A Cancer-Related microRNA Signature Shows Biomarker Utility in Multiple Myeloma
Abstract
:1. Introduction
2. Results
2.1. Development and Optimization of Real-Time qPCR Assays
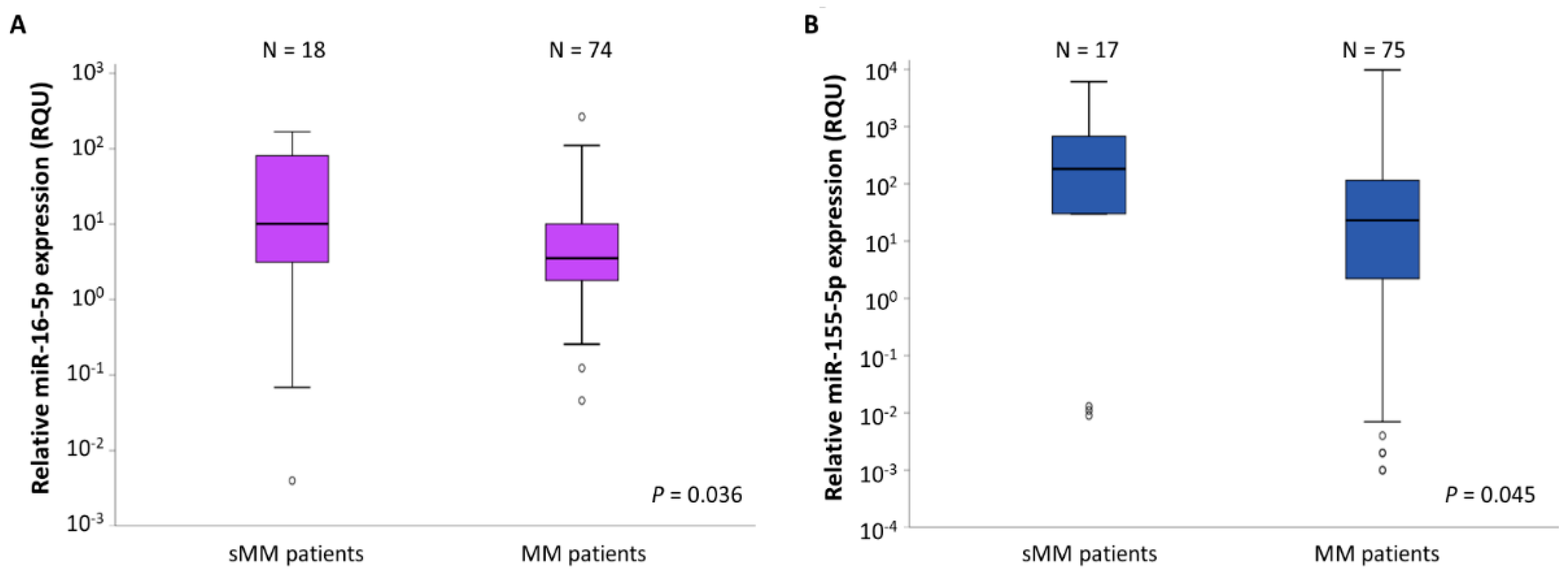
2.2. miR-16-5p and miR-155-5p Levels Are Significantly Lower in MM Patients Compared to sMM Patients
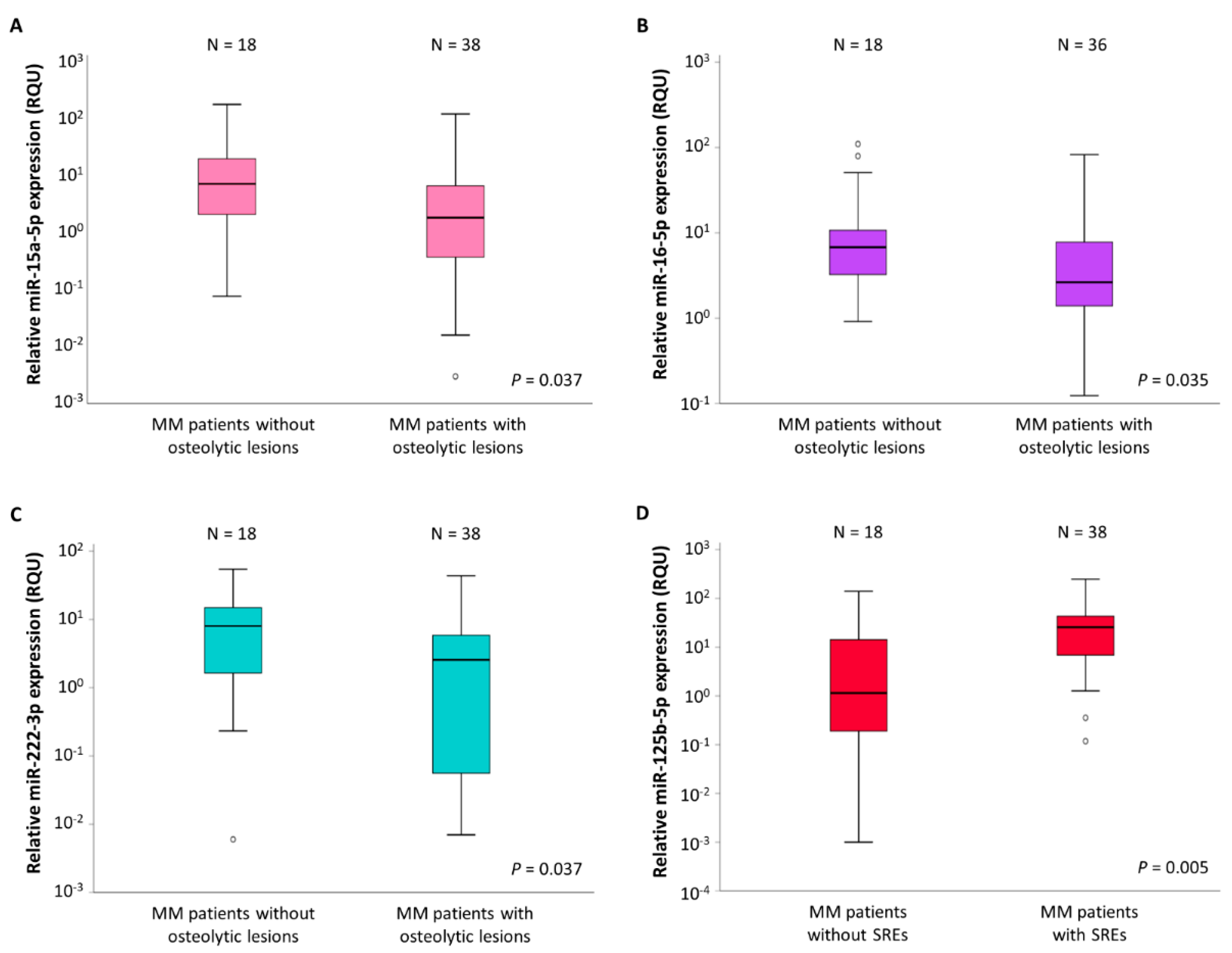
2.3. Association of the Expression of Three of the Investigated miRNAs with MM Bone Disease (MMBD)
2.4. miR-125b-5p Levels Are Associated with MMBD Severity
2.5. miR-223-3p Offers a Putative Prognostic Value in MM
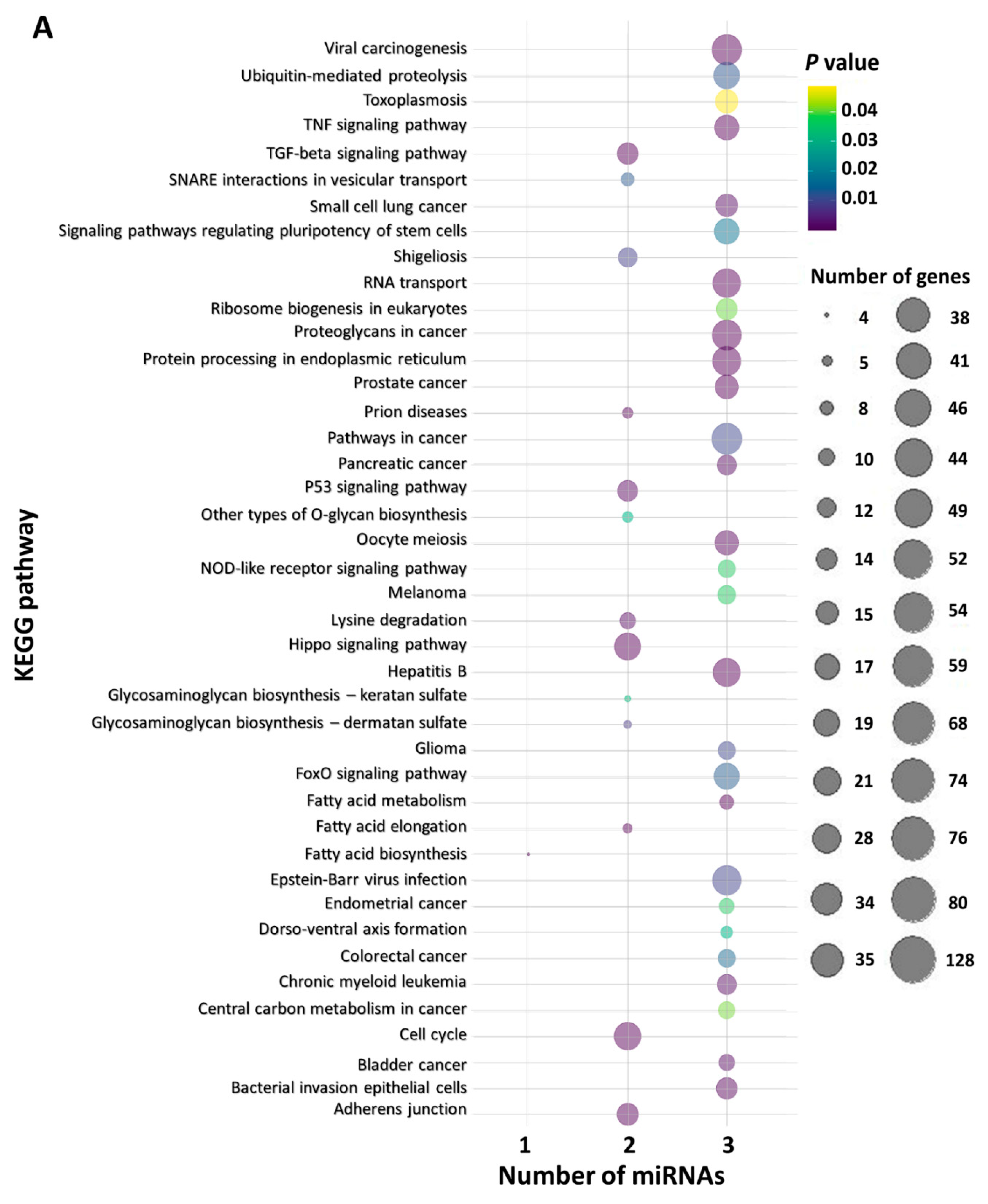
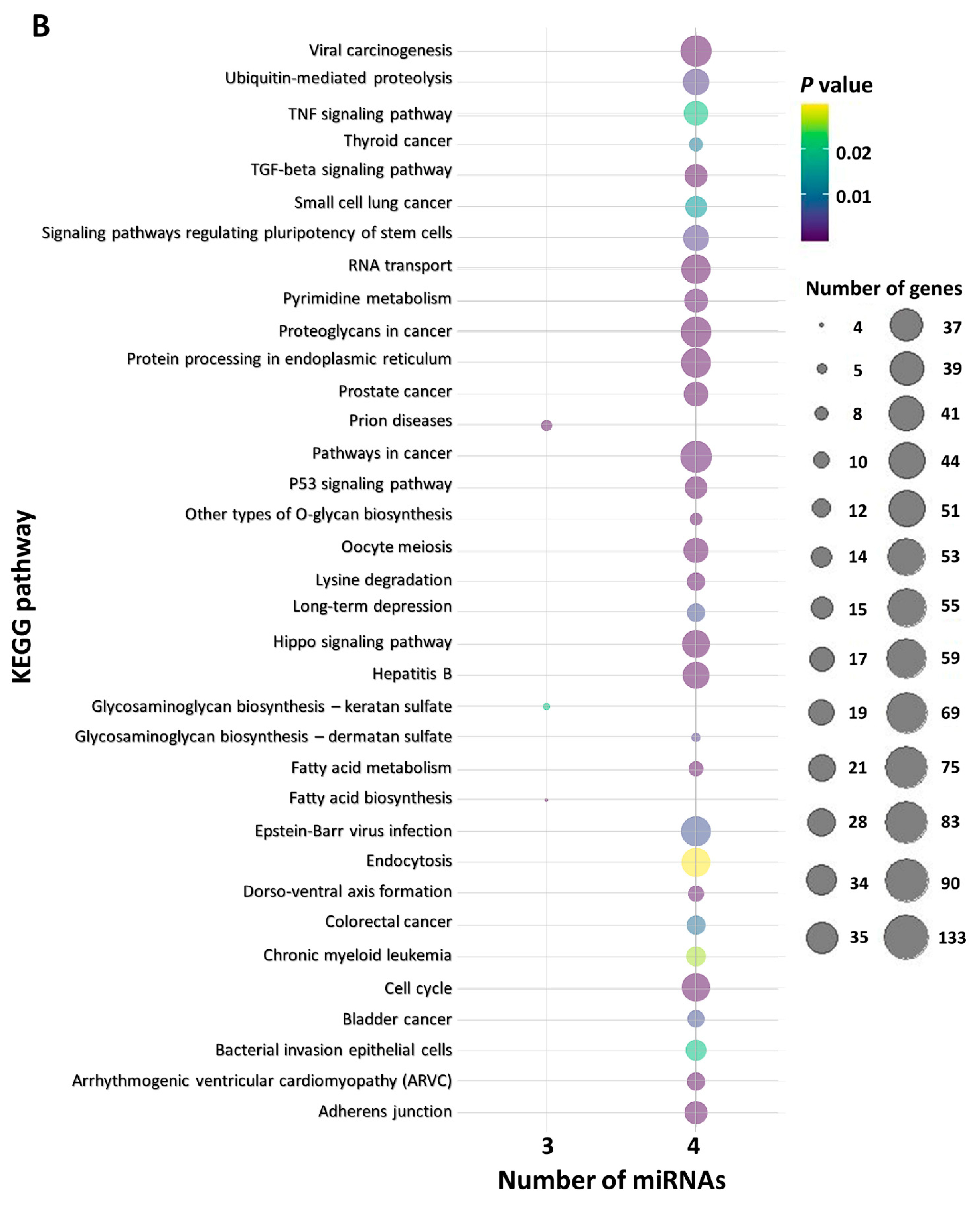
2.6. In Silico Functional miRNA Analysis
3. Discussion
4. Materials and Methods
4.1. Study Participants
4.2. CD138+ Plasma Cell Selection
4.3. RNA Isolation, In Vitro Polyadenylation, and Reverse Transcription
4.4. Quantification of miRNA Expression Using Real-Time qPCR
4.5. Biostatistics
4.6. Functional In Silico Analysis for miRNA Target Prediction and KEGG Pathway Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Usmani, S.Z.; Hoering, A.; Cavo, M.; Miguel, J.S.; Goldschimdt, H.; Hajek, R.; Turesson, I.; Lahuerta, J.J.; Attal, M.; Barlogie, B.; et al. Clinical predictors of long-term survival in newly diagnosed transplant eligible multiple myeloma—An IMWG Research Project. Blood Cancer J. 2018, 8, 123. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rajkumar, S.V.; Dimopoulos, M.A.; Palumbo, A.; Blade, J.; Merlini, G.; Mateos, M.V.; Kumar, S.; Hillengass, J.; Kastritis, E.; Richardson, P.; et al. International Myeloma Working Group updated criteria for the diagnosis of multiple myeloma. Lancet Oncol. 2014, 15, e538–e548. [Google Scholar] [CrossRef]
- Kumar, S.K.; Rajkumar, V.; Kyle, R.A.; van Duin, M.; Sonneveld, P.; Mateos, M.V.; Gay, F.; Anderson, K.C. Multiple myeloma. Nat. Rev. Dis. Primers 2017, 3, 17046. [Google Scholar] [CrossRef] [PubMed]
- Colombo, M.; Giannandrea, D.; Lesma, E.; Basile, A.; Chiaramonte, R. Extracellular Vesicles Enhance Multiple Myeloma Metastatic Dissemination. Int. J. Mol. Sci. 2019, 20, 3236. [Google Scholar] [CrossRef] [Green Version]
- Lee, Y.; Kim, M.; Han, J.; Yeom, K.H.; Lee, S.; Baek, S.H.; Kim, V.N. MicroRNA genes are transcribed by RNA polymerase II. EMBO J. 2004, 23, 4051–4060. [Google Scholar] [CrossRef]
- Liu, J.; Carmell, M.A.; Rivas, F.V.; Marsden, C.G.; Thomson, J.M.; Song, J.J.; Hammond, S.M.; Joshua-Tor, L.; Hannon, G.J. Argonaute2 is the catalytic engine of mammalian RNAi. Science 2004, 305, 1437–1441. [Google Scholar] [CrossRef] [Green Version]
