1. Introduction
Abnormal overexpression of mucin1 (MUC1) in various cancers and the its interactions with several proteins related to cancer transformation have led to MUC1 as a potential oncological target [
1,
2,
3]. MUC1 is a member of the mucin family and is composed of alpha and beta domains. The alpha domain, carrying highly repetitive sequences, is strongly glycosylated and is released in the process of mutation into cancer. In contrast, the beta domain is anchored to the cell membrane and faces the cytoplasm [
4].
The C-terminal domain of MUC1 (MUC1-C) serves as a crucial molecule in the cancerogenic process through its interactions with several signaling receptors, especially receptor tyrosine kinases such as all four members of the human epidermal growth factor receptor (HER) family receptors, the fibroblast growth factor receptor-3 (FGFR-3), and the platelet-derived growth factor β (PDGF β) [
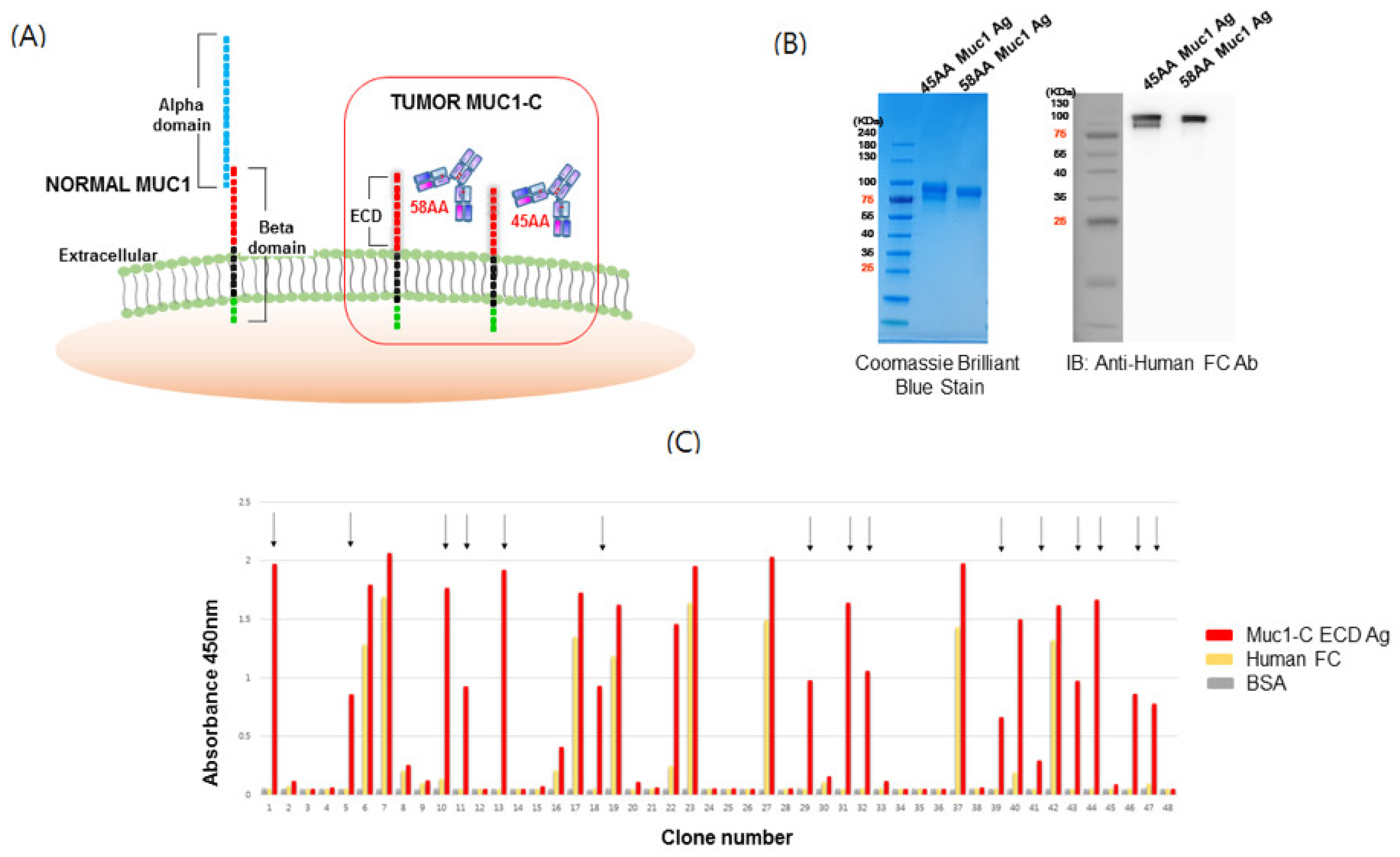
5]. MUC1-C consists of a small extracellular domain (ECD), a transmembrane domain (TM), and a cytoplasmic domain (CD) with 58, 28, and 72 amino acids, respectively. Galectin-3 binds to the N-glycosylation site at MUC1-C and acts as a bridge for receptor tyrosine kinases (RTKs) [
6]. The MUC1-C cytoplasmic domain plays a role in its intrinsic homodimerization and direct interactions with mediators of several signaling pathways. It is known that MUC1-C homodimers can be transported to the nucleus and mitochondria, resulting in diverse cellular behaviors such as inefficient transcription of p53 genes and the inhibition of apoptosis due to the direct binding of MUC1-C and p53 [
7]; the inhibition of cell adhesion and migration resulting from the reduction of the E-cadherin/β-catenin complex [
8,
9,
10,
11]; and the loss of apical-basal polarity in breast cancer cells. Also, recent reports suggest that MUC1-C-dependent gene signatures are associated with poor outcomes in breast and lung cancers, reinforcing the importance of MUC1-C in cancer transformation [
12]. There are various truncated forms of MUC1, and MUC1-C* is highly expressed in cancer. The release of pro-inflammatory cytokines such as interferon-gamma and tumor necrosis factor-alpha is known to activate several proteases (ADAM17, MMPs), whereby MUC1 is proteolytically cleaved into small extracellular domains, MUC1-C* [
13,
14]. MUC1-C* contains 45 amino acids (AAs) ECD (extracellular domains), and acts as a growth factor receptor for metastasis-associated proteins [
15,
16].
