Abstract
Inflammasomes are multimolecular complexes with potent inflammatory activity. As such, their activity is tightly regulated at the transcriptional and post-transcriptional levels. In this review, we present the transcriptional regulation of inflammasome genes from sensors (e.g., NLRP3) to substrates (e.g., IL-1β). Lineage-determining transcription factors shape inflammasome responses in different cell types with profound consequences on the responsiveness to inflammasome-activating stimuli. Pro-inflammatory signals (sterile or microbial) have a key transcriptional impact on inflammasome genes, which is largely mediated by NF-κB and that translates into higher antimicrobial immune responses. Furthermore, diverse intrinsic (e.g., circadian clock, metabolites) or extrinsic (e.g., xenobiotics) signals are integrated by signal-dependent transcription factors and chromatin structure changes to modulate transcriptionally inflammasome responses. Finally, anti-inflammatory signals (e.g., IL-10) counterbalance inflammasome genes induction to limit deleterious inflammation. Transcriptional regulations thus appear as the first line of inflammasome regulation to raise the defense level in front of stress and infections but also to limit excessive or chronic inflammation.
Keywords:
inflammasome; NF-κB; IRF; NLRP3; caspase-1; epigenetic modification; transcription factor; chromatin; promoter; enhancer 1. Introduction
A common theme in inflammatory signaling pathways is their tight control. The timely detection of pathogens and an appropriate magnitude of response are often dependent on the upregulation of specific sensors and/or their downstream adaptors/effectors. Yet, maintaining high expression of pattern recognition receptors or inflammatory mediators, in addition to being energetically costly, might trigger detection of endogenous molecules and spontaneous inflammation, causing autoinflammatory syndromes [1].
Two main categories of regulation act synergistically and in an intertwined manner to ensure appropriate responses; the first one at the transcriptional level and the second one at the post-transcriptional level (which includes post-transcriptional sensus-stricto, translational, and post-translational levels). In this review, we will focus on the first line of inflammasomes regulation: the transcriptional regulation. Post-transcriptional regulations of inflammasome components have been reviewed recently [2,3,4,5,6]. The roles of noncoding RNAs (which include microRNAs (miRNA) and long noncoding RNAs (lncRNA)) will not be described since miRNAs affect mRNA stability and translation rate downstream of transcription and since inflammasome regulation by lncRNA is either indirect [7] or mediated by a direct action onto inflammasome proteins [8].
After a brief introduction on inflammasomes and transcription regulation, we will focus on the transcriptional regulation of specific inflammasome genes illustrating whenever possible both the mechanisms and the functional consequences. Inflammasome regulation differs between humans and mice; we will thus present the regulations occurring in these two species. Of note, all uppercase letters are used for human GENES symbols and capitalized words for murine Genes symbols.
2. Overview of Inflammasome Complexes
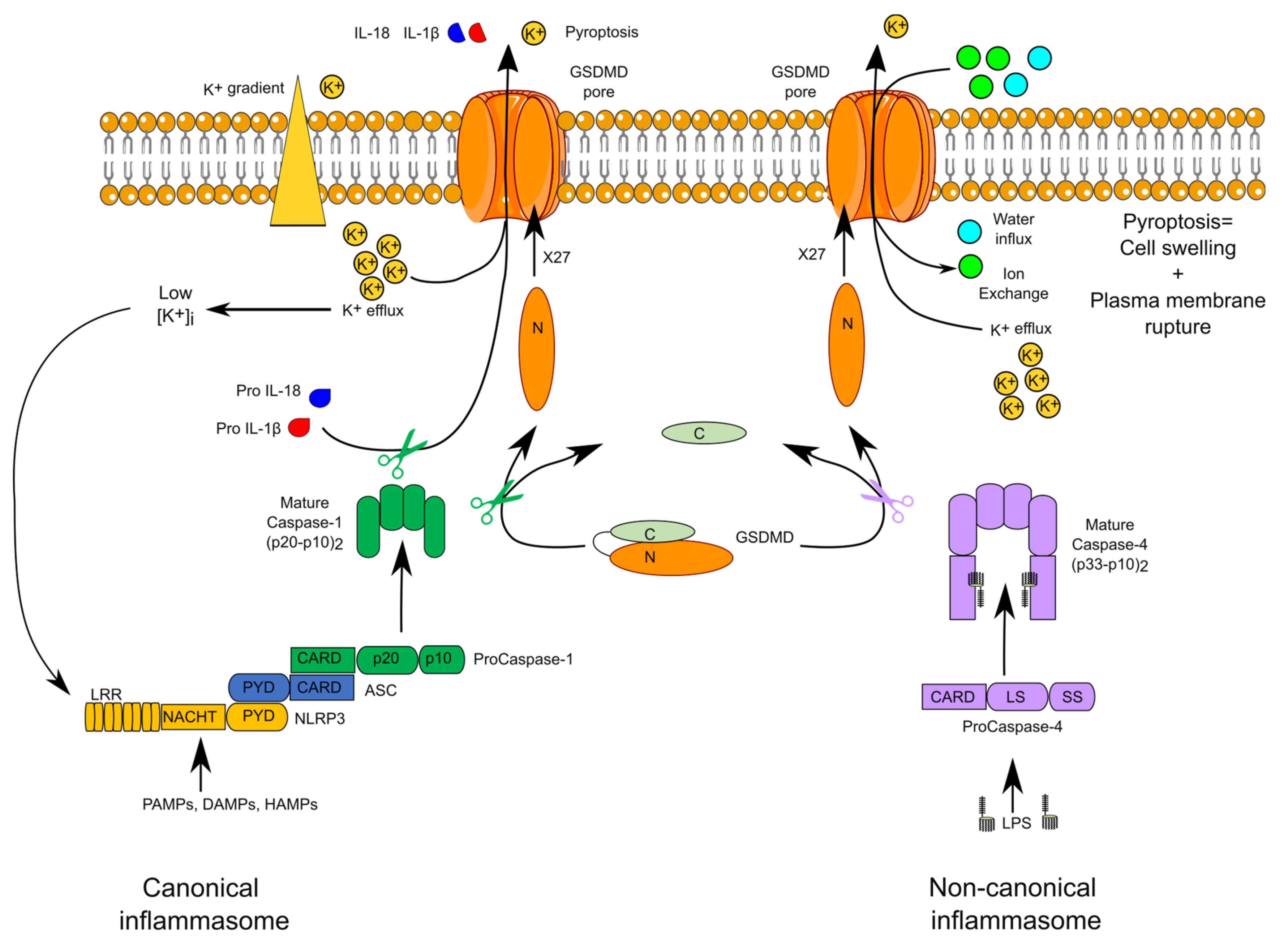
Inflammasomes are multimolecular protein complexes assembled in the cytosol in response to pathogen-associated molecular patterns (PAMPs), damage-associated molecular signals (DAMPs), or homeostasis-altering molecular processes (HAMPs) [9] (Figure 1). Formation of these complexes leads to the activation of inflammatory caspases. Canonical inflammasomes and noncanonical inflammasomes activate caspase-1 and caspase-4/caspase-5 in humans, whereas they activate caspase-1 and caspase-11 in mice. The adaptor ASC (Apoptosis-associated speck-like protein containing a CARD) is the central piece of canonical inflammasomes and connects inflammasome sensors with caspase-1. Inflammasome sensors include members of the NLR (nucleotide-binding domain and leucine-rich repeat containing) family (NLRP1, NLRP3, NAIP, and its adaptor NLRC4), AIM2 (Absent in Melanoma 2) and pyrin. The nature of the sensor defines the name of the inflammasome complex (e.g., the NLRP3 inflammasome).
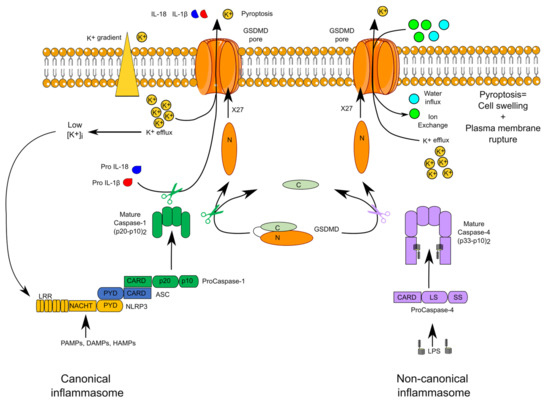
Figure 1.
Canonical and noncanonical inflammasomes. The schematic structure of the canonical NLRP3 and noncanonical caspase-4 inflammasomes are displayed. Mature self-cleaved caspase-1 and caspase-4 cleave gasdermin D (GSDMD) leading to oligomerization of the N-terminal fragment in the membrane. The GSDMD pore promotes IL-1β and IL-18 secretion and K+ efflux. The latter activates the NLRP3 inflammasome.
In noncanonical inflammasomes, caspase-4, or its murine homologue caspase-11, acts directly both as sensor of cytosolic lipopolysaccharide (LPS) and as the effector caspase [10]. The physiological role of caspase-5 remains unclear although caspase-5 senses cytosolic LPS and Pseudomonas aeruginosa (P. aeruginosa) outer membrane vesicles [11].
Upon activation within inflammasomes, caspase-1 triggers two main events: a fast inflammatory cell death, termed pyroptosis, and the cleavage of the proform of two cytokines (IL-1β and IL-18), resulting in the secretion of their active forms. Pyroptosis is due to the caspase-1-mediated cleavage of a single protein, termed gasdermin D (GSDMD). The cleaved N-terminal fragment of GSDMD oligomerizes into the plasma membrane to form a pore [12,13]. Structural work on GSDMA3, another GSDM family member, indicates that the mature GSDM pore is formed by 27 GSDM monomers creating a 180 Å diameter channel permitting the transport of IL-1 cytokines and of numerous ions, leading in most cases to an osmotic cell death [14]. In noncanonical inflammasomes, caspase-4 and caspase-11 cleave GSDMD and thus directly trigger pyroptosis [15]. The GSDMD pore is a nonselective pore mediating the exchange of numerous ions and molecules. Efflux of K+ is particularly important since it activates the NLRP3 inflammasome downstream of noncanonical inflammasomes [16,17].
Inflammasomes are implicated in the antimicrobial responses to numerous pathogens including Salmonella enterica, Legionella pneumophila, Francisella tularensis, and vaccinia virus [18]. Yet, inflammasome activation can also be deleterious as demonstrated in murine models of septic shock or Alzheimer’s disease [19] and in patients suffering from autoinflammatory syndromes due to gain of function mutations in inflammasome sensor genes [20]. Inflammasomes are thus tightly regulated with numerous positive and negative regulations ensuring, in most situations, a proper balance between immune defenses against infection and the absence of chronic sterile inflammation.
Here, we review the first level of inflammasome regulation, namely the transcriptional regulation. Importantly, transcriptional regulation affects most, if not all, inflammasome molecules from sensors to downstream inflammasome products (pro-inflammatory cytokines, GSDMD) and, has a profound impact on the inflammasome responses [21,22,23,24,25,26]. Before developing the specific transcriptional regulation of inflammasome genes, we will give a brief overview of transcription regulation.
3. Overview of Transcription Regulation
Transcription is the generation of an RNA molecule from a DNA template. Protein-encoding genes are transcribed by the RNA polymerase II (Pol II), whose activity is controlled at the recruitment phase, at the initiation of elongation (that requires to overcome Pol II pausing), and at its processivity level.
One key aspect of transcriptional regulation is mediated by regulating Pol II and transcription factors (TFs) access to DNA. DNA accessibility is highly controlled and limited in most of the genome due to high chromatin compaction. Nucleosomes (i.e., a ≈150 bp-long DNA piece wrapped around a histone octamer) represent the first level of chromatin organization. Nucleosomes can be located at the transcription start site (TSS), on the TATA box, or on specific transcription factor binding sites and can therefore block transcription. Chromatin remodeling allows TFs to access their target genes and control transcription. This process involves two families of enzymes: the ATP-dependent nucleosome remodeling complexes (e.g., from the switch/sucrose nonfermenting-SWI/SNF family) and the histone modifying enzymes (e.g., histone acetyl transferases-HAT). The histone-modifying enzymes add methyl or acetyl groups to histone tails. These covalent modifications change the affinity of histones for DNA and modify their ability to bind transcription coactivators or corepressors. As an example, histone H3 lysine 27 trimethylation-(H3K27me3) is a repressive epigenetic modification. In contrast, H3K4me3 and H3K9 acetylation-(H3K9ac) are activating modifications and H3K27ac is a mark of transcriptional activity. H3K4me1 is an enhancer-specific mark.
TFs binding sites are located in DNA sequences called promoter in the vicinity of the TSS and promoting Pol II recruitment. TFs binding sites can also be located megabases away from the TSS in regions called enhancers or silencers. These regions, often far away when considering the linear distance along the genome, can interact through DNA loops and be in close proximity when considering the three-dimensional chromatin structure. Transcription regulation thus integrates signals from both promoter and enhancer/silencer regions.
The cell type-specific and the signal-dependent transcriptional regulations are controlled by the hierarchical and coordinated binding of unique combinations of transcription factors. TFs can be classified in two categories: the pioneer TFs (e.g., PU.1) and the signal-dependent TFs (e.g., nuclear factor κB, NF-κB). Pioneer TFs bind to their specific recognition motifs, most often in enhancers, even in the context of highly compact chromatin. These pioneer TFs displace nucleosomes, open chromatin structure and promote DNA accessibility, revealing novel TFs binding sites for nonpioneer TFs. Pioneers TFs are expressed during lineage development and drive constitutive and signal-dependent, lineage-specific transcription programs. They are thus also named lineage-determining TFs. Binding of pioneer TFs on inactive enhancers transforms them into primed, poised (see IL1B enhancer below), or active enhancers [27] that present three different epigenetic states with specific characteristics. In contrast to inactive enhancers displaying highly compact chromatin, active enhancers present a wide nucleosome-free region, recruit Pol II, which results in strong expression of target genes. Primed enhancers display constitutive binding of lineage-determining/pioneer TFs, a nucleosome-free region of open chromatin but very low recruitment of paused Pol II. Poised enhancers are primed enhancers displaying the additional presence of repressive epigenetic marks (e.g., H3K27me3). During activation, they can rapidly change from poised to active state, a change associated with a switch from H3K27me3 to H3K27ac.
Productive transcription requires both Pol II recruitment and transition from a paused Pol II to an elongating Pol II. These two steps are indirectly controlled by TFs through histone modifications. Pol II recruitment can follow HAT recruitment and activation through direct interaction with TFs. The ensuing histone acetylation marks the chromatin for assembly of the preinitiation complex that includes Pol II bound to DNA close to the TSS. Transcription elongation is also mediated by TFs, which recruit the positive elongation factors bromodomain-containing protein 4 (Brd4) and P-TEFb (positive transcription elongation factor b). P-TEFb phosphorylates Pol II to promote productive elongation.
Transcription regulation is thus a complex process resulting from the hierarchical and collaborative actions of multiple lineage-determining TFs and signal-dependent TFs. These TFs act by modifying, in multiple stable or dynamic ways, chromatin structure to ensure Pol II recruitment and productive elongation in a timely controlled manner. Due to the high complexity of this process and the multitude of TFs binding sites affecting the transcription of a particular gene, almost each gene has its own spatiotemporal expression although common themes can be observed.
4. Transcriptional Regulation of Inflammasome Sensors
4.1. NLRP1
NLR Family Pyrin Domain Containing 1 (NLRP1) is an inflammasome sensor sensing cytosolic proteolytic activity. Anthrax lethal factor, a toxin with endoprotease activity, and IpaH7.8, a Shigella flexneri effector with ubiquitin ligase activity, trigger NLRP1B degradation and activation [28,29]. Human NLRP1, the three murine paralogues NLRP1A, B, and C, each one with different isoforms in different mouse strains, detect different stimuli [29].
In mice, Nlrp1a expression is restricted to the hematopoietic compartment and is expressed in hematopoietic stem cells, progenitor cells of both myeloid and lymphoid origins, and terminally differentiated cells such as macrophages [30].
Expression of Nlrp1a and Nlrp1c is positively regulated by SREBP-1A (Sterol regulatory element binding protein-1a), a basic helix–loop–helix leucine zipper (bHLH-LZ) TF [31]. SREBP-1A binds directly onto a canonical SREBP-1 binding site in the Nlrp1a proximal promoter [31]. Srebp-1a expression is itself under the direct control of NF-κB, in synergy with the monocyte/macrophage specific TF PU.1 (an Erythroblast Transformation Specific (ETS) family TF). NF-κB thus indirectly controls Nlrp1a expression in macrophages (Figure 2A) [31]. SREBP-1A also controls LPS-triggered lipogenesis in macrophages, an anabolic pathway required for optimal inflammasome responses. SREBP-1A regulation thus directly couples lipogenesis and control of the NLRP1A inflammasome [31]. SREBP TFs being key signaling nodes responding to metabolic clues, the control of Nlrp1a expression might thus participate to the control of metabolic inflammation. Interestingly, mice lacking the three murine Nlrp1 alleles (or Il18) develop metabolic syndrome and spontaneous obesity strengthening the functional link between metabolism and Nlrp1 sensors [32].
