Molecular and Functional Roles of MicroRNAs in the Progression of Hepatocellular Carcinoma—A Review
Abstract
1. Introduction
1.1. Hepatocellular Carcinoma
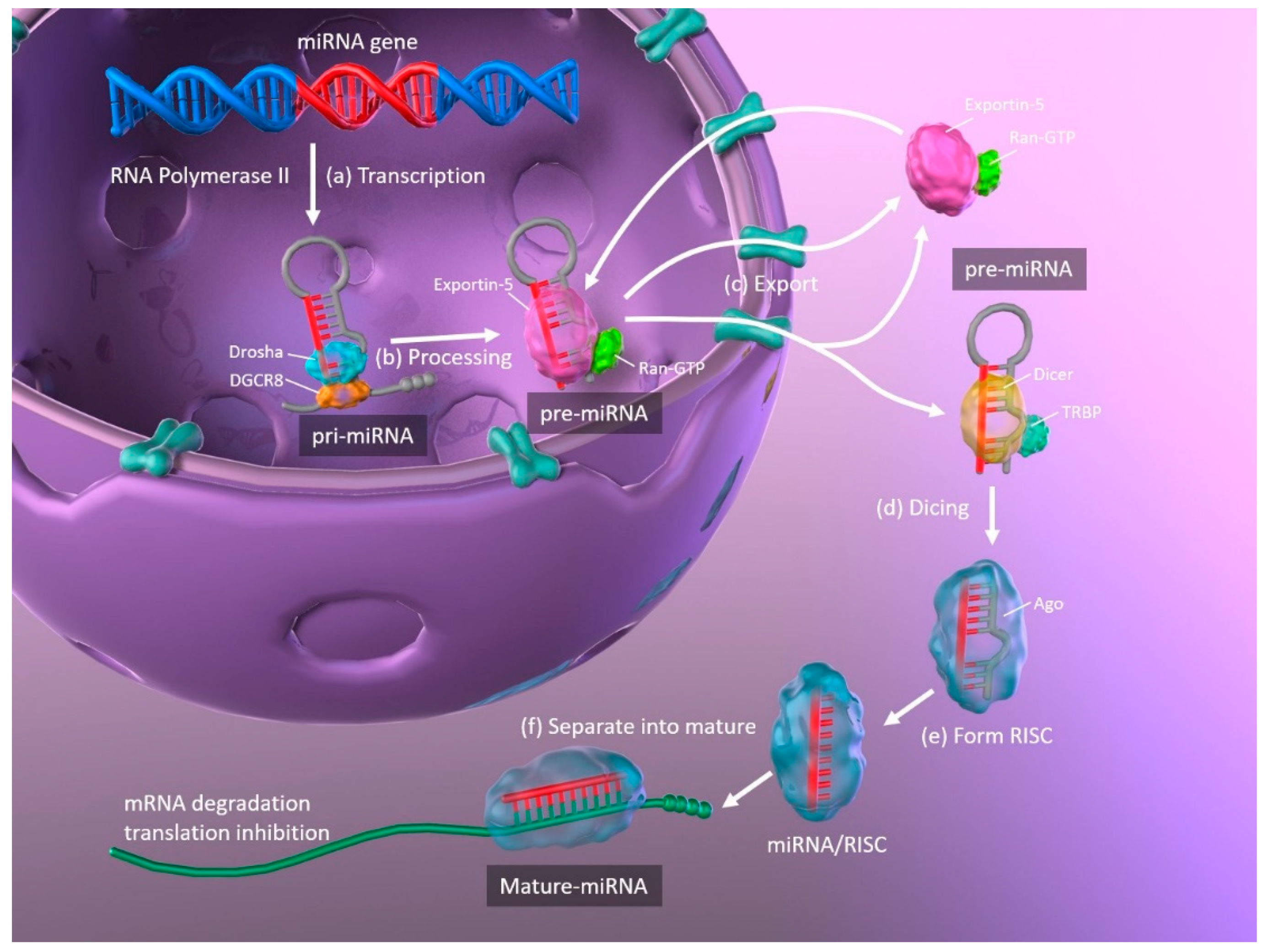
1.2. MicroRNAs
2. The Role of miRNAs in Liver Regeneration
2.1. Cell Proliferation in Hepatocytes
2.2. Apoptosis in Hepatocytes
3. The Role of miRNAs in Liver Diseases Associated with HCC
3.1. Hepatic Lipid Metabolism
3.2. Hepatic Inflammation
3.3. Hepatic Fibrosis
3.4. Hepatitis B Virus
3.5. Hepatitis C Virus
3.6. Alcoholic Liver Disease
3.7. Nonalcoholic Fatty Liver Disease
4. Dysregulation of miRNAs in HCC
4.1. Development of HCC
4.1.1. Oncogenic miRNAs
4.1.2. Tumor Suppressive miRNAs
4.2. miRNAs as Biomarkers for HCC
4.3. miRNAs as Therapeutic Targets for HCC
4.4. miRNAs in HCC Therapy Resistance
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| HCC | Hepatocellular carcinoma |
| miRNA | MicroRNA |
| siRNA | Small interfering RNA |
| priRNA | PIWI-Interacting RNA |
| mRNA | Messenger RNA |
| HBV | Viral hepatitis B |
| HCV | Viral hepatitis C |
| ALD | Alcoholic liver disease |
| NAFLD | Nonalcoholic fatty liver disease |
| UTR | Untranslated region |
| pri-miRNA | Primary microRNA |
| DGCR8 | DiGeorge syndrome critical region gene 8 |
| TRBP | Transactivation response element RNA-binding protein |
| RISC | RNA-Induced silencing complex |
| AGO | Argonaute protein |
| OncomiR | Oncogenic miRNA |
| IL-6 | Interleukin-6 |
| TNF-α | Tumor necrosis factor-α |
| HGF | Hepatocyte growth factor |
| PTEN | Phosphatase and tensin homolog deleted on chromosome |
| RhoB | Ras homolog family member B |
| PI3K | Phosphoinositide 3-kinase |
| mTOR | Mammalian target of rapamycin |
| SMAD3 | Small mother against decapentaplegic 3 |
| TRAIL | TNF related apoptosis-inducing ligand |
| PUMA | P53 upregulated modulator of apoptosis |
| STAT3 | Signal transducer and activator of transcription 3 |
| TIMP3 | Tissue inhibitor of metalloproteinase 3 |
| NASH | Nonalcoholic steatohepatitis |
| FASN | Fatty acid synthase |
| ACC1 | Acetyl-CoA carboxylase 1 |
| IGF | Insulin-like growth factor |
| HNF | Hepatocyte nuclear factor |
| AMPK | Adenosine 5′-monophosphate-activated protein kinase |
| SIRT1 | Sirtuin 1 |
| PPAR | Peroxisome proliferator activated receptor |
| LXR | Liver X receptor |
| SREBP | Sterol regulatory element-binding protein |
| HSC | Hepatic stellate cell |
| TGF | Transforming growth factor |
| ADAM17 | A disintegrin and metalloproteinase 17 |
| TLR2 | Toll-Like receptor 2 |
| WNT | Wingless-Related integration site |
| JAK | Janus kinase |
| MAPK | Mitogen-Activated protein kinase |
| SOCS3 | Suppressor of cytokine signaling 3 |
| HBe | Hepatitis Be |
| SHIP1 | SH2-Containg inositol phosphatase 1 |
| ULK1 | Unc-51 like autophagy activating kinase 1 |
| PCBP | Poly(rC)-Binding protein |
| DLC | Deleted in liver cancer |
| IRF1 | Interferon regulatory factor 1 |
| FN | Fibronectin |
| SCD | Stearoyl-CoA desaturase |
| CREB1 | cAMP responsive element binding protein 1 |
| TRIF | TIR-Domain-Containing adapter-inducing interferon |
| HMGA2 | High-Mobility group AT-hook 2 |
| C/EBP | CCAAT/enhancer binding protein |
| KLF5 | Kruppel-Like factor 5 |
| PDK1 | Pyruvate dehydrogenase kinase 1 |
| VEGF | Vascular endothelial growth factor |
| bFGF | Basic fibroblast growth factor |
| MMP2 | Matrix metallopeptidase 2 |
| HMGB | High-Mobility group box |
| DAMP | Damaged-Associated molecular pattern |
| TP53INP1 | Tumor protein p53 inducible nuclear protein 1 |
| VFL | von Hippel-Lindau tumor suppressor |
| MLH1 | MutL homolog 1 |
| CLDN | Claudin |
| DDIT | DNA damage-inducible transcript |
| IKK | I kappa B kinase |
| TAB3 | TGF-beta activated kinase 1 binding protein 3 |
| VAV2 | Vav guanine nucleotide exchange factor 2 |
| CDC42 | Cell division cycle 42 |
| PKB | Protein kinase B |
| ROCK | RHO-Associated protein kinase |
| EMT | Epithelial-Mesenchymal transition |
| PKM2 | Pyruvate kinase M2 |
| AFP | α-fetoprotein |
| PIVKA-II | Vitamin K absence or antagonists-II |
| AMO | AntimiRNA oligonucleotide |
| LNA | Locked-Nucleic-Acid antisense oligonucleotide |
| MTg-AMO | Antagomirs and multiple-target antimiRNA antisense oligodeoxyribonucleotide |
| AAV | Adeno-Associated virus |
| TAI | Transcatheter arterial infusion |
| ABC | ATP-Binding cassette |
| KEAP1 | Kelch-Like ECH-associated protein 1 |
| NFE2L2 | Nuclear factor erythroid 2 like 2 |
| DUSP6 | Dual specificity phosphatase 6 |
| EIF4E | Eukaryotic translation initiation factor 4E |
| BRAF | v-Raf murine sarcoma viral oncogene homolog B |
| PDGFR3 | Platelet-Derived growth factor receptor 3 |
| RASSF1 | Ras association domain family member 1 |
References
- Global Burden of Disease Cancer Collaboration; Fitzmaurice, C.; Allen, C.; Barber, R.M.; Barregard, L.; Bhutta, Z.A.; Brenner, H.; Dicker, D.J.; Chimed-Orchir, O.; Dandona, R.; et al. Global, regional, and national cancer incidence, mortality, years of life lost, years lived with disability, and disability-adjusted life-years for 32 cancer groups, 1990 to 2015: A systematic analysis for the global burden of disease study. JAMA Oncol. 2017, 3, 5245–5248. [Google Scholar]
- El-Serag, H.B.; Rudolph, K.L.L. Hepatocellular carcinoma: Epidemiology and molecular carcinogenesis. Gastroenterology 2007, 132, 2557–2576. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.D.; Hainaut, P.; Gores, G.J.; Amadou, A.; Plymoth, A.; Roberts, L.R.R. A global view of hepatocellular carcinoma: Trends, risk, prevention and management. Nat. Rev. Gastroenterol. Hepatol. 2019, 16, 589–604. [Google Scholar] [CrossRef] [PubMed]
- Park, J.W.; Chen, M.; Colombo, M.; Roberts, L.R.; Schwartz, M.; Chen, P.J.; Kudo, M.; Johnson, P.; Wagner, S.; Orsini, L.S.; et al. Global patterns of hepatocellular carcinoma management from diagnosis to death: The BRIDGE Study. Liver Int. 2015, 35, 2155–2166. [Google Scholar] [CrossRef]
- Younossi, Z.M.; Blissett, D.; Blissett, R.; Henry, L.; Stepanova, M.; Younossi, Y.; Racila, A.; Hunt, S.; Beckerman, R.R. The economic and clinical burden of nonalcoholic fatty liver disease in the United States and Europe. Hepatology 2016, 64, 1577–1586. [Google Scholar] [CrossRef]
- Altekruse, S.F.; Henley, S.J.; Cucinelli, J.E.; McGlynn, K.A.A. Changing hepatocellular carcinoma incidence and liver cancer mortality rates in the United States. Am. J. Gastroenterol. 2014, 109, 542–553. [Google Scholar] [CrossRef] [PubMed]
- Zhang, G.; Li, R.; Deng, Y.; Zhao, L.L. Conditional survival of patients with hepatocellular carcinoma: Results from the Surveillance, Epidemiology, and End Results registry. Expert Rev. Gastroenterol. Hepatol. 2018, 12, 515–523. [Google Scholar] [CrossRef]
- Xu, L.; Kim, Y.; Spolverato, G.; Gani, F.; Pawlik, T.M.M. Racial disparities in treatment and survival of patients with hepatocellular carcinoma in the United States. Hepatobiliary Surg. Nutr. 2016, 5, 43–52. [Google Scholar]
- Esquela-Kerscher, A.; Slack, F.J.J. Oncomirs—microRNAs with a role in cancer. Nat. Rev. Cancer 2006, 6, 2596–2599. [Google Scholar] [CrossRef]
- Bartel, D.P.P. MicroRNAs: Genomics, biogenesis, mechanism, and function. Cell 2004, 116, 281–297. [Google Scholar] [CrossRef]
- Kim, V.N.N. MicroRNA biogenesis: Coordinated cropping and dicing. Nat. Rev. Mol. Cell Biol. 2005, 6, 376–385. [Google Scholar] [CrossRef] [PubMed]
- Borchert, G.M.; Lanier, W.; Davidson, B.L.L. RNA polymerase III transcribes human microRNAs. Nat. Struct. Mol. Biol. 2006, 13, 1097–1101. [Google Scholar] [CrossRef]
- Garzon, R.; Fabbri, M.; Cimmino, A.; Calin, G.A.; Croce, C.M.M. MicroRNA expression and function in cancer. Trends Mol. Med. 2006, 12, 580–587. [Google Scholar] [CrossRef] [PubMed]
- Si, W.; Shen, J.; Zheng, H.; Fan, W.W. The role and mechanisms of action of microRNAs in cancer drug resistance. Clin. Epigenetics 2019, 11, 25. [Google Scholar] [CrossRef]
- Cheng, C.J.; Slack, F.J.J. The duality of oncomiR addiction in the maintenance and treatment of cancer. Cancer J. 2012, 18, 232–237. [Google Scholar] [CrossRef]
- Nedaeinia, R.; Manian, M.; Jazayeri, M.H.; Ranjbar, M.; Salehi, R.; Sharifi, M.; Mohaghegh, F.; Goli, M.; Jahednia, S.H.; Avan, A.; et al. Circulating exosomes and exosomal microRNAs as biomarkers in gastrointestinal cancer. Cancer Gene Ther. 2017, 24, 48–56. [Google Scholar] [CrossRef]
- Zheng, Q.; Chen, C.; Guan, H.; Kang, W.; Yu, C.C. Prognostic role of microRNAs in human gastrointestinal cancer: A systematic review and meta-analysis. Oncotarget 2017, 8, 46611–46623. [Google Scholar] [CrossRef]
- Santoni, G.; Morelli, M.B.; Amantini, C.; Battelli, N.N. Urinary markers in bladder cancer: An update. Front. Oncol. 2018, 8, 362. [Google Scholar] [CrossRef]
- Wang, J.; Ni, J.; Beretov, J.; Thompson, J.; Graham, P.; Li, Y.Y. Exosomal microRNAs as liquid biopsy biomarkers in prostate cancer. Crit. Rev. Oncol. Hematol. 2020, 145, 102860. [Google Scholar] [CrossRef]
- Miao, J.; Regenstein, J.M.; Xu, D.; Zhou, D.; Li, H.; Zhang, H.; Li, C.; Qiu, J.; Chen, X.X. The roles of microRNA in human cervical cancer. Arch. Biochem. Biophys. 2020, 690, 108480. [Google Scholar] [CrossRef]
