mRNA with Mammalian Codon Bias Accumulates in Yeast Mutants with Constitutive Stress Granules
Abstract
1. Introduction
2. Results
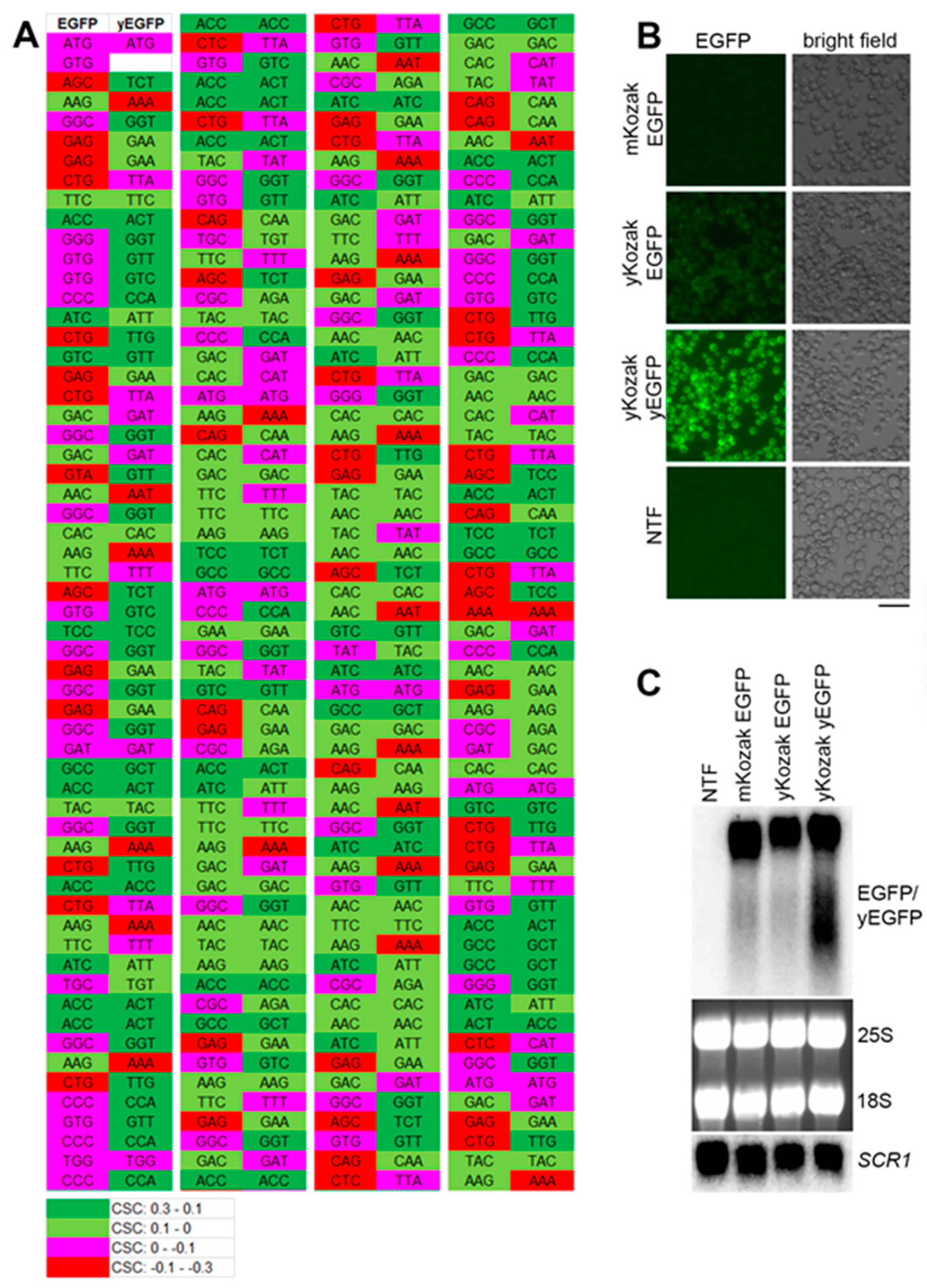
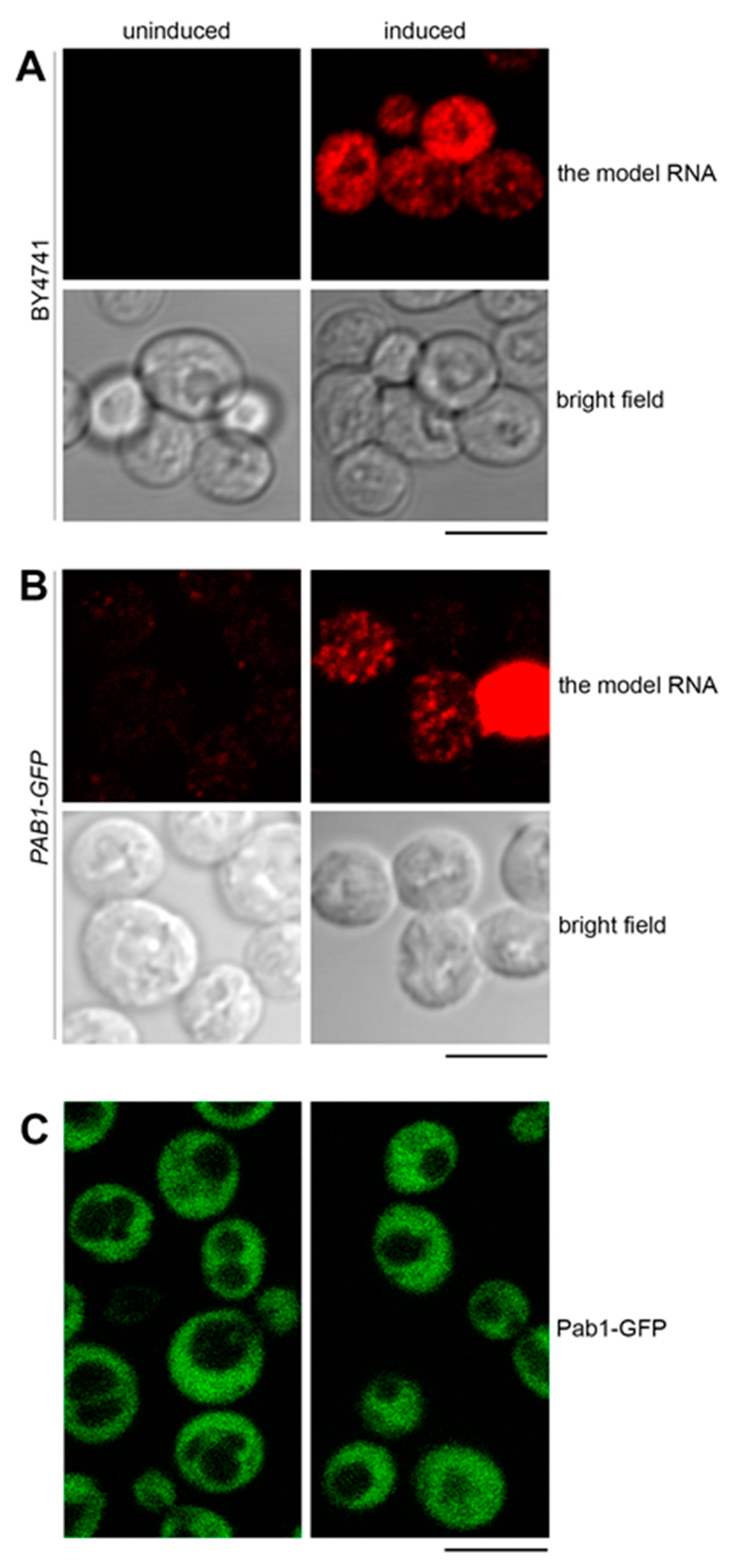
2.1. The Model RNA is not Downregulated due to Its Suboptimal Codon Bias
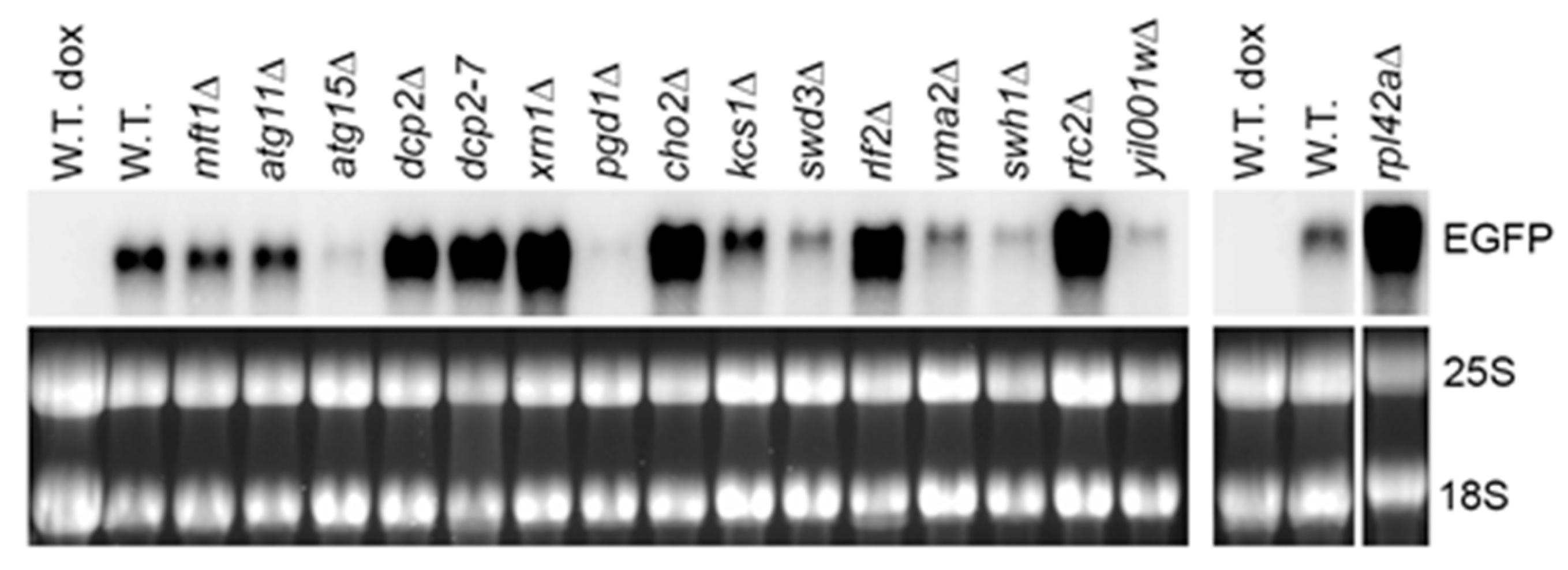
2.2. The Model RNA Accumulates in Yeast Mutants with Constitutive Stress Granules
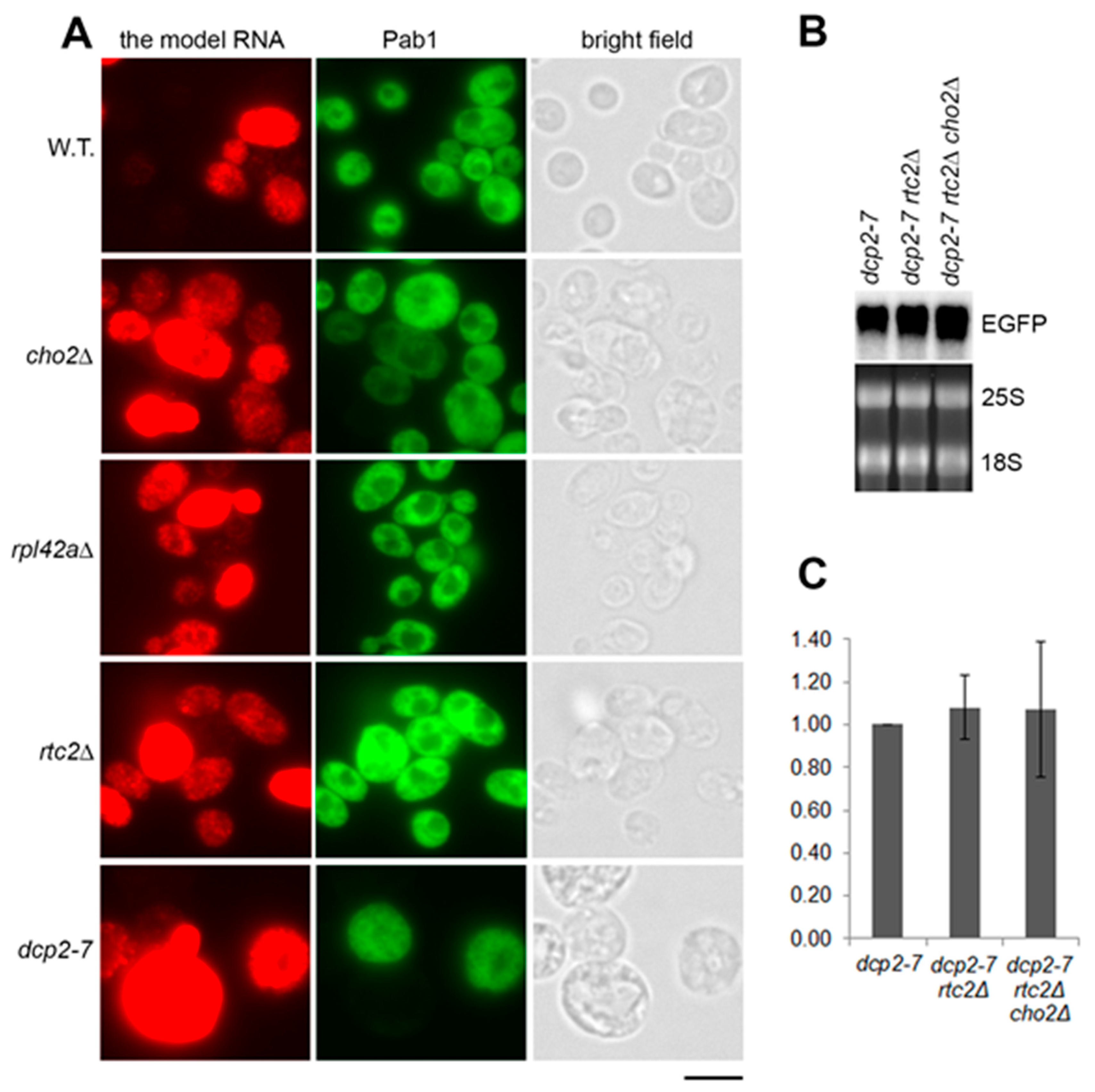
2.3. Accumulation of the Model RNA in rtc2Δ and cho2Δ Mutants Occurs due to Compromised Path Linked to the Activity of Dcp2
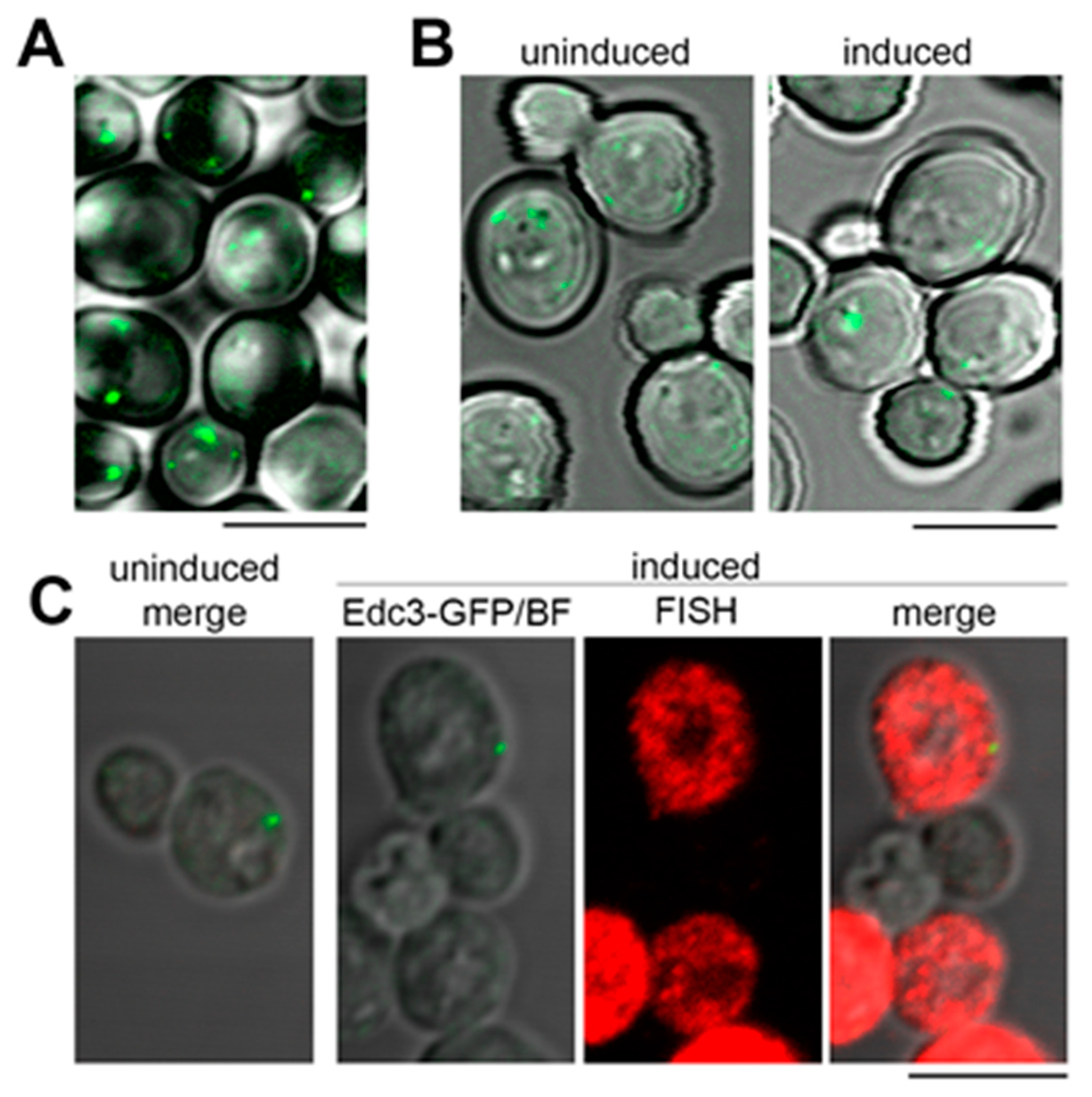
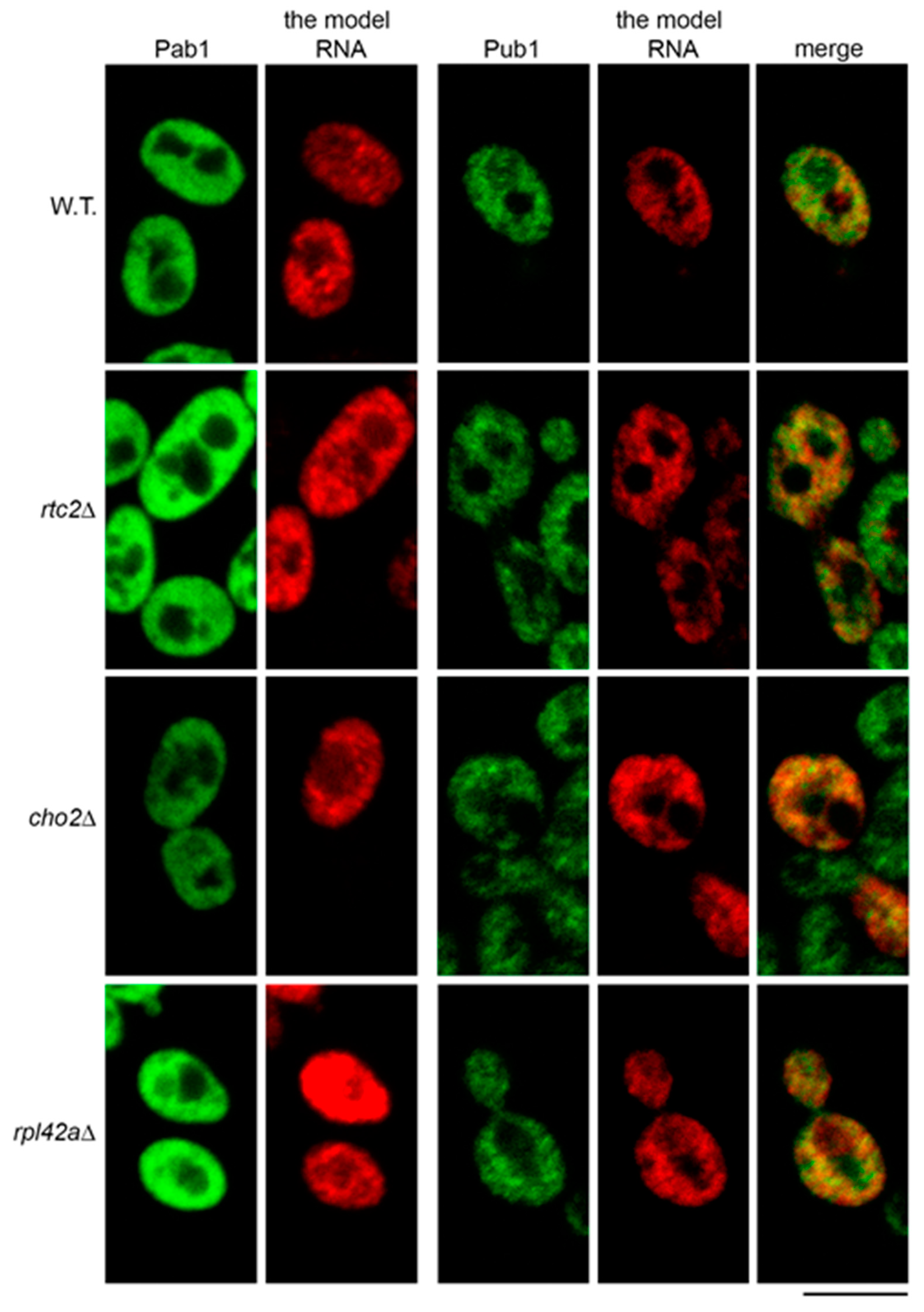
2.4. The Model RNA Forms Cytoplasmic Granules Distinct from Stress Granules and P Bodies
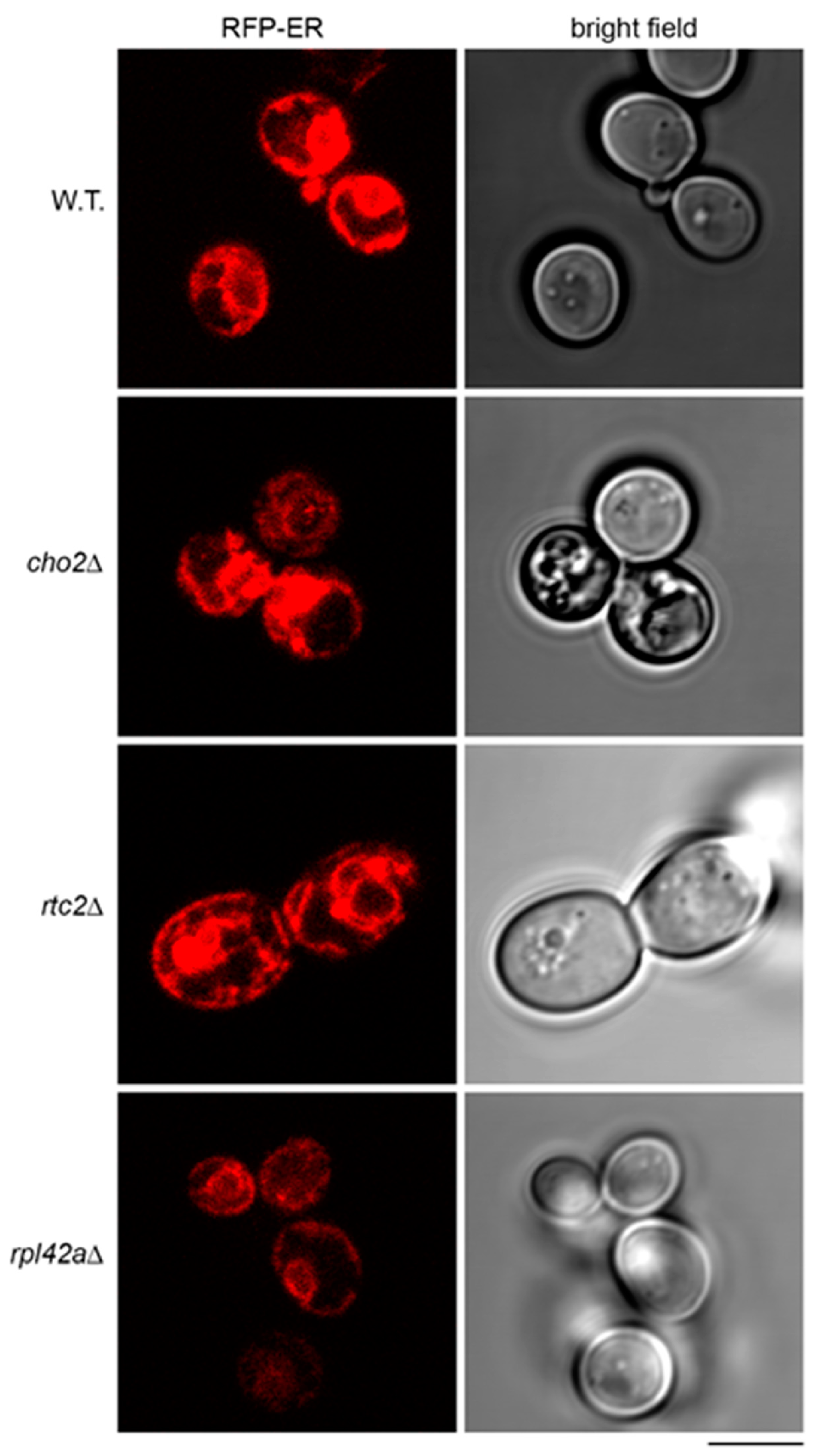
2.5. Accumulation of the Model RNA and Defects in Endoplasmic Reticulum
3. Discussion
4. Materials and Methods
4.1. Plasmids
4.2. Yeast Strains and Growth Conditions
4.3. Northern Blot
4.4. FISH and Immunofluorescence
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| mRNA | Messenger ribonucleic acid |
| PKA | Protein kinase A |
| HPLC | High-performance liquid chromatography |
| FPLC | Fast protein liquid chromatography |
| (y)(E)GFP | (yeast) (Enhanced) green fluorescent protein |
| ORF | Open reading frame |
| UTR | Untranslated region |
| CSC | Codon stabilization coefficient |
| NTF | Non-transformed |
| FISH | Fluorescent in situ hybridization |
| IF | Immunofluorescence |
| ER | Endoplasmic reticulum |
| RFP | Red fluorescent protein |
| PCR | Polymerase chain reaction |
| DNA | Deoxyribonucleic acid |
| FCC | Frozen competent cells |
| PBS | Phosphate-buffered saline |
| VRC | Ribonucleoside vanadyl complex |
| LSM | Laser scanning microscope |
