1. Introduction
In women, breast cancer is the most prevalent and second-deadliest type of cancer. [
1]. Depending on its oestrogen, progesterone and human epidermal receptor 2 (HER2) receptor status, it can be categorised into luminal-like A, luminal-like B, HER2-enriched and triple-negative breast cancer (TNBC) [
2]. Because TNBC lacks the expression of the receptors commonly targeted in breast cancer therapies, it has limited treatment options [
3]. The aggressive nature of TNBC is likely because of the high level of breast cancer stem cells (BCSCs) in the tumour bulk [
2]. BCSCs are a subset of cancer cells that can self-renew and differentiate into a variety of breast cancer cell lineages, supporting tumour growth [
4]. Our laboratory isolated BCSCs from primary human tumours of TNBC, which were described in recent publications [
5,
6,
7,
8]. BCSCs exhibit stem cell potential in vitro, as demonstrated by sphere formation assays and flow cytometry analysis of the CD49f and CD44 expression, and in vivo, because they form xenografts that are strikingly similar to the tumours that they were originally isolated from, both histologically and in terms of their gene expression profile [
5,
6,
7,
8].
We have previously performed a kinase screening of the bi-potential MDA-MB-468 TNBC cell line to detect kinases that prevent cancer stem cell differentiation (p. 1, [
