Diversity and Taxonomic Distribution of Endophytic Bacterial Community in the Rice Plant and Its Prospective
Abstract
:1. Introduction
2. Endophytic Bacterial Diversity in the Rice Plant
3. Factors Involving in Endophytic Bacterial Communities’ Variations
4. Method for Isolation and Identification of Endophytic Bacteria
5. Mechanism of Plant-Growth Promotion by Endophytes
5.1. Direct Plant-Growth Promotion by Endophytes
Biological Nitrogen Fixation
5.2. Availability of Phosphate
5.3. Phytohormones
5.3.1. Auxins
5.3.2. Cytokinins
5.3.3. Gibberellins
5.3.4. Ethylene and 1-Aminocyclopropane-1-Carboxylate Deaminase
5.3.5. Abscisic Acid
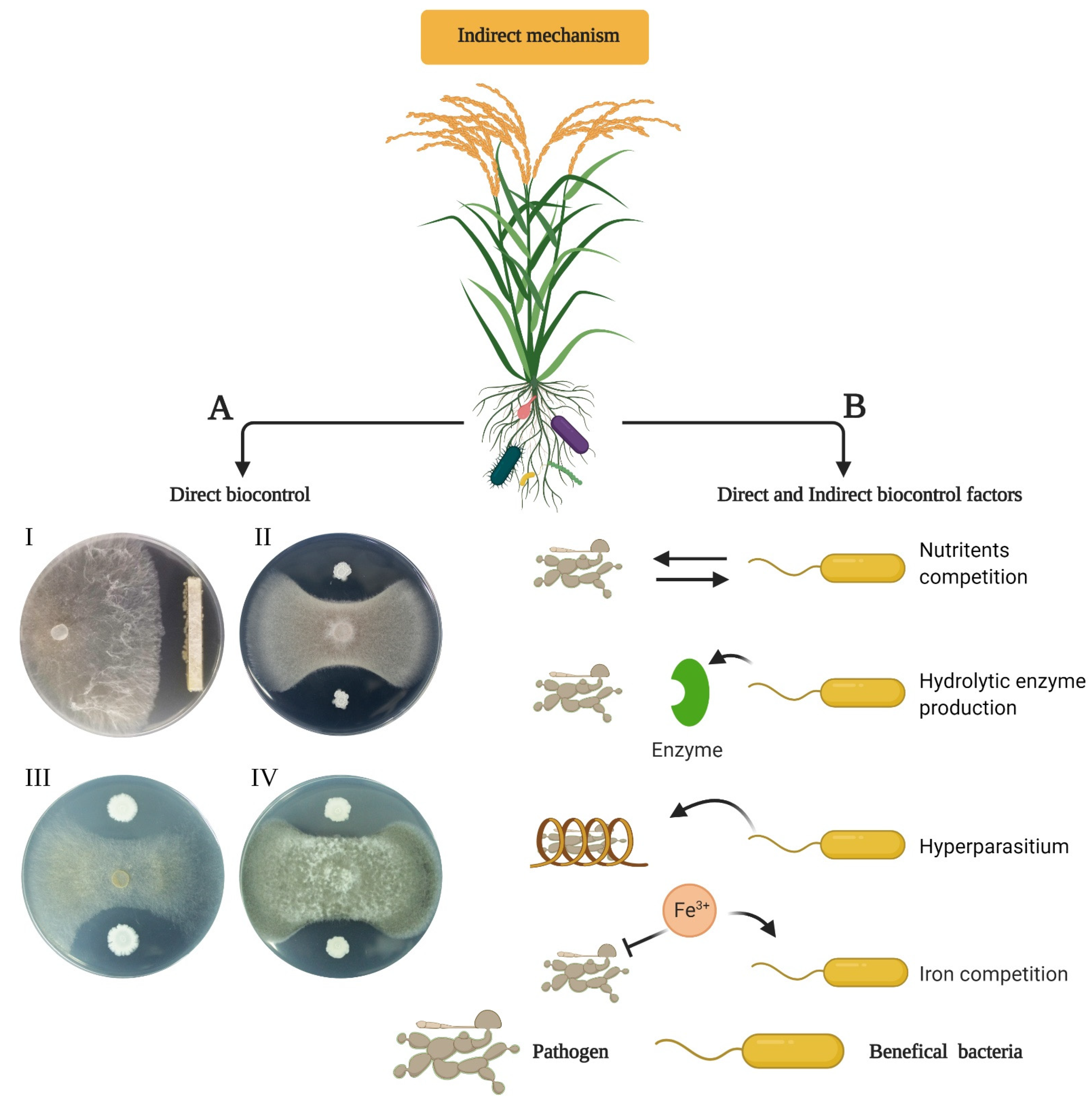
6. Indirect Plant-Growth Promotion by Endophytes
6.1. Biocontrol of Plant Pathogens by Endophytic Bacteria
6.2. Antibiosis
6.3. Signal Interference
6.4. Predation and Parasitism
6.5. Induced Systemic Resistance
7. Implementation of Bioinformatics in Endophytes Diversity Analysis
8. Endophytes-Induced Bioactive Compounds
9. Conclusions and Prospects
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Malfanova, N.V. Endophytic Bacteria with Plant Growth Promoting and Biocontrol Abilities. Ph.D. Thesis, Leiden University, Leiden, The Netherlands, 2013. [Google Scholar]
- Mundt, J.O.; Hinkle, N.F. Bacteria within ovules and seeds. Appl. Environ. Microbiol. 1976, 32, 694–698. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sessitsch, A.; Hardoim, P.; Döring, J.; Weilharter, A.; Krause, A.; Woyke, T.; Mitter, B.; Hauberg-Lotte, L.; Friedrich, F.; Rahalkar, M. Functional characteristics of an endophyte community colonizing rice roots as revealed by metagenomic analysis. Mol. Plant-Microbe Interact. 2012, 25, 28–36. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Krause, A.; Julich, H.; Mankar, M.; Reinhold-Hurek, B. The Regulatory Network controlling ethanol-induced expression of alcohol dehydrogenase in the endophyte Azoarcus sp. Strain BH72. Mol. Plant-Microbe Interact. 2017, 30, 778–785. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ahmad, Z.; Wu, J.; Chen, L.; Dong, W. Isolated Bacillus subtilis strain 330-2 and its antagonistic genes identified by the removing PCR. Sci. Rep. 2017, 7, 1–13. [Google Scholar]
- Ludueña, L.M.; Anzuay, M.S.; Angelini, J.G.; McIntosh, M.; Becker, A.; Rupp, O.; Goesmann, A.; Blom, J.; Fabra, A.; Taurian, T.J.G. Genome sequence of the endophytic strain Enterobacter sp. J49, a potential biofertilizer for peanut and maize. Genomics 2019, 111, 913–920. [Google Scholar] [CrossRef] [PubMed]
- Kandel, S.L.; Joubert, P.M.; Doty, S.L. Bacterial Endophyte colonization and distribution within plants. Microorganisms 2017, 5, 77. [Google Scholar] [CrossRef] [Green Version]
- Andreozzi, A.; Prieto, P.; Mercado-Blanco, J.; Monaco, S.; Zampieri, E.; Romano, S.; Valè, G.; Defez, R.; Bianco, C.J.E.M. Efficient colonization of the endophytes Herbaspirillum huttiense RCA24 and Enterobacter cloacae RCA25 influences the physiological parameters of Oryza sativa L. cv. Baldo rice. Environ. Microbiol. 2019, 21, 3489–3504. [Google Scholar] [CrossRef]
- Tavares, M.J.; Nascimento, F.X.; Glick, B.R.; Rossi, M.J. The expression of an exogenous ACC deaminase by the endophyte Serratia grimesii BXF1 promotes the early nodulation and growth of common bean. Lett. Appl. Microbiol. 2018, 66, 252–259. [Google Scholar] [CrossRef]
- Passari, A.; Upadhyaya, K.; Singh, G.; Abdel-Azeem, A.; Thankappan, S.; Uthandi, S.; Hashem, A.; Allah, E.F.A.; Malik, J.A.; As, A.; et al. Enhancement of disease resistance, growth potential, and photosynthesis in tomato (Solanum lycopersicum) by inoculation with an endophytic actinobacterium, Streptomyces thermocarboxydus strain BPSAC147. PLoS ONE 2019, 14, e0219014. [Google Scholar] [CrossRef] [Green Version]
- Gyaneshwar, P.; James, E.; Mathan, N.; Reddy, P.M.; Reinhold-Hurek, B.; Ladha, J.K. Endophytic Colonization of Rice by a Diazotrophic Strain of Serratia marcescens. J. Bacteriol. 2001, 183, 2634–2645. [Google Scholar] [CrossRef] [Green Version]
- Elferink, M.; Schierhorn, F. Global demand for food is rising. Can we meet it. Harv. Bus. Rev. 2016, 7, 2016. [Google Scholar]
- Vollset, S.E.; Goren, E.; Yuan, C.-W.; Cao, J.; E Smith, A.; Hsiao, T.; Bisignano, C.; Azhar, G.S.; Castro, E.; Chalek, J.; et al. Fertility, mortality, migration, and population scenarios for 195 countries and territories from 2017 to 2100: A forecasting analysis for the Global Burden of Disease Study. Lancet 2020, 396, 1285–1306. [Google Scholar] [CrossRef]
- Directive 2009/128/EC, R.E. Directive 2009/128/EC of the European Parliament and of the Council of 21 October 2009 establishing a framework for Community action to achieve the sustainable use of pesticides. Off. J. Eur. Union 2009, 309, 71–86. [Google Scholar]
- European Union Pesticides Database. Pesticides Database. Available online: http://ec.europa.eu/food/plant/protection/evaluation/database_act_subs_en.htm (accessed on 25 October 2020).
- Fang, L. Overview of Biofertilizer Registration in China. Available online: https://agrochemical.chemlinked.com/chempedia/overview-biofertilizer-registration-china (accessed on 20 October 2020).
