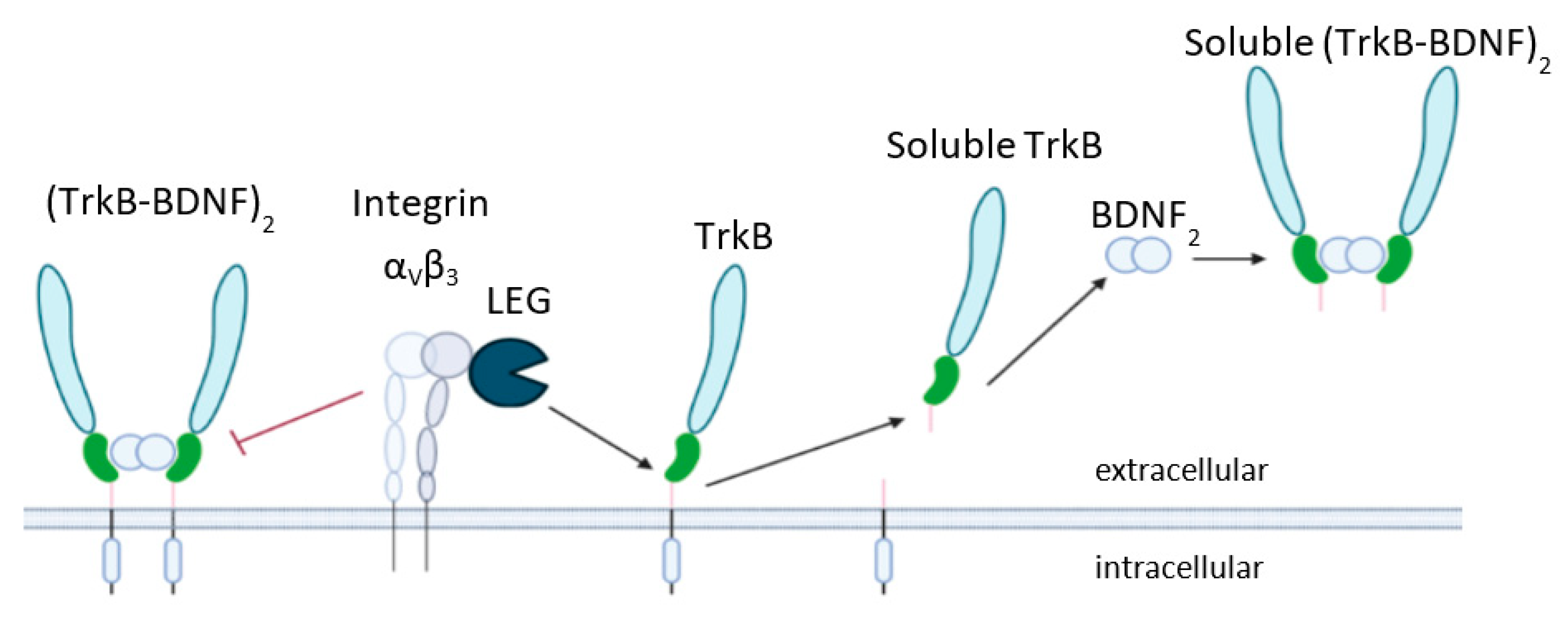
1. Introduction
Cell signaling allows the transfer of information between cells to coordinate different tissues and cell types [
1,
2].
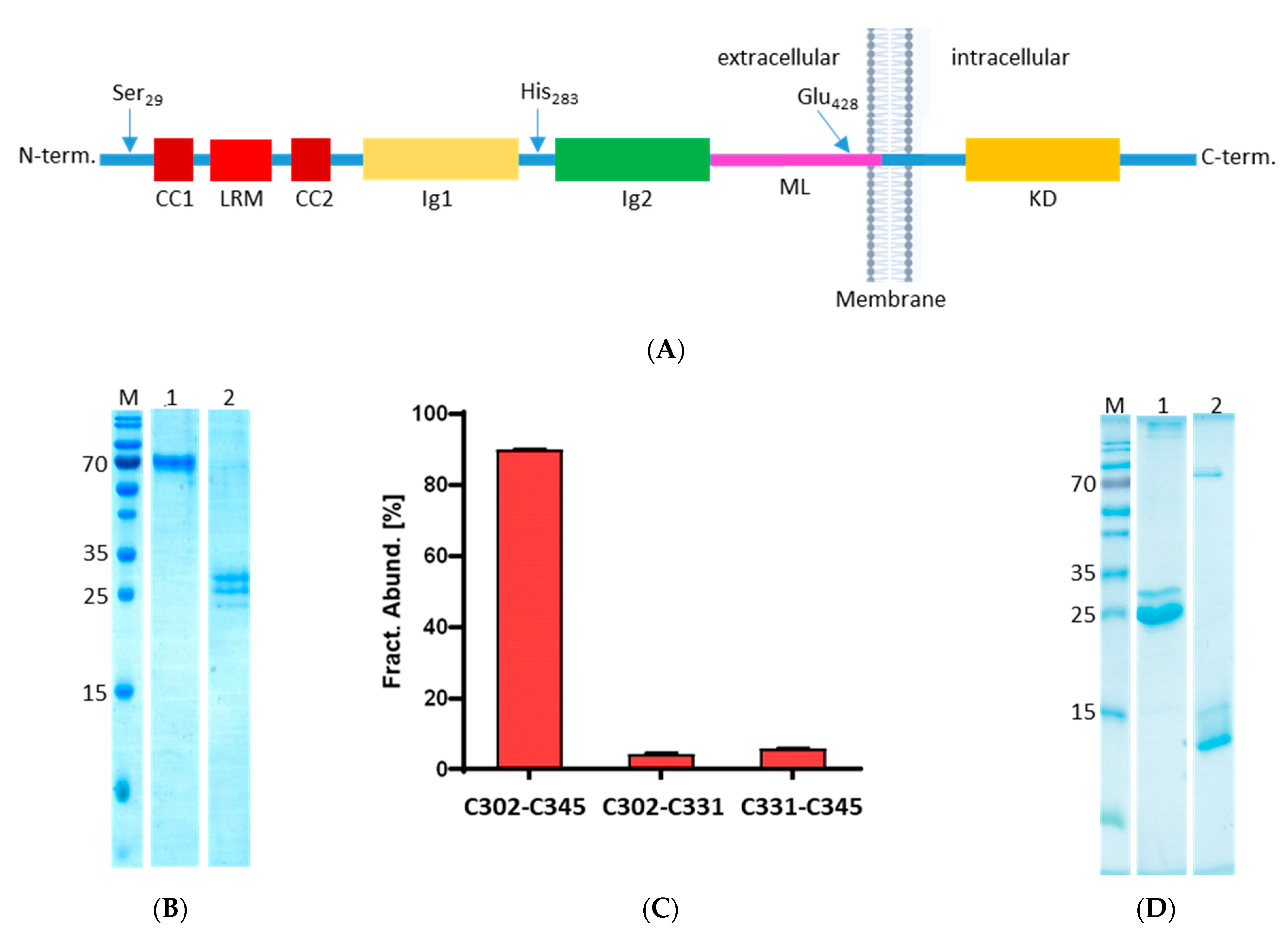
Of particular relevance are neuronal receptors such as the enzyme-coupled receptors p75NTR and Tyrosine-receptor-kinase B (TrkB), which transduce apoptosis and survival signals, respectively. These receptors work in concert with each other [
3]. By these interactions, TrkB determines the neuronal fate concerning proliferation and survival; axonal and dendritic growth as well as remodeling; development of neuronal cytoskeleton; membrane traffic and synapse formation; and neuronal function and plasticity [
4,
5]. An important downstream effector protein of TrkB signaling is legumain (LEG). Legumain is a clan CD, family C13 cysteine protease [
6], primarily located in the endo-lysosome [
7]. There it fulfills important immunologic functions, e.g., producing immunopeptide for MHCII loading [
7,
8]. Legumain can also escape the lysosome primarily during pathophysiologic conditions like cancer and neurodegenerative diseases. There it is found to be active in the cytoplasm and extracellular space [
9,
10,
11,
12,
13]. On the one hand, legumain’s expression is suppressed by TrkB signaling [
14]. On the other hand, in the absence of TrkB/BDNF (Brain-derived neurotrophic factor) signaling, the transcription factor C/EBPβ is upregulated, resulting in increased legumain expression [
14]. Furthermore, legumain-cleaved tau protein antagonizes TrkB signaling, [
15].
Known physiologic regulators of TrkB are the neurotrophins BDNF and NT4, which specifically bind TrkB and induce its dimerization [
5]. Additionally, TrkB is processed at its extracellular domain during excitotoxic events, whereby the exact identity of the involved sheddases remains unclear [
16]. While sheddase activity is mostly mediated by membrane-anchored proteases, also soluble proteases contribute to shedding activity. Indeed, legumain was previously shown to act as δ-secretase on amyloid precursor protein (APP) processing without being membrane-bound [