- Wadowska, K.; Bil-Lula, I.; Trembecki, L.; Sliwinska-Mosson, M.M. Genetic markers in lung cancer diagnosis: A review. Int. J. Mol. Sci. 2020, 21, 4569. [Google Scholar] [CrossRef]
- Xu, J.; An, P.; Winkler, C.A.; Yu, Y.Y. Dysregulated microRNAs in hepatitis B virus-related hepatocellular carcinoma: Potential as biomarkers and therapeutic targets. Front. Oncol. 2020, 10, 1271. [Google Scholar] [CrossRef]
- Lim, L.J.; Wong, S.Y.S.; Huang, F.; Lim, S.; Chong, S.S.; Ooi, L.L.; Kon, O.L.; Lee, C.G.G. Roles and regulation of long noncoding RNAs in hepatocellular carcinoma. Cancer Res. 2019, 79, 5131–5139. [Google Scholar] [CrossRef]
- Fausto, N.; Campbell, J.S.; Riehle, K.J.J. Liver regeneration. Hepatology 2006, 43 (Suppl. 1), S45–S53. [Google Scholar] [CrossRef]
- Shu, J.; Kren, B.T.; Xia, Z.; Wong, P.Y.; Li, L.; Hanse, E.A.; Min, M.X.; Li, B.; Albrecht, J.H.; Zeng, Y.; et al. Genomewide microRNA down-regulation as a negative feedback mechanism in the early phases of liver regeneration. Hepatology 2011, 54, 609–619. [Google Scholar] [CrossRef]
- Chen, X.; Murad, M.; Cui, Y.Y.; Yao, L.J.; Venugopal, S.K.; Dawson, K.; Wu, J.J. miRNA regulation of liver growth after 50% partial hepatectomy and small size grafts in rats. Transplantation 2011, 91, 293–399. [Google Scholar] [CrossRef] [PubMed]
- Yan-nan, B.; Zhao-yan, Y.; Li-xi, L.; Jiang, Y.; Qing-jie, X.; Yong, Z.Z. MicroRNA-21 accelerates hepatocyte proliferation in vitro via PI3K/Akt signaling by targeting PTEN. Biochem. Biophys. Res. Commun. 2014, 443, 802–807. [Google Scholar] [CrossRef]
- Chen, X.; Song, M.; Chen, W.; Dimitrova-Shumkovska, J.; Zhao, Y.; Cao, Y.; Song, Y.; Yang, W.; Wang, F.; Xiang, Y.; et al. MicroRNA-21 contributes to liver regeneration by targeting PTEN. Med. Sci. Monit. 2016, 22, 83–91. [Google Scholar] [CrossRef]
- Lv, T.; Kong, L.; Jiang, L.; Wu, H.; Wen, T.; Shi, Y.; Yang, J.J. Dicer1 facilitates liver regeneration in a manner dependent on the inhibitory effect of miR-21 on Pten and Rhob expression. Life Sci. 2019, 232, 116656. [Google Scholar] [CrossRef]
- Yuan, B.; Dong, R.; Shi, D.; Zhou, Y.; Zhao, Y.; Miao, M.; Jiao, B.B. Down-regulation of miR-23b may contribute to activation of the TGF-beta1/Smad3 signalling pathway during the termination stage of liver regeneration. FEBS Lett. 2011, 585, 9273–9274. [Google Scholar] [CrossRef]
- Chen, H.; Sun, Y.; Dong, R.; Yang, S.; Pan, C.; Xiang, D.; Miao, M.; Jiao, B.B. Mir-34a is upregulated during liver regeneration in rats and is associated with the suppression of hepatocyte proliferation. PLoS ONE 2011, 6, e20238. [Google Scholar] [CrossRef] [PubMed]
- Yi, P.S.; Zhang, M.; Xu, M.Q.Q. Role of microRNA in liver regeneration. Hepatobiliary Pancreat. Dis. Int. 2016, 15, 141–146. [Google Scholar] [CrossRef]
- Shirjang, S.; Mansoori, B.; Asghari, S.; Duijf, P.H.G.; Mohammadi, A.; Gjerstorff, M.; Baradaran, B.B. MicroRNAs in cancer cell death pathways: Apoptosis and necroptosis. Free Radic. Biol. Med. 2019, 139, 1–15. [Google Scholar] [CrossRef]
- An, F.; Gong, B.; Wang, H.; Yu, D.; Zhao, G.; Lin, L.; Tang, W.; Yu, H.; Bao, S.; Xie, Q.Q. miR-15b and miR-16 regulate TNF mediated hepatocyte apoptosis via BCL2 in acute liver failure. Apoptosis 2012, 17, 702–716. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Huang, F.; Wang, J.; Peng, L.; Luo, H.H. MiR-15b mediates liver cancer cells proliferation through targeting BCL-2. Int. J. Clin. Exp. Pathol. 2015, 8, 15677–15683. [Google Scholar]
- Gong, J.; Zhang, J.P.; Li, B.; Zeng, C.; You, K.; Chen, M.X.; Yuan, Y.; Zhuang, S.M.M. MicroRNA-125b promotes apoptosis by regulating the expression of Mcl-1, Bcl-w and IL-6R. Oncogene 2013, 32, 3071–3079. [Google Scholar] [CrossRef]
- Santhekadur, P.K.; Das, S.K.; Gredler, R.; Chen, D.; Srivastava, J.; Robertson, C.; Baldwin, A.S.S., Jr.; Fisher, P.B.; Sarkar, D.D. Multifunction protein staphylococcal nuclease domain containing 1 (SND1) promotes tumor angiogenesis in human hepatocellular carcinoma through novel pathway that involves nuclear factor kappaB and miR-221. J. Biol. Chem. 2012, 287, 13952–13958. [Google Scholar] [CrossRef]
- Li, Y.; Di, C.; Li, W.; Cai, W.; Tan, X.; Xu, L.; Yang, L.; Lou, G.; Yan, Y.Y. Oncomirs miRNA-221/222 and tumor suppressors miRNA-199a/195 are crucial miRNAs in liver cancer: A systematic analysis. Dig. Dis. Sci. 2016, 61, 2315–2327. [Google Scholar] [CrossRef]
- Garofalo, M.; Di Leva, G.; Romano, G.; Nuovo, G.; Suh, S.S.; Ngankeu, A.; Taccioli, C.; Pichiorri, F.; Alder, H.; Secchiero, P.; et al. miR-221&222 regulate TRAIL resistance and enhance tumorigenicity through PTEN and TIMP3 downregulation. Cancer Cell 2009, 16, 498–509. [Google Scholar] [PubMed]
- Sharma, A.D.; Narain, N.; Handel, E.M.; Iken, M.; Singhal, N.; Cathomen, T.; Manns, M.P.; Scholer, H.R.; Ott, M.; Cantz, T.T. MicroRNA-221 regulates FAS-induced fulminant liver failure. Hepatology 2011, 53, 1651–1661. [Google Scholar] [CrossRef]
- Jopling, C.C. Liver-specific microRNA-122: Biogenesis and function. RNA Biol. 2012, 9, 137–742. [Google Scholar] [CrossRef]
- Cheung, O.; Puri, P.; Eicken, C.; Contos, M.J.; Mirshahi, F.; Maher, J.W.; Kellum, J.M.; Min, H.; Luketic, V.A.; Sanyal, A.J.J. Nonalcoholic steatohepatitis is associated with altered hepatic MicroRNA expression. Hepatology 2008, 48, 1810–1820. [Google Scholar] [CrossRef]
- Wei, S.; Zhang, M.; Yu, Y.; Xue, H.; Lan, X.; Liu, S.; Hatch, G.; Chen, L.L. HNF-4alpha regulated miR-122 contributes to development of gluconeogenesis and lipid metabolism disorders in Type 2 diabetic mice and in palmitate-treated HepG2 cells. Eur. J. Pharm. 2016, 791, 254–263. [Google Scholar] [CrossRef] [PubMed]
- Braza-Boils, A.; Mari-Alexandre, J.; Molina, P.; Arnau, M.A.; Barcelo-Molina, M.; Domingo, D.; Girbes, J.; Giner, J.; Martinez-Dolz, L.; Zorio, E.E. Deregulated hepatic microRNAs underlie the association between non-alcoholic fatty liver disease and coronary artery disease. Liver Int. 2016, 36, 1221–1229. [Google Scholar] [CrossRef]
- Latorre, J.; Moreno-Navarrete, J.M.; Mercader, J.M.; Sabater, M.; Rovira, O.; Girones, J.; Ricart, W.; Fernandez-Real, J.M.; Ortega, F.J.J. Decreased lipid metabolism but increased FA biosynthesis are coupled with changes in liver microRNAs in obese subjects with NAFLD. Int. J. Obes. (Lond.) 2017, 41, 620–630. [Google Scholar] [CrossRef]
- Becker, P.P.; Rau, M.; Schmitt, J.; Malsch, C.; Hammer, C.; Bantel, H.; Mullhaupt, B.; Geier, A.A. Performance of Serum microRNAs -122, -192 and -21 as Biomarkers in Patients with Non-Alcoholic Steatohepatitis. PLoS ONE 2015, 10, e0142661. [Google Scholar] [CrossRef]
- Pirola, C.J.; Fernandez Gianotti, T.; Castano, G.O.; Mallardi, P.; San Martino, J.; Mora Gonzalez Lopez Ledesma, M.; Flichman, D.; Mirshahi, F.; Sanyal, A.J.; Sookoian, S.S. Circulating microRNA signature in non-alcoholic fatty liver disease: From serum non-coding RNAs to liver histology and disease pathogenesis. Gut 2015, 64, 8001–8002. [Google Scholar] [CrossRef]
- Akuta, N.; Kawamura, Y.; Suzuki, F.; Saitoh, S.; Arase, Y.; Fujiyama, S.; Sezaki, H.; Hosaka, T.; Kobayashi, M.; Suzuki, Y.; et al. Analysis of association between circulating miR-122 and histopathological features of nonalcoholic fatty liver disease in patients free of hepatocellular carcinoma. BMC Gastroenterol. 2016, 16, 141. [Google Scholar] [CrossRef]
- Esau, C.; Davis, S.; Murray, S.F.; Yu, X.X.; Pandey, S.K.; Pear, M.; Watts, L.; Booten, S.L.; Graham, M.; McKay, R.; et al. miR-122 regulation of lipid metabolism revealed by in vivo antisense targeting. Cell Metab. 2006, 3, 87–98. [Google Scholar] [CrossRef] [PubMed]
- Salvoza, N.C.; Klinzing, D.C.; Gopez-Cervantes, J.; Baclig, M.O.O. Association of circulating serum miR-34a and miR-122 with dyslipidemia among patients with non-alcoholic fatty liver disease. PLoS ONE 2016, 11, e0153497. [Google Scholar] [CrossRef] [PubMed]
- Ding, J.; Li, M.; Wan, X.; Jin, X.; Chen, S.; Yu, C.; Li, Y.Y. Effect of miR-34a in regulating steatosis by targeting PPARalpha expression in nonalcoholic fatty liver disease. Sci. Rep. 2015, 5, 13729. [Google Scholar] [CrossRef]
- Wan, Y.; McDaniel, K.; Wu, N.; Ramos-Lorenzo, S.; Glaser, T.; Venter, J.; Francis, H.; Kennedy, L.; Sato, K.; Zhou, T.; et al. Regulation of cellular senescence by miR-34a in alcoholic liver injury. Am. J. Pathol. 2017, 187, 2788–2798. [Google Scholar] [CrossRef] [PubMed]
- Rottiers, V.; Naar, A.M.M. MicroRNAs in metabolism and metabolic disorders. Nat. Rev. Mol. Cell Biol. 2012, 13, 239–250. [Google Scholar] [CrossRef] [PubMed]
- Hsu, S.H.; Wang, B.; Kota, J.; Yu, J.; Costinean, S.; Kutay, H.; Yu, L.; Bai, S.; La Perle, K.; Chivukula, R.R.; et al. Essential metabolic, anti-inflammatory, and anti-tumorigenic functions of miR-122 in liver. J. Clin. Investig. 2012, 122, 2871–2883. [Google Scholar] [CrossRef] [PubMed]
- Wen, J.; Friedman, J.R.R. miR-122 regulates hepatic lipid metabolism and tumor suppression. J. Clin. Investig. 2012, 122, 2773–2776. [Google Scholar] [CrossRef]
- Strum, J.C.; Johnson, J.H.; Ward, J.; Xie, H.; Feild, J.; Hester, A.; Alford, A.; Waters, K.M.M. MicroRNA 132 regulates nutritional stress-induced chemokine production through repression of SirT1. Mol. Endocrinol. 2009, 23, 1876–1884. [Google Scholar] [CrossRef]
