The Genetic Basis of Gene Expression Divergence in Antennae of Two Closely Related Moth Species, Helicoverpa armigera and Helicoverpa assulta
Abstract
:1. Introduction
2. Results
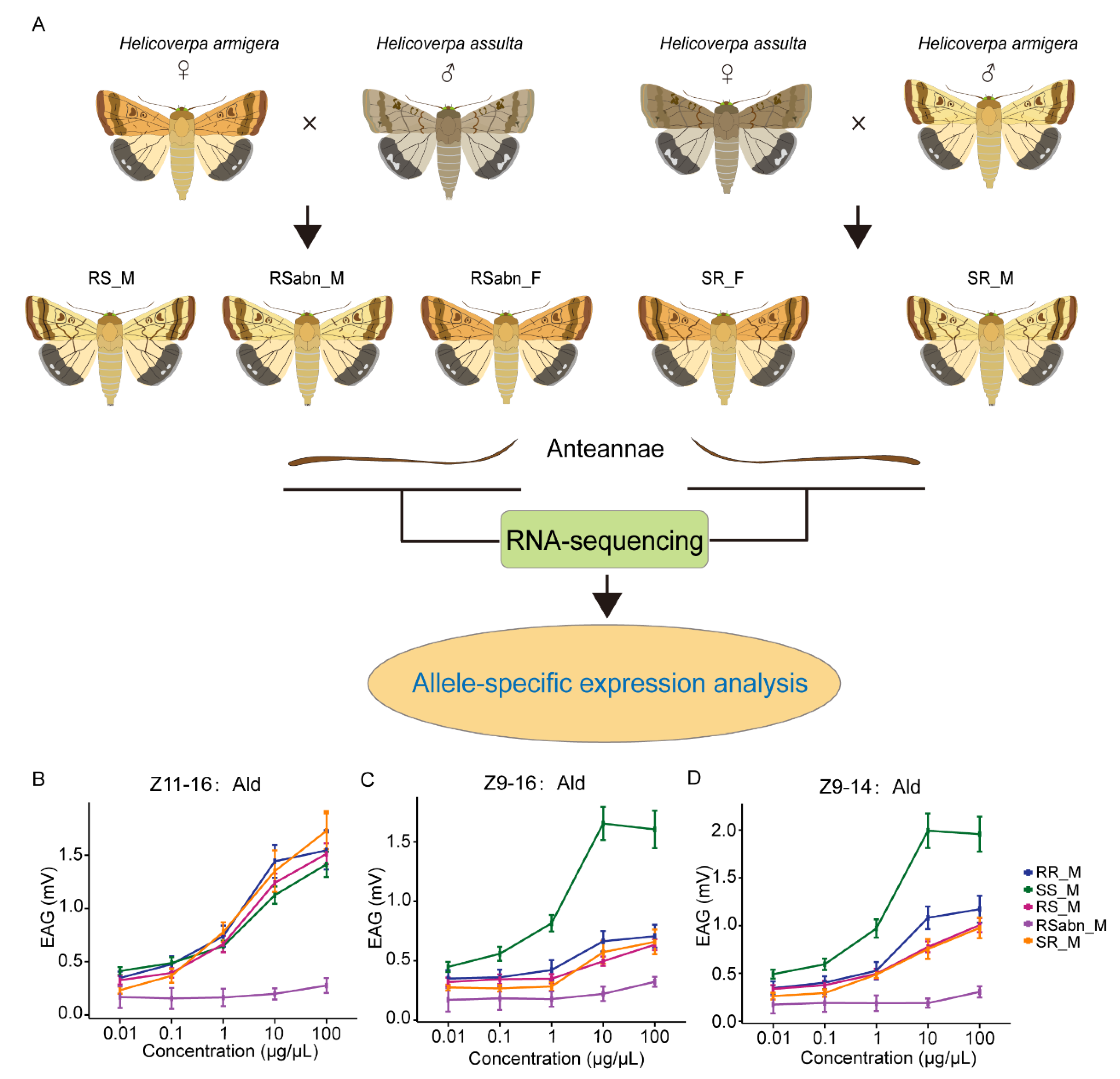
2.1. Interspecific Hybridization and Electrophysiological Responses of H. armigera and H. assulta
2.2. Transcriptome Analysis of H. armigera, H. assulta and F1 Hybrids
2.3. Principal Component Analysis (PCA) of H. armigera, H. assulta and F1 Hybrids
2.4. More Misexpressed Gene in the Antennae of Sterile and Reciprocal F1 Hybrids
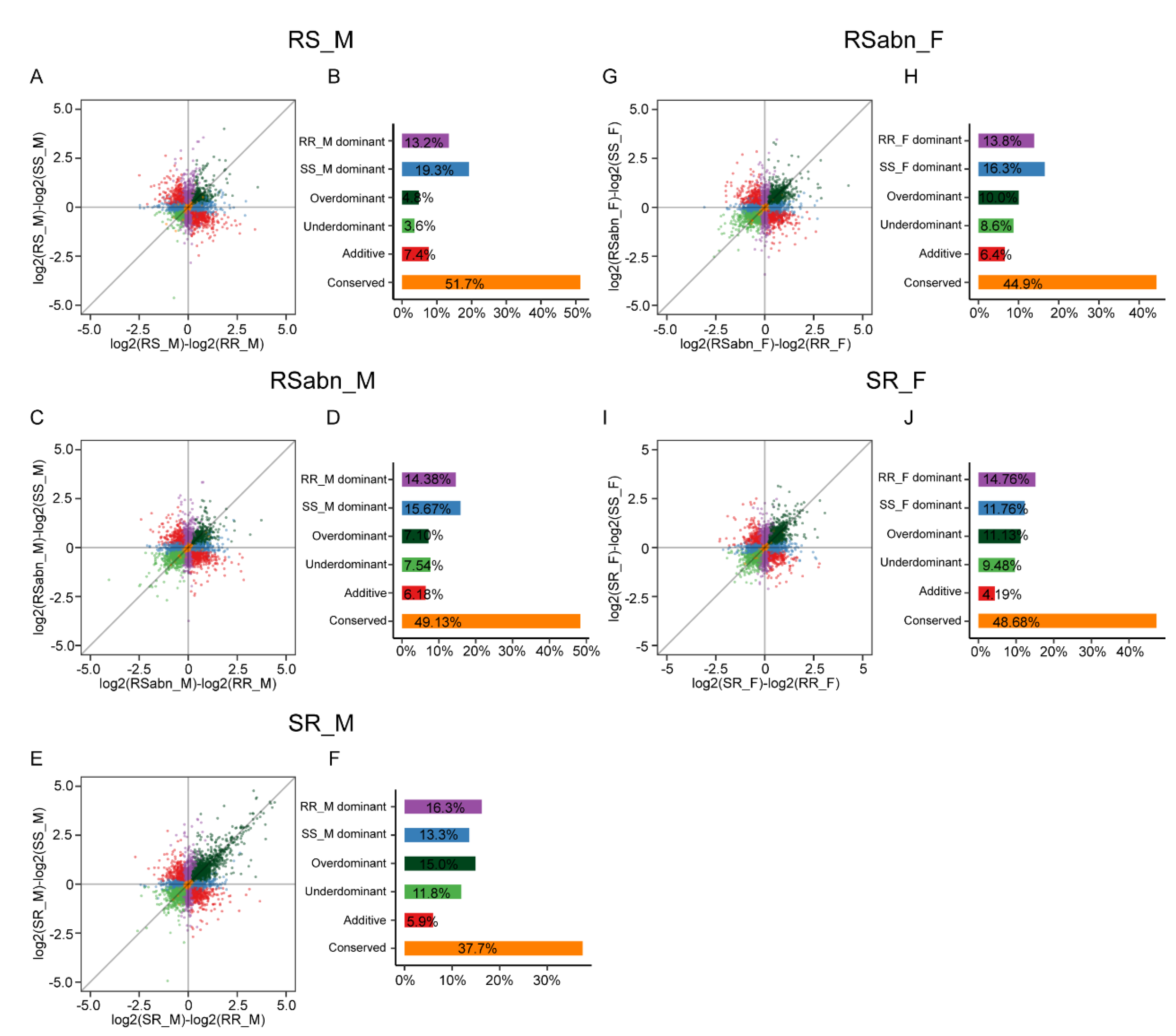
2.5. The Regulatory Patterns in Male Antennae Are Different from Those of Female Antennae
2.6. The Fertile Males in Initial F1 Hybrids Are More Reliable for Inferring Regulatory Patterns
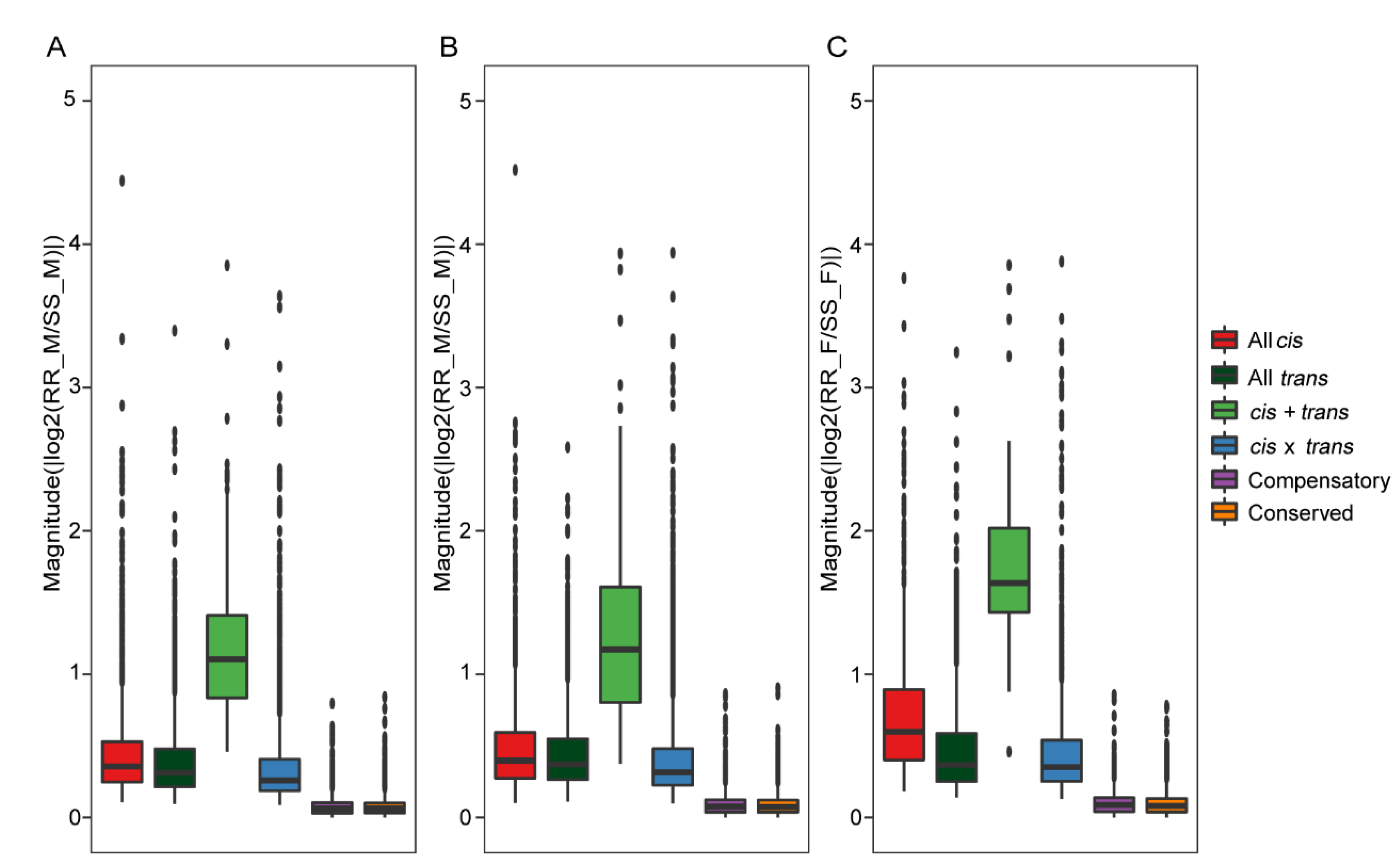
2.7. Contribution of cis- and trans-Regulatory Variants to Gene Expression Divergence in Antennae of H. armigera and H. assulta
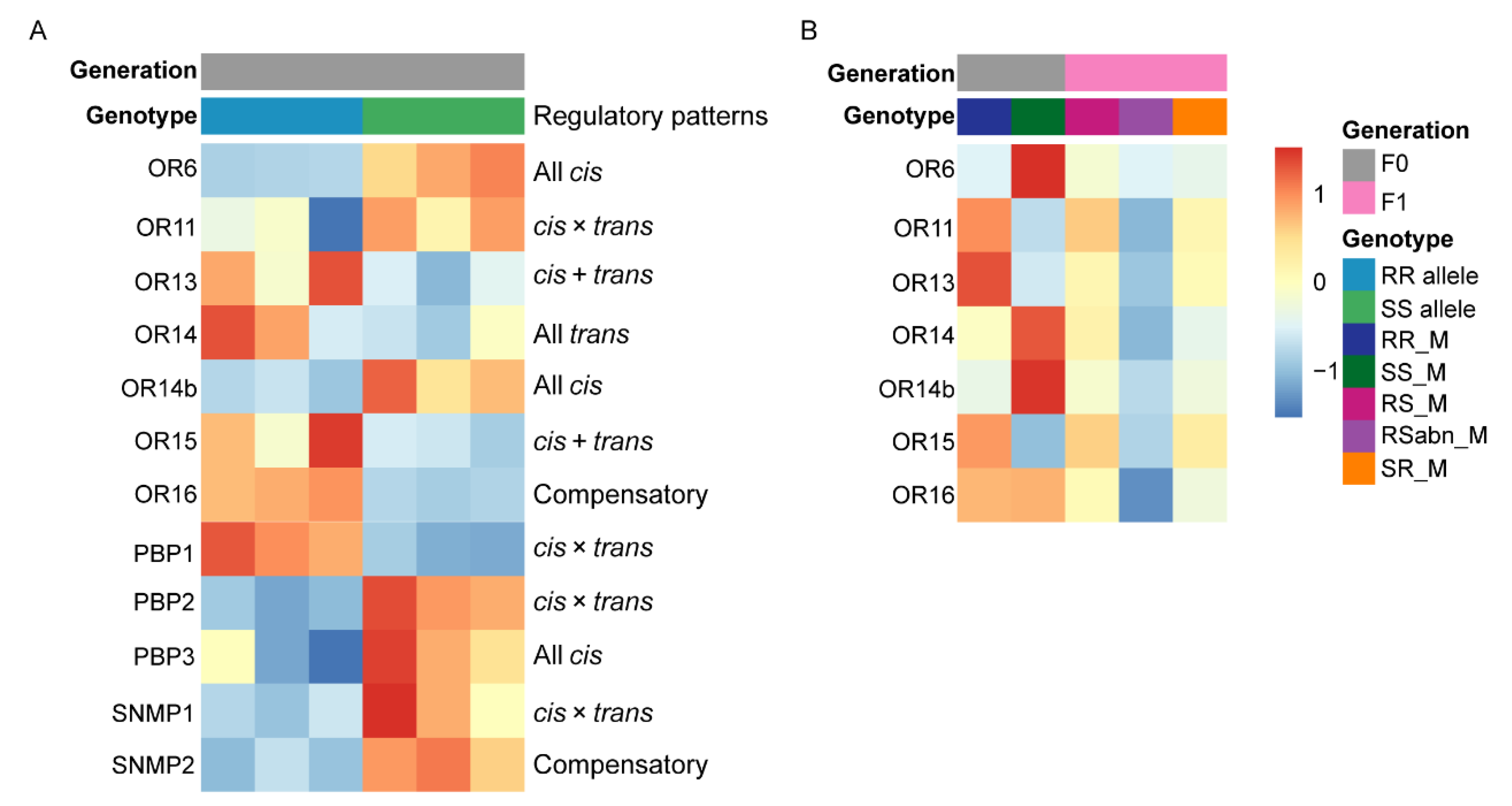
2.8. Regulatory Patterns of Olfactory-Related Protein Genes Expression in Antennae
3. Discussion
3.1. Genetic Basis of Gene Expression Divergence in Antennae of Related Insect Species
3.2. Effects of cis- and trans-Regulatory Variants on Antennal Dimorphism
3.3. Regulatory Patterns Inferred by RS_M Is More Accurate
3.4. cis-Related Regulations Play a Crucial Role in Gene Expression Divergence of Pheromone Perception Related Protein Genes
3.5. Relationships between the Expression Levels of PRs and the Electrophysiological Activities of Antennae
4. Materials and Methods
4.1. Insect Rearing and Interspecific Hybridization
4.2. Antennae Collection, RNA Extraction and Transcriptome Sequencing
4.3. Processing of RNA-seq Datasets
4.4. Measuring ASE and PCA Analysis
4.5. Classification of Inheritance Modes
4.6. Classification of Regulatory Patterns
4.7. Functional Annotation and Gene Ontology
4.8. DNA Extraction and Genomic PCR
4.9. EAG Recordings
4.10. Correlation Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Jiggins, C.D.; Naisbit, R.E.; Coe, R.L.; Mallet, J. Reproductive isolation caused by colour pattern mimicry. Nature 2001, 411, 302–305. [Google Scholar] [CrossRef] [PubMed]
- Mullen, S.P.; Shaw, K.L. Insect speciation rules: Unifying concepts in speciation research. Annu. Rev. Entomol. 2014, 59, 339–361. [Google Scholar] [CrossRef] [PubMed]
- Schluter, D. Evidence for ecological speciation and its alternative. Science 2009, 323, 737–741. [Google Scholar] [CrossRef] [PubMed]