17,
18].
Combining the above-mentioned information led us to explore the possibility of TrkB being processed by legumain in its ectodomain and its potential impact on TrkB binding to its specific binding partner BDNF. Here we identify legumain cleavage sites in the ectodomain of TrkB and show that this processing can create soluble TrkB isoforms with the ability to bind BDNF as well as the pro-form of BDNF. Additionally, we also show that the complex formation of TrkB and BDNF prevents the processing by legumain.
4. Materials and Methods
4.1. Materials
The following chemicals were bought from Merck Millipore (Darmstadt, Germany): MgSO4, ammonium-acetate, citric acid, KCl, Na2HPO4, KH2PO4, HCl, and EDTA. The following chemicals were bought from Applichem (Darmstadt, Germany): Triton X100, Guanidine, Glutathione, HEPES, CaCl2, Urea, and NaCl. The following chemicals were purchased from Sigma-Aldrich (Vienna, Austria): Tris-Base, β-mercaptoethanol, L-Arginine, and formic acid. Tween-20 was obtained from Roth (Karlsruhe, Germany), acetonitrile from VWR Chemicals (Radnor, PA, USA) and imidazole from NeoLab Migge (Heidelberg, Germany). The pHLsec plasmids were kindly provided by A. R. Aricescu (Cambridge, UK).
4.2. Expression of TrkB Variants in HEK293S Cells and Purification
Cloning. The coding sequences for the TrkB variants TrkB ECD and TrkB Ig2 were based on the according NCBI RefSeq NM_012731.2 and synthesized (Eurofins Genomics, Munich, Germany). The sequences were cloned into a pHLsec plasmid with restriction digestion by Age1 and Nde1 (New England Biolabs, Ipswich, MA, USA) and T4 DNA Ligase ligation (Thermo Fisher Scientific, Vienna, Austria). The pHLsec plasmid also adds a C-terminal His6-tag to the TrkB variants. The correct insertion and orientation of the coding sequence for TrkB ECD and TrkB Ig2 were confirmed by sequencing (Eurofins Genomics, Germany).