References
- Ashe, M.P.; De Long, S.K.; Sachs, A.B. Glucose depletion rapidly inhibits translation initiation in yeast. Mol. Biol. Cell 2000, 11, 833–848. [Google Scholar] [CrossRef] [PubMed]
- Buchan, J.R.; Muhlrad, D.; Parker, R. P bodies promote stress granule assembly in Saccharomyces cerevisiae. J. Cell Biol. 2008, 183, 441–455. [Google Scholar] [CrossRef] [PubMed]
- Zid, B.M.; O’Shea, E.K. Promoter sequences direct cytoplasmic localization and translation of mRNAs during starvation in yeast. Nature 2014, 514, 117–121. [Google Scholar] [CrossRef] [PubMed]
- Teixeira, D.; Sheth, U.; Valencia-Sanchez, M.A.; Brengues, M.; Parker, R. Processing bodies require RNA for assembly and contain nontranslating mRNAs. RNA 2005, 11, 371–382. [Google Scholar] [CrossRef]
- Buchan, J.R.; Parker, R. Eukaryotic stress granules: The ins and outs of translation. Mol. Cell 2009, 36, 932–941. [Google Scholar] [CrossRef]
- Decker, C.J.; Parker, R. P-bodies and stress granules: Possible roles in the control of translation and mRNA degradation. Cold Spring Harb. Perspect. Biol. 2012, 4, a012286. [Google Scholar] [CrossRef]
- Yoon, J.H.; Choi, E.J.; Parker, R. Dcp2 phosphorylation by Ste20 modulates stress granule assembly and mRNA decay in Saccharomyces cerevisiae. J. Cell Biol. 2010, 189, 813–827. [Google Scholar] [CrossRef]
- Teixeira, D.; Parker, R. Analysis of P-body assembly in Saccharomyces cerevisiae. Mol. Biol. Cell 2007, 18, 2274–2287. [Google Scholar] [CrossRef]
- Buchan, J.R.; Yoon, J.H.; Parker, R. Stress-specific composition, assembly and kinetics of stress granules in Saccharomyces cerevisiae. J. Cell Sci. 2011, 124, 228–239. [Google Scholar] [CrossRef]
