A First Glimpse of the Mexican Fruit Fly Anastrepha ludens (Diptera: Tephritidae) Antenna Morphology and Proteome in Response to a Proteinaceous Attractant
Abstract
:1. Introduction
2. Results
2.1. Morphological Features of Newly Emerged, Five- and 15-Day-Old Antennas
2.2. Proteome Profile of Antennas across Maturity Stages (Newly Emerged, Five-Day, and 15-Day Old Antennas)
2.3. Comparative Antenna Proteome Profile of A. ludens under Exposure of Ceratrap® by Tandem Mass Tags (TMT) and SPS-MS3 Approach
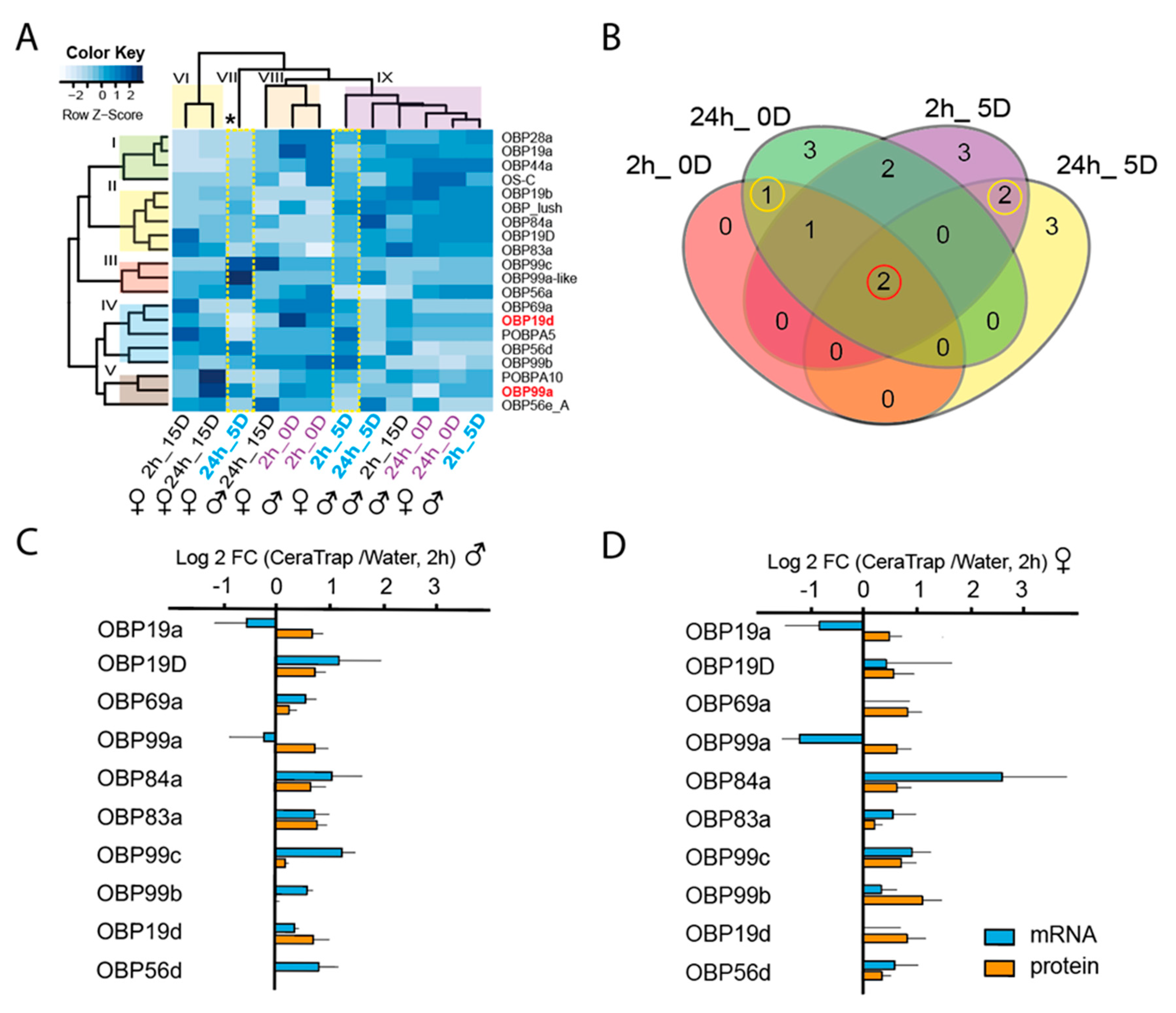
2.3.1. Odorant-Binding Proteins (OBPs)
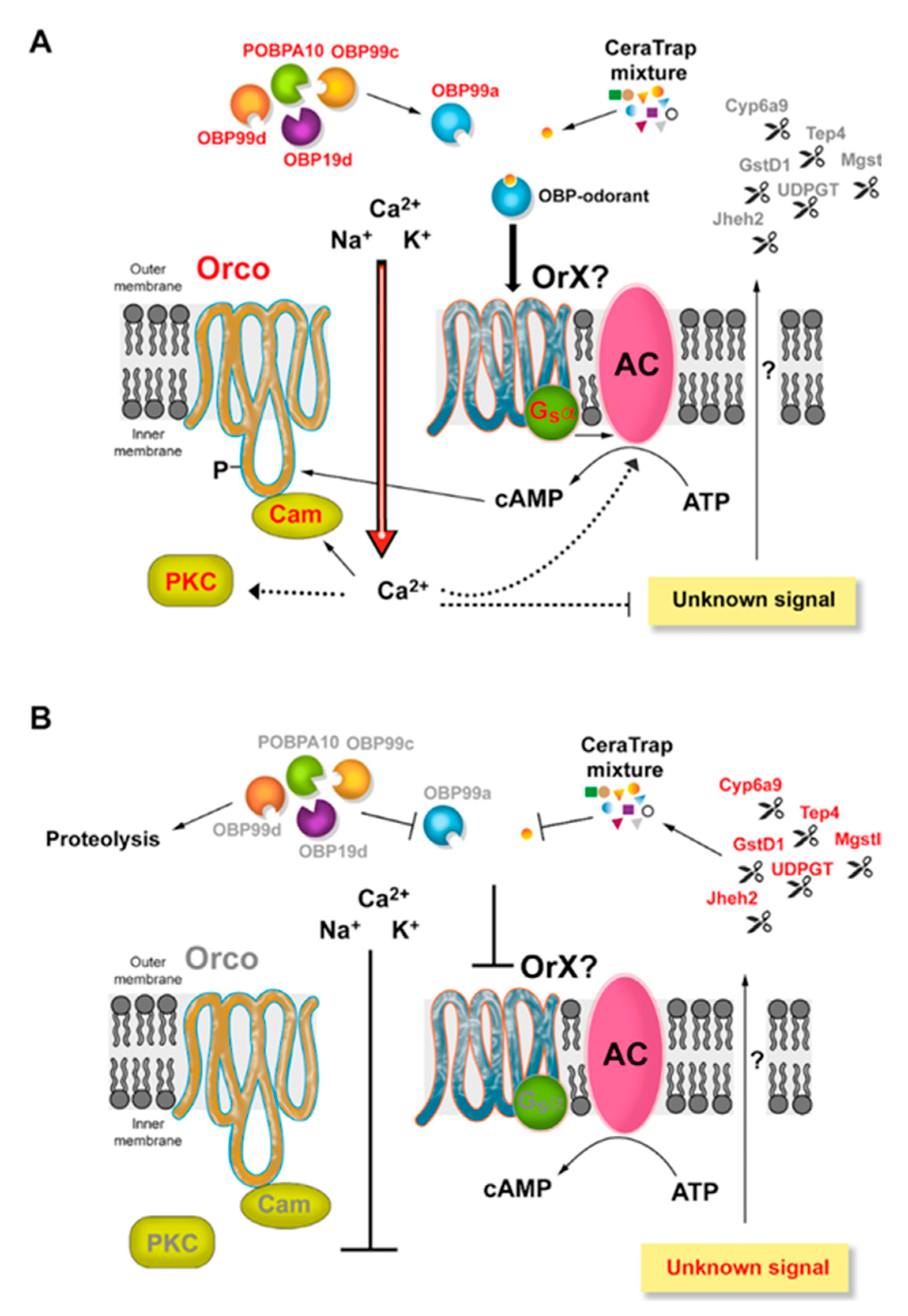
2.3.2. Signal Transduction
2.3.3. Odorant Degrading Enzymes (ODEs)
3. Discussion
3.1. Flushing Out the Secrets of Olfactory Perception in A. ludens via the Characterization of the Antennal Proteome
3.2. CeraTrap® an Efficient Lure that Induces the Upregulation of the Olfactory Machinery of A. ludens
3.3. Integration of Proteomic Data to Help the Establishment of a New Generation of Tools for the Biorational Management of A. ludens
4. Materials and Methods
4.1. Chemical Reagents
4.2. Insect Rearing and Study Site
4.3. Light Microscopy (LM) Studies
4.4. Confocal Scanning Laser Microscopy (CSLM)
4.5. Scanning Electron Microscopy (SEM)
4.6. Anastrepha ludens Exposure to CeraTrap®
4.7. Protein Extraction
4.8. Trypsin Digestion, TMT Labeling, Simple Fractionation and Desalting
4.9. Fractionation of Labeled Peptide Mixture
4.10. Nano-LC-MS/MS Analysis
4.11. Decision Tree-Driven MS/MS
4.12. Synchronous Precursor Selection (SPS)-MS3 for TMT Analysis
4.13. Data Analyses and Interpretation
4.14. Real Time-Quantitative Polymerase Chain Reaction (RT-qPCR) Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| Ar | Arista |
| AOX | Aldehyde oxidases |
| BP | Biological processes |
| Calr | Calreticulin |
| Cam | Calmodulin |
| CaMKII | Calcium/calmodulin-dependent protein kinase II |
| COX | Cytochrome c oxidases |
| Cpo | protein couch potato |
| CSLM | Confocal scanning laser microscopy |
| CSP | Chemosensory proteins |
| Cyp DDT | Cytochromes P450 1-chloro-4-[2,2,2-trichloro-1-(4-chlorophenyl)ethyl]benzene |
| EMR | Ezrin-moesin-radixin |
| EST | Expressed sequence tag |
| Flag | Flagellums |
| GO | Gene ontology |
| GRs | Gustatory receptors |
| Gst | Glutathione S transferases |
| IPM | Integrated pest management |
| IRs | Ionotropic receptors |
| iTRAQ | Isobaric tags for relative and absolute quantitation |
| LM | Light microscopy |
| ME | Methyl eugenol |
| Ncc69 | Sodium chloride cotransporter 69 |
| Nano-LC-MS/MS | Nanoscale liquid chromatography coupled to tandem mass spectrometry |
| OBPs | Odorant-binding proteins |
| ODE | Odorant-degrading enzymes |
| ODPs | Odorant-degrading proteins |
| ORs | Odorant receptors |
| Orco | Odorant receptor co-receptor |
| OSNs | Olfactory sensory neurons |
| PBS | Phosphate buffered saline |
| Ped | Pedicel |
| Pka-R2 | cAMP-dependent, regulatory subunit type 2 |
| PKC | Protein kinase C |
| POBPA | Putative odorant-binding protein |
| Pp2A-29B | Phosphatase 2A at 29B |
| qRT-PCR | Quantitative real-time PCR |
| Rtn | Reticulon-like protein |
| RpS | Ubiquitin-40S ribosomal protein |
| Sc | Scape |
| SCX | Strong cation exchange |
| SDS | Sodium dodecyl sulfate |
| SDS-PAGE | Sodium dodecyl sulfate polyacrylamide gel electrophoresis |
| SEM | Scanning electron microscopy |
| SNMPs | Sensory neuron membrane proteins |
| SPoCk | Secretory Pathway Calcium atpase |
| (SPS)-MS | Synchronous precursor selection |
| TMT | Tandem mass tags |
References
- Pelosi, P.; Lovinella, I.; Felicioli, A.; Dani, F.R. Soluble proteins of chemical communication: An overview across arthropods. Front. Physiol. 2014, 5, 320. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tegoni, M.; Ramoni, R.; Bignetti, E.; Spinelli, S.; Cambillau, C. Domain swapping creates a third putative combining site in bovine odorant binding protein dimer. Nat. Struct. Biol. 1996, 3, 863–867. [Google Scholar] [CrossRef]
- Tegoni, M.; Campanacci, V.; Cambillau, C. Structural aspects of sexual attraction and chemical communication in insects. Trends Biochem. Sci. 2004, 29, 257–264. [Google Scholar] [CrossRef]
- Ali, S.A.I.; Diakite, M.M.; Ali, S.A.I.; Wang, M.Q. Understanding insect behaviors and olfactory signal transduction. J. Genet. Mol. Cell. Biol. 2015, 2, 4. [Google Scholar]