Expression. Expression of the TrkB variants is based on the work of Aricescu et al. [
37]. In short, adherent HEK293S cells were passaged and expanded in T-175 flasks (Greiner Bio-One, Kremsmünster, Austria) until ~90% confluency. Cells were maintained in DMEM medium (Sigma Aldrich, Vienna, Austria) supplemented with 10% fetal bovine serum (Gibco, Carlsbad, CA, USA), non-essential amino acids (Sigma Aldrich, Vienna, Austria) and L-glutamine (Sigma Aldrich, Vienna, Austria) at 37 °C, 5% CO
2. 5 mL serum-free medium was mixed with 85 µg of TrkB plasmid DNA and 175 µL PEI stock (1 mg/mL) to allow DNA-PEI complex formation. Transfection was achieved by adding the mix to the confluent cells in 30 mL 2% FBS medium, briefly shaking the flask and putting it back into the incubator for 24 h. After that, the 2% FBS medium was removed and replaced by a 30 mL serum-free medium. Medium with secreted soluble TrkB variants was harvested after 72 h.
Purification. Harvested medium (30 mL per T-175 flask) was diluted 1:5 with wash buffer (50 mM Tris, 300 mM NaCl, pH 7.5) and incubated together with 3 mL Ni2+-NTA (Qiagen, Hilden, Germany) at 4 °C for 60 min. The flow-through was separated from the beads, and the beads were washed 2 times with 10 mL wash buffer. TrkB variants were eluted with 2–4 times 3 mL elution buffer (50 mM Tris, 300 mM NaCl, 250 mM Imidazole, pH 7.5). The elutions were pooled and concentrated with a 3 kDa Amicon (Merck Millipore, Darmstadt, Germany) to ~4 mg/mL.
4.3. Expression of Neurotrophin BDNF in E. coli as Inclusion Bodies, Folding and Purification
Cloning. The coding sequence for proBDNF was taken from the ENA database (#AAA63483) and synthesized (Eurofins Genomics, Germany). The sequences were cloned into a pET22b plasmid (Merck Millipore, Darmstadt, Germany) with restriction digestion by Xho1 and Nde1 (New England Biolabs, Ipswich, MA, USA) and T4 DNA Ligase ligation. The pET22b plasmid also adds a C-terminal His6-tag to proBDNF. The correct insertion and orientation of the coding sequence for proBDNF were confirmed by sequencing (Eurofins Genomics, Germany).
Expression. Rosetta 2 (DE3) E. coli (Merck Millipore, Darmstadt, Germany) were transformed with the plasmid and grown as a multiclonal culture in 50 mL standard LB medium (Roth, Karlsruhe, Germany) with 100 µg/mL Ampicillin and 20 µg/mL Chloramphenicol at 37 °C (both Applichem, Darmstadt, Germany). 600 mL LB-Ampicilin medium was inoculated with 3–4 mL of overnight culture. These cultures were grown at 37 °C to an OD600nm of 0.8–1.0. Expression was induced with 1 mM IPTG (ForMedium, Norfolk, UK), and the culture was further incubated at 37 °C for 3–4 h under continued shaking. Cells were harvested at 4000× g for 10 min at 4 °C. Pellets of 3–4 harvests were pooled and stored at −20 °C.
Inclusion Body Preparation. The pooled pellet was resuspended in at least 30 mL wash buffer 1 (50 mM Tris, 500 mM NaCl, 20 mM EDTA, pH 8.0) and sonicated (Bandelin, Berlin, Germany) at near 100% power, 50% duration, 9 intervals per 3 min on ice. A spatula tip of DNase (Applichem, Darmstadt, Germany) and 5 mM MgSO4 was added and incubated for 30 min at 4 °C on a rolling incubator. 2% Triton X-100 was added and manually shaken until the Triton was dissolved. The resuspended pellet was centrifuged at 17,500×
g for 15 min at 4 °C, and the supernatant was discarded. Next, the pellet was washed 2 times with 4 mL per gram pellet wash buffer 2 (50 mM Tris, 500 mM NaCl, 20 mM EDTA, pH 8.0, 2% Triton X-100) followed by 2 washes with wash buffer 1. Between each wash step, the sample was centrifuged at 17,500×
g for 15 min at 4 °C and resuspended with a Potter homogenizer. The final pellet was weight. Per gram pellet, 10 mL inclusion body solubilization buffer 1 (6 M guanidine-HCL, 50 mM Tris, 20 mM EDTA, 100 mM β-mercaptoethanol, pH 8.5) was added and stirred until the pellet was dissolved. After that, 2 M HCl was added until a pH of approximately 3.5 was reached (pH strips). The solution was centrifuged at 17,500×
g for 30 min at RT. Next, the supernatant was transferred into a 10 kDa cutoff dialysis membrane (Repligen, DG Breda, The Netherlands) and dialyzed overnight against 2 L dialysis buffer 1 (50 mM NaCl, 10 mM EDTA, pH 4.5). The dialysis buffer was changed 2 times during this time. The dialysis product was centrifuged at 17,500×
g for 30 min at RT, and the supernatant was discarded. The resulting inclusion body pellet is dissolved in 4–6 mL per gram pellet solubilization buffer 2 (6 M guanidine-HCl, 50 mM Tris, 20 mM EDTA, pH 3.5) followed by centrifugation at 17,500×
g for 20 min at RT. The supernatant was transferred into a new 15 mL tube, and the concentration was measured with a Bradford Assay [
38]. Preferred concentration for (re-) folding: 15–20 mg/mL or more.