- Schwander, T.; Leimar, O. Genes as leaders and followers in evolution. Trends Ecol. Evol. 2011, 26, 143–151. [Google Scholar] [CrossRef]
- Emerson, J.J.; Hsieh, L.C.; Sung, H.M.; Wang, T.Y.; Huang, C.J.; Lu, H.H.; Lu, M.Y.; Wu, S.H.; Li, W.H. Natural selection on cis and trans regulation in yeasts. Genome Res. 2010, 20, 826–836. [Google Scholar] [CrossRef]
- Hill, M.S.; Vande Zande, P.; Wittkopp, P.J. Molecular and evolutionary processes generating variation in gene expression. Nat. Rev. Genet. 2021, 22, 203–215. [Google Scholar] [CrossRef]
- Schaefke, B.; Emerson, J.J.; Wang, T.Y.; Lu, M.Y.; Hsieh, L.C.; Li, W.H. Inheritance of gene expression level and selective constraints on trans- and cis-regulatory changes in yeast. Mol. Biol. Evol. 2013, 30, 2121–2133. [Google Scholar] [CrossRef]
- Wittkopp, P.J.; Haerum, B.K.; Clark, A.G. Evolutionary changes in cis and trans gene regulation. Nature 2004, 430, 85–88. [Google Scholar] [CrossRef]
- Wittkopp, P.J.; Haerum, B.K.; Clark, A.G. Regulatory changes underlying expression differences within and between Drosophila species. Nat. Genet. 2008, 40, 346–350. [Google Scholar] [CrossRef]
- Shao, L.; Xing, F.; Xu, C.; Zhang, Q.; Che, J.; Wang, X.; Song, J.; Li, X.; Xiao, J.; Chen, L.L.; et al. Patterns of genome-wide allele-specific expression in hybrid rice and the implications on the genetic basis of heterosis. Proc. Natl. Acad. Sci. USA 2019, 116, 5653–5658. [Google Scholar] [CrossRef] [Green Version]
- Carroll, S.B. Evolution at two levels: On genes and form. PLoS Biol. 2005, 3, e245. [Google Scholar] [CrossRef] [PubMed]
- Wray, G.A. The evolutionary significance of cis-regulatory mutations. Nat. Rev. Genet. 2007, 8, 206–216. [Google Scholar] [CrossRef] [PubMed]
- Signor, S.A.; Nuzhdin, S.V. The Evolution of Gene Expression in cis and trans. Trends Genet. 2018, 34, 532–544. [Google Scholar] [CrossRef] [PubMed]
- Meiklejohn, C.D.; Coolon, J.D.; Hartl, D.L.; Wittkopp, P.J. The roles of cis- and trans-regulation in the evolution of regulatory incompatibilities and sexually dimorphic gene expression. Genome Res. 2014, 24, 84–95. [Google Scholar] [CrossRef]
- Mitter, C.; Poole, R.W.; Matthews, M. Biosystematics of the Heliothinae (Lepidoptera, Noctuidae). Annu. Rev. Entomol. 1993, 38, 207–225. [Google Scholar] [CrossRef]
- Wang, H.L.; Zhao, C.H.; Wang, C.Z. Comparative study of sex pheromone composition and biosynthesis in Helicoverpa armigera, H. assulta and their hybrid. Insect Biochem. Mol. Biol. 2005, 35, 575–583. [Google Scholar] [CrossRef]
