Comprehensive Comparative Analysis of the GATA Transcription Factors in Four Rosaceae Species and Phytohormonal Response in Chinese Pear (Pyrus bretschneideri) Fruit
Abstract
:1. Introduction
2. Results
2.1. Identification and Classification of GATA Gene Family in Four Rosaceae Species
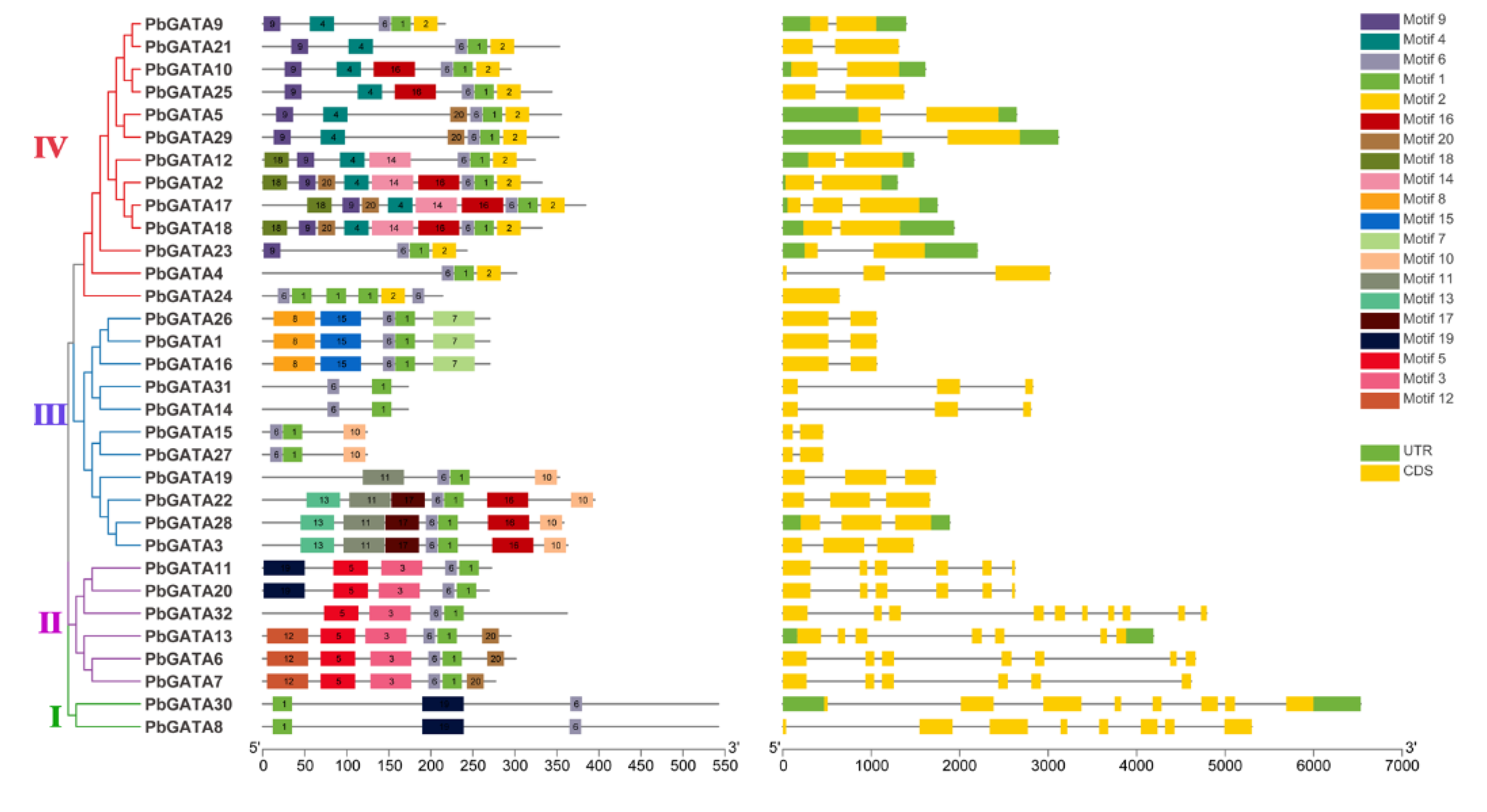
2.2. Gene Structure and Conserved Motifs of GATA Gene Family
2.3. GO Annotation Analysis
2.4. Chromosomal Distribution and Gene Duplication Events and Ka/Ks Analysis
2.5. Collinearity Relationships
2.6. Activity Analysis of PbGATA Promoter
2.7. Transcriptomic Pattern of the PbGATA Genes
2.8. Subcellular Localization Analysis
2.9. Gene Expression Pattern by qRT-PCR of PbGATA after Hormonal Treatment
3. Discussion
4. Materials and Methods
4.1. Database Search and Physicochemical Characterization of GATA Genes
4.2. Chromosomal Localization and Introns/Exons Analysis
4.3. Collinearity Relationships, Mode of Gene Duplications, and Synonymous (ks) and Non-Synonymous (ka) Substitution
4.4. Conserved Motifs and Promoter Sequence Analysis
4.5. In Silico Expression Analysis of Annotated PbGATA Genes
4.6. Comparative Phylogenetic Analysis
4.7. Plant Materials and Stress Treatment
4.8. Subcellular Localization of PbGATA Protein
4.9. Isolation of Total RNA and Quantitative Real-Time PCR
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Krishna, S.S.; Majumdar, I.; Grishin, N.V. SURVEY AND SUMMARY: Structural classification of zinc fingers. Nucleic Acids Res. 2003, 31, 532. [Google Scholar] [CrossRef] [Green Version]
- Nakashima, K.; Takasaki, H.; Mizoi, J.; Shinozaki, K.; Yamaguchi-Shinozaki, K. NAC transcription factors in plant abiotic stress responses. Biochim. Biophys. Acta BBA-Gene Regul. Mech. 2012, 1819, 97–103. [Google Scholar] [CrossRef]
- Ambawat, S.; Sharma, P.; Yadav, N.R.; Yadav, R.C. MYB transcription factor genes as regulators for plant responses: An overview. Physiol. Mol. Biol. Plants 2013, 19, 307–321. [Google Scholar] [CrossRef] [Green Version]
- Alves, M.S.; Dadalto, S.P.; Gonçalves, A.B.; De Souza, G.B.; Barros, V.A.; Fietto, L.G. Plant bZIP transcription factors responsive to pathogens: A review. Int. J. Mol. Sci. 2013, 14, 7815–7828. [Google Scholar] [CrossRef] [Green Version]
- Elrod-Erickson, M.; Pabo, C.O. Binding Studies with Mutants of Zif268: Contribution of individual side chains to binding affinity and specificity in the Zif268 zinc finger-DNA complex. J. Biol. Chem. 1999, 274, 19281–19285. [Google Scholar] [CrossRef] [Green Version]
- Merika, M.; Orkin, S.H. DNA-binding specificity of GATA family transcription factors. Mol. Cell. Biol. 1993, 13, 3999–4010. [Google Scholar] [PubMed] [Green Version]
- Behringer, C.; Schwechheimer, C. B-GATA transcription factors–insights into their structure, regulation, and role in plant development. Front. Plant Sci. 2015, 6, 90. [Google Scholar] [CrossRef] [Green Version]
