysis of TCP transcription factors in Setaria italica and Setaria viridis. Plant Signal. Behav. 2022, 17, 2075158. [Google Scholar] [CrossRef]
- Koyama, T.; Furutani, M.; Tasaka, M.; Ohme-Takagi, M. TCP Transcription Factors Control the Morphology of Shoot Lateral Organs via Negative Regulation of the Expression of Boundary-Specific Genes in Arabidopsis. Plant Cell 2007, 19, 473–484. [Google Scholar] [CrossRef]
- Koyama, T.; Mitsuda, N.; Seki, M.; Shinozaki, K.; Ohme-Takagi, M. TCP Transcription Factors Regulate the Activities of ASYMMETRIC LEAVES1 and MiR164, as Well as the Auxin Response, during Differentiation of Leaves in Arabidopsis. Plant Cell 2010, 22, 3574–3588. [Google Scholar] [CrossRef]
- Chen, L.; Chen, Y.Q.; Ding, A.M.; Chen, H.; Xia, F.; Wang, W.F.; Sun, Y.H. Genome-Wide Analysis of TCP Family in Tobacco. Genet. Mol. Res. 2016, 15, 15027728. [Google Scholar] [CrossRef]
- Song, C.-B.; Shan, W.; Yang, Y.-Y.; Tan, X.-L.; Fan, Z.-Q.; Chen, J.-Y.; Lu, W.-J.; Kuang, J.-F. Heterodimerization of MaTCP Proteins Modulates the Transcription of MaXTH10/11 Genes during Banana Fruit Ripening. Biochim. Biophys. Acta-Gene Regul. Mech. 2018, 1861, 613–622. [Google Scholar] [CrossRef] [PubMed]
- Liu, M.-M.; Wang, M.-M.; Yang, J.; Wen, J.; Guo, P.-C.; Wu, Y.-W.; Ke, Y.-Z.; Li, P.-F.; Li, J.-N.; Du, H. Evolutionary and Comparative Expression Analyses of TCP Transcription Factor Gene Family in Land Plants. Int. J. Mol. Sci. 2019, 20, 3591. [Google Scholar] [CrossRef] [PubMed]
- Xie, Y.-G.; Ma, Y.-Y.; Bi, P.-P.; Wei, W.; Liu, J.; Hu, Y.; Gou, Y.-J.; Zhu, D.; Wen, Y.-Q.; Feng, J.-Y. Transcription Factor FvTCP9 Promotes Strawberry Fruit Ripening by Regulating the Biosynthesis of Abscisic Acid and Anthocyanins. Plant Physiol. Biochem. 2020, 146, 374–383. [Google Scholar] [CrossRef]
- Wen, H.; Chen, Y.; Du, H.; Zhang, L.; Zhang, K.; He, H.; Pan, J.; Cai, R.; Wang, G. Genome-Wide Identification and Characterization of the TCP Gene Family in Cucumber (Cucumis sativus L.) and Their Transcriptional Responses to Different Treatments. Genes 2020, 11, 1379. [Google Scholar] [CrossRef]
- Shi, P.; Guy, K.M.; Wu, W.; Fang, B.; Yang, J.; Zhang, M.; Hu, Z. Genome-Wide Identification and Expression Analysis of the ClTCP Transcription Factors in Citrullus lanatus. BMC Plant Biol. 2016, 16, 85. [Google Scholar] [CrossRef] [PubMed]
- Wei, W.; Hu, Y.; Cui, M.-Y.; Han, Y.-T.; Gao, K.; Feng, J.-Y. Identification and Transcript Analysis of the TCP Transcription Factors in the Diploid Woodland Strawberry Fragaria Vesca. Front. Plant Sci. 2016, 7, 1937. [Google Scholar] [CrossRef]
- Yu, L.; Chen, Q.; Zheng, J.; Xu, F.; Ye, J.; Zhang, W.; Liao, Y.; Yang, X. Genome-Wide Identification and Expression Pattern Analysis of the TCP Transcription Factor Family in Ginkgo Biloba. Plant Signal. Behav. 2022, 17, 1994248. [Google Scholar] [CrossRef]
- Shang, X.; Han, Z.; Zhang, D.; Wang, Y.; Qin, H.; Zou, Z.; Zhou, L.; Zhu, X.; Fang, W.; Ma, Y. Genome-Wide Analysis of the TCP Gene Family and Their Expression Pattern Analysis in Tea Plant (Camellia sinensis). Front. Plant Sci. 2022, 13, 840350. [Google Scholar] [CrossRef]
- Zhou, H.; Hwarari, D.; Ma, H.; Xu, H.; Yang, L.; Luo, Y. Genomic Survey of TCP Transcription Factors in Plants: Phylogenomics, Evolution and Their Biology. Front. Genet. 2022, 13, 1060546. [Google Scholar] [CrossRef]
- Nicolas, M.; Cubas, P. TCP Factors: New Kids on the Signaling Block. Curr. Opin. Plant Biol. 2016, 33, 33–41. [Google Scholar] [CrossRef]
- Rath, M.; Challa, K.R.; Sarvepalli, K.; Nath, U. CINCINNATA-Like TCP Transcription Factors in Cell Growth–An Expanding Portfolio. Front. Plant Sci. 2022, 13, 825341. [Google Scholar] [CrossRef]
- Carrara, S.; Dornelas, M.C. TCP Genes and the Orchestration of Plant Architecture. Trop. Plant Biol. 2020, 14, 1–10. [Google Scholar] [CrossRef]
