Antimicrobial Resistance and Inorganic Nanoparticles
Abstract
1. Introduction
2. Methodology
3. Antibiotics
3.1. By Chemical Structure
3.2. By Effect
3.3. By Spectrum
3.4. By the Mechanism of Action
4. Antimicrobial Resistance
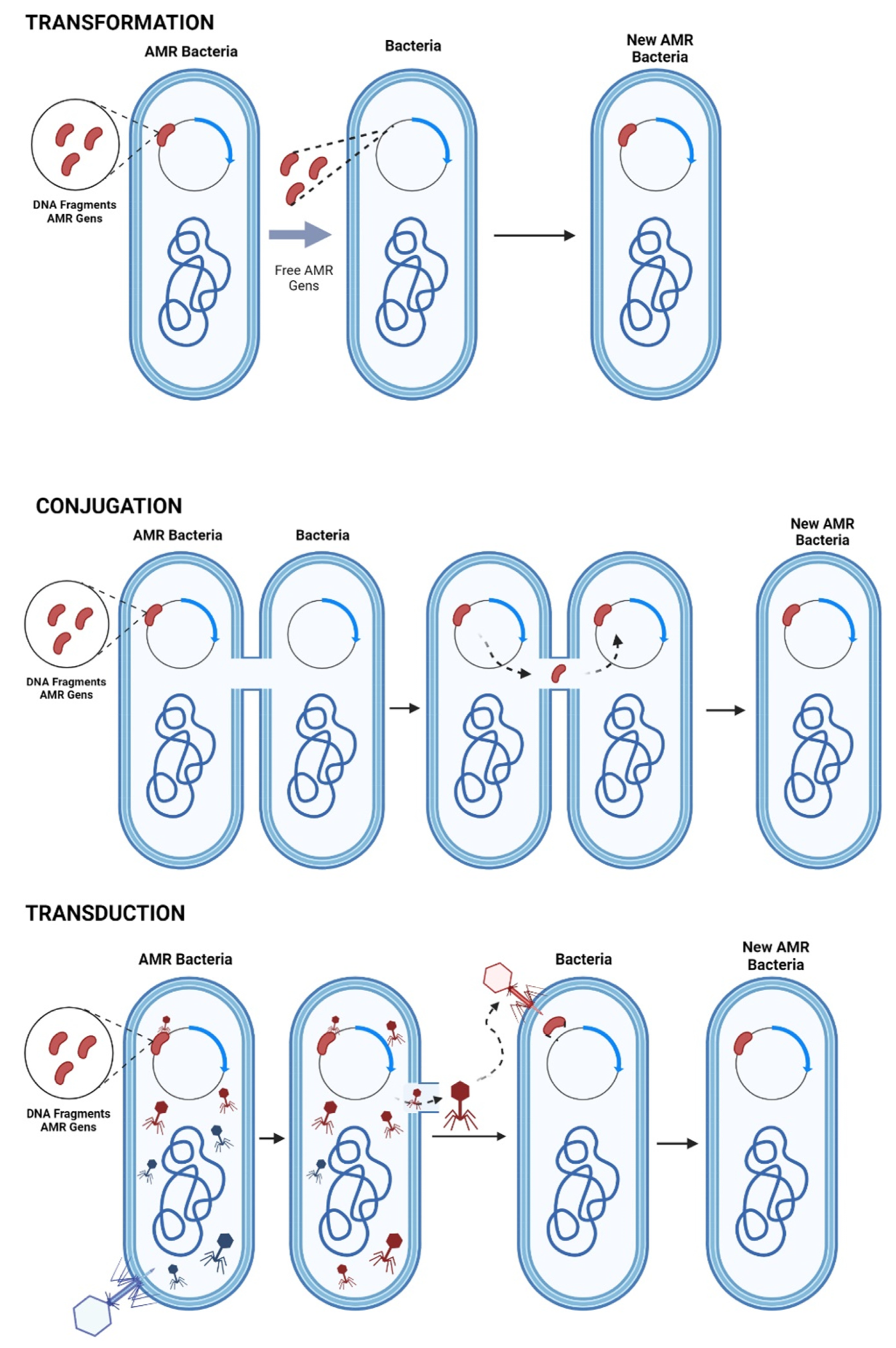
4.1. Mechanisms of AMR Gene Transfer
4.1.1. Chromosome Mutation
4.1.2. Extra Chromosomal Mutation
4.2. Biochemical Mechanisms of AMR
4.2.1. Inactivation of the Antibiotic
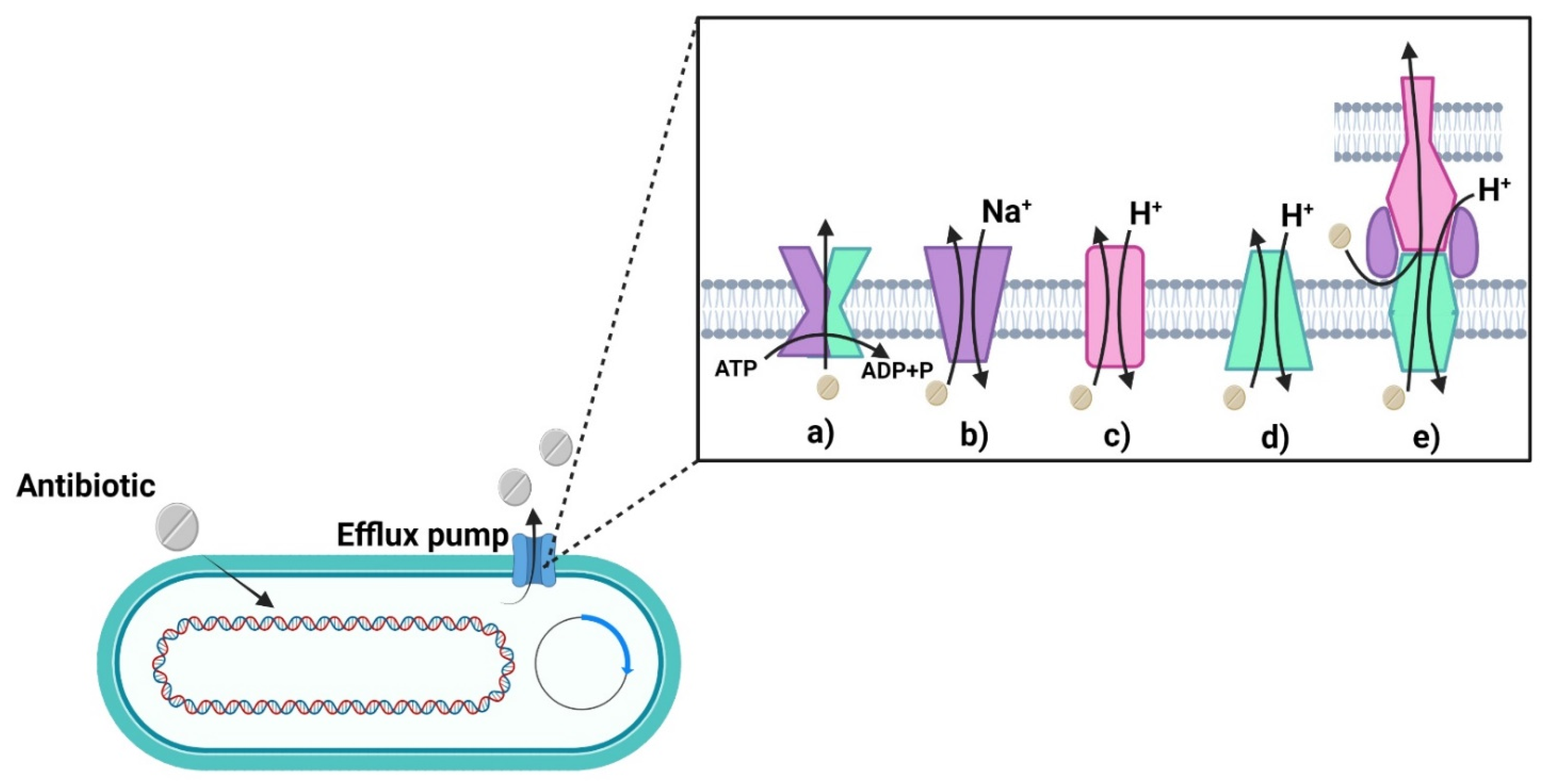
4.2.2. Antibiotic Excretion
4.2.3. Permeability of the Outer Membrane
4.2.4. Target Modification
5. Nanotechnology Applied to Antimicrobial Resistance
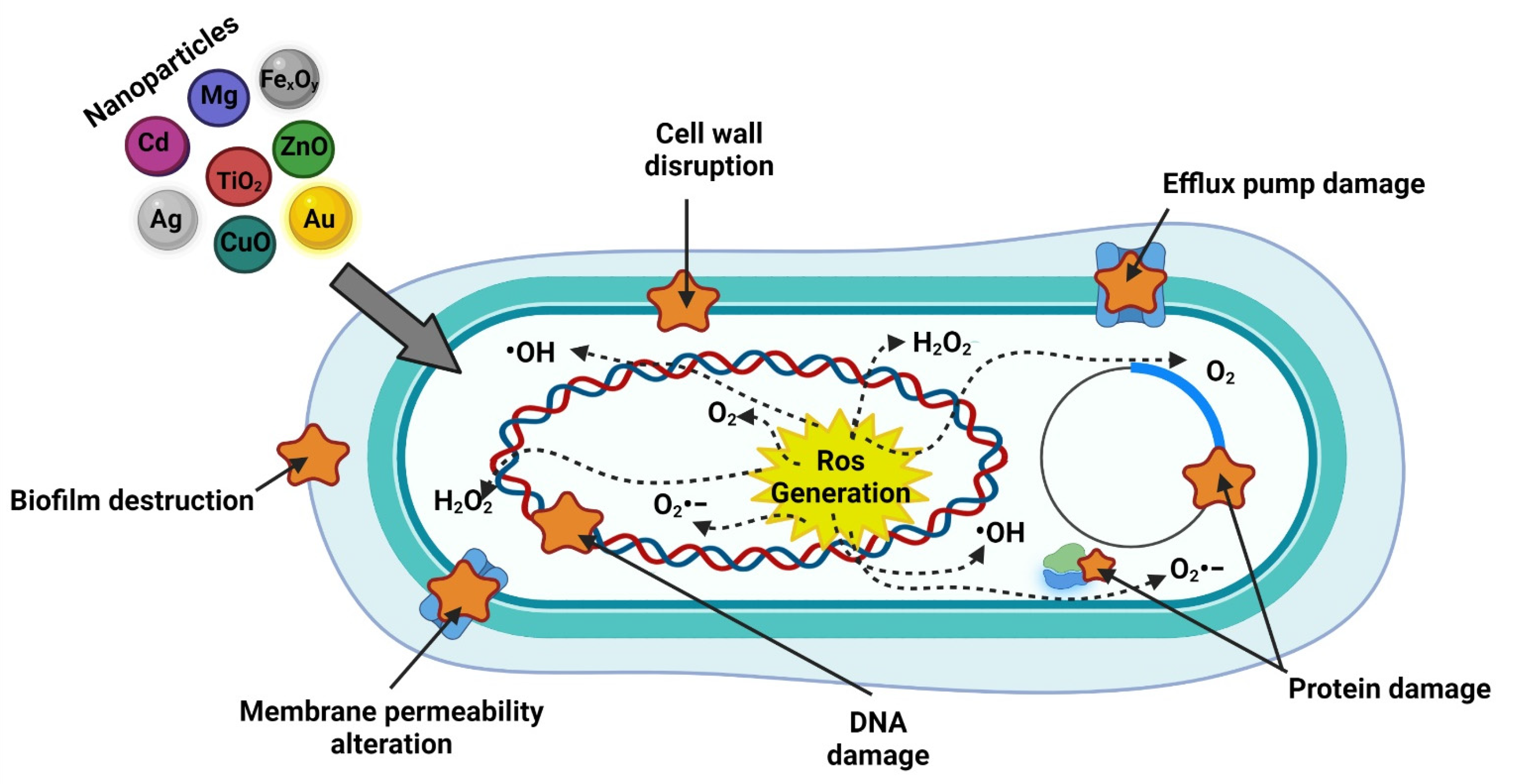
5.1. Nanoparticles Antimicrobial Effects
5.1.1. Interaction with the Cell Wall and Membrane
5.1.2. Generation of ROS
5.1.3. Penetration of the Cell Membrane
5.1.4. Inhibition of Protein Synthesis and DNA Damage
5.1.5. Damage to Metabolic Pathways
5.1.6. Biofilm Inhibition
5.2. Interaction between Antibiotics, Nanoparticles and Bacteria
5.2.1. Silver Nanoparticles
5.2.2. Zinc Oxide Nanoparticles
5.2.3. Gold Nanoparticles
5.2.4. TiO2 Nanoparticles
5.2.5. Other Nanoparticles
| Elemental Composition | Size and Morpho | Concentration (µg/mL) | Bacteria | Antibiotic | Inhibit | Mechanims | Author |
|---|---|---|---|---|---|---|---|
| AgNPs | 10 nm | 2.5 | P. aeruginosa | N/A | ~90% | AgNPs can enter cells and inhibits enzymatic systems in the respiratory chain, thereby altering their DNA synthesis | Salomoni et al. (2017) [4] |
| AgNPs | 35 ± 15 nm Spheroide | 0.35 0.5 0.05 8 | E. coli S. typhimurium S. aureus B. subtilis | Chloramphenicol | 50% | The combination of the AgNPs + antibiotic produced membrane damage | Vazquez-Muñoz et al. (2019) [237] |
| 0.05 0.1 16 0.12 | Kanamycin | 95% | |||||
| AgNPs | ~26 nm Spheras | 1 + 5 E 1 + 10 AMP 1 + 30 C 1 + 30 KF 1 + 2 DA 1 +30 TE 1 + 10 GEN 1 + 30 AMC 1 + 10 CFP 1 + 30 CXM | S. aureus MRSA E. coli P. aeruginosa A. actinomycetemcomitans | Erythromycin (E) Ampicillin (AMP) Chloramphenicol (C) Cephalothin (KF) Clindamycin (DA) Tetracycline (TE) Gentamycin (GEN) Amoxycillin (AMC) Cefpodoxime (CFP) Cefuroxime (CXM) | ~80% | ROS generation and mechanism of action of antibiotic | Ipe et al. (2020) [238] |
| AgNPs | 8–21 nm Spherical | 15.62 15.62 7.8 31.25 | S. epidermis S. haemolyticus | Ciprofloxacin Methicillin Gentamycin Rifampicin | 0.25 mm 0.06 mm 0.12 mm 1 mm | ROS generation and enhancement | Thomas et al. (2020) [239] |
| Mesoporous silica | 50–100 nm Spherical | 426 170 | A. baumannii | Cefepime Meropenem | 11 mm 11 mm | Antibiotic mechanims | Najafi et al. (2021) [240] |
| AuNPs | 33 ± 14 nm | 2/4 1/2 1/2 | E. coli S. aureus S. epidermis | Amoxicillin | 31 mm 30 mm 19 mm | The combination of antibiotic and NPs increase the concentration of antibiotic at the site of bacterium-antibiotic interaction; in additionthe multivalent presentation of amoxicillin blockade of the bacterial efflux pump | Kalita et al. (2016) [167] |
| AuNPs | 35 nm 200 nm | 0.72 0.73 | Klebsiella pneumaniae A. baumannii | Impinem Meropenem | 72 mm I/35 & 48 mm I/200 73 mm M/35 & 46 mm M/200 | The NPs improve the mechanism of action of antibiotic | Shaker et al. (2017) [241] |
| AuNPs | 8 ± 2 nm | 0.15 1.5 | S aureus MRSA | Amoxicillin | 85% | ROS generation by the antibiotic effect | Silvero et al. (2018) [242] |
| AuNPs | 30 ± 20 nm Irregular | 1.5 | P aeruginosa | Amoxicillin | 60–70% | Biofilm damage | Rocca et al. (2020) [243] |
| AuNPs | 5 nm | 1.18 0.23 | E. coli | Colistin | - 6.8% fold | N/A | Fuller et al. (2020) [244] |
| AuNPs | 25 nm | 62.5 | P. aeruginosa S. aureus | N/A | 19 mm 15.8 mm | AuNPs have a significant inhibitory effect on bacteria, to their ability to associate with the bacteria cell wall and rupture it, as well as disrupting bacterial metabolism by interfering with bacterial DNA | Abdulazeem et al. (2021) [245] |
| TiO2NPs | 64 ± 0.14 nm Irregular spheres | 8–64 | P. aeruginosa | Ceftriaxone Amikacin Ciprofloxacin Cefepime | 96% 88% 80% 100% | The antibiotic in combination with the nanostructure increases the synergistic effect of an antibiotic as can inhibit the cell | Youssef et al. (2020) [246] |
6. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Econom, D.E. NORMA MEXICANA NMX-R-80004-1-SCFI-2013 Nanotecnologías—Vocabulario—Parte 1: Conceptos Básicos Nanotechnologies—Vocabulary—Part 1: Core Terms, Dirección General de Normas (DGN); Dirección General de Normas (DGN): Cuauhtémoc, México, 2013. [Google Scholar]