- Pikkarainen, S.; Tokola, H.; Kerkelä, R.; Ruskoaho, H. GATA transcription factors in the developing and adult heart. Cardiovasc. Res. 2004, 63, 196–207. [Google Scholar] [CrossRef] [PubMed]
- Tong, Q.; Dalgin, G.; Xu, H.; Ting, C.-N.; Leiden, J.M.; Hotamisligil, G.S. Function of GATA transcription factors in preadipocyte-adipocyte transition. Science 2000, 290, 134–138. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Scazzocchio, C. The fungal GATA factors. Curr. Opin. Microbiol. 2000, 3, 126–131. [Google Scholar] [CrossRef]
- Xu, X.; Kim, S.K. The GATA transcription factor egl-27 delays aging by promoting stress resistance in Caenorhabditis elegans. PLoS Genet. 2012, 8, e1003108. [Google Scholar] [CrossRef] [Green Version]
- Crespo, J.L.; Daicho, K.; Ushimaru, T.; Hall, M.N. The GATA transcription factors GLN3 and GAT1 link TOR to salt stress in Saccharomyces cerevisiae. J. Biol. Chem. 2001, 276, 34441–34444. [Google Scholar] [CrossRef] [Green Version]
- Chi, Z.; Wang, X.-X.; Geng, Q.; Chi, Z.-M. Role of a GATA-type transcriptional repressor Sre1 in regulation of siderophore biosynthesis in the marine-derived Aureobasidium pullulans HN6. 2. Biometals 2013, 26, 955–967. [Google Scholar] [CrossRef]
- García-Salcedo, R.; Casamayor, A.; Ruiz, A.; González, A.; Prista, C.; Loureiro-Dias, M.C.; Ramos, J.; Ariño, J. Heterologous expression implicates a GATA factor in regulation of nitrogen metabolic genes and ion homeostasis in the halotolerant yeast Debaryomyces hansenii. Eukaryot. Cell 2006, 5, 1388–1398. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Daniel-Vedele, F.; Caboche, M. A tobacco cDNA clone encoding a GATA-1 zinc finger protein homologous to regulators of nitrogen metabolism in fungi. Mol. Gen. Genet. MGG 1993, 240, 365–373. [Google Scholar] [CrossRef]
- Zhang, C.; Hou, Y.; Hao, Q.; Chen, H.; Chen, L.; Yuan, S.; Shan, Z.; Zhang, X.; Yang, Z.; Qiu, D. Genome-wide survey of the soybean GATA transcription factor gene family and expression analysis under low nitrogen stress. PLoS ONE 2015, 10, e0125174. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chiang, Y.-H.; Zubo, Y.O.; Tapken, W.; Kim, H.J.; Lavanway, A.M.; Howard, L.; Pilon, M.; Kieber, J.J.; Schaller, G.E. Functional characterization of the GATA transcription factors GNC and CGA1 reveals their key role in chloroplast development, growth, and division in Arabidopsis. Plant Physiol. 2012, 160, 332–348. [Google Scholar] [CrossRef] [Green Version]
- Kobayashi, K.; Masuda, T. Transcriptional regulation of tetrapyrrole biosynthesis in Arabidopsis thaliana. Front. Plant Sci. 2016, 7, 1811. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kobayashi, K.; Ohnishi, A.; Sasaki, D.; Fujii, S.; Iwase, A.; Sugimoto, K.; Masuda, T.; Wada, H. Shoot removal induces chloroplast development in roots via cytokinin signaling. Plant Physiol. 2017, 173, 2340–2355. [Google Scholar] [CrossRef]