7]). Four to ten lentiviral short hairpin RNA (shRNA) constructs per target, each of which was tested in three replicates, were used to target 420 kinases; successfully transduced cells were immunocytochemically analysed for myoepithelial keratin 5 (K5) and luminal keratin 8 (K8) expression. This allowed the identification and validation of 11 kinases whose downregulation by at least two differential shRNAs induced a luminal phenotype (K5
−/K8
+). Further research revealed that the downregulation of endoplasmic reticulum to nucleus 1 (ERN1) and alpha kinase 1 in MDA-MB-468 inhibited cellular growth and tumour-forming potential (p. 1, [
7]). In the kinase screening, interleukin-1 receptor-associated kinase 2 (IRAK2) was identified.
IRAK2 is a pseudokinase that participates in the ‘Myddosome’ formation with other components of the IRAK family upon toll-like/interleukin-1 receptor (TLR/IL-1R) activation [
9,
10]. The IRAK2 pathway promotes the phosphorylation of mitogen-activated protein kinases (MAPKs) and nuclear factor-light chain enhancer of activated B cells (NF-κB) [
10]. A subfamily of MAPK, called extracellular signal-related kinase 1/2 (ERK1/ERK2), induces the expression of genes related to cellular proliferation, such as cyclin D1 [
11]. Through its interaction with ERN1, IRAK2 supports the unfolded protein response (UPR), a pathway that encourages cell survival and stress adaptation [
12,
13]. When ER-stress is present, ERN1 excises the mRNA of the protein known as IRE1-X-box-binding protein 1 (XBP1), which is subsequently translated into a transcription factor that promotes the expression of chaperones and protein degradation factors, thereby enhancing the endoplasmic reticulum’s (ER) capacity for folding proteins [