- Fitt, G.P. The ecology of Heliothis species in relation to agroecosystems. Annu. Rev. Entomol. 1989, 34, 17–52. [Google Scholar] [CrossRef]
- Wang, C.Z.; Dong, J.F. Interspecific hybridization of Helicoverpa armigera and H. assulta (Lepidoptera: Noctuidae). Chin. Sci. Bull. 2001, 46, 489–491. [Google Scholar] [CrossRef]
- Zhao, X.C.; Dong, J.F.; Tang, Q.B.; Yan, Y.H.; Gelbic, I.; Van Loon, J.J.A.; Wang, C.Z. Hybridization between Helicoverpa armigera and Helicoverpa assulta (Lepidoptera: Noctuidae): Development and morphological characterization of F1 hybrids. Bull. Entomol. Res. 2007, 95, 409–416. [Google Scholar] [CrossRef]
- Manoli, D.S.; Foss, M.; Villella, A.; Taylor, B.J.; Hall, J.C.; Baker, B.S. Male-specific fruitless specifies the neural substrates of Drosophila courtship behaviour. Nature 2005, 436, 395–400. [Google Scholar] [CrossRef]
- Schneider, D. Insect Antennae. Annu. Rev. Entomol. 1964, 9, 103–122. [Google Scholar] [CrossRef]
- Yuvaraj, J.K.; Corcoran, J.A.; Andersson, M.N.; Newcomb, R.D.; Anderbrant, O.; Lofstedt, C. Characterization of Odorant Receptors from a Non-ditrysian Moth, Eriocrania semipurpurella Sheds Light on the Origin of Sex Pheromone Receptors in Lepidoptera. Mol. Biol. Evol. 2017, 34, 2733–2746. [Google Scholar] [CrossRef] [PubMed]
- Hansson, B.S.; Stensmyr, M.C. Evolution of insect olfaction. Neuron 2011, 72, 698–711. [Google Scholar] [CrossRef] [PubMed]
- Unbehend, M.; Kozak, G.M.; Koutroumpa, F.; Coates, B.S.; Dekker, T.; Groot, A.T.; Heckel, D.G.; Dopman, E.B. bric a brac controls sex pheromone choice by male European corn borer moths. Nat. Commun. 2021, 12, 2818. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.; Hou, C.; Huang, L.Q.; Yan, F.S.; Wang, C.Z. Peripheral coding of sex pheromone blends with reverse ratios in two Helicoverpa species. PLoS ONE 2013, 8, e70078. [Google Scholar] [CrossRef]
- Zhang, J.; Wang, B.; Dong, S.; Cao, D.; Dong, J.; Walker, W.B.; Liu, Y.; Wang, G. Antennal transcriptome analysis and comparison of chemosensory gene families in two closely related noctuidae moths, Helicoverpa armigera and H. assulta. PLoS ONE 2015, 10, e0117054. [Google Scholar] [CrossRef]
- Wu, H.; Xu, M.; Hou, C.; Huang, L.Q.; Dong, J.F.; Wang, C.Z. Specific olfactory neurons and glomeruli are associated to differences in behavioral responses to pheromone components between two Helicoverpa species. Front. Behav. Neurosci. 2015, 9, 206. [Google Scholar] [CrossRef]
- Yang, K.; Huang, L.Q.; Ning, C.; Wang, C.Z. Two single-point mutations shift the ligand selectivity of a pheromone receptor between two closely related moth species. eLife 2017, 6, e29100. [Google Scholar] [CrossRef]