- Luo, X.-M.; Lin, W.-H.; Zhu, S.; Zhu, J.-Y.; Sun, Y.; Fan, X.-Y.; Cheng, M.; Hao, Y.; Oh, E.; Tian, M. Integration of light-and brassinosteroid-signaling pathways by a GATA transcription factor in Arabidopsis. Dev. Cell 2010, 19, 872–883. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Richter, R.; Behringer, C.; Zourelidou, M.; Schwechheimer, C. Convergence of auxin and gibberellin signaling on the regulation of the GATA transcription factors GNC and GNL in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 2013, 110, 13192–13197. [Google Scholar] [CrossRef] [Green Version]
- Qiao, X.; Li, M.; Li, L.; Yin, H.; Wu, J.; Zhang, S. Genome-wide identification and comparative analysis of the heat shock transcription factor family in Chinese white pear (Pyrus bretschneideri) and five other Rosaceae species. BMC Plant Biol. 2015, 15, 1–16. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yadav, S.; Modi, P.; Dave, A.; Vijapura, A.; Patel, D.; Patel, M. Effect of abiotic stress on crops. In Sustainable Crop Production; Intechopen: London, UK, 2020. [Google Scholar] [CrossRef]
- Cheng, X.; Li, M.; Li, D.; Zhang, J.; Jin, Q.; Sheng, L.; Cai, Y.; Lin, Y. Characterization and analysis of CCR and CAD gene families at the whole-genome level for lignin synthesis of stone cells in pear (Pyrus bretschneideri) fruit. Biol. Open 2017, 6, 1602–1613. [Google Scholar] [CrossRef] [Green Version]
- Su, X.; Zhao, Y.; Wang, H.; Li, G.; Cheng, X.; Jin, Q.; Cai, Y. Transcriptomic analysis of early fruit development in Chinese white pear (Pyrus bretschneideri Rehd.) and functional identification of PbCCR1 in lignin biosynthesis. BMC Plant Biol. 2019, 19, 1–19. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Finn, R.D.; Bateman, A.; Clements, J.; Coggill, P.; Eberhardt, R.Y.; Eddy, S.R.; Heger, A.; Hetherington, K.; Holm, L.; Mistry, J. Pfam: The protein families database. Nucleic Acids Res. 2014, 42, D222–D230. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mistry, J.; Finn, R.D.; Eddy, S.R.; Bateman, A.; Punta, M. Challenges in homology search: HMMER3 and convergent evolution of coiled-coil regions. Nucleic Acids Res. 2013, 41, e121. [Google Scholar] [CrossRef] [Green Version]
- Jin, J.; Zhang, H.; Kong, L.; Gao, G.; Luo, J. PlantTFDB 3.0: A portal for the functional and evolutionary study of plant transcription factors. Nucleic Acids Res. 2014, 42, D1182–D1187. [Google Scholar] [CrossRef] [Green Version]
- Punta, M.; Coggill, P.C.; Eberhardt, R.Y.; Mistry, J.; Tate, J.; Boursnell, C.; Pang, N.; Forslund, K.; Ceric, G.; Clements, J. The Pfam protein families database. Nucleic Acids Res. 2012, 40, D290–D301. [Google Scholar] [CrossRef]
- Letunic, I.; Doerks, T.; Bork, P. SMART 7: Recent updates to the protein domain annotation resource. Nucleic Acids Res. 2012, 40, D302–D305. [Google Scholar] [CrossRef]