- Wang, M.; Le Moigne, M.-A.; Bertheloot, J.; Crespel, L.; Perez-Garcia, M.-D.; Ogé, L.; Demotes-Mainard, S.; Hamama, L.; Davière, J.-M.; Sakr, S. BRANCHED1: A Key Hub of Shoot Branching. Front. Plant Sci. 2019, 10, 76. [Google Scholar] [CrossRef] [PubMed]
- Sarvepalli, K.; Nath, U. Critical Review CIN-TCP Transcription Factors: Transiting Cell Proliferation in Plants. IUBMB Life 2018, 70, 718–731. [Google Scholar] [CrossRef] [PubMed]
- Kalve, S.; De Vos, D.; Beemster, G.T.S. Leaf Development: A Cellular Perspective. Front. Plant Sci. 2014, 5, 362. [Google Scholar] [CrossRef]
- Nakayama, H.; Leichty, A.R.; Sinha, N.R. Molecular Mechanisms Underlying Leaf Development, Morphological Diversification, and Beyond. Plant Cell 2022, 34, 2534–2548. [Google Scholar] [CrossRef]
- Lang, L.; Schnittger, A. Endoreplication—A Means to an End in Cell Growth and Stress Response. Curr. Opin. Plant Biol. 2020, 54, 85–92. [Google Scholar] [CrossRef]
- Kieffer, M.; Master, V.; Waites, R.; Davies, B. TCP14 and TCP15 Affect Internode Length and Leaf Shape in Arabidopsis. Plant J. 2011, 68, 147–158. [Google Scholar] [CrossRef]
- Li, Z.Y.; Li, B.; Dong, A.W. The Arabidopsis Transcription Factor AtTCP15 Regulates Endoreduplication by Modulating Expression of Key Cell-Cycle Genes. Mol. Plant 2012, 5, 270–280. [Google Scholar] [CrossRef] [PubMed]
- Aguilar-Martínez, J.A.; Sinha, N. Analysis of the Role of Arabidopsis Class I TCP Genes AtTCP7, AtTCP8, AtTCP22, and AtTCP23 in Leaf Development. Front. Plant Sci. 2013, 4, 406. [Google Scholar] [CrossRef] [PubMed]
- Davière, J.-M.; Wild, M.; Regnault, T.; Baumberger, N.; Eisler, H.; Genschik, P.; Achard, P. Class I TCP-DELLA Interactions in Inflorescence Shoot Apex Determine Plant Height. Curr. Biol. 2014, 24, 1923–1928. [Google Scholar] [CrossRef] [PubMed]
- Peng, Y.; Chen, L.; Lu, Y.; Wu, Y.; Dumenil, J.; Zhu, Z.; Bevan, M.W.; Lia, Y. The Ubiquitin Receptors DA1, DAR1, and DAR2 Redundantly Regulate Endoreduplication by Modulating the Stability of TCP14/15 in Arabidopsis. Plant Cell 2015, 27, 649–662. [Google Scholar] [CrossRef] [PubMed]
- Zhang, N.; Wang, Z.; Bao, Z.; Yang, L.; Wu, D.; Shu, X.; Hua, J. MOS1 Functions Closely with TCP Transcription Factors to Modulate Immunity and Cell Cycle in Arabidopsis. Plant J. 2018, 93, 66–78. [Google Scholar] [CrossRef] [PubMed]
- Zhang, G.; Zhao, H.; Zhang, C.; Li, X.; Lyu, Y.; Qi, D.; Cui, Y.; Hu, L.; Wang, Z.; Liang, Z.; et al. TCP7 Functions Redundantly with Several Class I TCPs and Regulates Endoreplication in Arabidopsis. J. Integr. Plant Biol. 2019, 61, 1151–1170. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Cochet, F.; Ponnaiah, M.; Lebreton, S.; Matheron, L.; Pionneau, C.; Boudsocq, M.; Resentini, F.; Huguet, S.; Blázquez, M.; et al. The MPK8-TCP14 Pathway Promotes Seed Germination in Arabidopsis. Plant J. 2019, 100, 677–692. [Google Scholar] [CrossRef] [PubMed]
- Resentini, F.; Felipo-Benavent, A.; Colombo, L.; Blázquez, M.A.; Alabadí, D.; Masiero, S. TCP14 and TCP15 Mediate the Promotion of Seed Germination by Gibberellins in Arabidopsis thaliana. Mol. Plant 2015, 8, 482–485. [Google Scholar] [CrossRef] [PubMed]
- Camoirano, A.; Arce, A.L.; Ariel, F.D.; Alem, A.L.; Gonzalez, D.H.; Viola, I.L. Class I TCP Transcription Factors Regulate Trichome Branching and Cuticle Development in Arabidopsis. J. Exp. Bot. 2020, 71, 5438–5453. [Google Scholar] [CrossRef] [PubMed]
- Hiratsu, K.; Matsui, K.; Koyama, T.; Ohme-Takagi, M. Dominant Repression of Target Genes by Chimeric Repressors That Include the EAR Motif, a Repression Domain, in Arabidopsis. Plant J. 2003, 34, 733–739. [Google Scholar] [CrossRef]