- Bala, S.; Szabo, G.G. MicroRNA signature in alcoholic liver disease. Int. J. Hepatol. 2012, 2012, 498232. [Google Scholar] [CrossRef]
- Momen-Heravi, F.; Bala, S.; Kodys, K.; Szabo, G.G. Exosomes derived from alcohol-treated hepatocytes horizontally transfer liver specific miRNA-122 and sensitize monocytes to LPS. Sci. Rep. 2015, 5, 9991. [Google Scholar] [CrossRef]
- Bala, S.; Petrasek, J.; Mundkur, S.; Catalano, D.; Levin, I.; Ward, J.; Alao, H.; Kodys, K.; Szabo, G.G. Circulating microRNAs in exosomes indicate hepatocyte injury and inflammation in alcoholic, drug-induced, and inflammatory liver diseases. Hepatology 2012, 56, 1946–1957. [Google Scholar] [CrossRef]
- Ma, Z.; Liu, X.; Dong, H.; Xia, D.; Wang, L.; Chen, Y.; Xiong, Y.Y. Sorafenib and praziquantel synergistically attenuate Schistosoma japonicum-induced liver fibrosis in mice. Parasitol. Res. 2018, 117, 2831–2839. [Google Scholar] [CrossRef]
- Ying, H.Z.; Chen, Q.; Zhang, W.Y.; Zhang, H.H.; Ma, Y.; Zhang, S.Z.; Fang, J.; Yu, C.H.H. PDGF signaling pathway in hepatic fibrosis pathogenesis and therapeutics (Review). Mol. Med. Rep. 2017, 16, 7879–7889. [Google Scholar] [CrossRef] [PubMed]
- Wei, J.; Feng, L.; Li, Z.; Xu, G.; Fan, X.X. MicroRNA-21 activates hepatic stellate cells via PTEN/Akt signaling. Biomed. Pharm. 2013, 67, 387–392. [Google Scholar] [CrossRef]
- Marquez, R.T.; Bandyopadhyay, S.; Wendlandt, E.B.; Keck, K.; Hoffer, B.A.; Icardi, M.S.; Christensen, R.N.; Schmidt, W.N.; McCaffrey, A.P.P. Correlation between microRNA expression levels and clinical parameters associated with chronic hepatitis C viral infection in humans. Lab. Invest. 2010, 90, 1727–1736. [Google Scholar] [CrossRef]
- Chen, B.L.; Peng, J.; Li, Q.F.; Yang, M.; Wang, Y.; Chen, W.W. Exogenous bone morphogenetic protein-7 reduces hepatic fibrosis in Schistosoma japonicum-infected mice via transforming growth factor-beta/Smad signaling. World J. Gastroenterol. 2013, 19, 1405–1415. [Google Scholar] [CrossRef]
- Luo, X.; Zhang, D.; Xie, J.; Su, Q.; He, X.; Bai, R.; Gao, G.; Pan, W.W. MicroRNA-96 Promotes Schistosomiasis hepatic fibrosis in mice by suppressing Smad7. Mol. Methods Clin. Dev. 2018, 11, 73–82. [Google Scholar] [CrossRef]
- Singh, A.K.; Rooge, S.B.; Varshney, A.; Vasudevan, M.; Bhardwaj, A.; Venugopal, S.K.; Trehanpati, N.; Kumar, M.; Geffers, R.; Kumar, V.; et al. Global microRNA expression profiling in the liver biopsies of hepatitis B virus-infected patients suggests specific microRNA signatures for viral persistence and hepatocellular injury. Hepatology 2018, 67, 1695–1709. [Google Scholar] [CrossRef] [PubMed]
- Markovic, J.; Sharma, A.D.; Balakrishnan, A.A. MicroRNA-221: A fine tuner and potential biomarker of chronic liver injury. Cells 2020, 9, 1767. [Google Scholar] [CrossRef]
- Shaker, O.G.; Senousy, M.A.A. Serum microRNAs as predictors for liver fibrosis staging in hepatitis C virus-associated chronic liver disease patients. J. Viral. Hepat. 2017, 24, 636–644. [Google Scholar] [CrossRef]
- Park, J.K.; Kogure, T.; Nuovo, G.J.; Jiang, J.; He, L.; Kim, J.H.; Phelps, M.A.; Papenfuss, T.L.; Croce, C.M.; Patel, T.; et al. miR-221 silencing blocks hepatocellular carcinoma and promotes survival. Cancer Res. 2011, 71, 7608–7616. [Google Scholar] [CrossRef]
- Jiang, X.; Jiang, L.; Shan, A.; Su, Y.; Cheng, Y.; Song, D.; Ji, H.; Ning, G.; Wang, W.; Cao, Y.Y. Targeting hepatic miR-221/222 for therapeutic intervention of nonalcoholic steatohepatitis in mice. EBioMedicine 2018, 37, 307–321. [Google Scholar] [CrossRef] [PubMed]
- Menghini, R.; Federici, M.M. MicroRNA 221/222 cluster kicks out Timp-3 to inflame the liver. EBioMedicine 2018, 37, 7–8. [Google Scholar] [CrossRef] [PubMed]
- Bataller, R.; Brenner, D.A.A. Liver fibrosis. J. Clin. Invest. 2005, 115, 209–218. [Google Scholar] [CrossRef]
- Huang, Y.H.; Yang, Y.L.; Huang, F.C.; Tiao, M.M.; Lin, Y.C.; Tsai, M.H.; Wang, F.S.S. MicroRNA-29a mitigation of endoplasmic reticulum and autophagy aberrance counteracts in obstructive jaundice-induced fibrosis in mice. Exp. Biol. Med. (Maywood) 2018, 243, 13–21. [Google Scholar] [CrossRef]
- Huang, Y.H.; Yang, Y.L.; Wang, F.S.S. The Role of miR-29a in the Regulation, Function, and Signaling of Liver Fibrosis. Int. J. Mol. Sci. 2018, 19, 1889. [Google Scholar] [CrossRef]
- Wang, S.; Qiu, L.; Yan, X.; Jin, W.; Wang, Y.; Chen, L.; Wu, E.; Ye, X.; Gao, G.F.; Wang, F.; et al. Loss of microRNA 122 expression in patients with hepatitis B enhances hepatitis B virus replication through cyclin G(1) -modulated P53 activity. Hepatology 2012, 55, 730–741. [Google Scholar] [CrossRef] [PubMed]
- Lin, Y.; Deng, W.; Pang, J.; Kemper, T.; Hu, J.; Yin, J.; Zhang, J.; Lu, M.M. The microRNA-99 family modulates hepatitis B virus replication by promoting IGF-1R/PI3K/Akt/mTOR/ULK1 signaling-induced autophagy. Cell. Microbiol. 2017, 19, e12709. [Google Scholar] [CrossRef]
- Zhang, G.L.; Li, Y.X.; Zheng, S.Q.; Liu, M.; Li, X.; Tang, H.H. Suppression of hepatitis B virus replication by microRNA-199a-3p and microRNA-210. Antivir. Res. 2010, 88, 169–175. [Google Scholar] [CrossRef]
- Hayes, C.N.; Chayama, K.K. MicroRNAs as Biomarkers for Liver Disease and Hepatocellular Carcinoma. Int. J. Mol. Sci. 2016, 17, 280. [Google Scholar] [CrossRef]
- Banaudha, K.; Kaliszewski, M.; Korolnek, T.; Florea, L.; Yeung, M.L.; Jeang, K.T.; Kumar, A.A. MicroRNA silencing of tumor suppressor DLC-1 promotes efficient hepatitis C virus replication in primary human hepatocytes. Hepatology 2011, 53, 53–61. [Google Scholar] [CrossRef]
- Matsuura, K.; De Giorgi, V.; Schechterly, C.; Wang, R.Y.; Farci, P.; Tanaka, Y.; Alter, H.J.J. Circulating let-7 levels in plasma and extracellular vesicles correlate with hepatic fibrosis progression in chronic hepatitis C. Hepatology 2016, 64, 732–745. [Google Scholar] [CrossRef]
- Abdel-Al, A.; El-Ahwany, E.; Zoheiry, M.; Hassan, M.; Ouf, A.; Abu-Taleb, H.; Abdel Rahim, A.; El-Talkawy, M.D.; Zada, S.S. miRNA-221 and miRNA-222 are promising biomarkers for progression of liver fibrosis in HCV Egyptian patients. Virus Res. 2018, 253, 135–139. [Google Scholar] [CrossRef]
- McDaniel, K.; Huang, L.; Sato, K.; Wu, N.; Annable, T.; Zhou, T.; Ramos-Lorenzo, S.; Wan, Y.; Huang, Q.; Francis, H.; et al. The let-7/Lin28 axis regulates activation of hepatic stellate cells in alcoholic liver injury. J. Biol. Chem. 2017, 292, 11336–11347. [Google Scholar] [CrossRef] [PubMed]
- Brandon-Warner, E.; Feilen, N.A.; Culberson, C.R.; Field, C.O.; deLemos, A.S.; Russo, M.W.; Schrum, L.W.W. Processing of miR179–2 Cluster in Hepatic Stellate Cells Promotes Hepatic Fibrogenesis During Alcohol-Induced Injury. Alcohol. Clin. Exp. Res. 2016, 40, 1430–1442. [Google Scholar] [CrossRef] [PubMed]
- Torres, J.L.; Novo-Veleiro, I.; Manzanedo, L.; Alvela-Suarez, L.; Macias, R.; Laso, F.J.; Marcos, M.M. Role of microRNAs in alcohol-induced liver disorders and non-alcoholic fatty liver disease. World J. Gastroenterol. 2018, 24, 4104–4118. [Google Scholar] [CrossRef]
- Csak, T.; Bala, S.; Lippai, D.; Kodys, K.; Catalano, D.; Iracheta-Vellve, A.; Szabo, G.G. MicroRNA-155 deficiency attenuates liver steatosis and fibrosis without reducing inflammation in a mouse model of steatohepatitis. PLoS ONE 2015, 10, e0129251. [Google Scholar] [CrossRef]
- Wang, B.; Majumder, S.; Nuovo, G.; Kutay, H.; Volinia, S.; Patel, T.; Schmittgen, T.D.; Croce, C.; Ghoshal, K.; Jacob, S.T.T. Role of microRNA-155 at early stages of hepatocarcinogenesis induced by choline-deficient and amino acid-defined diet in C57BL/6 mice. Hepatology 2009, 50, 1152–1161. [Google Scholar] [CrossRef]
- Mak, D.; Babb de Villiers, C.; Chasela, C.; Urban, M.I.; Kramvis, A.A. Analysis of risk factors associated with hepatocellular carcinoma in black South Africans: 20002–012. PLoS ONE 2018, 13, e0196057. [Google Scholar] [CrossRef]
- Garzon, R.; Marcucci, G.; Croce, C.M.M. Targeting microRNAs in cancer: Rationale, strategies and challenges. Nat. Rev. Drug Discov. 2010, 9, 775–789. [Google Scholar] [CrossRef]
- Chen, Y.; Shen, A.; Rider, P.J.; Yu, Y.; Wu, K.; Mu, Y.; Hao, Q.; Liu, Y.; Gong, H.; Zhu, Y.; et al. A liver-specific microRNA binds to a highly conserved RNA sequence of hepatitis B virus and negatively regulates viral gene expression and replication. FASEB J. 2011, 25, 4511–4521. [Google Scholar] [CrossRef]
- Xu, J.; Zhu, X.; Wu, L.; Yang, R.; Yang, Z.; Wang, Q.; Wu, F.F. MicroRNA-122 suppresses cell proliferation and induces cell apoptosis in hepatocellular carcinoma by directly targeting Wnt/beta-catenin pathway. Liver Int. 2012, 32, 752–760. [Google Scholar] [CrossRef]
- Gao, D.; Zhai, A.; Qian, J.; Li, A.; Li, Y.; Song, W.; Zhao, H.; Yu, X.; Wu, J.; Zhang, Q.; et al. Down-regulation of suppressor of cytokine signaling 3 by miR-122 enhances interferon-mediated suppression of hepatitis B virus. Antivir. Res. 2015, 118, 2–8. [Google Scholar] [CrossRef]
- Xiao, C.; Rajewsky, K.K. MicroRNA control in the immune system: Basic principles. Cell 2009, 136, 26–36. [Google Scholar] [CrossRef]