With all these complex functions and diverse interaction partners, MUC1-C is one of the key oncogenic receptors; it contributes to the loss of polarity in cancer cells, cancer stemness, epithelial-mesenchymal transition, epigenetic programming of cancer cells, and functions as an immune suppressor by induction of PD-L1 (programmed death-ligand 1) expression [
17,
18,
19]. Therefore, the inhibition of MUC1-C function has been suggested as an effective anticancer strategy in diverse cancers [
20,
21]. Therapeutic strategies targeting MUC1-C are predominantly small molecules capable of penetrating into cells, for example, PMIP, a peptide that mimics the interaction domain of MUC1 with both epidermal growth factor receptor (EGFR) and beta-catenin [
22]. Other candidates, peptide GO-201 and GO-202, block the oligomerization of MUC1-beta, and thereby prevent its nuclear and mitochondrial import. These two small molecules have been shown to induce growth arrest and death of human breast cancer cells [
23]. However, the toxicity of PMIP and GO-201 peptides and their potential immunogenicity need to be investigated. A recent study used MUC1 ECD and part of alpha domain as an antigen to develop antibodies with specific anti-MUC1 activities. Most of the strategies employing therapeutic antibodies have targeted the variable number tandem repeat (VNTR) region and altered O-glycosylation sites of cancerous MUC1, and one of them failed in the phase II clinical study because of a lack of efficacy in breast cancer; however, the use of the AS1402 antibody as a drug candidate accelerated cancer research [