Folding. Folding of proBDNF from inclusion bodies was done by rapid dilution. For this, 50% of the inclusion body supernatant was dripped into 100 times the total supernatant volume of 4 °C cold folding buffer (0.75 M L-Arginine, 100 mM Tris, 1 mM EDTA, pH 9.0, GSH:GSSG 10:1) under constant slow stirring. After 4 h, the remaining 50% was slowly added, and stirring was continued for at least 4 more hours or overnight. Next, the folding solution was concentrated on ice with a peristaltic 5 kDa filter pump (Heidolph Instruments, Schwabach, Germany) and then dialyzed (Nadir membrane, Roth, Karlsruhe, Germany) against 5 L dialysis buffer 2 (20 mM HEPES, 100 mM NaCl, pH 7.0, 4 °C and RT). For thorough removal of guanidine, arginine, Tris, GSH, and GSSG, dialysis was carried out as follows: First dialysis for 2 h at RT, second dialysis overnight at 4 °C, last for 2 h at RT. For each dialysis step, the membrane tube containing the protein was shortly opened and mixed by pipetting and then transferred into a fresh dialysis buffer. After that, the dialysis product was centrifuged at 17,500× g for 15–30 min at 4 °C. The supernatant was transferred into a fitting tube or flask, and the concentration was measured with Bradford Assay.
proBDNF Purification. Purification of folded proBDNF was achieved by SP-sepharose cation exchange chromatography (Cytiva Lifesciences, Vienna, Austria). Approximately 2.5 mL of SP-sepharose beads were equilibrated with equilibration buffer (20 mM HEPES, 50 mM NaCl, pH 7.0) at 4 °C and transferred into the folding dialysis supernatant under constant slow stirring for 60 min. After, the suspension was transferred back into the column to collect the sepharose beads with bound proBDNF. At least 2 wash steps (20 mM HEPES, 100 mM NaCl, pH 7.0) with 4 times the column volume were carried out. Elution was achieved by adding 2.5 mL of elution buffer (20 mM HEPES, 500 mM NaCl, pH 7.0) and incubation on a roller incubator (CAT, Ballrechten-Dottingen, Germany) for 10 min at 4 °C. Elution steps were repeated at least 4 times. The concentration of each elution was measured with Bradford Assay. To decrease the high salt concentration of the elution buffer, the samples were rebuffered into the wash buffer over an NAP column (GE Healthcare, Vienna, Austria). The concentration was again measured with Bradford Assay to determine losses.
Activation. The rebuffered elutions were pooled, and 5 mM CaCl2 and hFurin in a ratio of 1:100 (m/m) were added. The mixture was incubated on a roller incubator for at least 3.5–4 h at RT. To stop the activation of proBDNF by Furin, 10 mM EDTA (pH 7.4) was added, and the mixture was centrifuged at 17,500× g for 15 min at 4 °C.
BDNF Purification. The activation assay was immediately loaded on a 1 mL SP-sepharose cation exchange column and incubated for 60 min at 4 °C. Washing, Elution and NAP-rebuffering of BDNF was carried out as described during proBDNF purification, except that the elution buffer contained 1 M salt instead of just 500 mM and the buffer for rebuffering was 1x PBS pH 7.4.
4.4. HPLC-MS Analysis for TrkB Ig2 and BDNF
General HPLC-MS parameters. For all chromatographic analyses, mobile phase A was H2O + 0.10% formic acid, and mobile phase B was acetonitrile + 0.10% formic acid. Chromatographic separations for intact Ig2, legumain-digested Ig2, peptide and disulfide mapping of Ig2 as well as intact proBDNF were carried out on a Discovery BIO wide pore C18 column (150 × 2.1 mm i.d., 3.0 μm particle size, 300 Å pore size, Supelco, Bellefonte, PE, USA). While queued for analysis, samples were stored in the autosampler at 4.0 °C. The mass spectrometers were mass calibrated using Pierce™ LTQ Velos ESI Positive Ion Calibration Solution from Life Technologies (Vienna, Austria).
Analysis of intact Ig2, legumain-digested Ig2, peptide and disulfide mapping of Ig2. For the MS analysis, Ig2 samples were precipitated by using ice-cold methanol and resuspended using 150 mM ammonium acetate. Trypsin was added in an enzyme-protein ratio of 1:20 and allowed to digest the protein at 37 °C overnight at 900 rpm. The reaction was stopped by the addition of 10% formic acid to a final concentration of 1%. Chromatographic separation was carried out on a capillary HPLC instrument (UltiMate™ U3000 RSLC, Thermo Fisher Scientific, Germering, Germany). Ten microliters of intact Ig2 and legumain-digested Ig2 samples [0.25 mg.mL−1] were injected using in-line split-loop mode and separated at a flow rate of 200 µL.min−1 and a column oven temperature of 70 °C applying the following linear gradient: 5.0% B for 10.0 min, 5.0–40.0% B for 20 min, 40.0–80.0% B for 5.0 min, 80.0% B for 5.0 min and 5.0% B for 20 min. UV detection was carried out at 214 nm using a 1.4 µL flow cell.
All samples were analyzed on a Thermo Scientific™ QExactive™ benchtop quadrupole-Orbitrap
® mass spectrometer equipped with an Ion Max™ source with heated electrospray ionization (HESI) probe, both from Thermo Fisher Scientific (Bremen, Germany), and an MXT715-000-MX Series II Switching Valve (IDEX Health & Science LLC, Oak Harbor, WA, USA). Mass spectrometric data were acquired between minute 10.0 and 55.0 in an
m/
z range of 500–3000 with instrument settings as specified earlier [
39]. Intact Ig2 was measured at a resolution of 17,500 at
m/
z 200, and legumain-digested Ig2 was analyzed at a resolution of 140,000 at
m/
z 200. MS/MS of intact and legumain-digested Ig2 was carried out applying all ion fragmentation (AIF) in the higher-energy collisional dissociation (HCD) cell at normalized collision energy (NCE) settings of 22.0 within a scan range of
m/
z 400–2500 and a resolution setting of 140,000 at
m/
z 200. Peptide and disulfide mapping was conducted using data-dependent MS/MS. Each scan cycle consisted of a full scan at a scan range of
m/
z 400–2000 with an AGC target of 3 × 10
6, a maximum injection time of 100 ms and a resolution setting of 70,000, followed by 5 data-dependent HCD scans at 30 NCE with an AGC target of 5 × 10
5, a maximum injection time of 200 ms and a resolution setting of 17,500. The dynamic exclusion was set to 10.0 s.