- Grousl, T.; Ivanov, P.; Frydlova, I.; Vasicova, P.; Janda, F.; Vojtova, J.; Malinska, K.; Malcova, I.; Novakova, L.; Janoskova, D.; et al. Robust heat shock induces eIF2alpha-phosphorylation-independent assembly of stress granules containing eIF3 and 40S ribosomal subunits in budding yeast, Saccharomyces cerevisiae. J. Cell Sci. 2009, 122, 2078–2088. [Google Scholar] [CrossRef]
- Kilchert, C.; Weidner, J.; Prescianotto-Baschong, C.; Spang, A. Defects in the secretory pathway and high Ca2+ induce multiple P-bodies. Mol. Biol. Cell 2010, 21, 2624–2638. [Google Scholar] [CrossRef] [PubMed]
- Shah, K.H.; Zhang, B.; Ramachandran, V.; Herman, P.K. Processing body and stress granule assembly occur by independent and differentially regulated pathways in Saccharomyces cerevisiae. Genetics 2013, 193, 109–123. [Google Scholar] [CrossRef] [PubMed]
- Ramachandran, V.; Shah, K.H.; Herman, P.K. The cAMP-dependent protein kinase signaling pathway is a key regulator of P body foci formation. Mol. Cell 2011, 43, 973–981. [Google Scholar] [CrossRef] [PubMed]
- Hilgers, V.; Teixeira, D.; Parker, R. Translation-independent inhibition of mRNA deadenylation during stress in Saccharomyces cerevisiae. RNA 2006, 12, 1835–1845. [Google Scholar] [CrossRef]
- Jona, G.; Choder, M.; Gileadi, O. Glucose starvation induces a drastic reduction in the rates of both transcription and degradation of mRNA in yeast. Biochim. Biophys. Acta 2000, 1491, 37–48. [Google Scholar] [CrossRef]
- Greatrix, B.W.; van Vuuren, H.J. Expression of the HXT13, HXT15 and HXT17 genes in Saccharomyces cerevisiae and stabilization of the HXT1 gene transcript by sugar-induced osmotic stress. Curr. Genet. 2006, 49, 205–217. [Google Scholar] [CrossRef]