24].
Antibodies that accurately target the extracellular domain of MUC1-C have not yet been developed clinically. So, we generated a recombinant MUC1-C ECD, which is critical for tumor progression, and a shorter antigen MUC1-C*. Phage display techniques were used to select antibodies that bind to the MUC1-C antigen and the surfaces of the MUC1-overexpressing cells. We confirmed their binding specificities via surface plasmon resonance (SPR), fluorescence-activated cell sorting (FACS), and confocal image analysis and compared them to those of the MUC1-C-targeting control antibody. We also evaluated the functionalities of these antibodies on the growth and invasion of MUC1-expressing breast cancer cell lines. Later, we examined the thermal stability and binding capacity of the selected antibodies as well. To understand MUC1 biology, which has diverse functions in tumor microenvironments [
25], antibodies that recognize MUC1-C represent essential tools. In this study, we found that our antibodies specifically bind to MUC1-C and explored their therapeutic functionalities.
3. Discussion
We generated antigens mimicking the ECD of MUC1-C (58 AA) and MUC1-C* (45 AA) as human Fc-fused forms to leverage their natural conformation in antibody screening, and antibody phage panning was performed to isolate MUC1-C specific antibodies. Phage display antibody panning resulted in the isolation of 24 human antibodies with binding specificities against the MUC1-C antigen. To confirm