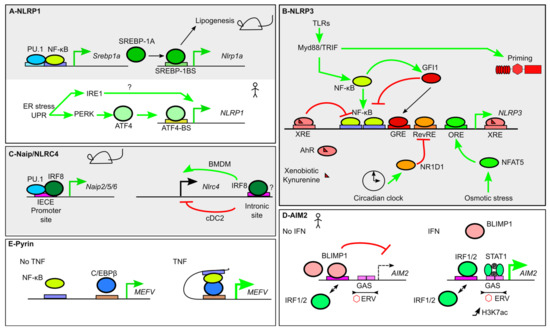
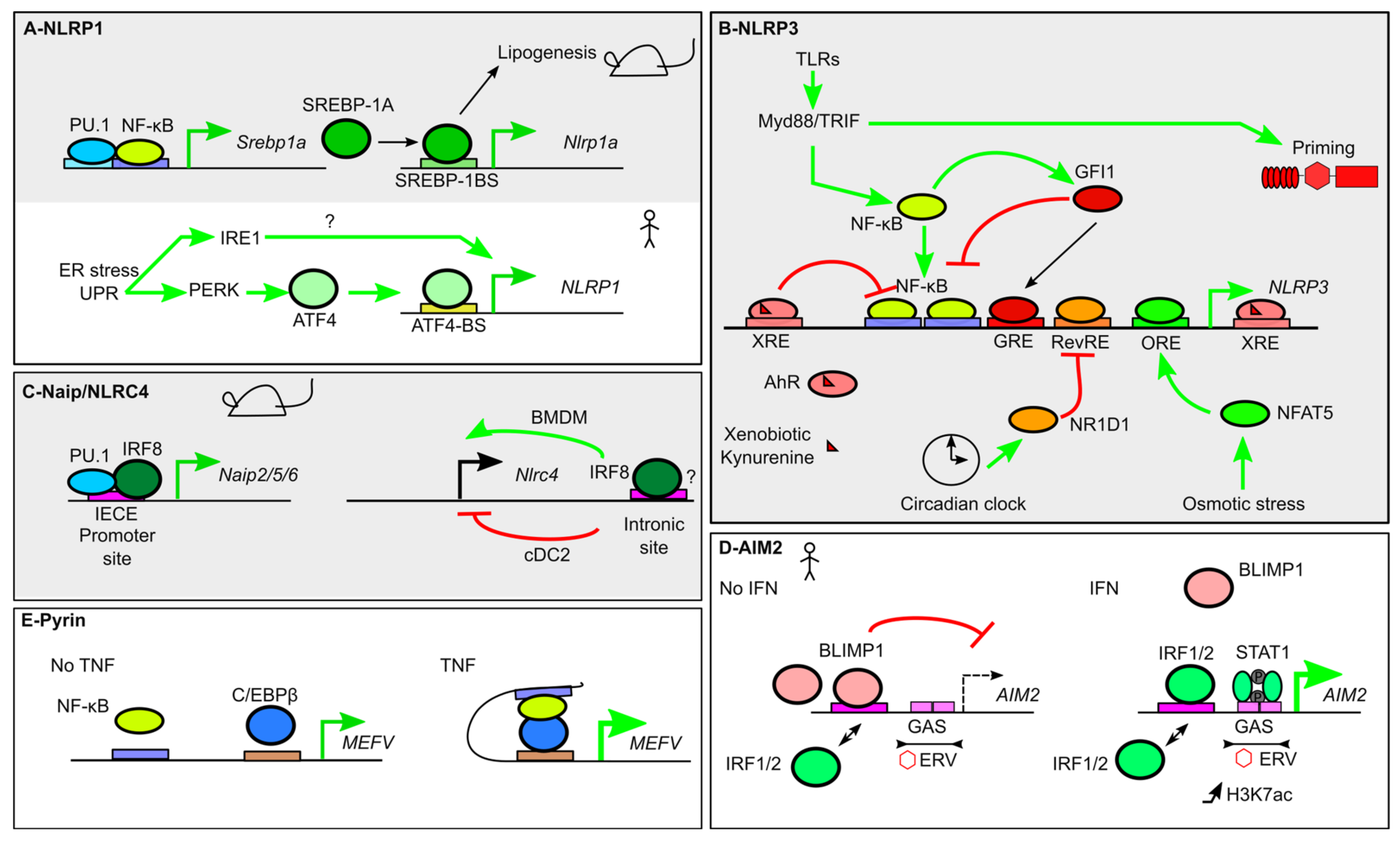
Figure 2.
Transcriptional regulation of inflammasome sensors. (A) In mice (gray background), Nlrp1a expression is indirectly induced by Nuclear factor κB (NF-κB), which controls Srebp1a expression. SREBP1A couples Nlrp1a expression and lipogenesis. In humans, NLRP1 expression is induced by ER stress and the Unfolded protein response (UPR) and two of its effectors, Inositol-requiring enzyme 1 (IRE1) and PKR-like ER protein kinase (PERK). PERK induces Activating Transcription Factor 4 (ATF4) expression that directly upregulates NLRP1 expression. (B) NLRP3 expression is under the control of numerous TFs. NF-κB activation downstream of Toll-like receptors (TLRs) induces NLRP3 expression, while NLRP3 inflammasome priming is independent on transcription. GFI1 negatively regulates NLRP3 expression. GFI1 expression is inducible by NF-κB, thus creating a negative feedback loop. Xenobiotics and anti-inflammatory metabolites promote Aryl hydrocarbon receptor (AhR) binding to Xenobiotic Response Elements (XRE) that blocks NF-κB-mediated NLRP3 induction. Similarly, NLRP3 promoter integrates signals from the circadian clock regulator Nuclear receptor subfamily 1 group d member 1 (NR1D1) and from Nuclear factor of activated T cells 5 (NFAT5), which binds to an Osmotic Response Element (ORE), following osmotic stress. (C) Several Naip are regulated at steady state by Interferon Regulatory Factor 8 (IRF8), which binds Ets-Interferon Regulatory Factors (IRF) composite Element (EICE), together with PU.1. Binding of IRF8 in the intronic region of Nlrc4 controls positively or negatively Nlrc4 expression in bone marrow-derived macrophages (BMDM) and type 2 conventional dendritic cells (cDC2), respectively. (D) At steady state (left panel) AIM2 expression is repressed due to B lymphocyte-induced maturation protein-1 (BLIMP1) binding to an IRF/BLIMP1 composite site. In the presence of IFN-γ, IRF1/2 displace BLIMP1. Furthermore, Signal Transducer and Activator of Transcription 1 (STAT1) binding to GAS triggers AIM2 expression, which correlates with a local increase in H3K7 acetylation (H3K7ac). The tandem GAS sites are present in an endogenous retrovirus sequence (ERV). (E) Pyrin encoded by the MEFV gene is under the regulation of NF-κB. In the presence of Tumor necrosis factor (TNF), a chromatin loop favors NF-κB and CCAAT/enhancer binding protein β (C/EBPβ) interaction to upregulate MEFV expression.
In humans, NLRP1 is broadly expressed but at particular high levels in keratinocytes, a feature that is not observed in mice [33]. The molecular basis of the differential expression of NLRP1 between humans and mice remains unknown. It is controlled at the transcriptional level since inflammasome transcripts are very low in murine keratinocytes [34].
Endoplasmic reticulum (ER) stress strongly induces NLRP1 expression in THP-1 monocytes and in various human cell lines. This induction is dependent on two effectors of the unfolded protein response (UPR): PKR-like ER protein kinase (PERK) and Inositol-requiring enzyme 1 (IRE1). Although the mechanisms downstream of IRE1 remain unsolved, PERK activation leads to Activating Transcription Factor 4 (ATF4) expression. ATF4 is a TF from the ATF/cAMP Response Element-binding protein (CREB) family. ATF4 binds to the promoter of human NLRP1 to induce its expression (Figure 2A) [35]. It is thus tempting to speculate that ER stress could prime the NLRP1 inflammasome to increase its ability to detect HAMPs and PAMPs.
4.2. NLRP3
NLRP3 is an inflammasome sensor detecting multiple cellular homeostasis perturbations such as membrane damage, mitochondrial defects, or perturbations of ionic concentrations. NLRP3 inflammasome activation is a two-step process and requires a signal 1 (priming signal, e.g., LPS) and a signal 2 (activation signal, e.g., nigericin). Priming is triggered by pro-inflammatory signals such as Toll-like receptor 4 (TLR4) engagement and has long been associated with transcriptional upregulation of NLRP3 [36,37]. It is now clear that NLRP3 priming is mediated by post-translational modifications independent of transcription (Figure 2B) [38,39,40]. Yet, Nlrp3 upregulation accelerates the kinetics and the level of caspase-1 activation following signal 2 addition [38]. The importance of this upregulation is likely enhanced by the low level of expression of Nlrp3 at steady state.
NLRP3 upregulation occurs within 2 h of LPS addition but can also be triggered by other TLR ligands, by the nucleotide-binding oligomerization domain 2 (Nod2) ligand (muramyl dipeptide) or by pro-inflammatory cytokines (Tumor necrosis factor (TNF), Interleukin (IL)-1α, IL-1β) in the absence of PAMPs [36,37]. This upregulation is dependent on NF-κB as first demonstrated using the BAY 11-7082 inhibitor. Two NF-κB binding sites are present in the NLRP3 promoter. Upon LPS treatment, the NF-κB subunit RelA/p65 binds the NLRP3 promoter. LPS-induced NLRP3 upregulation is lost upon mutation of these two binding sites indicating that these two sites control in a redundant manner the increase in NLRP3 promoter activity [41]. Upregulation of Nlrp3 (and Il1b) following NF-κB activation is partially dependent on an atypical IκBs (inhibitor of κB), a coactivator of NF-κB in bone marrow-derived macrophages (BMDMs) [42]. IκBζ binds to another NF-κB subunit, p50, in the Nlrp3 promoter. The NF-κB heterodimer RelA/p50 may thus be responsible for LPS-induced Nlrp3 expression. IκBζ recruitment increases H3K4me3, a mark of active transcription [43] suggesting that this coactivator promotes Nlrp3 induction through epigenetic modifications. Interestingly, the parasite Leishmania amazonensis subverts this process by targeting the epigenetic control of NF-κB-related pro-inflammatory genes. During infection, the promoters of these genes display hypoacetylation of the histone H3K9/14 and hypotrimethylation of histone H3K4. These inhibitory epigenetic modifications dampen Nlrp3 (and also Aim2, Nlrc4, Pycard, Il1b, and Il18) expression and promotes survival of the parasite within the host [44].
In addition to its control by pro-inflammatory signals, Nlrp3 expression is controlled by the circadian clock and presents a peak of expression during the night [45,46]. The circadian rhythm in Nlrp3 transcript level is dependent on a master regulator of the clock, the transcription factor NR1D1 (nuclear receptor subfamily 1 group D member 1, also known as Rev-erbα). Circadian oscillations of Nlrp3 and Nr1d1 expressions are in antiphase. Accordingly, NR1D1 binds directly to a Rev-response element (RevRE) in the Nlrp3 promoter to repress its expression (Figure 2B). Interestingly, an agonist of NR1D1 attenuates both dextran sodium sulfate (DSS)-induced colitis and D-galactosamine-induced fulminant hepatitis suggesting that NR1D1-mediated circadian control of Nlrp3 expression controls inflammation in vivo [45,46].
Besides NF-κB and NR1D1, direct binding of several TFs on the NLRP3 promoter have been reported with either negative or positive impact on NLRP3 expression. Downregulation of NLRP3 expression is controlled by the aryl hydrocarbon receptor (AhR) [47] and by B cell lymphoma 6 (BCL6), a transcriptional repressor that antagonizes NF-κB-mediated gene transcription [48]. AhR is a ligand-activated TF binding numerous environmental contaminants (e.g., dioxin) and endogenous ligands (e.g., the tryptophan derivative, kynurenine). AhR binds two XRE (Xenobiotic Responsive Element) sequences in the Nrlp3 promoter, and its binding is increased in a ligand-dependent manner. The two XRE sites surround the two NF-κB sites in the Nlrp3 promoter suggesting that AhR may interfere with NF-κB recruitment either directly or indirectly by modifying the local chromatin architecture. AhR and its ligands also negatively control Il1b expression indicating that environmental pollutants (and the anti-inflammatory tryptophan catabolites) dampen inflammasome responses (Figure 2B) [47].
Growth Factor Independence 1 (GFI1), a protein induced by NF-κB, binds the Nlrp3 promoter to inhibit Nlrp3 expression in a negative feedback loop [49]. In contrast, direct binding of Nuclear factor of activated T cells 5 (NFAT5—a transcription factor induced in response to high salt, hypoxia and mechanical stress) on an osmotic response element (ORE) in the promoter of Nlrp3 positively regulates Nlrp3 expression [50]. Overall, regulation of Nlrp3 expression emerges as a complex network primarily driven by pro-inflammatory signals but integrating environmental and intrinsic signals.
Nlrp3 expression has been mostly studied in macrophages and displays a cell-type-specific regulation in dendritic cells (DCs). Of note, the DC-specific gene regulation affects not only Nlrp3 but also other inflammasome genes as detailed below. Indeed, plasmacytoid dendritic cells (pDC) express neither Nlrp3 nor Il1b and are resistant to inflammasome activation [51]. Conventional DCs (cDC) have a limited ability to respond to inflammasome stimuli [21,51]. Interferon Regulatory Factors (IRFs) belong to a family of TFs, which includes 9 members in human and mice. pDCs express high levels of IRF8, while cDC1 and cDC2 express high levels of IRF8 and IRF4, respectively. IRF4 and IRF8 bind the promoter regions of Nlrp3, Il1b, and Aim2 and intronic regions of Pycard and Nlrc4, which correlate with the low expression of these genes in cDC1 and cDC2. Haploinsufficiency in Irf8 increases Nlrc4, Pycard, and Il1b expression in cDC1 and increases the inflammasome response to the NLRC4-engaging pathogen Salmonella typhimurium (S. typhimurium). In contrast, ectopic expression of Irf8 in macrophages decreases Nlrp3, Nlrc4, and il1b expression, while ectopic expression of Irf4 decreases Nlrc4, Il1b, and Pycard expression [21]. Importantly, the low inflammasome activity in cDC limits their pyroptosis in response to bacterial pathogens and favors antigen presentation and T cell priming.
4.3. NAIP and NLRC4
The Neuronal apoptosis inhibitory protein (NAIP)/NLRC4 inflammasome is atypical since NLRC4 is not a direct sensor but requires a NAIP protein to sense PAMPs. A single NAIP protein is encoded in the human genome and binds bacterial type III secretion system (T3SS) needle proteins [52] and flagellin [53]. Several NAIP paralogs are present in mice and bind T3SS needle proteins (NAIP1), T3SS inner rod proteins (NAIP2), or flagellin (NAIP5/6) [54,55].
IRF8 was identified in mice as a positive regulator of Naip2, 5, 6, and Nlrc4. IRF8 binds the promoter regions of Naip2, 5, 6, and an intronic region of Nlrc4. IRF8 has a very low intrinsic DNA-binding activity and, at steady state, binds with PU.1 at Ets-IRF composite elements (EICE) (Figure 2C) [56]. NAIP/NLRC4 inflammasomes are the major inflammasomes allowing mice to control S. typhimurium infection. Accordingly, Irf8−/− mice are highly susceptible to S. typhimurium [25]. Interestingly, although the effects of IRF8 were demonstrated at steady state, Irf8 is induced upon Legionella infection and could participate in the NAIP/NLRC4-dependent response against this flagellin-expressing pathogen [57]. Surprisingly, overexpression of Irf8 in bone marrow-derived macrophages decreases Nlrc4 expression. Furthermore, as presented above, in cDC1, IRF8 inhibits Nlrc4 expression (Figure 2C) [21]. IRF8 concentration modulates its ability to cooperate with other TFs and engage different binding sites [58] possibly explaining its reported opposed role on Nlrc4 at steady state and upon overexpression.
4.4. AIM2
AIM2 is a receptor of cytosolic DNA [59,60]. Since, in healthy cells, DNA is restricted to the nucleus, the presence of cytosolic DNA is either indicative of an infection or of a cellular stress associated with the loss of nuclear membrane integrity [61].
AIM2 is an IFN-inducible gene. In murine BMDMs, Aim2 is expressed at steady state and its expression is slightly induced by IFN, poly(dA:dT) treatment, or infections [60]. In mice, AIM2 levels are sufficient to promote its inflammasome functions, and the AIM2 inflammasome does not require a priming step [62,63]. Its role in myeloid human cells is less clear than in the murine context. Indeed, a cGAS-STING-lysosomal-NLRP3 pathway triggers cell death in response to cytosolic DNA in human myeloid cells [64], while in the presence of IFN-γ, AIM2 is involved in T. gondii responses. These results suggest that AIM2 functionality in human cells is variable depending on the context [65]. In contrast to murine macrophages, AIM2 levels are very low at steady state in human macrophages but are strongly induced by LPS [66] or IFN-γ [65]. The upregulation of AIM2 transcript in inflammatory conditions is likely relevant in human diseases since AIM2 expression level is increased in keratinocytes from psoriatic lesions compared to healthy skin [67].