- Romero-Santacreu, L.; Moreno, J.; Perez-Ortin, J.E.; Alepuz, P. Specific and global regulation of mRNA stability during osmotic stress in Saccharomyces cerevisiae. RNA 2009, 15, 1110–1120. [Google Scholar] [CrossRef]
- Garcia-Martinez, J.; Aranda, A.; Perez-Ortin, J.E. Genomic run-on evaluates transcription rates for all yeast genes and identifies gene regulatory mechanisms. Mol. Cell 2004, 15, 303–313. [Google Scholar] [CrossRef]
- Molina-Navarro, M.M.; Castells-Roca, L.; Belli, G.; Garcia-Martinez, J.; Marin-Navarro, J.; Moreno, J.; Perez-Ortin, J.E.; Herrero, E. Comprehensive transcriptional analysis of the oxidative response in yeast. J. Biol. Chem. 2008, 283, 17908–17918. [Google Scholar] [CrossRef]
- Buchan, J.R.; Kolaitis, R.M.; Taylor, J.P.; Parker, R. Eukaryotic stress granules are cleared by autophagy and Cdc48/VCP function. Cell 2013, 153, 1461–1474. [Google Scholar] [CrossRef]
- Pardi, N.; Hogan, M.J.; Porter, F.W.; Weissman, D. mRNA vaccines—A new era in vaccinology. Nat. Rev. Drug Discov. 2018, 17, 261–279. [Google Scholar] [CrossRef] [PubMed]
- Parker, R. RNA degradation in Saccharomyces cerevisae. Genetics 2012, 191, 671–702. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Z.; Reed, R. Purification of functional RNA-protein complexes using MS2-MBP. Curr. Protoc. Mol. Biol. 2003, 63, 27.3.1–27.3.7. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Z.; Sim, J.; Griffith, J.; Reed, R. Purification and electron microscopic visualization of functional human spliceosomes. Proc. Natl. Acad. Sci. USA 2002, 99, 12203–12207. [Google Scholar] [CrossRef] [PubMed]
- Presnyak, V.; Alhusaini, N.; Chen, Y.H.; Martin, S.; Morris, N.; Kline, N.; Olson, S.; Weinberg, D.; Baker, K.E.; Graveley, B.R.; et al. Codon optimality is a major determinant of mRNA stability. Cell 2015, 160, 1111–1124. [Google Scholar] [CrossRef] [PubMed]