14]. Apoptosis, UPR and autophagy are all tightly regulated pathways in which cells integrate signals and activate UPR and autophagy to block apoptosis if survival is favourable; if survival is unfavourable, apoptosis is no longer inhibited [
15].
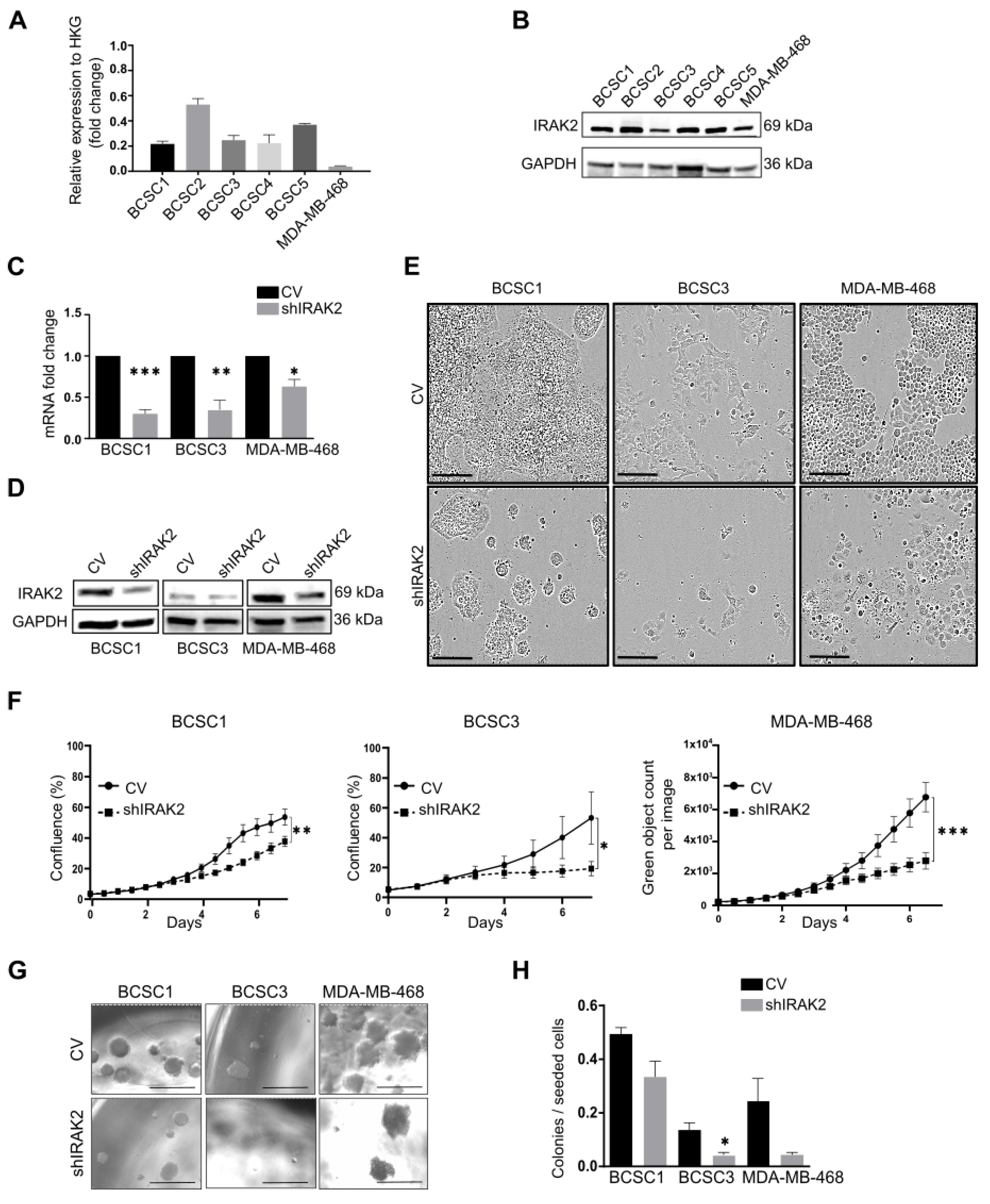
Because we previously identified IRAK2 in the kinase screening, we wondered if its downregulation could affect the growth of our primary BCSCs and the commercial TNBC cell line MDA-MB-468. We evaluated the impact of IRAK2 downregulation on cell phenotype, proliferation, sphere-forming ability and tumour development. Additionally, the impacted molecular pathways in the context of apoptosis, autophagy and UPR were investigated.
3. Discussion
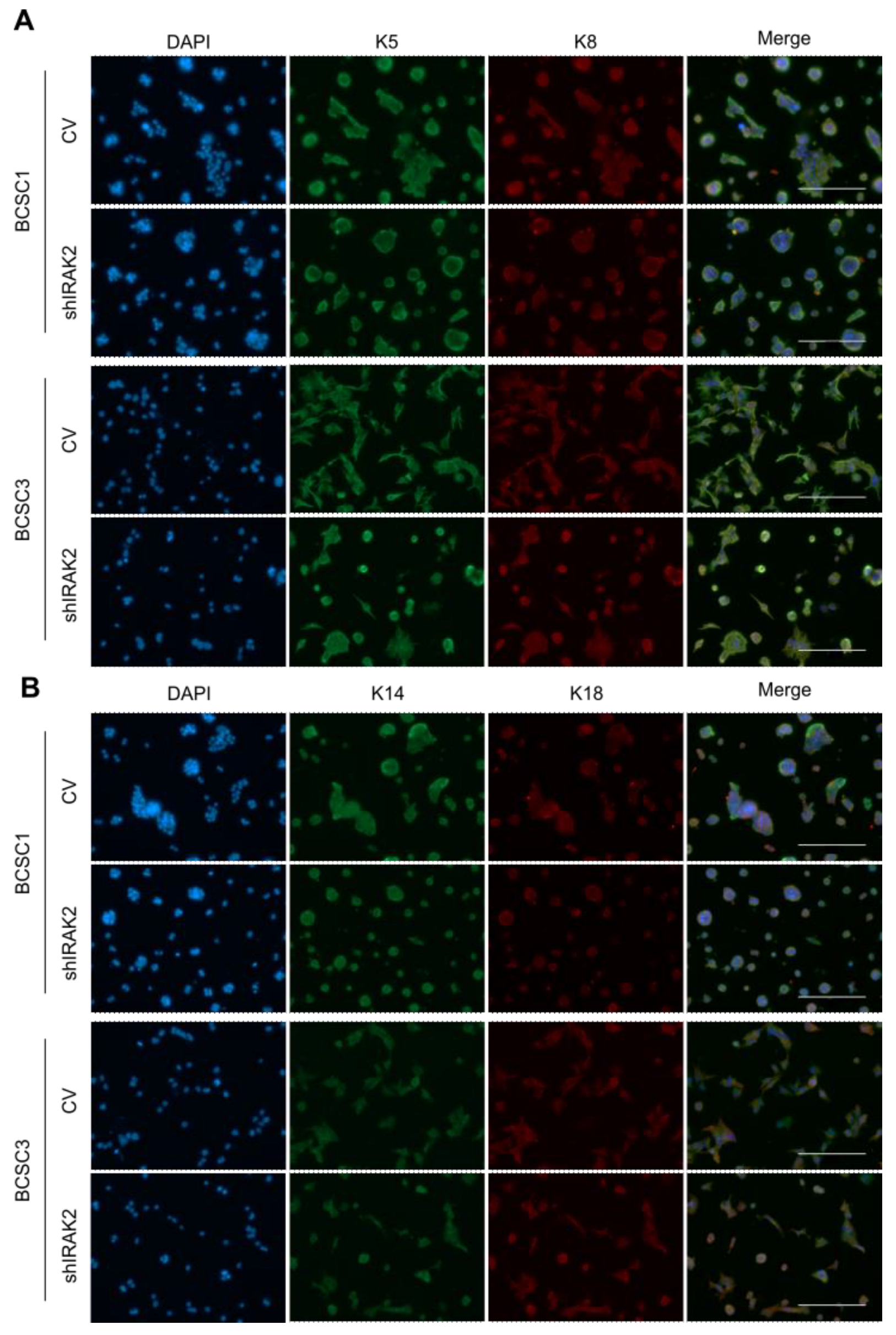
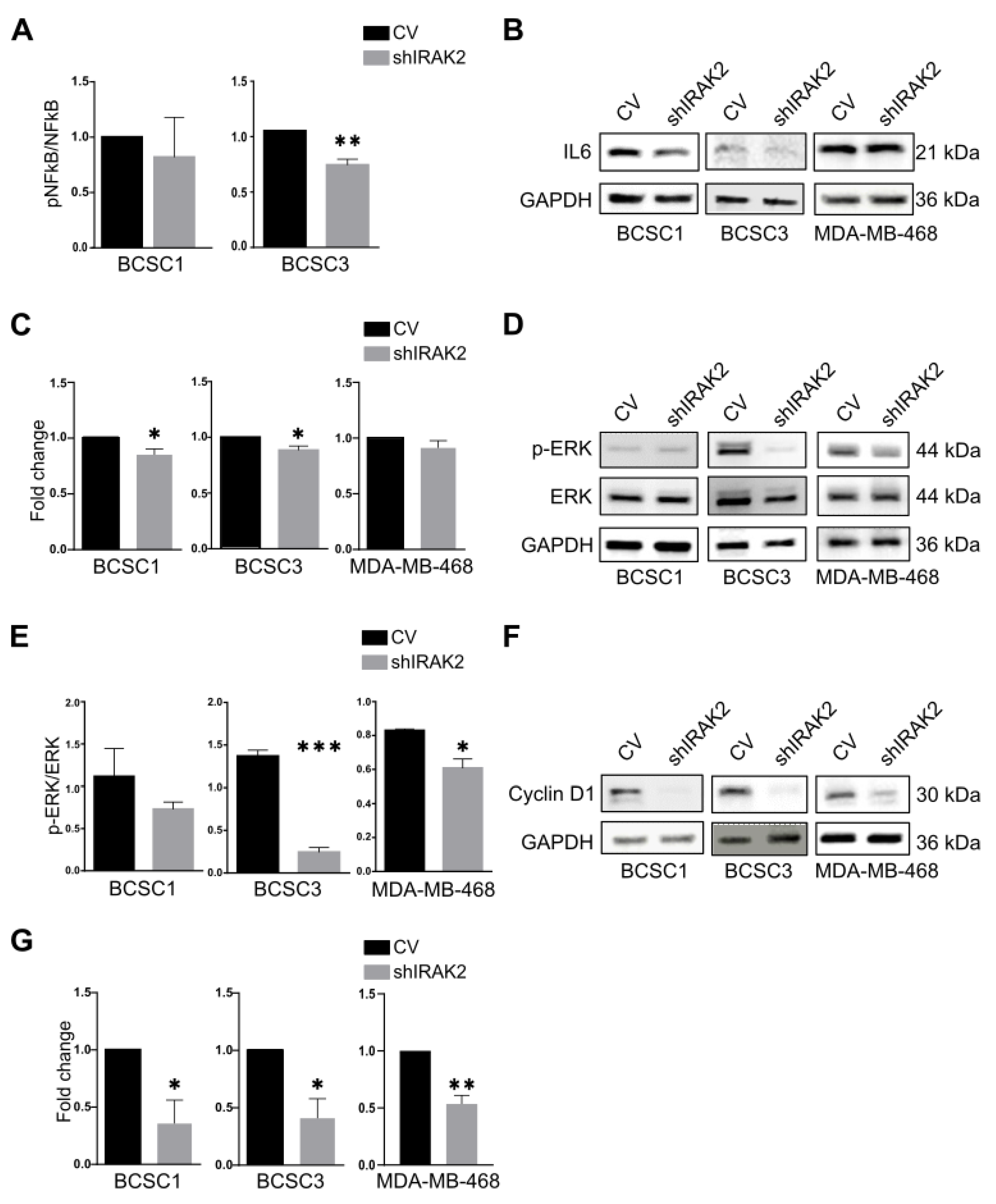
We reported that IRAK2 contributes to BCSCs and MDA-MB-468 self-renewal and likely has a pro-oncogenic role. The bi-potential phenotype of BCSCs did, however, continue after IRAK2 was downregulated because the expression of the keratins was unaffected, suggesting that there was no overt differentiation. When we examined the molecular mechanisms underlying the outcomes we observed, we discovered that the presence of IRAK2 knockdown decreased NF-κB and ERK phosphorylation as well as IL-6 and cyclin D1 expression. Considering that NF-κB, ERK and cyclin D1 pathways favour cellular growth [
19,
22,
23], their impairment upon IRAK2 knockdown could explain the decreased proliferation we had observed. Moreover, because cyclin D1 expression was impaired in shIRAK2 BCSC1 xenografts, it may be responsible for delayed tumour growth.