Isotopically resolved mass spectra were deconvoluted using the Xtract algorithm implemented in the Xcalibur™ software version 3.0.63 (Thermo Fisher Scientific, Waltham, MA, USA). Isotopically unresolved spectra were deconvoluted using the Respect algorithm implemented in Biopharma Finder 1.0 (Thermo Fisher Scientific, Waltham, MA, USA). AIF data were evaluated using the software ProSight Lite v1.4 Build 1.4.6 provided by the Kelleher Research Group (Northwestern University, Evanston, IL, USA); mass tolerance for annotation of b- and y-fragments was set to 25 ppm. Peptide mapping data were evaluated using Biopharma Finder 1.0.
Analysis of proBDNF and tryptic peptides thereof. For intact mass analysis, 300 µg of TCA precipitated proBDNF were dissolved in 175.0 mmol L−1 ammonium acetate. For disulfide mapping, 400 µg of TCA-precipitated proBDNF were dissolved in 100 µL 175.0 mmol L 1 ammonium acetate. Trypsin (1:20 enzyme protein ratio, Promega, Mannheim, Germany) was directly added and allowed to digest overnight at 37 °C and 900 rpm. The reaction was stopped by the addition of FA to a final concentration of 1%.
Intact proBDNF analysis was carried out in a Vanquish HPLC system (Thermo Fisher Scientific, Germering, Germany). Five microlitres of the sample were injected and separated during a linear gradient at 60.0 °C and a constant flow rate of 100.0 µL min−1: 15.0% B for 3.0 min, 15.0–90.0% B for 27.0 min, 90.0–99.0% B for 0.1 min, 99.0% B for 5.0 min, 99.0–5.0% B for 0.1 min, 5.0% B for 5.0 min.
Tryptic peptides were analyzed on a nano-HPLC system (UltiMate™ U3000 RSLCnano, Thermo Fisher Scientific, Germering, Germany) with an autosampler set to 4.0 °C and column oven to 50 °C. proBDNF-derived peptides were separated on an Acclaim PepMap® RSLC column (15 cm × 300 µm i.d., 2.0 µm particle size, 100 Å pore size, Thermo Scientific™). One microlitre of the sample was injected in full-loop mode. proBDNF-peptides were separated at 1.2 µL min 1. For peptide analyses, the linear gradient was as follows: 1.0% B for 5.0 min, 1.0–30% B for 30.0 min, 30.0–60.0% B for 5.0 min, 99.0% B for 5.0 min, 1.0% B for 10.0 min.
Intact proBDNF, as well as tryptic peptides thereof, were analyzed on a benchtop Q Exactive™ Plus Hybrid Quadrupole-Orbitrap™ Mass Spectrometer (Thermo Fisher Scientific, Bremen, Germany). Intact proteins were sprayed from a HESI source in positive ion mode. Ion source settings for proBDNF were as follows: source heater temperature 150 °C, spray voltage 4.5 kV, sheath, auxiliary, and spare gas flow 10, 8, and 0 arbitrary units, respectively. proBDNF was analyzed in full MS mode throughout the chromatographic run (0.0–50.0 min) over an
m/
z range of 1000–3500 at a resolution of 140,000 at
m/
z 200 using the following instrument settings: AGC target 3 × 10
6, maximum injection time of 150 ms, 10 microscans, capillary temperature 300 °C, and S-lens RF level 50. Tryptic peptides of proBDNF were ionized on a nano-ESI source under positive polarity at 1.5 kV. Peptides were acquired in full MS mode throughout the chromatographic run (0.0–55.0 min) within a scan range of
m/
z 400–3000 at a resolution of 70,000 at
m/
z 200 with an AGC of 3 × 10
6, maximum injection time of 100 ms. The capillary temperature was set to 250 °C, S-lens RF level to 60. For fragmentation, a top 10 method was utilized within a scan range of 200–2000
m/
z at a resolution of 17,500 at 200
m/
z with an AGC of 1 × 10
5 and a maximum injection time of 50 ms. The normalized collision energy was set to 28.0% and the dynamic exclusion to 10.0 s. All used HPLC-MS parameters are summarized in
Supplementary Tables S1 and S2.
For disulfide mapping, simulated isotope patterns were generated in Thermo Xcalibur Qual Browser software (v.4.2.28.14) with the following settings: the chemical formula was calculated in excel, 100% hydrogen adduct, most abundant charge state 2–4, output style profile (Gaussian, samples per peak 70), resolution 70,000, valley FWHM (full-width half maximum). Extracted ion current chromatograms were calculated by Thermo Xcalibur Qual Browser software using the following settings: enabled smoothing using Boxcar for 7 points, baseline subtraction enabled, mass tolerance 10 ppm. Peptide data was evaluated in Byonics (Protein Metrics Inc., Cupertino CA, USA). For proBDNF-peptides, lysine and arginine were defined as cleavage sites allowing 2 missed cleavages. Parameters for instrument settings were precursor mass tolerance of 20 ppm, fragmentation type CID low energy, and a fragment mass tolerance of 20 ppm. The protein false discovery rate (FDR) was set to 1%. Spectrum output options: charge state 1–6, maximum precursor mass 10,000 Da; protein output options: 2% FDR. Disulfide mapping was activated (−2.015650 Da at cysteine) was activated for proBDNF. The mass spectra of intact proBDNF were deconvoluted using BioPharma Finder 3.0 (Thermo Scientific) using the Xtract
® algorithm for isotopically resolved spectra. Detailed settings for evaluation parameters can be found in
Supplementary Table S3.