In human cells, AIM2 expression is dynamically controlled by B lymphocyte-induced maturation protein-1 (BLIMP1 also known as PR domain zinc finger protein 1-PRDM1), IRF1/2 and Signal Transducer and Activator of Transcription 1 (STAT1), a transcription factor activated by interferons [68,69]. BLIMP1 binding site overlaps with IRF1/2 binding site and these transcription factors compete to repress or activate AIM2 expression, respectively. Indeed, BLIMP1 knock-down increases IRF1/2 binding on the AIM2 promoter and AIM2 expression (Figure 2D) [68]. To our knowledge, it is unknown whether BLIMP1 modulation may be involved in licensing AIM2 function in certain human cell types. In addition, two tandem Gamma-activated site (GAS) sequences are present 220 nt upstream of AIM2 TSS. These GAS belong to the long terminal repeat (LTR) sequence of an endogenous retrovirus termed MER41. Treatment with IFN-γ triggers an increase in H3K27ac and STAT1 binding at the MER41.AIM2 site. Accordingly, MER41 sequence is required for AIM2 upregulation in HeLa cells exposed to IFN-γ. The conservation of the MER41.AIM2 site across anthropoid primates (but not in mice) suggests that this retrovirus sequence was co-opted for AIM2 regulation in an ancestor of anthropoid primates [69] and illustrates how past infections have shaped the species-specific regulation of inflammasome genes. Interestingly, a MER41 sequence displaying IRF1 and STAT1 binding in monocytes is also found in proximity of GSDMD, suggesting that the regulation of several inflammasome genes may result from retrovirus integration.
4.5. Pyrin
Pyrin is a sensor detecting Rho GTPase inhibition. Rho GTPases are altered by numerous bacterial toxins and bacterial effectors. Pyrin thus acts as a guard of this important cytoskeleton regulator [70]. Gain of function mutations in MEFV, the gene encoding Pyrin, cause several inflammatory syndromes, including Familial Mediterranean Fever (FMF) [71].
Pyrin is constitutively expressed in human neutrophils, monocytes, and M-CSF-differentiated monocyte-derived macrophages [72]. In monocytes, MEFV expression is positively regulated by IFN-α, IFN-γ, and TNF, while IL-4 and IL-10 repress it [73,74]. TNF-dependent upregulation of MEFV expression is due to the binding of CCAAT/enhancer binding protein β (C/EBPβ) in the MEFV promoter, which acts in synergy with NF-κB p65/RelA subunit. Indeed, NF-κB p65/RelA binds the MEFV promoter both through a canonical NF-κB binding site and indirectly through its binding to C/EBPβ (Figure 2E) [75]. The mechanism underlying IFN-mediated upregulation is unclear although IFN-stimulated response element (ISRE) and GAS consensus sequences have been identified in silico in the MEFV promoter [73].
In mice, Mefv expression is induced by TLR ligands and inflammatory cytokines (IFN-β + TNF). TNF signaling contribution to inflammation in a murine model of FMF strongly suggests that transcriptional regulation of inflammasome sensors is important not only for the detection of pathogens but also for the prevention of deleterious inflammatory reactions [76].
5. Transcriptional Regulation of ASC
The transcriptional regulation of ASC and its gene PYCARD was evidenced upon its discovery. Indeed, PYCARD was identified as a gene whose expression was lost in human breast cancer cells due to methylation of a CpG island in its promoter (in normal cells, CpG islands, in contrast to sparsely distributed CpG dinucleotides, remain unmethylated). Similarly, AIM2 was identified as a gene whose expression was lost in a melanoma cell line [77]. At the time, PYCARD was named “TMS1,” for “target of methylation-induced silencing 1” [78]. Overexpression of DNMT1 (DNA (cytosine-5)-methyltransferase) leads to aberrant hypermethylation of CpG islands in the 5′ untranslated region of PYCARD resulting in PYCARD silencing and recapitulating the transcriptional shutdown observed in numerous primary breast tumors. The pro- and antitumoral roles of inflammasomes are well established. Transcriptional silencing of inflammasome genes could thus promote tumor escape from immune surveillance.
Although PYCARD expression in nontumor cells is largely constitutive and insensitive to pro-inflammatory signals, invalidation of ifi205, a murine-specific gene from the IFN-inducible PYHIN (Pyrin and HIN domain) family leads to the loss of Pycard expression [79]. In addition, NF-κB p65/RelA and C/EBPβ have putative binding sites in the Pycard promoter, and ectopic expression of these transcription factors in a HEK293-reporter system increases Pycard promoter activity. Interestingly, IFI205 interacts with C/EBPβ and synergizes with p65/RelA to promote Pycard promoter activity [79]. IFI205 has no direct orthologues in humans [80]. Whether other members of the PYHIN/AIM2-like Receptor (ALR) family could regulate PYCARD expression in human cells remains unknown.
6. Transcriptional Regulation of Inflammatory Caspases
6.1. Caspase-1
Caspase-1 is the effector caspase of canonical inflammasomes. Its expression is largely constitutive but a loss of balance in CASP1 expression may be relevant in pathological situations. For instance, CASP1 expression is upregulated in the brain of multiple sclerosis (MS) patients compared to controls and may participate in MS lesions [81]. Conversely, CASP1 is downregulated in PBMCs from septic patients, a feature that could contribute to the severe immunodepression observed in these patients [82].
CASP1 promoter possesses an ISRE immediately upstream of its TSS participating in the control of CASP1 expression at steady state and in the presence of IFN. Indeed, IRF2 binds CASP1 promoter in primary human monocytes, and IRF2-deficient U937 monocytes present a strong decrease in CASP1 levels at steady state (Figure 2A) [22,83]. IFN-γ, which strongly induces IRF1, upregulates CASP1 expression [84]. In mice, a similar ISRE site is found in the promoter of Casp1, and Irf1−/− splenocytes are deficient in Casp1 induction following concanavalin-A stimulation [85]. In contrast, Irf2−/− mice and murine macrophages do not present a major defect in Casp1 level [23,86] suggesting differences in the basal regulation of CASP1 in mice and humans.
Another member of the IRFs family, IRF8, also controls CASP1 expression as exemplified by its role during Epstein–Barr virus (EBV) infection [87]. Invalidation of IRF8 in several EBV+ lymphoblastoid cell lines fully abolishes CASP1 expression. IRF8 binds the consensus ISRE sequence in the proximal promoter of CASP1, and its ability to transactivate CASP1 expression was validated using a luciferase reporter/minimal promoter assay. In this assay, IRF8 synergizes with IRF1 to control CASP1 promoter activity suggesting that IRF8 binds DNA in a ternary complex with IRF1 [56]. Interestingly, this IRF8/CASP1 cascade facilitates EBV lytic replication, likely through the cleavage of host factors involved in the maintenance of EBV latency (Figure 3A) [87].
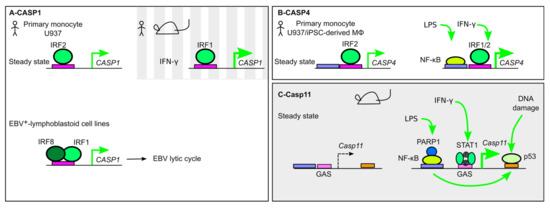
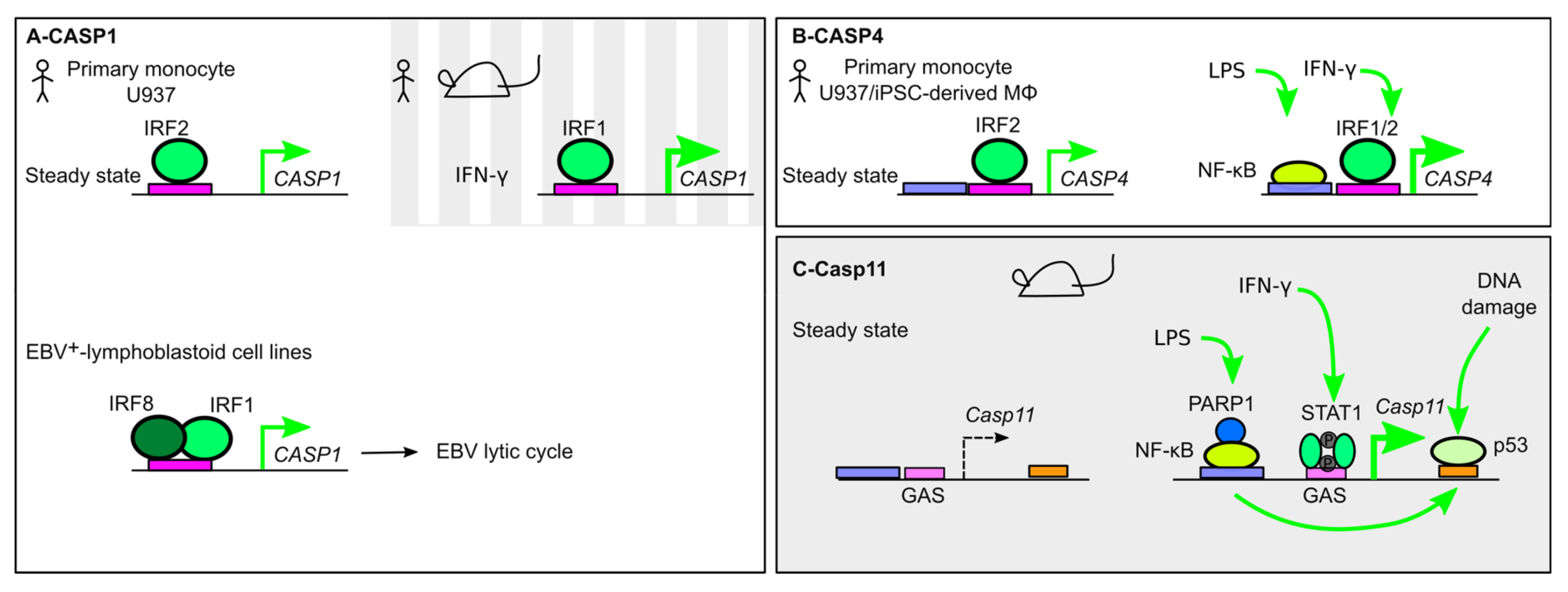
Figure 3.
Transcriptional regulation of inflammatory caspases. (A) In human monocytes, CASP1 expression is regulated at steady state by IRF2, while in the presence of IFN-γ, IRF1 regulates its expression. In Epstein–Barr virus (EBV+) lymphoblastoid cell lines, a ternary IRF1/8/DNA complex controls CASP1 expression and EBV lytic cycle. (B) CASP4 is constitutively expressed in human monocytes in an IRF2-dependent manner. IFN-γ and lipopolysaccharide (LPS) can upregulate CASP4 expression by promoting NF-κB and IRF1 binding to the CASP4 promoter. (C) Casp11 expression is undetectable at steady state but demonstrates a strong upregulation in the presence of LPS or IFN-γ, which promotes the recruitment of NF-κB and STAT1. Poly [ADP-ribose] polymerase 1 (PARP1) acts as a positive cofactor of NF-κB. DNA-damage triggers p53 binding in the Casp11 first intron and induction of its expression. p53 binding is dependent on NF-κB.
6.2. Caspase-4
Caspase-4 is an inflammatory protease, which binds intracellular LPS. Caspase-4 cleaves GSDMD, directly triggering pyroptosis and indirectly activating the canonical NLRP3 inflammasome.
IRF2 is a key regulator of CASP4 as identified through a genome-wide CRISPR-Cas9 screen performed in the premonocytic U937 cells [22]. IRF2−/− U937 are resistant to cytosolic LPS-induced pyroptosis and ectopic expression of CASP4 in IRF2−/− cells fully restores, even accelerates, pyroptosis in response to cytosolic LPS delivery. The regulatory role of IRF2 on CASP4 levels was confirmed in induced pluripotent stem cells (iPSCs)-derived macrophages at both mRNA and protein levels. IRF-2 binds CASP4 promoter in primary human monocytes. Interestingly, in the presence of IFN-γ, IRF1 can complement the absence of IRF2 to regulate CASP4 levels (Figure 3B). This regulation of CASP4 by IRF2 is not observed at the protein level in the cell line EA.hy926 (an hybrid of A549 and human umbilical vein endothelial cells), suggesting that the CASP1 expression dependency on IRFs might differ between different human cell types [23,88].
In addition to IRF1/2, the NF-κB subunit RelA upregulates CASP4 expression via its binding on a NF-κB binding site located ≈1000 bp upstream of CASP4 TSS [89].
6.3. Caspase-11
Caspase-11 is the murine homolog of human Caspase-4/5. Like caspase-4, caspase-11 directly binds intracytosolic LPS and cleaves GSDMD to trigger pyroptosis.
Naive mice present undetectable levels of Casp11 expression in tissues. Casp11 expression is induced in response to pro-inflammatory stimuli (e.g., LPS) or IFN [90,91]. Schauvliege et al. validated the presence of a NF-κB binding site in the Casp11 promoter required for Casp11 induction in response to LPS. cRel and likely other NF-κB subunits bind this site upon LPS stimulation. In the presence of IFN-γ, STAT1 binds a GAS motif in the Casp11 promoter. Mutation of this motif invalidates both IFN-γ- and LPS-induced Casp11 expression. Yet, since a putative NF-κB binding site overlaps with the GAS site, it is unclear whether STAT1 is required for both LPS- and IFN-γ-mediated Casp11 induction (Figure 3C) [90].
C/EBP homologous protein (CHOP) is a TF from the C/EBP family, whose expression is induced by ER stress. CHOP is required for Casp11 induction both in the lung of LPS-treated mice and in vitro in LPS-treated peritoneal macrophages. ER stress inducers (e.g., Thapsigargin) induce Casp11 expression in a CHOP-dependent manner in the absence of LPS [92]. Intratracheal LPS delivery induces ER stress in the lung and Chop−/− mice display attenuated LPS-induced lung inflammation. Although it is unclear whether CHOP acts directly on the Casp11 promoter, CHOP may connect ER stress and the noncanonical inflammasome as described above for ATF4 and NLRP1.
Poly(ADP-ribose) polymerase 1 (PARP-1) is one of the most abundant nuclear protein. PARP-1 is a multifunctional enzyme acting at numerous levels to regulate transcription [93]. Interestingly, Yoo et al. identified that PARP-1 is necessary for LPS-induced, but not IFNγ-induced, Casp11 expression. PARP-1 interacts with cRel and RelA and regulates NF-κB-dependent genes independently of its enzymatic activity [94]. PARP-1, as a NF-κB-co-activator, may thus regulate specifically NF-κB-mediated Casp11 induction [95].
Interestingly, Casp11 expression can also be induced in a p53-dependent manner following DNA damage [96]. Two p53 binding sites are present in the first intron of Casp11 and p53 binding on Casp11 promoter is observed by ChipSeq following gamma irradiation of thymocytes or etoposide treatment of mouse embryonic fibroblasts. NF-κB is required for p53 binding to Casp11 promoter [96]. CASP1 and NLRC4 promoters display a similar p53 binding site and a similar upregulation in response to DNA-damaging agents (e.g., etoposide) [97,98], suggesting that p53 (or p53 family members) and NF-κB function cooperatively to ensure an appropriate inflammasome response to sterile and infectious stresses.
The importance of Casp11 induction was demonstrated in murine models of septic shock. In contrast to humans, who express CASP4 constitutively, mice are highly resistant to LPS injection (lethal dose about 40 mg/kg). Yet, if mice are primed with a low dose of LPS (400 μg/kg) or with poly(I:C), a potent IFN inducer, they become highly sensitive to very low doses of LPS (10–100 ng/kg) [99,100]. Similarly, a transgenic mouse expressing human CASP4 under its native promoter displays a constitutive CASP4 expression in the spleen and in the intestine and was highly susceptible to LPS injection [101]. These experiments thus suggest that induction of Casp11 decreases the threshold of LPS detection. Although this induction is detrimental in a septic shock model, it contributes to the immune defenses against Gram-negative bacteria [24].
6.4. Caspase-5
Caspase-5 is an inflammatory caspase directly binding cytosolic LPS and triggering pyroptosis [10].
CASP5 is expressed at low levels in various organs with a more predominant expression in the spleen and in the colon [102]. Its expression is highly induced in vitro following LPS treatment or IFN treatment [102,103,104]. Induction of CASP5 is delayed in comparison to IL1B induction and is sensitive to the protein synthesis inhibitor cycloheximide, indicating that it is a secondary-response gene. Three putative binding sites for NF-κB are present in the CASP5 promoter but their functionality remains to be experimentally tested [105]. The relevance of CASP5 induction in diseases is still unclear, although a strong increase in CASP5 level (20-fold increase) is observed in lesional psoriatic skin [105].
7. Transcriptional Regulation of Downstream Targets
7.1. GSDMD
GSDMD is the pyroptosis executioner [12,15]. N-terminal fragments generated following inflammatory caspases cleavage oligomerize in the plasma membrane to form a transmembrane pore.
As a core component of inflammasomes, GSDMD is constitutively expressed in numerous tissues. Irf2 was identified in a forward genetic screen with a chemical mutagen, N-ethyl-N-nitrosourea (ENU), as a gene required for IL-1β release in response to Pam3CSK4 and ATP stimulation [23]. Similarly to the regulation of CASP4 described above, IRF2 regulates the expression of GSDMD at steady state [23]. IRF2 binds an ISRE sequence in the GSDMD promoter immediately upstream of the TSS. Irf2−/− mice do not express Gsdmd, and Irf2−/− BMDMs display a profound inflammasome defect. In human cells, IRF2 knockout in EA.hy926 cells confirmed the key role of IRF2 in controlling GSDMD expression [23]. We did not observe a major impact in the monocytic U937 cell line in the absence of IRF2 [22], suggesting that IRF2 regulates GSDMD in a cell-type-specific manner [88]. IRF1 and IRF2 have the same binding specificity for the ISRE sequence. Accordingly, and as described above for CASP4 regulation, in the absence of IRF2, IRF1 plays a compensatory role and partially controls GSDMD expression (Figure 4A) [23].