- Leal, W.S. Odorant reception in insects: Roles of receptors, binding proteins, and degrading enzymes. Annu. Rev. Entomol. 2013, 58, 373–391. [Google Scholar] [CrossRef]
- Kim, M.S.; Repp, A.; Smith, D.P. LUSH odorant-binding protein mediates chemosensory responses to alcohols in Drosophila melanogaster. Genetics 1998, 150, 711–721. [Google Scholar]
- Gomulski, L.M.; Dimopoulos, G.; Xi, Z.; Soares, M.B.; Bonaldo, M.F.; Malacrida, A.R.; Gasperi, G. Gene discovery in an invasive tephritid model pest species, the Mediterranean fruit fly, Ceratitis capitata. BMC Genom. 2008, 9, 243. [Google Scholar] [CrossRef] [Green Version]
- Siciliano, P.; Scolari, F.; Gomulski, L.M.; Falchetto, M.; Manni, M.; Gabrieli, P.; Field, L.M.; Zhou, J.-J.; Gasperi, G.; Malacrida, A.R. Sniffing out chemosensory genes from the mediterranean fruit fly, Ceratitis capitata. PLoS ONE 2014, 9, e85523. [Google Scholar] [CrossRef] [Green Version]
- Ramsdell, K.M.M.; Lyons-Sobaski, S.A.; Robertson, H.M.; Walden, K.K.O.; Feder, J.L.; Wanner, K.; Berlocher, S.H. Expressed sequence tags from cephalic chemosensory organs of the northern walnut husk fly, Rhagoletis suavis, including a putative canonical odorant receptor. J. Insect Sci. 2010, 10, 51. [Google Scholar] [CrossRef]
- Schwarz, D.; Robertson, H.M.; Feder, J.L.; Varala, K.; Hudson, M.E.; Ragland, G.J.; Hahn, D.A.; Berlocher, S.H. Sympatric ecological speciation meets pyrosequencing: Sampling the transcriptome of the apple maggot Rhagoletis pomonella. BMC Genom. 2009, 10, 633. [Google Scholar] [CrossRef] [Green Version]
- Elfekih, S.; Chen, C.Y.; Hsu, J.C.; Belcaid, M.; Haymer, D. Identification and preliminary characterization of chemosensory perception-associated proteins in the melon fly Bactrocera cucurbitae using RNA-seq. Sci. Rep. 2016, 6, 19112. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wu, Z.; Zhang, H.; Wang, Z.; Bin, S.; He, H.; Lin, J. Discovery of chemosensory genes in the oriental fruit fly, Bactrocera dorsalis. PLoS ONE 2015, 10, e0129794. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zheng, W.; Peng, W.; Zhu, C.; Zhang, Q.; Saccone, G.; Zhang, H. Identification and expression profile analysis of odorant binding proteins in the oriental fruit fly Bactrocera dorsalis. Int. J. Mol. Sci. 2013, 14, 14936–14949. [Google Scholar] [CrossRef] [Green Version]
- Nakamura, A.M.; Chahad-Ehlers, S.; Lima, A.L.A.; Taniguti, C.H.; Sobrinho, I., Jr.; Torres, F.R.; de Brito, R.A. Reference genes for accessing differential expression among developmental stages and analysis of differential expression of OBP genes in Anastrepha obliqua. Sci. Rep. 2016, 6, 17480. [Google Scholar] [CrossRef]
- Campanini, E.B.; De Brito, R.A. Molecular evolution of Odorant-binding proteins gene family in two closely related Anastrepha fruit flies. BMC Evol. Biol. 2016, 16, 198. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Campanini, E.B.; Congrains, C.; Torres, F.R.; De Brito, R.A. Odorant-binding proteins expression patterns in recently diverged species of Anastrepha fruit flies. Sci Rep. 2017, 7, 2194. [Google Scholar] [CrossRef] [Green Version]
- Aluja, M. Bionomics and management of Anastrepha. Annu. Rev. Entomol. 1994, 39, 155–178. [Google Scholar] [CrossRef]
- Birke, A.; Perez-Staples, D.; Greany, P.; Aluja, M. Interplay between foraging behaviour, adult density and fruit ripeness determines the effectiveness of gibberellic acid and host-marking pheromone in reducing susceptibility of grapefruit to infestation by the Mexican fruit-fly, Anastrepha ludens. Int. J. Pest Manag. 2011, 57, 321–328. [Google Scholar] [CrossRef]
- Aluja, M.; Rull, J. Managing pestiferous fruit flies (Diptera: Tephritidae) through environmental manipulation. In Biorational Tree Fruit Pest Management; Aluja, M., Leskey, T., Vincent, C., Eds.; CABI: Wallingford, UK, 2009; pp. 171–213. [Google Scholar]
- Badii, K.B.; Billah, M.K.; Afreh Nuamah, K.; Obeng Ofori, D.; Nyarko, G. Review of the pest status, economic impact and management of fruit-infesting flies (Diptera: Tephritidae) in Africa. Afr. J. Agric. Res. 2015, 10, 1488–1498. [Google Scholar] [CrossRef] [Green Version]