4.5. Legumain Fluorogenic Activity Assay Site
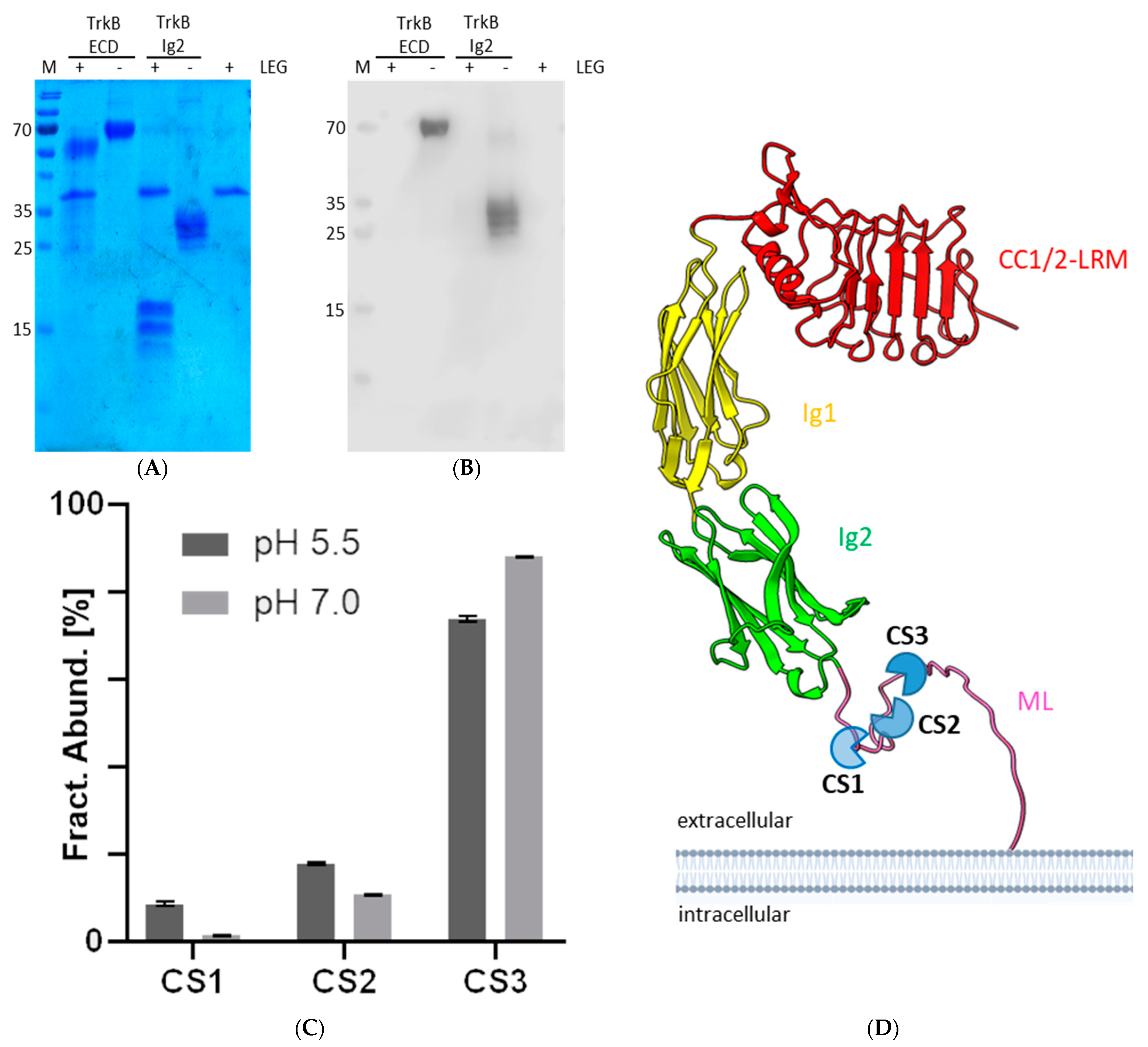
The enzymatic activity of legumain was investigated using the peptidic Z-Ala-Ala-fluorescence was measured at 460 nm upon excitation at 380 nm. Processing Asn-7-amino-4-methylcoumarin (Z-AAN-AMC; Bachem, Bubendorf, Switzerland) substrate. The activity was measured in assay buffers composed of 100 mM citric acid pH 5.5 and 50 mM NaCl or 100 mM Hepes pH 7.0 and 50 mM NaCl supplemented with 50 µM of the substrate. Additionally, the effect of the reducing agent DTT was tested upon the addition of 5 mM DTT to the assay buffers. Reactions were started by the addition of the enzyme at an 8 nM concentration. Assays were carried out at 37 °C in an infinite M200 plate reader (Tecan, Grödig, Austria). An increase in fluorescence was measured at 460 nm upon excitation at 380 nm.
4.6. Cleavage Assay to Determine Cleavage Site
Processing of TrkB ECD and Ig2 with legumain was done by mixing 0.5–1 mg/mL TrkB ECD/Ig2 and 0.007–0.055 mg/mL legumain in cleavage assay buffer (20 mM citric acid, 50 mM NaCl, pH 5.5) which was pH-fixed to pH 7 or 5.5 with either 1 M HEPES, 1 M Tris pH 7 or 1 M citric acid pH 5.5 (1/10th of total volume). The mixture was incubated for 10–15 min at RT. For MS measurements, legumain was inactivated by adding 1 M Ammonium acetate pH 8.6 at 1:1 ratio of total volume. An exemplary assay with respective ratios and volumes can be found in
Supplementary Table S4.