- Sweet, T.; Kovalak, C.; Coller, J. The DEAD-box protein Dhh1 promotes decapping by slowing ribosome movement. PLoS Biol. 2012, 10, e1001342. [Google Scholar] [CrossRef]
- Cormack, B.P.; Bertram, G.; Egerton, M.; Gow, N.A.; Falkow, S.; Brown, A.J. Yeast-enhanced green fluorescent protein (yEGFP): A reporter of gene expression in Candida albicans. Microbiology 1997, 143 Pt 2, 303–311. [Google Scholar] [CrossRef]
- Eshleman, N.; Liu, G.; McGrath, K.; Parker, R.; Buchan, J.R. Defects in THO/TREX-2 function cause accumulation of novel cytoplasmic mRNP granules that can be cleared by autophagy. RNA 2016, 22, 1200–1214. [Google Scholar] [CrossRef]
- Zopf, C.J.; Quinn, K.; Zeidman, J.; Maheshri, N. Cell-cycle dependence of transcription dominates noise in gene expression. PLoS Comput. Biol. 2013, 9, e1003161. [Google Scholar] [CrossRef]
- To, T.L.; Maheshri, N. Noise can induce bimodality in positive transcriptional feedback loops without bistability. Science 2010, 327, 1142–1145. [Google Scholar] [CrossRef]
- Maheshri, N.; O’Shea, E.K. Living with noisy genes: How cells function reliably with inherent variability in gene expression. Annu. Rev. Biophys. Biomol. Struct. 2007, 36, 413–434. [Google Scholar] [CrossRef] [PubMed]
- Zenklusen, D.; Larson, D.R.; Singer, R.H. Single-RNA counting reveals alternative modes of gene expression in yeast. Nat. Struct. Mol. Biol. 2008, 15, 1263–1271. [Google Scholar] [CrossRef] [PubMed]
- Raju, K.K.; Natarajan, S.; Kumar, N.S.; Kumar, D.A.; Raghavendra, N.M. Role of cytoplasmic deadenylation and mRNA decay factors in yeast apoptosis. FEMS Yeast Res. 2015, 15. [Google Scholar] [CrossRef] [PubMed]