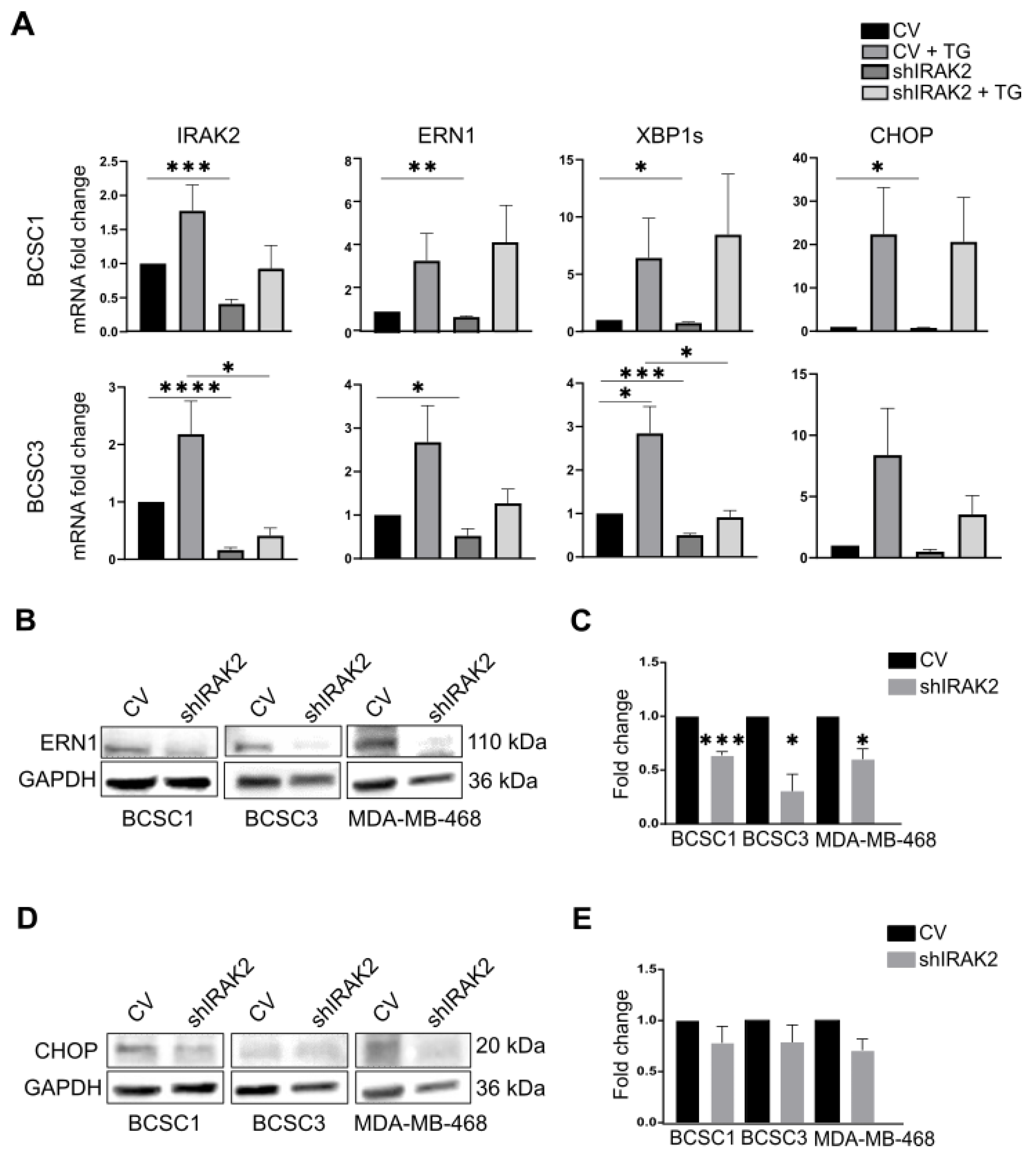
We reported that IRAK2 transcription is upregulated in BCSCs when an ER-stress inducer stimulates them, revealing an IRAK2 requirement in the presence of ER-stress; interestingly, IRAK2 upregulation in the presence of ER-stress agents was previously reported in a primary prostatic carcinoma cell line [
12]. IRAK2 knockdown per se affected ERN1 expression and XBP1 splicing, suggesting an interaction between IRAK2 and ERN1 signalling. These data opened up a new perspective related to the role of ERN1 in MDA-MB-468, because the cellular growth inhibition that we reported upon ERN1 depletion (p. 1, [
7]), as well as being caused by cellular differentiation, could be addressed in terms of ERN1 implications in the UPR.