- Bayda, S.; Adeel, M.; Tuccinardi, T.; Cordani, M.; Rizzolio, F.; Baeza, A. The History of Nanoscience and Nanotechnology: From Chemical-Physical Applications to Nanomedicine. Molecules 2020, 25, 112. [Google Scholar] [CrossRef] [PubMed]
- Kandi, V.; Kandi, S. Antimicrobial properties of nanomolecules: Potential candidates as antibiotics in the era of multi-drug resistance. Epidemiol. Health 2015, 37, e2015020. [Google Scholar] [CrossRef] [PubMed]
- Salomoni, R.; Léo, P.; Montemor, A.; Rinaldi, B.; Rodrigues, M. Antibacterial effect of silver nanoparticles in Pseudomonas aeruginosa. Nanotechnol. Sci. Appl. 2017, 10, 115–121. [Google Scholar] [CrossRef]
- NNI. Nanotechnology. Available online: www.nano.gov (accessed on 15 March 2021).
- Yougbare, S.; Chang, T.-K.; Tan, S.-H.; Kuo, J.-C.; Hsu, P.-H.; Su, C.-Y.; Kuo, T.-R. Antimicrobial Gold Nanoclusters: Recent Developments and Future Perspectives. Int. J. Mol. Sci. 2019, 20, 2924. [Google Scholar] [CrossRef]
- Contera, S.; Bernardino de la Serna, J.; Tetley, T.D. Biotechnology, nanotechnology and medicine. Emerg. Top. Life Sci. 2020, 4, 551–554. [Google Scholar] [CrossRef]
- Allahverdiyev, A.M.; Kon, K.V.; Abamor, E.S.; Bagirova, M.; Rafailovich, M. Coping with antibiotic resistance: Combining nanoparticles with antibiotics and other antimicrobial agents. Expert Rev. Anti. Infect. Ther. 2011, 9, 1035–1052. [Google Scholar] [CrossRef] [PubMed]
- Roy, J.; Chandra, S.; Maitra, S. Nanotechnology in castable refractory. Ceram. Int. 2019, 45, 19–29. [Google Scholar] [CrossRef]
- Peigneux, A.; Oltolina, F.; Colangelo, D.; Iglesias, G.R.; Delgado, A.V.; Prat, M.; Jimenez-Lopez, C. Functionalized Biomimetic Magnetic Nanoparticles as Effective Nanocarriers for Targeted Chemotherapy. Part. Part. Syst. Charact. 2019, 36, 1900057. [Google Scholar] [CrossRef]
- Kuo, Y.-L.; Wang, S.-G.; Wu, C.-Y.; Lee, K.-C.; Jao, C.-J.; Chou, S.-H.; Chen, Y.-C. Functional gold nanoparticle-based antibacterial agents for nosocomial and antibiotic-resistant bacteria. Nanomedicine 2016, 11, 2497–2510. [Google Scholar] [CrossRef] [PubMed]
- Kumari, A.; Singla, R.; Guliani, A.; Yadav, S.K. Nanoencapsulation for drug delivery. EXCLI J. 2014, 13, 265–286. [Google Scholar]
- Menon, S.; Rajeshkumar, S.; Kumar, V. A review on biogenic synthesis of gold nanoparticles, characterization, and its applications. Resour. Technol. 2017, 3, 516–527. [Google Scholar] [CrossRef]
- Kim, B.Y.S.; Rutka, J.T.; Chan, W.C.W. Nanomedicine. N. Engl. J. Med. 2010, 363, 2434–2443. [Google Scholar] [CrossRef] [PubMed]
- Blecher, K.; Nasir, A.; Friedman, A. The growing role of nanotechnology in combating infectious disease. Virulence 2011, 2, 395–401. [Google Scholar] [CrossRef] [PubMed]
- Ratner, B.D. Biomaterials: Been There, Done That, and Evolving into the Future. Annu. Rev. Biomed. Eng. 2019, 21, 171–191. [Google Scholar] [CrossRef] [PubMed]
- OMS. Global Priority List of Antibiotic-Resistant Bacteria to Guide Research, Discovery, and Development of New Antibiotics. Available online: https://www.who.int/medicines/publications/WHO-PPL-Short_Summary_25Feb-ET_NM_WHO.pdf?ua=1 (accessed on 25 February 2020).
- Li, B.; Webster, T.J. Bacteria antibiotic resistance: New challenges and opportunities for implant-associated orthopedic infections. J. Orthop. Res. 2017, 36, 22–32. [Google Scholar] [CrossRef]
- Kumar, L.; Verma, S.; Vaidya, B.; Mehra, N.K. Nanocarrier-Assisted Antimicrobial Therapy against Intracellular Pathogens; Chapter 13; Elsevier: Amsterdam, The Netherlands, 2017; pp. 293–324. ISBN 9780323461528. [Google Scholar] [CrossRef]
- Hutchings, M.I.; Truman, A.W.; Wilkinson, B. Antibiotics: Past, present and future. Curr. Opin. Microbiol. 2019, 51, 72–80. [Google Scholar] [CrossRef] [PubMed]
- Laxminarayan, R.; Duse, A.; Wattal, C.; Zaidi, A.K.M.; Wertheim, H.F.L.; Sumpradit, N.; Vlieghe, E.; Hara, G.L.; Gould, I.M.; Goossens, H.; et al. Antibiotic resistance-the need for global solutions. Lancet Infect. Dis. 2013, 13, 1057–1098. [Google Scholar] [CrossRef]
- Feng, Q.; Huang, Y.; Chen, M.; Li, G.; Chen, Y. Functional synergy of α-helical antimicrobial peptides and traditional antibiotics against Gram-negative and Gram-positive bacteria in vitro and in vivo. Eur. J. Clin. Microbiol. Infect. Dis. 2015, 34, 197–204. [Google Scholar] [CrossRef]
- CDC. Antimicrobial Resistance. Available online: https://www.cdc.gov/drugresistance/biggest-threats.html?CDC_AA_refVal=https%3A%2F%2Fwww.cdc.gov%2Fdrugresistance%2Fbiggest_threats.html (accessed on 15 March 2021).
- CDC. Antibiotic Resistance Threats in the United States; U.S. Centers for Disease Control and Prevention: Atlanta, GA, USA, 2019.