- Bi, Y.M.; Zhang, Y.; Signorelli, T.; Zhao, R.; Zhu, T.; Rothstein, S. Genetic analysis of Arabidopsis GATA transcription factor gene family reveals a nitrate-inducible member important for chlorophyll synthesis and glucose sensitivity. Plant J. 2005, 44, 680–692. [Google Scholar] [CrossRef]
- Bailey, T.L.; Johnson, J.; Grant, C.E.; Noble, W.S. The MEME suite. Nucleic Acids Res. 2015, 43, W39–W49. [Google Scholar] [CrossRef] [Green Version]
- Cao, Y.; Han, Y.; Li, D.; Lin, Y.; Cai, Y. MYB transcription factors in chinese pear (Pyrus bretschneideri Rehd.): Genome-wide identification, classification, and expression profiling during fruit development. Front. Plant Sci. 2016, 7, 577. [Google Scholar] [CrossRef] [Green Version]
- Cheng, X.; Li, G.; Muhammad, A.; Zhang, J.; Jiang, T.; Jin, Q.; Zhao, H.; Cai, Y.; Lin, Y. Molecular identification, phylogenomic characterization and expression patterns analysis of the LIM (LIN-11, Isl1 and MEC-3 domains) gene family in pear (Pyrus bretschneideri) reveal its potential role in lignin metabolism. Gene 2019, 686, 237–249. [Google Scholar] [CrossRef]
- Abdullah, M.; Cheng, X.; Cao, Y.; Su, X.; Manzoor, M.A.; Gao, J.; Cai, Y.; Lin, Y. Zinc finger-homeodomain transcriptional factors (ZHDs) in upland cotton (Gossypium hirsutum): Genome-wide identification and expression analysis in fiber development. Front. Genet. 2018, 9, 357. [Google Scholar] [CrossRef]
- Zhang, C.; Huang, Y.; Xiao, Z.; Yang, H.; Hao, Q.; Yuan, S.; Chen, H.; Chen, L.; Chen, S.; Zhou, X. A GATA transcription factor from soybean (Glycine max) regulates chlorophyll biosynthesis and suppresses growth in the transgenic Arabidopsis thaliana. Plants 2020, 9, 1036. [Google Scholar] [CrossRef]
- Peng, W.; Li, W.; Song, N.; Tang, Z.; Liu, J.; Wang, Y.; Pan, S.; Dai, L.; Wang, B. Genome-Wide Characterization, Evolution, and Expression Profile Analysis of GATA Transcription Factors in Brachypodium distachyon. Int. J. Mol. Sci. 2021, 22, 2026. [Google Scholar] [CrossRef] [PubMed]
- Jiang, C.; Lv, G.; Ge, J.; He, B.; Zhang, Z.; Hu, Z.; Zeng, B. Genome-wide identification of the GATA transcription factor family and their expression patterns under temperature and salt stress in Aspergillus oryzae. AMB Express 2021, 11, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Kim, M.; Xi, H.; Park, J. Genome-wide comparative analyses of GATA transcription factors among 19 Arabidopsis ecotype genomes: Intraspecific characteristics of GATA transcription factors. PLoS ONE 2021, 16, e0252181. [Google Scholar]
- Reyes, J.C.; Muro-Pastor, M.I.; Florencio, F.J. The GATA family of transcription factors in Arabidopsis and rice. Plant Physiol. 2004, 134, 1718–1732. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, Y.; Gao, M.; Singer, S.D.; Fei, Z.; Wang, H.; Wang, X. Genome-wide identification and analysis of the TIFY gene family in grape. PLoS ONE 2012, 7, e44465. [Google Scholar]