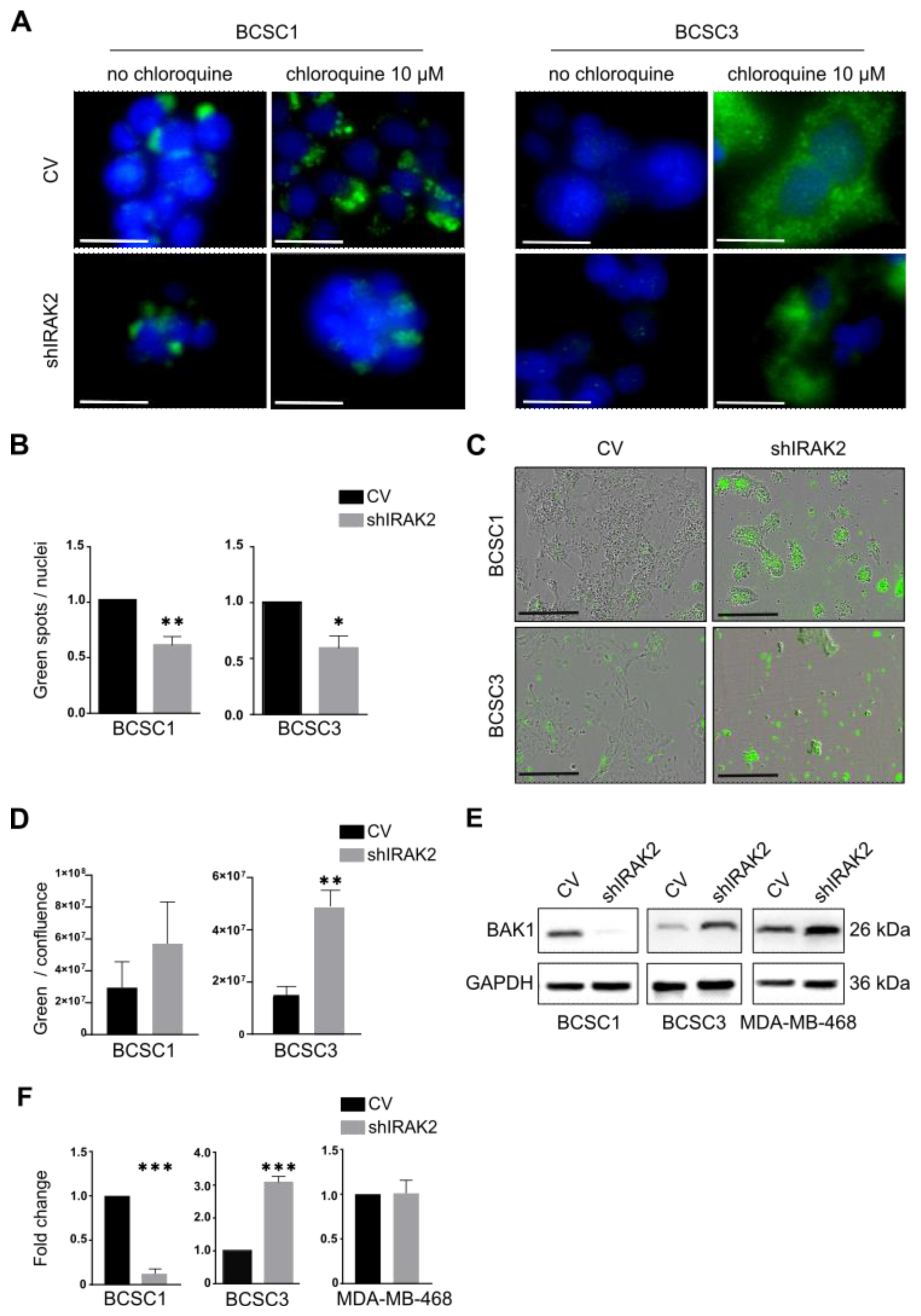
BCSC1 and BCSC3 presenting IRAK2 knockdown displayed a significantly decreased autophagy, and considering that ERN1 induces autophagy through the TRAF2/JNK/c-Jun pathway or XBP1 splicing [
24], the decreased autophagy could be due to IRAK2-mediated ERN1 downregulation. Moreover, BCSC1 and BCSC3 displayed increased apoptosis upon IRAK2 knockdown, as represented by increased PS externalisation. The downregulation of IL-6 and NF-κB mediated by IRAK2 knockdown may be responsible for the increased PS externalisation, because IL-6, as well as being a pro-inflammatory cytokine, is also an anti-apoptotic factor [
16], and NF-κB activation prevents apoptosis in several cell types [
23]. Therefore, BCSC3 characterised by IRAK2 knockdown may present increased apoptosis due to IL-6 and NF-κB downregulation by upregulating BAK1, whereas BCSC1 characterised by IRAK2 knockdown may activate apoptosis through different pathways that do not include BAK1 upregulation. These results are consistent with those of previous studies showing that the inhibition of other players in the IRAK2 pathway decreases NF-κB activation, thereby inducing apoptosis [
25]. Moreover, IRAK2 downregulation was previously associated with increased apoptosis in canine breast cancer [
26]. These findings confirmed that IRAK2 may help cells handle stress-related conditions by stimulating the UPR and autophagy while preventing apoptosis. Therefore, IRAK2 knockdown may impair both the ability of cells to proliferate and self-renew and their ability to resist hostile environments.
The cell lines considered were slightly differently affected by IRAK2 downregulation at the molecular level. Different factors may contribute to the cell line-specific molecular responses detected upon IRAK2 downregulation. MDA-MB-468 is a commercial cell line, whereas BCSCs were recently isolated from primary TNBC tumours [
5,
6,
8]. They present different mutations and genetic backgrounds, and they belong to different TNBC subtypes, given that MDA-MB 468 and BCSC3 are basal-like 1, whereas BCSC1 is basal-like 2. Furthermore, as depicted by our quantitative real-time polymerase chain reaction (qPCR) and Western blot results, the cells present different IRAK2 expression levels and degrees of its knockdown, representative of different residual IRAK2 activities that may differently influence their phenotype.
Cancer cell growth, migration, tumourigenesis and chemoresistance can all be impacted by targeting the proteins involved in the IRAK2 pathway, including IRAK1 and IRAK4 [
27,
28,
29,
30]. Targeting UPR-related proteins has also been demonstrated to be a promising anticancer method [
12,
31,
32]. Because we reported that IRAK2 downregulation impaired its pathway and we demonstrated IRAK2 participation in the UPR, IRAK2 could be considered an interesting target to affect the aggressive growth of TNBC.
We concluded that although IRAK2 downregulation affected the molecular pathways in the cell lines considered in this study slightly differently, overall, its downregulation was advantageous because it decreased cellular growth in vitro and delayed tumour progression in vivo. Moreover, IRAK2 downregulation compromised ERN1 signalling and autophagy, which are pathways that are frequently exploited by cancer cells to survive, further supporting the beneficial impact of IRAK2 downregulation in TNBC.