- Villalobos, J.; Flores, S.; Liedo, P.; Malo, E.A. Mass trapping is as effective as ground bait sprays for the control of Anastrepha (Diptera: Tephritidae) fruit flies in mango orchards. Pest Manag. Sci. 2017, 73, 2105–2110. [Google Scholar] [CrossRef]
- Epsky, N.D.; Kendra, P.E.; Schnell, E.Q. History and development of food-based attractants. In Trapping and the Detection, Control, and Regulation of Tephritid Fruit Flies: Lures, Area-Wide Programs, and Trade Implications; Shelly, T., Epsky, N., Jang, E., Reyes-Flores, J., Vargas, R., Eds.; Springer: Dordrecht, The Netherlands, 2014; pp. 75–118. [Google Scholar]
- Wei, D.; Zhang, M.Y.; Gu, P.M.; Smagghe, G.; Wang, J.J. Label-free based quantitative proteomic analysis identifies proteins involved in the testis maturation of Bactrocera dorsalis (Hendel). Comp. Biochem. Physiol. 2018, 25, 9–18. [Google Scholar] [CrossRef]
- Liu, H.; Zhao, X.F.; Fu, L.; Han, Y.Y.; Chen, J.; Lu, Y.Y. BdorOBP2 plays an indispensable role in the perception of methyl eugenol by mature males of Bactrocera dorsalis (Hendel). Sci. Rep. 2017, 7, 15894. [Google Scholar] [CrossRef] [Green Version]
- Wei, D.; Li, R.; Zhang, M.-Y.; Liu, Y.-W.; Zhang, Z.; Smagghe, G.; Wang, J.-J. Comparative proteomic profiling reveals molecular characteristics associated with oogenesis and oocyte maturation during ovarian development of Bactrocera dorsalis (Hendel). Int. J. Mol. Sci. 2017, 18, 1379. [Google Scholar] [CrossRef] [Green Version]
- Wei, D.; Li, H.M.; Tian, C.B.; Smagghe, G.; Jia, F.X.; Jiang, H.B.; Dou, W.; Wang, J.-J. Proteome analysis of male accessory gland secretions in oriental fruit flies reveals juvenile hormone-binding protein, suggesting impact on female reproduction. Sci. Rep. 2015, 5, 16845. [Google Scholar] [CrossRef] [Green Version]
- Cho, I.K.; Chang, C.L.; Li, Q.X. Diet-induced over-expression of flightless-I protein and its relation to flightlessness in Mediterranean fruit fly, Ceratitis capitata. PLoS ONE 2013, 8, e81099. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jacob, V.; Scolari, F.; Delatte, H.; Gasperi, G.; Jacquin-Joly, E.; Malacrida, A.R.; Duyck, P.-F. Current source density mapping of antennal sensory selectivity reveals conserved olfactory systems between tephritids and Drosophila. Sci. Rep. 2017, 7, 15304. [Google Scholar] [CrossRef] [Green Version]
- Fleischer, F.D.; Aluja, M. Behavior of tephritid flies: A Historical Perspective. In Fruit Flies (Tephritidae): Phylogeny and Evolution of Behavior; Aluja, M., Norrbom, A., Eds.; CRC Press: Boca Raton, FL, USA, 2000; pp. 57–88. [Google Scholar]
- Meats, A.; Leighton, S.M. Protein consumption by mated, unmated, sterile and fertile adults of the Queensland fruit fly, Bactrocera tryoni and its relation to egg production. Physiol. Entomol. 2004, 29, 176–182. [Google Scholar] [CrossRef]
- Aluja, M.; Díaz-Fleischer, F.; Papaj, D.R.; Lagunes, G.; Sivinski, J. Effects of age, diet, female density, and the host resource on egg load in Anastrepha ludens and Anastrepha obliqua (Diptera: Tephritidae). J. Insect Physiol. 2001, 47, 975–988. [Google Scholar] [CrossRef]
- Piñero, J.; Aluja, M.; Equihua, M.; Ojeda, M.M. Feeding history, age and sex influence the response of four economically important Anastrepha species (Diptera: Tephritidae) to human urine and hydrolyzed protein. Folia Entomol. Mex. 2002, 41, 283–298. [Google Scholar]
- Díaz-Fleischer, F.; Arredondo, J.; Flores, S.; Montoya, P.; Aluja, M. There is no magic fruit fly trap: Multiple biological factors influence the response of adult Anastrepha ludens and Anastrepha obliqua (Diptera: Tephritidae) individuals to MultiLure traps baited with BioLure or NuLure. J. Econ. Entomol. 2009, 102, 86–94. [Google Scholar] [CrossRef]
- Siciliano, P.; He, X.L.; Woodcock, C.; Pickett, J.A.; Field, L.M.; Birkett, M.A.; Kalinova, B.; Gomulski, L.M.; Scolari, F.; Gasperi, G.; et al. Identification of pheromone components and their binding affinity to the odorant binding protein CcapOBP83a-2 of the Mediterranean fruit fly, Ceratitis capitata. Insect Biochem. Mol. Biol. 2014, 48, 51–62. [Google Scholar] [CrossRef] [Green Version]