- Jiang, X.J.; Guo, H.; Di, C.; Yu, S.; Zhu, L.; Huang, L.Q.; Wang, C.Z. Sequence similarity and functional comparisons of pheromone receptor orthologs in two closely related Helicoverpa species. Insect Biochem. Mol. Biol. 2014, 48, 63–74. [Google Scholar] [CrossRef]
- Cui, W.C.; Wang, B.; Guo, M.B.; Liu, Y.; Jacquin-Joly, E.; Yan, S.C.; Wang, G.R. A receptor-neuron correlate for the detection of attractive plant volatiles in Helicoverpa assulta (Lepidoptera: Noctuidae). Insect Biochem. Mol. Biol. 2018, 97, 31–39. [Google Scholar] [CrossRef]
- Wu, H.; Li, R.T.; Dong, J.F.; Jiang, N.J.; Huang, L.Q.; Wang, C.Z. An odorant receptor and glomerulus responding to farnesene in Helicoverpa assulta (Lepidoptera: Noctuidae). Insect Biochem. Mol. Biol. 2019, 115, 103106. [Google Scholar] [CrossRef] [PubMed]
- Guo, H.; Huang, L.Q.; Gong, X.L.; Wang, C.Z. Comparison of functions of pheromone receptor repertoires in Helicoverpa armigera and Helicoverpa assulta using a Drosophila expression system. Insect Biochem. Mol. Biol. 2021, 141, 103702. [Google Scholar] [CrossRef] [PubMed]
- Guo, M.; Du, L.; Chen, Q.; Feng, Y.; Zhang, J.; Zhang, X.; Tian, K.; Cao, S.; Huang, T.; Jacquin-Joly, E.; et al. Odorant Receptors for Detecting Flowering Plant Cues Are Functionally Conserved across Moths and Butterflies. Mol. Biol. Evol. 2021, 38, 1413–1427. [Google Scholar] [CrossRef]
- Sahara, K.; Yoshido, A.; Shibata, F.; Fujikawa-Kojima, N.; Okabe, T.; Tanaka-Okuyama, M.; Yasukochi, Y. FISH identification of Helicoverpa armigera and Mamestra brassicae chromosomes by BAC and fosmid probes. Insect Biochem. Mol. Biol. 2013, 43, 644–653. [Google Scholar] [CrossRef]
- Deng, Z.; Zhang, Y.; Zhang, M.; Huang, J.; Li, C.; Ni, X.; Li, X. Characterization of the First W-Specific Protein-Coding Gene for Sex Identification in Helicoverpa armigera. Front. Genet. 2020, 11, 649. [Google Scholar] [CrossRef] [PubMed]
- Duchemin, W.; Dupont, P.Y.; Campbell, M.A.; Ganley, A.R.; Cox, M.P. HyLiTE: Accurate and flexible analysis of gene expression in hybrid and allopolyploid species. BMC Bioinform. 2015, 16, 8. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.C.; Yan, Y.H.; Wang, C.Z. Behavioral and electrophysiological responses of Helicoverpa assulta, H. armigera (Lepidoptera: Noctuidae), their F1 hybrids and backcross progenies to sex pheromone component blends. J. Comp. Physiol. A Neuroethol. Sens. Neural. Behav. Physiol. 2006, 192, 1037–1047. [Google Scholar] [CrossRef]