- OCDE. Stemming the Superbug Tide; OECD Health Policy Studies; OECD: Paris, France, 2018; ISBN 9789264307582. [Google Scholar]
- Eyler, R.F.; Shvets, K. Clinical Pharmacology of Antibiotics. Clin. J. Am. Soc. Nephrol. 2019, 14, 1080–1090. [Google Scholar] [CrossRef]
- Brunton, L.; Hilal-Dandan, R.; Knollmann BCMacDougall, C. Penicillins, cephalosporins and other β-lactam antibiotics. In Goodman & Gilman’s: The Pharmacological Basis of Therapeutics; Brunton, L., Hilal-Dandan, R., BC, K., Eds.; McGraw-Hill Education: New York, NY, USA, 2017; pp. 1023–1038. [Google Scholar]
- Lodise, T.P.; Lomaestro, B.; Drusano, G.L. Piperacillin-Tazobactam for Pseudomonas aeruginosa Infection: Clinical Implications of an Extended-Infusion Dosing Strategy. Clin. Infect. Dis. 2007, 44, 357–363. [Google Scholar] [CrossRef]
- Koomanachai, P.; Bulik, C.C.; Kuti, J.L.; Nicolau, D.P. Pharmacodynamic modeling of intravenous antibiotics against gram-negative bacteria collected in the United States. Clin. Ther. 2010, 32, 766–779. [Google Scholar] [CrossRef]
- Lingzhi, L.; Haojie, G.; Dan, G.; Hongmei, M.; Yang, L.; Mengdie, J.; Chengkun, Z.; Xiaohui, Z. The role of two-component regulatory system in β-lactam antibiotics resistance. Microbiol. Res. 2018, 215, 126–129. [Google Scholar] [CrossRef] [PubMed]
- Werth, B.J. Overview of Antibiotics. Available online: https://www.msdmanuals.com/home/infections/antibiotics/overview-of-antibiotics (accessed on 3 March 2020).
- Murray, P.R.; Rosenthal, K.S.; Pfaller, M.A. Bacteriology. In Medical Microbiology; Saunders, E., Ed.; Elsevier: Amsterdam, The Netherlands, 2020; p. 872. ISBN 9780323673228. [Google Scholar]
- Brown, E.D.; Wright, G.D. Antibacterial drug discovery in the resistance era. Nature 2016, 529, 336–343. [Google Scholar] [CrossRef] [PubMed]
- D’Costa, V.M.; King, C.E.; Kalan, L.; Morar, M.; Sung, W.W.L.; Schwarz, C.; Froese, D.; Zazula, G.; Calmels, F.; Debruyne, R.; et al. Antibiotic resistance is ancient. Nature 2011, 477, 457–461. [Google Scholar] [CrossRef] [PubMed]
- Banin, E.; Hughes, D.; Kuipers, O.P. Editorial: Bacterial pathogens, antibiotics and antibiotic resistance. FEMS Microbiol. Rev. 2017, 41, 450–452. [Google Scholar] [CrossRef]
- Luepke, K.H.; Suda, K.J.; Boucher, H.; Russo, R.L.; Bonney, M.W.; Hunt, T.D.; Mohr, J.F. Past, Present, and Future of Antibacterial Economics: Increasing Bacterial Resistance, Limited Antibiotic Pipeline, and Societal Implications. Pharmacother. J. Hum. Pharmacol. Drug Ther. 2017, 37, 71–84. [Google Scholar] [CrossRef] [PubMed]
- Chung, P.Y.; Khanum, R. Antimicrobial peptides as potential anti-biofilm agents against multidrug-resistant bacteria. J. Microbiol. Immunol. Infect. 2017, 50, 405–410. [Google Scholar] [CrossRef]
- Cárdenas, L.L.; Merchán, M.A.; López, D.P. New antibiotics against bacterial resistance. Infectio 2019, 23, 382. [Google Scholar] [CrossRef]
- MacFadden, D.R.; McGough, S.F.; Fisman, D.; Santillana, M.; Brownstein, J.S. Antibiotic resistance increases with local temperature. Nat. Clim. Chang. 2018, 8, 510–514. [Google Scholar] [CrossRef]
- Imran, M.; Das, K.R.; Naik, M.M. Co-selection of multi-antibiotic resistance in bacterial pathogens in metal and microplastic contaminated environments: An emerging health threat. Chemosphere 2019, 215, 846–857. [Google Scholar] [CrossRef]
- Aslam, B.; Wang, W.; Arshad, M.I.; Khurshid, M.; Muzammil, S.; Rasool, M.H.; Nisar, M.A.; Alvi, R.F.; Aslam, M.A.; Qamar, M.U.; et al. Antibiotic resistance: A rundown of a global crisis. Infect. Drug Resist. 2018, 11, 1645–1658. [Google Scholar] [CrossRef]
- Frieri, M.; Kumar, K.; Boutin, A. Antibiotic resistance. J. Infect. Public Health 2017, 10, 369–378. [Google Scholar] [CrossRef] [PubMed]
- Troncoso, C.; Pavez, M.; Santos, A.; Salazar, R.; Barrientos, L. Implicancias Estructurales y Fisiológicas de la Célula Bacteriana en los Mecanismos de Resistencia Antibiótica. Int. J. Morphol. 2017, 35, 1214–1223. [Google Scholar] [CrossRef]
- Bullington, W.; Hempstead, S.; Smyth, A.R.; Drevinek, P.; Saiman, L.; Waters, V.J.; Bell, S.C.; VanDevanter, D.R.; Flume, P.A.; Elborn, S.; et al. Antimicrobial resistance: Concerns of healthcare providers and people with CF. J. Cyst. Fibros. 2020, 20, 407–412. [Google Scholar] [CrossRef]
- Płusa, T.; Konieczny, R.; Baranowska, A.; Szymczak, Z. The growing resistance of bacterial strains to antibiotics. Pol. Merkur. Lek. 2019, 47, 106–110. [Google Scholar]
- Mobarki, N.; Almerabi, B.; Hattan, A. Antibiotic Resistance Crisis. Int. J. Med. Dev. Ctries 2019, 561–564. [Google Scholar] [CrossRef]
- Reygaert, W.C. An overview of the antimicrobial resistance mechanisms of bacteria. AIMS Microbiol. 2018, 4, 482–501. [Google Scholar] [CrossRef] [PubMed]
- Mandsberg, L.F.; Ciofu, O.; Kirkby, N.; Christiansen, L.E.; Poulsen, H.E.; Høiby, N. Antibiotic Resistance in Pseudomonas aeruginosa Strains with Increased Mutation Frequency Due to Inactivation of the DNA Oxidative Repair System. Antimicrob. Agents Chemother. 2009, 53, 2483–2491. [Google Scholar] [CrossRef] [PubMed]
- Kapoor, G.; Saigal, S.; Elongavan, A. Action and resistance mechanisms of antibiotics: A guide for clinicians. J. Anaesthesiol. Clin. Pharmacol. 2017, 33, 300. [Google Scholar] [CrossRef]
- Ruddaraju, L.K.; Pammi, S.V.N.; Guntuku, G.S.; Padavala, V.S.; Kolapalli, V.R.M. A review on anti-bacterials to combat resistance: From ancient era of plants and metals to present and future perspectives of green nano technological combinations. Asian J. Pharm. Sci. 2020, 15, 42–59. [Google Scholar] [CrossRef]
- Nas, F. Mechanisms of Bacterial Antibiotics Resistance: A Review. J. Adv. Microbiol. 2017, 7, 1–6. [Google Scholar] [CrossRef]
- Partridge, S.R.; Kwong, S.M.; Firth, N.; Jensen, S.O. Mobile Genetic Elements Associated with Antimicrobial Resistance. Clin. Microbiol. Rev. 2018, 31, e00088-17. [Google Scholar] [CrossRef]
- Majeed, H.T.; Aljanaby, A.A.J. Antibiotic Susceptibility Patterns and Prevalence of Some Extended Spectrum Beta-Lactamases Genes in Gram-Negative Bacteria Isolated from Patients Infected with Urinary Tract Infections in Al-Najaf City, Iraq. Avicenna J. Med. Biotechnol. 2019, 11, 192–201. [Google Scholar]
- Bassetti, M.; Vena, A.; Croxatto, A.; Righi, E.; Guery, B. How to manage Pseudomonas aeruginosa infections. Drugs Context 2018, 7, 1–18. [Google Scholar] [CrossRef]
- Henrichfreise, B.; Wiegand, I.; Pfister, W.; Wiedemann, B. Resistance Mechanisms of Multiresistant Pseudomonas aeruginosa Strains from Germany and Correlation with Hypermutation. Antimicrob. Agents Chemother. 2007, 51, 4062–4070. [Google Scholar] [CrossRef]
- Blair, J.M.A.; Webber, M.A.; Baylay, A.J.; Ogbolu, D.O.; Piddock, L.J.V. Molecular mechanisms of antibiotic resistance. Nat. Rev. Microbiol. 2015, 13, 42–51. [Google Scholar] [CrossRef] [PubMed]
- Arzanlou, M.; Chai, W.C.; Venter, H. Intrinsic, adaptive and acquired antimicrobial resistance in Gram-negative bacteria. Essays Biochem. 2017, 61, 49–59. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Mathieu, J.M.; Chattopadhyay, S.; Miller, J.T.; Wu, T.; Shibata, T.; Guo, W.; Alvarez, P.J.J. Defense Mechanisms of Pseudomonas aeruginosa PAO1 against Quantum Dots and Their Released Heavy Metals. ACS Nano 2012, 6, 6091–6098. [Google Scholar] [CrossRef]
- Hancock, R.E.W.; Brinkman, F.S.L. Function of Pseudomonas Porins in Uptake and Efflux. Annu. Rev. Microbiol. 2002, 56, 17–38. [Google Scholar] [CrossRef]
- Bradford, P.A. Epidemiology of Bacterial Resistance. In Antimicrobial Resistance in the 21st Century; Springer International Publishing: Cham, Switzerland, 2018; pp. 299–339. ISBN 978-3-319-78538-7. [Google Scholar]
- Bockstael, K.; Aerschot, A. Antimicrobial resistance in bacteria. Open Med. 2009, 4, 141–155. [Google Scholar] [CrossRef]
- Hasan Abusaiba, T.; AL-Harmoosh, R. Mechanisms of Antibiotics Resistance in Bacteria. Syst. Rev. Pharm. 2020, 11, 817–823. [Google Scholar] [CrossRef]
- Begum, S.; Begum, T.; Rahman, N.; Khan, R.A. A review on antibiotic resistance and way of combating antimicrobial resistance. GSC Biol. Pharm. Sci. 2021, 14, 087–097. [Google Scholar] [CrossRef]
- Orellana, J.; Contreras, R. Bacterial Resistance Algorithm. An Application to CVRP. In International Work-Conference on the Interplay between Natural and Artificial Computation; Springer: Cham, Switzerland, 2019; Volume 11487, pp. 204–211. [Google Scholar] [CrossRef]
- Stevenson, C.; Hall, J.P.; Harrison, E.; Wood, A.; Brockhurst, M.A. Gene mobility promotes the spread of resistance in bacterial populations. ISME J. 2017, 11, 1930–1932. [Google Scholar] [CrossRef] [PubMed]
- Gyles, C.; Boerlin, P. Horizontally Transferred Genetic Elements and Their Role in Pathogenesis of Bacterial Disease. Vet. Pathol. 2014, 51, 328–340. [Google Scholar] [CrossRef]
- Wang, X.; Yang, F.; Zhao, J.; Xu, Y.; Mao, D.; Zhu, X.; Luo, Y.; Alvarez, P.J.J. Bacterial exposure to ZnO nanoparticles facilitates horizontal transfer of antibiotic resistance genes. NanoImpact 2018, 10, 61–67. [Google Scholar] [CrossRef]
- Dodd, M.C. Potential impacts of disinfection processes on elimination and deactivation of antibiotic resistance genes during water and wastewater treatment. J. Environ. Monit. 2012, 14, 1754. [Google Scholar] [CrossRef]
- NHGRI. Available online: https://www.genome.gov/genetics-glossary/Plasmid (accessed on 10 April 2021).