4.7. Confirmation of Structural Integrity of TrkB after Processing by Gel Filtration
The TrkB cleavage assay was done as described above and shown in the
Supplementary Table S4. Only the total volume has been up-scaled to 500 µL.
Processed TrkB ECD was injected into the AKTA FPLC system (GE Healthcare, Vienna, Austria) with a Superdex 200 10/300 GL column (Cytiva Lifesciences) attached, while for TrkB Ig2, a Superdex 75 10/300 GL was used. Both columns were equilibrated with 1x PBS pH 7.4. Fractions with no visible legumain on SDS-PAGE were pooled and concentrated with a 5 kDa cutoff Amicon (Sartorius, Göttingen, Germany).
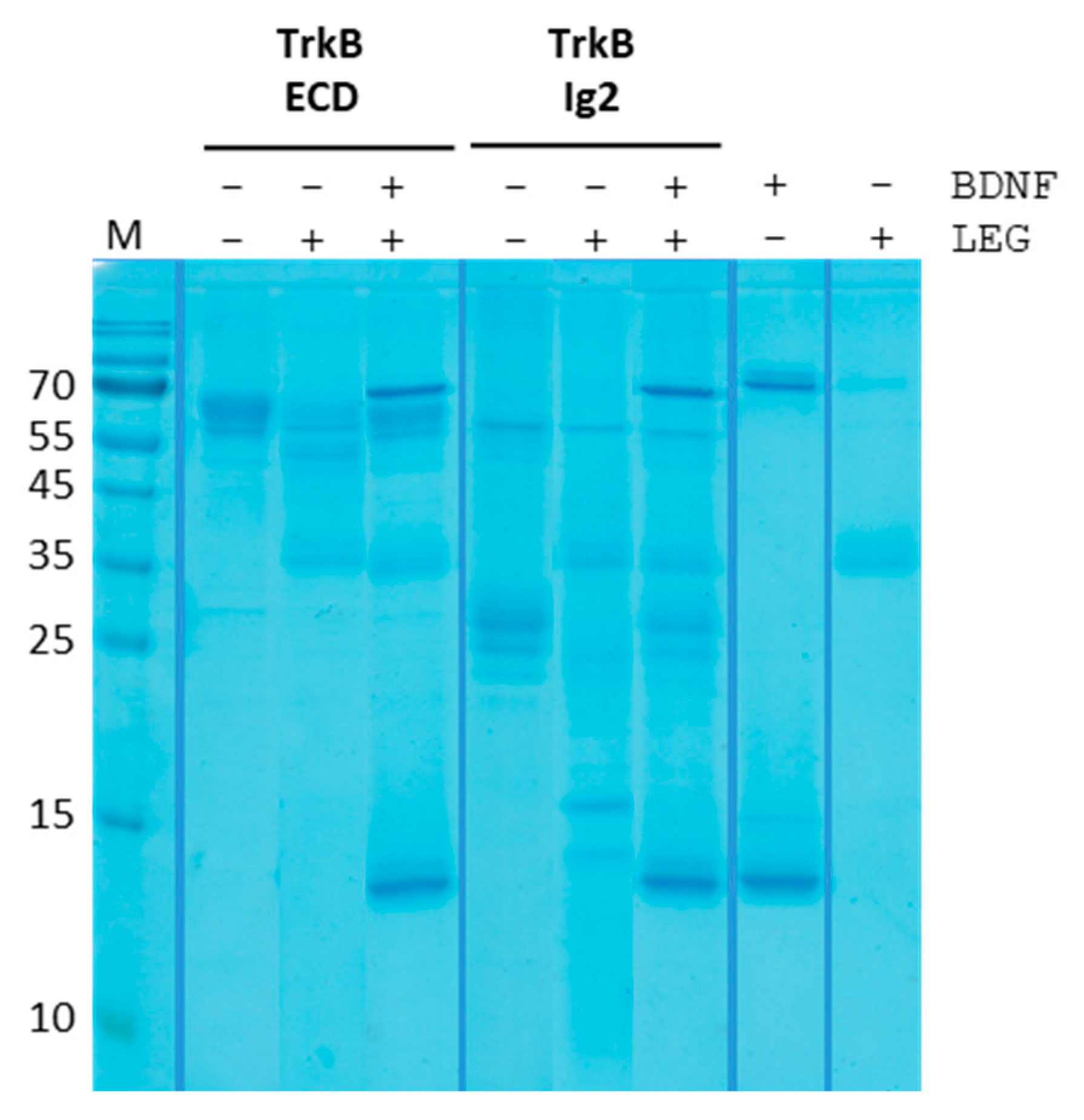
4.8. BDNF-TrkB Complex Cleavage Assay
Preparation. To allow complex formation, 47 µL of 0.3 mg/mL BDNF were mixed with 3 µL of ~4 mg/mL TrkB ECD/Ig2 for a total concentration of ~0.28 mg/mL of both in 1x PBS pH 7.4. The mixture was incubated at RT for 10 min. Legumain was diluted in cleavage assay buffer to a stock concentration of 1.875 mg/mL.
Cleavage assay. The complex cleavage assay was then mixed together similarly to the cleavage assay described before or as described in
Supplementary Table S4 and incubated at RT for 10 min. The stock solution for fixing the pH to 7 was 1 M Tris, and the assay buffer was the previously described cleavage assay buffer at pH 5.5. 10 µL samples were transferred onto a 15% SDS-PAGE.