- Zheng, W.; Peng, T.; He, W.; Zhang, H. High-throughput sequencing to reveal genes involved in reproduction and development in Bactrocera dorsalis (Diptera: Tephritidae). PLoS ONE 2012, 7, e36463. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, Z.; Smagghe, G.; Lei, Z.; Wang, J.J. Identification of male- and female-specific olfaction genes in antennae of the oriental fruit fly (Bactrocera dorsalis). PLoS ONE 2016, 11, e0147783. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, J.; Luo, D.; Wu, P.; Li, H.; Zhang, H.; Zheng, W. Identification and expression profiles of novel odorant binding proteins and functional analysis of OBP99a in Bactrocera dorsalis. Arch. Insect Biochem. Physiol. 2018, 98, e21452. [Google Scholar] [CrossRef] [PubMed]
- Lasa, R.; Cruz, A. Efficacy of new commercial traps and the lure Ceratrap® against Anastrepha obliqua (Diptera: Tephritidae). Fla. Entomol. 2014, 97, 1369–1377. [Google Scholar] [CrossRef]
- Lasa, R.; Herrera, F.; Miranda, E.; Gómez, E.; Antonio, S.; Aluja, M. Economic and highly effective trap-lure combination to monitor the Mexican fruit fly (Diptera: Tephritidae) at the orchard level. J. Econ. Entomol. 2015, 108, 1637–1645. [Google Scholar] [CrossRef]
- Rodríguez, C.; Tadeo, E.; Rull, J.; Lasa, R. Response of the sapote fruit fly, Anastrepha serpentina (Diptera: Tephritidae), to commercial lures and trap designs in sapodilla orchards. Fla. Entomol. 2015, 98, 1199–1203. [Google Scholar] [CrossRef]
- Andersson, M.N.; Löfstedt, C.; Newcomb, R.D. Insect olfaction and the evolution of receptor tuning. Front. Ecol. Evol. 2015, 3, 53. [Google Scholar] [CrossRef] [Green Version]
- Aluja, M.; Díaz-Fleischer, F.; Boller, E.F.; Hurter, J.; Edmunds, A.J.F.; Hagmann, L.; Patrian, B.; Reyes, J. Application of feces extracts and synthetic analogues of the host marking pheromone of Anastrepha ludens significantly reduces fruit infestation by A. obliqua in tropical plum and mango backyard orchards. J. Econ. Entomol. 2009, 102, 2268–2278. [Google Scholar] [CrossRef]
- Sayin, S.; Boehm, A.C.; Kobler, J.M.; De Backer, J.-F.; Grunwald Kadow, I.C. Internal state dependent odor processing and perception—The role of neuromodulation in the fly olfactory system. Front. Cell. Neurosci. 2018, 12, 11. [Google Scholar] [CrossRef] [Green Version]
- Brito, N.F.; Moreira, M.F.; Melo, A.C.A. A look inside odorant-binding proteins in insect chemoreception. J. Insect Physiol. 2016, 95, 51–65. [Google Scholar] [CrossRef]
- Vieira, F.G.; Rozas, J. Comparative genomics of the odorant-binding and chemosensory protein gene families across the Arthropoda: Origin and evolutionary history of the chemosensory system. Genome Biol. Evol. 2011, 3, 476–490. [Google Scholar] [CrossRef]
- Larter, N.K.; Sun, J.S.; Carlson, J.R. Organization and function of Drosophila odorant binding proteins. Elife 2016, 5, e20242. [Google Scholar] [CrossRef] [PubMed]
- Vosshall, L.B.; Amrein, H.; Morozov, P.S.; Rzhetsky, A.; Axel, R. A spatial map of olfactory receptor expression in the Drosophila antenna. Cell 1999, 96, 725–736. [Google Scholar] [CrossRef] [Green Version]
- Diaz-Fleischer, F.; Pinero, J.C.P.; Shelly, T.E. Interactions between tephritid fruit fly physiological state and stimuli from baits and traps: Looking for the pied piper of Hamelin to lure pestiferous fruit flies. In Trapping and the Detection, Control, and Regulation of Tephritid Fruit Flies; Shelly, T., Epsky, N., Jang, E.B., Reyes-Flores, J., Vargas, R.I., Eds.; Springer: Dordrecht, The Netherlands, 2014; pp. 145–172. [Google Scholar]
- Pérez-Staples, D.; Aluja, M. Anastrepha striata (Diptera: Tephritidae) females that mate with virgin males live longer. Ann. Entomol. Soc. Am. 2004, 97, 1336–1341. [Google Scholar] [CrossRef] [Green Version]
- Perea-Castellanos, C.; Pérez-Staples, D.; Liedo, P.; Díaz-Fleischer, F. Escape of Mexican fruit flies from traps baited with CeraTrap and effect of lure feeding on reproduction and survival. J. Econ. Entomol. 2015, 108, 1720–1727. [Google Scholar] [CrossRef] [PubMed]
- Aluja, M.; Jácome, I.; Macías-Ordóñez, R. Effect of adult nutrition on male sexual performance in four neotropical fruit fly species of the genus Anastrepha (Diptera: Tephritidae). J. Insect Behav. 2001, 14, 759–775. [Google Scholar] [CrossRef]
- Arredondo, J.; Flores, S.; Montoya, P.; Díaz-Fleischer, F. Effect of multiple endogenous biological factors on the response of the tephritids Anastrepha ludens and Anastrepha obliqua (Diptera: Tephritidae) to multilure traps baited with BioLure or NuLure in mango orchards. J. Econ. Entomol. 2014, 107, 1022–1031. [Google Scholar] [CrossRef] [Green Version]