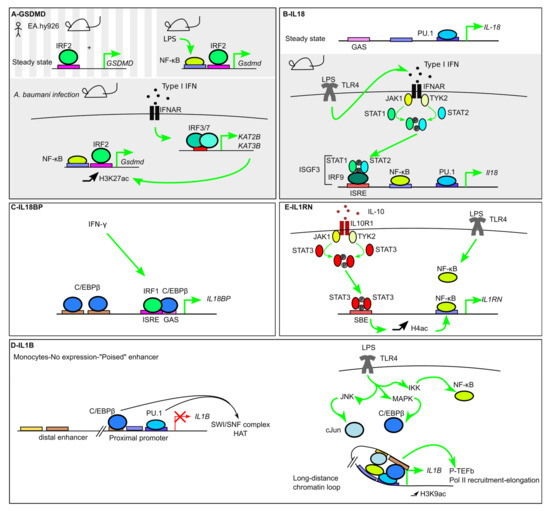
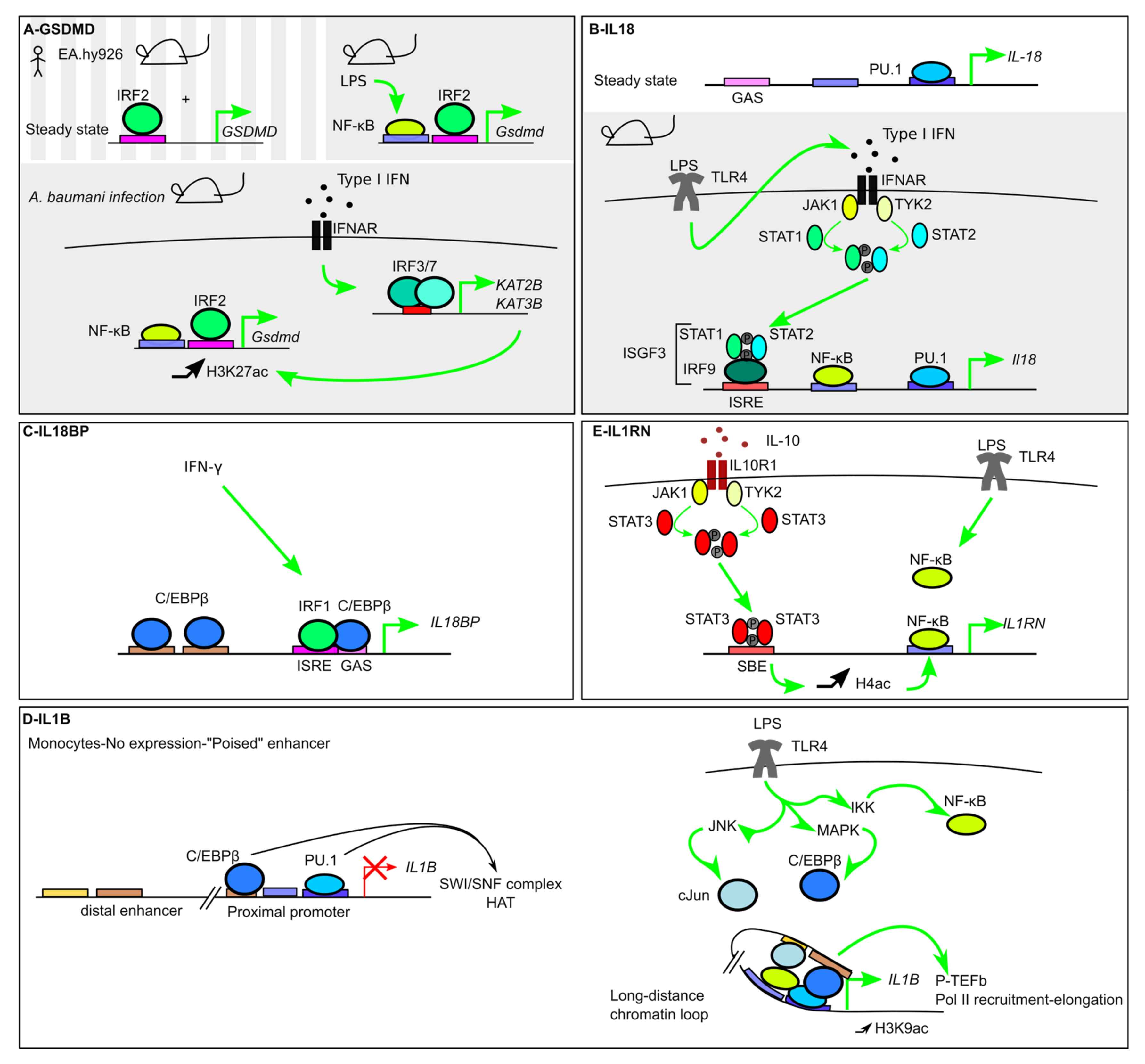
Figure 4.
Transcriptional regulation of inflammasome substrates. (A) In mice and in the human endothelial EA.hy926 cell line, GSDMD constitutive expression is controlled by IRF2. During infection, NF-κB increases Gsdmd expression. Furthermore, induction of the histone acetyl transferases KAT2B and KAT3B, in an Interferon-α/β Receptor-(IFNAR)/IRF3/7-dependent manner, correlates with an increase in H3K27ac epigenetic mark at the Gsdmd promoter and an increase in its transcription level. (B) IL18 is expressed at steady state and its promoter demonstrates PU.1 binding. LPS treatment leads to activation of STAT1 and STAT2 downstream of IFNAR, to the formation of the IFN-stimulated gene factor 3 (ISGF3) complex, containing IRF9, Phospho-STAT1, and Phospho-STAT2, and its binding to the Il18 promoter. (C) IL18BP is induced by IFN-γ. IRF1 binds C/EBPβ to promote its recruitment to a Gamma-activated site (GAS) site. (D) IL1b is not expressed at steady state although PU.1 and C/EBPβ cooperate in monocytes to recruit the Switch/sucrose nonfermenting (SWI/SNF) complex and Histone acetyl transferase (HAT) to render the enhancer in a “poised” state. LPS treatment triggers C/EBPβ and c-Jun recruitment at the distal enhancer and the formation of a long-distance loop allowing the TFs to cooperate to increase H3K9ac, recruit positive transcription elongation factor b (p-TEFb) and activate Pol II. (E) IL-10 induces IL1RN expression in a STAT3-dependent manner. STAT3 binding to STAT-Binding Element (SBE) increases Histone H4 acetylation and promotes NF-κB binding.
GSDMD expression is largely constitutive. Yet, infections or pro-inflammatory signals increase GSDMD expression. Indeed, Acinetobacter baumannii infection induces a 2–4-fold increase in Gsdmd transcript in the liver and in BMDMs in an IFNAR1- and IRF3/7-dependent manner [106]. This induction correlates with the type I IFN-mediated induction of two histone lysine (K)-acetyltransferases, KAT2B and KAT3B/P300, and with an increase in the epigenetic histone modification H3K27ac at the Gsdmd promoter [106]. In addition, NF-κB activation following LPS stimulation promotes a 2-fold induction of Gsdmd transcript in murine adipocytes. Two NF-κB binding sites are present in the proximal Gsdmd promoter, and RelA/p65 binding contributes to Gsdmd induction [107].
Melatonin, a hormone maintaining the circadian rhythm, blunts LPS-mediated NF-κB activation and downregulates the expression of several inflammasome genes including Gsdmd in vitro and in vivo [107]. Together with the role of NR1D1 in the regulation of Nlrp3 developed above, this regulation emphasizes the role of circadian rhythm in inflammasome transcriptional regulations.
7.2. IL-18 and IL-18BP
IL18 expression is largely constitutive in humans and in mice [108]. Il18 promoter includes a PU-box (a purine-rich sequence binding PU.1), NF-κB-recognition sequences, an ISRE site that was originally described to promote IRF-8-dependent induction of IL18 in macrophages [109] and GAS elements [110].
Accordingly, IL18 expression in human monocytes is inducible by LPS stimulation in a JAK/STAT-dependent manner. IL18 induction requires both NF-κB activation and type I IFN signaling, resulting in delayed IL18 induction kinetics compared to the kinetics of IL1b induction (an immediate early gene) [110]. Similarly, type I IFN signaling is critical for Il18 induction in murine BMDMs treated with LPS. STAT1 and IRF9 are specifically required for this activity. The involvement of STAT1 and IRF9 suggests that IFN-stimulated gene factor 3 (ISGF3), a ternary protein complex made of IRF9, STAT1, and STAT2 and functioning as a transcription factor downstream of type I IFN receptors, is a direct regulator of Il18 expression (Figure 4B) [111].
In murine epithelial cells, Il18 homeostatic expression is controlled by IL-22. IL-22 is a cytokine from the IL-10 family produced by T cells and innate lymphoid cells (ILC) that induces antimicrobial and tissue-protective responses in epithelia. Interestingly, IL-22 regulates homeostatic expression of Il18 in an organ-specific manner since Il22- and IL22R1-deficient mice display only a deficient Il18 mRNA expression in the ileum but not in the colon, spleen, and lung. IL-22 also increases Il18 expression during T. gondii infection [112]. Finally, Il18 expression in the colon and the liver is impacted by the microbiota. Indeed, intestinal dysbiosis (i.e., gut microbiome imbalance) triggers upregulation of Il18 in the liver. Conversely, germ-free mice have a down-regulation of Il18 mRNA levels in the colon [113].
IL-18 bioavailability is regulated by IL-18BP (IL-18 binding protein), a decoy receptor, whose expression is strongly inducible by IFN-γ. IL-18BP binds IL-18 limiting the free IL-18 able to bind IL-18R to trigger signaling. IL18BP induction corresponds to a negative feedback loop since IL-18 is a major activator of IFN-γ production. IL18BP promoter includes a GAS, an ISRE and two C/EBPβ sites [114]. IRF-1 was identified as the main IRF responsible for IL18BP induction downstream of IFN-γ signaling [114]. IRF-1 forms a complex with C/EBPβ and directs its binding to a GAS site (Figure 4C). Interestingly, STAT1 is directly involved in IL18BP regulation in specific cell types [115]. IL18BP induction in response to IFN-γ is higher in epithelial cells than in monocytes. This differential induction is linked to differential methylation of a CpG island in the IL18BP promoter resulting in variable histone H3K9 acetylation and Pol II recruitment. Epigenetic modifications thus control cell-specific IL18BP production, likely antagonizing the IL-18/IFN-γ axis with different kinetics at systemic and epithelial barrier sites [115].
7.3. IL-1b and IL-1RA
IL18 and IL1b display different regulations. Indeed, IL1b is not expressed at steady state but is strongly inducible following microbial (e.g., LPS) or sterile (e.g., TNF) pro-inflammatory signals.
IL1b transcription is regulated by three main regions: a proximal promoter, a distal one, and an enhancer site located ≈3 kb upstream of the TSS. Lineage-specific TFs control the cell-specific IL1b transcription and cooperate with signal-dependent TFs to trigger fast responses to pro-inflammatory signals. IL1b expression is most prominent in monocytes/macrophages, we will thus first present the regulation in these cells before presenting the atypical regulation that can occur in other cell types.
7.3.1. Roles of the Lineage-Specific/Pioneer TFs PU.1 and C/EBPβ
IL1B is primarily expressed in monocytes and macrophages [116]. The binding of the hematopoietic-specific transcription factor PU.1 on several PU-boxes in the proximal IL1B promoter and in the enhancer is largely responsible for the cell-specific expression of IL1B. Indeed, ectopic expression of PU.1 in HeLa cells is sufficient to drive IL1B expression. PU.1 is a pioneer TF, which recruits the SWI/SNF family chromatin remodeling complex [117]. Similarly, C/EBPβ initiates chromatin opening through its ability to bind the histone acetyl transferase p300/CBP and the SWI/SNF complex (Figure 4D) [118]. Like PU.1, C/EBPβ is predominantly expressed in monocytes/macrophages and its binding in the proximal promoter is required for IL1B expression [116]. PU.1 and C/EBPβ act in collaboration to generate a highly accessible IL1B promoter (without nucleosome blocking the access of Pol II), yet without Pol II binding at steady state [119]. This “poised” chromatin structure is likely responsible for the ability of IL1B promoter to be rapidly turned-on following pro-inflammatory signals [120]. c-Jun also binds constitutively on AP-1 (activator protein 1) sites in the proximal enhancer and likely acts in concert with PU.1 and C/EBPβ to remodel chromatin at this site [120]. IL1B is an immediate early gene meaning that its expression is upregulated immediately post pro-inflammatory signals independent of protein neosynthesis.
7.3.2. Signal-Dependent TFs (NF-κB)
These pioneer TFs act in synergy with multiple other TFs that are either bound constitutively or in a signal-dependent manner to the enhancer, the promoter, or the intronic sequences of IL1b. In addition to PU.1, C/EBPβ, and cJun, NF-κB, AP-1 (cJun/cFos heterodimer), STAT proteins, and IRF4/8 bind IL1B regulatory regions [21,120,121,122]. IRF4 and IRF8 participate in the dynamic transcription of IL1B following LPS treatment in human monocytes [120]. The relevance of these two TFs remains unclear since Irf8-deficient macrophages do not present any obvious Il1b defect [25] and since ectopic expression of Irf4 or Irf8 in murine macrophages inhibits Il1b expression [21].
The role of NF-κB is clearly established following treatments with microbial (e.g., LPS) or sterile pro-inflammatory stimuli (e.g., IL-1β in a feedforward loop). Il1b induction downstream of TLRs is largely mediated by MYD88, although a moderate TRIF-dependent induction is observed following TLR3 engagement by poly(I:C) [111]. p65/RelA is recruited to the IL1B promoter in a signal-dependent manner, while c-Jun, which expression is induced by MAPK downstream of TLRs, is both constitutively bound to the proximal IL1B enhancer and inducibly recruited to both the distal enhancer and the promoter [120].
The inducible expression of IL1B in response to pro-inflammatory signals is linked to an enhancer region located -3757 and -2729 bp upstream of the TSS. The induction is dependent on the binding of C/EBPβ at this enhancer site [123] and on a long-range chromatin looping that allows C/EBPβ to directly interact with PU.1, which is bound to the proximal promoter [124]. In addition, binding of NF-κB subunits (RelA/p65, NF-κB1 (p50), c-Rel (p85)) in the proximal promoter region contributes to IL1B induction [125]. NF-κB interacts with PU.1 [124]. LPS-induced IL1B transcription thus results from the intimate cooperation of the signal-dependent TF NF-κB with the pioneer TFs C/EBPβ and PU.1. C/EBPβ recruits the positive transcriptional elongation factor P-TEFb at the IL1B promoter. P-TEFb phosphorylates Pol II C-terminal tail and promotes transcription elongation [124].
IL1B promoter also includes a composite cAMP response element (CRE)/C/EBPβ site [126] located between -2755 and -2762. The CRE binds CREB and the related TF ATF-1 following LPS addition [127]. Increase in intracellular cAMP results in the activation of PKA and in the subsequent phosphorylation of CREB, which interacts with C/EBPβ [128] to induce IL1B expression. In agreement, Prostaglandin E2 (PGE2), by signaling through the Prostaglandin E2 receptors 2 and 4 (EP2 and EP4), induces an increase in intracellular cAMP and triggers IL1B induction [129,130]. Since IL-1β induces PGE2 production through the transcriptional control of COX2 (the inducible enzyme producing PGE2). This regulation illustrates another feedforward loop that could be key in mounting potent inflammatory responses [130].
7.3.3. Metabolic Regulation of IL1b
In addition to the direct NF-κB-mediated effect, LPS stimulation triggers IL1b induction through metabolic rewiring. Indeed, LPS treatment is associated with a macrophage metabolism shift from mitochondrial oxidative phosphorylation to aerobic glycolysis. This shift is associated with a strong increase in the intracellular concentration of succinate, an intermediate metabolite of the tricarboxylic acid (TCA) cycle. Succinate stabilizes hypoxia-inducible factor 1α (HIF1α) protein. Following LPS stimulation, HIF1α binds to the hypoxia responsive element (HRE) located ≈300 pb upstream of the Il1b TSS. Intracellular succinate thus acts as an endogenous danger signal connecting metabolic rewiring following chronic LPS stimulation and Il1b transcription (together with other inflammatory genes) [131]. HIF1α appears as a hub integrating numerous signals to induce Il1b. Indeed, in addition to sensing intracellular succinate concentration, HIF1α function requires its upregulation by NF-κB or by LXRα in human macrophages. LXR are nuclear receptors sensing cholesterol derivatives and activated in atherosclerotic plaques. Addition of an LXR agonist increases LXR binding at the HRE sites located in the IL1B and the HIF1α promoters, suggesting that an LXR/HIF1α complex binds at these sites [132]. In addition, hypoxia also increases LPS-induced IL-1β release, in agreement with IL1B being a direct target of HIF1α.