- Uberti-Manassero, N.G.; Lucero, L.E.; Viola, I.L.; Vegetti, A.C.; Gonzalez, D.H. The Class I Protein AtTCP15 Modulates Plant Development through a Pathway That Overlaps with the One Affected by CIN-like TCP Proteins. J. Exp. Bot. 2012, 63, 809–823. [Google Scholar] [CrossRef]
- Lucero, L.E.; Manavella, P.A.; Gras, D.E.; Ariel, F.D.; Gonzalez, D.H. Class I and Class II TCP Transcription Factors Modulate SOC1-Dependent Flowering at Multiple Levels. Mol. Plant 2017, 10, 1571–1574. [Google Scholar] [CrossRef]
- Li, X.; Zhang, G.; Liang, Y.; Hu, L.; Zhu, B.; Qi, D.; Cui, S.; Zhao, H. TCP7 Interacts with Nuclear Factor-Ys to Promote Flowering by Directly Regulating SOC1 in Arabidopsis. Plant J. 2021, 108, 1493–1506. [Google Scholar] [CrossRef]
- Ferrero, L.V.; Viola, I.L.; Ariel, F.D.; Gonzalez, D.H. Class I TCP Transcription Factors Target the Gibberellin Biosynthesis Gene GA20ox1 and the Growth-Promoting Genes HBI1 and PRE6 during Thermomorphogenic Growth in Arabidopsis. Plant Cell Physiol. 2019, 60, 1633–1645. [Google Scholar] [CrossRef]
- Ferrero, L.V.; Gastaldi, V.; Ariel, F.D.; Viola, I.L.; Gonzalez, D.H. Class I TCP Proteins TCP14 and TCP15 Are Required for Elongation and Gene Expression Responses to Auxin. Plant Mol. Biol. 2021, 105, 147–159. [Google Scholar] [CrossRef]
- Gastaldi, V.; Lucero, L.E.; Ferrero, L.V.; Ariel, F.D.; Gonzalez, D.H. Class-I TCP Transcription Factors Activate the SAUR63 Gene Subfamily in Gibberellin-Dependent Stamen Filament Elongation. Plant Physiol. 2020, 182, 2096–2110. [Google Scholar] [CrossRef]
- Xu, H.; Lantzouni, O.; Bruggink, T.; Benjamins, R.; Lanfermeijer, F.; Denby, K.; Schwechheimer, C.; Bassel, G.W. A Molecular Signal Integration Network Underpinning Arabidopsis Seed Germination. Curr. Biol. 2020, 30, 3703–3712.e4. [Google Scholar] [CrossRef]
- Alem, A.L.; Ariel, F.D.; Cho, Y.; Hong, J.C.; Gonzalez, D.H.; Viola, I.L. TCP15 Interacts with GOLDEN2-LIKE 1 to Control Cotyledon Opening in Arabidopsis. Plant J. 2022, 110, 748–763. [Google Scholar] [CrossRef]
- Gastaldi, V.; Alem, A.L.; Mansilla, N.; Ariel, F.D.; Viola, I.L.; Lucero, L.E.; Gonzalez, D.H. BREVIPEDICELLUS/KNAT1 Targets TCP15 to Modulate Filament Elongation during Arabidopsis Late Stamen Development. Plant Physiol. 2022, 191, 29–34. [Google Scholar] [CrossRef]
- Sun, T. Gibberellin-GID1-DELLA: A Pivotal Regulatory Module for Plant Growth and Development. Plant Physiol. 2010, 154, 567–570. [Google Scholar] [CrossRef]
- Ren, H.; Gray, W.M. SAUR Proteins as Effectors of Hormonal and Environmental Signals in Plant Growth. Mol. Plant 2015, 8, 1153–1164. [Google Scholar] [CrossRef]
- Bassel, G.W. To Grow or Not to Grow? Trends Plant Sci. 2016, 21, 498–505. [Google Scholar] [CrossRef]
- Tatematsu, K.; Nakabayashi, K.; Kamiya, Y.; Nambara, E. Transcription Factor AtTCP14 Regulates Embryonic Growth Potential during Seed Germination in Arabidopsis thaliana. Plant J. 2008, 53, 42–52. [Google Scholar] [CrossRef]
- Rueda-Romero, P.; Barrero-Sicilia, C.; Gómez-Cadenas, A.; Carbonero, P.; Oñate-Sánchez, L. Arabidopsis thaliana DOF6 Negatively Affects Germination in Non-after-Ripened Seeds and Interacts with TCP14. J. Exp. Bot. 2012, 63, 1937–1949. [Google Scholar] [CrossRef]
- Yeats, T.H.; Rose, J.K.C. The Formation and Function of Plant Cuticles. Plant Physiol. 2013, 163, 5–20. [Google Scholar] [CrossRef]
- Wagner, G.J.; Wang, E.; Shepherd, R.W. New Approaches for Studying and Exploiting an Old Protuberance, the Plant Trichome. Ann. Bot. 2004, 93, 3–11. [Google Scholar] [CrossRef]
- Serna, L.; Martin, C. Trichomes: Different Regulatory Networks Lead to Convergent Structures. Trends Plant Sci. 2006, 11, 274–280. [Google Scholar] [CrossRef]
- Tominaga-Wada, R.; Ishida, T.; Wada, T. New Insights into the Mechanism of Development of Arabidopsis Root Hairs and Trichomes. Int. Rev. Cell Mol. Biol. 2011, 286, 67–106. [Google Scholar] [CrossRef] [PubMed]