- McManus, C.J.; Coolon, J.D.; Duff, M.O.; Eipper-Mains, J.; Graveley, B.R.; Wittkopp, P.J. Regulatory divergence in Drosophila revealed by mRNA-seq. Genome Res. 2010, 20, 816–825. [Google Scholar] [CrossRef]
- Renaut, S.; Nolte, A.W.; Bernatchez, L. Gene expression divergence and hybrid misexpression between lake whitefish species pairs (Coregonus spp. Salmonidae). Mol. Biol. Evol. 2009, 26, 925–936. [Google Scholar] [CrossRef]
- Fraser, H.B. Improving Estimates of Compensatory cis-trans Regulatory Divergence. Trends Genet. 2019, 35, 3–5. [Google Scholar] [CrossRef]
- Vogt, N. Studying divergence in human-chimp hybrid cells. Nat. Methods 2021, 18, 444. [Google Scholar] [CrossRef] [PubMed]
- Coolon, J.D.; McManus, C.J.; Stevenson, K.R.; Graveley, B.R.; Wittkopp, P.J. Tempo and mode of regulatory evolution in Drosophila. Genome Res. 2014, 24, 797–808. [Google Scholar] [CrossRef] [PubMed]
- Bao, Y.; Hu, G.; Grover, C.E.; Conover, J.; Yuan, D.; Wendel, J.F. Unraveling cis and trans regulatory evolution during cotton domestication. Nat. Commun. 2019, 10, 5399. [Google Scholar] [CrossRef]
- Dahanukar, A.; Hallem, E.A.; Carlson, J.R. Insect chemoreception. Curr. Opin. Neurobiol. 2005, 15, 423–430. [Google Scholar] [CrossRef]
- Fleischer, J.; Pregitzer, P.; Breer, H.; Krieger, J. Access to the odor world: Olfactory receptors and their role for signal transduction in insects. Cell. Mol. Life Sci. 2018, 75, 485–508. [Google Scholar] [CrossRef] [PubMed]
- Benton, R.; Vannice, K.S.; Vosshall, L.B. An essential role for a CD36-related receptor in pheromone detection in Drosophila. Nature 2007, 450, 289–293. [Google Scholar] [CrossRef]
- Nozawa, M.; Nei, M. Evolutionary dynamics of olfactory receptor genes in Drosophila species. Proc. Natl. Acad. Sci. USA 2007, 104, 7122–7127. [Google Scholar] [CrossRef]
- Graze, R.M.; McIntyre, L.M.; Main, B.J.; Wayne, M.L.; Nuzhdin, S.V. Regulatory divergence in Drosophila melanogaster and D. simulans, a genomewide analysis of allele-specific expression. Genetics 2009, 183, 547–561. [Google Scholar] [CrossRef]
- Ramirez-Corona, B.A.; Fruth, S.; Ofoegbu, O.; Wunderlich, Z. The mode of expression divergence in Drosophila fat body is infection-specific. Genome Res. 2021, 31, 1024–1034. [Google Scholar] [CrossRef]
- Leal, W. Pheromone reception. Top. Curr. Chem. 2005, 240, 1–36. [Google Scholar]
- Williams, T.M.; Carroll, S.B. Genetic and molecular insights into the development and evolution of sexual dimorphism. Nat. Rev. Genet. 2009, 10, 797–804. [Google Scholar] [CrossRef]