- Caponigro, G.; Parker, R. Multiple functions for the poly(A)-binding protein in mRNA decapping and deadenylation in yeast. Genes Dev. 1995, 9, 2421–2432. [Google Scholar] [CrossRef]
- Anderson, J.T.; Paddy, M.R.; Swanson, M.S. PUB1 is a major nuclear and cytoplasmic polyadenylated RNA-binding protein in Saccharomyces cerevisiae. Mol. Cell. Biol. 1993, 13, 6102–6113. [Google Scholar] [CrossRef][Green Version]
- Hermesh, O.; Genz, C.; Yofe, I.; Sinzel, M.; Rapaport, D.; Schuldiner, M.; Jansen, R.P. Yeast phospholipid biosynthesis is linked to mRNA localization. J. Cell Sci. 2014, 127, 3373–3381. [Google Scholar] [CrossRef]
- Loewen, C.J.; Young, B.P.; Tavassoli, S.; Levine, T.P. Inheritance of cortical ER in yeast is required for normal septin organization. J. Cell Biol. 2007, 179, 467–483. [Google Scholar] [CrossRef]
- Rzeczkowski, K.; Beuerlein, K.; Muller, H.; Dittrich-Breiholz, O.; Schneider, H.; Kettner-Buhrow, D.; Holtmann, H.; Kracht, M. c-Jun N-terminal kinase phosphorylates DCP1a to control formation of P bodies. J. Cell Biol. 2011, 194, 581–596. [Google Scholar] [CrossRef]
- Aizer, A.; Kafri, P.; Kalo, A.; Shav-Tal, Y. The P body protein Dcp1a is hyper-phosphorylated during mitosis. PLoS ONE 2013, 8, e49783. [Google Scholar] [CrossRef]
- Boczkowski, D.; Nair, S.K.; Nam, J.H.; Lyerly, H.K.; Gilboa, E. Induction of tumor immunity and cytotoxic T lymphocyte responses using dendritic cells transfected with messenger RNA amplified from tumor cells. Cancer Res. 2000, 60, 1028–1034. [Google Scholar]
- Bonneaud, N.; Ozier-Kalogeropoulos, O.; Li, G.Y.; Labouesse, M.; Minvielle-Sebastia, L.; Lacroute, F. A family of low and high copy replicative, integrative and single-stranded S. cerevisiae/E. coli shuttle vectors. Yeast 1991, 7, 609–615. [Google Scholar] [CrossRef] [PubMed]