- Vision, T.J.; Brown, D.G.; Tanksley, S.D. The origins of genomic duplications in Arabidopsis. Science 2000, 290, 2114–2117. [Google Scholar] [CrossRef] [Green Version]
- Yang, S.; Zhang, X.; Yue, J.-X.; Tian, D.; Chen, J.-Q. Recent duplications dominate NBS-encoding gene expansion in two woody species. Mol. Genet. Genom. 2008, 280, 187–198. [Google Scholar] [CrossRef]
- Nawaz, M.A.; Rehman, H.M.; Baloch, F.S.; Ijaz, B.; Ali, M.A.; Khan, I.A.; Lee, J.D.; Chung, G.; Yang, S.H. Genome and transcriptome-wide analyses of cellulose synthase gene superfamily in soybean. J. Plant Physiol. 2017, 215, 163–175. [Google Scholar] [CrossRef]
- Soliman, E.R.; Meyer, P. Responsiveness and adaptation to salt stress of the redox-responsive transcription factor 1 (RRTF1) gene are controlled by its promoter. Mol. Biotechnol. 2019, 61, 254–260. [Google Scholar] [CrossRef]
- Li, G.; Liu, X.; Liang, Y.; Zhang, Y.; Cheng, X.; Cai, Y. Genome-wide characterization of the cellulose synthase gene superfamily in Pyrus bretschneideri and reveal its potential role in stone cell formation. Funct. Integr. Genom. 2020, 20, 723–738. [Google Scholar] [CrossRef]
- Lynch, M. Intron evolution as a population-genetic process. Proc. Natl. Acad. Sci. USA 2002, 99, 6118–6123. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Budak, H.; Zhang, B. MicroRNAs in Model and Complex Organisms; Springer: Berlin/Heidelberg, Germany, 2017. [Google Scholar]
- Thomas, J.; Bowman, M.J.; Vega, A.; Kim, H.R.; Mukherjee, A. Comparative transcriptome analysis provides key insights into gene expression pattern during the formation of nodule-like structures in Brachypodium. Funct. Integr. Genom. 2018, 18, 315–326. [Google Scholar] [CrossRef]
- Zhang, Y.; Li, Q.; Xu, L.; Qiao, X.; Liu, C.; Zhang, S. Comparative analysis of the P-type ATPase gene family in seven Rosaceae species and an expression analysis in pear (Pyrus bretschneideri Rehd.). Genomics 2020, 112, 2550–2563. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.-L.; Zhang, X.-N.; Lu, G.-L.; Wang, C.-R.; Wang, R. Regulation of gibberellin on gene expressions related with the lignin biosynthesis in ‘Wangkumbae’pear (Pyrus pyrifolia Nakai) fruit. Plant Growth Regul. 2015, 76, 127–134. [Google Scholar] [CrossRef]
- Zhang, Z.; Ren, C.; Zou, L.; Wang, Y.; Li, S.; Liang, Z. Characterization of the GATA gene family in Vitis vinifera: Genome-wide analysis, expression profiles, and involvement in light and phytohormone response. Genome 2018, 61, 713–723. [Google Scholar] [CrossRef] [Green Version]
- Shaikhali, J.; de Dios Barajas-Lopéz, J.; Ötvös, K.; Kremnev, D.; Garcia, A.S.; Srivastava, V.; Wingsle, G.; Bako, L.; Strand, Å. The CRYPTOCHROME1-dependent response to excess light is mediated through the transcriptional activators ZINC FINGER PROTEIN EXPRESSED IN INFLORESCENCE MERISTEM LIKE1 and ZML2 in Arabidopsis. Plant Cell 2012, 24, 3009–3025. [Google Scholar] [CrossRef] [Green Version]
