Identification and Evolutionary Analysis of the Auxin Response Factor (ARF) Family Based on Transcriptome Data from Caucasian Clover and Analysis of Expression Responses to Hormones
Abstract
:1. Introduction
2. Results
2.1. Transcriptome-Wide Identification of ARFs in Caucasian Clover
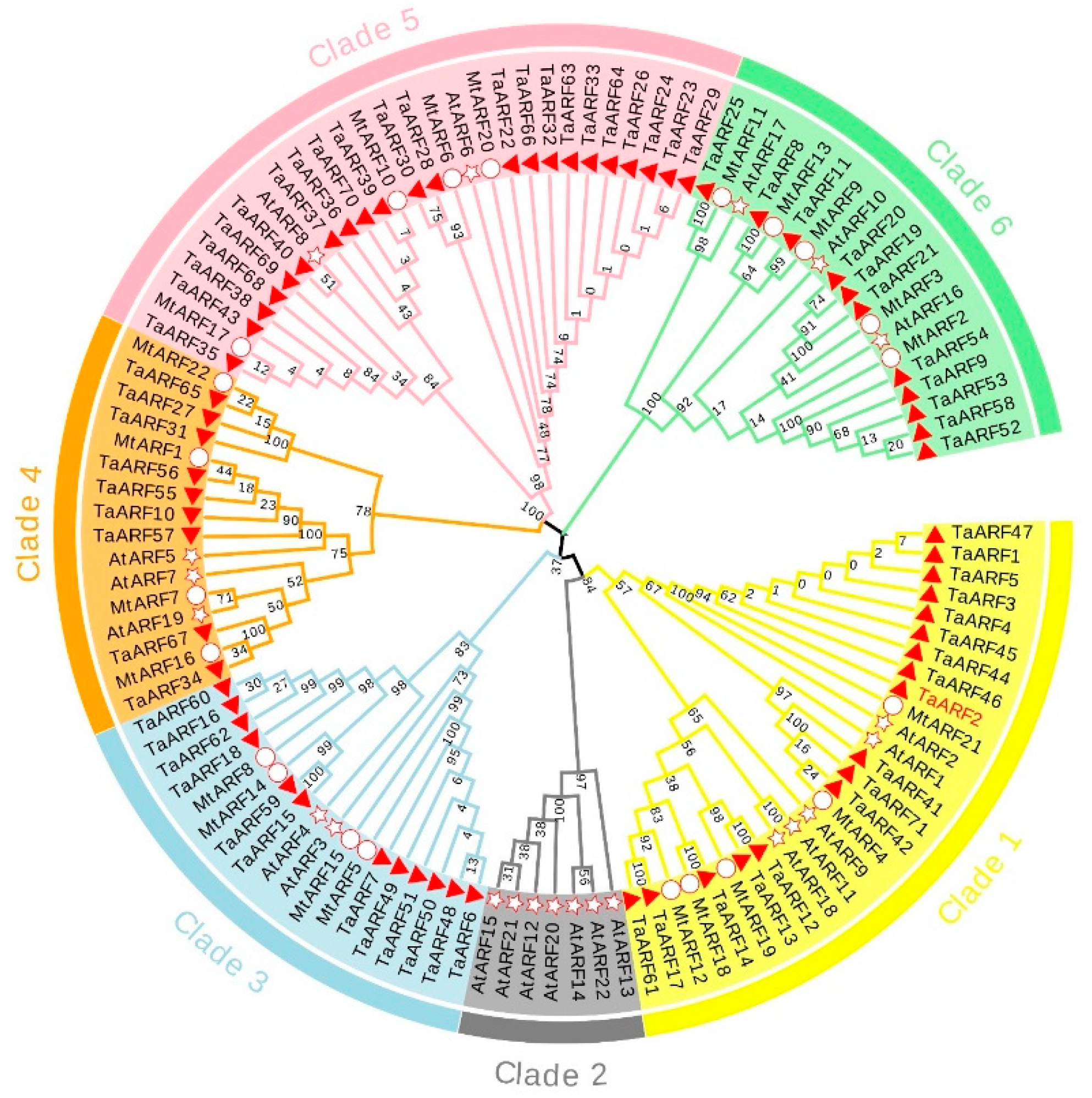
2.2. Phylogenetic Analysis of TaARFs
2.3. Transcriptome-Wide Identification of ARFs in Caucasian Clover
2.4. Expression Patterns of TaARF Genes in Different Tissues of the Caucasian Clover Rhizome
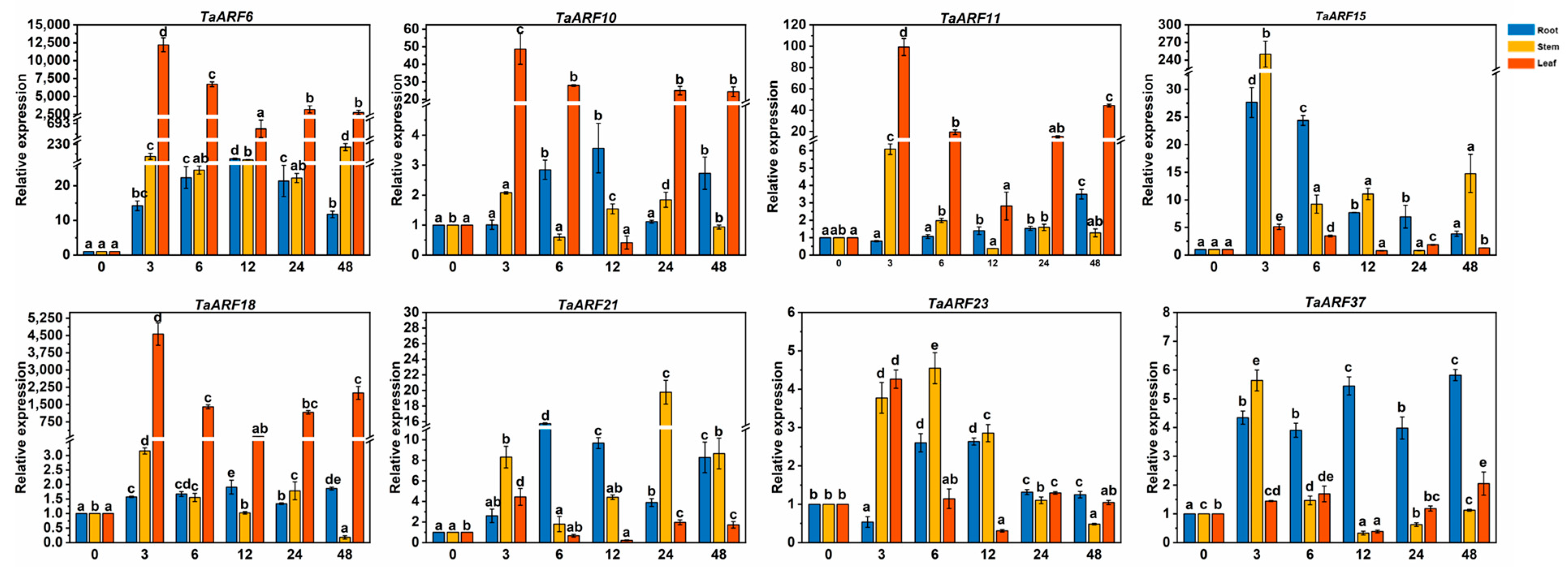
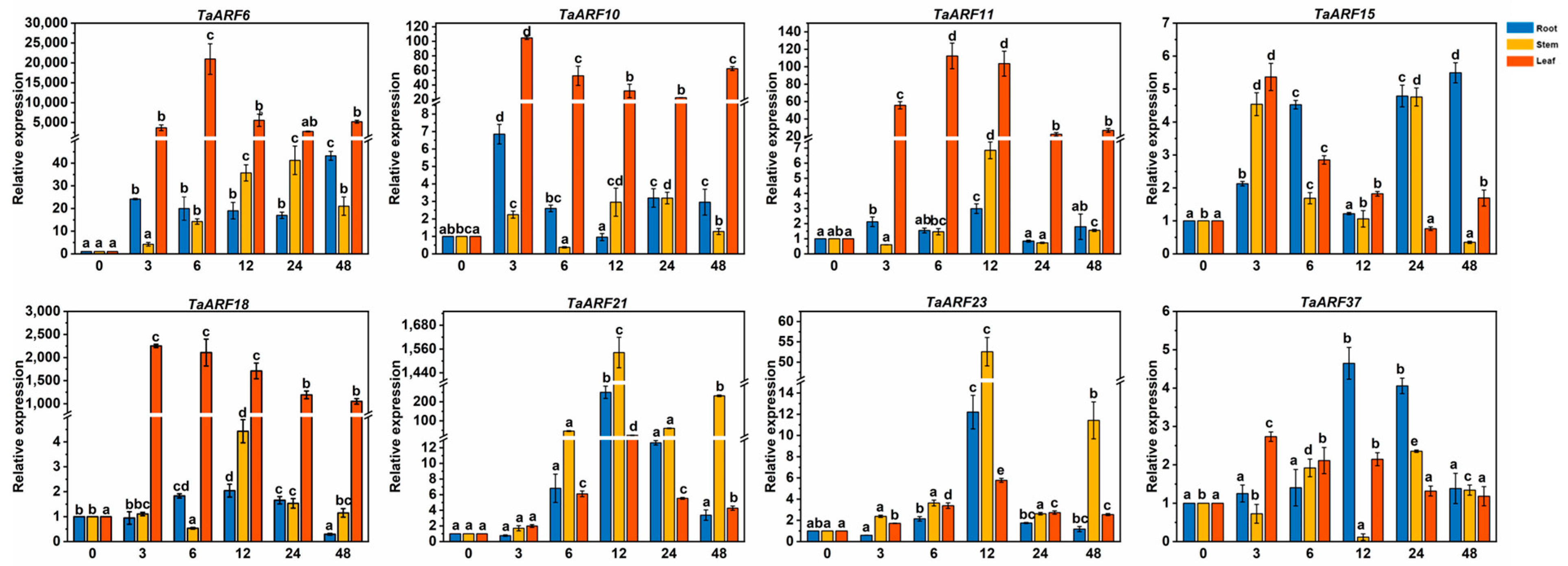
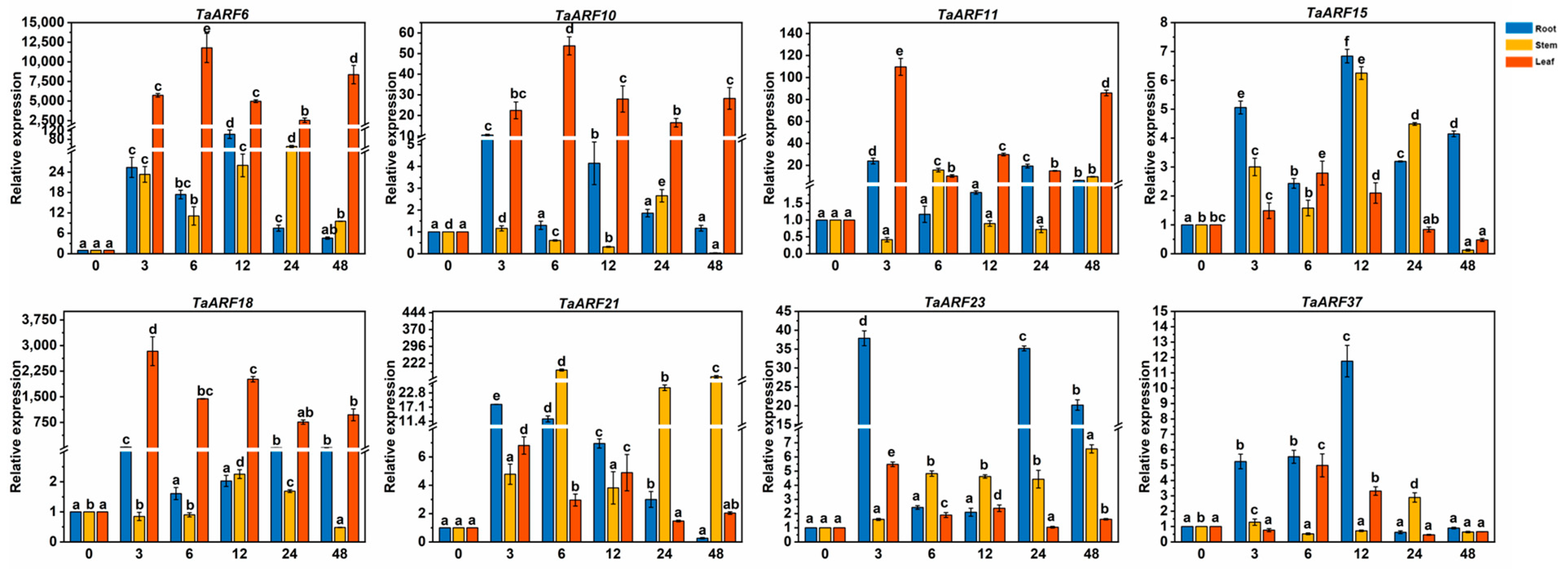
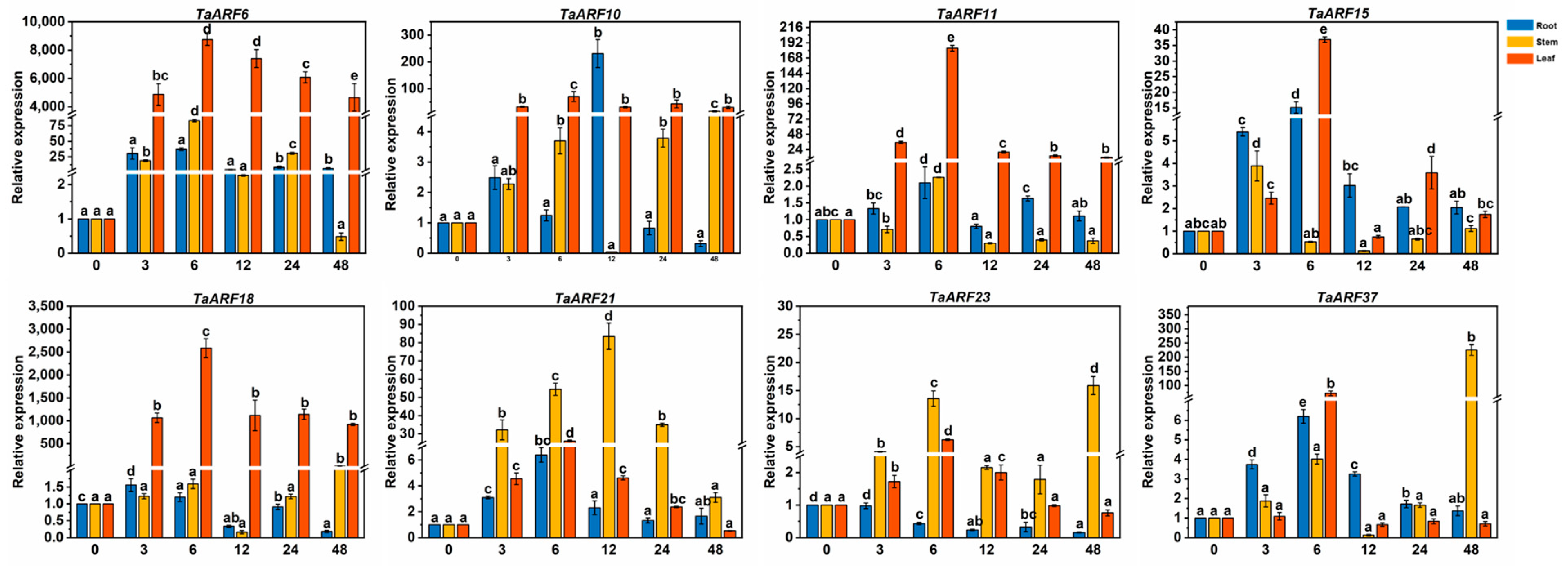
2.5. Expression Analysis of TaARF Genes in Response to Hormones
3. Discussion
4. Materials and Methods
4.1. Identification and Sequence Analysis of ARF Genes from Caucasian Clover
4.2. Physicochemical Properties and Conserved Motif Analysis
4.3. Multiple Sequence Alignment and Phylogenetic Analysis
4.4. Expression Analysis of Different Tissue Sites
4.5. Plant Materials
4.6. RNA Extraction, cDNA Synthesis and Quantitative RT–PCR
4.7. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Kim, B.W.; Nejad, J.G.; Sung, K.I.; Lee, B.H.; Albrecht, K.A. Effects of degrees of grass competition on spreading of Kura clover. Grassl. Sci. 2017, 63, 218–224. [Google Scholar] [CrossRef]
- Teutsch, C.D.; Collins, M.; Ditsch, D.C. Impact of Incorporation, Mulch, and Root Coating on the Establishment of Kura Clover from Rhizome Segments on Mine Spoils in Southeastern Kentucky. Forage Grazinglands 2004, 2, 1–8. [Google Scholar] [CrossRef]
- Baker, J.M. Vegetative propagation of kura clover: A field-scale test. Can. J. Plant Sci. 2012, 92, 1245–1251. [Google Scholar] [CrossRef]
- Riday, H.; Albrecht, K.A. Combining Kura Clover with Forage Legumes and Grasses to Optimize Pasture Forage Legume Content. Agron. J. 2012, 104, 353–362. [Google Scholar] [CrossRef]
- Xiujie, Y. Study on the Adaptation and Utilization of Trifolium ambiguum Bieb. Master’s Thesis, Inner Mongolia Agricultural University, Inner Mongolia, China, 2004. [Google Scholar]
- Yufa, Z. Honeyclover a fine legume species for making green and soil and water conservation legume. Pratacultural Sci. 1993, 06, 54–55. [Google Scholar]