their specificities and functionalities, they were produced as full-sized IgG1 antibodies and used in several immunological assays. For functional reference, the MIN-C2 antibody with specific binding to MUC1-C was used in every assay.
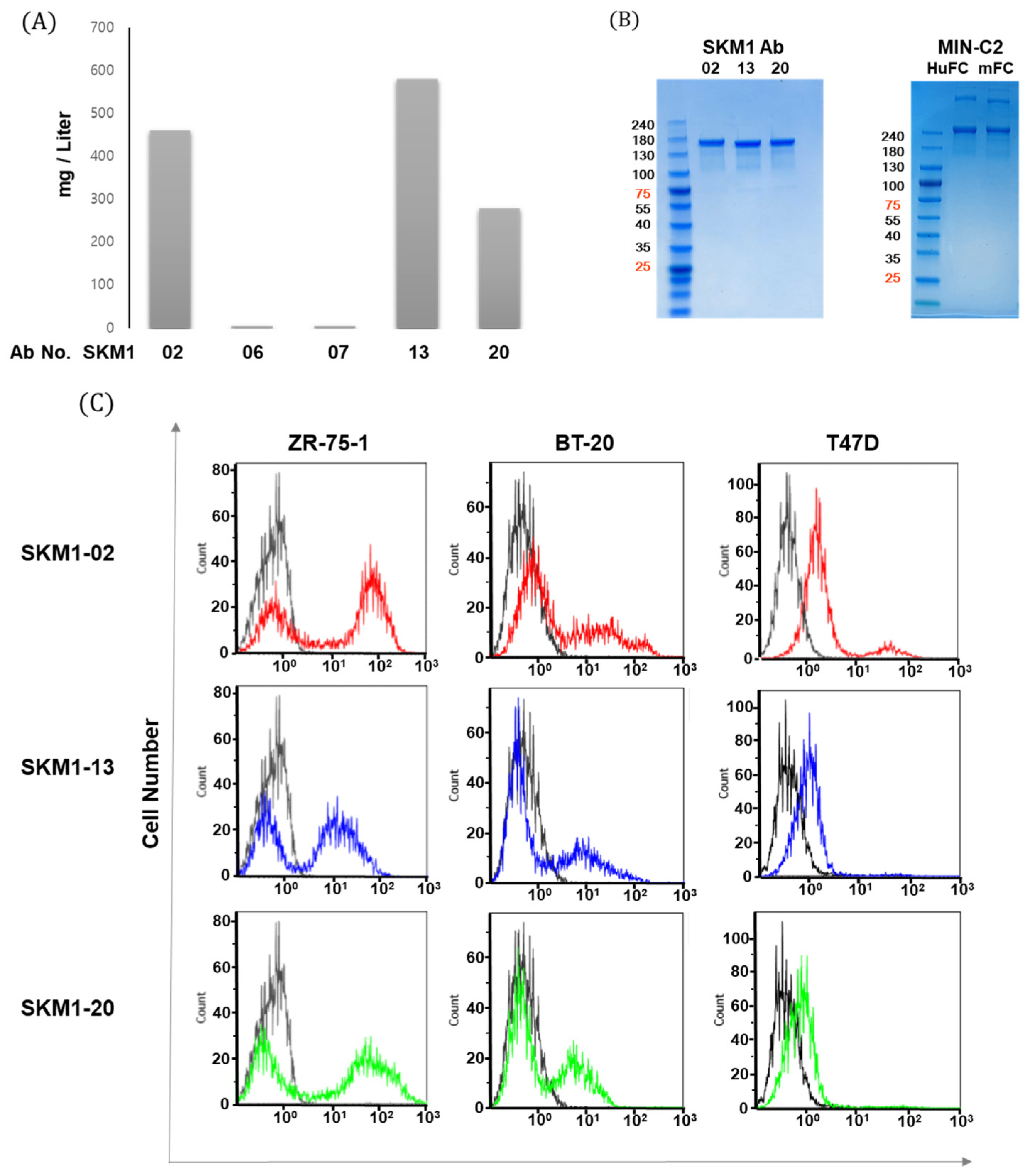
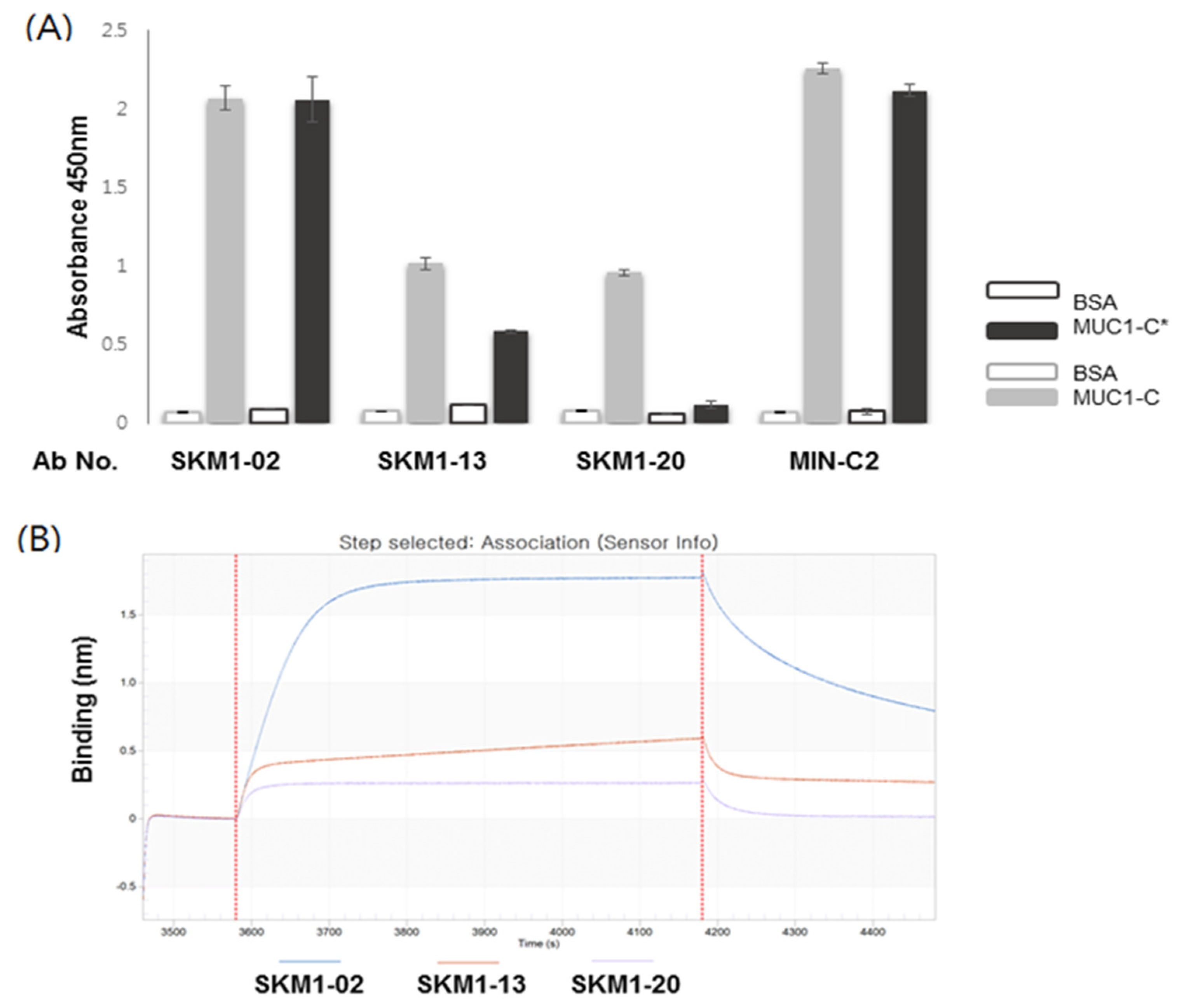
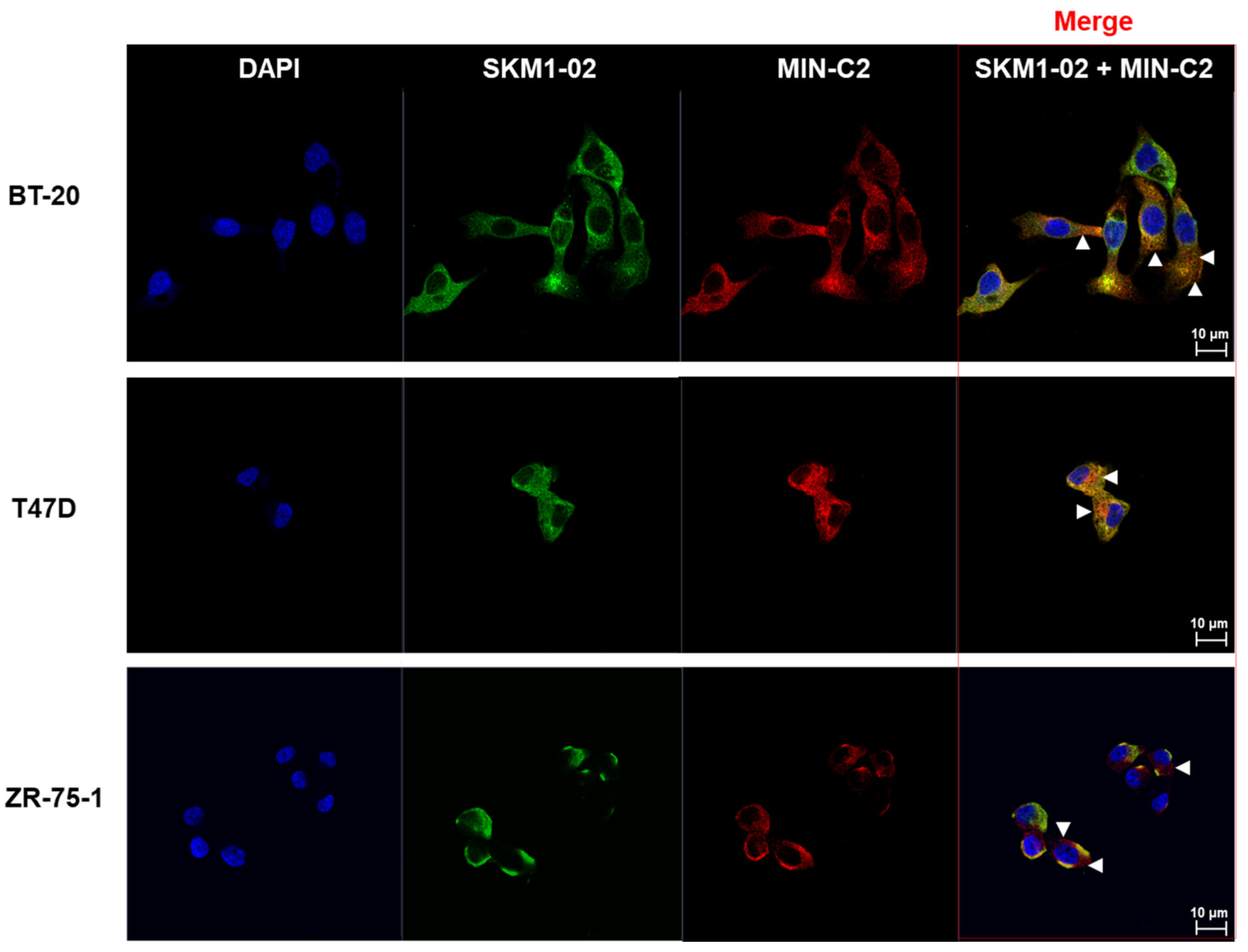
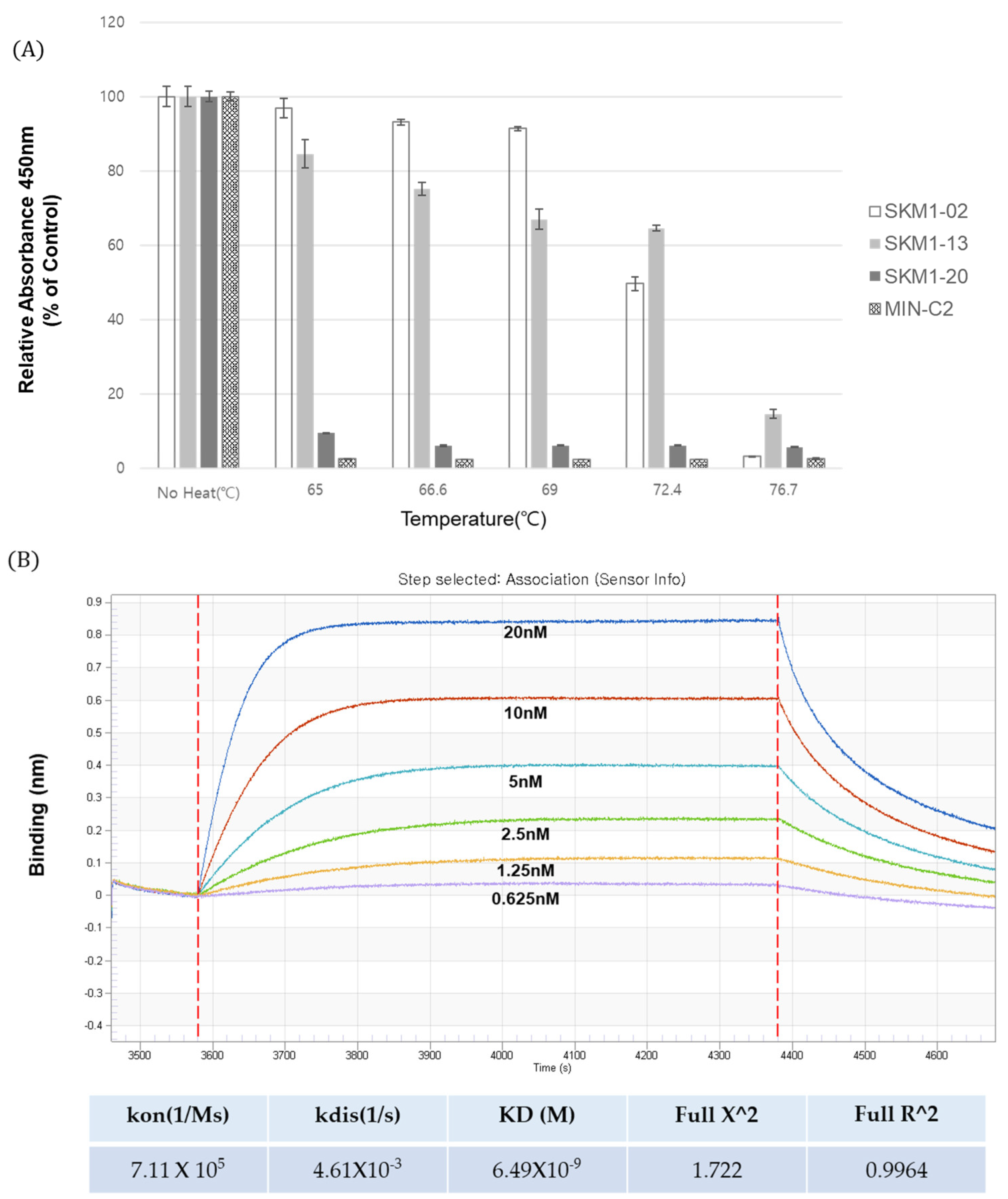
The top three clones including SKM1-02, -13, and -20 with a relatively high level of expression in a mammalian cell system reconfirmed MUC1-C-specific binding as full-size IgG in several immunological assays, such as ELISA, FACS assay, and BLI using the Octet system. Interestingly, SKM1-02 Ab showed similar binding with the 58 AA ECD and 45 AA ECD of MUC1-C; however, the -13 and -20 clones showed slightly different bindings with two antigens; the -20 clone showed an almost exclusive binding to the 58 AA ECD in the ELISA. Octet analysis also showed the same results as the ELISA. Confocal image analysis also showed the overlapped staining of SKM1-02 and MIN-C2 antibodies, which suggested that our MUC1-C-targeting antibodies bind to the MUC1 beta ECD domain and each antibody has slightly different epitopes within the MUC1-C ECD domains. The effects on MUC1-C function due to differences of the antibodies’ epitopes are not clear yet. However, since the results of the migration and growth inhibition assay showed certain differences between the antibodies, it is necessary to study more detailed mechanisms between these antibodies and MUC1-C.