- Guo, H.; Ingolia, N.T.; Weissman, J.S.; Bartel, D.P. Mammalian microRNAs predominantly act to decrease target mRNA levels. Nature 2010, 466, 835–840. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gebert, L.F.R.; MacRae, I.J. Regulation of microRNA function in animals. Nat. Rev. Mol. Cell Biol. 2019, 20, 21–37. [Google Scholar] [CrossRef]
- Peng, Y.; Croce, C.M. The role of MicroRNAs in human cancer. Signal. Transduct. Target. Ther. 2016, 1, 15004. [Google Scholar] [CrossRef] [Green Version]
- Soliman, A.M.; Lin, T.S.; Mahakkanukrauh, P.; Das, S. Role of microRNAs in Diagnosis, Prognosis and Management of Multiple Myeloma. Int. J. Mol. Sci. 2020, 21, 7539. [Google Scholar] [CrossRef]
- Papadimitriou, M.A.; Papanota, A.M.; Adamopoulos, P.G.; Pilala, K.M.; Liacos, C.I.; Malandrakis, P.; Mavrianou-Koutsoukou, N.; Patseas, D.; Eleutherakis-Papaiakovou, E.; Gavriatopoulou, M.; et al. miRNA-seq and clinical evaluation in multiple myeloma: miR-181a overexpression predicts short-term disease progression and poor post-treatment outcome. Br. J. Cancer 2021, 21, 1602. [Google Scholar] [CrossRef]
- Artemaki, P.I.; Letsos, P.A.; Zoupa, I.C.; Katsaraki, K.; Karousi, P.; Papageorgiou, S.G.; Pappa, V.; Scorilas, A.; Kontos, C.K. The Multifaceted Role and Utility of MicroRNAs in Indolent B-Cell Non-Hodgkin Lymphomas. Biomedicines 2021, 9, 333. [Google Scholar] [CrossRef] [PubMed]
- Papanota, A.M.; Karousi, P.; Kontos, C.K.; Ntanasis-Stathopoulos, I.; Scorilas, A.; Terpos, E. Multiple Myeloma Bone Disease: Implication of MicroRNAs in Its Molecular Background. Int. J. Mol. Sci. 2021, 22, 2375. [Google Scholar] [CrossRef]
- Condrat, C.E.; Thompson, D.C.; Barbu, M.G.; Bugnar, O.L.; Boboc, A.; Cretoiu, D.; Suciu, N.; Cretoiu, S.M.; Voinea, S.C. miRNAs as Biomarkers in Disease: Latest Findings Regarding Their Role in Diagnosis and Prognosis. Cells 2020, 9, 276. [Google Scholar] [CrossRef] [Green Version]
- Handa, H.; Murakami, Y.; Ishihara, R.; Kimura-Masuda, K.; Masuda, Y. The Role and Function of microRNA in the Pathogenesis of Multiple Myeloma. Cancers 2019, 11, 1738. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Papanota, A.M.; Tsiakanikas, P.; Kontos, C.K.; Malandrakis, P.; Liacos, C.I.; Ntanasis-Stathopoulos, I.; Kanellias, N.; Gavriatopoulou, M.; Kastritis, E.; Avgeris, M.; et al. A Molecular Signature of Circulating MicroRNA Can Predict Osteolytic Bone Disease in Multiple Myeloma. Cancers 2021, 13, 3877. [Google Scholar] [CrossRef] [PubMed]
- Karousi, P.; Papanota, A.-M.; Artemaki, P.I.; Liacos, C.-I.; Patseas, D.; Mavrianou-Koutsoukou, N.; Liosi, A.-A.; Kalioraki, M.-A.; Ntanasis-Stathopoulos, I.; Gavriatopoulou, M.; et al. tRNA Derivatives in Multiple Myeloma: Investigation of the Potential Value of a tRNA-Derived Molecular Signature. Biomedicines 2021, 9, 1811. [Google Scholar] [CrossRef]