- Wang, W.; Bian, H.; Li, F.; Li, X.; Zhang, D.; Sun, S.; Song, S.; Zhu, Q.; Ren, W.; Qin, C.; et al. HBeAg induces the expression of macrophage miR-155 to accelerate liver injury via promoting production of inflammatory cytokines. Cell. Mol. Life Sci. 2018, 75, 2627–2641. [Google Scholar] [CrossRef]
- Niepmann, M.; Shalamova, L.A.; Gerresheim, G.K.; Rossbach, O.O. Signals involved in regulation of hepatitis C virus RNA genome translation and replication. Front. Microbiol. 2018, 9, 395. [Google Scholar] [CrossRef]
- Niepmann, M.; Gerresheim, G.K.K. Hepatitis C virus translation regulation. Int. J. Mol. Sci. 2020, 21, 2328. [Google Scholar] [CrossRef]
- Ura, S.; Honda, M.; Yamashita, T.; Ueda, T.; Takatori, H.; Nishino, R.; Sunakozaka, H.; Sakai, Y.; Horimoto, K.; Kaneko, S.S. Differential microRNA expression between hepatitis B and hepatitis C leading disease progression to hepatocellular carcinoma. Hepatology 2009, 49, 1098–1112. [Google Scholar] [CrossRef]
- Peng, X.; Li, Y.; Walters, K.A.; Rosenzweig, E.R.; Lederer, S.L.; Aicher, L.D.; Proll, S.; Katze, M.G.G. Computational identification of hepatitis C virus associated microRNA-mRNA regulatory modules in human livers. BMC Genom. 2009, 10, 373. [Google Scholar] [CrossRef] [PubMed]
- Coll, M.; El Taghdouini, A.; Perea, L.; Mannaerts, I.; Vila-Casadesus, M.; Blaya, D.; Rodrigo-Torres, D.; Affo, S.; Morales-Ibanez, O.; Graupera, I.; et al. Integrative miRNA and gene expression profiling analysis of human quiescent hepatic stellate cells. Sci. Rep. 2015, 5, 11549. [Google Scholar] [CrossRef]
- Mandrekar, P.; Szabo, G.G. Signalling pathways in alcohol-induced liver inflammation. J. Hepatol. 2009, 50, 1258–1266. [Google Scholar] [CrossRef] [PubMed]
- Blaya, D.; Coll, M.; Rodrigo-Torres, D.; Vila-Casadesus, M.; Altamirano, J.; Llopis, M.; Graupera, I.; Perea, L.; Aguilar-Bravo, B.; Diaz, A.; et al. Integrative microRNA profiling in alcoholic hepatitis reveals a role for microRNA-182 in liver injury and inflammation. Gut 2016, 65, 1535–1545. [Google Scholar] [CrossRef]
- Leti, F.; Malenica, I.; Doshi, M.; Courtright, A.; Van Keuren-Jensen, K.; Legendre, C.; Still, C.D.; Gerhard, G.S.; DiStefano, J.K.K. High-throughput sequencing reveals altered expression of hepatic microRNAs in nonalcoholic fatty liver disease-related fibrosis. Transl. Res. 2015, 166, 304–314. [Google Scholar] [CrossRef]
- Guo, Y.; Xiong, Y.; Sheng, Q.; Zhao, S.; Wattacheril, J.; Flynn, C.R.R. A micro-RNA expression signature for human NAFLD progression. J. Gastroenterol. 2016, 51, 1022–1030. [Google Scholar] [CrossRef]
- Chen, Y.; Siegel, F.; Kipschull, S.; Haas, B.; Frohlich, H.; Meister, G.; Pfeifer, A.A. miR-155 regulates differentiation of brown and beige adipocytes via a bistable circuit. Nat. Commun. 2013, 4, 1769. [Google Scholar] [CrossRef]
- Khan, S.; Ayub, H.; Khan, T.; Wahid, F.F. MicroRNA biogenesis, gene silencing mechanisms and role in breast, ovarian and prostate cancer. Biochimie 2019, 167, 12–24. [Google Scholar] [CrossRef]
- Borel, F.; Konstantinova, P.; Jansen, P.L.L. Diagnostic and therapeutic potential of miRNA signatures in patients with hepatocellular carcinoma. J. Hepatol. 2012, 56, 1371–1383. [Google Scholar] [CrossRef]
- Morishita, A.; Iwama, H.; Fujihara, S.; Sakamoto, T.; Fujita, K.; Tani, J.; Miyoshi, H.; Yoneyama, H.; Himoto, T.; Masaki, T.T. MicroRNA profiles in various hepatocellular carcinoma cell lines. Oncol. Lett. 2016, 12, 1687–1692. [Google Scholar] [CrossRef]
- Morishita, A.; Masaki, T.T. MicroRNAs as possible biomarkers for hepatocellular carcinoma. Hepatol. Res. 2018, 48, 499–501. [Google Scholar] [CrossRef] [PubMed]
- Oura, K.; Fujita, K.; Morishita, A.; Iwama, H.; Nakahara, M.; Tadokoro, T.; Sakamoto, T.; Nomura, T.; Yoneyama, H.; Mimura, S.; et al. Serum microRNA-125a-5p as a potential biomarker of HCV-associated hepatocellular carcinoma. Oncol. Lett. 2019, 18, 8828–8890. [Google Scholar] [CrossRef]
- Zhu, Q.; Gong, L.; Wang, J.; Tu, Q.; Yao, L.; Zhang, J.R.; Han, X.J.; Zhu, S.J.; Wang, S.M.; Li, Y.H.; et al. miR-10b exerts oncogenic activity in human hepatocellular carcinoma cells by targeting expression of CUB and sushi multiple domains 1 (CSMD1). BMC Cancer 2016, 16, 806. [Google Scholar] [CrossRef]
- Wang, J.; Chu, Y.; Xu, M.; Zhang, X.; Zhou, Y.; Xu, M.M. miR-21 promotes cell migration and invasion of hepatocellular carcinoma by targeting KLF5. Oncol. Lett. 2019, 17, 2221–2227. [Google Scholar] [CrossRef]
- Koenig, A.B.; Barajas, J.M.; Guerrero, M.J.; Ghoshal, K.K. A Comprehensive Analysis of Argonaute-CLIP Data Identifies Novel, Conserved and Species-Specific Targets of miR-21 in Human Liver and Hepatocellular Carcinoma. Int. J. Mol. Sci. 2018, 19, 851. [Google Scholar] [CrossRef] [PubMed]
- Feng, X.; Jiang, J.; Shi, S.; Xie, H.; Zhou, L.; Zheng, S.S. Knockdown of miR-25 increases the sensitivity of liver cancer stem cells to TRAIL-induced apoptosis via PTEN/PI3K/Akt/Bad signaling pathway. Int. J. Oncol. 2016, 49, 2600–2610. [Google Scholar] [CrossRef]
- Yang, W.; Dou, C.; Wang, Y.; Jia, Y.; Li, C.; Zheng, X.; Tu, K.K. MicroRNA-92a contributes to tumor growth of human hepatocellular carcinoma by targeting FBXW7. Oncol. Rep. 2015, 34, 2576–2584. [Google Scholar] [CrossRef][Green Version]
- Iwai, N.; Yasui, K.; Tomie, A.; Gen, Y.; Terasaki, K.; Kitaichi, T.; Soda, T.; Yamada, N.; Dohi, O.; Seko, Y.; et al. Oncogenic miR-965–p inhibits apoptosis by targeting the caspase-9 gene in hepatocellular carcinoma. Int. J. Oncol. 2018, 53, 237–245. [Google Scholar]
- Wang, Y.; Chen, F.; Zhao, M.; Yang, Z.; Zhang, S.; Ye, L.; Gao, H.; Zhang, X.X. MiR-107 suppresses proliferation of hepatoma cells through targeting HMGA2 mRNA 3’UTR. Biochem. Biophys. Res. Commun. 2016, 480, 455–460. [Google Scholar] [CrossRef]
- Su, S.G.; Yang, M.; Zhang, M.F.; Peng, Q.Z.; Li, M.Y.; Liu, L.P.; Bao, S.Y.Y. miR-107-mediated decrease of HMGCS2 indicates poor outcomes and promotes cell migration in hepatocellular carcinoma. Int. J. Biochem. Cell Biol. 2017, 91, 53–59. [Google Scholar] [CrossRef]
- Zeng, Y.B.; Liang, X.H.; Zhang, G.X.; Jiang, N.; Zhang, T.; Huang, J.Y.; Zhang, L.; Zeng, X.C.C. miRNA-135a promotes hepatocellular carcinoma cell migration and invasion by targeting forkhead box O1. Cancer Cell Int. 2016, 16, 63. [Google Scholar] [CrossRef]
- Fu, X.; Wen, H.; Jing, L.; Yang, Y.; Wang, W.; Liang, X.; Nan, K.; Yao, Y.; Tian, T.T. MicroRNA-1555–p promotes hepatocellular carcinoma progression by suppressing PTEN through the PI3K/Akt pathway. Cancer Sci. 2017, 108, 620–631. [Google Scholar] [CrossRef]
- Yang, J.; He, Y.; Zhai, N.; Ding, S.; Li, J.; Peng, Z.Z. MicroRNA-181a inhibits autophagy by targeting Atg5 in hepatocellular carcinoma. Front. Biosci. (Landmark Ed.) 2018, 23, 388–396. [Google Scholar]
- Li, N.; Men, W.; Zheng, Y.; Wang, H.; Meng, X.X. Oroxin B induces apoptosis by down-regulating MicroRNA-221 resulting in the inactivation of the PTEN/PI3K/AKT pathway in liver cancer. Molecules 2019, 24, 4384. [Google Scholar] [CrossRef]
- Huo, W.; Du, M.; Pan, X.; Zhu, X.; Gao, Y.; Li, Z.Z. miR-203a-3p.1 targets IL-24 to modulate hepatocellular carcinoma cell growth and metastasis. FEBS Open Bio. 2017, 7, 1085–1091. [Google Scholar] [CrossRef]
- Yang, Y.; Zhang, J.; Xia, T.; Li, G.; Tian, T.; Wang, M.; Wang, R.; Zhao, L.; Yang, Y.; Lan, K.; et al. MicroRNA-210 promotes cancer angiogenesis by targeting fibroblast growth factor receptor-like 1 in hepatocellular carcinoma. Oncol. Rep. 2016, 36, 2553–2562. [Google Scholar] [CrossRef]
- Li, H.; Wang, H.; Ren, Z.Z. MicroRNA-2145–p Inhibits the invasion and migration of hepatocellular carcinoma cells by targeting wiskott-aldrich syndrome like. Cell. Physiol. Biochem. 2018, 46, 757–764. [Google Scholar] [CrossRef]
- Chen, Y.L.; Xu, Q.P.; Guo, F.; Guan, W.H.H. MicroRNA-302d downregulates TGFBR2 expression and promotes hepatocellular carcinoma growth and invasion. Exp. Ther. Med. 2017, 13, 681–687. [Google Scholar] [CrossRef][Green Version]
- Yu, Q.; Yang, X.; Duan, W.; Li, C.; Luo, Y.; Lu, S.S. miRNA-346 promotes proliferation, migration and invasion in liver cancer. Oncol. Lett. 2017, 14, 3255–3260. [Google Scholar] [CrossRef]
- Chang, R.M.; Xiao, S.; Lei, X.; Yang, H.; Fang, F.; Yang, L.Y.Y. miRNA-487a promotes proliferation and metastasis in hepatocellular carcinoma. Clin. Cancer Res. 2017, 23, 2593–2604. [Google Scholar] [CrossRef]
- Xie, B.H.; He, X.; Hua, R.X.; Zhang, B.; Tan, G.S.; Xiong, S.Q.; Liu, L.S.; Chen, W.; Yang, J.Y.; Wang, X.N.; et al. Mir-765 promotes cell proliferation by downregulating INPP4B expression in human hepatocellular carcinoma. Cancer Biomark. 2016, 16, 405–413. [Google Scholar] [CrossRef]
- Han, G.; Zhang, L.; Ni, X.; Chen, Z.; Pan, X.; Zhu, Q.; Li, S.; Wu, J.; Huang, X.; Wang, X.X. MicroRNA-873 promotes cell proliferation, migration, and invasion by directly targeting TSLC1 in hepatocellular carcinoma. Cell. Physiol. Biochem. 2018, 46, 2261–2270. [Google Scholar] [CrossRef]
- Jia, B.; Tan, L.; Jin, Z.; Jiao, Y.; Fu, Y.; Liu, Y.Y. MiR-892a promotes hepatocellular carcinoma cells proliferation and invasion through targeting CD226. J. Cell. Biochem. 2017, 118, 1489–1496. [Google Scholar] [CrossRef]