- Ying, C. Study on the Salt Tolerance and Cold Resistance of 3 Clovers. Master’s Thesis, Inner Mongolia Agricultural University, Inner Mongolia, China, 2006. [Google Scholar]
- Mou, D.; Ma, G.; Kazhuo, C.R.; Li, Z.; Zhong, K.; Xie, J.; Li, X. Comparative Analysis of Transcriptome of Caucasian Clover Trifolium ambiuum ambiguum Bieb. under Different Cooling Modes. Acta Agrestia Sin. 2021, 29, 1386–1396. [Google Scholar]
- Mou, D.; Yang, F.; Kazhuo, C.R.; Zhong, Q.; Hui, G.; Xiang, X.; Lai, L. Metabonomic Analysis of Caucasian Clover in Response to Low-Temperature Stress of Different Cooling Modes. Acta Agrestia Sin. 2021, 29, 1877–1884. [Google Scholar]
- Mou, D.; Yang, F.; Kana, C.R.; Hexiduojie; Xu, C.; Xie, J.; Li, X. Physiological responses of Caucasian clover to different cooling modes. Grassl. Turf 2022, 42, 3. [Google Scholar]
- Pengcheng, N.; Fan, H.; Mingjiu, W. Progress in breeding research on Caucasian triticale. J. Grassl. Forage Sci. 2022, 05, 1–9. [Google Scholar]
- Xiujie, Y.; Mingjiu, W.; Fengling, S. Physiologocal research on drought-resistance of Caucasian clover and white clover during seeding period. J. Inn. Mong. Agric. Univ. (Nat. Sci. Ed.) 2004, 02, 119–123. [Google Scholar]
- Xiaoyi, R.; Xu, L.; Kun, Y.; Mingyu, Z.; Taotao, H.; Xiujie, Y. Study of rhizome characters and endogenous hormones of Trifolium ambiguum at different growth stages. Pratacultural Sci. 2019, 36, 1734–1742. [Google Scholar]
- Yi, K.; Yihang, Z.; Yao, H.; Jiaxue, L.; Taotao, H.; Xiu, L.; Oeng, S.; Guowen, C.; Xiujie, Y. Effect of GA3 and 6-BA on rhizome segment growth and endogenous hormone content of Caucasian clover. Acta Prataculturae Sin. 2020, 29, 22–30. [Google Scholar]
- Mingyu, Z.; Xiaoyi, R.; Xu, L.; Boyang, S.; Xiaohan, D.; Xiujie, Y. Effect of IBA and NAA on Trifolium ambiguum seed germination and growth of creeping-root. Pratacultural Sci. 2019, 36, 93–100. [Google Scholar]
- Yunhu, Z.; Mingjiu, W.; Chunqiu, W.; Haijun, C. Stydt on root growing of Trifolium ambiguum under different fertilizer levels. J. Inn. Mong. Agric. Univ. (Nat. Sci. Ed.) 2010, 31, 81–85. [Google Scholar]
- Guilfoyle, T.J.; Hagen, G. Auxin response factors. Curr. Opin. Plant Biol. 2007, 10, 453–460. [Google Scholar] [CrossRef]