- Ye, J.; Hu, T.; Yang, C.; Li, H.; Yang, M.; Ijaz, R.; Ye, Z.; Zhang, Y. Transcriptome profiling of tomato fruit development reveals transcription factors associated with ascorbic acid, carotenoid and flavonoid biosynthesis. PLoS ONE 2015, 10, e0130885. [Google Scholar] [CrossRef]
- Wu, J.; Fu, L.; Yi, H. Genome-wide identification of the transcription factors involved in citrus fruit ripening from the transcriptomes of a late-ripening sweet orange mutant and its wild type. PLoS ONE 2016, 11, e0154330. [Google Scholar] [CrossRef]
- Zhu, W.; Guo, Y.; Chen, Y.; Wu, D.; Jiang, L. Genome-wide identification, phylogenetic and expression pattern analysis of GATA family genes in Brassica napus. BMC Plant Biol. 2020, 20, 1–12. [Google Scholar] [CrossRef]
- Manzoor, M.A.; Guohui, L.; Muhammad, A.; Han, W.; Wenlong, H.; Yang, Z.; Xinya, W.; Yu, Z.; Xiaofeng, F.; Qing, J. Genome-wide investigation and comparative analysis of MATE gene family in Rosaceae species and their regulatory role in abiotic stress responses in Chinese pear (Pyrus bretschneideri). Physiol. Plant. 2021, 173, 1163–1178. [Google Scholar] [CrossRef] [PubMed]
- Manzoor, M.A.; Cheng, X.; Li, G.; Su, X.; Abdullah, M.; Cai, Y. Gene structure, evolution and expression analysis of the P-ATPase gene family in Chinese pear (Pyrus bretschneideri). Comput. Biol. Chem. 2020, 88, 107346. [Google Scholar] [CrossRef]
- Marchler-Bauer, A.; Derbyshire, M.K.; Gonzales, N.R.; Lu, S.; Chitsaz, F.; Geer, L.Y.; Geer, R.C.; He, J.; Gwadz, M.; Hurwitz, D.I. CDD: NCBI’s conserved domain database. Nucleic Acids Res. 2015, 43, D222–D226. [Google Scholar] [CrossRef] [Green Version]
- Artimo, P.; Jonnalagedda, M.; Arnold, K.; Baratin, D.; Csardi, G.; De Castro, E.; Duvaud, S.; Flegel, V.; Fortier, A.; Gasteiger, E. ExPASy: SIB bioinformatics resource portal. Nucleic Acids Res. 2012, 40, W597–W603. [Google Scholar] [CrossRef] [PubMed]
- Carlevaro-Fita, J.; Lanzós, A.; Feuerbach, L.; Hong, C.; Mas-Ponte, D.; Pedersen, J.S.; Johnson, R. Cancer LncRNA Census reveals evidence for deep functional conservation of long noncoding RNAs in tumorigenesis. Commun. Biol. 2020, 3, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Hu, B.; Jin, J.; Guo, A.-Y.; Zhang, H.; Luo, J.; Gao, G. GSDS 2.0: An upgraded gene feature visualization server. Bioinformatics 2015, 31, 1296–1297. [Google Scholar] [CrossRef] [Green Version]
- Wang, Y.; Tang, H.; DeBarry, J.D.; Tan, X.; Li, J.; Wang, X.; Lee, T.-h.; Jin, H.; Marler, B.; Guo, H. MCScanX: A toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 2012, 40, e49. [Google Scholar] [CrossRef] [Green Version]