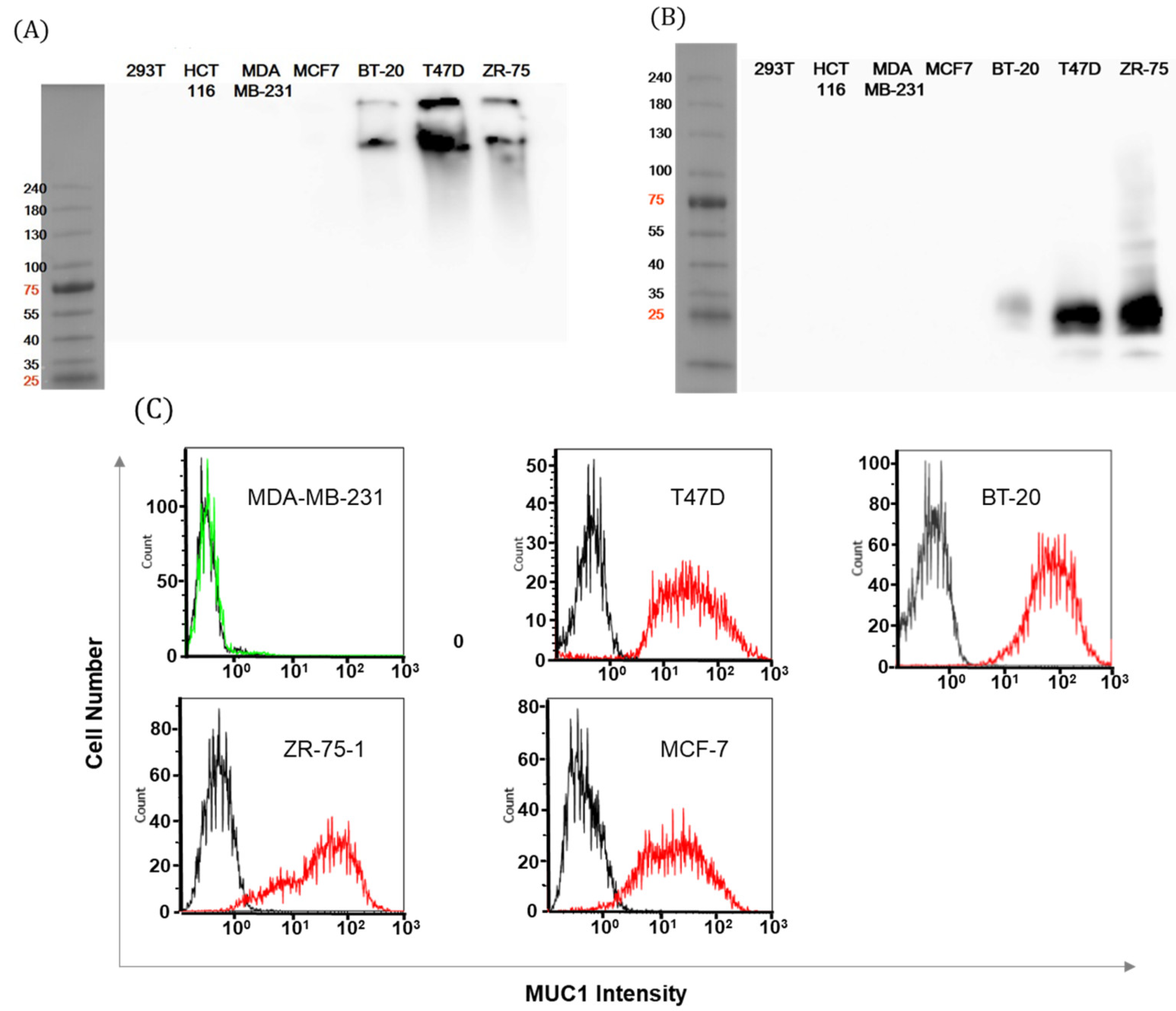
In addition, the MUC1-C binding antibodies to some cancer cells resulted in two separate peaks. The T47D cell line showed no peak separation but ZR-75-1 and BT-20 cell lines showed dual peaks. This dual peak staining pattern was also shown from the FACS staining of the reference antibody, MIN-C2. We tried the characterization of individual populations through cellular sorting by a BD FACSAria instrument, which revealed that the two distinct peaks reappeared from each population (data not shown), which suggested that these peaks represent two groups of cells engaged in cell proliferation. Further, the two distinct peaks may represent a mixture of cancer stem cells and normal cancer cells; however, no match was found with stem cell markers. Further experiments are needed to understand the high and low peaks in the MUC1-C-specific antibody staining.