- Mockaitis, K.; Estelle, M. Auxin Receptors and Plant Development: A New Signaling Paradigm. Annu. Rev. Cell Dev. Biol. 2008, 24, 55–80. [Google Scholar] [CrossRef]
- Su, Y.H.; Liu, Y.B.; Bai, B.; Zhang, X.S. Establishment of embryonic shoot-root axis is involved in auxin and cytokinin response during Arabidopsis somatic embryogenesis. Front. Plant Sci. 2015, 5, 792. [Google Scholar] [CrossRef]
- Hou, L. Study on the Function of Poplar Auxin Response Factor ARF7 Gene Abstract. Master’s Thesis, Shandong Agricultural University, Tai’an, China, 2021. [Google Scholar]
- Oeller, P.W.; Keller, J.A.; Parks, J.E.; Silbert, J.E.; Theologis, A. Structural characterization of the early indoleacetic acid-inducible genes, PS-IAA4/5 and PS-IAA6, of pea (Pisum sativum L.). J. Mol. Biol. 1993, 233, 789–798. [Google Scholar] [CrossRef]
- Oeller, P.W.; Theologis, A. Induction kinetics of the nuclear proteins encoded by the early indoleacetic acid-inducible genes, PS-IAA4/5 and PS-IAA6, in pea (Pisum sativum L.). Plant J. Cell Mol. Biol. 1995, 7, 37–48. [Google Scholar] [CrossRef]
- Ren, H.; Gray, W.M. SAUR Proteins as Effectors of Hormonal and Environmental Signals in Plant Growth. Mol. Plant 2015, 8, 1153–1164. [Google Scholar] [CrossRef]
- Wojtaczka, P.; Ciarkowska, A.; Starzynska, E.; Ostrowski, M. The GH3 amidosynthetases family and their role in metabolic crosstalk modulation of plant signaling compounds. Phytochemistry 2022, 194, 113039. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.; Kong, Y.; Wang, J.; Kong, Y.Y.; Wang, J.H. Research advances in auxin-responsive SAUR genes. Chin. Bull. Life Sci. 2014, 26, 407–413. [Google Scholar]
- Tiwari, S.B.; Hagen, G.; Guilfoyle, T. The roles of auxin response factor domains in auxin-responsive transcription. Plant Cell 2003, 15, 533–543. [Google Scholar] [CrossRef] [PubMed]
- Liu, K.; Yuan, C.; Li, H.; Lin, W.; Yang, Y.; Shen, C.; Zheng, X. Genome-wide identification and characterization of auxin response factor (ARF) family genes related to flower and fruit development in papaya (Carica papaya L.). BMC Genom. 2015, 16, 901. [Google Scholar] [CrossRef]
- Cance, C.; Martin-Arevalillo, R.; Boubekeur, K.; Dumas, R. Auxin response factors are keys to the many auxin doors. New Phytol. 2022, 235, 402–419. [Google Scholar] [CrossRef]