- Montesinos, E. Development, registration and commercialization of microbial pesticides for plant protection. Int. Microbiol. 2003, 6, 245–252. [Google Scholar] [CrossRef]
- Glick, B.R.; Karaturovic, D.M.; Newell, P.C. A novel procedure for rapid isolation of plant growth promoting pseudomonads. Can. J. Microbiol. 1995, 41, 533–536. [Google Scholar] [CrossRef]
- Khare, E.; Mishra, J.; Arora, N.K. Multifaceted interactions between endophytes and plant: Developments and prospects. Front. Microbiol. 2018, 9, 2732. [Google Scholar] [CrossRef]
- Glick, B.R. Beneficial Plant-Bacterial Interactions; Springer: New York, NY, USA, 2015. [Google Scholar]
- Van Rossum, G.; Drake, F.L. PYTHON 2.6 Reference Manual; CreateSpace: Scotts Valley, CA, USA, 2009. [Google Scholar]
- Letunic, I.; Bork, P.J.N.A.R. Interactive tree of life (iTOL) v4: Recent updates and new developments. Nucleic Acids Res. 2019, 47, W256–W259. [Google Scholar] [CrossRef] [Green Version]
- Kaga, H.; Mano, H.; Tanaka, F.; Watanabe, A.; Kaneko, S.; Morisaki, H. Rice seeds as sources of endophytic bacteria. Microbes Environ. 2009, 24, 154–162. [Google Scholar] [CrossRef] [PubMed]
- Shahzad, R.; Waqas, M.; Khan, A.L.; Asaf, S.; Khan, M.A.; Kang, S.-M.; Yun, B.-W.; Lee, I.-J. Seed-borne endophytic Bacillus amyloliquefaciens RWL-1 produces gibberellins and regulates endogenous phytohormones of Oryza sativa. Plant Physiol. Biochem. 2016, 106, 236–243. [Google Scholar] [CrossRef] [PubMed]
- Khaskheli, M.A.; Wu, L.; Chen, G.; Chen, L.; Hussain, S.; Song, D.; Liu, S.; Feng, G. Isolation and Characterization of Root-Associated Bacterial Endophytes and Their Biocontrol Potential against Major Fungal Phytopathogens of Rice (Oryza sativa L.). Pathogens 2020, 9, 172. [Google Scholar] [CrossRef] [PubMed]
- Valdez-Nuñez, R.; Ríos-Ruiz, W.; Ormeño-Orrillo, E.; Torres-Chávez, E.; Torres-Delgado, J. Genetic characterization of rice endophytic bacteria (Oryza sativa L.) with antimicrobial activity against Burkholderia glumae. Rev. Argent. Microbiol. 2020, 52, 315–327. [Google Scholar]
- Nasrollahi, M.; Pourbabaei, A.A.; Etesami, H.; Talebi, K. Diazinon degradation by bacterial endophytes in rice plant (Oryzia sativa L.): A possible reason for reducing the efficiency of diazinon in the control of the rice stem–borer. Chemosphere 2020, 246, 125759. [Google Scholar] [CrossRef]
- Defez, R.; Andreozzi, A.; Bianco, C. The overproduction of indole-3-acetic acid (IAA) in endophytes upregulates nitrogen fixation in both bacterial cultures and inoculated rice plants. Microb. Ecol. 2017, 74, 441–452. [Google Scholar] [CrossRef]
- Zhao, J.; Zhao, X.; Wang, J.; Gong, Q.; Zhang, X.; Zhang, G. Isolation, Identification and Characterization of Endophytic Bacterium Rhizobium oryzihabitans sp. nov., from Rice Root with Biotechnological potential in agriculture. Microorganisms 2020, 8, 608. [Google Scholar] [CrossRef] [PubMed]
- Banik, A.; Mukhopadhaya, S.K.; Dangar, T.K. Characterization of N 2-fixing plant growth promoting endophytic and epiphytic bacterial community of Indian cultivated and wild rice (Oryza spp.) genotypes. Planta 2016, 243, 799–812. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.-J.; Zhang, J.; Sun, L.; Zhang, R.-J.; Zhang, C.-W.; Yin, H.-Q.; Zhang, X.-X. Rhizobium oryziradicis sp. nov., isolated from rice roots. Int. J. Syst. Evol. Microbiol. 2017, 67, 963–968. [Google Scholar] [CrossRef]
- Kwak, M.-J.; Song, J.Y.; Kim, S.-Y.; Jeong, H.; Kang, S.G.; Kim, B.K.; Kwon, S.-K.; Lee, C.H.; Yu, D.S.; Park, S.-H.; et al. Complete Genome Sequence of the Endophytic Bacterium Burkholderia sp. Strain KJ006. Am. Soc. Microbiol. 2012, 194, 4432–4433. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Raj, G.; Shadab, M.; Deka, S.; Das, M.; Baruah, J.; Bharali, R.; Talukdar, N.C. Seed interior microbiome of rice genotypes indigenous to three agroecosystems of Indo-Burma biodiversity hotspot. BMC Genom. 2019, 20, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Zhang, L.; Meng, A.; Zhang, J.; Xie, M.; Qin, Y.; Faulk, D.C.; Zhang, B.; Yang, S.; Qiu, L. Isolation and molecular identification of endophytic diazotrophs from seeds and stems of three cereal crops. PLoS ONE 2017, 12, e0187383. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Aljuraifani, A.; Aldosary, S.; Ababutain, I. In vitro antimicrobial activity of endophytes, isolated from Moringa peregrina growing in eastern region of Saudi Arabia. Natl. Acad. Sci. Lett. 2019, 42, 75–80. [Google Scholar] [CrossRef] [Green Version]
- Correa-Galeote, D.; Bedmar, E.J.; Arone, G. Maize endophytic bacterial diversity as affected by soil cultivation history. Front. Microbiol. 2018, 9, 484. [Google Scholar] [CrossRef]
- Marag, P.S.; Suman, A. Growth stage and tissue specific colonization of endophytic bacteria having plant growth promoting traits in hybrid and composite maize (Zea mays L.). Microbiol. Res. 2018, 214, 101–113. [Google Scholar] [CrossRef] [PubMed]
- Rania, A.B.A.; Jabnoun-Khiareddine, H.; Nefzi, A.; Mokni-Tlili, S.; Daami-Remadi, M. Endophytic bacteria from Datura metel for plant growth promotion and bioprotection against Fusarium wilt in tomato. Biocontrol Sci. Technol. 2016, 26, 1139–1165. [Google Scholar] [CrossRef]
- Zhang, Q.; Li, Y.; Xu, F.; Zheng, M.; Xi, X.; Zhang, X.; Han, C. Optimization of submerged fermentation medium for matrine production by Aspergillus terreus, an endophytic fungus harboring seeds of Sophora flavescens, using response surface methodology. Mycobiology 2017, 45, 90–96. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Proença, D.N.; Francisco, R.; Kublik, S.; Schöler, A.; Vestergaard, G.; Schloter, M.; Morais, P.V. The microbiome of endophytic, wood colonizing bacteria from pine trees as affected by pine wilt disease. Sci. Rep. 2017, 7, 1–9. [Google Scholar] [CrossRef] [Green Version]
- Pereira, G.V.D.M.; Magalhães, K.T.; Lorenzetii, E.R.; Souza, T.P.; Schwan, R.F. A Multiphasic approach for the identification of endophytic bacterial in strawberry fruit and their potential for plant growth promotion. Microb. Ecol. 2011, 63, 405–417. [Google Scholar] [CrossRef]
- Kukkurainen, S.; Leino, A.; Vähämiko, S.; Kärkkäinen, H.R.; Ahanen, K.; Sorvari, S.; Rugienius, R.; Toldi, O. Occurrence and location of endophytic bacteria in garden and wild strawberry. HortScience 2005, 40, 348–352. [Google Scholar] [CrossRef] [Green Version]
- Villalobos, S.D.L.S.; Robles, R.I.; Cota, F.I.P.; Larsen, J.; Lozano, P.; Tiedje, J.M. Bacillus cabrialesii sp. nov., an endophytic plant growth promoting bacterium isolated from wheat (Triticum turgidum subsp. durum) in the Yaqui Valley, Mexico. Int. J. Syst. Evol. Microbiol. 2019, 69, 3939–3945. [Google Scholar] [CrossRef]
- Paz, I.C.P.; Santin, R.C.M.; Guimaraes, A.M.; Rosa, O.P.P.; Dias, A.C.F.; Quecine, M.; Azevedo, J.; Matsumura, A.T.S. Eucalyptus growth promotion by endophytic Bacillus spp. Genet. Mol. Res. 2012, 11, 3711–3720. [Google Scholar] [CrossRef]
- Hazarika, D.J.; Goswami, G.; Gautom, T.; Parveen, A.; Das, P.; Barooah, M.; Boro, R.C. Lipopeptide mediated biocontrol activity of endophytic Bacillus subtilis against fungal phytopathogens. BMC Microbiol. 2019, 19, 71. [Google Scholar] [CrossRef]
- Bruisson, S.; Zufferey, M.; L’Haridon, F.; Trutmann, E.; Anand, A.; Dutartre, A.; De Vrieze, M.; Weisskopf, L. Endophytes and epiphytes from the grapevine leaf microbiome as potential biocontrol agents against phytopathogens. Front. Microbiol. 2019, 10, 2726. [Google Scholar] [CrossRef] [Green Version]
- Bell, C.; Dickie, G.A.; Harvey, W.; Chan, J. Endophytic bacteria in grapevine. Can. J. Microbiol. 2011, 41, 46–53. [Google Scholar] [CrossRef]
- Schmidt, C.; Lovecká, P.; Mrnka, L.; Vychodilová, A.; Strejček, M.; Fenclová, M.; Demnerová, K.J.M.E. Distinct communities of poplar endophytes on an unpolluted and a risk element-polluted site and their plant growth-promoting potential in vitro. Microb. Ecol. 2018, 75, 955–969. [Google Scholar] [CrossRef] [PubMed]
- Sziderics, A.H.; Rasche, F.; Trognitz, F.; Sessitsch, A.; Wilhelm, E. Bacterial endophytes contribute to abiotic stress adaptation in pepper plants (Capsicum annuum L.). Can. J. Microbiol. 2007, 53, 1195–1202. [Google Scholar] [CrossRef]