- Lerner, M.; Harada, M.; Loven, J.; Castro, J.; Davis, Z.; Oscier, D.; Henriksson, M.; Sangfelt, O.; Grander, D.; Corcoran, M.M. DLEU2, frequently deleted in malignancy, functions as a critical host gene of the cell cycle inhibitory microRNAs miR-15a and miR-16-1. Exp. Cell Res. 2009, 315, 2941–2952. [Google Scholar] [CrossRef] [PubMed]
- Binder, M.; Rajkumar, S.V.; Ketterling, R.P.; Greipp, P.T.; Dispenzieri, A.; Lacy, M.Q.; Gertz, M.A.; Buadi, F.K.; Hayman, S.R.; Hwa, Y.L.; et al. Prognostic implications of abnormalities of chromosome 13 and the presence of multiple cytogenetic high-risk abnormalities in newly diagnosed multiple myeloma. Blood Cancer J. 2017, 7, e600. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gao, X.; Zhang, R.; Qu, X.; Zhao, M.; Zhang, S.; Wu, H.; Jianyong, L.; Chen, L. MiR-15a, miR-16-1 and miR-17-92 cluster expression are linked to poor prognosis in multiple myeloma. Leuk. Res. 2012, 36, 1505–1509. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Xu, Y.; Deng, S.; Li, Z.; Zou, D.; Yi, S.; Sui, W.; Hao, M.; Qiu, L. MicroRNA-15a/16-1 cluster located at chromosome 13q14 is down-regulated but displays different expression pattern and prognostic significance in multiple myeloma. Oncotarget 2015, 6, 38270–38282. [Google Scholar] [CrossRef] [PubMed]
- Roccaro, A.M.; Sacco, A.; Thompson, B.; Leleu, X.; Azab, A.K.; Azab, F.; Runnels, J.; Jia, X.; Ngo, H.T.; Melhem, M.R.; et al. microRNAs 15a and 16 regulate tumor proliferation in multiple myeloma. Blood 2009, 113, 6669–6680. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Zhou, L.; Shi, M.; Kuang, Y.; Fang, L. Downregulation of miRNA-15a and miRNA-16 promote tumor proliferation in multiple myeloma by increasing CABIN1 expression. Oncol. Lett. 2018, 15, 1287–1296. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sun, C.Y.; She, X.M.; Qin, Y.; Chu, Z.B.; Chen, L.; Ai, L.S.; Zhang, L.; Hu, Y. miR-15a and miR-16 affect the angiogenesis of multiple myeloma by targeting VEGF. Carcinogenesis 2013, 34, 426–435. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, W.; Corrigan-Cummins, M.; Barber, E.A.; Saleh, L.M.; Zingone, A.; Ghafoor, A.; Costello, R.; Zhang, Y.; Kurlander, R.J.; Korde, N.; et al. Aberrant Levels of miRNAs in Bone Marrow Microenvironment and Peripheral Blood of Myeloma Patients and Disease Progression. J. Mol. Diagn. JMD 2015, 17, 669–678. [Google Scholar] [CrossRef] [PubMed]
- Gutierrez, N.C.; Sarasquete, M.E.; Misiewicz-Krzeminska, I.; Delgado, M.; De Las Rivas, J.; Ticona, F.V.; Ferminan, E.; Martin-Jimenez, P.; Chillon, C.; Risueno, A.; et al. Deregulation of microRNA expression in the different genetic subtypes of multiple myeloma and correlation with gene expression profiling. Leukemia 2010, 24, 629–637. [Google Scholar] [CrossRef] [PubMed]