- Mathur, J. Trichome Cell Morphogenesis in Arabidopsis: A Continuum of Cellular DecisionsThis Review Is One of a Selection of Papers Published in the Special Issue on Plant Cell Biology. Can. J. Bot. 2006, 84, 604–612. [Google Scholar] [CrossRef]
- Ding, A.-M.; Xu, C.-T.; Xie, Q.; Zhang, M.-J.; Yan, N.; Dai, C.-B.; Lv, J.; Cui, M.-M.; Wang, W.-F.; Sun, Y.-H. ERF4 Interacts with and Antagonizes TCP15 in Regulating Endoreduplication and Cell Growth in Arabidopsis. J. Integr. Plant Biol. 2022, 64, 1673–1689. [Google Scholar] [CrossRef]
- Camoirano, A.; Alem, A.L.; Gonzalez, D.H.; Viola, I.L. Arabidopsis Thaliana TCP15 Interacts with the MIXTA-like Transcription Factor MYB106/NOECK. Plant Signal. Behav. 2021, 16, 1938432. [Google Scholar] [CrossRef]
- Wang, M.Y.; Zhao, P.M.; Cheng, H.Q.; Han, L.B.; Wu, X.M.; Gao, P.; Wang, H.Y.; Yang, C.L.; Zhong, N.Q.; Zuo, J.R.; et al. The Cotton Transcription Factor TCP14 Functions in Auxin-Mediated Epidermal Cell Differentiation and Elongation1. Plant Physiol. 2013, 162, 1669–1680. [Google Scholar] [CrossRef]
- Ma, Y.N.; Xu, D.B.; Yan, X.; Wu, Z.K.; Kayani, S.I.; Shen, Q.; Fu, X.Q.; Xie, L.H.; Hao, X.L.; Hassani, D.; et al. Jasmonate- and Abscisic Acid-Activated AaGSW1-AaTCP15/AaORA Transcriptional Cascade Promotes Artemisinin Biosynthesis in Artemisia annua. Plant Biotechnol. J. 2021, 19, 1412–1428. [Google Scholar] [CrossRef]
- Ma, Y.-N.; Xu, D.-B.; Li, L.; Zhang, F.; Fu, X.-Q.; Shen, Q.; Lyu, X.-Y.; Wu, Z.-K.; Pan, Q.-F.; Shi, P.; et al. Jasmonate Promotes Artemisinin Biosynthesis by Activating the TCP14-ORA Complex in Artemisia annua. Sci. Adv. 2018, 4, eaas9357. [Google Scholar] [CrossRef]
- Hou, X.; Zhou, J.; Liu, C.; Liu, L.; Shen, L.; Yu, H. Nuclear Factor Y-Mediated H3K27me3 Demethylation of the SOC1 Locus Orchestrates Flowering Responses of Arabidopsis. Nat. Commun. 2014, 5, 4601. [Google Scholar] [CrossRef]
- Ho, W.W.H.; Weigel, D. Structural Features Determining Flower-Promoting Activity of Arabidopsis FLOWERING LOCUS T. Plant Cell 2014, 26, 552–564. [Google Scholar] [CrossRef] [PubMed]
- Kubota, A.; Ito, S.; Shim, J.S.; Johnson, R.S.; Song, Y.H.; Breton, G.; Goralogia, G.S.; Kwon, M.S.; Cintró, D.L.; Koyama, T.; et al. TCP4-Dependent Induction of CONSTANS Transcription Requires GIGANTEA in Photoperiodic Flowering in Arabidopsis. PLoS Genet. 2017, 13, e1006856. [Google Scholar] [CrossRef]
- Liu, J.; Cheng, X.; Liu, P.; Li, D.; Chen, T.; Gu, X.; Sun, J. MicroRNA319-Regulated TCPs Interact with FBHs and PFT1 to Activate CO Transcription and Control Flowering Time in Arabidopsis. PLoS Genet. 2017, 13, e1006833. [Google Scholar] [CrossRef]
- Balsemão-Pires, E.; Andrade, L.R.; Sachetto-Martins, G. Functional Study of TCP23 in Arabidopsis Thaliana during Plant Development. Plant Physiol. Biochem. 2013, 67, 120–125. [Google Scholar] [CrossRef]
- Wu, J.-F.; Tsai, H.-L.; Joanito, I.; Wu, Y.-C.; Chang, C.-W.; Li, Y.-H.; Wang, Y.; Hong, J.C.; Chu, J.-W.; Hsu, C.-P.; et al. LWD–TCP Complex Activates the Morning Gene CCA1 in Arabidopsis. Nat. Commun. 2016, 7, 13181. [Google Scholar] [CrossRef]
- Wang, X.; Xu, X.; Mo, X.; Zhong, L.; Zhang, J.; Mo, B.; Kuai, B. Overexpression of TCP8 Delays Arabidopsis Flowering through a FLOWERING LOCUS C-Dependent Pathway. BMC Plant Biol. 2019, 19, 534. [Google Scholar] [CrossRef]
- Camoirano, A.; Alem, A.L.; Gonzalez, D.H.; Viola, I.L. The N-Terminal Region Located Upstream of the TCP Domain Is Responsible for the Antagonistic Action of the Arabidopsis thaliana TCP8 and TCP23 Transcription Factors on Flowering Time. Plant Sci. 2022, 328, 111571. [Google Scholar] [CrossRef]