- Julin, D. Plasmid Cloning Vectors. In Molecular Life Sciences; Springer: New York, NY, USA, 2014; pp. 1–12. [Google Scholar]
- Santos López, A. Importancia de los Plásmidos ColE1 en la Resistencia a Antibióticos. Ph.D. Thesis, Universidad Complutense de Madrid, Madrid, Spain, 2017. [Google Scholar]
- Thomas, C.M.; Frost, L.S. Plasmid Genomes, Introduction to. In Molecular Life Sciences; Springer: New York, NY, USA, 2014; pp. 1–20. [Google Scholar]
- Redondo-Salvo, S.; Fernández-López, R.; Ruiz, R.; Vielva, L.; de Toro, M.; Rocha, E.P.C.; Garcillán-Barcia, M.P.; de la Cruz, F. Pathways for horizontal gene transfer in bacteria revealed by a global map of their plasmids. Nat. Commun. 2020, 11, 3602. [Google Scholar] [CrossRef]
- Bennett, P.M. Plasmid encoded antibiotic resistance: Acquisition and transfer of antibiotic resistance genes in bacteria. Br. J. Pharmacol. 2008, 153, S347–S357. [Google Scholar] [CrossRef]
- Cain, A.K.; Barquist, L.; Goodman, A.L.; Paulsen, I.T.; Parkhill, J.; van Opijnen, T. A decade of advances in transposon-insertion sequencing. Nat. Rev. Genet. 2020, 21, 526–540. [Google Scholar] [CrossRef] [PubMed]
- Asif, M.; Alvi, I.A.; Rehman, S.U. Insight into Acinetobacter baumannii: Pathogenesis, global resistance, mechanisms of resistance, treatment options, and alternative modalities. Infect. Drug Resist. 2018, 11, 1249–1260. [Google Scholar] [CrossRef]
- Fan, C.; Wu, Y.-H.; Decker, C.M.; Rohani, R.; Gesell Salazar, M.; Ye, H.; Cui, Z.; Schmidt, F.; Huang, W.E. Defensive Function of Transposable Elements in Bacteria. ACS Synth. Biol. 2019, 8, 2141–2151. [Google Scholar] [CrossRef]
- Pournajaf, A.; Razavi, S.; Irajian, G.; Ardebili, A.; Erfani, Y.; Solgi, S.; Yaghoubi, S.; Rasaeian, A.; Yahyapour, Y.; Kafshgari, R.; et al. Integron types, antimicrobial resistance genes, virulence gene profile, alginate production and biofilm formation in Iranian cystic fibrosis Pseudomonas aeruginosa isolates. Infez. Med. 2018, 26, 226–236. [Google Scholar] [PubMed]
- Barlow, R.S.; Gobius, K.S. Diverse class 2 integrons in bacteria from beef cattle sources. J. Antimicrob. Chemother. 2006, 58, 1133–1138. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; Lan, F.; Lu, Y.; Li, B. Characterization of Integrons and Antimicrobial Resistance in Escherichia coli Sequence Type 131 Isolates. Can. J. Infect. Dis. Med. Microbiol. 2020, 2020, 3826186. [Google Scholar] [CrossRef] [PubMed]
- Kaushik, M.; Kumar, S.; Kapoor, R.K.; Virdi, J.S.; Gulati, P. Integrons in Enterobacteriaceae: Diversity, distribution and epidemiology. Int. J. Antimicrob. Agents 2018, 51, 167–176. [Google Scholar] [CrossRef]
- Khosravi, A.D.; Motahar, M.; Abbasi Montazeri, E. The frequency of class1 and 2 integrons in Pseudomonas aeruginosa strains isolated from burn patients in a burn center of Ahvaz, Iran. PLoS ONE 2017, 12, e0183061. [Google Scholar] [CrossRef]
- Zhao, X.; Hu, M.; Zhang, Q.; Zhao, C.; Zhang, Y.; Li, L.; Qi, J.; Luo, Y.; Zhou, D.; Liu, Y. Characterization of integrons and antimicrobial resistance in Salmonella from broilers in Shandong, China. Poult. Sci. 2020, 99, 7046–7054. [Google Scholar] [CrossRef]
- Bello-López, J.M.; Cabrero-Martínez, O.A.; Ibáñez-Cervantes, G.; Hernández-Cortez, C.; Pelcastre-Rodríguez, L.I.; Gonzalez-Avila, L.U.; Castro-Escarpulli, G. Horizontal Gene Transfer and Its Association with Antibiotic Resistance in the Genus Aeromonas spp. Microorganisms 2019, 7, 363. [Google Scholar] [CrossRef]
- Lerminiaux, N.A.; Cameron, A.D.S. Horizontal transfer of antibiotic resistance genes in clinical environments. Can. J. Microbiol. 2019, 65, 34–44. [Google Scholar] [CrossRef]
- Schneider, C.L. Bacteriophage-Mediated Horizontal Gene Transfer: Transduction. In Bacteriophages; Springer International Publishing: Cham, Switzerland, 2021; pp. 151–192. [Google Scholar]
- Melander, R.J.; Melander, C. The Challenge of Overcoming Antibiotic Resistance: An Adjuvant Approach? ACS Infect. Dis. 2017, 3, 559–563. [Google Scholar] [CrossRef]
- Baker, S.; Thomson, N.; Weill, F.-X.; Holt, K.E. Genomic insights into the emergence and spread of antimicrobial-resistant bacterial pathogens. Science 2018, 360, 733–738. [Google Scholar] [CrossRef]
- Holmes, A.H.; Moore, L.S.P.; Sundsfjord, A.; Steinbakk, M.; Regmi, S.; Karkey, A.; Guerin, P.J.; Piddock, L.J. V Understanding the mechanisms and drivers of antimicrobial resistance. Lancet 2016, 387, 176–187. [Google Scholar] [CrossRef]
- Blair, J.M.; Richmond, G.E.; Piddock, L.J. Multidrug efflux pumps in Gram-negative bacteria and their role in antibiotic resistance. Future Microbiol. 2014, 9, 1165–1177. [Google Scholar] [CrossRef] [PubMed]
- Zgurskaya, H.I.; Rybenkov, V.V. Permeability barriers of Gram-negative pathogens. Ann. N. Y. Acad. Sci. 2020, 1459, 5–18. [Google Scholar] [CrossRef]
- Hernando-Amado, S.; Blanco, P.; Alcalde-Rico, M.; Corona, F.; Reales-Calderón, J.A.; Sánchez, M.B.; Martínez, J.L. Multidrug efflux pumps as main players in intrinsic and acquired resistance to antimicrobials. Drug Resist. Updat. 2016, 28, 13–27. [Google Scholar] [CrossRef] [PubMed]
- Li, N.; Wu, J.-X.; Ding, D.; Cheng, J.; Gao, N.; Chen, L. Structure of a Pancreatic ATP-Sensitive Potassium Channel. Cell 2017, 168, 101–110.e10. [Google Scholar] [CrossRef]
- Johnson, Z.L.; Chen, J. Structural Basis of Substrate Recognition by the Multidrug Resistance Protein MRP1. Cell 2017, 168, 1075–1085.e9. [Google Scholar] [CrossRef]
- Verhalen, B.; Dastvan, R.; Thangapandian, S.; Peskova, Y.; Koteiche, H.A.; Nakamoto, R.K.; Tajkhorshid, E.; Mchaourab, H.S. Energy transduction and alternating access of the mammalian ABC transporter P-glycoprotein. Nature 2017, 543, 738–741. [Google Scholar] [CrossRef]
- Mousa, J.J.; Yang, Y.; Tomkovich, S.; Shima, A.; Newsome, R.C.; Tripathi, P.; Oswald, E.; Bruner, S.D.; Jobin, C. MATE transport of the E. coli-derived genotoxin colibactin. Nat. Microbiol. 2016, 1, 15009. [Google Scholar] [CrossRef] [PubMed]
- Radchenko, M.; Symersky, J.; Nie, R.; Lu, M. Structural basis for the blockade of MATE multidrug efflux pumps. Nat. Commun. 2015, 6, 7995. [Google Scholar] [CrossRef]
- Radchenko, M.; Nie, R.; Lu, M. Disulfide Cross-linking of a Multidrug and Toxic Compound Extrusion Transporter Impacts Multidrug Efflux. J. Biol. Chem. 2016, 291, 9818–9826. [Google Scholar] [CrossRef]
- Zwama, M.; Yamasaki, S.; Nakashima, R.; Sakurai, K.; Nishino, K.; Yamaguchi, A. Multiple entry pathways within the efflux transporter AcrB contribute to multidrug recognition. Nat. Commun. 2018, 9, 124. [Google Scholar] [CrossRef]
- Sun, J.; Deng, Z.; Yan, A. Bacterial multidrug efflux pumps: Mechanisms, physiology and pharmacological exploitations. Biochem. Biophys. Res. Commun. 2014, 453, 254–267. [Google Scholar] [CrossRef]
- Radestock, S.; Forrest, L.R. The Alternating-Access Mechanism of MFS Transporters Arises from Inverted-Topology Repeats. J. Mol. Biol. 2011, 407, 698–715. [Google Scholar] [CrossRef] [PubMed]
- Kaback, H.R. A chemiosmotic mechanism of symport. Proc. Natl. Acad. Sci. USA 2015, 112, 1259–1264. [Google Scholar] [CrossRef]
- Heng, J.; Zhao, Y.; Liu, M.; Liu, Y.; Fan, J.; Wang, X.; Zhao, Y.; Zhang, X.C. Substrate-bound structure of the E. coli multidrug resistance transporter MdfA. Cell Res. 2015, 25, 1060–1073. [Google Scholar] [CrossRef]
- Su, C.-C.; Yin, L.; Kumar, N.; Dai, L.; Radhakrishnan, A.; Bolla, J.R.; Lei, H.-T.; Chou, T.-H.; Delmar, J.A.; Rajashankar, K.R.; et al. Structures and transport dynamics of a Campylobacter jejuni multidrug efflux pump. Nat. Commun. 2017, 8, 171. [Google Scholar] [CrossRef]
- Poonsuk, K.; Tribuddharat, C.; Chuanchuen, R. Simultaneous overexpression of multidrug efflux pumps in Pseudomonas aeruginosa non-cystic fibrosis clinical isolates. Can. J. Microbiol. 2014, 60, 437–443. [Google Scholar] [CrossRef] [PubMed]
- Saxena, P.; Joshi, Y.; Rawat, K.; Bisht, R. Biofilms: Architecture, Resistance, Quorum Sensing and Control Mechanisms. Indian J. Microbiol. 2019, 59, 3–12. [Google Scholar] [CrossRef]
- Felden, B.; Cattoir, V. Bacterial Adaptation to Antibiotics through Regulatory RNAs. Antimicrob. Agents Chemother. 2018, 62, e02503-17. [Google Scholar] [CrossRef] [PubMed]
- Hall, C.W.; Mah, T.-F. Molecular mechanisms of biofilm-based antibiotic resistance and tolerance in pathogenic bacteria. FEMS Microbiol. Rev. 2017, 41, 276–301. [Google Scholar] [CrossRef]
- Gou, N.; Onnis-Hayden, A.; Gu, A.Z. Mechanistic Toxicity Assessment of Nanomaterials by Whole-Cell-Array Stress Genes Expression Analysis. Environ. Sci. Technol. 2010, 44, 5964–5970. [Google Scholar] [CrossRef]
- Nevius, B.A.; Chen, Y.P.; Ferry, J.L.; Decho, A.W. Surface-functionalization effects on uptake of fluorescent polystyrene nanoparticles by model biofilms. Ecotoxicology 2012, 21, 2205–2213. [Google Scholar] [CrossRef]
- Tkachenko, A.G. Stress Responses of Bacterial Cells as Mechanism of Development of Antibiotic Tolerance (Review). Appl. Biochem. Microbiol. 2018, 54, 108–127. [Google Scholar] [CrossRef]