- Demir, E.; Dickson, B.J. Fruitless Splicing Specifies Male Courtship Behavior in Drosophila. Cell 2005, 121, 785–794. [Google Scholar] [CrossRef]
- Wittkopp, P.J.; Kalay, G. Cis-regulatory elements: Molecular mechanisms and evolutionary processes underlying divergence. Nat. Rev. Genet. 2011, 13, 59–69. [Google Scholar] [CrossRef] [PubMed]
- Cooper, R.D.; Shaffer, H.B. Allele-specific expression and gene regulation help explain transgressive thermal tolerance in non-native hybrids of the endangered California tiger salamander (Ambystoma californiense). Mol. Ecol. 2021, 30, 987–1004. [Google Scholar] [CrossRef] [PubMed]
- Gokhman, D.; Agoglia, R.M.; Kinnebrew, M.; Gordon, W.; Sun, D.; Bajpai, V.K.; Naqvi, S.; Chen, C.; Chan, A.; Chen, C.; et al. Human-chimpanzee fused cells reveal cis-regulatory divergence underlying skeletal evolution. Nat. Genet. 2021, 53, 467–476. [Google Scholar] [CrossRef]
- Liu, X.; Li, Y.I.; Pritchard, J.K. Trans Effects on Gene Expression Can Drive Omnigenic Inheritance. Cell 2019, 177, 1022–1034. [Google Scholar] [CrossRef] [PubMed]
- Ngumbi, E.; Chen, L.; Fadamiro, H.Y. Comparative GC-EAD responses of a specialist (Microplitis croceipes) and a generalist (Cotesia marginiventris) parasitoid to cotton volatiles induced by two caterpillar species. J. Chem. Ecol. 2009, 35, 1009–1020. [Google Scholar] [CrossRef]
- Sun, J.G.; Huang, L.Q.; Wang, C.Z. Electrophysiological and behavioral responses of Helicoverpa assulta (Lepidoptera: Noctuidae) to tobacco volatiles. Arthropod-Plant Interact. 2012, 6, 375–384. [Google Scholar] [CrossRef]
- Dobzhansky, T. Genetics and the Origin of Species; Columbia University Press: New York, NY, USA, 1937. [Google Scholar]
- Muller, H.J. Bearings of the Drosophila work on systematics. New Syst. Oxf. 1940, 1940, 185–268. [Google Scholar]
- Kerwin, R.E.; Sweigart, A.L. Rampant Misexpression in a Mimulus (Monkeyflower) Introgression Line Caused by Hybrid Sterility, Not Regulatory Divergence. Mol. Biol. Evol. 2020, 37, 2084–2098. [Google Scholar] [CrossRef]
- Mack, K.L.; Nachman, M.W. Gene Regulation and Speciation. Trends Genet. 2017, 33, 68–80. [Google Scholar] [CrossRef] [PubMed]
- Wei, K.H.; Clark, A.G.; Barbash, D.A. Limited gene misregulation is exacerbated by allele-specific upregulation in lethal hybrids between Drosophila melanogaster and Drosophila simulans. Mol. Biol. Evol. 2014, 31, 1767–1778. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Leary, G.P.; Allen, J.E.; Bunger, P.L.; Luginbill, J.B.; Linn, C.E., Jr.; Macallister, I.E.; Kavanaugh, M.P.; Wanner, K.W. Single mutation to a sex pheromone receptor provides adaptive specificity between closely related moth species. Proc. Natl. Acad. Sci. USA 2012, 109, 14081–14086. [Google Scholar] [CrossRef] [PubMed]