4. Materials and Methods
4.1. Cell Culture
BCSCs, as previously described [
5,
6,
7,
8], were isolated in 2014 from TNBC specimens by the mechanical and enzymatic disruption of the tissue, obtaining single cells that are seeded in 50% matrigel (Corning, 354230, Bedford, MA, USA) at 37 °C and under low-oxygen conditions (3% oxygen, 5% carbon dioxide, and 92% dinitrogen), and subsequently expanded in 2D (2% matrigel) [
5,
6,
8]. BCSCs were authenticated by performing a Multiplex human Cell line Authentication Test. MDA-MB-468 cells (ATCC, RRID:CVCL_0419, Manassas, VA, USA) presented a green fluorescent protein nuclear tag and were cultured in Dulbecco’s modified Eagle’s medium (DMEM, Gibco, 41966-029, Grand Island, NY, USA), 10% foetal bovine serum (without tetracycline to avoid uncontrolled gene downregulation, Gibco, 15140-122, Grand Island, NY, USA) and 1% penicillin/streptomycin (Gibco, 15140-122, Grand Island, NY, USA). All experiments were performed using mycoplasma-free cells. All cell lines were authenticated using single nucleotide polymorphisms profiling within the last 3 years.
4.2. Lentiviral Production and Knockdown
One thousand nanograms of the vector of interest were incubated for 30 min in six-well plates (Falcon, Corning, 353046, Durham, NC, USA) with 400 µL of Opti-MEM (Life Technologies, 11058021, Carlsbad, CA), 4 µL of X-tremeGENE™ transfection reagent (Sigma-Aldrich, 6366236001, Taufkirchen, Germany), 700 ng of pCMVdR8.74 (Addgene plasmid, 22036) and 350 ng of pMDVSVG (Addgene plasmid, 8454). Two million 293FT cells (Invitrogen, RRID:CVCL_6911, Thermo Fisher, Waltham, MA, USA) were added to each well, and the medium was replaced with fresh UltraCULTURE™ (Lonza, 12-725F, Basel, Switzerland) 24 h later. The lentivirus-containing medium was harvested and centrifuged at 500×
g for 10 min 48 and 72 h later. The supernatant was filtered with 0.45-µm filters (Cytiva, WH10462100, Marlborough, MA, USA), overlayed at a 4:1 ratio with 10% sucrose-containing buffer [
33] and centrifuged at 10,000×
g for 4 h at 4 °C. Pellets were resuspended in UltraCULTURE™. The lentivirus was serially diluted 1:2 in DMEM containing 15-μg/mL polybrene (Sigma-Aldrich, 107689, Taufkirchen, Germany) and titrated on 3 × 10
3 293FT cells/well of a 96-well culture plate, which was centrifuged at 1000×
g for 1 h. The selection was performed using 2-μg/mL puromycin (Sigma-Aldrich, P8833-10MG, Taufkirchen, Germany). The highest viral dilution infecting cells was chosen as the multiplicity of infection (MOI) equal to one. Different MOIs were tested on each cell line before proceeding with their infection. Cells were infected with lentivirus at the optimal MOI in a medium containing 15-μg/mL polybrene, and plates were centrifuged at 1000×
g for 1 h. pLKO-Tet-On vectors were used to perform inducible knockdown [
34,
35], and the knockdown was induced by supplementing cellular media with 100-ng/mL doxycycline (Sigma-Aldrich, D9891, Taufkirchen, Germany), which was refreshed every 48 h.
4.3. Proliferation Assay
Control and shIRAK2 cells were seeded at 3 × 103 cells/well in 96-well culture plates (Falcon, Corning, 353072, Durham, NC, USA), and their growth was observed using the IncuCyte® Live-Cell analysis system (Sartorius, Ann Arbor, MI, USA). After 1-week, growth curves were built considering the ‘cells percentage of confluency’ for BCSCs or ‘green object count per image’ for MDA-MB-468.
4.4. Sphere-Forming Capacity
Control and shIRAK2 cells were seeded at 2 × 102 cells/well in 96-well low-attachment flat-bottom plates (Corning, CLS7007-25EA, Durham, NC, USA) in a mixture of 10-µL medium and 10-µL matrigel. Thirty minutes after seeding, 80 µL of the medium was added to each well, and the cells were left to grow for 2 weeks. The doxycycline-containing medium was added to the wells every 2 days. To maintain a constant 100-ng/mL doxycycline concentration, exactly 5 µL of the medium was added to each well every time; however, the amount of doxycycline added was calculated considering the total volume of the medium contained in the well. Sphere-forming capacity was quantified as the ratio between the spheres counted after 2 weeks and the number of cells seeded per well.
4.5. Immunofluorescence Staining
Control and shIRAK2 BCSC1 and BCSC3 were seeded at 1 × 104 cells/well in 96-well culture plates. After 5 days, the cells were washed twice with 150-μL/well phosphate buffered saline (PBS), fixed with 100 μL of ice-cold methanol at 4 °C for 15 min, rinsed twice with 150-μL/well PBS (Gibco, 70011044, Grand Island, NY, USA), permeabilised with 150 μL/well of 1× tris buffered saline with tween (TBST) and blocked with 100 μL/well of 1-mg/mL albumin/PBS (blocking solution). The cells were incubated overnight at 4 °C with 40 μL/well of primary antibody diluted in blocking solution; the following day, the cells were rinsed and incubated for 1 h with 50 μL/well of secondary antibody diluted in blocking solution. The secondary antibody was washed out, and nuclei were stained with 4′,6-diamidino-2-phenylindole (DAPI, Sigma-Aldrich, D9542, Taufkirchen, Germany). The following were the primary antibodies used: anti-keratin 5 (Covance PRB, 160P, Princeton, NJ, USA, 1:250), anti-keratin 8 (BioLegend, C5301, San Diego, CA, USA, 1:250), anti-keratin 14 (Covance, PRB-155P, Princeton, NJ, USA, 1:250), anti-keratin 18 (Dako Agilent, M7010, Santa Clara, CA, USA, 1:250) and anti-LC3-II (Cell Signalling, 38689, Danvers, MA, USA, 1:1600). The following were the secondary antibodies used: Alexa Fluor 488 donkey anti-rabbit (Thermo Fisher, A21206, Waltham, MA, USA, 1:500) and Alexa Fluor 568 donkey anti-mouse (Invitrogen, A10037, Thermo Fisher, Waltham, MA, USA, 1:500).