- Hurst, L.D. The Ka/Ks ratio: Diagnosing the form of sequence evolution. Trends Genet. TIG 2002, 18, 486. [Google Scholar] [CrossRef]
- Gaut, B.S.; Morton, B.R.; McCaig, B.C.; Clegg, M.T. Substitution rate comparisons between grasses and palms: Synonymous rate differences at the nuclear gene Adh parallel rate differences at the plastid gene rbcL. Proc. Natl. Acad. Sci. USA 1996, 93, 10274–10279. [Google Scholar] [CrossRef] [Green Version]
- Letunic, I.; Bork, P. Interactive tree of life (iTOL) v3: An online tool for the display and annotation of phylogenetic and other trees. Nucleic Acids Res. 2016, 44, W242–W245. [Google Scholar] [CrossRef]
- Yang, S.; Xiong, X.; Arif, S.; Gao, L.; Zhao, L.; Shah, I.H.; Zhang, Y. A calmodulin-like CmCML13 from Cucumis melo improved transgenic Arabidopsis salt tolerance through reduced shoot’s Na+, and also improved drought resistance. Plant Physiol. Biochem. 2020, 155, 271–283. [Google Scholar] [CrossRef] [PubMed]
- Chun, Y.; Fang, J.; Zafar, S.A.; Shang, J.; Zhao, J.; Yuan, S.; Li, X. MINI SEED 2 (MIS2) encodes a receptor-like kinase that controls grain size and shape in rice. Rice 2020, 13, 7. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2− ΔΔCT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]












Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Manzoor, M.A.; Sabir, I.A.; Shah, I.H.; Wang, H.; Yu, Z.; Rasool, F.; Mazhar, M.Z.; Younas, S.; Abdullah, M.; Cai, Y. Comprehensive Comparative Analysis of the GATA Transcription Factors in Four Rosaceae Species and Phytohormonal Response in Chinese Pear (Pyrus bretschneideri) Fruit. Int. J. Mol. Sci. 2021, 22, 12492. https://doi.org/10.3390/ijms222212492
Manzoor MA, Sabir IA, Shah IH, Wang H, Yu Z, Rasool F, Mazhar MZ, Younas S, Abdullah M, Cai Y. Comprehensive Comparative Analysis of the GATA Transcription Factors in Four Rosaceae Species and Phytohormonal Response in Chinese Pear (Pyrus bretschneideri) Fruit. International Journal of Molecular Sciences. 2021; 22(22):12492. https://doi.org/10.3390/ijms222212492
Chicago/Turabian StyleManzoor, Muhammad Aamir, Irfan Ali Sabir, Iftikhar Hussain Shah, Han Wang, Zhao Yu, Faiz Rasool, Muhammad Zaid Mazhar, Shoaib Younas, Muhammad Abdullah, and Yongping Cai. 2021. "Comprehensive Comparative Analysis of the GATA Transcription Factors in Four Rosaceae Species and Phytohormonal Response in Chinese Pear (Pyrus bretschneideri) Fruit" International Journal of Molecular Sciences 22, no. 22: 12492. https://doi.org/10.3390/ijms222212492
APA StyleManzoor, M. A., Sabir, I. A., Shah, I. H., Wang, H., Yu, Z., Rasool, F., Mazhar, M. Z., Younas, S., Abdullah, M., & Cai, Y. (2021). Comprehensive Comparative Analysis of the GATA Transcription Factors in Four Rosaceae Species and Phytohormonal Response in Chinese Pear (Pyrus bretschneideri) Fruit. International Journal of Molecular Sciences, 22(22), 12492. https://doi.org/10.3390/ijms222212492