- Gari, E.; Piedrafita, L.; Aldea, M.; Herrero, E. A set of vectors with a tetracycline-regulatable promoter system for modulated gene expression in Saccharomyces cerevisiae. Yeast 1997, 13, 837–848. [Google Scholar] [CrossRef]
- Gerami-Nejad, M.; Zacchi, L.F.; McClellan, M.; Matter, K.; Berman, J. Shuttle vectors for facile gap repair cloning and integration into a neutral locus in Candida albicans. Microbiology 2013, 159, 565–579. [Google Scholar] [CrossRef] [PubMed]
- Longtine, M.S.; McKenzie, A., 3rd; Demarini, D.J.; Shah, N.G.; Wach, A.; Brachat, A.; Philippsen, P.; Pringle, J.R. Additional modules for versatile and economical PCR-based gene deletion and modification in Saccharomyces cerevisiae. Yeast 1998, 14, 953–961. [Google Scholar] [CrossRef]
- Janke, C.; Magiera, M.M.; Rathfelder, N.; Taxis, C.; Reber, S.; Maekawa, H.; Moreno-Borchart, A.; Doenges, G.; Schwob, E.; Schiebel, E.; et al. A versatile toolbox for PCR-based tagging of yeast genes: New fluorescent proteins, more markers and promoter substitution cassettes. Yeast 2004, 21, 947–962. [Google Scholar] [CrossRef]
- Wach, A.; Brachat, A.; Pohlmann, R.; Philippsen, P. New heterologous modules for classical or PCR-based gene disruptions in Saccharomyces cerevisiae. Yeast 1994, 10, 1793–1808. [Google Scholar] [CrossRef]
- Sheff, M.A.; Thorn, K.S. Optimized cassettes for fluorescent protein tagging in Saccharomyces cerevisiae. Yeast 2004, 21, 661–670. [Google Scholar] [CrossRef]
- Strathern, J.N.; Higgins, D.R. Recovery of plasmids from yeast into Escherichia coli: Shuttle vectors. Methods Enzymol. 1991, 194, 319–329. [Google Scholar]