- Amodio, N.; Gallo Cantafio, M.E.; Botta, C.; Agosti, V.; Federico, C.; Caracciolo, D.; Ronchetti, D.; Rossi, M.; Driessen, C.; Neri, A.; et al. Replacement of miR-155 Elicits Tumor Suppressive Activity and Antagonizes Bortezomib Resistance in Multiple Myeloma. Cancers 2019, 11, 236. [Google Scholar] [CrossRef] [Green Version]
- Misso, G.; Zarone, M.R.; Lombardi, A.; Grimaldi, A.; Cossu, A.M.; Ferri, C.; Russo, M.; Vuoso, D.C.; Luce, A.; Kawasaki, H.; et al. miR-125b Upregulates miR-34a and Sequentially Activates Stress Adaption and Cell Death Mechanisms in Multiple Myeloma. Mol. Ther. Nucleic Acids 2019, 16, 391–406. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Di Martino, M.T.; Gulla, A.; Cantafio, M.E.; Lionetti, M.; Leone, E.; Amodio, N.; Guzzi, P.H.; Foresta, U.; Conforti, F.; Cannataro, M.; et al. In vitro and in vivo anti-tumor activity of miR-221/222 inhibitors in multiple myeloma. Oncotarget 2013, 4, 242–255. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xu, J.; Su, Y.; Xu, A.; Fan, F.; Mu, S.; Chen, L.; Chu, Z.; Zhang, B.; Huang, H.; Zhang, J.; et al. miR-221/222-Mediated Inhibition of Autophagy Promotes Dexamethasone Resistance in Multiple Myeloma. Mol. Ther. J. Am. Soc. Gene Ther. 2019, 27, 559–570. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, W.L.; Chi, C.T.; Meng, X.H.; Liang, S.D. miRNA15a5p facilitates the bone marrow stem cell apoptosis of femoral head necrosis through the Wnt/betacatenin/PPARgamma signaling pathway. Mol. Med. Rep. 2019, 19, 4779–4787. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Duan, L.; Zhao, H.; Xiong, Y.; Tang, X.; Yang, Y.; Hu, Z.; Li, C.; Chen, S.; Yu, X. miR-16-2* Interferes with WNT5A to Regulate Osteogenesis of Mesenchymal Stem Cells. Cell Physiol. Biochem. 2018, 51, 1087–1102. [Google Scholar] [CrossRef]
- Yu, T.; You, X.; Zhou, H.; He, W.; Li, Z.; Li, B.; Xia, J.; Zhu, H.; Zhao, Y.; Yu, G.; et al. MiR-16-5p regulates postmenopausal osteoporosis by directly targeting VEGFA. Aging 2020, 12, 9500–9514. [Google Scholar] [CrossRef]
- Yoshizuka, M.; Nakasa, T.; Kawanishi, Y.; Hachisuka, S.; Furuta, T.; Miyaki, S.; Adachi, N.; Ochi, M. Inhibition of microRNA-222 expression accelerates bone healing with enhancement of osteogenesis, chondrogenesis, and angiogenesis in a rat refractory fracture model. J. Orthop. Sci. 2016, 21, 852–858. [Google Scholar] [CrossRef] [Green Version]
- Wang, H.; Xie, Z.; Hou, T.; Li, Z.; Huang, K.; Gong, J.; Zhou, W.; Tang, K.; Xu, J.; Dong, S. MiR-125b Regulates the Osteogenic Differentiation of Human Mesenchymal Stem Cells by Targeting BMPR1b. Cell Physiol. Biochem. 2017, 41, 530–542. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.; Zhang, L.; Yan, C.; Wang, F.; Zhang, Y. Overexpression of miR125b Promotes Osteoporosis Through miR-125b-TRAF6 Pathway in Postmenopausal Ovariectomized Rats. Diabetes Metab. Syndr. Obes. 2021, 14, 671–682. [Google Scholar] [CrossRef] [PubMed]