- Ye, Y.; Wei, Y.; Xu, Y.; Li, Y.; Wang, R.; Chen, J.; Zhou, Y.; Fu, Z.; Chen, Y.; Wang, X.; et al. Induced MiR-1249 expression by aberrant activation of Hedegehog signaling pathway in hepatocellular carcinoma. Exp. Cell Res. 2017, 355, 9–17. [Google Scholar] [CrossRef]
- Liu, Z.; Wang, Y.; Dou, C.; Sun, L.; Li, Q.; Wang, L.; Xu, Q.; Yang, W.; Liu, Q.; Tu, K.K. MicroRNA-1468 promotes tumor progression by activating PPAR-gamma-mediated AKT signaling in human hepatocellular carcinoma. J. Exp. Clin. Cancer Res. 2018, 37, 49. [Google Scholar] [CrossRef]
- Cheng, L.; Wang, H.; Han, S.S. MiR-3910 promotes the growth and migration of cancer cells in the progression of hepatocellular carcinoma. Dig. Dis. Sci. 2017, 62, 2812–2820. [Google Scholar] [CrossRef] [PubMed]
- Song, L.; Zhang, W.; Chang, Z.; Pan, Y.; Zong, H.; Fan, Q.; Wang, L.L. miR-4417 targets tripartite motif-containing 35 (TRIM35) and regulates pyruvate kinase muscle 2 (PKM2) phosphorylation to promote proliferation and suppress apoptosis in hepatocellular carcinoma cells. Med. Sci. Monit. 2017, 23, 1741–1750. [Google Scholar] [CrossRef] [PubMed]
- Jin, F.; Wang, Y.; Li, M.; Zhu, Y.; Liang, H.; Wang, C.; Wang, F.; Zhang, C.Y.; Zen, K.; Li, L.L. MiR-26 enhances chemosensitivity and promotes apoptosis of hepatocellular carcinoma cells through inhibiting autophagy. Cell Death Dis. 2017, 8, e2540. [Google Scholar] [CrossRef] [PubMed]
- Mahati, S.; Xiao, L.; Yang, Y.; Mao, R.; Bao, Y.Y. miR-29a suppresses growth and migration of hepatocellular carcinoma by regulating CLDN1. Biochem. Biophys. Res. Commun. 2017, 486, 732–737. [Google Scholar] [CrossRef]
- Zhao, G.; Han, C.; Zhang, Z.; Wang, L.; Xu, J.J. Increased expression of microRNA-315–p inhibits cell proliferation, migration, and invasion via regulating Sp1 transcription factor in HepG2 hepatocellular carcinoma cell line. Biochem. Biophys. Res. Commun. 2017, 490, 371–377. [Google Scholar] [CrossRef]
- Tian, Q.; Xiao, Y.; Wu, Y.; Liu, Y.; Song, Z.; Gao, W.; Zhang, J.; Yang, J.; Zhang, Y.; Guo, T.; et al. MicroRNA-33b suppresses the proliferation and metastasis of hepatocellular carcinoma cells through the inhibition of Sal-like protein 4 expression. Int. J. Mol. Med. 2016, 38, 1587–1595. [Google Scholar] [CrossRef]
- Zhang, J.J.; Chen, J.T.; Hua, L.; Yao, K.H.; Wang, C.Y.Y. miR-98 inhibits hepatocellular carcinoma cell proliferation via targeting EZH2 and suppressing Wnt/beta-catenin signaling pathway. Biomed. Pharm. 2017, 85, 472–478. [Google Scholar] [CrossRef]
- Jin, Y.; Wang, J.; Han, J.; Luo, D.; Sun, Z.Z. MiR-122 inhibits epithelial-mesenchymal transition in hepatocellular carcinoma by targeting Snail1 and Snail2 and suppressing WNT/beta-cadherin signaling pathway. Exp. Cell Res. 2017, 360, 210–217. [Google Scholar] [CrossRef]
- Li, G.; Zhang, W.; Gong, L.; Huang, X.X. MicroRNA 125a-5p inhibits cell proliferation and induces apoptosis in hepatitis b virus-related hepatocellular carcinoma by downregulation of ErbB3. Oncol. Res. 2019, 27, 449–458. [Google Scholar] [CrossRef]
- Kim, J.K.; Noh, J.H.; Jung, K.H.; Eun, J.W.; Bae, H.J.; Kim, M.G.; Chang, Y.G.; Shen, Q.; Park, W.S.; Lee, J.Y.; et al. Sirtuin7 oncogenic potential in human hepatocellular carcinoma and its regulation by the tumor suppressors MiR-125a-5p and MiR-125b. Hepatology 2013, 57, 1055–1067. [Google Scholar] [CrossRef]
- Jing, B.Q.; Ou, Y.; Zhao, L.; Xie, Q.; Zhang, Y.X.X. Experimental study on the prevention of liver cancer angiogenesis via miR-126. Eur. Rev. Med. Pharm. Sci. 2017, 21, 5096–5100. [Google Scholar]
- Cui, S.; Sun, Y.; Liu, Y.; Liu, C.; Wang, J.; Hao, G.; Sun, Q.Q. MicroRNA137 has a suppressive role in liver cancer via targeting EZH2. Mol. Med. Rep. 2017, 16, 9494–9502. [Google Scholar] [CrossRef]
- Yu, Q.; Xiang, L.; Yin, L.; Liu, X.; Yang, D.; Zhou, J.J. Loss-of-function of miR-142 by hypermethylation promotes TGF-beta-mediated tumour growth and metastasis in hepatocellular carcinoma. Cell Prolif. 2017, 50, e12384. [Google Scholar] [CrossRef]
- Su, F.; Zhao, J.; Qin, S.; Wang, R.; Li, Y.; Wang, Q.; Tan, Y.; Jin, H.; Zhu, F.; Ou, Y.; et al. Over-expression of Thrombospondin 4 correlates with loss of miR-142 and contributes to migration and vascular invasion of advanced hepatocellular carcinoma. Oncotarget 2017, 8, 23277–23288. [Google Scholar] [CrossRef] [PubMed]
- Hua, S.; Liu, C.; Liu, L.; Wu, D.D. miR-1423–p inhibits aerobic glycolysis and cell proliferation in hepatocellular carcinoma via targeting LDHA. Biochem. Biophys. Res. Commun. 2018, 496, 947–954. [Google Scholar] [CrossRef]
- Bao, H.; Li, X.; Li, H.; Xing, H.; Xu, B.; Zhang, X.; Liu, Z.Z. MicroRNA-144 inhibits hepatocellular carcinoma cell proliferation, invasion and migration by targeting ZFX. J. Biosci. 2017, 42, 103–111. [Google Scholar] [CrossRef]
- Dou, C.; Liu, Z.; Xu, M.; Jia, Y.; Wang, Y.; Li, Q.; Yang, W.; Zheng, X.; Tu, K.; Liu, Q.Q. miR-1873–p inhibits the metastasis and epithelial-mesenchymal transition of hepatocellular carcinoma by targeting S100A4. Cancer Lett. 2016, 381, 380–390. [Google Scholar] [CrossRef]
- Zhao, Y.; Li, F.; Zhang, X.; Liu, A.; Qi, J.; Cui, H.; Zhao, P.P. MicroRNA-194 acts as a prognostic marker and inhibits proliferation in hepatocellular carcinoma by targeting MAP4K4. Int. J. Clin. Exp. Pathol. 2015, 8, 12446–12454. [Google Scholar] [PubMed]
- Ding, J.; Huang, S.; Wang, Y.; Tian, Q.; Zha, R.; Shi, H.; Wang, Q.; Ge, C.; Chen, T.; Zhao, Y.; et al. Genome-wide screening reveals that miR-195 targets the TNF-alpha/NF-kappaB pathway by down-regulating IkappaB kinase alpha and TAB3 in hepatocellular carcinoma. Hepatology 2013, 58, 654–666. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.; Zhao, N.; Li, S.; Fang, J.H.; Chen, M.X.; Yang, J.; Jia, W.H.; Yuan, Y.; Zhuang, S.M.M. MicroRNA-195 suppresses angiogenesis and metastasis of hepatocellular carcinoma by inhibiting the expression of VEGF, VAV2, and CDC42. Hepatology 2013, 58, 642–653. [Google Scholar] [CrossRef]
- Ghosh, A.; Dasgupta, D.; Ghosh, A.; Roychoudhury, S.; Kumar, D.; Gorain, M.; Butti, R.; Datta, S.; Agarwal, S.; Gupta, S.; et al. MiRNA199a-3p suppresses tumor growth, migration, invasion and angiogenesis in hepatocellular carcinoma by targeting VEGFA, VEGFR1, VEGFR2, HGF and MMP2. Cell Death Dis. 2017, 8, e2706. [Google Scholar] [CrossRef]
- Ren, K.; Li, T.; Zhang, W.; Ren, J.; Li, Z.; Wu, G.G. miR-199a-3p inhibits cell proliferation and induces apoptosis by targeting YAP1, suppressing Jagged1-Notch signaling in human hepatocellular carcinoma. J. Biomed. Sci. 2016, 23, 79. [Google Scholar] [CrossRef]
- Zhou, S.J.; Liu, F.Y.; Zhang, A.H.; Liang, H.F.; Wang, Y.; Ma, R.; Jiang, Y.H.; Sun, N.F.F. MicroRNA-199b-5p attenuates TGF-beta1-induced epithelial-mesenchymal transition in hepatocellular carcinoma. Br. J. Cancer 2017, 117, 233–244. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Song, W.; Shen, W.; Yang, X.; Sun, W.; Qu, S.; Shang, R.; Ma, B.; Pu, M.; Tao, K.; et al. MicroRNA-200a suppresses cell invasion and migration by directly targeting GAB1 in hepatocellular carcinoma. Oncol. Res. 2017, 25, 1–10. [Google Scholar] [CrossRef]
- Zheng, X.B.; Chen, X.B.; Xu, L.L.; Zhang, M.; Feng, L.; Yi, P.S.; Tang, J.W.; Xu, M.Q.Q. miR-203 inhibits augmented proliferation and metastasis of hepatocellular carcinoma residual in the promoted regenerating liver. Cancer Sci. 2017, 108, 338–346. [Google Scholar] [CrossRef]
- Wu, H.; Tao, J.; Li, X.; Zhang, T.; Zhao, L.; Wang, Y.; Zhang, L.; Xiong, J.; Zeng, Z.; Zhan, N.; et al. MicroRNA-206 prevents the pathogenesis of hepatocellular carcinoma by modulating expression of met proto-oncogene and cyclin-dependent kinase 6 in mice. Hepatology 2017, 66, 1952–1967. [Google Scholar] [CrossRef]
- Jia, P.; Wei, G.; Zhou, C.; Gao, Q.; Wu, Y.; Sun, X.; Li, X.X. Upregulation of MiR-212 inhibits migration and tumorigenicity and inactivates wnt/beta-catenin signaling in human hepatocellular carcinoma. Technol. Cancer Res. Treat. 2018, 17, 1533034618765221. [Google Scholar] [CrossRef]
- Wang, L.; Bo, X.; Zheng, Q.; Xiao, X.; Wu, L.; Li, B.B. miR-296 inhibits proliferation and induces apoptosis by targeting FGFR1 in human hepatocellular carcinoma. FEBS Lett. 2016, 590, 4252–4262. [Google Scholar] [CrossRef]
- Wang, L.; Yao, J.; Shi, X.; Hu, L.; Li, Z.; Song, T.; Huang, C.C. MicroRNA-302b suppresses cell proliferation by targeting EGFR in human hepatocellular carcinoma SMMC-7721 cells. BMC Cancer 2013, 13, 448. [Google Scholar] [CrossRef]
- Wang, L.; Yao, J.; Sun, H.; Sun, R.; Chang, S.; Yang, Y.; Song, T.; Huang, C.C. miR-302b suppresses cell invasion and metastasis by directly targeting AKT2 in human hepatocellular carcinoma cells. Tumour Biol. 2016, 37, 847–855. [Google Scholar] [CrossRef]
- Cui, H.; Song, R.; Wu, J.; Wang, W.; Chen, X.; Yin, J.J. MicroRNA-337 regulates the PI3K/AKT and Wnt/beta-catenin signaling pathways to inhibit hepatocellular carcinoma progression by targeting high-mobility group AT-hook 2. Am. J. Cancer Res. 2018, 8, 405–421. [Google Scholar] [PubMed]
- Zhang, T.; Liu, W.; Zeng, X.C.; Jiang, N.; Fu, B.S.; Guo, Y.; Yi, H.M.; Li, H.; Zhang, Q.; Chen, W.J.; et al. Down-regulation of microRNA-3383–p promoted angiogenesis in hepatocellular carcinoma. Biomed. Pharm. 2016, 84, 583–591. [Google Scholar] [CrossRef]