- Tuo, L.; Yan, X.R.; Li, F.N.; Bao, Y.X.; Shi, H.C.; Li, H.Y.; Sun, C.H. Brachybacterium endophyticum sp. nov., a novel endophytic actinobacterium isolated from bark of Scutellaria baicalensis Georgi. Int. J. Syst. Evol. Microbiol. 2018, 68, 3563–3568. [Google Scholar] [CrossRef]
- Shaikh, A.; Parmar, P.; Katagi, R.; Patel, D.; Desai, H.R.; Solanki, B.G. Bioprospecting Potential of Endophytic Bacteria from Leaves of Gossypium hirsutum. Int. J. Curr. Microbiol. Appl. Sci. 2017, 6, 1718–1730. [Google Scholar] [CrossRef]
- Boiu-sicuia, O.-A.; Constantinescu, F.; Cornea, C.P.J.S.B. Selection and characterization of new endophytic bacterial strains isolated from potato tuber useful in biocontrol strategies. Sci. Bull. 2017, 21, 23–28. [Google Scholar]
- Mahmood, A.; Takagi, K.; Ito, K.; Kataoka, R. Changes in endophytic bacterial communities during different growth stages of cucumber (Cucumis sativus L.). World J. Microbiol. Biotechnol. 2019, 35, 104. [Google Scholar] [CrossRef] [PubMed]
- Abdallah, R.A.B.; Mokni-Tlili, S.; Nefzi, A.; Jabnoun-Khiareddine, H.; Daami-Remadi, M. Biocontrol of Fusarium wilt and growth promotion of tomato plants using endophytic bacteria isolated from Nicotiana glauca organs. Biol. Control 2016, 97, 80–88. [Google Scholar] [CrossRef]
- Wang, X.; Liang, G. Control efficacy of an endophytic Bacillus amyloliquefaciens strain BZ6-1 against peanut bacterial wilt, Ralstonia solanacearum. BioMed Res. Int. 2014, 2014, 1–11. [Google Scholar]
- Murugappan, R.; Begum, S.B.; Roobia, R.R. Symbiotic influence of endophytic Bacillus pumilus on growth promotion and probiotic potential of the medicinal plant Ocimum sanctum. Symbiosis 2013, 60, 91–99. [Google Scholar] [CrossRef]
- Tariq, M.; Hameed, S.; Yasmeen, T.; Zahid, M.; Zafar, M. Molecular characterization and identification of plant growth promoting endophytic bacteria isolated from the root nodules of pea (Pisum sativum L.). World J. Microbiol. Biotechnol. 2014, 30, 719–725. [Google Scholar] [CrossRef]
- Molina, M.A.; Ramos, J.L.; Espinosa-Urgel, M.J.E.M. A two-partner secretion system is involved in seed and root colonization and iron uptake by Pseudomonas putida KT2440. Environ. Microbiol. 2006, 8, 639–647. [Google Scholar] [CrossRef]
- Borah, M.; Das, S.; Boruah, H.; Boro, R.; Barooah, M.J.B. Diversity of culturable endophytic bacteria from wild and cultivated rice showed potential plant growth promoting activities. bioRxiv 2018. [Google Scholar] [CrossRef] [Green Version]
- Zhou, J.; Li, P.; Meng, D.; Gu, Y.; Zheng, Z.; Yin, H.; Zhou, Q.; Li, J. Isolation, characterization and inoculation of Cd tolerant rice endophytes and their impacts on rice under Cd contaminated environment. Environ. Pollut. 2020, 260, 113990. [Google Scholar] [CrossRef] [PubMed]
- Walitang, D.I.; Kim, C.G.; Jeon, S.; Kang, Y.; Sa, T. Conservation and transmission of seed bacterial endophytes across generations following crossbreeding and repeated inbreeding of rice at different geographic locations. MicrobiologyOpen 2019, 8, e00662. [Google Scholar] [CrossRef] [PubMed]
- Hardoim, P.; Hardoim, C.; Van Overbeek, L.S.; Van Elsas, J.D. Dynamics of Seed-Borne Rice Endophytes on Early Plant Growth Stages. PLoS ONE 2012, 7, e30438. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mitter, B.; Pfaffenbichler, N.; Flavell, R.; Compant, S.; Antonielli, L.; Petric, A.; Berninger, T.; Naveed, M.; Sheibani-Tezerji, R.; von Maltzahn, G.; et al. A New Approach to modify plant microbiomes and traits by introducing beneficial bacteria at flowering into progeny seeds. Front. Microbiol. 2017, 8, 11. [Google Scholar] [CrossRef] [Green Version]
- Manirajan, B.A.; Ratering, S.; Rusch, V.; Schwiertz, A.; Geissler-Plaum, R.; Cardinale, M.; Schnell, S. Bacterial microbiota associated with flower pollen is influenced by pollination type, and shows a high degree of diversity and species-specificity. Environ. Microbiol. 2016, 18, 5161–5174. [Google Scholar] [CrossRef]
- Agarwal, V.K.; Sinclair, J.B. Principles of Seed Pathology; CRC Press: Boca Raton, FL, USA, 1996. [Google Scholar]
- Pfeiffer, B.; Fender, A.-C.; Lasota, S.; Hertel, D.; Jungkunst, H.F.; Daniel, R. Leaf litter is the main driver for changes in bacterial community structures in the rhizosphere of ash and beech. Appl. Soil Ecol. 2013, 72, 150–160. [Google Scholar] [CrossRef]
- Breidenbach, B.; Pump, J.; Dumont, M.G. Microbial community structure in the rhizosphere of rice plants. Front. Microbiol. 2016, 6, 1537. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Berendsen, R.L.; Pieterse, C.M.; Bakker, P.A. The rhizosphere microbiome and plant health. Trends Plant Sci. 2012, 17, 478–486. [Google Scholar] [CrossRef]
- Liu, H.; Carvalhais, L.C.; Crawford, M.; Singh, E.; Dennis, P.G.; Pieterse, C.; Schenk, P.M. Inner plant values: Diversity, colonization and benefits from endophytic bacteria. Front. Microbiol. 2017, 8, 2552. [Google Scholar] [CrossRef] [PubMed]
- Okunishi, S.; Sako, K.; Mano, H.; Imamura, A.; Morisaki, H. Bacterial flora of endophytes in the maturing seed of cultivated rice (Oryza sativa). Microbes Environ. 2005, 20, 168–177. [Google Scholar] [CrossRef] [Green Version]
- Durán, P.; Acuña, J.J.; Tapia, M.; Azcón, R.; Paredes, C.; Rengel, Z.; Mora, M.L. Endophytic bacteria from selenium-supplemented wheat plants could be useful for plant-growth promotion, biofortification and Gaeumannomyces graminis biocontrol in wheat production. Biol. Fertil. Soils 2014, 50, 983–990. [Google Scholar] [CrossRef]
- Ren, J.-H.; Li, H.; Wang, Y.-F.; Ye, J.-R.; Yan, A.-Q.; Wu, X.-Q. Biocontrol potential of an endophytic Bacillus pumilus JK-SX001 against poplar canker. Biol. Control 2013, 67, 421–430. [Google Scholar] [CrossRef]
- Mohamad, O.A.; Li, L.; Ma, J.-B.; Hatab, S.; Xu, L.; Guo, J.-W.; Rasulov, B.A.; Liu, Y.-H.; Hedlund, B.P.; Li, W.-J. Evaluation of the antimicrobial activity of endophytic bacterial populations from Chinese traditional medicinal plant licorice and characterization of the bioactive secondary metabolites produced by Bacillus atrophaeus against Verticillium dahliae. Front. Microbiol. 2018, 9, 924. [Google Scholar] [CrossRef]
- Shi, Y.; Yang, H.; Zhang, T.; Sun, J.; Lou, K. Illumina-based analysis of endophytic bacterial diversity and space-time dynamics in sugar beet on the north slope of Tianshan mountain. Appl. Microbiol. Biotechnol. 2014, 98, 6375–6385. [Google Scholar] [CrossRef]
- Elbeltagy, A.; Nishioka, K.; Suzuki, H.; Sato, T.; Sato, Y.-I.; Morisaki, H.; Mitsui, H.; Minamisawa, K. Isolation and characterization of endophytic bacteria from wild and traditionally cultivated rice varieties. Soil Sci. Plant Nutr. 2000, 46, 617–629. [Google Scholar] [CrossRef]
- Compant, S.; Van Der Heijden, M.G.A.; Sessitsch, A. Climate change effects on beneficial plant–microorganism interactions. FEMS Microbiol. Ecol. 2010, 73, 197–214. [Google Scholar] [CrossRef] [PubMed]
- Penuelas, J.; Rico, L.; Ogaya, R.; Jump, A.; Terradas, J.J.P.B. Summer season and long-term drought increase the richness of bacteria and fungi in the foliar phyllosphere of Quercus ilex in a mixed Mediterranean forest. Plant Biol. 2012, 14, 565–575. [Google Scholar] [CrossRef]
- Ren, G.; Zhang, H.; Lin, X.; Zhu, J.; Jia, Z. Response of leaf endophytic bacterial community to elevated CO2 at different growth stages of rice plant. Front. Microbiol. 2015, 6, 855. [Google Scholar] [CrossRef]
- Walitang, D.I.; Kim, C.-G.; Kim, K.; Kang, Y.; Kim, Y.K.; Sa, T. The influence of host genotype and salt stress on the seed endophytic community of salt-sensitive and salt-tolerant rice cultivars. BMC Plant Biol. 2018, 18, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Bogas, A.C.; Ferreira, A.J.; Araújo, W.L.; Astolfi-Filho, S.; Kitajima, E.W.; Lacava, P.T.; Azevedo, J.L. Endophytic bacterial diversity in the phyllosphere of Amazon Paullinia cupana associated with asymptomatic and symptomatic anthracnose. Springerplus 2015, 4, 258. [Google Scholar] [CrossRef] [Green Version]