- Hillengass, J.; Usmani, S.; Rajkumar, S.V.; Durie, B.G.M.; Mateos, M.V.; Lonial, S.; Joao, C.; Anderson, K.C.; Garcia-Sanz, R.; Riva, E.; et al. International myeloma working group consensus recommendations on imaging in monoclonal plasma cell disorders. Lancet Oncol. 2019, 20, e302–e312. [Google Scholar] [CrossRef]
- Palumbo, A.; Avet-Loiseau, H.; Oliva, S.; Lokhorst, H.M.; Goldschmidt, H.; Rosinol, L.; Richardson, P.; Caltagirone, S.; Lahuerta, J.J.; Facon, T.; et al. Revised International Staging System for Multiple Myeloma: A Report From International Myeloma Working Group. J. Clin. Oncol. Off. J. Am. Soc. Clin. Oncol. 2015, 33, 2863–2869. [Google Scholar] [CrossRef] [PubMed]
- Obeng, E.A.; Carlson, L.M.; Gutman, D.M.; Harrington, W.J., Jr.; Lee, K.P.; Boise, L.H. Proteasome inhibitors induce a terminal unfolded protein response in multiple myeloma cells. Blood 2006, 107, 4907–4916. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shen, J.; Fu, B.; Li, Y.; Wu, Y.; Sang, H.; Zhang, H.; Lin, H.; Liu, H.; Huang, W. E3 Ubiquitin Ligase-Mediated Regulation of Osteoblast Differentiation and Bone Formation. Front. Cell Dev. Biol. 2021, 9, 706395. [Google Scholar] [CrossRef] [PubMed]
- Flynt, E.; Bisht, K.; Sridharan, V.; Ortiz, M.; Towfic, F.; Thakurta, A. Prognosis, Biology, and Targeting of TP53 Dysregulation in Multiple Myeloma. Cells 2020, 9, 287. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sawamura, M.; Murakami, H.; Tsuchiya, J. Tumor necrosis factor-alpha and interleukin 4 in myeloma cell precursor differentiation. Leuk. Lymphoma 1996, 21, 31–36. [Google Scholar] [CrossRef]
- Jurisic, V.; Colovic, M. Correlation of sera TNF-alpha with percentage of bone marrow plasma cells, LDH, beta2-microglobulin, and clinical stage in multiple myeloma. Med. Oncol. 2002, 19, 133–139. [Google Scholar] [CrossRef]
- Hong, Y.; Yu, J.; Wang, G.; Qiao, W. Association between tumor necrosis factor alpha gene polymorphisms and multiple myeloma risk: An updated meta-analysis. Hematology 2019, 24, 216–224. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Terpos, E.; Christoulas, D.; Gavriatopoulou, M. Biology and treatment of myeloma related bone disease. Metabolism 2018, 80, 80–90. [Google Scholar] [CrossRef]
- Urashima, M.; Ogata, A.; Chauhan, D.; Hatziyanni, M.; Vidriales, M.B.; Dedera, D.A.; Schlossman, R.L.; Anderson, K.C. Transforming growth factor-beta1: Differential effects on multiple myeloma versus normal B cells. Blood 1996, 87, 1928–1938. [Google Scholar] [CrossRef] [Green Version]