- Quint, M.; Gray, W.M. Auxin signaling. Curr. Opin. Plant Biol. 2006, 9, 448–453. [Google Scholar] [CrossRef]
- Mengya, S.; Yang, L.; Wei, Z.; Yueping, L. Progress in Mechanism of Auxin Response Factors. Biotechnol. Bull. 2012, 08, 24–28. [Google Scholar]
- Finet, C.; Berne-Dedieu, A.; Scutt, C.P.; Marletaz, F. Evolution of the ARF Gene Family in Land Plants: Old Domains, New Tricks. Mol. Biol. Evol. 2013, 30, 45–56. [Google Scholar] [CrossRef]
- Korasick, D.A.; Jez, J.M.; Strader, L.C. Refining the nuclear auxin response pathway through structural biology. Curr. Opin. Plant Biol. 2015, 27, 22–28. [Google Scholar] [CrossRef]
- Li, S.-B.; Xie, Z.-Z.; Hu, C.-G.; Zhang, J.-Z. A Review of Auxin Response Factors (ARFs) in Plants. Front. Plant Sci. 2016, 7, 47. [Google Scholar] [CrossRef]
- Reed, J.W. Roles and activities of Aux/IAA proteins in Arabidopsis. Trends Plant Sci. 2001, 6, 420–425. [Google Scholar] [CrossRef] [PubMed]
- Ulmasov, T.; Hagen, G.; Guilfoyle, T.J. Dimerization and DNA binding of auxin response factors. Plant J. Cell Mol. Biol. 1999, 19, 309–319. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; Pei, K.; Fu, Y.; Sun, Z.; Li, S.; Liu, H.; Tang, K.; Han, B.; Tao, Y. Genome-wide analysis of the auxin response factors (ARF) gene family in rice (Oryza sativa). Gene 2007, 394, 13–24. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Jiang, H.; Chen, W.; Qian, Y.; Ma, Q.; Cheng, B.; Zhu, S. Genome-wide analysis of the auxin response factor (ARF) gene family in maize (Zea mays). Plant Growth Regul. 2011, 63, 225–234. [Google Scholar] [CrossRef]
- Xing, H.; Pudake, R.N.; Guo, G.; Xing, G.; Hu, Z.; Zhang, Y.; Sun, Q.; Ni, Z. Genome-wide identification and expression profiling of auxin response factor (ARF) gene family in maize. BMC Genom. 2011, 12, 178. [Google Scholar] [CrossRef]
- Kumar, R.; Tyagi, A.K.; Sharma, A.K. Genome-wide analysis of auxin response factor (ARF) gene family from tomato and analysis of their role in flower and fruit development. Mol. Genet. Genom. 2011, 285, 245–260. [Google Scholar] [CrossRef]
- Luo, X.-C.; Sun, M.-H.; Xu, R.-R.; Shu, H.-R.; Wang, J.-W.; Zhang, S.-Z. Genomewide identification and expression analysis of the ARF gene family in apple. J. Genet. 2014, 93, 785–797. [Google Scholar] [CrossRef]