- Piñero, J.; Aluja, M.; Vázquez, A.; Equihua, M.; Varón, J. Human urine and chicken feces as fruit fly (Diptera: Tephritidae) attractants for resource-poor fruit growers. J. Econ. Entomol. 2003, 96, 334–340. [Google Scholar] [CrossRef]
- Aluja, M.; Pérez-Staples, D.; Sivinski, J.; Sánchez, A.; Piñero, J. Effects of male condition on fitness in two tropical tephritid flies with contrasting life histories. Anim. Behav. 2008, 76, 1997–2009. [Google Scholar] [CrossRef]
- Harwood, J.F.; Chen, K.; Liedo, P.; Müller, H.-G.; Wang, J.-L.; Morice, A.E.; Carey, J.R. Female access and diet affect insemination success, senescence, and the cost of reproduction in male Mexican fruit flies Anastrepha ludens. Physiol. Entomol. 2015, 40, 65–71. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liedo, P.; Orozco, D.; Cruz-López, L.; Quintero, J.L.; Becerra-Pérez, C.; del Refugio Hernández, M.; Oropeza, A.; Toledo, J. Effect of post-teneral diets on the performance of sterile Anastrepha ludens and Anastrepha obliqua fruit flies. J. App. Entomol. 2013, 137, 49–60. [Google Scholar] [CrossRef]
- Taylor, P.W.; Pérez-Staples, D.; Weldon, C.W.; Collins, S.R.; Fanson, B.G.; Yap, S.; Smallridge, C. Post-teneral nutrition as an influence on reproductive development, sexual performance and longevity of Queensland fruit flies. J. App. Entomol. 2013, 137, 113–125. [Google Scholar] [CrossRef]
- Pikielny, C.W.; Hasan, G.; Rouyer, F.; Rosbash, M. Members of a family of Drosophila putative odorant-binding proteins are expressed in different subsets of olfactory hairs. Neuron 1994, 12, 35–49. [Google Scholar] [CrossRef]
- Galindo, K.; Smith, D.P. A large family of divergent Drosophila odorant-binding proteins expressed in gustatory and olfactory sensilla. Genetics 2001, 159, 1059–1072. [Google Scholar]
- Butler, M.J. Discovery of genes with highly restricted expression patterns in the Drosophila wing disc using DNA oligonucleotide microarrays. Development 2003, 130, 659–670. [Google Scholar] [CrossRef] [Green Version]
- Maitra, S.; Dombrowski, S.M.; Waters, L.C.; Ganguly, R. Three second chromosome-linked clustered Cyp6 genes show differential constitutive and barbital-induced expression in DDT-resistant and susceptible strains of Drosophila melanogaster. Gene 1996, 180, 165–171. [Google Scholar] [CrossRef]
- Low, W.Y.; Feil, S.C.; Ng, H.L.; Gorman, M.A.; Morton, C.J.; Pyke, J.; McConville, M.J.; Bieri, M.; Mok, Y.-F.; Robin, C.; et al. Recognition and detoxification of the insecticide DDT by Drosophila melanogaster glutathione S-transferase D1. J. Mol. Biol. 2010, 399, 358–366. [Google Scholar] [CrossRef]
- Wicher, D.; Schäfer, R.; Bauernfeind, R.; Stensmyr, M.C.; Heller, R.; Heinemann, S.H.; Hansson, B. Drosophila odorant receptors are both ligand-gated and cyclic-nucleotide-activated cation channels. Nature 2008, 452, 1007–1011. [Google Scholar] [CrossRef]
- Mukunda, L.; Lavista-Llanos, S.; Hansson, B.S.; Wicher, D. Dimerisation of the Drosophila odorant coreceptor Orco. Front. Cell. Neurosci. 2014, 8, 261. [Google Scholar] [CrossRef] [Green Version]
- Nichols, A.S.; Luetje, C.W. Transmembrane segment 3 of Drosophila melanogaster odorant receptor subunit 85b contributes to ligand-receptor interactions. J. Biol. Chem. 2010, 285, 11854–11862. [Google Scholar] [CrossRef] [Green Version]
- Nichols, A.S.; Chen, S.; Luetje, C.W. Subunit contributions to insect olfactory receptor function: Channel block and odorant recognition. Chem. Sens. 2011, 36, 781–790. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mukunda, L.; Miazzi, F.; Kaltofen, S.; Hansson, B.S.; Wicher, D. Calmodulin modulates insect odorant receptor function. Cell Calcium 2014, 55, 191–199. [Google Scholar] [CrossRef]
- Sargsyan, V.; Getahun, M.N.; Llanos, S.L.; Olsson, S.B.; Hansson, B.S.; Wicher, D. Phosphorylation via PKC regulates the function of the Drosophila odorant co-Receptor. Front. Cell. Neurosci. 2011, 5, 5. [Google Scholar] [CrossRef] [Green Version]
- Wang, Z.; Wilson, G.F.; Griffith, L.C. Calcium/calmodulin-dependent protein kinase II phosphorylates and regulates the Drosophila eag potassium channel. J. Biol. Chem. 2002, 277, 24022–24029. [Google Scholar] [CrossRef] [Green Version]