No antibody targeting of MUC1 has been successful in clinical studies so far, most of which targeted the alpha domain of MUC1, because cancer-specific underglycosylated MUC1 plays a significant role in carcinogenesis and most of the antibodies obtained using hybridoma approaches against MUC1 generated alpha-domain-targeting antibodies. Since the first pioneered candidate AS1402 failed in a phase II clinical study because of its cancer-promoting effect [
24], alternative approaches such as drug-conjugates, peptides, or vaccines have been investigated, but few have used MUC1-C*-targeting monoclonal antibodies.
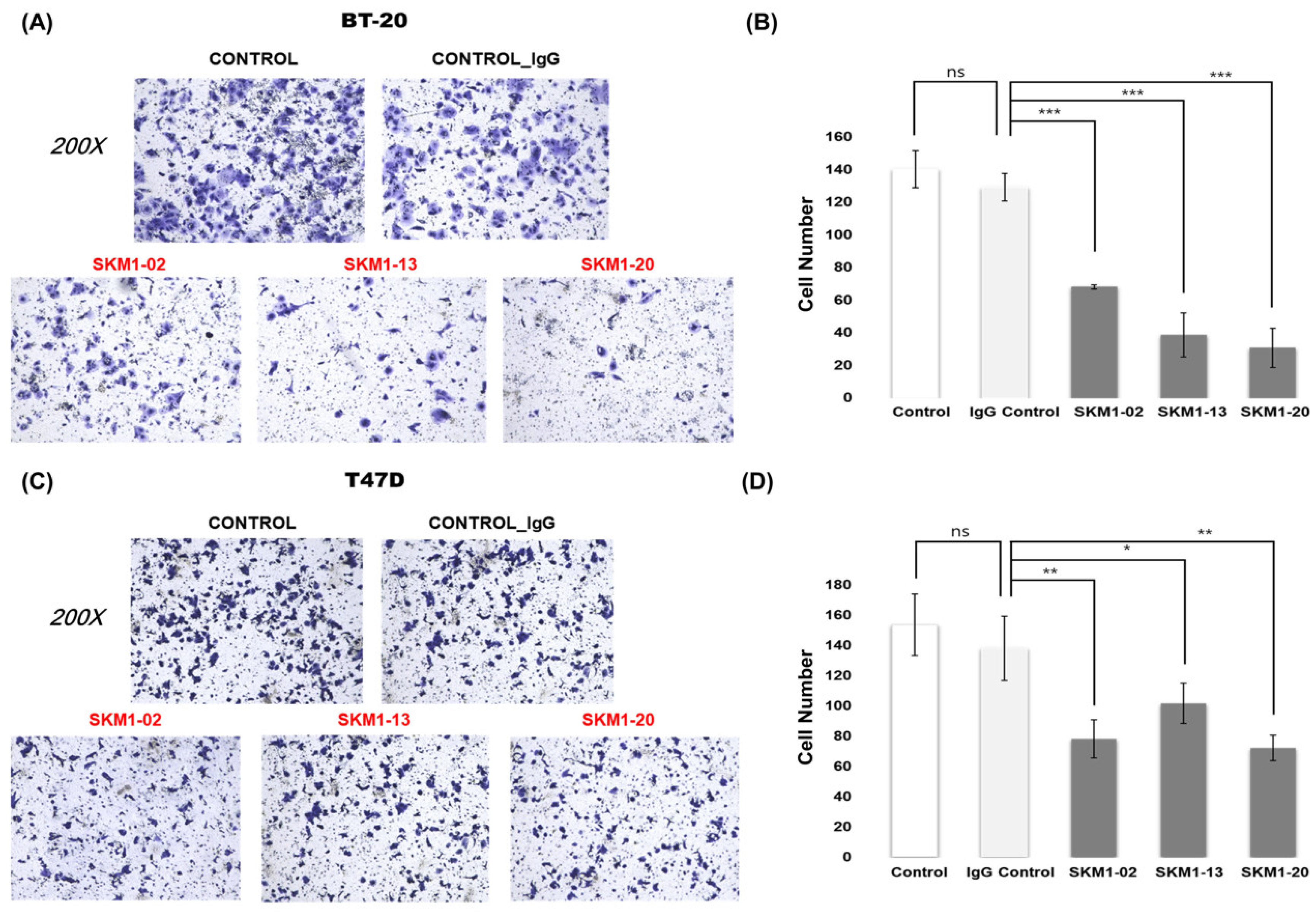
To address the potential of therapeutic antibody development targeting the MUC1-C extracellular domain, we tested the functionality of the top three clones in assays for cancer cell invasion and cancer growth inhibition. All three antibodies showed high levels of migration inhibition, which varied in cell lines and individual antibodies. It is known that MUC1-C translocation to the nucleus in association with beta-catenin represses E-cadherin expression, which then destabilizes the adherens junctions and induces cytoskeletal rearrangement resulting in loss of cell–cell contact inhibition. Therefore, MUC1-C antibody treatment affects all these events and leads to anti-metastatic outcomes [
31]. The results of the invasion assay against the BT-20 cell line (TNBC, triple-negative breast cancer) and T47D cell line (her2 negative cell line) showed that these antibodies represent potential anti-metastatic therapeutic candidates. In addition, the proliferation assay for SKM1-02 showed promising results. Because MUC1-C is involved in multiple mechanisms of cell proliferation, such as extracellular bridging through galectin 3 and RTKs, homodimer formation and interaction with signal mediators at cytoplasmic domain, and activation of transcription factors during the nuclear and mitochondrial translocation of MUC1-C, the results of the SKM1-02 proliferation inhibition assay cannot be explained currently; however, based on one of these mechanisms, it exerts growth inhibition. From a drug development perspective, with a dissociation constant of 6.5 nM against MUC1-C and a binding capability even at 72.4 °C, SKM1-02 represents a potential antibody drug candidate.