- Gietz, R.D.; Schiestl, R.H. Frozen competent yeast cells that can be transformed with high efficiency using the LiAc/SS carrier DNA/PEG method. Nat. Protoc. 2007, 2, 1–4. [Google Scholar] [CrossRef]
- Schmitt, M.E.; Brown, T.A.; Trumpower, B.L. A rapid and simple method for preparation of RNA from Saccharomyces cerevisiae. Nucleic Acids Res. 1990, 18, 3091–3092. [Google Scholar] [CrossRef]
- Rosen, K.M.; Lamperti, E.D.; Villa-Komaroff, L. Optimizing the northern blot procedure. BioTechniques 1990, 8, 398–403. [Google Scholar] [PubMed]
- Trcek, T.; Chao, J.A.; Larson, D.R.; Park, H.Y.; Zenklusen, D.; Shenoy, S.M.; Singer, R.H. Single-mRNA counting using fluorescent in situ hybridization in budding yeast. Nat. Protoc. 2012, 7, 408–419. [Google Scholar] [CrossRef] [PubMed]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kozlova, N.V.; Pichon, C.; Rahmouni, A.R. mRNA with Mammalian Codon Bias Accumulates in Yeast Mutants with Constitutive Stress Granules. Int. J. Mol. Sci. 2020, 21, 1234. https://doi.org/10.3390/ijms21041234
Kozlova NV, Pichon C, Rahmouni AR. mRNA with Mammalian Codon Bias Accumulates in Yeast Mutants with Constitutive Stress Granules. International Journal of Molecular Sciences. 2020; 21(4):1234. https://doi.org/10.3390/ijms21041234
Chicago/Turabian StyleKozlova, Natalia V., Chantal Pichon, and A. Rachid Rahmouni. 2020. "mRNA with Mammalian Codon Bias Accumulates in Yeast Mutants with Constitutive Stress Granules" International Journal of Molecular Sciences 21, no. 4: 1234. https://doi.org/10.3390/ijms21041234
APA StyleKozlova, N. V., Pichon, C., & Rahmouni, A. R. (2020). mRNA with Mammalian Codon Bias Accumulates in Yeast Mutants with Constitutive Stress Granules. International Journal of Molecular Sciences, 21(4), 1234. https://doi.org/10.3390/ijms21041234