- Lu, C.S.; Hodge, J.J.L.; Mehren, J.; Sun, X.X.; Griffith, L.C. Regulation of the Ca2+/CaM-responsive pool of CaMKII by scaffold-dependent autophosphorylation. Neuron 2003, 40, 1185–1197. [Google Scholar] [CrossRef] [Green Version]
- Perry, T.; Batterham, P. Harnessing model organisms to study insecticide resistance. Curr. Opin. Insect Sci. 2018, 27, 61–67. [Google Scholar] [CrossRef] [PubMed]
- Beck, J.J.; Light, D.M.; Gee, W.S. Electroantennographic bioassay as a screening tool for host plant volatiles. J. Vis. Exp. 2012, 63, e3931. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gonzalez, F.; Witzgall, P.; Walker, W.B. Protocol for heterologous expression of insect odourant receptors in Drosophila. Front. Ecol. Evol. 2016, 4, 24. [Google Scholar] [CrossRef] [Green Version]
- Backman, A.C.; Bengtsson, M.; Borg-Karlsson, A.K.; Liblikas, I.; Witzgall, P. Volatiles from apple (Malus domestica) eliciting antennal responses in female codling moth Cydia pomonella (L.) (Lepidoptera: Tortricidae): Effect of plant injury and sampling technique. Z. Nat. C 2001, 56, 262–268. [Google Scholar] [CrossRef] [Green Version]
- Ansebo, L.; Coracini, M.; Bengtsson, M.; Liblikas, I.; Ramírez, M.; Borg-Karlson, A.-K.; Tasin, M.; Witzgall, P. Antennal and behavioural response of codling moth Cydia pomonella to plant volatiles. J. App. Entomol. 2004, 128, 488–493. [Google Scholar] [CrossRef]
- Coracini, M.; Bengtsson, M.; Liblikas, L.; Witzgall, P. Attraction of codling moth males to apple volatiles. Entomol. Exp. Appl. 2004, 110, 1–10. [Google Scholar] [CrossRef]
- Dobritsa, A.A.; Van Der Goes Van Naters, W.; Warr, C.G.; Steinbrecht, R.A.; Carlson, J.R. Integrating the molecular and cellular basis of odor coding in the Drosophila antenna. Neuron 2003, 37, 827–841. [Google Scholar] [CrossRef] [Green Version]
- Hallem, E.A.; Ho, M.G.; Carlson, J.R. The molecular basis of odor coding in the Drosophila antenna. Cell 2004, 117, 965–979. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bozzola, J.; Russel, L. Electron Microscopy: Principles and Techniques for Biologists; Jones and Bartlett Publishers: Burlington, MA, USA, 1992. [Google Scholar]
- Puente-Rivera, J.; Villalpando, J.L.; Villalobos-Osnaya, A.; Vázquez-Carrillo, L.I.; León-Ávila, G.; Ponce-Regalado, M.D.; López-Camarillo, C.; Elizalde-Contreras, J.M.; Ruiz-May, E.; Arroyo, R.; et al. The 50 kDa metalloproteinase TvMP50 is a zinc-mediated Trichomonas vaginalis virulence factor. Mol. Biochem. Parasitol. 2017, 217, 32–41. [Google Scholar] [CrossRef] [PubMed]
- Serrano-Maldonado, C.E.; García-Cano, I.; González-Canto, A.; Ruiz-May, E.; Elizalde-Contreras, J.M.; Quirasco, M. Cloning and characterization of a novel N-acetylglucosaminidase (AtlD) from Enterococcus faecalis. J. Mol. Microbiol. Biotechnol. 2018, 28, 14–27. [Google Scholar] [CrossRef]
- Swaney, D.L.; McAlister, G.C.; Coon, J.J. Decision tree-driven tandem mass spectrometry for shotgun proteomics. Nat. Methods 2008, 5, 959–964. [Google Scholar] [CrossRef] [PubMed]
- Frese, C.K.; Altelaar, A.F.M.; Hennrich, M.L.; Nolting, D.; Zeller, M.; Griep-Raming, J.; Heck, A.J.R.; Mohammed, S. Improved peptide identification by targeted fragmentation using CID, HCD and ETD on an LTQ-Orbitrap velos. J. Proteome Res. 2011, 10, 2377–2388. [Google Scholar] [CrossRef]
- McAlister, G.C.; Nusinow, D.P.; Jedrychowski, M.P.; Wühr, M.; Huttlin, E.L.; Erickson, B.K.; Rad, R.; Haas, W.; Gygi, S.P. MultiNotch MS3 enables accurate, sensitive, and multiplexed detection of differential expression across cancer cell line proteomes. Anal. Chem. 2014, 86, 7150–7158. [Google Scholar] [CrossRef]
- Eng, J.K.; McCormack, A.L.; Yates, J.R. An approach to correlate tandem mass spectral data of peptides with amino acid sequences in a protein database. J. Am. Soc. Mass Spectrom. 1994, 5, 976–989. [Google Scholar] [CrossRef] [Green Version]
- Dorfer, V.; Pichler, P.; Stranzl, T.; Stadlmann, J.; Taus, T.; Winkler, S.; Mechtler, K. MS Amanda, a universal identification algorithm optimized for high accuracy tandem mass spectra. J. Proteome Res. 2014, 13, 3679–3684. [Google Scholar] [CrossRef] [PubMed]