- Ryoo, H.M.; Lee, M.H.; Kim, Y.J. Critical molecular switches involved in BMP-2-induced osteogenic differentiation of mesenchymal cells. Gene 2006, 366, 51–57. [Google Scholar] [CrossRef] [PubMed]
- Cottini, F.; Hideshima, T.; Xu, C.; Sattler, M.; Dori, M.; Agnelli, L.; ten Hacken, E.; Bertilaccio, M.T.; Antonini, E.; Neri, A.; et al. Rescue of Hippo coactivator YAP1 triggers DNA damage-induced apoptosis in hematological cancers. Nat. Med. 2014, 20, 599–606. [Google Scholar] [CrossRef] [Green Version]
- Rui, H.B.; Zheng, X.Q.; Lin, M.Y.; Yang, A.P. Sirtuin 6 promotes cell aging of myeloma cell line KM-HM_(31) by via Hippo signal pathway. Eur. Rev. Med. Pharmacol. Sci. 2018, 22, 6880–6884. [Google Scholar] [CrossRef]
- Pan, J.X.; Xiong, L.; Zhao, K.; Zeng, P.; Wang, B.; Tang, F.L.; Sun, D.; Guo, H.H.; Yang, X.; Cui, S.; et al. YAP promotes osteogenesis and suppresses adipogenic differentiation by regulating beta-catenin signaling. Bone Res. 2018, 6, 18. [Google Scholar] [CrossRef] [Green Version]
- Matsumoto, Y.; La Rose, J.; Kent, O.A.; Wagner, M.J.; Narimatsu, M.; Levy, A.D.; Omar, M.H.; Tong, J.; Krieger, J.R.; Riggs, E.; et al. Reciprocal stabilization of ABL and TAZ regulates osteoblastogenesis through transcription factor RUNX2. J. Clin. Investig. 2016, 126, 4482–4496. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, B.; Shi, M.; Li, J.; Zhang, H.; Chen, B.; Chen, L.; Gao, W.; Giuliani, N.; Zhao, R.C. Elevated tumor necrosis factor-alpha suppresses TAZ expression and impairs osteogenic potential of Flk-1+ mesenchymal stem cells in patients with multiple myeloma. Stem Cells Dev. 2007, 16, 921–930. [Google Scholar] [CrossRef]
- Eda, H.; Aoki, K.; Kato, S.; Okawa, Y.; Takada, K.; Tanaka, T.; Marumo, K.; Ohkawa, K. The proteasome inhibitor bortezomib inhibits FGF-2-induced reduction of TAZ levels in osteoblast-like cells. Eur. J. Haematol 2010, 85, 68–75. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.Y.; Li, Y.C.; Geng, C.Y.; Wang, H.J.; Chen, W.M. Potential Relationship between Clinical Significance and Serum Exosomal miRNAs in Patients with Multiple Myeloma. BioMed Res. Int. 2019, 2019, 1575468. [Google Scholar] [CrossRef] [PubMed]
- Standal, T.; Seidel, C.; Hjertner, O.; Plesner, T.; Sanderson, R.D.; Waage, A.; Borset, M.; Sundan, A. Osteoprotegerin is bound, internalized, and degraded by multiple myeloma cells. Blood 2002, 100, 3002–3007. [Google Scholar] [CrossRef]
- Piatopoulou, D.; Avgeris, M.; Marmarinos, A.; Xagorari, M.; Baka, M.; Doganis, D.; Kossiva, L.; Scorilas, A.; Gourgiotis, D. miR-125b predicts childhood acute lymphoblastic leukaemia poor response to BFM chemotherapy treatment. Br. J. Cancer 2017, 117, 801–812. [Google Scholar] [CrossRef] [PubMed]