4.6. Immunohistochemistry
Pieces of xenografts were fixed in paraformaldehyde (Carl Roth, 0335.1, Karlsruhe, Germany) 4% for 24 h at 4 °C, embedded in paraffin, sliced into 2-μm thick sections and mounted on glass slides. The slides were immersed in xylene for 5 min, then in a descending concentration of ethanol for a few seconds and rinsed with distilled water. The slides were immersed in citrate buffer at a pH of 6 (Dako Agilent, S236984, Santa Clara, CA, USA) for 1 h at 96 °C, allowed to chill for 30 min, washed with distilled water for 5 min and immersed in 5% hydrogen peroxide (Sigma-Aldrich, SA31642, Taufkirchen, Germany)/methanol for 15 min. The slides were incubated overnight at 4 °C with 200 µL of antibody diluent (Dako Agilent, S3022, Santa Clara, CA, USA) containing the primary antibody; the following day, they were washed with Tris-buffered saline 1× for 5 min and incubated for 40 min with 200 µL of the antibody diluent containing the secondary antibody. Next, 200 µL of 3,3′-diaminobenzidine (DAB, Dako Agilent, K3468, Santa Clara, CA, USA) solution was incubated on each slide for 10 min and subsequently washed for 5 min in mineral water and 5 min in distilled water. The slides were subsequently stained by immersing them in hematoxylin (Dako Agilent, C5700, Santa Clara, CA, USA) for 5 min, washing them in mineral water for 5 min and washing them in distilled water for 5 min. The slides were immersed in ascending concentrations of ethanol, bathed in xylene for 5 min and covered with the mounting medium and a coverslip. DAB staining was quantified using the software ImageJ (1.53n,
imagej.nih.gov/ij, accessed on 11 January 2023), selecting HDAB in ‘Color Deconvolution’ option. The OD was evaluated using the following equation:
with max intensity = 255 and mean intensity = mean grey value calculated by ImageJ. The primary antibody anti-cyclin D1 (Invitrogen, PA532373, Thermo Fisher, Waltham, MA, USA, 1:100) was used. The secondary antibody polyclonal goat anti-rabbit (Dako Agilent, P044801, Santa Clara, CA, USA, 1:100) was used.
4.7. RNA Isolation and qPCR
RNA was extracted from cell pellets using the miRNeasy
® mini kit (Qiagen, 1038703, Hilden, Germany), and reverse transcription was performed using the EvoScript reverse transcriptase kit (Roche, 07912323001, Mannheim, Germany), following the manufacturer’s instructions. For qPCR, a mixture containing 6.25 μL of TaqMan universal master mix II (Thermo Fisher, 4444965, Waltham, MA, USA), 0.5 μL of forward primer, 0.5 μL of reverse primer (
Table 1) and 0.125 μL of the corresponding universal probe library (UPL) probe (Roche, 4683633001, Mannheim, Germany) (
Table 1) for each reaction was prepared and dispatched in a 384-well plate. Then, 12.5 ng of cDNA in 2.5 μL was added to each well. The samples were run in Roche’s LightCycler 480, with an activation step of 10 min at 95 °C, an amplification step repeated 50 times with each step consisting of 15 s at 95 °C and 1 min at 60 °C and a last cooling step at 40 °C. The primers and UPL probes used are indicated in
Table 1. qPCR data were analysed considering the cycle threshold values obtained from the run, and the relative expression was assessed using the 2
−ΔΔCT method [
36].