- Gould, F.; Estock, M.; Hillier, N.K.; Powell, B.; Groot, A.T.; Ward, C.M.; Emerson, J.L.; Schal, C.; Vickers, N.J. Sexual isolation of male moths explained by a single pheromone response QTL containing four receptor genes. Proc. Natl. Acad. Sci. USA 2010, 107, 8660–8665. [Google Scholar] [CrossRef] [PubMed]
- Yang, K.; Wang, C.Z. Review of pheromone receptors in heliothine species: Expression, function, and evolution. Entomol. Exp. Appl. 2020, 169, 156–171. [Google Scholar] [CrossRef]
- Prieto-Godino, L.L.; Rytz, R.; Cruchet, S.; Bargeton, B.; Abuin, L.; Silbering, A.F.; Ruta, V.; Dal Peraro, M.; Benton, R. Evolution of Acid-Sensing Olfactory Circuits in Drosophilids. Neuron 2017, 93, 661–676.e6. [Google Scholar] [CrossRef]
- Pickrell, J.K.; Marioni, J.C.; Pai, A.A.; Degner, J.F.; Engelhardt, B.E.; Nkadori, E.; Veyrieras, J.B.; Stephens, M.; Gilad, Y.; Pritchard, J.K. Understanding mechanisms underlying human gene expression variation with RNA sequencing. Nature 2010, 464, 768–772. [Google Scholar] [CrossRef]
- Tehranchi, A.; Hie, B.; Dacre, M.; Kaplow, I.; Pettie, K.; Combs, P.; Fraser, H.B. Fine-mapping cis-regulatory variants in diverse human populations. eLife 2019, 8, e39595. [Google Scholar] [CrossRef]
- Fraser, H.B. Genome-wide approaches to the study of adaptive gene expression evolution. Bioessays 2011, 33, 469–477. [Google Scholar] [CrossRef]
- Groot, A.T.; Dekker, T.; Heckel, D.G. The Genetic Basis of Pheromone Evolution in Moths. Annu. Rev. Entomol. 2016, 61, 99–117. [Google Scholar] [CrossRef]
- Chang, H.; Guo, M.; Wang, B.; Liu, Y.; Dong, S.; Wang, G. Sensillar expression and responses of olfactory receptors reveal different peripheral coding in two Helicoverpa species using the same pheromone components. Sci. Rep. 2016, 6, 18742. [Google Scholar] [CrossRef] [PubMed]
- Wu, K.; Gong, P. A new and practical artificial diet for the cotton bollworm. Entomol. Sin. 1997, 4, 277–282. [Google Scholar]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lunter, G.; Goodson, M. Stampy: A statistical algorithm for sensitive and fast mapping of Illumina sequence reads. Genome Res. 2011, 21, 936–939. [Google Scholar] [CrossRef]