- Singh, R.K.; Mishra, R.P.N.; Jaiswal, H.K.; Kumar, V.; Pandey, S.P.; Rao, S.B.; Annapurna, K. Isolation and Identification of Natural Endophytic Rhizobia from Rice (Oryza sativa L.) Through rDNA PCR-RFLP and Sequence Analysis. Curr. Microbiol. 2006, 52, 345–349. [Google Scholar] [CrossRef]
- Mano, H.; Tanaka, F.; Watanabe, A.; Kaga, H.; Okunishi, S.; Morisaki, H. Culturable surface and endophytic bacterial flora of the maturing seeds of rice plants (Oryza sativa) cultivated in a paddy field. Microbes Environ. 2006, 21, 86–100. [Google Scholar] [CrossRef] [Green Version]
- Ji, S.H.; Gururani, M.; Chun, S.-C. Isolation and characterization of plant growth promoting endophytic diazotrophic bacteria from Korean rice cultivars. Microbiol. Res. 2013, 169, 83–98. [Google Scholar] [CrossRef]
- Ferrando, L.; Mañay, J.F.; Scavino, A.F. Molecular and culture-dependent analyses revealed similarities in the endophytic bacterial community composition of leaves from three rice (Oryza sativa) varieties. FEMS Microbiol. Ecol. 2012, 80, 696–708. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, H.; Sun, X.; Song, W.; Wang, Y.; Cai, M. Screening, identification and distribution of endophytic associative diazotrophs isolated from rice plants. Acta Bot. Sin. 1999, 41, 927–931. [Google Scholar]
- Zhang, C.-W.; Zhang, J.; Zhao, J.-J.; Zhao, X.; Zhao, D.-F.; Yin, H.-Q.; Zhang, X.-X. Serratia oryzae sp. nov., isolated from rice stems. Int. J. Syst. Evol. Microbiol. 2017, 67, 2928–2933. [Google Scholar] [CrossRef] [PubMed]
- Xiong, X.Q.; Liao, H.D.; Ma, J.S.; Liu, X.; Zhang, L.; Shi, X.; Yang, X.; Lu, X.; Zhu, Y. Isolation of a rice endophytic bacterium, P antoea sp. S d-1, with ligninolytic activity and characterization of its rice straw degradation ability. Lett. Appl. Microbiol. 2014, 58, 123–129. [Google Scholar] [CrossRef]
- Wang, X.; He, S.-W.; Guo, H.-B.; Thin, K.K.; Gao, J.-S.; Wang, Y.; Zhang, X.-X. Pseudomonas rhizoryzae sp. nov., isolated from rice. Int. J. Syst. Evol. Microbiol. 2020, 70, 944–950. [Google Scholar] [CrossRef] [PubMed]
- Mosquito, S.; Bertani, I.; Licastro, D.; Compant, S.; Myers, M.P.; Hinarejos, E.; Levy, A.; Venturi, V. In planta colonization and role of T6SS in two rice Kosakonia endophytes. Mol. Plant-Microbe Interact. 2020, 33, 349–363. [Google Scholar] [CrossRef] [PubMed]
- Eevers, N.; Gielen, M.; Sánchez-López, A.; Jaspers, S.; White, J.C.; Vangronsveld, J.; Weyens, N. Optimization of isolation and cultivation of bacterial endophytes through addition of plant extract to nutrient media. Microb. Biotechnol. 2015, 8, 707–715. [Google Scholar] [CrossRef]
- Tholozan, J.; Cappelier, J.; Tissier, J.; Delattre, G.; Federighi, M.J.A.; Microbiology, E. Physiological Characterization of Viable-but-Nonculturable Campylobacter Jejuni cells. Appl. Environ. Microbiol. 1999, 65, 1110–1116. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ma, Y.; Prasad, M.; Rajkumar, M.; Freitas, H. Plant growth promoting rhizobacteria and endophytes accelerate phytoremediation of metalliferous soils. Biotechnol. Adv. 2011, 29, 248–258. [Google Scholar] [CrossRef]
- Schleifer, K.H. Classification of Bacteria and Archaea: Past, present and future. Syst. Appl. Microbiol. 2009, 32, 533–542. [Google Scholar] [CrossRef]
- Girard, G.; Traag, B.A.; Sangal, V.; Mascini, N.; Hoskisson, P.; Goodfellow, M.; van Wezel, G.P. A novel taxonomic marker that discriminates between morphologically complex actinomycetes. Open Biol. 2013, 3, 130073. [Google Scholar] [CrossRef] [Green Version]
- Ventura, M.; Canchaya, C.; Tauch, A.; Chandra, G.; Fitzgerald, G.F.; Chater, K.F.; van Sinderen, D. Genomics of Actinobacteria: Tracing the Evolutionary History of an Ancient Phylum. Microbiol. Mol. Biol. Rev. 2007, 71, 495–548. [Google Scholar] [CrossRef] [Green Version]
- Nouioui, I.; Carro, L.; García-López, M.; Meier-Kolthoff, J.P.; Woyke, T.; Kyrpides, N.; Pukall, R.; Klenk, H.-P.; Goodfellow, M.; Göker, M. Genome-based taxonomic classification of the phylum actinobacteria. Front. Microbiol. 2018, 9, 2007. [Google Scholar] [CrossRef] [Green Version]
- Goris, J.; Konstantinidis, K.T.; Klappenbach, J.A.; Coenye, T.; Vandamme, P.; Tiedje, J.M. DNA–DNA hybridization values and their relationship to whole-genome sequence similarities. Int. J. Syst. Evol. Microbiol. 2007, 57, 81–91. [Google Scholar] [CrossRef] [Green Version]
- Maiden, M.C. Multilocus Sequence Typing of Bacteria. Annu. Rev. Microbiol. 2006, 60, 561–588. [Google Scholar] [CrossRef]
- Saito, A.; Ikeda, S.; Ezura, H.; Minamisawa, K. Microbial Community Analysis of the Phytosphere Using Culture-Independent Methodologies. Microbes Environ. 2007, 22, 93–105. [Google Scholar] [CrossRef]
- Fisher, M.M.; Triplett, E.W. Automated Approach for Ribosomal Intergenic Spacer Analysis of Microbial Diversity and Its Application to Freshwater Bacterial Communities. Appl. Environ. Microbiol. 1999, 65, 4630–4636. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ma, Y.; Rajkumar, M.; Zhang, C.; Freitas, H. Beneficial role of bacterial endophytes in heavy metal phytoremediation. J. Environ. Manag. 2016, 174, 14–25. [Google Scholar] [CrossRef] [PubMed]
- Reiter, B.; Pfeifer, U.; Schwab, H.; Sessitsch, A. Response of Endophytic Bacterial Communities in Potato Plants to Infection with Erwinia carotovora subsp. atroseptica. Appl. Environ. Microbiol. 2002, 68, 2261–2268. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Thijs, S.; van Dillewijn, P.; Sillen, W.; Truyens, S.; Holtappels, M.; D’haen, J.; Carleer, R.; Weyens, N.; Ameloot, M.; Ramos, J.L.; et al. Exploring the rhizospheric and endophytic bacterial communities of Acer pseudoplatanus growing on a TNT-contaminated soil: Towards the development of a rhizocompetent TNT-detoxifying plant growth promoting consortium. Plant Soil 2014, 385, 15–36. [Google Scholar] [CrossRef]
- Annapurna, K.; Govindasamy, V.; Sharma, M.; Ghosh, A.; Chikara, S.K. Whole genome shotgun sequence of Bacillus paralicheniformis strain KMS 80, a rhizobacterial endophyte isolated from rice (Oryza sativa L.). 3 Biotech 2018, 8, 223. [Google Scholar] [CrossRef]
- Tian, X.; Cao, L.; Tan, H.; Han, W.-Q.; Chen, M.; Liu, Y.; Zhou, S. Diversity of Cultivated and Uncultivated Actinobacterial Endophytes in the Stems and Roots of Rice. Microb. Ecol. 2007, 53, 700–707. [Google Scholar] [CrossRef]
- Ren, G.; Zhu, C.; Alam, M.S.; Tokida, T.; Sakai, H.; Nakamura, H.; Usui, Y.; Zhu, J.; Hasegawa, T.; Jia, Z. Response of soil, leaf endosphere and phyllosphere bacterial communities to elevated CO2 and soil temperature in a rice paddy. Plant Soil 2015, 392, 27–44. [Google Scholar] [CrossRef]
- Battu, L.; Reddy, M.M.; Goud, B.S.; Ulaganathan, K.; Kandasamy, U. Genome inside genome: NGS based identification and assembly of endophytic Sphingopyxis granuli and Pseudomonas aeruginosa genomes from rice genomic reads. Genomics 2017, 109, 141–146. [Google Scholar] [CrossRef] [PubMed]
- Manter, D.K.; Delgado, J.A.; Holm, D.G.; Stong, R.A. Pyrosequencing reveals a highly diverse and cultivar-specific bacterial endophyte community in potato roots. Microb. Ecol. 2010, 60, 157–166. [Google Scholar] [CrossRef] [PubMed]
- He, Y.; Zhou, B.-J.; Deng, G.-H.; Jiang, X.-T.; Zhang, H.; Zhou, H.-W. Comparison of microbial diversity determined with the same variable tag sequence extracted from two different PCR amplicons. BMC Microbiol. 2013, 13, 208. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Akinsanya, M.; Goh, J.K.; Lim, S.P.; Ting, A.S.Y. Metagenomics study of endophytic bacteria in Aloe vera using next-generation technology. Genom. Data 2015, 6, 159–163. [Google Scholar] [CrossRef] [Green Version]
- Ma, Y.; Oliveira, R.S.; Nai, F.; Rajkumar, M.; Luo, Y.; Rocha, I.D.S.; Freitas, H. The hyperaccumulator Sedum plumbizincicola harbors metal-resistant endophytic bacteria that improve its phytoextraction capacity in multi-metal contaminated soil. J. Environ. Manag. 2015, 156, 62–69. [Google Scholar] [CrossRef] [Green Version]
- Miliute, I.; Buzaite, O.; Baniulis, D.; Stanys, V. Bacterial endophytes in agricultural crops and their role in stress tolerance: A review. Zemdirb. Agric. 2015, 102, 465–478. [Google Scholar] [CrossRef] [Green Version]