4.8. Protein Isolation and Western Blot
The lysis solution was prepared following the cOmplete™ lysis-M solution (Roche, 11697498001, Mannheim, Germany) manufacturer’s instructions and supplemented with PhosSTOP (Sigma-Aldrich, 93106075, Taufkirchen, Germany). The cell pellet was resuspended in 50 µL of the lysis solution, incubated on ice for 30 min and centrifuged at 4 °C for 15 min at 13,000 rpm; the supernatants were stored at −80 °C. Protein quantification was performed using the DC™ Protein assay kit II (Bio-Rad Laboratories, 5000112, Hercules, CA, USA). Then, 20 µg of proteins were loaded in the Mini-PROTEAN TGX precast gels (Bio-Rad Laboratories, 456-9036, Hercules, CA, USA), transferred on the Trans-Blot® transfer system membranes (Bio-Rad laboratories, 1704156, Hercules, CA, USA) that were inserted in 15-mL conical tubes containing 5 mL of 1× TBST and rolled on a roller mixer for 5 min. The membranes were blocked for 1 h with 5 mL of blocking solution (2.5 g of albumin, 50-mL 1× TBST) and subsequently incubated overnight at 4 °C with the blocking solution containing the primary antibody. The following day, the membranes were washed out with 1× TBST for 5 min and incubated for 1 h with the blocking solution containing the secondary antibody. The membranes were washed with 1× TBST three times for 5 min and developed using the WEST-ZOL® plus detection system (iNtRON Biotechnology, 16024, Seongnam, Kyonggi-do, South Korea). The membranes were occasionally stripped, incubating them for 45 min in stripping buffer (20-mL SDS 10%, 12.5-mL Tris HCl, 67.5-mL water, 0.8-mL ß-mercaptoethanol and a pH of 6.8). The following primary antibodies were used: anti-IRAK2 (Cell Signalling, 4367S, Danvers, MA, USA, 1:1000), anti-IL6 (Biozol, LS-C 165212, Eching, Bayern, Germany, 1:1000), anti-cyclin D1 (Invitrogen, PA532373, Thermo Fisher, Waltham, MA, USA, 1:1000), anti-ERN1 (Proteintech, 27528-1, Rosemont, IL, USA, 1:1000), anti-CHOP (Sigma-Aldrich, SAB5700602, Taufkirchen, Germany, 1:1000), anti-BAK1 (Thermo Fisher, MA5-32111, Waltham, MA, USA, 1:1000), anti-ERK (Cell Signalling, 4696S, Danvers, MA, USA, 1:1000) and anti-P-ERK (Cell Signalling, 9101S, Danvers, MA, USA, 1:1000). The following secondary antibodies were used: polyclonal goat anti-rabbit (Dako Agilent, P044801, Santa Clara, CA, USA, 1:2000) and polyclonal rabbit anti-mouse (Dako Agilent, P0260, Santa Clara, CA, USA, 1:2000).
4.9. ER-Stress Induction
Control and shIRAK2 BCSC1 and BCSC3 were seeded at 1 × 105 cells/well in a 6-well plate, and the medium was supplemented with 100-ng/mL doxycycline for 3 days. The cells were subsequently treated for 3 h with 5-µM thapsigargin in 2 mL of medium (Sigma-Aldrich, T9033, Taufkirchen, Germany). The cells were detached and centrifuged, and RNA was subsequently isolated from them.
4.10. Autophagy Evaluation
IRAK2 knockdown was induced with 100-ng/mL doxycycline in the control and shIRAK2 BCSC1 and BCSC3 for 4 days. Next, 1 × 104 cells/well were seeded in a 96-well culture plate, treated overnight with 10-μM chloroquine (Sigma-Aldrich, C6628-25G, Taufkirchen, Germany) and subsequently fixed. LC3-II immunofluorescence staining was quantified using the ImageJ plugin automatic nuclei counter ITCN. Channel images were split, green images (LC3-II staining) were inverted and, using the plugin ITCN, the width (5 for BCSC1 and 6 for BCSC3), minimum distance (2.5 for BCSC1 and 3 for BCSC3) and threshold (1.5) to consider were defined. ‘Detect dark peaks’ was selected, and spots (representing LC3-II staining) were counted. Subsequently, nuclei in the blue images (DAPI staining) were manually counted. Autophagy was quantified by evaluating the ratio between the counted dark peaks over the number of nuclei per picture.
4.11. Apoptosis Assay
Control and shIRAK2 BCSC1 and BCSC3 were seeded at 1 × 104 cells/well in 96-well plates in the presence of 100-ng/mL doxycycline for 5 days. Next, apoptosis was assessed using the apoptosis assay kit (Abcam, ab176749, Cambridge, UK) following the manufacturer’s instructions. Apoptosis was quantified considering the green fluorescence measured at Ex/Em = 490/525 nm over cellular confluence, both measured using the IncuCyte® Live-Cell analysis system.
4.12. NF-κB Phosphorylation Assay
Control and shIRAK2 BCSC1 and BCSC3 were treated with 100-ng/mL doxycycline for 4 days and seeded at 3 × 104 cells/well in 96-well culture plates. The EnzyFluo phosphorylation assay kit (BioAssay Systems, ABIN5691837, Hayward, CA, USA) was used to quantify the phosphorylated and total NF-κB, following the manufacturer’s instructions. NF-κB activation was defined by normalising the signal of the measured phosphorylated NF-κB to the NF-κB total protein content.
4.13. Orthotopic Breast Cancer Xenografts
Here, 1 × 105 BCSC1 or 5 × 105 MDA-MB-468 were mixed with one million irradiated Hs27 fibroblasts (ATCC, RRID:CVCL_0335, Manassas, VA, USA). Matrigel was added at a 1:1 ratio to the mixture for a total volume of 40 μL, and the mixtures were transplanted into the mammary fat pads of non-obese diabetic/severe combined immunodeficiency female mice. Two injections were performed for each mouse, one on each side. When the xenografts reached 3 mm in diameter, IRAK2 knockdown was induced by feeding the animals three times a week with gelatin containing 2-mg/mL doxycycline and 0.08-g/mL sugar. The animals were sacrificed when the xenografts reached a diameter close to 15 mm. Three mice received control cells and another three received IRAK2 knockdown cells.
4.14. Statistical Analysis
Data are expressed as the mean of the group plus the SEM, and statistical analysis was performed using the software GraphPad Prism 9 (GraphPad Software Inc., San Diego, CA, USA). Normality was assessed using the Shapiro–Wilk test. In the case of normal distribution, the unpaired Student’s t-test was performed, or else the non-parametric Mann–Whitney U test was used. The paired Student’s t-test was used to compare the growth of control and IRAK2 knockdown xenografts. Proliferation data were analysed using the Wilcoxon test. p-values less than 0.05 were considered as significant (*, p ≤ 0.05; **, p ≤ 0.01; ***, p ≤ 0.001; ****, p ≤ 0.0001).