- Kalluri, U.C.; DiFazio, S.P.; Brunner, A.M.; Tuskan, G.A. Genome-wide analysis of Aux/IAA and ARF gene families in Populus trichocarpa. BMC Plant Biol. 2007, 7, 59. [Google Scholar] [CrossRef]
- Le, B.; Nawaz, M.A.; Rehman, H.M.; Le, T.; Yang, S.H.; Golokhvast, K.S.; Son, E.; Chung, G. Genome-wide characterization and expression pattern of auxin response factor (ARF) gene family in soybean and common bean. Genes Genom. 2016, 38, 1165–1178. [Google Scholar] [CrossRef]
- Shen, C.; Yue, R.; Sun, T.; Zhang, L.; Xu, L.; Tie, S.; Wang, H.; Yang, Y. Genome-wide identification and expression analysis of auxin response factor gene family in Medicago truncatula. Front. Plant Sci. 2015, 6, 73. [Google Scholar] [CrossRef]
- Li, Q.Q.; Zhang, Z.; Wang, Y.L.; Zhong, L.Y.; Chao, Z.F.; Gao, Y.Q.; Han, M.L.; Xu, L.; Chao, D.Y. Phytochrome B inhibits darkness-induced hypocotyl adventitious root formation by stabilizing IAA14 and suppressing ARF7 and ARF19. Plant J. 2021, 105, 1689–1702. [Google Scholar] [CrossRef] [PubMed]
- Tang, Y.Y.; Du, G.I.; Xiang, J.; Hu, C.L.; Li, X.T.; Wang, W.H.; Zhu, H.; Qiao, L.X.; Zhao, C.M.; Wang, J.S.; et al. Genome-wide identification of auxin response factor (ARF) gene family and the miR160-ARF18-mediated response to salt stress in peanut (Arachis hypogaea L.). Genomics 2022, 114, 171–184. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Xu, J.; Zhang, Y.; Cui, J.; Hu, H. Transcriptome-wide identification, characterization, and expression analysis of R2R3-MYB gene family during lignin biosynthesis in Chinese cedar (Cryptomeria fortunei Hooibrenk). Ind. Crops Prod. 2022, 182, 114883. [Google Scholar] [CrossRef]
- Burks, D.J.; Azad, R.K. Identification and Network-Enabled Characterization of Auxin Response Factor Genes in Medicago truncatula. Front. Plant Sci. 2016, 7, 1857. [Google Scholar] [CrossRef]
- De Smet, I.; Juergens, G. Patterning the axis in plants-auxin in control. Curr. Opin. Genet. Dev. 2007, 17, 337–343. [Google Scholar] [CrossRef]
- Li, X.; Zhang, X.; Shi, T.; Chen, M.; Jia, C.; Wang, J.; Hou, Z.; Han, J.; Bian, S. Identification of ARF family in blueberry and its potential involvement of fruit development and pH stress response. BMC Genom. 2022, 23, 329. [Google Scholar] [CrossRef]