- Ivleva, N.B.; Groat, J.; Staub, J.M.; Stephens, M. Expression of Active Subunit of Nitrogenase via Integration into Plant Organelle Genome. PLoS ONE 2016, 11, e0160951. [Google Scholar] [CrossRef] [Green Version]
- Bhattacharjee, R.B.; Singh, A.; Mukhopadhyay, S.N. Use of nitrogen-fixing bacteria as biofertiliser for non-legumes: Prospects and challenges. Appl. Microbiol. Biotechnol. 2008, 80, 199–209. [Google Scholar] [CrossRef]
- Kim, J.; Rees, D.C. Nitrogenase and biological nitrogen fixation. Biochemistry 1994, 33, 389–397. [Google Scholar] [CrossRef]
- Pallai, R. Effect of Plant Growth-Promoting Rhizobacteria on Canola (Brassica napus L) and Lentil (Lens culinaris Medik) Plants; University of Saskatchewan: Saskatoon, SK, Canada, 2005. [Google Scholar]
- Ladha, J.; De Bruijn, F.; Malik, K. Introduction: Assessing opportunities for nitrogen fixation in rice-a frontier project. Plant Soil 1997, 194, 1–10. [Google Scholar] [CrossRef]
- Holguin, G.; Patten, C.L. Biochemical and Genetic Mechanisms Used by Plant Growth Promoting Bacteria; Imperial College Press: London, UK, 1999. [Google Scholar]
- Wang, L.; Zhang, L.; Liu, Z.; Zhao, D.; Liu, X.; Zhang, B.; Xie, J.; Hong, Y.; Li, P.; Chen, S. A minimal nitrogen fixation gene cluster from Paenibacillus sp. WLY78 enables expression of active nitrogenase in Escherichia coli. PLoS Genet. 2013, 9, e1003865. [Google Scholar] [CrossRef]
- Bishop, P.E.; Joerger, R.D. Genetics and molecular biology of alternative nitrogen fixation systems. Annu. Rev. Plant Biol. 1990, 41, 109–125. [Google Scholar] [CrossRef]
- Cocking, E.C. Endophytic colonization of plant roots by nitrogen-fixing bacteria. Plant Soil 2003, 252, 169–175. [Google Scholar] [CrossRef]
- Ding, T.; Su, B.; Chen, X.; Xie, S.; Gu, S.; Wang, Q.; Huang, D.; Jiang, H. An endophytic bacterial strain isolated from Eucommia ulmoides inhibits southern corn leaf blight. Front. Microbiol. 2017, 8, 903. [Google Scholar] [CrossRef]
- Kandel, S.L.; Firrincieli, A.; Joubert, P.M.; Okubara, P.A.; Leston, N.D.; McGeorge, K.M.; Mugnozza, G.S.; Harfouche, A.; Kim, S.-H.; Doty, S.L. An in vitro study of bio-control and plant growth promotion potential of Salicaceae endophytes. Front. Microbiol. 2017, 8, 386. [Google Scholar] [CrossRef] [Green Version]
- Amaral, F.P.D.; Bueno, J.C.F.; Hermes, V.S.; Arisi, A.C.M. Gene expression analysis of maize seedlings (DKB240 variety) inoculated with plant growth promoting bacterium Herbaspirillum seropedicae. Symbiosis 2014, 62, 41–50. [Google Scholar] [CrossRef]
- Carrell, A.A.; Frank, A.C. Pinus flexilis and Picea engelmannii share a simple and consistent needle endophyte microbiota with a potential role in nitrogen fixation. Front. Microbiol. 2014, 5, 333. [Google Scholar] [CrossRef] [Green Version]
- Ahemad, M. Phosphate-solubilizing bacteria-assisted phytoremediation of metalliferous soils: A review. 3 Biotech 2014, 5, 111–121. [Google Scholar] [CrossRef] [Green Version]
- Afzal, I.; Shinwari, Z.K.; Sikandar, S.; Shahzad, S. Plant beneficial endophytic bacteria: Mechanisms, diversity, host range and genetic determinants. Microbiol. Res. 2019, 221, 36–49. [Google Scholar] [CrossRef]
- Maheshwari, R.; Bhutani, N.; Suneja, P. Screening and characterization of siderophore producing endophytic bacteria from Cicer arietinum and Pisum sativum plants. J. Appl. Biol. Biotechnol. 2019, 7, 7–14. [Google Scholar]
- Abedinzadeh, M.; Etesami, H.; Alikhani, H.A. Characterization of rhizosphere and endophytic bacteria from roots of maize (Zea mays L.) plant irrigated with wastewater with biotechnological potential in agriculture. Biotechnol. Rep. 2019, 21, e00305. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Wang, W.; Jin, Z.; Du, B.; Ding, Y.; Ni, T.; Jiao, F. Screening and diversity of plant growth promoting endophytic bacteria from peanut. Afr. J. Microbiol. Res. 2013, 7, 875–884. [Google Scholar]
- Gharu, A.; Tarafdar, J. Influence of organic acids on mobilization of inorganic and organic phosphorus in soil. J. Indian Soc. Soil Sci. 2004, 52, 248–253. [Google Scholar]
- Chen, Y.; Rekha, P.; Arun, A.; Shen, F.; Lai, W.-A.; Young, C. Phosphate solubilizing bacteria from subtropical soil and their tricalcium phosphate solubilizing abilities. Appl. Soil Ecol. 2006, 34, 33–41. [Google Scholar] [CrossRef]
- Khan, K.S.; Joergensen, R.G. Changes in microbial biomass and P fractions in biogenic household waste compost amended with inorganic P fertilizers. Bioresour. Technol. 2009, 100, 303–309. [Google Scholar] [CrossRef]
- Palaniappan, P.; Chauhan, P.S.; Saravanan, V.; Anandham, R.; Sa, T. Isolation and characterization of plant growth promoting endophytic bacterial isolates from root nodule of Lespedeza sp. Biol. Fertil. Soils 2010, 46, 807–816. [Google Scholar] [CrossRef]
- Puente, M.E.; Li, C.Y.; Bashan, Y. Rock-degrading endophytic bacteria in cacti. Environ. Exp. Bot. 2009, 66, 389–401. [Google Scholar] [CrossRef]
- Goswami, D.; Vaghela, H.; Parmar, S.; Dhandhukia, P.; Thakker, J.N. Plant growth promoting potentials of Pseudomonas spp. strain OG isolated from marine water. J. Plant Interact. 2013, 8, 281–290. [Google Scholar] [CrossRef] [Green Version]
- Patel, K.; Goswami, D.; Dhandhukia, P.; Thakker, J. Techniques to study microbial phytohormones. In Bacterial Metabolites in Sustainable Agroecosystem. Sustainable Development and Biodiversity; Maheshwari, D., Ed.; Springer: Cham, Switzerland, 2015; Volume 12, pp. 1–27. [Google Scholar]
- Khan, M.M.A.; Haque, E.; Paul, N.C.; Khaleque, M.A.; Al-Garni, S.M.; Rahman, M.; Islam, M.T. Enhancement of growth and grain yield of rice in nutrient deficient soils by rice probiotic bacteria. Rice Sci. 2017, 24, 264–273. [Google Scholar] [CrossRef]
- Phetcharat, P.; Duangpaeng, A. Screening of endophytic bacteria from organic rice tissue for indole acetic acid production. Procedia Eng. 2012, 32, 177–183. [Google Scholar] [CrossRef] [Green Version]
- Xu, T.; Cao, L.; Zeng, J.; Franco, C.; Yang, Y.; Hu, X.; Liu, Y.; Wang, X.; Gao, Y.; Bu, Z.; et al. The antifungal action mode of the rice endophyte Streptomyces hygroscopicus OsiSh-2 as a potential biocontrol agent against the rice blast pathogen. Pestic. Biochem. Physiol. 2019, 160, 58–69. [Google Scholar] [CrossRef] [PubMed]
- Spaepen, S.; Vanderleyden, J. Auxin and plant-microbe interactions. Cold Spring Harb. Perspect. Biol. 2011, 3, a001438. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Glick, B.R. Plant Growth-Promoting Bacteria: Mechanisms and Applications. Scientifica 2012, 2012, 1–15. [Google Scholar] [CrossRef] [Green Version]
- Mole, B.M.; Baltrus, D.A.; Dangl, J.L.; Grant, S.R. Global virulence regulation networks in phytopathogenic bacteria. Trends Microbiol. 2007, 15, 363–371. [Google Scholar] [CrossRef] [PubMed]
- Spaepen, S.; Vanderleyden, J.; Remans, R. Indole-3-acetic acid in microbial and microorganism-plant signaling. FEMS Microbiol. Rev. 2007, 31, 425–448. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xu, T.; Li, Y.; Zeng, X.; Yang, X.; Yang, Y.; Yuan, S.; Hu, X.; Zeng, J.; Wang, Z.; Liu, Q. Isolation and evaluation of endophytic Streptomyces endus OsiSh-2 with potential application for biocontrol of rice blast disease. J. Sci. Food Agric. 2017, 97, 1149–1157. [Google Scholar] [CrossRef] [PubMed]
- Leveau, J.H.J.; Lindow, S.E. Utilization of the Plant Hormone Indole-3-Acetic Acid for Growth by Pseudomonas putida Strain 1290. Appl. Environ. Microbiol. 2005, 71, 2365–2371. [Google Scholar] [CrossRef] [Green Version]
- Asaf, S.; Khan, M.A.; Khan, A.L.; Waqas, M.; Shahzad, R.; Kim, A.-Y.; Kang, S.-M.; Lee, I.-J. Bacterial endophytes from arid land plants regulate endogenous hormone content and promote growth in crop plants: An example of Sphingomonas sp. and Serratia marcescens. J. Plant Interact. 2017, 12, 31–38. [Google Scholar] [CrossRef] [Green Version]
- Herrera, S.D.; Grossi, C.; Zawoznik, M.; Groppa, M.D. Wheat seeds harbour bacterial endophytes with potential as plant growth promoters and biocontrol agents of Fusarium graminearum. Microbiol. Res. 2016, 186, 37–43. [Google Scholar] [CrossRef]
- Ortiz-Castro, R.; Contreras-Cornejo, H.A.; Macías-Rodríguez, L.; López-Bucio, J. The role of microbial signals in plant growth and development. Plant Signal. Behav. 2009, 4, 701–712. [Google Scholar] [CrossRef] [Green Version]