- Tajadini, M.; Panjehpour, M.; Javanmard, S.H. Comparison of SYBR Green and TaqMan methods in quantitative real-time polymerase chain reaction analysis of four adenosine receptor subtypes. Adv. Biomed. Res. 2014, 3, 85. [Google Scholar] [CrossRef] [PubMed]
- Piatopoulou, D.; Avgeris, M.; Drakaki, I.; Marmarinos, A.; Xagorari, M.; Baka, M.; Pourtsidis, A.; Kossiva, L.; Gourgiotis, D.; Scorilas, A. Clinical utility of miR-143/miR-182 levels in prognosis and risk stratification specificity of BFM-treated childhood acute lymphoblastic leukemia. Ann. Hematol. 2018, 97, 1169–1182. [Google Scholar] [CrossRef]
- Schmittgen, T.D.; Livak, K.J. Analyzing real-time PCR data by the comparative C(T) method. Nat. Protoc. 2008, 3, 1101–1108. [Google Scholar] [CrossRef]
- Camp, R.L.; Dolled-Filhart, M.; Rimm, D.L. X-tile: A new bio-informatics tool for biomarker assessment and outcome-based cut-point optimization. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 2004, 10, 7252–7259. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vlachos, I.S.; Kostoulas, N.; Vergoulis, T.; Georgakilas, G.; Reczko, M.; Maragkakis, M.; Paraskevopoulou, M.D.; Prionidis, K.; Dalamagas, T.; Hatzigeorgiou, A.G. DIANA miRPath v.2.0: Investigating the combinatorial effect of microRNAs in pathways. Nucleic Acids Res. 2012, 40, W498–W504. [Google Scholar] [CrossRef] [PubMed]


| Variable | Mean ± S.E. 1 | Range | Percentiles | ||
|---|---|---|---|---|---|
| 25th | 50th (Median) | 75th | |||
| miR-16-5p levels (RQU 2) | |||||
| in sMM patients | 44.58 ± 14.04 | 0.004–167.3 | 2.94 | 10.16 | 82.20 |
| in MM patients | 13.77 ± 4.10 | 0.046–264.5 | 1.77 | 3.54 | 10.26 |
| miR-155-5p levels (RQU 2) | |||||
| in sMM patients | 1285.7± 539.8 | <0.001–6097.5 | 15.06 | 182.8 | 1402.7 |
| in MM patients | 366.0 ± 167.6 | <0.001–9738.0 | 2.21 | 23.05 | 130.3 |
| miR-15a-5p levels (RQU 2) | |||||
| in MM patients without osteolytic lesions | 29.93 ± 12.66 | 0.008–181.0 | 1.76 | 7.30 | 22,29 |
| in MM patients with osteolytic lesions | 7.71 ± 3.25 | 0.003–122.9 | 0.37 | 1.85 | 6.76 |
| miR-16-5p levels (RQU 2) | |||||
| in MM patients without osteolytic lesions | 19.27 ± 7.18 | 0.92–109.9 | 3.07 | 7.06 | 15.68 |
| in MM patients with osteolytic lesions | 7.49 ± 2.39 | 0.12–82.77 | 1.39 | 2.64 | 8.50 |
| miR-222-3p levels (RQU 2) | |||||
| in MM patients without osteolytic lesions | 10.95 ± 3.17 | 0.006–54.28 | 1.56 | 8.03 | 15.57 |
| in MM patients with osteolytic lesions | 6.03 ± 1.65 | 0.007–43.54 | 0.06 | 2.56 | 6.55 |
| miR-125b-5p levels (RQU 2) | |||||
| in MM patients without SREs 3 | 15.00 ± 8.12 | 0.001–139.8 | 0.17 | 1.17 | 15.62 |
| in MM patients with SREs 3 | 39.39 ± 12.72 | 0.12–247.7 | 5.96 | 26.28 | 44.75 |
| Univariate Analysis | Multivariate Analysis | |||||
|---|---|---|---|---|---|---|