- Luo, J.; Zhou, J.-J.; Zhang, J.-Z. Aux/IAA Gene Family in Plants: Molecular Structure, Regulation, and Function. Int. J. Mol. Sci. 2018, 19, 259. [Google Scholar] [CrossRef]
- Liscum, E.; Reed, J.W. Genetics of Aux/IAA and ARF action in plant growth and development. Plant Mol. Biol. 2002, 49, 387–400. [Google Scholar] [CrossRef]
- Li, Y.; Qi, Y. Advances in Biological Functions of Aux/IAA Gene Family in Plants. Chin. Bull. Bot. 2022, 57, 30–41. [Google Scholar]
- The Arabidopsis Genome Initiative. Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature 2000, 408, 796–815. [Google Scholar] [CrossRef]
- Liu, Y.; Wang, R.; Yu, J.; Huang, S.; Zhang, Y.; Wei, H.; Wei, Z. Genome-Wide Identification and Characterization of Auxin Response Factor (ARF) Gene Family Involved in Wood Formation and Response to Exogenous Hormone Treatment in Populus trichocarpa. Int. J. Mol. Sci. 2023, 24, 740. [Google Scholar] [CrossRef] [PubMed]
- Ali, S.; Wang, W.; Zhang, Z.; Xie, L.; Boer, D.R.; Khan, N. Genome-Wide Identification, Expression and Interaction Analysis of ARF and AUX/IAA Gene Family in Soybean. Front. Biosci. Landmark 2022, 27, 251. [Google Scholar] [CrossRef] [PubMed]
- Li, P.; Ma, Q.; Qu, C.; Zhu, S.; Zhao, K.; Ma, X.; Li, Z.; Zhang, X.; Gong, F.; Yin, D. Genome-wide identification and expression analysis of auxin response factors in peanut (Arachis hypogaea L.). PeerJ 2021, 9, e12319. [Google Scholar] [CrossRef] [PubMed]
- Okushima, Y.; Overvoorde, P.J.; Arima, K.; Alonso, J.M.; Chan, A.; Chang, C.; Ecker, J.R.; Hughes, B.; Lui, A.; Nguyen, D.; et al. Functional genomic analysis of the AUXIN RESPONSE FACTOR gene family members in Arabidopsis thaliana: Unique and overlapping functions of ARF7 and ARF19. Plant Cell 2005, 17, 444–463. [Google Scholar] [CrossRef]
- Mei, M.; Ai, W.; Liu, L.; Xu, X.; Lu, X. Genome-wide identification of the auxin response factor (ARF) gene family in Magnolia sieboldii and functional analysis of MsARF5. Front. Plant Sci. 2022, 13, 958816. [Google Scholar] [CrossRef]
- Zhang, X.; Jiang, J.; Yang, Y.; Ma, Z.; Meng, L.; Cui, G.; Yin, X. Identification and responding to exogenous hormone of HB-KNOX family based on transcriptome data of Caucasian clover. Gene 2022, 828, 146469. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Harter, K.; Theologis, A. Protein-protein interactions among the Aux/IAA proteins. Proc. Natl. Acad. Sci. USA 1997, 94, 11786–11791. [Google Scholar] [CrossRef]