NRF2 (NF-E2-related factor-2) is a transcription factor activated by oxidative stress. At steady state, NRF2 is degraded due to its binding to KEAP1, a E3 ubiquitin ligase adaptor. During oxidative stress, KEAP1 dissociates from NRF2 allowing NRF2 to accumulate and translocate into the nucleus. In the nucleus, NRF2 predominantly binds ARE (antioxidant responsive element) to positively regulate transcription. NRF2 binds DNA upstream of Il1b TSS, in the enhancer region. Surprisingly, NRF2 binding is independent of ARE sites, suggesting that NRF2 might be recruited via interaction with another TF. Furthermore, NRF2 activation correlates with a decrease in Pol II recruitment at the Il1b TSS [133]. NRF2 may thus have an atypical activity (ARE-independent repression) at the Il1b promoter to decrease inflammation in conditions of high oxidative stress.
7.3.4. Negative Regulation of IL1b
Il1b expression is negatively regulated by IL-10 in a STAT3-dependent manner. Type I IFNs, which induce IL-10, also negatively regulate IL1b transcription in murine BMDMs and in human primary monocytes [26].
IL-19, -20, and -24 are cytokines belonging to the IL-10 family, produced by myeloid and epithelial cells and signaling through the type I IL-20 receptor (IL-20R). These cytokines negatively regulate Il1b transcription in murine keratinocytes exposed to Staphylococcus aureus. IL20R signaling leads to a decrease in STAT3 and in C/EBPβ active isoforms. Meanwhile, C/EBPβ displays increasing inhibitory sumoylation. The shift in the balance between C/EBPβ active and inactive isoforms correlates with a decrease in C/EBPβ binding at the Il1b promoter [134].
7.3.5. IL1b Regulation in T Cells
Several studies [135,136,137] have demonstrated a production of IL-1β in CD4+ T cells using elegant cell-type-specific gene ablation and bone-marrow transfer experiments [136,137]. IL-1β produced by T cells contributes to chronic inflammatory diseases such as experimental autoimmune encephalomyelitis [136]. IL1B expression in CD4+ T cells (and particularly the CCR5+ subset) is upregulated following TCR engagement with different costimulation signals. CD4+ T cells do not present detectable amounts of PU.1 protein [138] even after TCR engagement. Accordingly, IL1B promoter in resting T cells is not accessible and does not present the poised architecture observed in monocytes [119]. In agreement with this difference, Il1b expression is much lower in activated T cells than in activated monocytes (≈1000-fold at the transcript level). The mechanism leading to TCR-mediated and PU.1-independent induction of Il1b in CD4+ T cells remains unclear. This mechanism is associated with an increase in the activating epigenetic histone modifications (H3K4me3 and H3K9ac), a corresponding decrease in the inhibiting modification (H3K27me3) at the Il1b promoter site and Pol II enrichment throughout Il1b gene upon T cell activation. In contrast to housekeeping genes transcribed at high levels, inhibitory H3K27me3 marks persist on the Il1b promoter resulting in a bivalent H3K4me3+/H3K27me3+ low-activity promoter [138].
7.3.6. IL1b Regulation in DCs
As discussed above, conventional DCs are largely deficient in inflammasome responses due to IRF4/IRF8-mediated negative regulation of inflammasome genes transcription [21,51]. IL-21 is a cytokine produced by follicular helper (Tfh), Th17, and NK cells that triggers STAT3 activation. In cDCs, but not in BMDMs, IL-21 (or IL-10) increases Il1b transcription in a Stat3-dependent manner. Although Il1b induction (≈3-fold) remains modest compared to Il1b induction induced by NF-κB activation in BMDMs, IL-21 contributes to Il1b expression in vivo during Pneumonia Virus of Mice (PVM) infection, indicating that this production is likely relevant in specific contexts [139]. This IL-21/IL-10-Stat3-dependent increase in Il1b expression illustrates how one cytokine can have a differential impact in two different cell types (cDCs, BMDMs).
Overall, IL1B enhancers and promoters integrate signals from multiple cytokines either directly (e.g., IL-1β autoamplification loop) or indirectly (type I IFN inducing IL-10), from intrinsic (e.g., circadian clock), environmental (xenobiotics), metabolic (e.g., succinate) or infectious (LPS) signals. IL1B regulation has been mostly studied in monocytes/macrophages, in which IL1B is an immediate gene strongly upregulated following pro-inflammatory signals detection. Yet, there is now clear evidence that low expression of IL1b in DCs or CD4+ T cells, although controlled by different transcriptional mechanisms, is relevant in chronic inflammatory diseases [140].
7.3.7. IL1RN
IL1RN encodes IL-1 receptor antagonist, a secreted cytokine that competes with IL-1α and IL-1β for their binding to the IL-1 receptor. In contrast to IL1α/β, IL1Ra does not trigger IL-1 receptor signaling. The expression level of IL1RN thus balances IL-1 bioactivity [141].
IL1RN expression is inducible by LPS [142], IL-1α/β (acting as a negative feedback loop) [143] and IFN treatment [144]. Like IL1B, IL1RN expression is controlled by C/EBPα/β, PU.1, and NF-κB that bind to the IL1RN promoter [143]. IL-10 also induces IL1RN expression in a STAT3-dependent manner to dampen inflammation. IL-10 triggers recruitment of STAT3 to the IL1RN promoter and synergizes with LPS to promote the recruitment of NF-κB p50/p65 and Pol II. The increased NF-κB recruitment following IL-10 addition correlates with histone H4 acetylation suggesting that STAT3 recruitment at the IL1RN promoter modifies chromatin structure and accessibility to NF-κB binding sites [145] (Figure 4E).
8. Conclusions
While there is no doubt that post-transcriptional regulations of inflammasomes are intimately linked to their activation, these regulations take place in a transcriptional landscape (Table 1) that shapes the output responses (Figure 5). At steady state, the transcriptional regulation of inflammasome genes is the result of a complex cascade of events taking place during lineage differentiation in a species-specific manner. As observed in cDCs, these regulations have a profound impact on the biological responses by limiting pyroptosis and favoring T cell priming and adaptive immune responses [21]. Furthermore, in inflammatory conditions, the upregulation of inflammasome genes (as exemplified by Casp11 and the sensitivity to LPS [99]) strongly modifies the ability of the host to detect and respond to PAMPs. Importantly, signal-induced transcriptional regulations can be partly fixed in time due to epigenetic reprogramming/immunological imprinting. These long-term changes are responsible for the trained immunity (also called innate immune memory). They affect inflammasome genes [146] and contribute to the protective effect of trained immunity against pathogens [147] or possibly to chronic inflammation [148].

Table 1.
Main Transcription Factors (TFs) Regulating Inflammasome Genes.
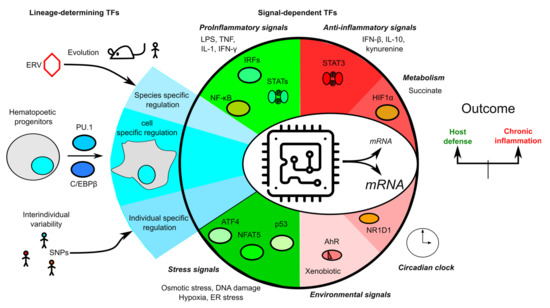
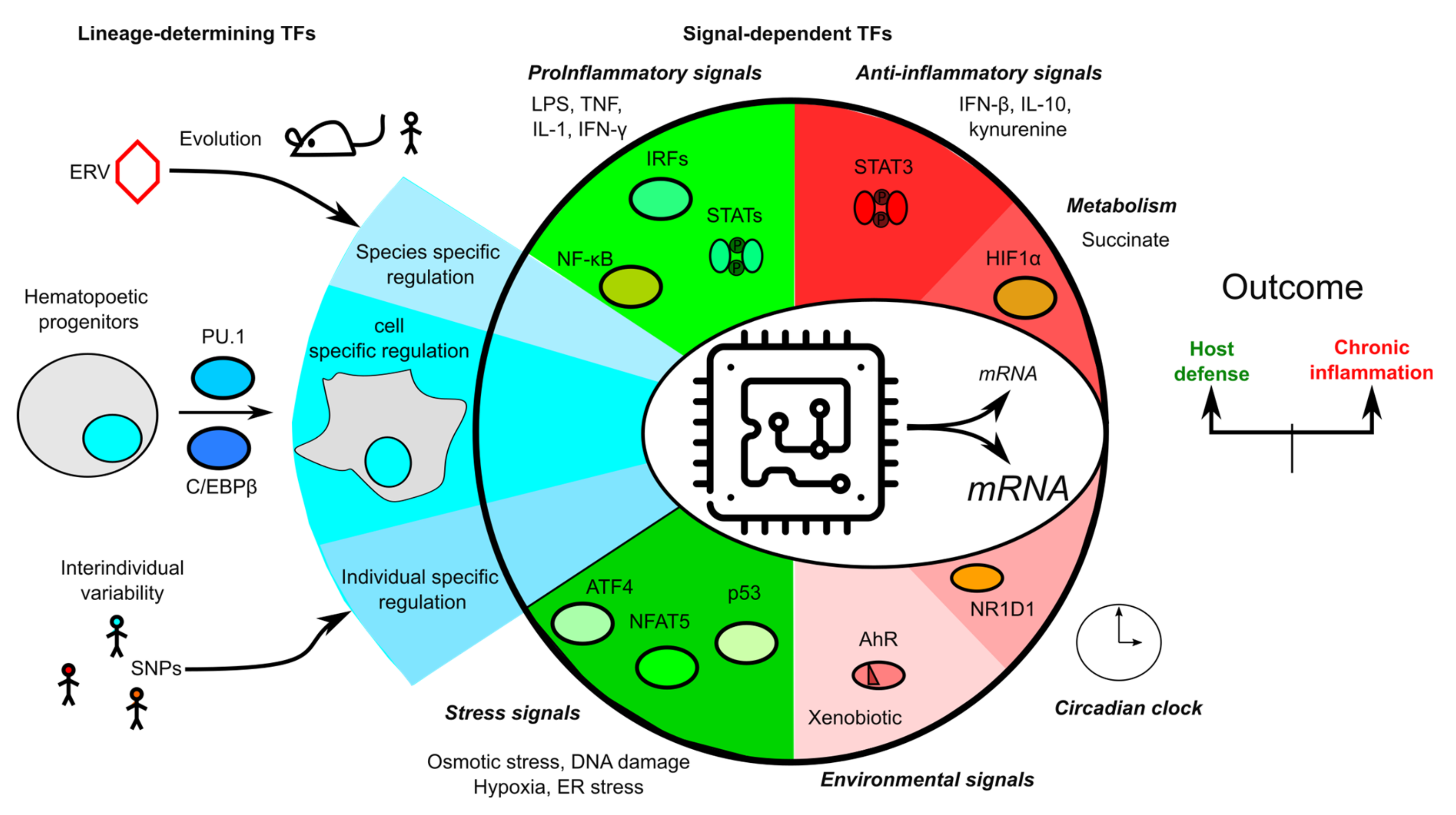
Figure 5.
Overview of the transcriptional regulation of inflammasome genes. Transcription is schematized as a microchip integrating multiple signals from evolution (e.g., endogenous retroviruses), lineage engagement and differentiation, individual single nucleotide polymorphisms (SNPs), the environment, metabolism, stress responses, and pro- and anti-inflammatory immune signaling pathways to control inflammasome transcripts levels.
Although specific inflammasome genes have specific regulations, there are also common regulations that globally affect the inflammasome responses either positively (e.g., NF-κB activation) or negatively (IRF4/8 in cDCs). These specific inflammasome gene regulations likely shape the biological inflammasome response (balance between IL-1β and IL-18, balance between cytokine release and pyroptosis, and balance between apoptosis and pyroptosis). Similarly, the clinical phenotypes resulting from gain of function mutations in inflammasome genes differ based on the mutated sensor. The prevalence of organ-specific phenotypes (e.g., cutaneous phenotypes in NLRP1-associated syndromes [149,150]) or cytokine-driven diseases (IL-18 in NLRC4-associated syndromes [151] compared to IL-1 in NLRP3/Cryopyrin-Associated periodic Syndromes [152]) may be largely due to differential transcriptional regulations. In addition to monogenic diseases, the fine understanding of the networks of DNA motifs, chromatin structure, and transcription factors that regulate inflammasome genes may help us understanding how single nucleotide polymorphisms (SNPs) could impact inflammasome regulation [153] and predispose human beings to various inflammatory diseases. Selective inhibitors of histone modifiers have been developed [154] with an impact on the pro-inflammatory response of human macrophages. Developing specific therapeutic intervention targeting transcription regulation is a fascinating challenge for the field.
Funding
This research was funded by the Agence Nationale de la Recherche, grant number 16-CE15-0011.
Conflicts of Interest
The authors declare no conflict of interest.