- Käll, L.; Canterbury, J.D.; Weston, J.; Noble, W.S.; MacCoss, M.J. Semi-supervised learning for peptide identification from shotgun proteomics datasets. Nat. Methods 2007, 4, 923–925. [Google Scholar] [CrossRef]
- Huang, D.W.; Sherman, B.T.; Lempicki, R.A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 2009, 4, 44–57. [Google Scholar] [CrossRef] [PubMed]
- Huang, D.W.; Sherman, B.T.; Lempicki, R.A. Bioinformatics enrichment tools: Paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 2009, 37, 1–13. [Google Scholar] [CrossRef] [Green Version]
- Supek, F.; Bošnjak, M.; Škunca, N.; Šmuc, T. Revigo summarizes and visualizes long lists of gene ontology terms. PLoS ONE 2011, 6, e21800. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Perez-Riverol, Y.; Csordas, A.; Bai, J.; Bernal-Llinares, M.; Hewapathirana, S.; Kundu, D.J.; Inuganti, A.; Griss, J.; Mayer, G.; Eisenacher, M.; et al. The PRIDE database and related tools and resources in 2019: Improving support for quantification data. Nucleic Acids Res. 2019, 47, D442–D450. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]



| Name. | Search Engine Score | |||
|---|---|---|---|---|
| Coverage | Mascot | Sequest | Amanda | |
| Response to Pheromone (GO:0019236) | ||||
| Odorant-binding protein 99b(Obp99b) | 20.3 | 41.6 | 8.9 | 1005.1 |
| Odorant-binding protein 19a(Obp19a) | 5.4 | 60.4 | 2.2 | 329.1 |
| Odorant-binding protein 99a(Obp99a) | 20.2 | 289.5 | 50.3 | 3044.6 |
| Odorant-binding protein 28a(Obp28a) | 23.9 | 195.2 | 24.6 | 2393.9 |
| General odorant-binding protein Lush (OBP Lush) | 18.9 | 58.2 | 10.5 | 1392.3 |
| Odorant-binding protein 19d(Obp19d) | 8.8 | 261 | 22.4 | 873.6 |
| Odorant-binding protein 99c, isoform B(Obp99c_B) | 11.6 | 128.1 | 10.9 | 1270.6 |
| Odorant-binding protein 83a(Obp83a) | 42.9 | 460.1 | 59.4 | 2532.2 |
| Odorant-binding protein 69a(Obp69a) | 23.9 | 218 | 28.1 | 500.4 |
| Odorant-binding protein 44a(Obp44a) | 22.5 | 158.9 | 16.5 | 2334.1 |
| Putative odorant-binding protein A10 | 51.6 | 586.3 | 87.2 | 7396 |
| Sensory neuron membrane protein 1(Snmp1) | 12.1 | 68.3 | 9.7 | 1222.2 |
| Odorant receptor co-receptor (Orco) | 13.5 | 58.2 | 7.9 | 948.7 |
| Cellular Calcium Ion Homeostasis (GO:0006874) | ||||
| Cation-transporting ATPase | 6.5 | 62.3 | 9.1 | 1085.3 |
| Ryanodine receptor (RyR) | 1.1 | 94.8 | 8.8 | 800.4 |
| upheld(up) | 32.2 | 564.1 | 105.7 | 10436 |
| Plasma membrane calcium ATPase (PMCA) | 14.3 | 349.3 | 68 | 3815.5 |
| Inositol 1,4,5,-tris-phosphate receptor(Itp-r83A) | 4.4 | 182.8 | 19.2 | 1969.8 |
| Calreticulin (Calr) | 18.1 | 185.7 | 44.2 | 4205.3 |
| Calnexin 99A(Cnx99A) | 25.2 | 269 | 49.2 | 5100.4 |
| Calmodulin (Cam) | 79.1 | 513.6 | 73.1 | 5233.8 |
| Calcium/calmodulin-dependent protein kinase II(CaMKII) | 18.8 | 193.5 | 22.4 | 2524 |
| Secretory pathway calcium atpase (SPoCk) | 8.5 | 238.2 | 30.9 | 1887 |
| Calmodulin-binding protein related to a Rab3 GDP/GTP exchange protein (Crag) | 2.9 | 56.6 | 3.2 | 551.3 |
| Calcium-independent phospholipase A2 VIA (iPLA2-VIA) | 8.3 | 116.2 | 18.1 | 1589.9 |
| Putative Odorant-Degrading Enzymes (ODE) | ||||
| Cytochrome c oxidase subunit 5B(COX5B) | 10.8 | 56.3 | 5.4 | 471.5 |
| Cytochrome c oxidase subunit 5A(COX5A) | 16 | 247 | 34 | 2052.4 |
| Cytochrome c oxidase subunit 5A(COX5A) | 16 | 247 | 34 | 2052.4 |
| Cytochrome c oxidase subunit 7A(COX7A) | 9.2 | 25 | 1.9 | 181.7 |
| Glutathione S transferase E7(GstE7) | 26.1 | 241.4 | 39.3 | 3324.3 |
| Juvenile hormone epoxide hydrolase 2(Jheh2) | 22.8 | 189.1 | 32.4 | 2632.6 |
| Thioester-containing protein 4(Tep4) | 16 | 502.5 | 68.1 | 6495.8 |
| Cytochrome P450-4d2(Cyp4d2) | 6.4 | 72.7 | 7.2 | 728.7 |
| Cytochrome P450-9b2(Cyp9b2) | 15.5 | 170.6 | 35 | 3672.4 |
| Cytochrome P450 reductase (Cpr) | 21.2 | 512.2 | 89 | 7761.4 |
| Cytochrome P450-4e1(Cyp4e1) | 27.7 | 230.8 | 40.6 | 3214.5 |
| Cytochrome P450-6a8(Cyp6a8) | 13.8 | 162.4 | 21.2 | 1942.7 |
| Cytochrome P450 6a2 | 14.8 | 40.8 | 2.7 | 216.2 |
| Probable cytochrome P450 6a23 | 7.6 | 119.3 | 11 | 1013.7 |
| Glutathione S transferase S1(GstS1) | 22.5 | 179.7 | 32.6 | 3020.1 |
| Glutathione S transferase E9(GstE9) | 19 | 281.4 | 30.7 | 3201.6 |
| Glutathione S transferase D11(GstD11) | 30.6 | 300.9 | 36.5 | 3926.5 |
| Glutathione S-transferase D1 | 38.1 | 740.2 | 136.1 | 8744.6 |
| Microsomal glutathione S-transferase-like (Mgstl) | 8.4 | 54.22 | 3.3 | 398.9 |