- Persello-Cartieaux, F.; Nussaume, L.; Robaglia, C. Tales from the underground: Molecular plant–rhizobacteria interactions. Plant Cell Environ. 2003, 26, 189–199. [Google Scholar] [CrossRef]
- Nieto, K.; Frankenberger, W. Influence of adenine, isopentyl alcohol and Azotobacter chroococcum on the growth of Raphanus sativus. Plant Soil 1990, 127, 147–156. [Google Scholar] [CrossRef]
- Da, K.; Nowak, J.; Flinn, B. Potato cytosine methylation and gene expression changes induced by a beneficial bacterial endophyte, Burkholderia phytofirmans strain PsJN. Plant Physiol. Biochem. 2012, 50, 24–34. [Google Scholar] [CrossRef]
- Bhore, S.J.; Ravichantar, N.; Loh, C.Y. Screening of endophytic bacteria isolated from leaves of Sambung Nyawa [Gynura procumbens (Lour.) Merr.] for cytokinin-like compounds. Bioinformation 2010, 5, 191–197. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dalal, J.; Kulkarni, N. Antagonistic and Plant Growth Promoting Potentials of Indigenous Endophytic Actinomycetes of Soybean (Glycine max (L) Merril). CIBTech J. Microbiol. 2014, 3, 1–12. [Google Scholar]
- Dodd, I.; Zinovkina, N.; Safronova, V.; Belimov, A. Rhizobacterial mediation of plant hormone status. Ann. Appl. Biol. 2010, 157, 361–379. [Google Scholar] [CrossRef]
- Tsavkelova, E.; Klimova, S.Y.; Cherdyntseva, T.; Netrusov, A. Microbial producers of plant growth stimulators and their practical use: A review. Appl. Biochem. Microbiol. 2006, 42, 117–126. [Google Scholar] [CrossRef]
- Kang, S.-M.; Joo, G.-J.; Hamayun, M.; Na, C.-I.; Shin, D.-H.; Kim, H.Y.; Hong, J.-K.; Lee, I.-J. Gibberellin production and phosphate solubilization by newly isolated strain of Acinetobacter calcoaceticus and its effect on plant growth. Biotechnol. Lett. 2009, 31, 277–281. [Google Scholar] [CrossRef]
- Ishak, Z.; Mohd Iswadi, M.; Russman Nizam, A.; Ahmad Kamil, M.; Ernie Eileen, R.; Wan Syaidatul, A.; Ainon, H. Plant growth hormones produced by endophytic Bacillus subtilis strain LKM-BK isolated from cocoa. Malays. Cocoa J. 2016, 9, 127–133. [Google Scholar]
- Dugardeyn, J.; Van Der Straeten, D. Ethylene: Fine-tuning plant growth and development by stimulation and inhibition of elongation. Plant Sci. 2008, 175, 59–70. [Google Scholar] [CrossRef]
- Belimov, A.; Hontzeas, N.; Safronova, V.; Demchinskaya, S.; Piluzza, G.; Bullitta, S.; Glick, B. Cadmium-tolerant plant growth-promoting bacteria associated with the roots of Indian mustard (Brassica juncea L. Czern.). Soil Biol. Biochem. 2005, 37, 241–250. [Google Scholar] [CrossRef]
- Giovanelli, J.; Mudd, S.H.; Datko, A.H. Sulfur amino acids in plants. In Amino Acids and Derivatives; Elsevier Inc.: Amsterdam, The Netherlands, 1980; Volume 5, pp. 453–505. [Google Scholar]
- Glick, B.R.; Todorovic, B.; Czarny, J.; Cheng, Z.; Duan, J.; McConkey, B. Promotion of plant growth by bacterial ACC deaminase. Crit. Rev. Plant Sci. 2007, 26, 227–242. [Google Scholar] [CrossRef]
- Shaharoona, B.; Arshad, M.; Zahir, Z. Effect of plant growth promoting rhizobacteria containing ACC-deaminase on maize (Zea mays L.) growth under axenic conditions and on nodulation in mung bean (Vigna radiata L.). Lett. Appl. Microbiol. 2006, 42, 155–159. [Google Scholar] [CrossRef]
- Grichko, V.P.; Glick, B.R. Amelioration of flooding stress by ACC deaminase-containingplant growth-promoting bacteria. Plant Physiol. Biochem. 2001, 39, 11–17. [Google Scholar] [CrossRef]
- Mayak, S.; Tirosh, T.; Glick, B.R. Plant growth-promoting bacteria that confer resistance to water stress in tomatoes and peppers. Plant Sci. 2004, 166, 525–530. [Google Scholar] [CrossRef]
- Zahir, Z.A.; Ghani, U.; Naveed, M.; Nadeem, S.M.; Asghar, H.N. Comparative effectiveness of Pseudomonas and Serratia sp. containing ACC-deaminase for improving growth and yield of wheat (Triticum aestivum L.) under salt-stressed conditions. Arch. Microbiol. 2009, 191, 415–424. [Google Scholar] [CrossRef] [PubMed]
- Berg, G. Plant–microbe interactions promoting plant growth and health: Perspectives for controlled use of microorganisms in agriculture. Appl. Microbiol. Biotechnol. 2009, 84, 11–18. [Google Scholar] [CrossRef] [PubMed]
- Smyth, E.; McCarthy, J.; Nevin, R.; Khan, M.; Dow, M.; O’Gara, F.; Doohan, F. In vitro analyses are not reliable predictors of the plant growth promotion capability of bacteria; a Pseudomonas fluorescens strain that promotes the growth and yield of wheat. J. Appl. Microbiol. 2011, 111, 683–692. [Google Scholar] [CrossRef] [PubMed]
- Perrig, D.; Boiero, M.; Masciarelli, O.; Penna, C.; Ruiz, O.; Cassán, F.; Luna, M. Plant-growth-promoting compounds produced by two agronomically important strains of Azospirillum brasilense, and implications for inoculant formulation. Appl. Microbiol. Biotechnol. 2007, 75, 1143–1150. [Google Scholar] [CrossRef]
- Spaepen, S.; Das, F.; Luyten, E.; Michiels, J.; Vanderleyden, J. Indole-3-acetic acid-regulated genes in Rhizobium etli CNPAF512. FEMS Microbiol. Lett. 2009, 291, 195–200. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Malfanova, N.; Kamilova, F.; Validov, S.; Shcherbakov, A.; Chebotar, V.; Tikhonovich, I.; Lugtenberg, B. Characterization of Bacillus subtilis HC8, a novel plant-beneficial endophytic strain from giant hogweed. Microb. Biotechnol. 2011, 4, 523–532. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Melnick, R.L.; Zidack, N.K.; Bailey, B.A.; Maximova, S.N.; Guiltinan, M.; Backman, P.A. Bacterial endophytes: Bacillus spp. from annual crops as potential biological control agents of black pod rot of cacao. Biol. Control 2008, 46, 46–56. [Google Scholar] [CrossRef]
- Liu, B.; Huang, L.; Buchenauer, H.; Kang, Z. Isolation and partial characterization of an antifungal protein from the endophytic Bacillus subtilis strain EDR4. Pestic. Biochem. Physiol. 2010, 98, 305–311. [Google Scholar] [CrossRef]
- Li, H.; Wang, X.; Han, M.; Zhao, Z.; Wang, M.; Tang, Q.; Liu, C.; Kemp, B.; Gu, Y.; Shuang, J. Endophytic Bacillus subtilis ZZ120 and its potential application in control of replant diseases. Afr. J. Biotechnol. 2012, 11, 231–242. [Google Scholar]
- Villa-Rodríguez, E.; Parra-Cota, F.; Castro-Longoria, E.; López-Cervantes, J.; de los Santos-Villalobos, S. Bacillus subtilis TE3: A promising biological control agent against Bipolaris sorokiniana, the causal agent of spot blotch in wheat (Triticum turgidum L. subsp. durum). Biol. Control 2019, 132, 135–143. [Google Scholar] [CrossRef]
- Rangjaroen, C.; Lumyong, S.; Sloan, W.T.; Sungthong, R. Herbicide-tolerant endophytic bacteria of rice plants as the biopriming agents for fertility recovery and disease suppression of unhealthy rice seeds. BMC Plant Biol. 2019, 19, 580. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chung, E.J.; Hossain, M.T.; Khan, A.; Kim, K.H.; Jeon, C.O.; Chung, Y.R. Bacillus oryzicola sp. nov., an endophytic bacterium isolated from the roots of rice with antimicrobial, plant growth promoting, and systemic resistance inducing activities in rice. Plant Pathol. J. 2015, 31, 152. [Google Scholar] [CrossRef] [PubMed]
- Verma, S.K.; Kingsley, K.; Irizarry, I.; Bergen, M.; Kharwar, R.; White, J.F., Jr. Seed-vectored endophytic bacteria modulate development of rice seedlings. J. Appl. Microbiol. 2017, 122, 1680–1691. [Google Scholar] [CrossRef]
- Ruiza, D.; Agaras, B.C.; De Werrab, P.; Wall, L.G.; Valverde, C. Characterization and screening of plant probiotic traits of bacteria isolated from rice seeds cultivated in Argentina. J. Microbiol. 2011, 49, 902–912. [Google Scholar] [CrossRef] [PubMed]
- Santoyo, G.; Moreno-Hagelsieb, G.; Orozco-Mosqueda, M.D.C.; Glick, B.R. Plant growth-promoting bacterial endophytes. Microbiol. Res. 2016, 183, 92–99. [Google Scholar] [CrossRef]