Abbreviations
| AhR | Aryl hydrocarbon receptor |
| AIM2 | Absent in melanoma 2 |
| AP-1 | Activator protein-1 |
| ARE | Antioxidant responsive element |
| ASC | Apoptosis-associated speck-like protein containing a card |
| ATF | Activating transcription factor |
| BLIMP1 | B lymphocyte-induced maturation protein-1 |
| BMDM | Bone marrow-derived macrophage |
| BCL6 | B cell lymphoma 6 |
| Brd4 | Bromodomain-containing protein 4 |
| CHOP | C/ebp homologous protein |
| CRE | Camp-response element |
| CREB | Camp response element-binding protein |
| DC | Dendritic cell |
| DSS | Dextran sodium sulfate |
| EBV | Epstein–Barr virus |
| EICE | Ets-irf composite element |
| ENU | N-ethyl-n-nitrosourea |
| ERV | Endogenous retrovirus |
| ETS | Erythroblast transformation specific |
| GAS | Gamma-activated site |
| GFI1 | Growth factor independence 1 |
| GRE1 | Gli-responsive element |
| HAMPs | Homeostasis-altering molecular processes |
| HAT | Histone acetyl transferase |
| HIF1α | Hypoxia-inducible factor 1α |
| HRE | Hypoxia response element |
| IFNAR | Interferon-α/β receptor |
| IRE-1 | Inositol-requiring enzyme 1 |
| IRF | Interferon regulatory factor |
| ISRE | Ifn-stimulated response element |
| LTR | Long terminal repeat |
| NAIP | Neuronal apoptosis inhibitory protein |
| NFAT5 | Nuclear factor of activated t cells 5 |
| NF-κB | Nuclear factor κb |
| NLR | Nucleotide-binding domain and leucine-rich repeat containing |
| NLRP | Nlr family pyrin domain containing |
| Nod2 | Nucleotide-binding oligomerization domain 2 |
| NR1D1 | Nuclear receptor subfamily 1 group d member 1 |
| Nrf2 | Nf-e2-related factor 2 |
| ORE | Osmotic response element |
| PAMP | Pathogen-associated molecular pattern |
| PARP1 | Poly [ADP-ribose] polymerase 1 |
| PERK | Pkr-like er protein kinase |
| P-TEFb SBE | Positive transcription elongation factor b STAT-Binding Element |
| SREBP-1a | Sterol regulatory element binding protein-1a |
| STAT1 | Signal Transducer and Activator of Transcription 1 |
| SWI/SNF | Switch/sucrose nonfermenting |
| TCA | Tricarboxylic acid |
| TF | Transcription factor |
| TLR | Toll-like receptor |
| TNF | Tumor necrosis factor |
| TSS | Transcription start site |
| UPR | Unfolded protein response |
| XRE | Xenobiotic response elements |
References
- Paludan, S.R.; Pradeu, T.; Masters, S.L.; Mogensen, T.H. Constitutive immune mechanisms: Mediators of host defence and immune regulation. Nat. Rev. Immunol. 2020, 1–14. [Google Scholar] [CrossRef]
- Song, N.; Li, T. Regulation of NLRP3 Inflammasome by Phosphorylation. Front. Immunol. 2018, 9, 2305. [Google Scholar] [CrossRef] [PubMed]
- Samir, P.; Kanneganti, T.-D. Hidden Aspects of Valency in Immune System Regulation. Trends Immunol. 2019, 40, 1082–1094. [Google Scholar] [CrossRef] [PubMed]
- Lopez-Castejon, G. Control of the inflammasome by the ubiquitin system. FEBS J. 2019, 287, 11–26. [Google Scholar] [CrossRef] [PubMed]
- Moretti, J.; Blander, J.M. Increasing complexity of NLRP3 inflammasome regulation. J. Leukoc. Biol. 2020. [Google Scholar] [CrossRef] [PubMed]
- Taabazuing, C.Y.; Griswold, A.R.; Bachovchin, D.A. The NLRP1 and CARD8 inflammasomes. Immunol. Rev. 2020, 297, 13–25. [Google Scholar] [CrossRef] [PubMed]
- Cai, L.-J.; Tu, L.; Huang, X.-M.; Huang, J.; Qiu, N.; Xie, G.-H.; Liao, J.-X.; Du, W.; Zhang, Y.-Y.; Tian, J.-Y. LncRNA MALAT1 facilitates inflammasome activation via epigenetic suppression of Nrf2 in Parkinson’s disease. Mol. Brain 2020, 13, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Zhang, P.; Cao, L.; Zhou, R.; Yang, X.; Wu, M. The lncRNA Neat1 promotes activation of inflammasomes in macrophages. Nat. Commun. 2019, 10, 1–17. [Google Scholar] [CrossRef]
- Zhong, F.L.; Robinson, K.; Teo, D.E.T.; Tan, K.-Y.; Lim, C.; Harapas, C.R.; Yu, C.-H.; Xie, W.H.; Sobota, R.M.; Au, V.B.; et al. Human DPP9 represses NLRP1 inflammasome and protects against autoinflammatory diseases via both peptidase activity and FIIND domain binding. J. Biol. Chem. 2018, 293, 18864–18878. [Google Scholar] [CrossRef]
- Shi, J.; Zhao, Y.; Wang, Y.; Gao, W.; Ding, J.; Li, P.; Hu, L.; Shao, F. Inflammatory caspases are innate immune receptors for intracellular LPS. Nat. Cell Biol. 2014, 514, 187–192. [Google Scholar] [CrossRef]
- Bitto, N.J.; Baker, P.J.; Dowling, J.K.; Wray-McCann, G.; De Paoli, A.; Tran, L.S.; Leung, P.L.; Stacey, K.J.; Mansell, A.; Masters, S.L.; et al. Membrane vesicles from Pseudomonas aeruginosa activate the noncanonical inflammasome through caspase-5 in human monocytes. Immunol. Cell Biol. 2018, 96, 1120–1130. [Google Scholar] [CrossRef] [PubMed]
- Shi, J.; Zhao, Y.; Wang, K.; Shi, X.; Wang, Y.; Huang, H.; Zhuang, Y.; Cai, T.; Wang, F.; Shao, F. Cleavage of GSDMD by inflammatory caspases determines pyroptotic cell death. Nat. Cell Biol. 2015, 526, 660–665. [Google Scholar] [CrossRef] [PubMed]
- Ding, J.; Wang, K.; Liu, W.; She, Y.; Sun, Q.; Shi, J.; Sun, H.; Wang, D.-C.; Shao, F. Pore-forming activity and structural autoinhibition of the gasdermin family. Nat. Cell Biol. 2016, 535, 111–116. [Google Scholar] [CrossRef] [PubMed]
- Ruan, J.; Xia, S.; Liu, X.; Lieberman, J.; Wu, H. Cryo-EM structure of the gasdermin A3 membrane pore. Nat. Cell Biol. 2018, 557, 62–67. [Google Scholar] [CrossRef] [PubMed]
- Kayagaki, N.; Stowe, I.B.; Lee, B.L.; O’Rourke, K.; Anderson, K.; Warming, S.; Cuellar, T.L.; Haley, B.; Roose-Girma, M.; Phung, Q.T.; et al. Caspase-11 cleaves gasdermin D for non-canonical inflammasome signalling. Nature 2015, 526, 666–671. [Google Scholar] [CrossRef]
- Rühl, S.; Broz, P. Caspase-11 activates a canonical NLRP3 inflammasome by promoting K+efflux. Eur. J. Immunol. 2015, 45, 2927–2936. [Google Scholar] [CrossRef]
- Schmid-Burgk, J.L.; Gaidt, M.M.; Schmidt, T.; Ebert, T.S.; Bartok, E.; Hornung, V. Caspase-4 mediates non-canonical activation of the NLRP3 inflammasome in human myeloid cells. Eur. J. Immunol. 2015, 45, 2911–2917. [Google Scholar] [CrossRef]
- Broz, P.; Monack, D.M. Molecular mechanisms of inflammasome activation during microbial infections. Immunol. Rev. 2011, 243, 174–190. [Google Scholar] [CrossRef]
- Ising, C.; Venegas, C.; Zhang, S.; Scheiblich, H.; Schmidt, S.V.; Vieira-Saecker, A.; Schwartz, S.; Albasset, S.; McManus, R.M.; Tejera, D.; et al. NLRP3 inflammasome activation drives tau pathology. Nat. Cell Biol. 2019, 575, 669–673. [Google Scholar] [CrossRef]
- Pathak, S.; McDermott, M.F.; Savic, S. Autoinflammatory diseases: Update on classification diagnosis and management. J. Clin. Pathol. 2016, 70, 1–8. [Google Scholar] [CrossRef]
- McDaniel, M.M.; Kottyan, L.C.; Singh, H.; Pasare, C. Suppression of Inflammasome Activation by IRF8 and IRF4 in cDCs Is Critical for T Cell Priming. Cell Rep. 2020, 31, 107604. [Google Scholar] [CrossRef]
- Benaoudia, S.; Martin, A.; Gamez, M.P.; Gay, G.; Lagrange, B.; Cornut, M.; Krasnykov, K.; Claude, J.; Bourgeois, C.F.; Hughes, S.; et al. A genome-wide screen identifies IRF2 as a key regulator of caspase-4 in human cells. EMBO Rep. 2019, 20. [Google Scholar] [CrossRef] [PubMed]
- Kayagaki, N.; Lee, B.L.; Stowe, I.B.; Kornfeld, O.S.; O’Rourke, K.; Mirrashidi, K.M.; Haley, B.; Watanabe, C.; Roose-Girma, M.; Modrusan, Z.; et al. IRF2 transcriptionally induces GSDMD expression for pyroptosis. Sci. Signal. 2019, 12, eaax4917. [Google Scholar] [CrossRef] [PubMed]
- Aachoui, Y.; Kajiwara, Y.; Leaf, I.A.; Mao, D.; Ting, J.P.-Y.; Coers, J.; Aderem, A.; Buxbaum, J.D.; Miao, E.A. Canonical Inflammasomes Drive IFN-γ to Prime Caspase-11 in Defense against a Cytosol-Invasive Bacterium. Cell Host Microbe 2015, 18, 320–332. [Google Scholar] [CrossRef]
- Karki, R.; Lee, E.; Place, D.; Samir, P.; Mavuluri, J.; Sharma, B.R.; Balakrishnan, A.; Malireddi, R.S.; Geiger, R.; Zhu, Q.; et al. IRF8 Regulates Transcription of Naips for NLRC4 Inflammasome Activation. Cell 2018, 173, 920–933. [Google Scholar] [CrossRef] [PubMed]
- Guarda, G.; Braun, M.; Staehli, F.; Tardivel, A.; Mattmann, C.; Förster, I.; Farlik, M.; Decker, T.; Du Pasquier, R.A.; Romero, P.; et al. Type I Interferon Inhibits Interleukin-1 Production and Inflammasome Activation. Immunology 2011, 34, 213–223. [Google Scholar] [CrossRef]
- Heinz, S.; Romanoski, C.E.; Benner, C.; Glass, C.K. The selection and function of cell type-specific enhancers. Nat. Rev. Mol. Cell Biol. 2015, 16, 144–154. [Google Scholar] [CrossRef]
- Sandstrom, A.; Mitchell, P.S.; Goers, L.; Mu, E.W.; Lesser, C.F.; Vance, R.E. Functional degradation: A mechanism of NLRP1 inflammasome activation by diverse pathogen enzymes. Science 2019, 364, eaau1330. [Google Scholar] [CrossRef]
- Boyden, E.D.; Dietrich, W.F. Nalp1b controls mouse macrophage susceptibility to anthrax lethal toxin. Nat. Genet. 2006, 38, 240–244. [Google Scholar] [CrossRef]
- Masters, S.L.; Gerlic, M.; Metcalf, D.; Preston, S.; Pellegrini, M.; O’Donnell, J.A.; McArthur, K.; Baldwin, T.M.; Chevrier, S.; Nowell, C.J.; et al. NLRP1 Inflammasome Activation Induces Pyroptosis of Hematopoietic Progenitor Cells. Immunology 2012, 37, 1009–1023. [Google Scholar] [CrossRef]
- Im, S.-S.; Yousef, L.; Blaschitz, C.; Liu, J.Z.; Edwards, R.A.; Young, S.G.; Raffatellu, M.; Osborne, T.F. Linking Lipid Metabolism to the Innate Immune Response in Macrophages through Sterol Regulatory Element Binding Protein-1a. Cell Metab. 2011, 13, 540–549. [Google Scholar] [CrossRef]
- Murphy, A.J.; Kraakman, M.J.; Kammoun, H.L.; Dragoljevic, D.; Lee, M.K.; Lawlor, K.E.; Wentworth, J.M.; VasanthaKumar, A.; Gerlic, M.; Whitehead, L.W.; et al. IL-18 Production from the NLRP1 Inflammasome Prevents Obesity and Metabolic Syndrome. Cell Metab. 2016, 23, 155–164. [Google Scholar] [CrossRef] [PubMed]
- Fenini, G.; Grossi, S.; Contassot, E.; Biedermann, T.; Reichmann, E.; French, L.E.; Beer, H.-D. Genome Editing of Human Primary Keratinocytes by CRISPR/Cas9 Reveals an Essential Role of the NLRP1 Inflammasome in UVB Sensing. J. Investig. Dermatol. 2018, 138, 2644–2652. [Google Scholar] [CrossRef]
- Sand, J.; Haertel, E.; Biedermann, T.; Contassot, E.; Reichmann, E.; French, L.E.; Werner, S.; Beer, H.-D. Expression of inflammasome proteins and inflammasome activation occurs in human, but not in murine keratinocytes. Cell Death Dis. 2018, 9, 24. [Google Scholar] [CrossRef]
- D’Osualdo, A.; Anania, V.G.; Yu, K.; Lill, J.R.; Kaufman, R.J.; Matsuzawa, S.-I.; Reed, J.C. Transcription Factor ATF4 Induces NLRP1 Inflammasome Expression during Endoplasmic Reticulum Stress. PLoS ONE 2015, 10, e0130635. [Google Scholar] [CrossRef]
- Bauernfeind, F.G.; Horvath, G.; Stutz, A.; Alnemri, E.S.; Macdonald, K.L.; Speert, D.P.; Fernandes-Alnemri, T.; Wu, J.; Monks, B.G.; Fitzgerald, K.A.; et al. Cutting Edge: NF-κB Activating Pattern Recognition and Cytokine Receptors License NLRP3 Inflammasome Activation by Regulating NLRP3 Expression. J. Immunol. 2009, 183, 787–791. [Google Scholar] [CrossRef] [PubMed]
- Franchi, L.; Eigenbrod, T.; Núñez, G. Cutting Edge: TNF-α Mediates Sensitization to ATP and Silica via the NLRP3 Inflammasome in the Absence of Microbial Stimulation. J. Immunol. 2009, 183, 792–796. [Google Scholar] [CrossRef]
- Juliana, C.; Fernandes-Alnemri, T.; Kang, S.; Farias, A.; Qin, F.; Alnemri, E.S. Non-transcriptional Priming and Deubiquitination Regulate NLRP3 Inflammasome Activation. J. Biol. Chem. 2012, 287, 36617–36622. [Google Scholar] [CrossRef] [PubMed]
- Fernandes-Alnemri, T.; Kang, S.; Anderson, C.; Sagara, J.; Fitzgerald, K.A.; Alnemri, E.S. Cutting Edge: TLR Signaling Licenses IRAK1 for Rapid Activation of the NLRP3 Inflammasome. J. Immunol. 2013, 191, 3995–3999. [Google Scholar] [CrossRef] [PubMed]
- Ghonime, M.G.; Shamaa, O.R.; Das, S.; Eldomany, R.A.; Fernandes-Alnemri, T.; Alnemri, E.S.; Gavrilin, M.A.; Wewers, M.D. Inflammasome Priming by Lipopolysaccharide Is Dependent upon ERK Signaling and Proteasome Function. J. Immunol. 2014, 192, 3881–3888. [Google Scholar] [CrossRef]
- Qiao, Y.; Wang, P.; Qi, J.; Zhang, L.; Gao, C. TLR-induced NF-κB activation regulates NLRP3 expression in murine macrophages. FEBS Lett. 2012, 586, 1022–1026. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Ahn, H.; Yu, S.; Ahn, J.-H.; Ko, H.-J.; Kweon, M.-N.; Hong, E.-J.; An, B.-S.; Lee, E.; Lee, G.-S. IκBζ controls NLRP3 inflammasome activation via upregulation of the Nlrp3 gene. Cytokine 2020, 127, 154983. [Google Scholar] [CrossRef]
- Hildebrand, D.G.; Alexander, E.; Hörber, S.; Lehle, S.; Obermayer, K.; Münck, N.-A.; Rothfuss, O.; Frick, J.-S.; Morimatsu, M.; Schmitz, I.; et al. IκBζ Is a Transcriptional Key Regulator of CCL2/MCP-1. J. Immunol. 2013, 190, 4812–4820. [Google Scholar] [CrossRef] [PubMed]
- Lecoeur, H.; Prina, E.; Rosazza, T.; Kokou, K.; N’Diaye, P.; Aulner, N.; Varet, H.; Bussotti, G.; Xing, Y.; Milon, G.; et al. Targeting Macrophage Histone H3 Modification as a Leishmania Strategy to Dampen the NF-κB/NLRP3-Mediated Inflammatory Response. Cell Rep. 2020, 30, 1870–1882. [Google Scholar] [CrossRef]
- Pourcet, B.; Zecchin, M.; Ferri, L.; Beauchamp, J.; Sitaula, S.; Billon, C.; Delhaye, S.; Vanhoutte, J.; Mayeuf-Louchart, A.; Thorel, Q.; et al. Nuclear Receptor Subfamily 1 Group D Member 1 Regulates Circadian Activity of NLRP3 Inflammasome to Reduce the Severity of Fulminant Hepatitis in Mice. Gastroenterology 2018, 154, 1449–1464. [Google Scholar] [CrossRef]
- Wang, S.; Lin, Y.; Yuan, X.; Li, F.; Guo, L.; Wu, B. REV-ERBα integrates colon clock with experimental colitis through regulation of NF-κB/NLRP3 axis. Nat. Commun. 2018, 9, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Huai, W.; Zhao, R.; Song, H.; Zhao, J.; Zhang, L.; Zhang, L.; Gao, C.; Han, L.; Zhao, W. Aryl hydrocarbon receptor negatively regulates NLRP3 inflammasome activity by inhibiting NLRP3 transcription. Nat. Commun. 2014, 5, 4738. [Google Scholar] [CrossRef] [PubMed]
- Chen, D.; Xiong, X.-Q.; Zang, Y.-H.; Tong, Y.; Zhou, B.; Chen, Q.; Li, Y.-H.; Gao, X.-Y.; Kang, Y.-M.; Zhu, G.-Q. BCL6 attenuates renal inflammation via negative regulation of NLRP3 transcription. Cell Death Dis. 2017, 8, e3156. [Google Scholar] [CrossRef] [PubMed]