- Santajit, S.; Indrawattana, N. Mechanisms of Antimicrobial Resistance in ESKAPE Pathogens. Biomed. Res. Int. 2016, 2016, 2475067. [Google Scholar] [CrossRef]
- Miller, K.P.; Wang, L.; Benicewicz, B.C.; Decho, A.W. Inorganic nanoparticles engineered to attack bacteria. Chem. Soc. Rev. 2015, 44, 7787–7807. [Google Scholar] [CrossRef] [PubMed]
- Piñón-Castillo, H.A.; Martínez-Chamarro, R.; Reyes-Martínez, R.; Salinas-Vera, Y.M.; Manjarrez-Nevárez, L.A.; Muñoz-Castellanos, L.N.; López-Camarillo, C.; Orrantia-Borunda, E. Palladium Nanoparticles Functionalized with PVP-Quercetin Inhibits Cell Proliferation and Activates Apoptosis in Colorectal Cancer Cells. Appl. Sci. 2021, 11, 1988. [Google Scholar] [CrossRef]
- Shrestha, A.; Kishen, A. Antibacterial Nanoparticles in Endodontics: A Review. J. Endod. 2016, 42, 1417–1426. [Google Scholar] [CrossRef] [PubMed]
- Bi, J.; Li, T.; Ren, H.; Ling, R.; Wu, Z.; Qin, W. Capillary electrophoretic determination of heavy-metal ions using 11-mercaptoundecanoic acid and 6-mercapto-1-hexanol co-functionalized gold nanoparticle as colorimetric probe. J. Chromatogr. A 2019, 1594, 208–215. [Google Scholar] [CrossRef]
- Thakkar, K.N.; Mhatre, S.S.; Parikh, R.Y. Biological synthesis of metallic nanoparticles. Nanomed. Nanotechnol. Biol. Med. 2010, 6, 257–262. [Google Scholar] [CrossRef]
- Brewer, M.; Zhang, T.; Dong, W.; Rutherford, M.; Tian, Z.R. Future Approaches of Nanomedicine in Clinical Science. Med. Clin. N. Am. 2007, 91, 963–1016. [Google Scholar] [CrossRef]
- McNamara, K.; Tofail, S.A.M. Nanosystems: The use of nanoalloys, metallic, bimetallic, and magnetic nanoparticles in biomedical applications. Phys. Chem. Chem. Phys. 2015, 17, 27981–27995. [Google Scholar] [CrossRef]
- Kaweeteerawat, C.; Na Ubol, P.; Sangmuang, S.; Aueviriyavit, S.; Maniratanachote, R. Mechanisms of antibiotic resistance in bacteria mediated by silver nanoparticles. J. Toxicol. Environ. Health Part. A 2017, 80, 1276–1289. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Xu, H.; Chen, Z.-S.; Chen, G. Biosynthesis of Nanoparticles by Microorganisms and Their Applications. J. Nanomater. 2011, 2011, 270974. [Google Scholar] [CrossRef]
- Liu, J.; Qiao, S.Z.; Hu, Q.H.; Max Lu, G.Q. Magnetic Nanocomposites with Mesoporous Structures: Synthesis and Applications. Small 2011, 7, 425–443. [Google Scholar] [CrossRef]
- Spirescu, V.A.; Niculescu, A.-G.; Slave, Ș.; Bîrcă, A.C.; Dorcioman, G.; Grumezescu, V.; Holban, A.M.; Oprea, O.-C.; Vasile, B.Ș.; Grumezescu, A.M.; et al. Anti-Biofilm Coatings Based on Chitosan and Lysozyme Functionalized Magnetite Nanoparticles. Antibiotics 2021, 10, 1269. [Google Scholar] [CrossRef]
- Motelica, L.; Ficai, D.; Oprea, O.; Ficai, A.; Trusca, R.D.; Andronescu, E.; Holban, A.M. Biodegradable alginate films with ZnO nanoparticles and citronella essential oil-a novel antimicrobial structure. Pharmaceutics 2021, 13, 1020. [Google Scholar] [CrossRef] [PubMed]
- Vasile, B.S.; Chircov, C.; Matei, M.; Neacs, I.A. Iron Oxide—Silica Core—Shell Nanoparticles Functionalized with Essential Oils for Antimicrobial Therapies. Antibiotics 2021, 10, 1138. [Google Scholar]
- Puiu, R.A.; Balaure, P.C.; Constantinescu, E.; Grumezescu, A.M.; Andronescu, E.; Oprea, O.C.; Vasile, B.S.; Grumezescu, V.; Negut, I.; Nica, I.C.; et al. Anti-cancer nanopowders and maple-fabricated thin coatings based on spions surface modified with paclitaxel loaded β-cyclodextrin. Pharmaceutics 2021, 13, 1356. [Google Scholar] [CrossRef]
- Isela Ruvalcaba Ontiveros, R.; Alberto Duarte Moller, J.; Rocío Carrasco Hernandez, A.; Esperanza Esparza-Ponce, H.; Orrantia Borunda, E.; Deisy Gómez Esparza, C.; Manuel Olivares Ramírez, J. A Simple Way to Produce Gold Nanoshells for Cancer Therapy. In Current Topics in Biochemical Engineering; IntechOpen: London, UK, 2019. [Google Scholar] [CrossRef]
- Khameneh, B.; Diab, R.; Ghazvini, K.; Fazly Bazzaz, B.S. Breakthroughs in bacterial resistance mechanisms and the potential ways to combat them. Microb. Pathog. 2016, 95, 32–42. [Google Scholar] [CrossRef]
- Slavin, Y.N.; Asnis, J.; Häfeli, U.O.; Bach, H. Metal nanoparticles: Understanding the mechanisms behind antibacterial activity. J. Nanobiotechnol. 2017, 15, 65. [Google Scholar] [CrossRef]
- AlMatar, M.; Makky, E.A.; Var, I.; Koksal, F. The Role of Nanoparticles in the Inhibition of Multidrug-Resistant Bacteria and Biofilms. Curr. Drug Deliv. 2018, 15, 470–484. [Google Scholar] [CrossRef]
- Franci, G.; Falanga, A.; Galdiero, S.; Palomba, L.; Rai, M.; Morelli, G.; Galdiero, M. Silver Nanoparticles as Potential Antibacterial Agents. Molecules 2015, 20, 8856–8874. [Google Scholar] [CrossRef]
- Nisar, P.; Ali, N.; Rahman, L.; Ali, M.; Shinwari, Z.K. Antimicrobial activities of biologically synthesized metal nanoparticles: An insight into the mechanism of action. JBIC J. Biol. Inorg. Chem. 2019, 24, 929–941. [Google Scholar] [CrossRef]
- Abdal Dayem, A.; Hossain, M.; Lee, S.; Kim, K.; Saha, S.; Yang, G.-M.; Choi, H.; Cho, S.-G. The Role of Reactive Oxygen Species (ROS) in the Biological Activities of Metallic Nanoparticles. Int. J. Mol. Sci. 2017, 18, 120. [Google Scholar] [CrossRef]
- Snezhkina, A.V.; Kudryavtseva, A.V.; Kardymon, O.L.; Savvateeva, M.V.; Melnikova, N.V.; Krasnov, G.S.; Dmitriev, A.A. ROS Generation and Antioxidant Defense Systems in Normal and Malignant Cells. Oxid. Med. Cell. Longev. 2019, 2019, 6175804. [Google Scholar] [CrossRef] [PubMed]
- Gupta, A.; Mumtaz, S.; Li, C.-H.; Hussain, I.; Rotello, V.M. Combatting antibiotic-resistant bacteria using nanomaterials. Chem. Soc. Rev. 2019, 48, 415–427. [Google Scholar] [CrossRef] [PubMed]
- Ulloa-Ogaz, A.L.; Piñón-Castillo, H.A.; Muñoz-Castellanos, L.N.; Athie-García, M.S.; Ballinas-Casarrubias, M.D.L.; Murillo-Ramirez, J.G.; Flores-Ongay, L.Á.; Duran, R.; Orrantia-Borunda, E. Oxidative damage to Pseudomonas aeruginosa ATCC 27833 and Staphylococcus aureus ATCC 24213 induced by CuO-NPs. Environ. Sci. Pollut. Res. 2017, 24, 22048–22060. [Google Scholar] [CrossRef] [PubMed]
- Amini, S.M. Preparation of antimicrobial metallic nanoparticles with bioactive compounds. Mater. Sci. Eng. C 2019, 103, 109809. [Google Scholar] [CrossRef]
- Buzea, C.; Pacheco, I.I.; Robbie, K. Nanomaterials and nanoparticles: Sources and toxicity. Biointerphases 2007, 2, MR17–MR71. [Google Scholar] [CrossRef]
- El-Ansary, A.; Al-Daihan, S. On the Toxicity of Therapeutically Used Nanoparticles: An Overview. J. Toxicol. 2009, 2009, 754810. [Google Scholar] [CrossRef]
- Pérez-Díaz, M.A.; Boegli, L.; James, G.; Velasquillo, C.; Sánchez-Sánchez, R.; Martínez-Martínez, R.-E.; Martínez-Castañón, G.A.; Martinez-Gutierrez, F. Silver nanoparticles with antimicrobial activities against Streptococcus mutans and their cytotoxic effect. Mater. Sci. Eng. C 2015, 55, 360–366. [Google Scholar] [CrossRef] [PubMed]
- Padmavathy, N.; Vijayaraghavan, R. Interaction of ZnO Nanoparticles with Microbes—A Physio and Biochemical Assay. J. Biomed. Nanotechnol. 2011, 7, 813–822. [Google Scholar] [CrossRef]
- Yu, J.; Zhang, W.; Li, Y.; Wang, G.; Yang, L.; Jin, J.; Chen, Q.; Huang, M. Synthesis, characterization, antimicrobial activity and mechanism of a novel hydroxyapatite whisker/nano zinc oxide biomaterial. Biomed. Mater. 2014, 10, 015001. [Google Scholar] [CrossRef]
- Lesniak, A.; Salvati, A.; Santos-Martinez, M.J.; Radomski, M.W.; Dawson, K.A.; Åberg, C. Nanoparticle Adhesion to the Cell Membrane and Its Effect on Nanoparticle Uptake Efficiency. J. Am. Chem. Soc. 2013, 135, 1438–1444. [Google Scholar] [CrossRef] [PubMed]
- Sarwar, A.; Katas, H.; Samsudin, S.N.; Zin, N.M. Regioselective Sequential Modification of Chitosan via Azide-Alkyne Click Reaction: Synthesis, Characterization, and Antimicrobial Activity of Chitosan Derivatives and Nanoparticles. PLoS ONE 2015, 10, e0123084. [Google Scholar] [CrossRef]
- Ahmed, B.; Hashmi, A.; Khan, M.S.; Musarrat, J. ROS mediated destruction of cell membrane, growth and biofilms of human bacterial pathogens by stable metallic AgNPs functionalized from bell pepper extract and quercetin. Adv. Powder Technol. 2018, 29, 1601–1616. [Google Scholar] [CrossRef]
- Yang, B.; Chen, Y.; Shi, J. Reactive Oxygen Species (ROS)-Based Nanomedicine. Chem. Rev. 2019, 119, 4881–4985. [Google Scholar] [CrossRef] [PubMed]
- Shaikh, S.; Nazam, N.; Rizvi, S.M.D.; Ahmad, K.; Baig, M.H.; Lee, E.J.; Choi, I. Mechanistic Insights into the Antimicrobial Actions of Metallic Nanoparticles and Their Implications for Multidrug Resistance. Int. J. Mol. Sci. 2019, 20, 2468. [Google Scholar] [CrossRef]
- Pan, W.-Y.; Huang, C.-C.; Lin, T.-T.; Hu, H.-Y.; Lin, W.-C.; Li, M.-J.; Sung, H.-W. Synergistic antibacterial effects of localized heat and oxidative stress caused by hydroxyl radicals mediated by graphene/iron oxide-based nanocomposites. Nanomed. Nanotechnol. Biol. Med. 2016, 12, 431–438. [Google Scholar] [CrossRef]
- Zhang, W.; Li, Y.; Niu, J.; Chen, Y. Photogeneration of Reactive Oxygen Species on Uncoated Silver, Gold, Nickel, and Silicon Nanoparticles and Their Antibacterial Effects. Langmuir 2013, 29, 4647–4651. [Google Scholar] [CrossRef] [PubMed]