- Van Rij, E.T.; Wesselink, M.; Chin-A-Woeng, T.F.; Bloemberg, G.V.; Lugtenberg, B.J. Influence of environmental conditions on the production of phenazine-1-carboxamide by Pseudomonas chlororaphis PCL1391. Mol. Plant-Microbe Interact. 2004, 17, 557–566. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jousset, A.; Rochat, L.; Lanoue, A.; Bonkowski, M.; Keel, C.; Scheu, S. Plants respond to pathogen infection by enhancing the antifungal gene expression of root-associated bacteria. Mol. Plant-Microbe Interact. 2011, 24, 352–358. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dekkers, L.C.; Mulders, I.H.M.; Phoelich, C.C.; Chin-A-Woeng, T.F.C.; Wijfjes, A.H.M.; Lugtenberg, B.J.J. The sss Colonization Gene of the Tomato-Fusarium oxysporum f. sp. radicis-lycopersici Biocontrol strain pseudomonas fluorescens wcs365 can improve root colonization of other wild-type Pseudomonas spp. bacteria. Mol. Plant-Microbe Interactions 2000, 13, 1177–1183. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Haas, D.; Défago, G. Biological control of soil-borne pathogens by fluorescent pseudomonads. Nat. Rev. Microbiol. 2005, 3, 307–319. [Google Scholar] [CrossRef] [PubMed]
- Benizri, E.; Baudoin, E.; Guckert, A. Root Colonization by inoculated plant growth-promoting rhizobacteria. Biocontrol Sci. Technol. 2001, 11, 557–574. [Google Scholar] [CrossRef]
- Zhang, L.H.; Dong, Y.H. Quorum sensing and signal interference: Diverse implications. Mol. Microbiol. 2004, 53, 1563–1571. [Google Scholar] [CrossRef] [PubMed]
- Uroz, S.; Dessaux, Y.; Oger, P. Quorum sensing and quorum quenching: The yin and yang of bacterial communication. ChemBioChem 2009, 10, 205–216. [Google Scholar] [CrossRef] [PubMed]
- Müller, H.; Westendorf, C.; Leitner, E.; Chernin, L.; Riedel, K.; Schmidt, S.; Eberl, L.; Berg, G. Quorum-sensing effects in the antagonistic rhizosphere bacterium Serratia plymuthica HRO-C48. FEMS Microbiol. Ecol. 2009, 67, 468–478. [Google Scholar] [CrossRef] [Green Version]
- Dandurishvili, N.; Toklikishvili, N.; Ovadis, M.; Eliashvili, P.; Giorgobiani, N.; Keshelava, R.; Tediashvili, M.; Vainstein, A.; Khmel, I.; Szegedi, E. Broad-range antagonistic rhizobacteria Pseudomonas fluorescens and Serratia plymuthica suppress Agrobacterium crown gall tumours on tomato plants. J. Appl. Microbiol. 2011, 110, 341–352. [Google Scholar] [CrossRef]
- Chernin, L.; Toklikishvili, N.; Ovadis, M.; Kim, S.; Ben-Ari, J.; Khmel, I.; Vainstein, A. Quorum-sensing quenching by rhizobacterial volatiles. Environ. Microbiol. Rep. 2011, 3, 698–704. [Google Scholar] [CrossRef] [PubMed]
- Berg, G.; Hallmann, J. Control of plant pathogenic fungi with bacterial endophytes. In Microbial Root Endophytes; Springer: New York, NY, USA, 2006; pp. 53–69. [Google Scholar]
- El-Tarabily, K.A. Rhizosphere-competent isolates of streptomycete and non-streptomycete actinomycetes capable of producing cell-wall-degrading enzymes to control Pythium aphanidermatum damping-off disease of cucumber. Botany 2006, 84, 211–222. [Google Scholar] [CrossRef]
- Ordentlich, A.; Elad, Y.; Chet, I. The role of chitinase of Serratia marcescens in biocontrol of Sclerotium rolfsii. Phytopathology 1988, 78, 84–88. [Google Scholar]
- Araújo, W.L.; Marcon, J.; Maccheroni, W.; van Elsas, J.D.; van Vuurde, J.W.; Azevedo, J.L. Diversity of endophytic bacterial populations and their interaction with Xylella fastidiosa in citrus plants. Appl. Environ. Microbiol. 2002, 68, 4906–4914. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Alvin, A.; Miller, K.I.; Neilan, B.A. Exploring the potential of endophytes from medicinal plants as sources of antimycobacterial compounds. Microbiol. Res. 2014, 169, 483–495. [Google Scholar] [CrossRef] [PubMed]
- Pieterse, C.M.; Van der Does, D.; Zamioudis, C.; Leon-Reyes, A.; Van Wees, S.C. Hormonal modulation of plant immunity. Annu. Rev. Cell Dev. Biol. 2012, 28, 489–521. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Akhtar, M.S.; Siddiqui, Z.A. Role of plant growth promoting rhizobacteria in biocontrol of plant diseases and sustainable agriculture. In Plant Growth and Health Promoting Bacteria; Springer: New York, NY, USA, 2010; pp. 157–195. [Google Scholar]
- Kamilova, F.; Validov, S.; Azarova, T.; Mulders, I.; Lugtenberg, B. Enrichment for enhanced competitive plant root tip colonizers selects for a new class of biocontrol bacteria. Environ. Microbiol. 2005, 7, 1809–1817. [Google Scholar] [CrossRef] [PubMed]
- Van Loon, L.; Bakker, P. Induced systemic resistance as a mechanism of disease suppression by rhizobacteria. In PGPR: Biocontrol and Biofertilization; Siddiqui, Z.A., Ed.; Springer: Dordrecht, The Netherlands, 2003; pp. 39–66. [Google Scholar]
- Van Loon, L.; Bakker, P. Root-associated bacteria inducing systemic resistance. In Plant-Associated Bacteria; Gnanamanickam, S.S., Ed.; Springer: Dordrecht, The Netherlands, 2007; pp. 269–316. [Google Scholar]
- Bolyen, E.; Rideout, J.R.; Dillon, M.R.; Bokulich, N.A.; Abnet, C.C.; Al-Ghalith, G.A.; Alexander, H.; Alm, E.J.; Arumugam, M.; Asnicar, F.; et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 2019, 37, 852–857. [Google Scholar] [CrossRef] [PubMed]
- Caporaso, J.G.; Bittinger, K.; Bushman, F.; DeSantis, T.Z.; Andersen, G.; Knight, R. PyNAST: A flexible tool for aligning sequences to a template alignment. Bioinformatics 2009, 26, 266–267. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, Q.; Garrity, G.M.; Tiedje, J.M.; Cole, J.R. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl. Environ. Microbiol. 2007, 73, 5261–5267. [Google Scholar] [CrossRef] [Green Version]
- Kwak, M.-J.; Kong, H.G.; Choi, K.; Kwon, S.-K.; Song, J.Y.; Lee, J.; Lee, P.A.; Choi, S.Y.; Seo, M.; Lee, H.J.; et al. Rhizosphere microbiome structure alters to enable wilt resistance in tomato. Nat. Biotechnol. 2018, 36, 1100–1109. [Google Scholar] [CrossRef]
- Zhang, J.; Liu, Y.-X.; Zhang, N.; Hu, B.; Jin, T.; Xu, H.; Qin, Y.; Yan, P.; Zhang, X.; Guo, X.; et al. NRT1.1B is associated with root microbiota composition and nitrogen use in field-grown rice. Nat. Biotechnol. 2019, 37, 676–684. [Google Scholar] [CrossRef]
- Edgar, R.C. UPARSE: Highly accurate OTU sequences from microbial amplicon reads. Nat. Methods 2013, 10, 996–998. [Google Scholar] [CrossRef]
- Rideout, J.R.; He, Y.; Navas-Molina, J.A.; Walters, W.A.; Ursell, L.K.; Gibbons, S.M.; Chase, J.; McDonald, D.; Gonzalez, A.; Robbins-Pianka, A.; et al. Subsampled open-reference clustering creates consistent, comprehensive OTU definitions and scales to billions of sequences. PeerJ 2014, 2, e545. [Google Scholar] [CrossRef] [Green Version]
- Edgar, R.C.J.B. Updating the 97% identity threshold for 16S ribosomal RNA OTUs. Bioinformatics 2018, 34, 2371–2375. [Google Scholar] [CrossRef] [PubMed]
- Louca, S.; Parfrey, L.W.; Doebeli, M. Decoupling function and taxonomy in the global ocean microbiome. Science 2016, 353, 1272–1277. [Google Scholar] [CrossRef] [PubMed]
- Omojate, G.C.; Enwa, F.O.; Jewo, A.O.; Eze, C.O. Mechanisms of antimicrobial actions of phytochemicals against enteric pathogens—A review. J. Pharm. Chem. Biol. Sci. 2014, 2, 77–85. [Google Scholar]
- Pimentel, M.R.; Molina, G.; Dionísio, A.P.; Junior, M.R.M.; Pastore, G.M. The Use of Endophytes to Obtain Bioactive Compounds and Their Application in Biotransformation Process. Biotechnol. Res. Int. 2011, 2011, 1–11. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shukla, S.; Habbu, P.; Kulkarni, V.; Jagadish, K.; Pandey, A.; Sutariya, V. Endophytic microbes: A novel source for biologically/pharmacologically active secondary metabolites. Asian J. Pharm. Toxicol. 2014, 2, 1–6. [Google Scholar]
- Strobel, G.A. Endophytes as sources of bioactive products. Microbes Infect. 2003, 5, 535–544. [Google Scholar] [CrossRef]
- Liarzi, O.; Bucki, P.; Miyara, S.B.; Ezra, D. Bioactive Volatiles from an Endophytic Daldinia cf. concentrica Isolate Affect the Viability of the Plant Parasitic Nematode Meloidogyne javanica. PLoS ONE 2016, 11, e0168437. [Google Scholar] [CrossRef]
- Nongkhlaw, F.M.; Joshi, S.R. Investigation on the bioactivity of culturable endophytic and epiphytic bacteria associated with ethnomedicinal plants. J. Infect. Dev. Ctries. 2015, 9, 954–961. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gouda, S.; Das, G.; Sen, S.K.; Shin, H.-S.; Patra, J.K. Endophytes: A treasure house of bioactive compounds of medicinal importance. Front. Microbiol. 2016, 7, 1538. [Google Scholar] [CrossRef] [PubMed] [Green Version]