- Zhu, L.; Meng, Q.; Liang, S.; Ma, Y.; Li, R.; Li, G.; Zeng, H. The transcription factor GFI1 negatively regulates NLRP3 inflammasome activation in macrophages. FEBS Lett. 2014, 588, 4513–4519. [Google Scholar] [CrossRef]
- Ma, P.; Zha, S.; Shen, X.; Zhao, Y.; Li, L.; Yang, L.; Lei, M.; Liu, W. NFAT5 mediates hypertonic stress-induced atherosclerosis via activating NLRP3 inflammasome in endothelium. Cell Commun. Signal. 2019, 17, 1–13. [Google Scholar] [CrossRef]
- Erlich, Z.; Shlomovitz, I.; Edry-Botzer, L.; Cohen, H.; Frank, D.; Wang, H.; Lew, A.M.; Lawlor, K.E.; Zhan, Y.; Vince, J.E.; et al. Macrophages, rather than DCs, are responsible for inflammasome activity in the GM-CSF BMDC model. Nat. Immunol. 2019, 20, 397–406. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Zhao, Y.; Shi, J.; Shao, F. Human NAIP and mouse NAIP1 recognize bacterial type III secretion needle protein for inflammasome activation. Proc. Natl. Acad. Sci. USA 2013, 110, 14408–14413. [Google Scholar] [CrossRef] [PubMed]
- Kortmann, J.; Brubaker, S.W.; Monack, D.M. Cutting Edge: Inflammasome Activation in Primary Human Macrophages Is Dependent on Flagellin. J. Immunol. 2015, 195, 815–819. [Google Scholar] [CrossRef]
- Kofoed, E.M.; Vance, R.E. Innate immune recognition of bacterial ligands by NAIPs determines inflammasome specificity. Nat. Cell Biol. 2011, 477, 592–595. [Google Scholar] [CrossRef]
- Zhao, Y.; Shi, J.; Shi, X.; Wang, Y.; Wang, F.; Shao, F. Genetic functions of the NAIP family of inflammasome receptors for bacterial ligands in mice. J. Exp. Med. 2016, 213, 647–656. [Google Scholar] [CrossRef]
- Langlais, D.; Barreiro, L.B.; Gros, P. The macrophage IRF8/IRF1 regulome is required for protection against infections and is associated with chronic inflammation. J. Exp. Med. 2016, 213, 585–603. [Google Scholar] [CrossRef] [PubMed]
- Fortier, A.; Doiron, K.; Saleh, M.; Grinstein, S.; Gros, P. Restriction of Legionella pneumophila Replication in Macrophages Requires Concerted Action of the Transcriptional Regulators Irf1 and Irf8 and Nod-Like Receptors Naip5 and Nlrc4. Infect. Immun. 2009, 77, 4794–4805. [Google Scholar] [CrossRef]
- Kim, S.; Bagadia, P.; Anderson, D.A.; Liu, T.-T.; Huang, X.; Theisen, D.J.; O’Connor, K.W.; Ohara, R.A.; Iwata, A.; Murphy, T.L.; et al. High Amount of Transcription Factor IRF8 Engages AP1-IRF Composite Elements in Enhancers to Direct Type 1 Conventional Dendritic Cell Identity. Immunity 2020, 53, 759–774. [Google Scholar] [CrossRef]
- Fernandes-Alnemri, T.; Yu, J.-W.; Datta, P.; Wu, J.; Alnemri, E.S. AIM2 activates the inflammasome and cell death in response to cytoplasmic DNA. Nat. Cell Biol. 2009, 458, 509–513. [Google Scholar] [CrossRef]
- Hornung, V.; Ablasser, A.; Charrel-Dennis, M.; Bauernfeind, F.G.; Horvath, G.; Caffrey, D.R.; Latz, E.; Fitzgerald, K.A. AIM2 recognizes cytosolic dsDNA and forms a caspase-1-activating inflammasome with ASC. Nat. Cell Biol. 2009, 458, 514–518. [Google Scholar] [CrossRef]
- Di Micco, A.; Frera, G.; Lugrin, J.; Jamilloux, Y.; Hsu, E.-T.; Tardivel, A.; De Gassart, A.; Zaffalon, L.; Bujisic, B.; Siegert, S.; et al. AIM2 inflammasome is activated by pharmacological disruption of nuclear envelope integrity. Proc. Natl. Acad. Sci. USA 2016, 113, E4671–E4680. [Google Scholar] [CrossRef] [PubMed]
- Man, S.M.; Karki, R.; Malireddi, R.S.; Neale, G.; Vogel, P.; Yamamoto, M.; Lamkanfi, M.; Kanneganti, T.-D. The transcription factor IRF1 and guanylate-binding proteins target activation of the AIM2 inflammasome by Francisella infection. Nat. Immunol. 2015, 16, 467–475. [Google Scholar] [CrossRef] [PubMed]
- Meunier, E.; Wallet, P.; Dreier, R.F.; Costanzo, S.; Anton, L.; Rühl, S.; Dussurgey, S.; Dick, M.S.; Kistner, A.; Rigard, M.; et al. Guanylate-binding proteins promote activation of the AIM2 inflammasome during infection with Francisella novicida. Nat. Immunol. 2015, 16, 476–484. [Google Scholar] [CrossRef] [PubMed]
- Gaidt, M.M.; Ebert, T.S.; Chauhan, D.; Ramshorn, K.; Pinci, F.; Zuber, S.; O’Duill, F.; Schmid-Burgk, J.L.; Hoss, F.; Buhmann, R.; et al. The DNA Inflammasome in Human Myeloid Cells Is Initiated by a STING-Cell Death Program Upstream of NLRP3. Cell 2017, 171, 1110–1124. [Google Scholar] [CrossRef]
- Fisch, D.; Bando, H.; Clough, B.; Hornung, V.; Yamamoto, M.; Shenoy, A.R.; Frickel, E. Human GBP 1 is a microbe-specific gatekeeper of macrophage apoptosis and pyroptosis. EMBO J. 2019, 38, e100926. [Google Scholar] [CrossRef] [PubMed]
- Schroder, K.; Sagulenko, V.; Zamoshnikova, A.; Richards, A.A.; Cridland, J.A.; Irvine, K.M.; Stacey, K.J.; Sweet, M.J. Acute lipopolysaccharide priming boosts inflammasome activation independently of inflammasome sensor induction. Immunobiology 2012, 217, 1325–1329. [Google Scholar] [CrossRef]
- Dombrowski, Y.; Peric, M.; Koglin, S.; Kammerbauer, C.; Göss, C.; Anz, D.; Simanski, M.; Gläser, R.; Harder, J.; Hornung, V.; et al. Cytosolic DNA Triggers Inflammasome Activation in Keratinocytes in Psoriatic Lesions. Sci. Transl. Med. 2011, 3, 82ra38. [Google Scholar] [CrossRef]
- Doody, G.M.; Care, M.A.; Burgoyne, N.J.; Bradford, J.R.; Bota, M.; Bonifer, C.; Westhead, D.R.; Tooze, R.M. An extended set of PRDM1/BLIMP1 target genes links binding motif type to dynamic repression. Nucleic Acids Res. 2010, 38, 5336–5350. [Google Scholar] [CrossRef]
- Chuong, E.B.; Elde, N.C.; Feschotte, C. Regulatory evolution of innate immunity through co-option of endogenous retroviruses. Science 2016, 351, 1083–1087. [Google Scholar] [CrossRef]
- Xu, H.; Yang, J.; Gao, W.; Li, L.; Li, P.; Zhang, L.; Gong, Y.-N.; Peng, X.; Xi, J.J.; Chen, S.; et al. Innate immune sensing of bacterial modifications of Rho GTPases by the Pyrin inflammasome. Nat. Cell Biol. 2014, 513, 237–241. [Google Scholar] [CrossRef]
- Jamilloux, Y.; Magnotti, F.; Belot, A.; Henry, T. The pyrin inflammasome: From sensing RhoA GTPases-inhibiting toxins to triggering autoinflammatory syndromes. Pathog. Dis. 2018, 76. [Google Scholar] [CrossRef] [PubMed]
- Gavrilin, M.A.; Mitra, S.; Seshadri, S.; Nateri, J.; Berhe, F.; Hall, M.W.; Wewers, M.D. Pyrin Critical to Macrophage IL-1β Response toFrancisellaChallenge. J. Immunol. 2009, 182, 7982–7989. [Google Scholar] [CrossRef] [PubMed]
- Centola, M.; Wood, G.; Frucht, D.M.; Galon, J.; Aringer, M.; Farrell, C.; Kingma, D.W.; Horwitz, M.E.; Mansfield, E.; Holland, S.M.; et al. The gene for familial Mediterranean fever, MEFV, is expressed in early leukocyte development and is regulated in response to inflammatory mediators. Blood 2000, 95, 3223–3231. [Google Scholar] [CrossRef] [PubMed]
- Grandemange, S.; Aksentijevich, I.; Jeru, I.; Gül, A.; Touitou, I. The regulation of MEFV expression and its role in health and familial Mediterranean fever. Genes Immun. 2011, 12, 497–503. [Google Scholar] [CrossRef]
- Papin, S.; Cazeneuve, C.; Duquesnoy, P.; Jéru, I.; Sahali, D.; Amselem, S. The Tumor Necrosis Factor α-dependent Activation of the Human Mediterranean Fever (MEFV) Promoter Is Mediated by a Synergistic Interaction between C/EBPβ and NFκB p65. J. Biol. Chem. 2003, 278, 48839–48847. [Google Scholar] [CrossRef] [PubMed]
- Sharma, D.; Malik, A.; Guy, C.; Vogel, P.; Kanneganti, T.-D. TNF/TNFR axis promotes pyrin inflammasome activation and distinctly modulates pyrin inflammasomopathy. J. Clin. Investig. 2018, 129, 150–162. [Google Scholar] [CrossRef]
- Deyoung, K.L.; E Ray, M.; A Su, Y.; Anzick, S.L.; Johnstone, R.W.; A Trapani, J.; Meltzer, P.S.; Trent, J.M. Cloning a novel member of the human interferon-inducible gene family associated with control of tumorigenicity in a model of human melanoma. Oncogene 1997, 15, 453–457. [Google Scholar] [CrossRef]
- E Conway, K.; McConnell, B.B.; E Bowring, C.; Donald, C.D.; Warren, S.T.; Vertino, P.M. TMS1, a novel proapoptotic caspase recruitment domain protein, is a target of methylation-induced gene silencing in human breast cancers. Cancer Res. 2000, 60, 6236–6242. [Google Scholar]
- Ghosh, S.; Wallerath, C.; Covarrubias, S.; Hornung, V.; Carpenter, S.B.; Fitzgerald, K.A.; Wallareth, C. The PYHIN Protein p205 Regulates the Inflammasome by Controlling Asc Expression. J. Immunol. 2017, 199, 3249–3260. [Google Scholar] [CrossRef]
- Brunette, R.L.; Young, J.M.; Whitley, D.G.; Brodsky, I.E.; Malik, H.S.; Stetson, D.B. Extensive evolutionary and functional diversity among mammalian AIM2-like receptors. J. Exp. Med. 2012, 209, 1969–1983. [Google Scholar] [CrossRef]
- Ming, X.; Li, W.; Maeda, Y.; Blumberg, B.; Raval, S.; Cook, S.D.; Dowling, P.C. Caspase-1 expression in multiple sclerosis plaques and cultured glial cells. J. Neurol. Sci. 2002, 197, 9–18. [Google Scholar] [CrossRef]
- Giamarellos-Bourboulis, E.J.; Van De Veerdonk, F.L.; Mouktaroudi, M.; Raftogiannis, M.; Antonopoulou, A.; Joosten, L.A.; Pickkers, P.; Savva, A.; Georgitsi, M.; Van Der Meer, J.W.M.; et al. Inhibition of caspase-1 activation in gram-negative sepsis and experimental endotoxemia. Crit. Care 2011, 15, R27. [Google Scholar] [CrossRef] [PubMed]
- Fairfax, B.P.; Humburg, P.; Makino, S.; Naranbhai, V.; Wong, D.; Lau, E.; Jostins, L.; Plant, K.; Andrews, R.; McGee, C.; et al. Innate Immune Activity Conditions the Effect of Regulatory Variants upon Monocyte Gene Expression. Science 2014, 343, 1246949. [Google Scholar] [CrossRef]
- Dai, C.; Krantz, S.B. Interferon gamma induces upregulation and activation of caspases 1, 3, and 8 to produce apoptosis in human erythroid progenitor cells. Blood 1999, 93, 3309–3316. [Google Scholar] [CrossRef] [PubMed]
- Tamura, T.; Ishihara, M.; Lamphier, M.S.; Tanaka, N.; Oishi, I.; Aizawa, S.; Matsuyama, T.; Mak, T.W.; Taki, S.; Taniguchi, T. An IRF-1-dependent pathway of DNA damage-induced apoptosis in mitogen-activated T lymphocytes. Nat. Cell Biol. 1995, 376, 596–599. [Google Scholar] [CrossRef] [PubMed]
- Cuesta, N.; Nhu, Q.M.; Zudaire, E.; Polumuri, S.; Cuttitta, F.; Vogel, S.N. IFN regulatory factor-2 regulates macrophage apoptosis through a STAT1/3- and caspase-1-dependent mechanism. J. Immunol. (Baltim. MD 1950) 2007, 178, 3602–3611. [Google Scholar] [CrossRef]
- Lv, D.; Zhang, K.; Li, R. Interferon regulatory factor 8 regulates caspase-1 expression to facilitate Epstein-Barr virus reactivation in response to B cell receptor stimulation and chemical induction. PLoS Pathog. 2018, 14, e1006868. [Google Scholar] [CrossRef]
- Thygesen, S.J.; Stacey, K.J. IRF 1 and IRF 2 regulate the non-canonical inflammasome. EMBO Rep. 2019, 20, e48891. [Google Scholar] [CrossRef]
- Yang, H.-J.; Wang, M.; Wang, L.; Cheng, B.-F.; Lin, X.-Y.; Feng, Z.-W. NF-κB Regulates Caspase-4 Expression and Sensitizes Neuroblastoma Cells to Fas-Induced Apoptosis. PLoS ONE 2015, 10, e0117953. [Google Scholar] [CrossRef]
- Schauvliege, R.; Vanrobaeys, J.; Schotte, P.; Beyaert, R. Caspase-11 Gene Expression in Response to Lipopolysaccharide and Interferon-γ Requires Nuclear Factor-κB and Signal Transducer and Activator of Transcription (STAT) 1. J. Biol. Chem. 2002, 277, 41624–41630. [Google Scholar] [CrossRef]
- Crowley, S.M.; Han, X.; Allaire, J.M.; Stahl, M.; Rauch, I.; Knodler, L.A.; Vallance, B.A. Intestinal restriction of Salmonella Typhimurium requires caspase-1 and caspase-11 epithelial intrinsic inflammasomes. PLoS Pathog. 2020, 16, e1008498. [Google Scholar] [CrossRef] [PubMed]
- Endo, M.; Mori, M.; Akira, S.; Gotoh, T. C/EBP Homologous Protein (CHOP) Is Crucial for the Induction of Caspase-11 and the Pathogenesis of Lipopolysaccharide-Induced Inflammation. J. Immunol. 2006, 176, 6245–6253. [Google Scholar] [CrossRef] [PubMed]
- Eleazer, R.; Fondufe-Mittendorf, Y.N. The multifaceted role of PARP1 in RNA biogenesis. Wiley Interdiscip. Rev. RNA 2020, e12607. [Google Scholar] [CrossRef] [PubMed]
- Paul, O.H.; Covic, M.; Hasan, S.; Imhof, R.; Hottiger, M.O. The Enzymatic and DNA Binding Activity of PARP-1 Are Not Required for NF-κB Coactivator Function. J. Biol. Chem. 2001, 276, 45588–45597. [Google Scholar] [CrossRef]
- Yoo, L.; Hong, S.; Shin, K.S.; Kang, S.J. PARP-1 regulates the expression of caspase-11. Biochem. Biophys. Res. Commun. 2011, 408, 489–493. [Google Scholar] [CrossRef]
- Frank, A.K.; Leu, J.I.-J.; Zhou, Y.; Devarajan, K.; Nedelko, T.; Klein-Szanto, A.; Hollstein, M.C.; Murphy, M.E. The Codon 72 Polymorphism of p53 Regulates Interaction with NF-κB and Transactivation of Genes Involved in Immunity and Inflammation. Mol. Cell. Biol. 2011, 31, 1201–1213. [Google Scholar] [CrossRef]
- Gupta, S.; Radha, V.; Furukawa, Y.; Swarup, G. Direct Transcriptional Activation of Human Caspase-1 by Tumor Suppressor p53. J. Biol. Chem. 2001, 276, 10585–10588. [Google Scholar] [CrossRef]
- Sadasivam, S.; Gupta, S.; Radha, V.; Batta, K.; Kundu, T.K.; Swarup, G. Caspase-1 activator Ipaf is a p53-inducible gene involved in apoptosis. Oncogene 2004, 24, 627–636. [Google Scholar] [CrossRef]
- Hagar, J.A.; Powell, D.A.; Aachoui, Y.; Ernst, R.K.; Miao, E.A. Cytoplasmic LPS Activates Caspase-11: Implications in TLR4-Independent Endotoxic Shock. Science 2013, 341, 1250–1253. [Google Scholar] [CrossRef]
- Kayagaki, N.; Wong, M.T.; Stowe, I.B.; Ramani, S.R.; Gonzalez, L.C.; Akashi-Takamura, S.; Miyake, K.; Zhang, J.; Lee, W.P.; Muszyński, A.; et al. Noncanonical Inflammasome Activation by Intracellular LPS Independent of TLR4. Science 2013, 341, 1246–1249. [Google Scholar] [CrossRef]
- Kajiwara, Y.; Schiff, T.; Voloudakis, G.; Sosa, M.A.G.; Elder, G.; Bozdagi, O.; Buxbaum, J.D. A Critical Role for Human Caspase-4 in Endotoxin Sensitivity. J. Immunol. 2014, 193, 335–343. [Google Scholar] [CrossRef]
- Lin, X.Y.; Choi, M.S.K.; Porter, A.G. Expression Analysis of the Human Caspase-1 Subfamily Reveals Specific Regulation of theCASP5Gene by Lipopolysaccharide and Interferon-γ. J. Biol. Chem. 2000, 275, 39920–39926. [Google Scholar] [CrossRef] [PubMed]
- Casson, C.N.; Yu, J.; Reyes, V.M.; Taschuk, F.O.; Yadav, A.; Copenhaver, A.M.; Nguyen, H.T.; Collman, R.G.; Shin, S. Human caspase-4 mediates noncanonical inflammasome activation against gram-negative bacterial pathogens. Proc. Natl. Acad. Sci. USA 2015, 112, 6688–6693. [Google Scholar] [CrossRef] [PubMed]