Currently, several antibodies against cancer-specific antigens are being tested on the CAR-T cell platform, which showed that the epitope of the antibody is one of the key factors contributing to robust anticancer activities. The MUC1 antigen has been analyzed comprehensively in CAR-T therapy, but the antibody used in CAR-T cell targeting yielded different efficacies. Therefore, MUC1-C-targeting antibodies such as SKM1-02 that contain membrane-proximal epitopes for CAR-T cells represent the optimum antibody candidate for the development of CAR-T therapy.
4. Materials and Methods
4.1. Phage Display
Phage display experiments were mainly performed with the protocols that are well-described in detail in the “Phage Display: A Laboratory Manual” [
32]. Briefly, biopanning for the enrichment of antibody binders was performed using immunotubes (maxisorp, Nunc, Rochester, NY, USA) or magnetic beads (Dynabeads M-270 epoxy, Invitrogen, Invitrogen, Carlsbad, CA, USA) that were either coated with human normal IgG or recombinant MUC1-C protein. Phages were pre-adsorbed on human IgG coated beads to remove human Fc binders, then the phages in the supernatant were incubated at 4 °C for 1 h with immobilized antigens. The phage-antigens were washed 5 times with 0.1% Tween 20/PBS and the bound phages were eluted with 0.25% trypsin (Gibco, Waltham, MA, USA). The eluted phages were amplified by direct infection to
E. coli ER2738 in the exponential growth phase. The phages from an overnight culture were purified by PEG precipitation (final 5%
w/
v polyethylene glycol 8000 (PEG8000), 0.5 M NaCl) and used in the next panning cycle. Four rounds of panning were performed to enrich the antibody binders. Individual colonies from the 3rd and 4th rounds of panning were picked randomly and tested by enzyme-linked immunosorbent assay (ELISA) for MUC1-C antigen binding and the clones with positive ELISA signal were applied for DNA sequencing.
4.2. ScFv ELISA
A total of 600 phage clones were tested for reactivity with MUC1-C and MUC1-C* by ELISA. Single colonies from 3rd and 4th panning output plates were cultured in a 96-deep-well plate (300 μL media/well with ampicillin) with 1M IPTG induction and shaking at 25 °C overnight. The next day, bacterial cells were spun down, and the supernatant was removed. The pellet was suspended with 80 μL ice-cold 1X TES buffer (20% w/v sucrose, 50 mM Tris, 1 mM EDTA, pH 8.0) and incubated on ice for 30 min. Then, 120 μL of ice-cold 0.2X TES buffer was added to each well, mixed thoroughly, and incubated on ice for 30 min. After spinning down, the supernatant contained periplasmic fraction. Periplasmic extracts containing scFv were added to ELISA plates coated with 2 µg/mL antigen or control human normal IgG in phosphate-buffered saline (PBS). The plate was incubated at room temperature for 1 h and washed 5 times with 0.05% Tween/PBS, followed by incubation with anti-Myc antibody (3E10 clone) in 5% skim milk/PBS for 1 hr. After washing and incubation with anti-mouse IgG-HRP secondary antibody for 1hr, the well with antibody binding was visualized with a general TMB substrate colorization method.
4.3. Immunoblotting
Cells were lysed in RIPA buffer (50 mM Tris-HCl, 150 mM NaCl, 1% NP-40, and 0.25% Na-deoxycholate) with a protease and phosphatase inhibitor cocktail (Genedepot, Katy, TX, USA). The concentration of total protein was determined using the BCA Assay kit (Thermo Scientific, Rockford, IL, USA). Whole-cell lysates were separated by SDS-PAGE and transferred to PVDF membranes (Bio-Rad, Hercules, CA, USA). The primary antibodies used were anti-MUC1 (BD Bioscience, Franklin Lakes, NJ, USA) and anti-MUC1-C (AbCam, Cambridge, UK). Antibodies were routinely used at 1∶1000 dilution with 5% skim milk/PBS for primary antigen binding, then secondary anti-mouse IgG conjugated with horseradish peroxidase were incubated at 1:5000 dilution with 5% skim milk/PBS. This immunoblot was visualized with a chemiluminescent Western blot detection kit (Ab Frontier, Seoul, Korea) and Luminescent Image Analyzer (GE Healthcare Bio-sciences, Chicago, IL, USA) according to the manufacturer’s instructions.
4.4. SPR (OCTET)
Binding affinity of the anti-hMUC1 monoclonal antibody was determined using an Octet RED96 system (ForteBio, Pall Life Sciences, Westborough, MA, USA) and the experiment was performed in accordance to manufacturer’s protocol. All samples were diluted in running buffer. Wells of a black flat bottom polypropylene plate were loaded with each sample and the reaction buffers. Recombinant hMUC1-antigen was immobilized onto amine-reactive biosensors (AR2G, ForteBio, Fremont, CA, USA). The MUC1-C antibody was serially diluted two-fold (0.626–20 nM) in a kinetic buffer. Kinetic parameters and dissociation constant of the antibody were calculated with data analysis software (ver9) provided by the Octet system of ForteBio, in which antigen/antibody binding was modeled as 1:2 binding.
4.5. Cell Culture
Cancer cell lines were maintained at 37 °C with 5% CO2. ZR-75-1 was obtained from American Type Culture Collection (Manassas, VA, USA) and T47D, BT-20, MCF-7, HCT116, and MDA-MB-231 were obtained from KCLB (Korean Cell Bank, Seoul, Korea). ZR-75-1 and T47D cell lines were cultured in RPMI 1640 (Hyclone, Logan, UT, USA) medium supplemented with 100 units/mL penicillin, 100 μg/mL streptomycin, and 2 mmol/L l-glutamine. All the above cell culture reagents were purchased from Invitrogen (Carlsbad, CA, USA). BT-20 (Korean Cell Bank, Seoul, Korea) cells were cultured in Eagle’s minimum essential medium (Gibco, Gaithersburg, MD, USA) supplemented with 10% (v/v) fetal bovine serum (FBS; Gibco) as guided by ATCC. HEK293F cells were cultured in FreestyleTM expression medium (Gibco) using Multitron incubation shaker (Infors HT, Bottmingen, Switzerland) with 8% CO2, 37 °C.