- Yuan, J.; Ji, H.; Xiao, F.; Lin, Z.; Zhao, X.; Wang, Z.; Zhao, J.; Lu, J.J. MicroRNA-340 inhibits the proliferation and invasion of hepatocellular carcinoma cells by targeting JAK1. Biochem. Biophys. Res. Commun. 2017, 483, 578–584. [Google Scholar] [CrossRef]
- Yu, M.; Xue, H.; Wang, Y.; Shen, Q.; Jiang, Q.; Zhang, X.; Li, K.; Jia, M.; Jia, J.; Xu, J.; et al. miR-345 inhibits tumor metastasis and EMT by targeting IRF1-mediated mTOR/STAT3/AKT pathway in hepatocellular carcinoma. Int. J. Oncol. 2017, 50, 975–983. [Google Scholar] [CrossRef]
- Pan, X.P.; Wang, H.X.; Tong, D.M.; Li, Y.; Huang, L.H.; Wang, C.C. miRNA-370 acts as a tumor suppressor via the downregulation of PIM1 in hepatocellular carcinoma. Eur. Rev. Med. Pharm. Sci. 2017, 21, 1254–1263. [Google Scholar]
- Liu, X.; Zhang, A.; Xiang, J.; Lv, Y.; Zhang, X.X. miR-451 acts as a suppressor of angiogenesis in hepatocellular carcinoma by targeting the IL-6R-STAT3 pathway. Oncol. Rep. 2016, 36, 1385–1392. [Google Scholar] [CrossRef]
- Zhang, M.; Wu, J.; Zhang, R.; Yang, J.; Zhang, Q.; Liu, B.B. miR-497 inhibits the carcinogenesis of hepatocellular carcinoma by targeting the Rictor/Akt signal pathway. Int. J. Clin. Exp. Pathol. 2019, 12, 1992–2000. [Google Scholar]
- Ye, Y.; Zhuang, J.; Wang, G.; He, S.; Zhang, S.; Wang, G.; Ni, J.; Wang, J.; Xia, W.W. MicroRNA-495 suppresses cell proliferation and invasion of hepatocellular carcinoma by directly targeting insulin-like growth factor receptor-1. Exp.Ther. Med. 2018, 15, 1150–1158. [Google Scholar] [CrossRef]
- Liu, Y.; Hong, W.; Zhou, C.; Jiang, Z.; Wang, G.; Wei, G.; Li, X.X. miR-539 inhibits FSCN1 expression and suppresses hepatocellular carcinoma migration and invasion. Oncol. Rep. 2017, 37, 2593–2602. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Zhang, D.; Jiang, J.; Dong, L.L. Loss of miR-638 promotes invasion and epithelial-mesenchymal transition by targeting SOX2 in hepatocellular carcinoma. Oncol. Rep. 2017, 37, 323–332. [Google Scholar] [CrossRef]
- Cheng, J.; Chen, Y.; Zhao, P.; Liu, X.; Dong, J.; Li, J.; Huang, C.; Wu, R.; Lv, Y.Y. Downregulation of miRNA-638 promotes angiogenesis and growth of hepatocellular carcinoma by targeting VEGF. Oncotarget 2016, 7, 30702–30711. [Google Scholar] [CrossRef]
- Huang, W.; Li, J.; Guo, X.; Zhao, Y.; Yuan, X.X. miR-663a inhibits hepatocellular carcinoma cell proliferation and invasion by targeting HMGA2. Biomed. Pharm. 2016, 81, 431–438. [Google Scholar] [CrossRef]
- Zhang, Y.; Wei, Y.; Li, X.; Liang, X.; Wang, L.; Song, J.; Zhang, X.; Zhang, C.; Niu, J.; Zhang, P.; et al. microRNA-874 suppresses tumor proliferation and metastasis in hepatocellular carcinoma by targeting the DOR/EGFR/ERK pathway. Cell Death Dis. 2018, 9, 130. [Google Scholar] [CrossRef]
- Ding, D.; Zhang, Y.; Yang, R.; Wang, X.; Ji, G.; Huo, L.; Shao, Z.; Li, X.X. miR-940 suppresses tumor cell invasion and migration via regulation of CXCR2 in hepatocellular carcinoma. Biomed. Res. Int. 2016, 2016, 7618342. [Google Scholar] [CrossRef]
- Lin, M.F.; Yang, Y.F.; Peng, Z.P.; Zhang, M.F.; Liang, J.Y.; Chen, W.; Liu, X.H.; Zheng, Y.L.L. FOXK2, regulted by miR-12715–p, promotes cell growth and indicates unfavorable prognosis in hepatocellular carcinoma. Int. J. Biochem. Cell Biol. 2017, 88, 155–161. [Google Scholar] [CrossRef] [PubMed]
- Zhu, H.; Wang, G.; Zhou, X.; Song, X.; Gao, H.; Ma, C.; Chang, H.; Li, H.; Liu, F.F.; Lu, J.; et al. miR-1299 suppresses cell proliferation of hepatocellular carcinoma (HCC) by targeting CDK6. Biomed. Pharm. 2016, 83, 792–797. [Google Scholar] [CrossRef]
- Yang, C.; Xu, Y.; Cheng, F.; Hu, Y.; Yang, S.; Rao, J.; Wang, X.X. miR-1301 inhibits hepatocellular carcinoma cell migration, invasion, and angiogenesis by decreasing Wnt/beta-catenin signaling through targeting BCL9. Cell Death Dis. 2017, 8, e2999. [Google Scholar] [CrossRef]
- Slack, F.J.; Chinnaiyan, A.M.M. The role of non-coding RNAs in oncology. Cell 2019, 179, 1033–1055. [Google Scholar] [CrossRef]
- Wang, X.W.; Heegaard, N.H.; Orum, H.H. MicroRNAs in liver disease. Gastroenterology 2012, 142, 1431–1443. [Google Scholar] [CrossRef]
- Zhang, T.; Yang, Z.; Kusumanchi, P.; Han, S.; Liangpunsakul, S.S. Critical role of microRNA-21 in the pathogenesis of liver diseases. Front. Med. (Lausanne) 2020, 7, 7. [Google Scholar] [CrossRef]
- Mishra, P.J.; Bertino, J.R.R. MicroRNA polymorphisms: The future of pharmacogenomics, molecular epidemiology and individualized medicine. Pharmacogenomics 2009, 10, 399–416. [Google Scholar] [CrossRef] [PubMed]
- Guo, X.; Lv, X.; Lv, X.; Ma, Y.; Chen, L.; Chen, Y.Y. Circulating miR-21 serves as a serum biomarker for hepatocellular carcinoma and correlated with distant metastasis. Oncotarget 2017, 8, 44050–44058. [Google Scholar] [CrossRef]
- Yoon, J.S.; Kim, G.; Lee, Y.R.; Park, S.Y.; Tak, W.Y.; Kweon, Y.O.; Park, J.G.; Lee, H.W.; Han, Y.S.; Ha, H.T.; et al. Clinical significance of microRNA-21 expression in disease progression of patients with hepatocellular carcinoma. Biomark. Med. 2018, 12, 1105–1114. [Google Scholar] [CrossRef]
- Zhou, Y.; Ren, H.; Dai, B.; Li, J.; Shang, L.; Huang, J.; Shi, X.X. Hepatocellular carcinoma-derived exosomal miRNA-21 contributes to tumor progression by converting hepatocyte stellate cells to cancer-associated fibroblasts. J. Exp. Clin. Cancer Res. 2018, 37, 324. [Google Scholar] [CrossRef] [PubMed]
- Chen, M.; Liu, Y.; Varley, P.; Chang, Y.; He, X.X.; Huang, H.; Tang, D.; Lotze, M.T.; Lin, J.; Tsung, A.A. High-mobility group box 1 promotes hepatocellular carcinoma progression through miR-21-mediated matrix metalloproteinase activity. Cancer Res. 2015, 75, 1645–1656. [Google Scholar] [CrossRef]
- Blaya, D.; Aguilar-Bravo, B.; Hao, F.; Casacuberta-Serra, S.; Coll, M.; Perea, L.; Vallverdu, J.; Graupera, I.; Pose, E.; Llovet, L.; et al. Expression of microRNA-155 in inflammatory cells modulates liver injury. Hepatology 2018, 68, 691–706. [Google Scholar] [CrossRef]
- Chen, Z.; Ma, T.; Huang, C.; Hu, T.; Li, J.J. The pivotal role of microRNA-155 in the control of cancer. J. Cell. Physiol. 2014, 229, 545–550. [Google Scholar] [CrossRef]
- Yan, X.L.; Jia, Y.L.; Chen, L.; Zeng, Q.; Zhou, J.N.; Fu, C.J.; Chen, H.X.; Yuan, H.F.; Li, Z.W.; Shi, L.; et al. Hepatocellular carcinoma-associated mesenchymal stem cells promote hepatocarcinoma progression: Role of the S100A4-miR155-SOCS1-MMP9 axis. Hepatology 2013, 57, 2274–2286. [Google Scholar] [CrossRef]
- Pineau, P.; Volinia, S.; McJunkin, K.; Marchio, A.; Battiston, C.; Terris, B.; Mazzaferro, V.; Lowe, S.W.; Croce, C.M.; Dejean, A.A. miR-221 overexpression contributes to liver tumorigenesis. Proc. Natl. Acad. Sci. USA 2010, 107, 264–269. [Google Scholar] [CrossRef]
- Johnson, S.M.; Grosshans, H.; Shingara, J.; Byrom, M.; Jarvis, R.; Cheng, A.; Labourier, E.; Reinert, K.L.; Brown, D.; Slack, F.J.J. RAS is regulated by the let-7 microRNA family. Cell 2005, 120, 635–647. [Google Scholar] [CrossRef]
- Winkler, I.; Bitter, C.; Winkler, S.; Weichenhan, D.; Thavamani, A.; Hengstler, J.G.; Borkham-Kamphorst, E.; Kohlbacher, O.; Plass, C.; Geffers, R.; et al. Identification of Ppargamma-modulated miRNA hubs that target the fibrotic tumor microenvironment. Proc. Natl. Acad. Sci. USA 2020, 117, 454–463. [Google Scholar] [CrossRef] [PubMed]
- Ji, J.; Zhao, L.; Budhu, A.; Forgues, M.; Jia, H.L.; Qin, L.X.; Ye, Q.H.; Yu, J.; Shi, X.; Tang, Z.Y.; et al. Let-7g targets collagen type I alpha2 and inhibits cell migration in hepatocellular carcinoma. J. Hepatol. 2010, 52, 690–697. [Google Scholar] [CrossRef]
- Kwon, J.J.; Factora, T.D.; Dey, S.; Kota, J.J. A systematic review of miR-29 in Cancer. Mol. Ther. Oncolytics 2019, 12, 173–194. [Google Scholar] [CrossRef]
- Xiong, Y.; Fang, J.H.; Yun, J.P.; Yang, J.; Zhang, Y.; Jia, W.H.; Zhuang, S.M.M. Effects of microRNA-29 on apoptosis, tumorigenicity, and prognosis of hepatocellular carcinoma. Hepatology 2010, 51, 836–845. [Google Scholar] [CrossRef]
- Su, H.; Yang, J.R.; Xu, T.; Huang, J.; Xu, L.; Yuan, Y.; Zhuang, S.M.M. MicroRNA-101, down-regulated in hepatocellular carcinoma, promotes apoptosis and suppresses tumorigenicity. Cancer Res. 2009, 69, 1135–1142. [Google Scholar] [CrossRef] [PubMed]
- Weidle, U.H.; Schmid, D.; Birzele, F.; Brinkmann, U.U. MicroRNAs Involved in Metastasis of Hepatocellular Carcinoma: Target Candidates, Functionality and Efficacy in Animal Models and Prognostic Relevance. Cancer Genom. Proteom. 2020, 17, 1–21. [Google Scholar] [CrossRef]
- Cheng, D.; Deng, J.; Zhang, B.; He, X.; Meng, Z.; Li, G.; Ye, H.; Zheng, S.; Wei, L.; Deng, X.; et al. LncRNA HOTAIR epigenetically suppresses miR-122 expression in hepatocellular carcinoma via DNA methylation. EBioMedicine 2018, 36, 159–170. [Google Scholar] [CrossRef]
- Tomimaru, Y.; Eguchi, H.; Nagano, H.; Wada, H.; Kobayashi, S.; Marubashi, S.; Tanemura, M.; Tomokuni, A.; Takemasa, I.; Umeshita, K.; et al. Circulating microRNA-21 as a novel biomarker for hepatocellular carcinoma. J. Hepatol. 2012, 56, 167–175. [Google Scholar] [CrossRef]