- Lin, J.; Zhang, H.; Chen, Z.; Zheng, Y. Penetration of Lipid Membranes by Gold Nanoparticles: Insights into Cellular Uptake, Cytotoxicity, and Their Relationship. ACS Nano 2010, 4, 5421–5429. [Google Scholar] [CrossRef] [PubMed]
- Yamanaka, M.; Hara, K.; Kudo, J. Bactericidal Actions of a Silver Ion Solution on Escherichia coli, Studied by Energy-Filtering Transmission Electron Microscopy and Proteomic Analysis. Appl. Environ. Microbiol. 2005, 71, 7589–7593. [Google Scholar] [CrossRef]
- Cui, Y.; Zhao, Y.; Tian, Y.; Zhang, W.; Lü, X.; Jiang, X. The molecular mechanism of action of bactericidal gold nanoparticles on Escherichia coli. Biomaterials 2012, 33, 2327–2333. [Google Scholar] [CrossRef] [PubMed]
- Su, G.; Zhang, X.; Giesy, J.P.; Musarrat, J.; Saquib, Q.; Alkhedhairy, A.A.; Yu, H. Comparison on the molecular response profiles between nano zinc oxide (ZnO) particles and free zinc ion using a genome-wide toxicogenomics approach. Environ. Sci. Pollut. Res. 2015, 22, 17434–17442. [Google Scholar] [CrossRef]
- Nagy, A.; Harrison, A.; Sabbani, S.; Munson, R.S., Jr.; Dutta, P.K.; Waldman, W.J. Silver nanoparticles embedded in zeolite membranes: Release of silver ions and mechanism of antibacterial action. Int. J. Nanomed. 2011, 6, 1833. [Google Scholar] [CrossRef]
- Qiu, W.; Zheng, X.; Wei, Y.; Zhou, X.; Zhang, K.; Wang, S.; Cheng, L.; Li, Y.; Ren, B.; Xu, X.; et al. D-Alanine metabolism is essential for growth and biofilm formation of Streptococcus mutans. Mol. Oral Microbiol. 2016, 31, 435–444. [Google Scholar] [CrossRef] [PubMed]
- Su, H.-L.; Chou, C.-C.; Hung, D.-J.; Lin, S.-H.; Pao, I.-C.; Lin, J.-H.; Huang, F.-L.; Dong, R.-X.; Lin, J.-J. The disruption of bacterial membrane integrity through ROS generation induced by nanohybrids of silver and clay. Biomaterials 2009, 30, 5979–5987. [Google Scholar] [CrossRef] [PubMed]
- Niño-Martínez, N.; Salas Orozco, M.F.; Martínez-Castañón, G.-A.; Torres Méndez, F.; Ruiz, F. Molecular Mechanisms of Bacterial Resistance to Metal and Metal Oxide Nanoparticles. Int. J. Mol. Sci. 2019, 20, 2808. [Google Scholar] [CrossRef] [PubMed]
- Salas-Orozco, M.; Niño-Martínez, N.; Martínez-Castañón, G.-A.; Méndez, F.T.; Jasso, M.E.C.; Ruiz, F. Mechanisms of Resistance to Silver Nanoparticles in Endodontic Bacteria: A Literature Review. J. Nanomater. 2019, 2019, 7630316. [Google Scholar] [CrossRef]
- Webster, T.J.; Seil, I. Antimicrobial applications of nanotechnology: Methods and literature. Int. J. Nanomed. 2012, 7, 2767. [Google Scholar] [CrossRef]
- Poyedinok, N.; Mykhaylova, O.; Sergiichuk, N.; Tugay, T.; Tugay, A.; Lopatko, S.; Matvieieva, N. Effect of Colloidal Metal Nanoparticles on Biomass, Polysaccharides, Flavonoids, and Melanin Accumulation in Medicinal Mushroom Inonotus obliquus (Ach.:Pers.) Pilát. Appl. Biochem. Biotechnol. 2020, 191, 1315–1325. [Google Scholar] [CrossRef] [PubMed]
- Courtois, P.; Rorat, A.; Lemiere, S.; Levard, C.; Pradas, A.; Lors, C.; Vandenbulcke, F. Accumulation and Localization of Silver Nanoparticles in Eisenia Fetida Earthworms. Environ. Sci. Pollut. Res. 2021, 28, 3756–3765. [Google Scholar] [CrossRef] [PubMed]
- Yarjanli, Z.; Ghaedi, K.; Esmaeili, A.; Rahgozar, S.; Zarrabi, A. Iron oxide nanoparticles may damage to the neural tissue through iron accumulation, oxidative stress, and protein aggregation. BMC Neurosci. 2017, 18, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Matter, M.T.; Li, J.H.; Lese, I.; Schreiner, C.; Bernard, L.; Scholder, O.; Hubeli, J.; Keevend, K.; Tsolaki, E.; Bertero, E.; et al. Multiscale Analysis of Metal Oxide Nanoparticles in Tissue: Insights into Biodistribution and Biotransformation. Adv. Sci. 2020, 7, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Morales-Dalmau, J.; Vilches, C.; Sanz, V.; De Miguel, I.; Rodríguez-Fajardo, V.; Berto, P.; Martínez-Lozano, M.; Casanovas, O.; Durduran, T.; Quidant, R. Quantification of gold nanoparticle accumulation in tissue by two-photon luminescence microscopy. Nanoscale 2019, 11, 11331–11339. [Google Scholar] [CrossRef] [PubMed]
- Arya, S.S.; Sharma, M.M.; Das, R.K.; Rookes, J.; Cahill, D.; Lenka, S.K. Heliyon Vanillin mediated green synthesis and application of gold nanoparticles for reversal of antimicrobial resistance in Pseudomonas aeruginosa clinical isolates. Heliyon 2019, 5, e02021. [Google Scholar] [CrossRef]
- Chavan, C.; Kamble, S.; Murthy, A.V.R.; Kale, S.N. Ampicillin-mediated functionalized gold nanoparticles against ampicillin-resistant bacteria: Strategy, preparation and interaction studies. Nanotechnology 2020, 31, 215604. [Google Scholar] [CrossRef]
- Kalita, S.; Kandimalla, R.; Sharma, K.K.; Kataki, A.C.; Deka, M.; Kotoky, J. Amoxicillin functionalized gold nanoparticles reverts MRSA resistance. Mater. Sci. Eng. C 2016, 61, 720–727. [Google Scholar] [CrossRef]
- Armentano, I.; Arciola, C.R.; Fortunati, E.; Ferrari, D.; Mattioli, S.; Amoroso, C.F.; Rizzo, J.; Kenny, J.M.; Imbriani, M.; Visai, L. The Interaction of Bacteria with Engineered Nanostructured Polymeric Materials: A Review. Sci. World J. 2014, 2014, 410423. [Google Scholar] [CrossRef]
- Gao, W.; Thamphiwatana, S.; Angsantikul, P.; Zhang, L. Nanoparticle approaches against bacterial infections. Wiley Interdiscip. Rev. Nanomed. Nanobiotechnol. 2014, 6, 532–547. [Google Scholar] [CrossRef]
- Li, H.; Chen, Q.; Zhao, J.; Urmila, K. Enhancing the antimicrobial activity of natural extraction using the synthetic ultrasmall metal nanoparticles. Sci. Rep. 2015, 5, 11033. [Google Scholar] [CrossRef] [PubMed]
- Luan, B.; Huynh, T.; Zhou, R. Complete wetting of graphene by biological lipids. Nanoscale 2016, 8, 5750–5754. [Google Scholar] [CrossRef]
- McBee, M.E.; Chionh, Y.H.; Sharaf, M.L.; Ho, P.; Cai, M.W.L.; Dedon, P.C. Production of Superoxide in Bacteria Is Stress- and Cell State-Dependent: A Gating-Optimized Flow Cytometry Method that Minimizes ROS Measurement Artifacts with Fluorescent Dyes. Front. Microbiol. 2017, 8, 459. [Google Scholar] [CrossRef]
- Ye, Q.; Chen, W.; Huang, H.; Tang, Y.; Wang, W.; Meng, F.; Wang, H.; Zheng, Y. Iron and zinc ions, potent weapons against multidrug-resistant bacteria. Appl. Microbiol. Biotechnol. 2020, 104, 5213–5227. [Google Scholar] [CrossRef] [PubMed]
- Zou, L.; Wang, J.; Gao, Y.; Ren, X.; Rottenberg, M.E.; Lu, J.; Holmgren, A. Synergistic antibacterial activity of silver with antibiotics correlating with the upregulation of the ROS production. Sci. Rep. 2018, 8, 11131. [Google Scholar] [CrossRef]
- Xu, R.; Wang, D.; Zhang, J.; Li, Y. Shape-Dependent Catalytic Activity of Silver Nanoparticles for the Oxidation of Styrene. Chem.-Asian J. 2006, 1, 888–893. [Google Scholar] [CrossRef]
- Agnihotri, S.; Mukherji, S.; Mukherji, S. Size-controlled silver nanoparticles synthesized over the range 5–100 nm using the same protocol and their antibacterial efficacy. RSC Adv. 2014, 4, 3974–3983. [Google Scholar] [CrossRef]
- Hachicho, N.; Hoffmann, P.; Ahlert, K.; Heipieper, H.J. Effect of silver nanoparticles and silver ions on growth and adaptive response mechanisms of Pseudomonas putida mt-2. FEMS Microbiol. Lett. 2014, 355, 71–77. [Google Scholar] [CrossRef]
- Graves, J.L.; Tajkarimi, M.; Cunningham, Q.; Campbell, A.; Nonga, H.; Harrison, S.H.; Barrick, J.E. Rapid evolution of silver nanoparticle resistance in Escherichia coli. Front. Genet. 2015, 6, 42. [Google Scholar] [CrossRef]
- Panáček, A.; Kvítek, L.; Smékalová, M.; Večeřová, R.; Kolář, M.; Röderová, M.; Dyčka, F.; Šebela, M.; Prucek, R.; Tomanec, O.; et al. Bacterial resistance to silver nanoparticles and how to overcome it. Nat. Nanotechnol. 2018, 13, 65–71. [Google Scholar] [CrossRef]
- Lellouche, J.; Friedman, A.; Lahmi, R.; Gedanken, A.; Banin, E. Antibiofilm surface functionalization of catheters by magnesium fluoride nanoparticles. Int. J. Nanomed. 2012, 7, 1175. [Google Scholar] [CrossRef][Green Version]
- Yun, S.; Huang, J.J. Routes for Drug Delivery: Sustained-Release Devices. Retinal Pharmacotherapeut. 2016, 55, 84–92. [Google Scholar]
- Peng, Z.; Ni, J.; Zheng, K.; Shen, Y.; Wang, X.; He, G.; Jin, S.; Tang, T. Dual effects and mechanism of TiO2 nanotube arrays in reducing bacterial colonization and enhancing C3H10T1/2 cell adhesion. Int. J. Nanomed. 2013, 8, 3093. [Google Scholar] [CrossRef]
- Wunderink, R.G.; Giamarellos-Bourboulis, E.J.; Rahav, G.; Mathers, A.J.; Bassetti, M.; Vazquez, J.; Cornely, O.A.; Solomkin, J.; Bhowmick, T.; Bishara, J.; et al. Effect and Safety of Meropenem–Vaborbactam versus Best-Available Therapy in Patients with Carbapenem-Resistant Enterobacteriaceae Infections: The TANGO II Randomized Clinical Trial. Infect. Dis. Ther. 2018, 7, 439–455. [Google Scholar] [CrossRef]
- Jijie, R.; Barras, A.; Teodorescu, F.; Boukherroub, R.; Szunerits, S. Advancements on the molecular design of nanoantibiotics: Current level of development and future challenges. Mol. Syst. Des. Eng. 2017, 2, 349–369. [Google Scholar] [CrossRef]