4.6. Flow Cytometry (FACS)
Flow cytometry was performed in a FACS calibur (Becton-Dickinson, BD Biosciences) equipped with a 488 nm argon laser and a 615 nm red diode laser. Cells were harvested and cleaned with FACS buffer (2% FBS in PBS). The cells were incubated with the recommended IgG (4 µg/mL) for 1 h at room temperature. After washing the unbound antibody, followed by incubation with Alexa Fluor 488-conjugated goat anti-Human IgG antibody or 647 conjugated goat anti-mouse IgG antibody for 1 h, the samples were analyzed using Kaluza software (Beckman Coulter, Brea, CA, USA).
4.7. ELISA
A 50 µl aliquot of antigen (2 µg/mL) in PBS buffer was added to the individual wells of a microtiter plate and incubated overnight at 4 °C. The coating solution was removed and the plate was washed three times by filling the wells with 100 μL PBS containing 0.05% Tween 20 (PBS-T). The remaining protein-binding sites in the coated wells were blocked by adding 100 μL of the blocking buffer, 3% BSA in PBS to each well, and incubated for 1 h at RT with gentle shaking. The plates were then incubated with 1 µg of IgG or anti-MUC1-C clones in blocking buffer for 1 h at 37 °C, then were washed three times with PBS-T, followed by secondary antibody incubation with (HRP)-conjugated goat anti-human IgG antibody for 1 h. The bound antibody was detected by the addition of the 3,3′,5,5′-tetramethylbenzidine substrate solution (TMB; BD Biosciences, San Jose, CA, USA). After incubation for 5 min, the color development was stopped with 1N HCl. The absorbance (optical density at 450 nm) of each well was determined with a plate reader.
4.8. Proliferation Assay
A total of approximately 4 × 103 cells were plated in 96-well plates in triplicate and cultured in 90 μL medium containing 10% FBS. After 1, 3, 6, and 9 days of incubation with monoclonal antibody, a 10 μL aliquot of cell proliferation assay reagent from Cell Counting Kit-8 (Dojindo, Kumamoto, Japan) was added to each well and they were left for 1–4 hrs of additional incubation in a CO2 incubator. Final absorbance was measured at 450 nm using the microplate reader Epoch (Thermo Fisher, Waltham, MA, USA). Each experiment was repeated three times.
4.9. Purification of Recombinant Human MUC1-C Protein and Full IgG Antibodies
The human Muc1 antigen containing 45 and 58 amino acids (45 AA: Glycine from 1110 to alanine 1154, 58 AA: serine from 1097 to alanine 1154; GenBank Accession No. P15941) was synthesized and amplified by PCR using the following primer sets: 58 AA sense 5ʹ-AGGCCCAGGCGGCCTCTGTGGTGGTACAA-3′ and anti-sense 5ʹ-AGGCCAGCCAGGCCCCCAGCCCCAGACTG-3′, 45 AA sense 5ʹ-GGGGCCC AGGCGGCCGGTACCATCAATGTCCACGAC-3′ and anti-sense 5ʹ- GCTGGCCAGCCAGGCCAGC CCCAGACTGGGCAGAG-3′. Using the Sfi I restriction enzyme, the amplified DNA fragments were cloned into the expression vector pCEP4-FC. The MUC1-pCEP4 vector was transiently expressed in 293F cells and after 12 days, the culture medium was harvested.
Each set of VH and VL primers for the 24 MUC1-C binding clones were synthesized and used for PCR-amplification of individual VH and VL fragments (
Table S2). Each VH primer was designed to have restriction sites for KpnI or BamHI at 5′ ends and NheI at 3′ ends. For VL primers, BamHI and BsiWI restriction sites were incorporated at the 5′ and 3′ ends, respectively. VH and VL fragments of each clone were subcloned into the pCEP4 mammalian expression vector (Invitrogen) as a fused form either with CH1-CH2-CH3 for heavy chain expression or with C
kappa for light chain expression. Matching heavy and light chain vectors for each clone were co-transfected into Expi-CHO cells according to manufacturer’s protocol and after 12 days, the culture medium was harvested. The MUC1 antigens and full IgG antibodies were purified by affinity column chromatography using protein A agarose beads (GenScript, Piscataway, NJ, USA).
After dialysis with phosphate-buffered saline at pH 7.4 (PBS), antibody concentration was quantified using a NanoDrop 2000 spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA), and the purity of each antibody was evaluated by SDS/polyacrylamide gel electrophoresis (PAGE) and Coomassie brilliant blue staining.
4.10. Immunocytochemistry and Confocal Microscopy
The cells were seeded on sterile glass coverslips in 12-well culture plates and incubated in freshly prepared 4% paraformaldehyde-PBS at room temperature for 10 min. The MUC1-C antibody dilutions were prepared in 1% normal BSA. The coverslips were incubated for 1 h at room temperature. After rinsing the unbound monoclonal antibody, the slides were incubated with Alexa Fluor-conjugated secondary antibodies, such as Alexa Fluor 488-conjugated goat anti-Human IgG antibody or 647 conjugated goat anti-mouse IgG antibody, for 1 h. All the samples were mounted and observed using confocal laser scanning microscopy (CLSM system; LSM 700, Carl Zeiss, Jena, Germany).
4.11. Invasion Assay
BD Matrigel Basement Membrane Matrix (BD BioScience, concentration approx. 10 mg/mL) was mixed 1∶4 with PBS and allowed to polymerize in transwell inserts (Corning, Corning, NY, USA) overnight and 3×105 cells were seeded directly onto the Matrigel-coated inserts. Transwell inserts were finally placed in serum-free medium, and the lower compartment was filled with a medium supplemented with 10% FBS (to obtain a chemotactic gradient). Human MUC1-C candidate Abs were added to the upper compartment of the invasion chamber, and then incubated overnight. At least three independent experiments were performed in duplicate for each sample. After incubation for 48 h, the chambers were washed with phosphate-buffered saline (PBS) and fixed with 4% paraformaldehyde in PBS. Samples were then washed and stained with crystal violet. The cells passing through the filter into the lower chamber were counted using a phase-contrast microscope.