- Gibbings, D.J.; Ciaudo, C.; Erhardt, M.; Voinnet, O.O. Multivesicular bodies associate with components of miRNA effector complexes and modulate miRNA activity. Nat. Cell Biol. 2009, 11, 1143–1149. [Google Scholar] [CrossRef] [PubMed]
- Mitchell, P.S.; Parkin, R.K.; Kroh, E.M.; Fritz, B.R.; Wyman, S.K.; Pogosova-Agadjanyan, E.L.; Peterson, A.; Noteboom, J.; O’Briant, K.C.; Allen, A.; et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proc. Natl. Acad. Sci. USA 2008, 105, 10513–10518. [Google Scholar] [CrossRef]
- Hung, C.H.; Hu, T.H.; Lu, S.N.; Kuo, F.Y.; Chen, C.H.; Wang, J.H.; Huang, C.M.; Lee, C.M.; Lin, C.Y.; Yen, Y.H.; et al. Circulating microRNAs as biomarkers for diagnosis of early hepatocellular carcinoma associated with hepatitis B virus. Int. J. Cancer 2016, 138, 714–720. [Google Scholar] [CrossRef]
- Xie, Y.; Yao, Q.; Butt, A.M.; Guo, J.; Tian, Z.; Bao, X.; Li, H.; Meng, Q.; Lu, J.J. Expression profiling of serum microRNA-101 in HBV-associated chronic hepatitis, liver cirrhosis, and hepatocellular carcinoma. Cancer Biol. Ther. 2014, 15, 1248–1255. [Google Scholar] [CrossRef] [PubMed]
- Giray, B.G.; Emekdas, G.; Tezcan, S.; Ulger, M.; Serin, M.S.; Sezgin, O.; Altintas, E.; Tiftik, E.N.N. Profiles of serum microRNAs; miR-125b-5p and miR2233–p serve as novel biomarkers for HBV-positive hepatocellular carcinoma. Mol. Biol. Rep. 2014, 41, 4513–4519. [Google Scholar] [CrossRef] [PubMed]
- Zekri, A.N.; Youssef, A.S.; El-Desouky, E.D.; Ahmed, O.S.; Lotfy, M.M.; Nassar, A.A.; Bahnassey, A.A.A. Serum microRNA panels as potential biomarkers for early detection of hepatocellular carcinoma on top of HCV infection. Tumour Biol. 2016, 37, 12273–12286. [Google Scholar] [CrossRef]
- Chen, F.; Li, X.F.; Fu, D.S.; Huang, J.G.; Yang, S.E.E. Clinical potential of miRNA-221 as a novel prognostic biomarker for hepatocellular carcinoma. Cancer Biomark. 2017, 18, 209–214. [Google Scholar] [CrossRef] [PubMed]
- Xie, R.T.; Cong, X.L.; Zhong, X.M.; Luo, P.; Yang, H.Q.; Lu, G.X.; Luo, P.; Chang, Z.Y.; Sun, R.; Wu, T.M.; et al. MicroRNA-33a downregulation is associated with tumorigenesis and poor prognosis in patients with hepatocellular carcinoma. Oncol. Lett. 2018, 15, 4571–4577. [Google Scholar] [CrossRef]
- Fu, X.; Liu, M.; Qu, S.; Ma, J.; Zhang, Y.; Shi, T.; Wen, H.; Yang, Y.; Wang, S.; Wang, J.; et al. Exosomal microRNA-325–p induces multidrug resistance in hepatocellular carcinoma via the PI3K/Akt pathway. J. Exp. Clin. Cancer Res. 2018, 37, 52. [Google Scholar] [CrossRef]
- Shi, M.; Jiang, Y.; Yang, L.; Yan, S.; Wang, Y.G.; Lu, X.J.J. Decreased levels of serum exosomal miR-638 predict poor prognosis in hepatocellular carcinoma. J. Cell. Biochem. 2018, 119, 4711–4716. [Google Scholar] [CrossRef]
- Wang, Y.; Zhang, C.; Zhang, P.; Guo, G.; Jiang, T.; Zhao, X.; Jiang, J.; Huang, X.; Tong, H.; Tian, Y.Y. Serum exosomal microRNAs combined with alpha-fetoprotein as diagnostic markers of hepatocellular carcinoma. Cancer Med. 2018, 7, 1670–1679. [Google Scholar] [CrossRef]
- Chung, H.J.; Choi, Y.E.; Kim, E.S.; Han, Y.H.; Park, M.J.; Bae, I.H.H. miR-29b attenuates tumorigenicity and stemness maintenance in human glioblastoma multiforme by directly targeting BCL2L2. Oncotarget 2015, 6, 18429–18444. [Google Scholar] [CrossRef]
- Kong, Y.W.; Ferland-McCollough, D.; Jackson, T.J.; Bushell, M.M. microRNAs in cancer management. Lancet Oncol. 2012, 13, e249–e258. [Google Scholar] [CrossRef]
- Amodeo, V.; Bazan, V.; Fanale, D.; Insalaco, L.; Caruso, S.; Cicero, G.; Bronte, G.; Rolfo, C.; Santini, D.; Russo, A.A. Effects of anti-miR-182 on TSP-1 expression in human colon cancer cells: There is a sense in antisense? Expert Opin. Ther. Targets 2013, 17, 1249–1261. [Google Scholar] [CrossRef]
- Nedaeinia, R.; Sharifi, M.; Avan, A.; Kazemi, M.; Rafiee, L.; Ghayour-Mobarhan, M.; Salehi, R.R. Locked nucleic acid anti-miR-21 inhibits cell growth and invasive behaviors of a colorectal adenocarcinoma cell line: LNA-anti-miR as a novel approach. Cancer Gene Ther. 2016, 23, 246–253. [Google Scholar] [CrossRef] [PubMed]
- Kota, J.; Chivukula, R.R.; O’Donnell, K.A.; Wentzel, E.A.; Montgomery, C.L.; Hwang, H.W.; Chang, T.C.; Vivekanandan, P.; Torbenson, M.; Clark, K.R.; et al. Therapeutic microRNA delivery suppresses tumorigenesis in a murine liver cancer model. Cell 2009, 137, 1005–1017. [Google Scholar] [CrossRef]
- Hsu, S.H.; Yu, B.; Wang, X.; Lu, Y.; Schmidt, C.R.; Lee, R.J.; Lee, L.J.; Jacob, S.T.; Ghoshal, K.K. Cationic lipid nanoparticles for therapeutic delivery of siRNA and miRNA to murine liver tumor. Nanomedicine 2013, 9, 1169–1180. [Google Scholar] [CrossRef]
- Inchingolo, R.; Posa, A.; Mariappan, M.; Spiliopoulos, S.S. Locoregional treatments for hepatocellular carcinoma: Current evidence and future directions. World J. Gastroenterol. 2019, 25, 4614–4628. [Google Scholar] [CrossRef]
- Gottesman, M.M.M. Mechanisms of cancer drug resistance. Annu. Rev. Med. 2002, 53, 615–627. [Google Scholar] [CrossRef] [PubMed]
- Korita, P.V.; Wakai, T.; Shirai, Y.; Matsuda, Y.; Sakata, J.; Takamura, M.; Yano, M.; Sanpei, A.; Aoyagi, Y.; Hatakeyama, K.; et al. Multidrug resistance-associated protein 2 determines the efficacy of cisplatin in patients with hepatocellular carcinoma. Oncol. Rep. 2010, 23, 965–972. [Google Scholar]
- Wang, X.Q.; Ongkeko, W.M.; Chen, L.; Yang, Z.F.; Lu, P.; Chen, K.K.; Lopez, J.P.; Poon, R.T.; Fan, S.T.T. Octamer 4 (Oct4) mediates chemotherapeutic drug resistance in liver cancer cells through a potential Oct4-AKT-ATP-binding cassette G2 pathway. Hepatology 2010, 52, 528–539. [Google Scholar] [CrossRef]
- Meng, W.; Tai, Y.; Zhao, H.; Fu, B.; Zhang, T.; Liu, W.; Li, H.; Yang, Y.; Zhang, Q.; Feng, Y.; et al. Downregulation of miR-33a-5p in hepatocellular carcinoma: A possible mechanism for chemotherapy resistance. Med. Sci. Monit. 2017, 23, 1295–1304. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Wei, L.; Wang, X.; Lv, L.; Liu, J.; Xing, H.; Song, Y.; Xie, M.; Lei, T.; Zhang, N.; Yang, M.M. The emerging role of microRNAs and long noncoding RNAs in drug resistance of hepatocellular carcinoma. Mol. Cancer 2019, 18, 147. [Google Scholar] [CrossRef] [PubMed]
- Ou, Y.; Zhai, D.; Wu, N.; Li, X.X. Downregulation of miR-363 increases drug resistance in cisplatin-treated HepG2 by dysregulating Mcl-1. Gene 2015, 572, 116–122. [Google Scholar] [CrossRef]
- Ma, J.; Wang, T.; Guo, R.; Yang, X.; Yin, J.; Yu, J.; Xiang, Q.; Pan, X.; Tang, H.; Lei, X.X. Involvement of miR-133a and miR-326 in ADM resistance of HepG2 through modulating expression of ABCC1. J. Drug Target. 2015, 23, 519–524. [Google Scholar] [CrossRef]
- Shi, L.; Wu, L.; Chen, Z.; Yang, J.; Chen, X.; Yu, F.; Zheng, F.; Lin, X.X. MiR-141 activates Nrf2-dependent antioxidant pathway via down-regulating the expression of keap1 conferring the resistance of hepatocellular carcinoma cells to 5-fluorouracil. Cell. Physiol. Biochem. 2015, 35, 2333–2348. [Google Scholar] [CrossRef]
- Lee, H.; Kim, C.; Kang, H.; Tak, H.; Ahn, S.; Yoon, S.K.; Kuh, H.J.; Kim, W.; Lee, E.K.K. microRNA-200a-3p increases 5-fluorouracil resistance by regulating dual specificity phosphatase 6 expression. Exp. Mol. Med. 2017, 49, e327. [Google Scholar] [CrossRef]
- Wang, X.J.; Zhang, D.L.; Fu, C.; Wei, B.Z.; Li, G.J.J. MiR-183 modulates multi-drug resistance in hepatocellular cancer (HCC) cells via miR-183-IDH2/SOCS6-HIF-1alpha feedback loop. Eur. Rev. Med. Pharm. Sci. 2016, 20, 2020–2027. [Google Scholar]
- Yang, X.; Zang, J.; Pan, X.; Yin, J.; Xiang, Q.; Yu, J.; Gan, R.; Lei, X.X. miR-503 inhibits proliferation making human hepatocellular carcinoma cells susceptible to 5fluorouracil by targeting EIF4E. Oncol. Rep. 2017, 37, 563–570. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Xia, H.; Ooi, L.L.; Hui, K.M.M. MicroRNA-216a/217-induced epithelial-mesenchymal transition targets PTEN and SMAD7 to promote drug resistance and recurrence of liver cancer. Hepatology 2013, 58, 629–641. [Google Scholar] [CrossRef]
- Azumi, J.; Tsubota, T.; Sakabe, T.; Shiota, G.G. miR-181a induces sorafenib resistance of hepatocellular carcinoma cells through downregulation of RASSF1 expression. Cancer Sci. 2016, 107, 1256–1262. [Google Scholar] [CrossRef]
- Liu, K.; Liu, S.; Zhang, W.; Jia, B.; Tan, L.; Jin, Z.; Liu, Y.Y. miR-494 promotes cell proliferation, migration and invasion, and increased sorafenib resistance in hepatocellular carcinoma by targeting PTEN. Oncol. Rep. 2015, 34, 1003–1010. [Google Scholar] [CrossRef]
| Liver Disease | miRNA | Function | Reference |
|---|---|---|---|
| HBV infection | miR-122, miR-99 family | Promote HBV replication | [75,76] |
| miR-199-3p, miR-201 | Suppress HBV replication | [77] | |
| HCV infection | miR-141 | Promote HCV replication | [78,79] |
| let-7 family, miR-21 | Suppress liver fibrosis | [63,80] | |
| miR-16, miR-146a, miR-221, miR-222 | Biomarker for liver fibrosis | [81] | |
| ALD | let-7 family, miR-19b | Suppress liver fibrosis | [82,83] |
| NAFLD/NASH | miR-155 | Promote carcinogenesis | [84,85,86] |
| miRNA | Targets | Mechanism | Reference |
|---|---|---|---|
| miR-10b | CUB and sushi multiple domains 1 (CSMD1) | Division, migration, invasion | [109] |