- Kim, S.H.; Son, G.H.; Bhattacharjee, S.; Kim, H.J.; Nam, J.C.; Nguyen, P.D.T.; Hong, J.C.; Gassmann, W. The Arabidopsis Immune Adaptor SRFR1 Interacts with TCP Transcription Factors That Redundantly Contribute to Effector-Triggered Immunity. Plant J. 2014, 78, 978–989. [Google Scholar] [CrossRef]
- Li, M.; Chen, H.; Chen, J.; Chang, M.; Palmer, I.A.; Gassmann, W.; Liu, F.; Fu, Z.Q. Tcp Transcription Factors Interact with Npr1 and Contribute Redundantly to Systemic Acquired Resistance. Front. Plant Sci. 2018, 9, 1153. [Google Scholar] [CrossRef]
- Spears, B.J.; Howton, T.C.; Gao, F.; Garner, C.M.; Shahid Mukhtar, M.; Gassmann, W. Direct Regulation of the EFR-Dependent Immune Response by Arabidopsis TCP Transcription Factors. Mol. Plant Microbe Interact. 2019, 32, 540–549. [Google Scholar] [CrossRef]
- Viola, I.L.; Camoirano, A.; Gonzalez, D.H. Redox-Dependent Modulation of Anthocyanin Biosynthesis by the TCP Transcription Factor TCP15 during Exposure to High Light Intensity Conditions in Arabidopsis. Plant Physiol. 2016, 170, 74–85. [Google Scholar] [CrossRef]
- Viola, I.L.; Güttlein, L.N.; Gonzalez, D.H. Redox Modulation of Plant Developmental Regulators from the Class I TCP Transcription Factor Family. Plant Physiol. 2013, 162, 1434–1447. [Google Scholar] [CrossRef] [PubMed]
- Waters, M.T.; Wang, P.; Korkaric, M.; Capper, R.G.; Saunders, N.J.; Langdale, J.A. GLK Transcription Factors Coordinate Expression of the Photosynthetic Apparatus in Arabidopsis. Plant Cell 2009, 21, 1109–1128. [Google Scholar] [CrossRef]
- Sun, T.; Zhou, F.; Huang, X.-Q.; Chen, W.-C.; Kong, M.-J.; Zhou, C.-F.; Zhuang, Z.; Li, L.; Lu, S. ORANGE Represses Chloroplast Biogenesis in Etiolated Arabidopsis Cotyledons via Interaction with TCP14. Plant Cell 2019, 31, 2996–3014. [Google Scholar] [CrossRef] [PubMed]
- Zhao, H.; Bao, Y. PIF4: Integrator of Light and Temperature Cues in Plant Growth. Plant Sci. 2021, 313, 111086. [Google Scholar] [CrossRef] [PubMed]
- Mo, W.; Zhang, J.; Zhang, L.; Yang, Z.; Yang, L.; Yao, N.; Xiao, Y.; Li, T.; Li, Y.; Zhang, G.; et al. Arabidopsis Cryptochrome 2 Forms Photobodies with TCP22 under Blue Light and Regulates the Circadian Clock. Nat. Commun. 2022, 13, 2631. [Google Scholar] [CrossRef] [PubMed]
- Mazur, M.J.; Spears, B.J.; Djajasaputra, A.; Van Der Gragt, M.; Vlachakis, G.; Beerens, B.; Gassmann, W.; Van Den Burg, H.A. Arabidopsis TCP Transcription Factors Interact with the SUMO Conjugating Machinery in Nuclear Foci. Front. Plant Sci. 2017, 8, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Perez, M.; Guerringue, Y.; Ranty, B.; Pouzet, C.; Jauneau, A.; Robe, E.; Mazars, C.; Galaud, J.P.; Aldon, D. Specific TCP Transcription Factors Interact with and Stabilize PRR2 within Different Nuclear Sub-Domains. Plant Sci. 2019, 287, 110197. [Google Scholar] [CrossRef]
- Guan, P.; Ripoll, J.J.; Wang, R.; Vuong, L.; Bailey-Steinitz, L.J.; Ye, D.; Crawford, N.M. Interacting TCP and NLP Transcription Factors Control Plant Responses to Nitrate Availability. Proc. Natl. Acad. Sci. USA 2017, 114, 2419–2424. [Google Scholar] [CrossRef]
- Guan, P.; Wang, R.; Nacry, P.; Breton, G.; Kay, S.A.; Pruneda-Paz, J.L.; Davani, A.; Crawford, N.M. Nitrate Foraging by Arabidopsis Roots Is Mediated by the Transcription Factor TCP20 through the Systemic Signaling Pathway. Proc. Natl. Acad. Sci. USA 2014, 111, 15267–15272. [Google Scholar] [CrossRef]
- Chu, X.; Li, M.; Zhang, S.; Fan, M.; Han, C.; Xiang, F.; Li, G.; Wang, Y.; Xiang, C.; Wang, J.; et al. HBI1-TCP20 Interaction Positively Regulates the CEPs-mediated Systemic Nitrate Acquisition. J. Integr. Plant Biol. 2021, 63, 902–912. [Google Scholar] [CrossRef] [PubMed]
- Andrés-Colás, N.; Carrió-Seguí, A.; Abdel-Ghany, S.E.; Pilon, M.; Peñarrubia, L. Expression of the Intracellular COPT3-Mediated Cu Transport Is Temporally Regulated by the TCP16 Transcription Factor. Front. Plant Sci. 2018, 9, 910. [Google Scholar] [CrossRef]