| Endophytes | Host | Antagonistic Activity | References |
|---|---|---|---|
| Bacillus wiedmannii strain C-1CL-2 | Japonica Rice (Oryza sativa L.) cultivar Xiushui 48 | Fusarium graminearum and Rhizoctonia solani | [25] |
| Bacillus wiedmannii strain C-1CL-4 | Fusarium graminearum and Rhizoctonia solani | ||
| Bacillus altitudinis strain C-1W-5 | Magnaporthe oryzae | ||
| Lysinibacillus fusiformis strain C-1WF-3 | Fusarium moniliforme | ||
| Fictibacillus phosphorivorans strain C-1Y-9 | Fusarium graminearum and Rhizoctonia solani | ||
| Fictibacillus phosphorivorans strain C-1Y-10 | Magnaporthe oryzae | ||
| Fictibacillus phosphorivorans strain C-1Y-13 | Fusarium moniliforme | ||
| Fictibacillus phosphorivorans strain C-1Y-16 | Fusarium moniliforme | ||
| Fictibacillus phosphorivorans strain C-1Y-17 | Fusarium graminearum, Fusarium moniliforme and Rhizoctonia solani | ||
| Bacillus altitudinis strain C-2B-2 1 | Rice (Oryza sativa L.) cultivar Y-003 | Rhizoctonia solani | |
| Bacillus altitudinis strain C-2D-1 1 | Rhizoctonia solani | ||
| Bacillus aryabhattai strain C-2HW-1 | Fusarium graminearum and Rhizoctonia solani | ||
| Lysinibacillus mangiferihumi strain C-2HW-3 | Fusarium moniliforme | ||
| Bacillus marisflavi strain C-2LY-2 | Fusarium graminearum and Rhizoctonia solani | ||
| Bacillus marisflavi strain C-2LY-5 | Fusarium graminearum and Rhizoctonia solani | ||
| Bacillus aryabhattai strain C-2LY-6 | Fusarium graminearum, Fusarium moniliforme and Rhizoctonia solani | ||
| Bacillus altitudinis strain C-2RO-1 | Fusarium graminearum, Rhizoctonia solani and Magnaporthe oryzae | ||
| Bacillus altitudinis strain C-2RO-3 | Fusarium graminearum, Rhizoctonia solani and Magnaporthe oryzae | ||
| Bacillus altitudinis strain C-2RO-4 | Rhizoctonia solani | ||
| Bacillus altitudinis strain C-2S-1 | Rhizoctonia solani | ||
| Bacillus altitudinis strain C-2SN-3 | Fusarium moniliforme | ||
| Bacillus cereus strain C-2W-1 | Fusarium moniliforme | ||
| Bacillus tequilensis strain C-2W-3 | Rhizoctonia solani | ||
| Bacillus indicus strain C-2Y-1 | Fusarium graminearum and Rhizoctonia solani | ||
| Bacillus marisflavi strain C-2Y-2 | Fusarium graminearum, Fusarium moniliforme and Rhizoctonia solani | ||
| Bacillus marisflavi strain C-2Y-6 | Fusarium graminearum and Rhizoctonia solani | ||
| Bacillus cereus strain C-3CL-6 | Rice (Oryza sativa L.) cultivar CO-39 | Fusarium moniliforme | |
| Bacillus toyonensis strain C-3CL-7 | Rhizoctonia solani | ||
| Bacillus wiedmannii strain C-3CL-8 | Fusarium graminearum and Rhizoctonia solani | ||
| Lysinibacillus mangiferihumi strain C-3F-5 | Fusarium moniliforme | ||
| Bacillus altitudinis strain C-3R-1 | Fusarium graminearum and Rhizoctonia solani | ||
| Bacillus altitudinis strain C-3R-3 | Fusarium graminearum, Rhizoctonia solani and Magnaporthe oryzae | ||
| Bacillus altitudinis strain C-3R-8 | Fusarium graminearum and Rhizoctonia solani | ||
| Bacillus cereus strain C-3R-9 | Fusarium moniliforme | ||
| Bacillus altitudinis strain C-3R-10 | Fusarium graminearum, Rhizoctonia solani and Magnaporthe oryzae | ||
| Cupriavidus metallidurans strain C-3SP-3 | Magnaporthe oryzae | ||
| Paenibacillus cucumis strain C-3T-7 | Fusarium moniliforme | ||
| Bacillus altitudinis strain C-3WA-8 | Rhizoctonia solani and Magnaporthe oryzae | ||
| Bacillus marisflavi strain C-3Y-2 | Fusarium graminearum and Fusarium moniliforme | ||
| Bacillus marisflavi strain C-3Y-5 | Fusarium graminearum, Fusarium moniliforme and Rhizoctonia solani | ||
| Fictibacillus phosphorivorans strain C-3Y-10 | Magnaporthe oryzae | ||
| Bacillus sp. (in: Bacteria) strain CPS003 | Rice Oryza sativa L. var. indica cv. RD41 | Bipolaris sp. isolate KPS3, Bipolaris sp. isolate KPS5, Curvularia sp. isolate KPS41, Nigrospora sp. isolate KPS45, Fusarium sp. isolate KPS91 and Curvularia sp. isolate KPS102 | [176] |
| Bacillus sp. (in: Bacteria) strain CZR007 | |||
| Bacillus sp. (in: Bacteria) strain CZS004 | |||
| Bacillus sp. (in: Bacteria) strain CZS006 | |||
| Streptomyces endus strain OsiSh -2 | Rice (Oryza sativa L.) cultivar Gumei 4 | Magnaporthe oryzae, Colletitrichum karstii, Colletitrichum siamense, Colletitrichum camelliae, Colletitrichum frocticola, Colletitrichum glosporioides, Valsa mali, Guignardia bidwellii, Physalospora piricola, Botryosphaeria ribis, Rhizopus nigricans, Phomopsis vexans, Cladosporium fulvum, Stempblium consortiale, Botrytis cinerea, Colletrichum orbiculare, Helminthosporium maydis, Nigrospora oryzae and Rhizoctonia solani | [140,145] |
| Bacillus velezensis strain YC7010 | Rice (Oryza sativa L.) | Fusarium fujikuroi | [177] |
| Bacillus velezensis strain YC7007 | Fusarium fujikuroi KACC 44022, Magnaporthe grisea isolate KACC 40415, Bipolaris oryzae isolate KACC 40853, Rhizoctonia solani isolate KCTC 40101, Sclerotinia sclerotiorum isolate GSCC 50501, Botrytis cinerea isolate KCTC 6973, Fusarium oxysporum isolate KCTC 16909, Botryosphaeria dothidea isolate GSCC 50201, Pythium ultimum isolate GSCC 50651 and Alternaria panax isolate KACC 42461 | ||
| Enterobacter asburiae strain VWB1 | Rice (Oryza sativa L.) cultivar rex | Fusarium oxysporum | [178] |
| Pantoea dispersa strain VWB2 | Fusarium oxysporum | ||
| Pseudomonas putida strain VWB3 | Fusarium oxysporum | ||
| Pantoea agglomerans strain CT1 | Rice (Oryza sativa L.) variety CT6919 | Pythium ultimum | [179] |
| Pantoea agglomerans strain CT2 | Curvularia sp and Fusarium oxysporum var. radicis-lycopersici strain 22 | ||
| Pantoea ananatis strain CT10 | Pythium ultimum, Curvularia sp and Fusarium oxysporum var. radicis-lycopersici strain 27 | ||
| Paenibacillus sp. CT14 | Curvularia sp | ||
| Microbacterium sp. CT28 | Pythium ultimum | ||
| Curtobacterium sp.CT30 | Pythium ultimum | ||
| Paenibacillus kribbensis strain HS-R01 | Rice cultivars (Oryza sativa var. Japonica c.v. Chilbo, Chuchung, Haiami, Ilpum | Fusarium oxysporum and Rhizoctonia solani | [83] |
| Paenibacillus kribbensis strain HS-R14 | Fusarium oxysporum and Rhizoctonia solani | ||
| Bacillus aryabhattai strain HS-S05 | Fusarium oxysporum | ||
| Bacillus megaterium strain KW7-R08 | Fusarium oxysporum | ||
| Klebsiella pneumoniae strain KW7-S06 | Fusarium oxysporum and Rhizoctonia solani | ||
| Klebsiella pneumoniae strain KW7-S22 | Fusarium oxysporum and Rhizoctonia solani | ||
| Klebsiella pneumoniae strain KW7-S27 | Fusarium oxysporum and Rhizoctonia solani | ||
| Klebsiella pneumoniae strain KW7-S33 | Fusarium oxysporum and Rhizoctonia solani | ||
| Bacillus subtilis strain CB-R05 | Fusarium oxysporum and Rhizoctonia solani | ||
| Microbacterium binotii strain CB-S18 | Fusarium oxysporum and Rhizoctonia solani | ||
| Microbacterium trichotecenolyticum strain SW521-L21 | Fusarium oxysporum and Rhizoctonia solani | ||
| Microbacterium trichotecenolyticum strain SW521-L37 | Fusarium oxysporum and Rhizoctonia solani |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ali, M.; Ali, Q.; Sohail, M.A.; Ashraf, M.F.; Saleem, M.H.; Hussain, S.; Zhou, L. Diversity and Taxonomic Distribution of Endophytic Bacterial Community in the Rice Plant and Its Prospective. Int. J. Mol. Sci. 2021, 22, 10165. https://doi.org/10.3390/ijms221810165
Ali M, Ali Q, Sohail MA, Ashraf MF, Saleem MH, Hussain S, Zhou L. Diversity and Taxonomic Distribution of Endophytic Bacterial Community in the Rice Plant and Its Prospective. International Journal of Molecular Sciences. 2021; 22(18):10165. https://doi.org/10.3390/ijms221810165
Chicago/Turabian StyleAli, Mohsin, Qurban Ali, Muhammad Aamir Sohail, Muhammad Furqan Ashraf, Muhammad Hamzah Saleem, Saddam Hussain, and Lei Zhou. 2021. "Diversity and Taxonomic Distribution of Endophytic Bacterial Community in the Rice Plant and Its Prospective" International Journal of Molecular Sciences 22, no. 18: 10165. https://doi.org/10.3390/ijms221810165
APA StyleAli, M., Ali, Q., Sohail, M. A., Ashraf, M. F., Saleem, M. H., Hussain, S., & Zhou, L. (2021). Diversity and Taxonomic Distribution of Endophytic Bacterial Community in the Rice Plant and Its Prospective. International Journal of Molecular Sciences, 22(18), 10165. https://doi.org/10.3390/ijms221810165