| miR-21 | Kruppel like factors 5 (KLF1), Calmodulin regulated spectrin associated protein family member 1 (CAMSAP1), DEAD-box helicase 1 (DDX1), MARCKS like 1 (MARCKSL1) | Division, cell growth, migration, invasion | [110,111] |
| miR-25 | TNF related apoptosis-inducing ligand (TRAIL) | Apoptosis | [112] |
| miR-92a | F-box and WD repeat domain containing 7 (FBXW7) | Proliferation, cell cycle transition, apoptosis | [113] |
| miR-96-5p | Caspase-9 | Apoptosis | [114] |
| miR-107 | High mobility group AT-hock 2 (HMGA2), 3-hydroxy-3-methylglutaryl-CoA synthase 2 (HMGCS2) | Cell growth, Epithelial-mesenchymal transition (EMT) | [115,116] |
| miR-135a | Forkhead box O1 (FOX O1) | Migration, invasion | [117] |
| miR-155-5p | Phosphatase and tensin homolog (PTEN) | Proliferation, invasion, migration, apoptosis | [118] |
| miR-181a | Autophagy related 5 (ATG5) | Autophagy | [119] |
| miR-221 | P53, P53 upregulated modulator of apoptosis (PUMA), NF-κB, signal transducer and activator of transcription 3 (STAT3), adeno-associated virus serotype 8 (AAV8), PTEN, metalloproteinase inhibitor 3 (TIMP3), TRAIL | Apoptosis, proliferation | [37,38,39,40,120] |
| miR-203a-3p | Interleukin 24(IL-24) | Cell growth, metastasis | [121] |
| miR-210 | Fibroblast growth factor receptor like 1 (FGFRL1) | Angiogenesis | [122] |
| miR-214-5p | Wiskott-Aldrich syndrome like (WASL) | Invasion, migration | [123] |
| miR-302d | Transforming growth factor beta receptor 2 (TGFBR2) | Cell growth, invasion | [124] |
| miR-346 | F-box and leucine rich repeat protein 2 (FBXL2) | Proliferation, migration, invasion | [125] |
| miR-487a | EVH1 domain containing 2 (SPRED2) 2, phosphoinositide-3-Kinase regulatory 1 (PIK3R1) | Proliferation, metastasis | [126] |
| miR-765 | Inositol polyphosphate-4-phosphatase type II B (INPP4B) | Proliferation | [127] |
| miR-873 | Tumor suppressor in lung cancer 1 (TSLC1) | Proliferation, migration, invasion | [128] |
| miR-892a | CD266 molecule | Proliferation, invasion | [129] |
| miR-1249 | Patched 1 (PTCH1) | Cell growth, migration, invasion | [130] |
| miR-1468 | Cbp/p300 interacting transactivator with Glu/Asp rich carboxy-terminal domain 2 (CITED2), Up-frameshift protein 1 (UPF1) | Proliferation, apoptosis | [131] |
| miR-3910 | Macrophage stimulating 1 (MST1) | Cell growth, migration | [132] |
| miR-4417 | Tripartite motif containing 35 (TRIM35), pyruvate kinase muscle 2 (PKM2) | Proliferation, apoptosis | [133] |
| miRNA | Targets | Mechanism | Reference |
|---|---|---|---|
| miR-15b | B cell CLL/lymphoma 2 (BCL2) | Proliferation, apoptosis | [35] |
| miR-26 | unc-51 like autophagy activating kinase 1 (ULK1) | Autophagy | [134] |
| miR-29a | Claudin 1 (CLDN1) | Cell growth, migration | [135] |
| miR-31-5p | Sp1 transcription factor (SP1) | Proliferation, migration, invasion | [136] |
| miR-33b | Spalt-like transcription factor 4 (SALL4) | Proliferation, metastasis | [137] |
| miR-98 | Enhancer of zeste homolog 2 (EZH2) | Proliferation | [138] |
| miR-122 | A disintegrin and metalloproteinase domain 10 (ADAM10), ADAM17, insulin like growth factor 1 receptor (IGF1R), Serum response factor (SRF), Cyclin G1, Snail family transcriptional repressor 1 (SNAl1), SNAl2 | Proliferation, invasion, EMT | [41,139] |
| miR-125a-5p | Sirtuin 7 (SIRT7), erb-b2 receptor tyrosine kinase 3 (ERBB3) | Proliferation, apoptosis | [140,141] |
| miR-125b | Myeloid cell leukemia 1 (MCL1), BCLw, IL-6R, SIRT7 | Apoptosis, proliferation | [35,141] |
| miR-126 | Vascular endothelial growth factor (VEGF) | Angiogenesis | [142] |
| miR-137 | EZH2 | Proliferation | [143] |
| miR-142 | Transforming growth factor β (TGFβ), Thrombospondin 4 (THBS4) | Cell growth, metastasis, migration, invasion | [144,145] |
| miR-142-3p | Lactate dehydrogenase A (LDHA) | Proliferation | [146] |
| miR-144 | Zinc finger protein X-linked (ZFX) | Proliferation, invasion, migration | [147] |
| miR-187-3p | S100 calcium binding protein A4 (S100A4) | Metastasis, EMT | [148] |
| miR-194 | Mitogen-activated protein kinase kinase kinase kinase 4 (MAP4K4) | Proliferation | [149] |
| miR-195 | Cyclin D1/3, cyclin-dependent kinase 4/6 (CDK4/6), Cell division cycle 42 (CDC42), Vav guanine nucleotide exchange factor 2 (VAV2), E2F2, BCL2, BCLw, VEGF | Proliferation, apoptosis, angiogenesis, metastasis | [150,151] |
| miR-199a-3p | VEGFA, VEGFR1-2, Matrix metallopeptidase 2 (MMP2), Hepatocyte growth factor (HGF), Yes 1 associated transcriptional regulator 1 (YAP1) | Angiogenesis, proliferation, apoptosis | [152,153] |
| miR-199b-5p | TGFβ | EMT | [154] |
| miR-200a | GRB2 associated binding protein 1 (GAB1) | Invasion, migration | [155] |
| miR-203 | IL1β, SNAl1, Twist family bHLH transcription factor 1 (TWIST1) | Proliferation, metasis | [156] |
| miR-206 | Cyclin D1, CDK6 | Proliferation | [157] |
| miR-212 | FOXM1 | Migration, cell growth | [158] |
| miR-296 | Fibroblast growth factor receptor 1 (FGFR1) | proliferation, apoptosis | [159] |
| miR-302b | Epidermal growth factor receptor (EGFR), AKT serine/threonine kinase 2 (AKT2) | proliferation, invasion, metastasis | [160,161] |
| miR-337 | HMGA3 | Proliferation, apoptosis | [162] |
| miR-338-3p | Metastasis associated in colon cancer 1 (MACC1), β-catenin, VEGF | Angiogenesis | [163] |
| miR-340 | Janus kinase 1 (JAK1) | Proliferation, invasion | [164] |
| miR-345 | Interferon regulatory factor 1 (INF1) | Metastasis, EMT | [165] |
| miR-370 | Pim-1 proto-oncogene, serine/threonine kinase (PIM1) | Cell growth, invasion | [166] |
| miR-451 | IL6R | Angiogenesis | [167] |
| miR-497 | RPTOR independent companion of MTOR (RICTOR) | proliferation, migration and invasion | [168] |
| miR-495 | IGF1R | Proliferation, invasion | [169] |
| miR-539 | Fascin actin-bundling protein 1 (FSCN1) | Migration, invasion | [170] |
| miR-638 | SRY-box transcription factor 2 (SOX2), VEGF | Invasion, EMT, angiogenesis | [171,172] |
| miR-663a | HMGA2 | Proliferation, invasion | [173] |
| miR-874 | δ opioid receptor (DOR) | Proliferation, metastasis | [174] |
| miR-940 | C-X-C motif chemokine receptor 2 (CXCR2) | Migration, invasion | [175] |
| miR-1271-5p | FOXK2 | Cell growth | [176] |
| miR-1299 | CDK6 | Proliferation | [177] |
| miR-1301 | BCL9, β-catenin, VEGFA | Migration, invasion, angiogenesis | [178] |
| miRNA | Expression | Background | Reference | |
|---|---|---|---|---|
| Diagnosis | let-7 | up | HBV infection | [202] |
| miR-101 | down | HBV infection | [203] | |
| miR-122 | up | HBV infection, HCV infection | [202,205] | |
| miR-125a-5p | up | HBV infection, HCV infection | [204] | |
| miR-223-3p | down | HBV infection | [204] | |
| Poor prognosis | miR-32-5p | up | HBV infection and others | [208] |
| miR-33a | down | Unknown | [207] | |
| miR-92a | up | HBV infection and others | [113] | |
| miR-122 | up | HBV infection, HCV infection | [210] | |
| miR-137 | down | HBV infection and others | [143] | |
| miR-148a | up | HBV infection, HCV infection | [210] | |
| miR-194 | down | Unknown | [149] | |
| miR-221 | up | HBV infection and others | [206] | |
| miR-296 | down | HBV infection and others | [159] | |
| miR-487a | up | HBV infection and others | [126] | |
| miR-638 | down | HBV infection, HCV infection and others | [209] | |
| miR-940 | down | HBV infection and others | [175] | |
| miR-1246 | up | HBV infection, HCV infection | [210] | |
| miR-1468 | up | HBV infection and others | [131] |
| Drug | miRNA | Target | Reference |
|---|---|---|---|
| Cisplatin | miR-33a-5p | SOCS3 | [220,221] |
| miR-363 | MCL1 | [223] | |
| Doxorubicin | miR-133a | ABC subfamily C1 | [224] |
| miR-326 | ABC subfamily C1 | [224] | |
| 5-FU | miR-141 | Kelch-like ECH-associated protein 1 (KEAP1), Nuclear factor erythroid 2 like 2 (NFE2L2) | [225] |
| miR-125b | BCL2 | [14] | |
| miR-195 | BCL2 | [14] | |
| miR-503 | eukaryotic translation initiation factor 4E (EIF4E) | [228] | |
| Sorafenib | miR-93 | p21Cip/Waf1 | [222] |
| miR-181 | Ras association domain family 1 (RASSF1) | [230] | |
| miR-216a | p21Cip/Waf1 | [222] | |
| miR-217 | p21Cip/Waf1 | [222] | |
| miR-494 | PTEN, mTOR | [231] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Oura, K.; Morishita, A.; Masaki, T. Molecular and Functional Roles of MicroRNAs in the Progression of Hepatocellular Carcinoma—A Review. Int. J. Mol. Sci. 2020, 21, 8362. https://doi.org/10.3390/ijms21218362
Oura K, Morishita A, Masaki T. Molecular and Functional Roles of MicroRNAs in the Progression of Hepatocellular Carcinoma—A Review. International Journal of Molecular Sciences. 2020; 21(21):8362. https://doi.org/10.3390/ijms21218362
Chicago/Turabian StyleOura, Kyoko, Asahiro Morishita, and Tsutomu Masaki. 2020. "Molecular and Functional Roles of MicroRNAs in the Progression of Hepatocellular Carcinoma—A Review" International Journal of Molecular Sciences 21, no. 21: 8362. https://doi.org/10.3390/ijms21218362
APA StyleOura, K., Morishita, A., & Masaki, T. (2020). Molecular and Functional Roles of MicroRNAs in the Progression of Hepatocellular Carcinoma—A Review. International Journal of Molecular Sciences, 21(21), 8362. https://doi.org/10.3390/ijms21218362