- Perea-García, A.; Garcia-Molina, A.; Andrés-Colás, N.; Vera-Sirera, F.; Pérez-Amador, M.A.; Puig, S.; Peñarrubia, L. Arabidopsis Copper Transport Protein COPT2 Participates in the Cross Talk between Iron Deficiency Responses and Low-Phosphate Signaling. Plant Physiol. 2013, 162, 180–194. [Google Scholar] [CrossRef]
- Weßling, R.; Epple, P.; Altmann, S.; He, Y.; Yang, L.; Henz, S.R.; McDonald, N.; Wiley, K.; Bader, K.C.; Gläßer, C.; et al. Convergent Targeting of a Common Host Protein-Network by Pathogen Effectors from Three Kingdoms of Life. Cell Host Microbe 2014, 16, 364–375. [Google Scholar] [CrossRef] [PubMed]
- Lopez, J.A.; Sun, Y.; Blair, P.B.; Mukhtar, M.S. TCP Three-Way Handshake: Linking Developmental Processes with Plant Immunity. Trends Plant Sci. 2015, 20, 238–245. [Google Scholar] [CrossRef]
- Mukhtar, M.S.; Carvunis, A.-R.; Dreze, M.; Epple, P.; Steinbrenner, J.; Moore, J.; Tasan, M.; Galli, M.; Hao, T.; Nishimura, M.T.; et al. Independently Evolved Virulence Effectors Converge onto Hubs in a Plant Immune System Network. Science 2011, 333, 596–601. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Cui, D.; Liu, C.; Zhao, J.; Liu, J.; Liu, N.; Tang, D.; Hu, Y. TCP transcription factors interact with ZED1-related kinases as components of the temperature-regulated immunity. Plant Cell Environ. 2019, 42, 2045–2056. [Google Scholar] [CrossRef]
- Wang, X.; Gao, J.; Zhu, Z.; Dong, X.; Wang, X.; Ren, G.; Zhou, X.; Kuai, B. TCP Transcription Factors Are Critical for the Coordinated Regulation of ISOCHORISMATE SYNTHASE 1 Expression in Arabidopsis thaliana. Plant J. 2015, 82, 151–162. [Google Scholar] [CrossRef] [PubMed]
- Danisman, S.; Van Der Wal, F.; Dhondt, S.; Waites, R.; De Folter, S.; Bimbo, A.; Dj Van Dijk, A.; Muino, J.M.; Cutri, L.; Dornelas, M.C.; et al. Arabidopsis Class I and Class II TCP Transcription Factors Regulate Jasmonic Acid Metabolism and Leaf Development Antagonistically. Plant Physiol. 2012, 159, 1511–1523. [Google Scholar] [CrossRef]
- Yang, L.; Teixeira, P.J.P.L.; Biswas, S.; Finkel, O.M.; He, Y.; Salas-Gonzalez, I.; English, M.E.; Epple, P.; Mieczkowski, P.; Dangl, J.L. Pseudomonas Syringae Type III Effector HopBB1 Promotes Host Transcriptional Repressor Degradation to Regulate Phytohormone Responses and Virulence. Cell Host Microbe 2017, 21, 156–168. [Google Scholar] [CrossRef]
- Xiao, O.; Lin, W.; Feng, E.; Wu, C.; Ou, X. Genome-Wide Identification of Binding Sites for SmTCP7a Transcription Factors of Eggplant during Bacterial Wilt Resistance by ChIP-Seq. Int. J. Mol. Sci. 2022, 23, 6844. [Google Scholar] [CrossRef]
- Willig, J.J.; Guarneri, N.; van Steenbrugge, J.J.M.; de Jong, W.; Chen, J.; Goverse, A.; Lozano Torres, J.L.; Sterken, M.G.; Bakker, J.; Smant, G. The Arabidopsis Transcription Factor TCP9 Modulates Root Architectural Plasticity, Reactive Oxygen Species-Mediated Processes, and Tolerance to Cyst Nematode Infections. Plant J. 2022, 112, 1070–1083. [Google Scholar] [CrossRef] [PubMed]
- Ceulemans, E.; Ibrahim, H.M.M.; De Coninck, B.; Goossens, A. Pathogen Effectors: Exploiting the Promiscuity of Plant Signaling Hubs. Trends Plant Sci. 2021, 26, 780–795. [Google Scholar] [CrossRef]
- Sugio, A.; Kingdom, H.N.; MacLean, A.M.; Grieve, V.M.; Hogenhout, S.A. Phytoplasma Protein Effector SAP11 Enhances Insect Vector Reproduction by Manipulating Plant Development and Defense Hormone Biosynthesis. Proc. Natl. Acad. Sci. USA 2011, 108, E1254–E1263. [Google Scholar] [CrossRef] [PubMed]