- Gidhi, A.; Kumar, M.; Mukhopadhyay, K. The auxin response factor gene family in wheat (Triticum aestivum L.): Genome-wide identification, characterization and expression analyses in response to leaf rust. S. Afr. J. Bot. 2021, 140, 312–325. [Google Scholar] [CrossRef]
- Zhang, Y.; Zeng, Z.; Chen, C.; Li, C.; Xia, R.; Li, J. Genome-wide characterization of the auxin response factor (ARF) gene family of litchi (Litchi chinensis Sonn.): Phylogenetic analysis, miRNA regulation and expression changes during fruit abscission. PeerJ 2019, 7, e6677. [Google Scholar] [CrossRef]
- Wang, S.C.; Tiwari, S.B.; Hagen, G.; Guilfoyle, T.J. AUXIN RESPONSE FACTOR7 restores the expression of auxin-responsive genes in mutant Arabidopsis leaf mesophyll protoplasts. Plant Cell 2005, 17, 1979–1993. [Google Scholar] [CrossRef]
- O’Neill, D.P.; Davidson, S.E.; Clarke, V.C.; Yamauchi, Y.; Yamaguchi, S.; Kamiya, Y.; Reid, J.B.; Ross, J.J. Regulation of the gibberellin pathway by auxin and DELLA proteins. Planta 2010, 232, 1141–1149. [Google Scholar] [CrossRef]
- Weston, D.E.; Reid, J.B.; Ross, J.J. Auxin regulation of gibberellin biosynthesis in the roots of pea (Pisum sativum). Funct. Plant Biol. 2009, 36, 362–369. [Google Scholar] [CrossRef] [PubMed]
- Vert, G.; Walcher, C.L.; Chory, J.; Nemhauser, J.L. Integration of auxin and brassinosteroid pathways by Auxin Response Factor 2. Proc. Natl. Acad. Sci. USA 2008, 105, 9829–9834. [Google Scholar] [CrossRef] [PubMed]
- Schruff, M.C.; Spielman, M.; Tiwari, S.; Adams, S.; Fenby, N.; Scott, R.J. The AUXIN RESPONSE FACTOR 2 gene of Arabidopsis links auxin signalling, cell division, and the size of seeds and other organs. Development 2006, 133, 251–261. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Zhang, J.; Li, X.; Zhu, L.; Lian, Z.; Fang, H.; Lu, L.; Lu, Y.; Shi, J.; Chen, J.; et al. The Identification and Expression Analysis of the Nitraria sibirica Pall. Auxin-Response Factor (ARF) Gene Family. Int. J. Mol. Sci. 2022, 23, 11122. [Google Scholar] [CrossRef]
- Chen, J.; Wang, S.; Wu, F.; Wei, M.; Li, J.; Yang, F. Genome-Wide Identification and Functional Characterization of Auxin Response Factor (ARF) Genes in Eggplant. Int. J. Mol. Sci. 2022, 23, 6219. [Google Scholar] [CrossRef]