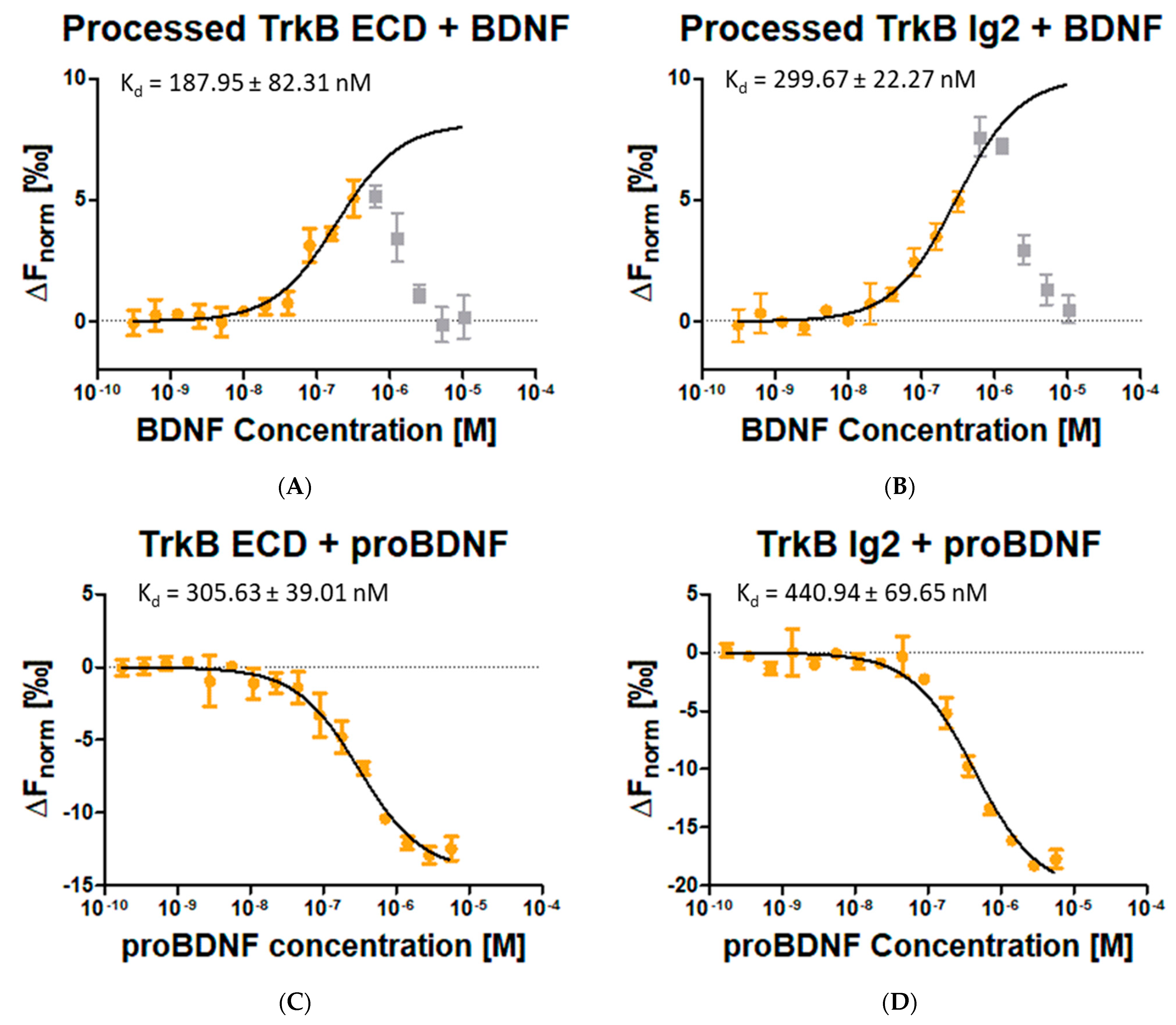
4.9. Determination of Complex Formation Affinity by Microscale Thermophoresis (MST)
Protein labeling. BDNF (20 µM), intact TrkB ECD/Ig2 (20 µM/20 µM), processed TrkB ECD/Ig2 (4.9 µM/15 µM) were labeled with NT-647 RED dye (NanoTemper, Munich, Germany) according to the protocol provided by NanoTemper with the following change: The buffer was not exchanged with the protocols exchange buffer, but all proteins were kept in either 1x PBS pH 7.4 or 20 mM HEPES, 100 mM NaCl, pH 7.5.
Incubation. For the following combinations of labeled and unlabeled proteins, each preparation step was the same: labeled intact TrkB ECD/Ig2 (0.65 µM/0.79 µM) with unlabeled BDNF (20.5 µM), labeled processed TrkB ECD/Ig2 (0.25 µM/0.52 µM) with unlabeled BDNF (20.5 µM) and labeled BDNF (1.1 µM) with unlabeled TrkB ECD/Ig2 (20 µM/20 µM). Absorption at 280 and 650 nm was measured for each labeled protein to determine the concentration and degree-of-labeling (DOL) before being diluted 1:67 or 1:100 in 180 µL ice cold MST buffer (1x PBS, pH 7.4, 0.005% Tween 20). Next, 10 µL of unlabeled protein was diluted 1:2 per step in a 16x dilution series with MST buffer at RT. To each 10 µL of the dilution series, 10 µL of diluted labeled protein was added and mixed by pipetting. The mix was incubated in the dark at RT for 10 min and centrifuged for 5 min at 13,000 rpm at 4 °C (Heraeus biofuge fresco, Hanau, Germany). Thereafter, each of the 16 dilution steps was loaded into a capillary (Monolith NT.115, Nanotemper, Munich, Germany) and placed into the NT.115-Monolith (NanoTemper, Munich, Germany) thermophoresis device.
Measurement and data generation. MST signals were measured with the Monolith NT.115 MST device and recorded with MO.Control Software (v1.6.1, NanoTemper). Excitation Power was set to 40% for Nano-RED, and MST-Power was set to medium. Acquired data were evaluated and affinities were determined with MO.Affinity Software (v2.3, NanoTemper).