- González-Fuente, M.; Carrère, S.; Monachello, D.; Marsella, B.G.; Cazalé, A.C.; Zischek, C.; Mitra, R.M.; Rezé, N.; Cottret, L.; Mukhtar, M.S.; et al. EffectorK, a Comprehensive Resource to Mine for Ralstonia, Xanthomonas, and Other Published Effector Interactors in the Arabidopsis Proteome. Mol. Plant Pathol. 2020, 21, 1257–1270. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Zhao, Y.; Luo, X.; Hong, H.; Yang, T.; Huang, S.; Wang, C.; Chen, H.; Qian, X.; Feng, M.; et al. NLR Surveillance of Pathogen Interference with Hormone Receptors Induces Immunity. Nature 2022, 613, 145–152. [Google Scholar] [CrossRef]
- Liu, H.; Gao, Y.; Wu, M.; Shi, Y.; Wang, H.; Wu, L.; Xiang, Y. TCP10, a TCP Transcription Factor in Moso Bamboo (Phyllostachys edulis), Confers Drought Tolerance to Transgenic Plants. Environ. Exp. Bot. 2020, 172, 104002. [Google Scholar] [CrossRef]
- Xu, Y.; Liu, H.; Gao, Y.; Xiong, R.; Wu, M.; Zhang, K.; Xiang, Y. The TCP Transcription Factor PeTCP10 Modulates Salt Tolerance in Transgenic Arabidopsis. Plant Cell Rep. 2021, 40, 1971–1987. [Google Scholar] [CrossRef]
- Mishra, S.; Sahu, G.; Prasad Shaw, B.; Sagarika Mishra, C. Insight into the Cellular and Physiological Regulatory Modulations of Class-I TCP9 to Enhance Drought and Salinity Stress Tolerance in Cowpea. Physiol. Plant. 2022, 174, e13542. [Google Scholar] [CrossRef]
- Wang, K.; Zhang, N.; Fu, X.; Zhang, H.; Liu, S.; Pu, X.; Wang, X.; Si, H. StTCP15 Regulates Potato Tuber Sprouting by Modulating the Dynamic Balance between Abscisic Acid and Gibberellic Acid. Front. Plant Sci. 2022, 13, 1009552. [Google Scholar] [CrossRef]
- Almeida, D.M.; Gregorio, G.B.; Oliveira, M.M.; Saibo, N.J.M. Five Novel Transcription Factors as Potential Regulators of OsNHX1 Gene Expression in a Salt Tolerant Rice Genotype. Plant Mol. Biol. 2017, 93, 61–77. [Google Scholar] [CrossRef]
- Wu, J.; Wu, W.; Liang, J.; Jin, Y.; Gazzarrini, S.; He, J.; Yi, M. GhTCP19 Transcription Factor Regulates Corm Dormancy Release by Repressing GhNCED Expression in Gladiolus. Plant Cell Physiol. 2019, 60, 52–62. [Google Scholar] [CrossRef]
- Dong, H.; Dumenil, J.; Lu, F.H.; Na, L.; Vanhaeren, H.; Naumann, C.; Klecker, M.; Prior, R.; Smith, C.; McKenzie, N.; et al. Ubiquitylation Activates a Peptidase That Promotes Cleavage and Destabilization of Its Activating E3 Ligases and Diverse Growth Regulatory Proteins to Limit Cell Proliferation in Arabidopsis. Genes Dev. 2017, 31, 197–208. [Google Scholar] [CrossRef]
- Steiner, E.; Efroni, I.; Gopalraj, M.; Saathoff, K.; Tseng, T.S.; Kieffer, M.; Eshed, Y.; Olszewski, N.; Weiss, D. The Arabidopsis O-Linked N-Acetylglucosamine Transferase SPINDLY Interacts with Class I TCPs to Facilitate Cytokinin Responses in Leaves and Flowers. Plant Cell 2012, 24, 96–108. [Google Scholar] [CrossRef]
- Steiner, E.; Livne, S.; Kobinson-Katz, T.; Tal, L.; Pri-Tal, O.; Mosquna, A.; Tarkowská, D.; Mueller, B.; Tarkowski, P.; Weiss, D. The Putative O-Linked N-Acetylglucosamine Transferase SPINDLY Inhibits Class I TCP Proteolysis to Promote Sensitivity to Cytokinin. Plant Physiol. 2000, 171, 1485–1494. [Google Scholar] [CrossRef]
- Steiner, E.; Triana, M.R.; Kubasi, S.; Blum, S.; Paz-Ares, J.; Rubio, V.; Weiss, D. KISS ME DEADLY F-Box Proteins Modulate Cytokinin Responses by Targeting the Transcription Factor TCP14 for Degradation. Plant Physiol. 2021, 185, 1495–1499. [Google Scholar] [CrossRef]
- Zentella, R.; Sui, N.; Barnhill, B.; Hsieh, W.P.; Hu, J.; Shabanowitz, J.; Boyce, M.; Olszewski, N.E.; Zhou, P.; Hunt, D.F.; et al. The Arabidopsis O-Fucosyltransferase SPINDLY Activates Nuclear Growth Repressor Della. Nat. Chem. Biol. 2017, 13, 479–485. [Google Scholar] [CrossRef]
- Wang, Y.; He, Y.; Su, C.; Zentella, R.; Sun, T.; Wang, L. Nuclear Localized O-Fucosyltransferase SPY Facilitates PRR5 Proteolysis to Fine-Tune the Pace of Arabidopsis Circadian Clock. Mol. Plant 2020, 13, 446–458. [Google Scholar] [CrossRef] [PubMed]