| Covariate | HR 1 | BCa 4 Bootstrap 5 95% CI 2 | Bootstrap5 p Value 3 | HR 1 | BCa 4 Bootstrap 5 95% CI 2 | Bootstrap5 p Value 3 |
| miR-223-3p expression status | ||||||
| Positive | 1.00 | 1.00 | ||||
| Negative | 3.11 | 0.95–22.15 | 0.034 | 3.34 | 0.71–2.6 × 105 | 0.046 |
| R-ISS 6 (ordinal) | 3.31 | 1.05–13.22 | 0.025 | 3.14 | 1.05–21.17 | 0.021 |
| Variable | Number of MM Patients (%) |
|---|---|
| Gender | |
| Male | 44 (57.9%) |
| Female | 32 (42.1%) |
| Myeloma type | |
| IgG | 44 (57.9%) |
| IgA | 17 (22.4%) |
| IgD | 2 (2.6%) |
| Light chain | 10 (13.2%) |
| Non-secretory | 2 (2.6%) |
| Missing data | 1 (1.3%) |
| ISS 1 stage | |
| I | 15 (19.7%) |
| II | 25 (32.9%) |
| III | 34 (44.8%) |
| Missing data | 2 (2.6%) |
| R-ISS 2 stage | |
| I | 11 (14.5%) |
| II | 40 (52.6%) |
| III | 18 (23.7%) |
| Missing data | 7 (9.2%) |
| Bone disease | |
| No | 22 (28.9%) |
| Yes | 50 (65.8%) |
| Missing data | 4 (5.3%) |
| WBLDCT 3 osteolysis | |
| No | 18 (23.7%) |
| Yes | 38 (50.0%) |
| Missing data | 20 (26.3%) |
| SREs 4 (38 MM patients) | |
| No | 18 (47.4%) |
| Yes | 20 (52.6%) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Papanota, A.-M.; Karousi, P.; Kontos, C.K.; Artemaki, P.I.; Liacos, C.-I.; Papadimitriou, M.-A.; Bagratuni, T.; Eleutherakis-Papaiakovou, E.; Malandrakis, P.; Ntanasis-Stathopoulos, I.; et al. A Cancer-Related microRNA Signature Shows Biomarker Utility in Multiple Myeloma. Int. J. Mol. Sci. 2021, 22, 13144. https://doi.org/10.3390/ijms222313144
Papanota A-M, Karousi P, Kontos CK, Artemaki PI, Liacos C-I, Papadimitriou M-A, Bagratuni T, Eleutherakis-Papaiakovou E, Malandrakis P, Ntanasis-Stathopoulos I, et al. A Cancer-Related microRNA Signature Shows Biomarker Utility in Multiple Myeloma. International Journal of Molecular Sciences. 2021; 22(23):13144. https://doi.org/10.3390/ijms222313144
Chicago/Turabian StylePapanota, Aristea-Maria, Paraskevi Karousi, Christos K. Kontos, Pinelopi I. Artemaki, Christine-Ivy Liacos, Maria-Alexandra Papadimitriou, Tina Bagratuni, Evangelos Eleutherakis-Papaiakovou, Panagiotis Malandrakis, Ioannis Ntanasis-Stathopoulos, and et al. 2021. "A Cancer-Related microRNA Signature Shows Biomarker Utility in Multiple Myeloma" International Journal of Molecular Sciences 22, no. 23: 13144. https://doi.org/10.3390/ijms222313144
APA StylePapanota, A.-M., Karousi, P., Kontos, C. K., Artemaki, P. I., Liacos, C.-I., Papadimitriou, M.-A., Bagratuni, T., Eleutherakis-Papaiakovou, E., Malandrakis, P., Ntanasis-Stathopoulos, I., Gavriatopoulou, M., Kastritis, E., Avgeris, M., Dimopoulos, M.-A., Scorilas, A., & Terpos, E. (2021). A Cancer-Related microRNA Signature Shows Biomarker Utility in Multiple Myeloma. International Journal of Molecular Sciences, 22(23), 13144. https://doi.org/10.3390/ijms222313144