- Qi, G.; Li, L.; Yu, F.; Wang, H. Vancomycin-Modified Mesoporous Silica Nanoparticles for Selective Recognition and Killing of Pathogenic Gram-Positive Bacteria Over Macrophage-Like Cells. ACS Appl. Mater. Interfaces 2013, 5, 10874–10881. [Google Scholar] [CrossRef]
- Tang, S.; Zheng, J. Antibacterial Activity of Silver Nanoparticles: Structural Effects. Adv. Healthc. Mater. 2018, 7, 1701503. [Google Scholar] [CrossRef]
- Cheon, J.Y.; Kim, S.J.; Rhee, Y.H.; Kwon, O.H.; Park, W.H. Shape-dependent antimicrobial activities of silver nanoparticles. Int. J. Nanomed. 2019, 14, 2773–2780. [Google Scholar] [CrossRef] [PubMed]
- Burdușel, A.-C.; Gherasim, O.; Grumezescu, A.M.; Mogoantă, L.; Ficai, A.; Andronescu, E. Biomedical Applications of Silver Nanoparticles: An Up-to-Date Overview. Nanomaterials 2018, 8, 681. [Google Scholar] [CrossRef]
- Lee, S.; Jun, B.-H. Silver Nanoparticles: Synthesis and Application for Nanomedicine. Int. J. Mol. Sci. 2019, 20, 865. [Google Scholar] [CrossRef] [PubMed]
- Helmlinger, J.; Sengstock, C.; Groß-Heitfeld, C.; Mayer, C.; Schildhauer, T.A.; Köller, M.; Epple, M. Silver nanoparticles with different size and shape: Equal cytotoxicity, but different antibacterial effects. RSC Adv. 2016, 6, 18490–18501. [Google Scholar] [CrossRef]
- Raza, M.; Kanwal, Z.; Rauf, A.; Sabri, A.; Riaz, S.; Naseem, S. Size- and Shape-Dependent Antibacterial Studies of Silver Nanoparticles Synthesized by Wet Chemical Routes. Nanomaterials 2016, 6, 74. [Google Scholar] [CrossRef]
- Pal, S.; Tak, Y.K.; Song, J.M. Does the Antibacterial Activity of Silver Nanoparticles Depend on the Shape of the Nanoparticle? A Study of the Gram-Negative Bacterium Escherichia coli. Appl. Environ. Microbiol. 2007, 73, 1712–1720. [Google Scholar] [CrossRef]
- Nanda, A.; Saravanan, M. Biosynthesis of silver nanoparticles from Staphylococcus aureus and its antimicrobial activity against MRSA and MRSE. Nanomed. Nanotechnol. Biol. Med. 2009, 5, 452–456. [Google Scholar] [CrossRef]
- Lara, H.H.; Ayala-Núñez, N.V.; del Carmen Ixtepan Turrent, L.; Rodríguez Padilla, C. Bactericidal effect of silver nanoparticles against multidrug-resistant bacteria. World J. Microbiol. Biotechnol. 2010, 26, 615–621. [Google Scholar] [CrossRef]
- Ashraf, N.; Ahmad, F.; Lu, Y.; Yin, D.-C. Bacterial extracellular protein interacts with silver ions to produce protein-encapsulated bactericidal AgNPs. Process. Biochem. 2021, 106, 120–129. [Google Scholar] [CrossRef]
- Shirzadi-Ahodashti, M.; Mizwari, Z.M.; Hashemi, Z.; Rajabalipour, S.; Ghoreishi, S.M.; Mortazavi-Derazkola, S.; Ebrahimzadeh, M.A. Discovery of high antibacterial and catalytic activities of biosynthesized silver nanoparticles using C. fruticosus (CF-AgNPs) against multi-drug resistant clinical strains and hazardous pollutants. Environ. Technol. Innov. 2021, 23, 101607. [Google Scholar] [CrossRef]
- Gudkov, S.V.; Burmistrov, D.E.; Serov, D.A.; Rebezov, M.B.; Semenova, A.A.; Lisitsyn, A.B. A Mini Review of Antibacterial Properties of ZnO Nanoparticles. Front. Phys. 2021, 9, 641481. [Google Scholar] [CrossRef]
- Rabani, I.; Lee, S.; Kim, H.; Yoo, J.; Hussain, S.; Maqbool, T.; Seo, Y. Journal of Environmental Chemical Engineering Engineering-safer-by design ZnO nanoparticles incorporated cellulose nanofiber hybrid for high UV protection and low photocatalytic activity with mechanism. J. Environ. Chem. Eng. 2021, 9, 105845. [Google Scholar] [CrossRef]
- Sevcik, J.; Urbanek, P.; Skoda, D.; Jamatia, T.; Nadazdy, V.; Urbanek, M.; Antos, J.; Munster, L.; Kuritka, I. Materials & Design Energy resolved-electrochemical impedance spectroscopy investigation of the role of Al-doped ZnO nanoparticles in electronic structure modification of polymer nanocomposite LEDs. Mater. Des. 2021, 205, 109738. [Google Scholar] [CrossRef]
- Bankar, D.B.; Hawaldar, R.R.; Arbuj, S.S.; Shinde, S.T.; Gadde, J.R.; Rakshe, D.S.; Amalnerkar, D.P.; Kanade, K.G. Palladium loaded on ZnO nanoparticles: Synthesis, characterization and application as heterogeneous catalyst for Suzuki–Miyaura cross-coupling reactions under ambient and ligand-free conditions. Mater. Chem. Phys. 2019, 243, 122561. [Google Scholar] [CrossRef]
- Sathish, P.; Dineshbabu, N.; Ravichandran, K.; Arun, T.; Karuppasamy, P. Combustion synthesis, characterization and antibacterial properties of pristine ZnO and Ga doped ZnO nanoparticles. Ceram. Int. 2021, 47, 27934–27941. [Google Scholar] [CrossRef]
- Anwar, M.A.; Aqib, A.I.; Ashfaq, K.; Deeba, F.; Khan, M.K.; Khan, S.R.; Muzammil, I.; Shoaib, M.; Naseer, M.A.; Riaz, T.; et al. Antimicrobial resistance modulation of MDR E. coli by antibiotic coated ZnO nanoparticles. Microb. Pathog. 2020, 148, 104450. [Google Scholar] [CrossRef] [PubMed]
- Prasad, A.R.; Williams, L.; Garvasis, J.; Shamsheera, K.O.; Basheer, S.M.; Kuruvilla, M.; Joseph, A. Applications of phytogenic ZnO nanoparticles: A review on recent advancements. J. Mol. Liq. 2021, 331, 115805. [Google Scholar] [CrossRef]
- Benitez-salazar, M.I.; Ni, V.E.; Rodríguez-p, J.E.; Caldas-arias, L.; Fern, I. Chemical synthesis versus green synthesis to obtain ZnO powders: Evaluation of the antibacterial capacity of the nanoparticles obtained by the chemical method. J. Environ. Chem. Eng. 2021, 9, 106544. [Google Scholar] [CrossRef]
- Gold, K.; Slay, B.; Knackstedt, M.; Gaharwar, A.K. Antimicrobial Activity of Metal and Metal-Oxide Based Nanoparticles. Adv. Ther. 2018, 1, 1700033. [Google Scholar] [CrossRef]
- Saha, R.K.; Debanath, M.K.; Paul, B.; Medhi, S.; Saikia, E. Antibacterial and nonlinear dynamical analysis of flower and hexagon-shaped ZnO microstructures. Sci. Rep. 2020, 10, 2598. [Google Scholar] [CrossRef]
- Turner, R.J. Metal-based antimicrobial strategies. Microb. Biotechnol. 2017, 10, 1062–1065. [Google Scholar] [CrossRef]
- Banoee, M.; Seif, S.; Nazari, Z.E.; Jafari-Fesharaki, P.; Shahverdi, H.R.; Moballegh, A.; Moghaddam, K.M.; Shahverdi, A.R. ZnO nanoparticles enhanced antibacterial activity of ciprofloxacin against Staphylococcus aureus and Escherichia coli. J. Biomed. Mater. Res. Part B Appl. Biomater. 2010, 93B, 557–561. [Google Scholar] [CrossRef]
- Patra, P.; Mitra, S.; Debnath, N.; Pramanik, P.; Arunava, G. Ciprofloxacin conjugated zinc oxide nanoparticle: A camouflage towards multidrug resistant bacteria. Bull. Mater. Sci. 2014, 3, 199–206. [Google Scholar] [CrossRef]
- Zhong, Q.; Tian, J.; Liu, T.; Guo, Z.; Ding, S.; Li, H. Preparation and antibacterial properties of carboxymethyl chitosan/ZnO nanocomposite microspheres with enhanced biocompatibility. Mater. Lett. 2018, 212, 58–61. [Google Scholar] [CrossRef]
- Eleraky, N.E.; Allam, A.; Hassan, S.B.; Omar, M.M. Nanomedicine Fight against Antibacterial Resistance: An Overview of the Recent Pharmaceutical Innovations. Pharmaceutics 2020, 12, 142. [Google Scholar] [CrossRef]
- Petros, R.A.; DeSimone, J.M. Strategies in the design of nanoparticles for therapeutic applications. Nat. Rev. Drug Discov. 2010, 9, 615–627. [Google Scholar] [CrossRef] [PubMed]
- Pati, R.; Mehta, R.K.; Mohanty, S.; Padhi, A.; Sengupta, M.; Vaseeharan, B.; Goswami, C.; Sonawane, A. Topical application of zinc oxide nanoparticles reduces bacterial skin infection in mice and exhibits antibacterial activity by inducing oxidative stress response and cell membrane disintegration in macrophages. Nanomed. Nanotechnol. Biol. Med. 2014, 10, 1195–1208. [Google Scholar] [CrossRef] [PubMed]
- Premanathan, M.; Karthikeyan, K.; Jeyasubramanian, K.; Manivannan, G. Selective toxicity of ZnO nanoparticles toward Gram-positive bacteria and cancer cells by apoptosis through lipid peroxidation. Nanomed. Nanotechnol. Biol. Med. 2011, 7, 184–192. [Google Scholar] [CrossRef]
- Alex, S.; Tiwari, A. Functionalized gold nanoparticles: Synthesis, properties and applications-A review. J. Nanosci. Nanotechnol. 2015, 15, 1869–1894. [Google Scholar] [CrossRef]
- Xu, G.; Hou, J.; Zhao, Y.; Bao, J.; Yang, M.; Fa, H.; Yang, Y.; Li, L.; Huo, D.; Hou, C. Dual-signal aptamer sensor based on polydopamine-gold nanoparticles and exonuclease I for ultrasensitive malathion detection. Sens. Actuators B Chem. 2019, 428–436. [Google Scholar] [CrossRef]
- Yang, H.; Xu, W.; Zhou, Y. Signal amplification in immunoassays by using noble metal nanoparticles: A review. Microchim. Acta 2019, 186, 859. [Google Scholar] [CrossRef] [PubMed]
- Elbialy, N.S.; Fathy, M.M.; Khalil, W.M. Doxorubicin loaded magnetic gold nanoparticles for in vivo targeted drug delivery. Int. J. Pharm. 2015, 490, 190–199. [Google Scholar] [CrossRef]
- Khandelwal, P.; Singh, D.K.; Poddar, P. Advances in the Experimental and Theoretical Understandings of Antibiotic Conjugated Gold Nanoparticles for Antibacterial Applications. ChemistrySelect 2019, 4, 6719–6738. [Google Scholar] [CrossRef]
- Mu, H.; Liu, Q.; Niu, H.; Sun, Y.; Duan, J. Gold nanoparticles make chitosan–streptomycin conjugates effective towards Gram-negative bacterial biofilm. RSC Adv. 2016, 6, 8714–8721. [Google Scholar] [CrossRef]