- Liang, L.; Wang, Q.; Song, Z.; Wu, Y.; Liang, Q.; Wang, Q.; Yang, J.; Bi, Y.; Zhou, W.; Fan, L.M. O-Fucosylation of CPN20 by SPINDLY Derepresses Abscisic Acid Signaling During Seed Germination and Seedling Development. Front. Plant Sci. 2021, 12, 724144. [Google Scholar] [CrossRef]
- Xu, S.L.; Chalkley, R.J.; Maynard, J.C.; Wang, W.; Ni, W.; Jiang, X.; Shin, K.; Cheng, L.; Savage, D.; Hühmer, A.F.R.; et al. Proteomic Analysis Reveals O-GlcNAc Modification on Proteins with Key Regulatory Functions in Arabidopsis. Proc. Natl. Acad. Sci. USA 2017, 114, E1536–E1543. [Google Scholar] [CrossRef] [PubMed]
- Purayannur, S.; Kumar, K.; Kaladhar, V.C.; Verma, P.K. Phylogenomic Analysis of MKKs and MAPKs from 16 Legumes and Detection of Interacting Pairs in Chickpea Divulge MAPK Signalling Modules. Sci. Rep. 2017, 7, 5026. [Google Scholar] [CrossRef] [PubMed]
- Masuda, H.P.; Cabral, L.M.; De Veylder, L.; Tanurdzic, M.; De Almeida Engler, J.; Geelen, D.; Inzé, D.; Martienssen, R.A.; Ferreira, P.C.G.; Hemerly, A.S. ABAP1 Is a Novel Plant Armadillo BTB Protein Involved in DNA Replication and Transcription. EMBO J. 2008, 27, 2746–2756. [Google Scholar] [CrossRef]
- Pruneda-Paz, J.L.; Breton, G.; Para, A.; Kay, S.A. A Functional Genomics Approach Reveals CHE as a Component of the Arabidopsis Circadian Clock. Science 2009, 323, 1481–1485. [Google Scholar] [CrossRef]
- Chen, G.-H.; Sun, J.-Y.; Liu, M.; Liu, J.; Yang, W.-C. SPOROCYTELESS Is a Novel Embryophyte-Specific Transcription Repressor That Interacts with TPL and TCP Proteins in Arabidopsis. J. Genet. Genom. 2014, 41, 617–625. [Google Scholar] [CrossRef]
- Schippers, J.H.; Foyer, C.H.; van Dongen, J.T. Redox Regulation in Shoot Growth, SAM Maintenance and Flowering. Curr. Opin. Plant Biol. 2016, 29, 121–128. [Google Scholar] [CrossRef]
- Eljebbawi, A.; del Guerrero, Y.C.R.; Dunand, C.; Estevez, J.M. Highlighting Reactive Oxygen Species as Multitaskers in Root Development. iScience 2021, 24, 101978. [Google Scholar] [CrossRef]
- Hammani, K.; Gobert, A.; Hleibieh, K.; Choulier, L.; Small, I.; Giegé, P. An Arabidopsis Dual-Localized Pentatricopeptide Repeat Protein Interacts with Nuclear Proteins Involved in Gene Expression Regulation. Plant Cell 2011, 23, 730–740. [Google Scholar] [CrossRef] [PubMed]
- Spears, B.J.; McInturf, S.A.; Collins, C.; Chlebowski, M.; Cseke, L.J.; Su, J.; Mendoza-Cózatl, D.G.; Gassmann, W. Class I TCP Transcription Factor AtTCP8 Modulates Key Brassinosteroid-Responsive Genes. Plant Physiol. 2022, 190, 1457–1473. [Google Scholar] [CrossRef] [PubMed]
- Karaaslan, E.S.; Wang, N.; Faiß, N.; Liang, Y.; Montgomery, S.A.; Laubinger, S.; Berendzen, K.W.; Berger, F.; Breuninger, H.; Liu, C. Marchantia TCP Transcription Factor Activity Correlates with Three-Dimensional Chromatin Structure. Nat. Plants 2020, 6, 1250–1261. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Viola, I.L.; Alem, A.L.; Jure, R.M.; Gonzalez, D.H. Physiological Roles and Mechanisms of Action of Class I TCP Transcription Factors. Int. J. Mol. Sci. 2023, 24, 5437. https://doi.org/10.3390/ijms24065437
Viola IL, Alem AL, Jure RM, Gonzalez DH. Physiological Roles and Mechanisms of Action of Class I TCP Transcription Factors. International Journal of Molecular Sciences. 2023; 24(6):5437. https://doi.org/10.3390/ijms24065437
Chicago/Turabian StyleViola, Ivana L., Antonela L. Alem, Rocío M. Jure, and Daniel H. Gonzalez. 2023. "Physiological Roles and Mechanisms of Action of Class I TCP Transcription Factors" International Journal of Molecular Sciences 24, no. 6: 5437. https://doi.org/10.3390/ijms24065437
APA StyleViola, I. L., Alem, A. L., Jure, R. M., & Gonzalez, D. H. (2023). Physiological Roles and Mechanisms of Action of Class I TCP Transcription Factors. International Journal of Molecular Sciences, 24(6), 5437. https://doi.org/10.3390/ijms24065437