- Pearce, S.L.; Clarke, D.F.; East, P.D.; Elfekih, S.; Gordon, K.H.J.; Jermiin, L.S.; McGaughran, A.; Oakeshott, J.G.; Papanicolaou, A.; Perera, O.P.; et al. Genomic innovations, transcriptional plasticity and gene loss underlying the evolution and divergence of two highly polyphagous and invasive Helicoverpa pest species. BMC Biol. 2017, 15, 63. [Google Scholar] [CrossRef]
- Wang, L.; Israel, J.W.; Edgar, A.; Raff, R.A.; Raff, E.C.; Byrne, M.; Wray, G.A. Genetic basis for divergence in developmental gene expression in two closely related sea urchins. Nat. Ecol. Evol. 2020, 4, 831–840. [Google Scholar] [CrossRef]
- Kolde, R. Pheatmap: Pretty Heatmaps. R Package Version 1.0.12. 2019. Available online: https://rdrr.io/cran/pheatmap/ (accessed on 1 May 2019).
- Buchfink, B.; Reuter, K.; Drost, H.G. Sensitive protein alignments at tree-of-life scale using DIAMOND. Nat. Methods 2021, 18, 366–368. [Google Scholar] [CrossRef]
- Wu, T.; Hu, E.; Xu, S.; Chen, M.; Guo, P.; Dai, Z.; Feng, T.; Zhou, L.; Tang, W.; Zhan, L.; et al. clusterProfiler 4.0: A universal enrichment tool for interpreting omics data. Innovation 2021, 2, 100141. [Google Scholar] [CrossRef]


Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Guo, P.-P.; Li, G.-C.; Dong, J.-F.; Gong, X.-L.; Wang, L.; Yang, K.; Yang, J.; Huang, L.-Q.; Wang, C.-Z. The Genetic Basis of Gene Expression Divergence in Antennae of Two Closely Related Moth Species, Helicoverpa armigera and Helicoverpa assulta. Int. J. Mol. Sci. 2022, 23, 10050. https://doi.org/10.3390/ijms231710050
Guo P-P, Li G-C, Dong J-F, Gong X-L, Wang L, Yang K, Yang J, Huang L-Q, Wang C-Z. The Genetic Basis of Gene Expression Divergence in Antennae of Two Closely Related Moth Species, Helicoverpa armigera and Helicoverpa assulta. International Journal of Molecular Sciences. 2022; 23(17):10050. https://doi.org/10.3390/ijms231710050
Chicago/Turabian StyleGuo, Ping-Ping, Guo-Cheng Li, Jun-Feng Dong, Xin-Lin Gong, Lingyu Wang, Ke Yang, Jun Yang, Ling-Qiao Huang, and Chen-Zhu Wang. 2022. "The Genetic Basis of Gene Expression Divergence in Antennae of Two Closely Related Moth Species, Helicoverpa armigera and Helicoverpa assulta" International Journal of Molecular Sciences 23, no. 17: 10050. https://doi.org/10.3390/ijms231710050
APA StyleGuo, P.-P., Li, G.-C., Dong, J.-F., Gong, X.-L., Wang, L., Yang, K., Yang, J., Huang, L.-Q., & Wang, C.-Z. (2022). The Genetic Basis of Gene Expression Divergence in Antennae of Two Closely Related Moth Species, Helicoverpa armigera and Helicoverpa assulta. International Journal of Molecular Sciences, 23(17), 10050. https://doi.org/10.3390/ijms231710050