- Eckhart, L.; Kittel, C.; Gawlas, S.; Gruber, F.; Mildner, M.; Jilma, B.; Tschachler, E. Identification of a novel exon encoding the amino-terminus of the predominant caspase-5 variants. Biochem. Biophys. Res. Commun. 2006, 348, 682–688. [Google Scholar] [CrossRef]
- Salskov-Iversen, M.L.; Johansen, C.; Kragballe, K.; Iversen, L. Caspase-5 Expression Is Upregulated in Lesional Psoriatic Skin. J. Investig. Dermatol. 2011, 131, 670–676. [Google Scholar] [CrossRef]
- Li, Y.; Guo, X.; Hu, C.; Du, Y.; Guo, C.; Wang, D.; Zhao, W.; Huang, G.; Li, C.; Lu, Q.; et al. Type I IFN operates pyroptosis and necroptosis during multidrug-resistant A. baumannii infection. Cell Death Differ. 2018, 25, 1304–1318. [Google Scholar] [CrossRef]
- Liu, Z.; Gan, L.; Xu, Y.; Luo, D.; Ren, Q.; Wu, S.; Sun, C. Melatonin alleviates inflammasome-induced pyroptosis through inhibiting NF-κB/GSDMD signal in mice adipose tissue. J. Pineal Res. 2017, 63, e12414. [Google Scholar] [CrossRef] [PubMed]
- Dinarello, C.A. Immunological and Inflammatory Functions of the Interleukin-1 Family. Annu. Rev. Immunol. 2009, 27, 519–550. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.M.; Kang, H.S.; Paik, S.G.; Pyun, K.H.; Anderson, K.L.; E Torbett, B.; Choi, I. Roles of IFN consensus sequence binding protein and PU.1 in regulating IL-18 gene expression. J. Immunol. 1999, 163, 2000–2007. [Google Scholar]
- Verweyen, E.; Holzinger, D.; Weinhage, T.; Hinze, C.; Wittkowski, H.; Pickkers, P.; Albeituni, S.; Verbist, K.; Nichols, K.E.; Schulert, G.; et al. Synergistic Signaling of TLR and IFNα/β Facilitates Escape of IL-18 Expression from Endotoxin Tolerance. Am. J. Respir. Crit. Care Med. 2020, 201, 526–539. [Google Scholar] [CrossRef]
- Zhu, Q.; Kanneganti, T.-D. Cutting Edge: Distinct Regulatory Mechanisms Control Proinflammatory Cytokines IL-18 and IL-1β. J. Immunol. 2017, 198, 4210–4215. [Google Scholar] [CrossRef] [PubMed]
- Muñoz, M.; Eidenschenk, C.; Ota, N.; Wong, K.; Lohmann, U.; Kühl, A.A.; Wang, X.; Manzanillo, P.; Li, Y.; Rutz, S.; et al. Interleukin-22 Induces Interleukin-18 Expression from Epithelial Cells during Intestinal Infection. Immunity 2015, 42, 321–331. [Google Scholar] [CrossRef]
- Levy, M.; Thaiss, C.A.; Zeevi, D.; Dohnalová, L.; Zilberman-Schapira, G.; Mahdi, J.A.; David, E.; Savidor, A.; Korem, T.; Herzig, Y.; et al. Microbiota-Modulated Metabolites Shape the Intestinal Microenvironment by Regulating NLRP6 Inflammasome Signaling. Cell 2015, 163, 1428–1443. [Google Scholar] [CrossRef] [PubMed]
- Hurgin, V.; Novick, D.; Rubinstein, M. The promoter of IL-18 binding protein: Activation by an IFN-/-induced complex of IFN regulatory factor 1 and CCAAT/enhancer binding protein. Proc. Natl. Acad. Sci. USA 2002, 99, 16957–16962. [Google Scholar] [CrossRef] [PubMed]
- Bachmann, M.; Paulukat, J.; Pfeilschifter, J.; Mühl, H. Molecular mechanisms of IL-18BP regulation in DLD-1 cells: Pivotal direct action of the STAT1/GAS axis on the promoter level. J. Cell. Mol. Med. 2008, 13, 1987–1994. [Google Scholar] [CrossRef]
- Kominato, Y.; Galson, D.; Waterman, W.R.; Webb, A.C.; E Auron, P. Monocyte expression of the human prointerleukin 1 beta gene (IL1B) is dependent on promoter sequences which bind the hematopoietic transcription factor Spi-1/PU.1. Mol. Cell. Biol. 1995, 15, 58–68. [Google Scholar] [CrossRef] [PubMed]
- Minderjahn, J.; Schmidt, A.; Fuchs, A.; Schill, R.; Raithel, J.; Babina, M.; Schmidl, C.; Gebhard, C.; Schmidhofer, S.; Mendes, K.; et al. Mechanisms governing the pioneering and redistribution capabilities of the non-classical pioneer PU.1. Nat. Commun. 2020, 11, 1–16. [Google Scholar] [CrossRef]
- Plachetka, A.; Chayka, O.; Wilczek, C.; Melnik, S.; Bonifer, C.; Klempnauer, K.-H. C/EBPβ Induces Chromatin Opening at a Cell-Type-Specific Enhancer. Mol. Cell. Biol. 2008, 28, 2102–2112. [Google Scholar] [CrossRef]
- Liang, M.D.; Zhang, Y.; McDevit, D.; Marecki, S.; Nikolajczyk, B.S. The Interleukin-1β Gene Is Transcribed from a Poised Promoter Architecture in Monocytes. J. Biol. Chem. 2006, 281, 9227–9237. [Google Scholar] [CrossRef]
- Zhang, Y.; Saccani, S.; Shin, H.; Nikolajczyk, B.S. Dynamic Protein Associations Define Two Phases of IL-1β Transcriptional Activation. J. Immunol. 2008, 181, 503–512. [Google Scholar] [CrossRef]
- Serkkola, E.; Hurme, M. Synergism between protein-kinase C and cAMP-dependent pathways in the expression of the interleukin-1beta gene is mediated via the activator-protein-1 (AP-1) enhancer activity. JBIC J. Biol. Inorg. Chem. 1993, 213, 243–249. [Google Scholar] [CrossRef]
- Unlu, S.; Kumar, A.; Waterman, W.R.; Tsukada, J.; Wang, K.Z.; Galson, D.L.; Auron, P.E. Phosphorylation of IRF8 in a pre-associated complex with Spi-1/PU.1 and non-phosphorylated Stat1 is critical for LPS induction of the IL1B gene. Mol. Immunol. 2007, 44, 3364–3379. [Google Scholar] [CrossRef]
- Shirakawa, F.; Saito, K.; A Bonagura, C.; Galson, D.L.; Fenton, M.J.; Webb, A.C.; E Auron, P. The human prointerleukin 1 beta gene requires DNA sequences both proximal and distal to the transcription start site for tissue-specific induction. Mol. Cell. Biol. 1993, 13, 1332–1344. [Google Scholar] [CrossRef] [PubMed]
- Adamik, J.; Wang, K.Z.Q.; Unlu, S.; Su, A.-J.A.; Tannahill, G.M.; Galson, D.L.; O’Neill, L.A.; Auron, P.E. Distinct Mechanisms for Induction and Tolerance Regulate the Immediate Early Genes Encoding Interleukin 1β and Tumor Necrosis Factor α. PLoS ONE 2013, 8, e70622. [Google Scholar] [CrossRef]
- Hiscott, J.; Marois, J.; Garoufalis, J.; D’Addario, M.; Roulston, A.; Kwan, I.; Pepin, N.; Lacoste, J.; Nguyen, H.; Bensi, G. Characterization of a functional NF-kappa B site in the human interleukin 1 beta promoter: Evidence for a positive autoregulatory loop. Mol. Cell. Biol. 1993, 13, 6231–6240. [Google Scholar] [CrossRef] [PubMed]
- Tsukada, J.; Saito, K.; Waterman, W.R.; Webb, A.C.; E Auron, P. Transcription factors NF-IL6 and CREB recognize a common essential site in the human prointerleukin 1 beta gene. Mol. Cell. Biol. 1994, 14, 7285–7297. [Google Scholar] [CrossRef][Green Version]
- Chandra, G.; Cogswell, J.P.; Miller, L.R.; Godlevski, M.M.; Stinnett, S.W.; Noel, S.L.; Kadwell, S.H.; A Kost, T.; Gray, J.G. Cyclic AMP signaling pathways are important in IL-1 beta transcriptional regulation. J. Immunol. 1995, 155, 4535–4543. [Google Scholar]
- Chen, Y.; Zhuang, S.; Cassenaer, S.; Casteel, D.E.; Gudi, T.; Boss, G.R.; Pilz, R.B. Synergism between Calcium and Cyclic GMP in Cyclic AMP Response Element-Dependent Transcriptional Regulation Requires Cooperation between CREB and C/EBP-β. Mol. Cell. Biol. 2003, 23, 4066–4082. [Google Scholar] [CrossRef] [PubMed]
- Park, Y.-G.; Kang, S.-K.; Noh, S.-H.; Park, K.-K.; Chang, Y.-C.; Lee, Y.-C.; Kim, C.-H. PGE2 induces IL-1β gene expression in mouse osteoblasts through a cAMP–PKA signaling pathway. Int. Immunopharmacol. 2004, 4, 779–789. [Google Scholar] [CrossRef] [PubMed]
- Zasłona, Z.; Pålsson-McDermott, E.M.; Menon, D.; Haneklaus, M.; Flis, E.; Prendeville, H.; Corcoran, S.E.; Peters-Golden, M.; O’Neill, L.A. The Induction of Pro–IL-1β by Lipopolysaccharide Requires Endogenous Prostaglandin E2Production. J. Immunol. 2017, 198, 3558–3564. [Google Scholar] [CrossRef]
- Tannahill, G.M.; Curtis, A.M.; Adamik, J.; Palsson-McDermott, E.M.; McGettrick, A.F.; Goel, G.; Frezza, C.; Bernard, N.J.; Kelly, B.T.; Foley, N.H.; et al. Succinate is an inflammatory signal that induces IL-1β through HIF-1α. Nat. Cell Biol. 2013, 496, 238–242. [Google Scholar] [CrossRef]
- Ménégaut, L.; Thomas, C.; Jalil, A.; Julla, J.B.; Magnani, C.; Ceroi, A.; Basmaciyan, L.; Dumont, A.; Le Goff, W.; Mathew, M.J.; et al. Interplay between Liver X Receptor and Hypoxia Inducible Factor 1α Potentiates Interleukin-1β Production in Human Macrophages. Cell Rep. 2020, 31, 107665. [Google Scholar] [CrossRef]
- Kobayashi, E.H.; Suzuki, T.; Funayama, R.; Nagashima, T.; Hayashi, M.; Sekine, H.; Tanaka, N.; Moriguchi, T.; Motohashi, H.; Nakayama, K.; et al. Nrf2 suppresses macrophage inflammatory response by blocking proinflammatory cytokine transcription. Nat. Commun. 2016, 7, 11624. [Google Scholar] [CrossRef] [PubMed]
- Myles, I.A.; Fontecilla, N.M.; Valdez, P.A.; Vithayathil, P.J.; Naik, S.; Belkaid, Y.; Ouyang, W.; Datta, S.K. Signaling via the IL-20 receptor inhibits cutaneous production of IL-1β and IL-17A to promote infection with methicillin-resistant Staphylococcus aureus. Nat. Immunol. 2013, 14, 804–811. [Google Scholar] [CrossRef] [PubMed]
- Doitsh, G.; Galloway, N.L.K.; Geng, X.; Yang, Z.; Monroe, K.M.; Zepeda, O.; Hunt, P.W.; Hatano, H.; Sowinski, S.; Muñoz-Arias, I.; et al. Cell death by pyroptosis drives CD4 T-cell depletion in HIV-1 infection. Nat. Cell Biol. 2013, 505, 509–514. [Google Scholar] [CrossRef] [PubMed]
- Martin, B.N.; Wang, C.; Zhang, C.-J.; Kang, Z.; Gulen, M.F.; Zepp, J.A.; Zhao, J.; Bian, G.; Do, J.-S.; Min, B.; et al. T cell–intrinsic ASC critically promotes TH17-mediated experimental autoimmune encephalomyelitis. Nat. Immunol. 2016, 17, 583–592. [Google Scholar] [CrossRef] [PubMed]
- Arbore, G.; West, E.E.; Spolski, R.; Robertson, A.A.B.; Klos, A.; Rheinheimer, C.; Dutow, P.; Woodruff, T.M.; Yu, Z.X.; O’Neill, L.A.; et al. T helper 1 immunity requires complement-driven NLRP3 inflammasome activity in CD4+ T cells. Science 2016, 352, aad1210. [Google Scholar] [CrossRef] [PubMed]
- Pulugulla, S.H.; Packard, T.A.; Galloway, N.L.; Grimmett, Z.W.; Doitsh, G.; Adamik, J.; Galson, D.L.; Greene, W.C.; Auron, P.E. Distinct mechanisms regulate IL1B gene transcription in lymphoid CD4 T cells and monocytes. Cytokine 2018, 111, 373–381. [Google Scholar] [CrossRef]
- Wan, C.-K.; Li, P.; Spolski, R.; Oh, J.; Andraski, A.B.; Du, N.; Yu, Z.-X.; Dillon, C.P.; Green, D.R.; Leonard, W.J. IL-21-mediated non-canonical pathway for IL-1β production in conventional dendritic cells. Nat. Commun. 2015, 6, 7988. [Google Scholar] [CrossRef][Green Version]
- Huang, X.; Feng, Z.; Jiang, Y.; Li, J.; Xiang, Q.; Guo, S.; Yang, C.; Fei, L.; Guo, G.; Zheng, L.; et al. VSIG4 mediates transcriptional inhibition of Nlrp3 and Il-1β in macrophages. Sci. Adv. 2019, 5, eaau7426. [Google Scholar] [CrossRef]
- Hirsch, E.; Irikura, V.M.; Paul, S.M.; Hirsh, D. Functions of interleukin 1 receptor antagonist in gene knockout and overproducing mice. Proc. Natl. Acad. Sci. USA 1996, 93, 11008–11013. [Google Scholar] [CrossRef]
- Arend, W.P.; Smith, M.F.; Janson, R.W.; Joslin, F.G. IL-1 receptor antagonist and IL-1 beta production in human monocytes are regulated differently. J. Immunol. 1991, 147, 1530–1536. [Google Scholar] [PubMed]
- La, E.; Fischer, S.M. Transcriptional Regulation of Intracellular IL-1 Receptor Antagonist Gene by IL-1α in Primary Mouse Keratinocytes. J. Immunol. 2001, 166, 6149–6155. [Google Scholar] [CrossRef] [PubMed]
- Tilg, H.; Mier, J.W.; Vogel, W.; E Aulitzky, W.; Wiedermann, C.J.; Vannier, E.; Huber, C.; A Dinarello, C. Induction of circulating IL-1 receptor antagonist by IFN treatment. J. Immunol. 1993, 150, 4687–4692. [Google Scholar] [PubMed]
- Tamassia, N.; Castellucci, M.; Rossato, M.; Gasperini, S.; Bosisio, D.; Giacomelli, M.; Badolato, R.; Cassatella, M.A.; Bazzoni, F. Uncovering an IL-10-dependent NF-KB recruitment to the IL-1ra promoter that is impaired in STAT3 functionally defective patients. FASEB J. 2009, 24, 1365–1375. [Google Scholar] [CrossRef]
- Camilli, G.; Bohm, M.; Piffer, A.C.; Lavenir, R.; Williams, D.L.; Neven, B.; Grateau, G.; Georgin-Lavialle, S.; Quintin, J. β-Glucan–induced reprogramming of human macrophages inhibits NLRP3 inflammasome activation in cryopyrinopathies. J. Clin. Investig. 2020, 130, 4561–4573. [Google Scholar] [CrossRef]
- Quintin, J.; Saeed, S.; Martens, J.H.; Giamarellos-Bourboulis, E.J.; Ifrim, D.C.; Logie, C.; Jacobs, L.; Jansen, T.; Kullberg, B.-J.; Wijmenga, C.; et al. Candida albicans Infection Affords Protection against Reinfection via Functional Reprogramming of Monocytes. Cell Host Microbe 2012, 12, 223–232. [Google Scholar] [CrossRef]
- Christ, A.; Günther, P.; Lauterbach, M.A.; Duewell, P.; Biswas, D.; Pelka, K.; Scholz, C.J.; Oosting, M.; Haendler, K.; Baßler, K.; et al. Western Diet Triggers NLRP3-Dependent Innate Immune Reprogramming. Cell 2018, 172, 162–175. [Google Scholar] [CrossRef]
- Grandemange, S.; Sanchez, E.; Louis-Plence, P.; Mau-Them, F.T.; Bessis, D.; Coubes, C.; Frouin, E.; Seyger, M.; Girard, M.; Puechberty, J.; et al. A new autoinflammatory and autoimmune syndrome associated with NLRP1 mutations: NAIAD (NLRP1-associated autoinflammation with arthritis and dyskeratosis). Ann. Rheum. Dis. 2016, 76, 1191–1198. [Google Scholar] [CrossRef]
- Zhong, F.L.; Mamaï, O.; Sborgi, L.; Boussofara, L.; Hopkins, R.; Robinson, K.; Szeverényi, I.; Takeichi, T.; Balaji, R.; Lau, A.; et al. Germline NLRP1 Mutations Cause Skin Inflammatory and Cancer Susceptibility Syndromes via Inflammasome Activation. Cell 2016, 167, 187–202. [Google Scholar] [CrossRef]
- Weiss, E.S.; Girard-Guyonvarc’H, C.; Holzinger, D.; De Jesus, A.A.; Tariq, Z.; Picarsic, J.; Schiffrin, E.J.; Foell, D.; Grom, A.A.; Ammann, S.; et al. Interleukin-18 diagnostically distinguishes and pathogenically promotes human and murine macrophage activation syndrome. Blood 2018, 131, 1442–1455. [Google Scholar] [CrossRef] [PubMed]
- Hawkins, P.N.; Lachmann, H.J.; McDermott, M.F. Interleukin-1–Receptor Antagonist in the Muckle–Wells Syndrome. N. Engl. J. Med. 2003, 348, 2583–2584. [Google Scholar] [CrossRef]
- Notarnicola, C.; Boizet-Bonhoure, B.; Barbara, P.D.S.; A Osta, M.; Cattan, D.; Touitou, I. Characterization of new mutations in the 5′-flanking region of the familial Mediterranean fever gene. Genes Immun. 2009, 10, 273–279. [Google Scholar] [CrossRef][Green Version]
- Kruidenier, L.; Chung, C.-W.; Cheng, Z.; Liddle, J.; Che, K.; Joberty, G.; Bantscheff, M.; Bountra, C.; Bridges, A.; Diallo, H.; et al. A selective jumonji H3K27 demethylase inhibitor modulates the proinflammatory macrophage response. Nat. Cell Biol. 2012, 488, 404–408. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).