- Hu, D.; Li, H.; Wang, B.; Ye, Z.; Lei, W.; Jia, F.; Jin, Q.; Ren, K.-F.; Ji, J. Surface-Adaptive Gold Nanoparticles with Effective Adherence and Enhanced Photothermal Ablation of Methicillin-Resistant Staphylococcus aureus Biofilm. ACS Nano 2017, 11, 9330–9339. [Google Scholar] [CrossRef] [PubMed]
- Xie, Y.; Liu, Y.; Yang, J.; Liu, Y.; Hu, F.; Zhu, K.; Jiang, X. Gold Nanoclusters for Targeting Methicillin-Resistant Staphylococcus aureus In Vivo. Angew. Chem. Int. Ed. 2018, 57, 3958–3962. [Google Scholar] [CrossRef]
- Rad, M.R.; Kazemian, H.; Yazdani, F.; Monfared, M.R.Z.; Rahdar, H.; Javadi, A.; Kodori, M. Antibacterial Activity of Gold Nanoparticles Conjugated by Aminoglycosides against A. baumannii Isolates from Burn Patients. Recent Pat. Antiinfect. Drug Discov. 2019, 13, 256–264. [Google Scholar] [CrossRef]
- Khan, F.; Lee, J.-W.; Manivasagan, P.; Pham, D.T.N.; Oh, J.; Kim, Y.-M. Synthesis and characterization of chitosan oligosaccharide-capped gold nanoparticles as an effective antibiofilm drug against the Pseudomonas aeruginosa PAO1. Microb. Pathog. 2019, 135, 103623. [Google Scholar] [CrossRef] [PubMed]
- Riaz, S.; Fatima Rana, N.; Hussain, I.; Tanweer, T.; Nawaz, A.; Menaa, F.; Janjua, H.A.; Alam, T.; Batool, A.; Naeem, A.; et al. Effect of Flavonoid-Coated Gold Nanoparticles on Bacterial Colonization in Mice Organs. Nanomaterials 2020, 10, 1769. [Google Scholar] [CrossRef] [PubMed]
- Stoyanova, A.; Hitkova, H.; Bachvarova-Nedelcheva, A.; Iordanova, R.; Ivanova, N.; Sredkova, M. Synthesis and antibacterial activity of TiO2/ZnO nanocomposites prepared via nonhydrolytic route. J. Chem. Technol. Metall. 2013, 48, 154–161. [Google Scholar]
- Menazea, A.A.; Awwad, N.S. Antibacterial activity of TiO2 doped ZnO composite synthesized via laser ablation route for antimicrobial application. J. Mater. Res. Technol. 2020, 9, 9434–9441. [Google Scholar] [CrossRef]
- Torbat, T.V.; Javanbakht, V. Fabrication of TiO2/Zn2TiO4/Ag Nanocomposite for Synergic Effects of UV Radiation Protection and Antibacterial Activity in Sunscreen; Elsevier: Amsterdam, The Netherlands, 2019; ISBN 8415683111. [Google Scholar]
- Häffner, S.M.; Parra-ortiz, E.; Skoda, M.W.A.; Saerbeck, T.; Browning, K.L.; Malmsten, M. Composition effects on photooxidative membrane destabilization by TiO2 nanoparticles. J. Colloid Interface Sci. 2021, 584, 19–33. [Google Scholar] [CrossRef]
- Huang, Y.-Y.; Rajda, P.J.; Szewczyk, G.; Bhayana, B.; Chiang, L.Y.; Sarna, T.; Hamblin, M.R. Sodium nitrite potentiates antimicrobial photodynamic inactivation: Possible involvement of peroxynitrate. Photochem. Photobiol. Sci. 2019, 18, 505–515. [Google Scholar] [CrossRef] [PubMed]
- Mohammed, L.; Gomaa, H.G.; Ragab, D.; Zhu, J. Magnetic nanoparticles for environmental and biomedical applications: A review. Particuology 2017, 30, 1–14. [Google Scholar] [CrossRef]
- Flores-González, M.; Talavera-Rojas, M.; Soriano-Vargas, E.; Rodríguez-González, V. Practical mediated-assembly synthesis of silver nanowires using commercial Camellia sinensis extracts and their antibacterial properties. New J. Chem. 2018, 42, 2133–2139. [Google Scholar] [CrossRef]
- Davarpanah, A.M.; Rahdar, A.; Dastnae, M.A.; Zeybek, O.; Beyzaei, H. (1-x)BaFe12O19/xCoFe2O4 hard/soft magnetic nanocomposites: Synthesis, physical characterization, and antibacterial activities study. J. Mol. Struct. 2019, 1175, 445–449. [Google Scholar] [CrossRef]
- Madubuonu, N.; Aisida, S.O.; Ali, A.; Ahmad, I.; Zhao, T.; Botha, S.; Maaza, M.; Ezema, F.I. Biosynthesis of iron oxide nanoparticles via a composite of Psidium guavaja-Moringa oleifera and their antibacterial and photocatalytic study. J. Photochem. Photobiol. B Biol. 2019, 199, 111601. [Google Scholar] [CrossRef]
- Khalil, A.T.; Ovais, M.; Ullah, I.; Ali, M.; Shinwari, Z.K.; Maaza, M. Physical properties, biological applications and biocompatibility studies on biosynthesized single phase cobalt oxide (Co3O4) nanoparticles via Sageretia thea (Osbeck.). Arab. J. Chem. 2020, 13, 606–619. [Google Scholar] [CrossRef]
- Dogra, V.; Kaur, G.; Jindal, S.; Kumar, R.; Kumar, S.; Singhal, N.K. Bactericidal effects of metallosurfactants based cobalt oxide/hydroxide nanoparticles against Staphylococcus aureus. Sci. Total Environ. 2019, 681, 350–364. [Google Scholar] [CrossRef]
- Vazquez-Muñoz, R.; Meza-Villezcas, A.; Fournier, P.G.J.; Soria-Castro, E.; Juarez-Moreno, K.; Gallego-Hernández, A.L.; Bogdanchikova, N.; Vazquez-Duhalt, R.; Huerta-Saquero, A. Enhancement of antibiotics antimicrobial activity due to the silver nanoparticles impact on the cell membrane. PLoS ONE 2019, 14, e0224904. [Google Scholar] [CrossRef] [PubMed]
- Ipe, D.S.; Kumar, P.T.S.; Love, R.M.; Hamlet, S.M. Silver Nanoparticles at Biocompatible Dosage Synergistically Increases Bacterial Susceptibility to Antibiotics. Front. Microbiol. 2020, 11, 1074. [Google Scholar] [CrossRef] [PubMed]
- Thomas, R.; Jishma, P.; Snigdha, S.; Soumya, K.R.; Mathew, J.; Radhakrishnan, E.K. Enhanced antimicrobial efficacy of biosynthesized silver nanoparticle based antibiotic conjugates. Inorg. Chem. Commun. 2020, 117, 107978. [Google Scholar] [CrossRef]
- Najafi, A.; Khosravian, P.; Validi, M.; Porgham Daryasari, M.; Drees, F.; Gholipour, A. Antimicrobial action of mesoporous silica nanoparticles loaded with cefepime and meropenem separately against multidrug-resistant (MDR) Acinetobacter baumannii. J. Drug Deliv. Sci. Technol. 2021, 65, 102757. [Google Scholar] [CrossRef]
- Shaker, M.A.; Shaaban, M.I. Formulation of carbapenems loaded gold nanoparticles to combat multi-antibiotic bacterial resistance: In vitro antibacterial study. Int. J. Pharm. 2017, 525, 71–84. [Google Scholar] [CrossRef]
- Silvero C, M.J.; Rocca, D.M.; de la Villarmois, E.A.; Fournier, K.; Lanterna, A.E.; Perez, M.F.; Becerra, M.C.; Scaiano, J.C. Selective Photoinduced Antibacterial Activity of Amoxicillin-Coated Gold Nanoparticles: From One-Step Synthesis to in Vivo Cytocompatibility. ACS Omega 2018, 3, 1220–1230. [Google Scholar] [CrossRef] [PubMed]
- Rocca, D.M.; Silvero, C.M.J.; Aiassa, V.; Cecilia Becerra, M. Rapid and effective photodynamic treatment of biofilm infections using low doses of amoxicillin-coated gold nanoparticles. Photodiagnosis Photodyn. Ther. 2020, 31, 101811. [Google Scholar] [CrossRef] [PubMed]
- Fuller, M.; Whiley, H.; Köper, I. Antibiotic delivery using gold nanoparticles. SN Appl. Sci. 2020, 2, 1022. [Google Scholar] [CrossRef]
- Abdulazeem, L.; Abdalkareem Jasim, S.; Rajab, W.J. Anti-bacterial activity of gold nanoparticles against two type of antibiotic resistance pathogenic bacteria in Al-Hilla city. Mater. Today Proc. 2021, in press. [Google Scholar] [CrossRef]
- Youssef, F.; Farghaly, U.; Abd El-Baky, R.M.; Waly, N. Comparative Study of Antibacterial Effects of Titanium Dioxide Nanoparticles Alone and in Combination with Antibiotics on MDR Pseudomonas aeruginosa Strains. Int. J. Nanomed. 2020, 15, 3393–3404. [Google Scholar] [CrossRef]
| Priority | Pathogenic Bacteria | Antibiotics for Which There is Resistance |
|---|---|---|
| Critical | Acinetobacter baumannii | Carbapenem |
| Pseudomonas aeruginosa | ||
| Enterobacteriaceae | ||
| Mycobacteria | Carbapenem and 3rd generation cephalosporins | |
| Mycobacterium tuberculosis | 3rd generation cephalosporins | |
| High | Enterococcus faecium | Vancomycin and methicillin |
| Staphylococcus aureus | ||
| Helicobacter pylori | Vancomycin | |
| Campylobacter | Clarithromycin | |
| Salmonella spp. | Fluoroquinolones | |
| Neisseria gonorrhoeae | 3rd generation fluoroquinolone | |
| Medium | Streptococcus pneumoniae | |
| Haemophilus influenza | Non-sensible to penicillin | |
| Shigella spp. | Ampicillin and fluroquinolones |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Balderrama-González, A.-S.; Piñón-Castillo, H.-A.; Ramírez-Valdespino, C.-A.; Landeros-Martínez, L.-L.; Orrantia-Borunda, E.; Esparza-Ponce, H.-E. Antimicrobial Resistance and Inorganic Nanoparticles. Int. J. Mol. Sci. 2021, 22, 12890. https://doi.org/10.3390/ijms222312890
Balderrama-González A-S, Piñón-Castillo H-A, Ramírez-Valdespino C-A, Landeros-Martínez L-L, Orrantia-Borunda E, Esparza-Ponce H-E. Antimicrobial Resistance and Inorganic Nanoparticles. International Journal of Molecular Sciences. 2021; 22(23):12890. https://doi.org/10.3390/ijms222312890
Chicago/Turabian StyleBalderrama-González, Andrea-Sarahí, Hilda-Amelia Piñón-Castillo, Claudia-Adriana Ramírez-Valdespino, Linda-Lucila Landeros-Martínez, Erasmo Orrantia-Borunda, and Hilda-Esperanza Esparza-Ponce. 2021. "Antimicrobial Resistance and Inorganic Nanoparticles" International Journal of Molecular Sciences 22, no. 23: 12890. https://doi.org/10.3390/ijms222312890
APA StyleBalderrama-González, A.-S., Piñón-Castillo, H.-A., Ramírez-Valdespino, C.-A., Landeros-Martínez, L.-L., Orrantia-Borunda, E., & Esparza-Ponce, H.-E. (2021). Antimicrobial Resistance and Inorganic Nanoparticles. International Journal of Molecular Sciences, 22(23), 12890. https://doi.org/10.3390/ijms222312890