- Yin, X.J.; Yi, K.; Zhao, Y.H.; Hu, Y.; Li, X.; He, T.T.; Liu, J.X.; Cui, G.W. Revealing the full-length transcriptome of Caucasian clover rhizome development. BMC Plant Biol. 2020, 20, 429. [Google Scholar] [CrossRef]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An Integrative Toolkit Developed for Interactive Analyses of Big Biological Data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef]
- Meng, L.; Yang, Y.; Ma, Z.; Jiang, J.; Zhang, X.; Chen, Z.; Cui, G.; Yin, X. Integrated physiological, transcriptomic and metabolomic analysis of the response of Trifolium pratense L. to Pb toxicity. J. Hazard. Mater. 2022, 436, 129128. [Google Scholar] [CrossRef]


Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jiang, J.; Wang, Z.; Chen, Z.; Wu, Y.; Mu, M.; Nie, W.; Zhao, S.; Cui, G.; Yin, X. Identification and Evolutionary Analysis of the Auxin Response Factor (ARF) Family Based on Transcriptome Data from Caucasian Clover and Analysis of Expression Responses to Hormones. Int. J. Mol. Sci. 2023, 24, 15357. https://doi.org/10.3390/ijms242015357
Jiang J, Wang Z, Chen Z, Wu Y, Mu M, Nie W, Zhao S, Cui G, Yin X. Identification and Evolutionary Analysis of the Auxin Response Factor (ARF) Family Based on Transcriptome Data from Caucasian Clover and Analysis of Expression Responses to Hormones. International Journal of Molecular Sciences. 2023; 24(20):15357. https://doi.org/10.3390/ijms242015357
Chicago/Turabian StyleJiang, Jingwen, Zicheng Wang, Zirui Chen, Yuchen Wu, Meiqi Mu, Wanting Nie, Siwen Zhao, Guowen Cui, and Xiujie Yin. 2023. "Identification and Evolutionary Analysis of the Auxin Response Factor (ARF) Family Based on Transcriptome Data from Caucasian Clover and Analysis of Expression Responses to Hormones" International Journal of Molecular Sciences 24, no. 20: 15357. https://doi.org/10.3390/ijms242015357
APA StyleJiang, J., Wang, Z., Chen, Z., Wu, Y., Mu, M., Nie, W., Zhao, S., Cui, G., & Yin, X. (2023). Identification and Evolutionary Analysis of the Auxin Response Factor (ARF) Family Based on Transcriptome Data from Caucasian Clover and Analysis of Expression Responses to Hormones. International Journal of Molecular Sciences, 24(20), 15357. https://doi.org/10.3390/ijms242015357