| Alpha-Est10 | 5.4 | 35.5 | 2.3 | 465.8 |
| Thioester-containing protein 2(Tep2) | 3.2 | 48 | 10.4 | 912.2 |
| Thioester-containing protein 2(Tep2) | 18.8 | 877.7 | 180 | 14738.1 |
| Thioester-containing protein 3(Tep3) | 5.4 | 86.9 | 22.1 | 1787.1 |
| Aldehyde dehydrogenase (Aldh) | 28.4 | 358.5 | 61.6 | 4366.4 |
| Aldehyde oxidase 3(AOX3) | 11.4 | 705.6 | 112.3 | 11329.5 |
| UDP-glycosyltransferase 35b(Ugt35b) | 5.8 | 25.5 | 1.9 | 325.1 |
| Aldehyde oxidase 1 (AOX1) | 6.8 | 236.5 | 37.3 | 2716 |
| Proteins | Code | 0D | 5D | 15D | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 2 h | 24 h | 2 h | 24 h | 2 h | 24 h | ||||||||
| F | M | F | M | F | M | F | M | F | M | F | M | ||
| Abundance Ratio:(CeraTrap®)/(Water) | |||||||||||||
| Ubiquitin-40S ribosomal protein S27a (RpS27A) | P15357 | 2.0 | 1.4 | 2.1 | 1.7 | 2.4 | 2.1 | 0 | 1.2 | 0 | 1.1 | 2.2 | 1.2 |
| Reticulon-like protein (Rtn1_c) | Q7KTP4 | 0 | 1.5 | 1.4 | 1.7 | 1.6 | 1.7 | 1.3 | 1.4 | 0 | 1.0 | 1.9 | 1.0 |
| Sodium/potassium-transporting ATPase subunit alpha (Atpalpha) | P13607 | 0.9 | 1.7 | 1.7 | 1.7 | 1.5 | 1.7 | 1.5 | 1.5 | 0.3 | 1.0 | 1.9 | 1.1 |
| PRL-1 phosphatase (PRL-1) | Q95VY8 | 0 | 0 | 1.6 | 1.7 | 1.3 | 1.6 | 1.3 | 1.5 | 0 | 0 | 0 | 1.3 |
| Moesin/ezrin/radixin homolog 1, Ezrin-moesin-radixin 1 (EMR1) | P46150 | 1.8 | 1.4 | 1.6 | 1.6 | 1.5 | 1.7 | 1.4 | 1.4 | 0.6 | 1.3 | 2.1 | 1.1 |
| Juvenile hormone-inducible protein 26 | Q7K0P0 | 0.7 | 0 | 1.4 | 1.6 | 1.3 | 1.4 | 1.5 | 1.3 | 0 | 1.4 | 0 | 1.1 |
| Innexin inx2 (Inx2) | Q9V427 | 1.4 | 0 | 1.5 | 1.6 | 1.3 | 1.6 | 1.3 | 1.4 | 0 | 1.1 | 0 | 0 |
| Voltage-dependent anion-selective channel (Porin) | Q94920 | 1.0 | 1.7 | 1.6 | 1.6 | 1.5 | 1.7 | 1.4 | 1.6 | 0.1 | 1.0 | 3.0 | 1.1 |
| G protein alpha s subunit | P20354 | 0 | 0 | 1.6 | 1.6 | 1.1 | 1.5 | 1.4 | 1.4 | 0 | 1.1 | 0 | 1.6 |
| plasma membrane calcium ATPase (PMCA) | Q59DP9 | 0.7 | 1.3 | 1.4 | 1.6 | 1.3 | 1.6 | 1.5 | 1.3 | 1.2 | 1.2 | 0 | 1.0 |
| Secretory Pathway Calcium atpase (SPoCk) | Q9VNR2 | 0 | 0 | 1.5 | 1.6 | 1.1 | 1.7 | 1.3 | 1.4 | 0 | 1.3 | 0 | 0 |
| Scribbled, isoform S (Scrib) | A0A0B4KHN3 | 0 | 0 | 1.1 | 1.6 | 1.8 | 1.7 | 1.3 | 1.2 | 0 | 0 | 0 | 1.1 |
| Protein lap4 (LAP4) | Q7KRY7 | 0 | 1.3 | 1.7 | 1.6 | 1.0 | 1.4 | 1.3 | 1.4 | 0 | 1.3 | 2.3 | 1.3 |
| Transient receptor potential cation channel protein painless (Pain) | Q9W0Y6 | 2.3 | 1.4 | 1.7 | 1.6 | 1.3 | 1.5 | 1.3 | 1.4 | 0 | 1.2 | 0 | 1.0 |
| C-terminal Binding Protein | A4V2S3 | 0.7 | 0 | 1.4 | 1.6 | 1.2 | 1.4 | 1.5 | 1.3 | 0 | 1.1 | 2.7 | 1.3 |
| NADPH:adrenodoxin oxidoreductase (Dare) | Q9V3T9 | 0 | 0 | 1.5 | 1.6 | 1.2 | 1.4 | 1.4 | 1.3 | 0 | 1.1 | 0 | 1.1 |
| Calreticulin (Calr) | P29413 | 1.8 | 1.3 | 1.7 | 1.6 | 1.4 | 1.5 | 1.5 | 1.2 | 0.6 | 1.4 | 0.8 | 1.1 |
| Protein phosphatase 2A at 29B (Pp2A-29B) | P36179 | 0 | 1.5 | 1.5 | 1.6 | 1.0 | 1.5 | 1.1 | 1.2 | 0 | 1.1 | 0 | 1.4 |
| Sodium chloride cotransporter 69 (Ncc69) | Q9VTW8 | 0.8 | 0 | 1.6 | 1.5 | 1.4 | 1.7 | 1.3 | 1.4 | 0 | 1.1 | 0 | 1.2 |
| Reticulon-like protein (Rtn1) | Q9VMV9 | 1.8 | 1.5 | 1.5 | 1.5 | 1.5 | 1.4 | 1.4 | 1.3 | 0.3 | 1.1 | 0 | 1.0 |
| Reticulon-like protein (Rtn1_b) | E1JHT6 | 0 | 0 | 1.5 | 1.5 | 1.5 | 1.4 | 1.4 | 1.3 | 0.1 | 1.1 | 0 | 1.0 |
| Receptor of activated protein kinase C 1n (PKC) | M9PCC1 | 0.7 | 1.5 | 1.5 | 1.5 | 1.4 | 1.7 | 1.4 | 1.6 | 0 | 1.4 | 0 | 1.0 |
| Protein couch potato (Cpo) | Q01617 | 0 | 0 | 1.6 | 1.5 | 1.4 | 1.5 | 1.3 | 1.4 | 0 | 1.5 | 0 | 1.0 |
| Protein kinase, cAMP-dependent, catalytic subunit 1 | P12370 | 0 | 0 | 0 | 1.5 | 1.2 | 1.4 | 1.3 | 1.4 | 0 | 1.0 | 0 | 1.2 |
| Protein kinase, cAMP-dependent, regulatory subunit type 2 | P81900 | 0 | 1.6 | 1.6 | 1.5 | 1.4 | 1.5 | 1.3 | 0 | 0 | 1.1 | 0 | 1.7 |
| G protein beta-subunit 13F | P26308 | 0 | 1.2 | 1.3 | 1.5 | 1.4 | 1.7 | 1.3 | 0 | 0 | 1.2 | 0 | 1.0 |
| Calcium/calmodulin-dependent protein kinase type II (CaMKII) | Q00168 | 0 | 0 | 1.6 | 1.5 | 1.2 | 1.5 | 1.5 | 1.4 | 0 | 1.1 | 0 | 0 |
| Sensory neuron membrane protein 1 (Snmp1) | Q9VDD3 | 1.6 | 1.4 | 1.8 | 1.5 | 1.1 | 1.5 | 1.4 | 1.5 | 0.9 | 1.3 | 0 | 1.0 |
| Calmodulin | P62152 | 1.0 | 1.5 | 1.5 | 1.4 | 1.2 | 1.5 | 1.4 | 1.2 | 0 | 1.6 | 0 | 1.1 |
| AMP deaminase (AMPdeam) | Q9VY76 | 0 | 0 | 1.4 | 1.4 | 1.3 | 1.6 | 1.4 | 1.4 | 0.4 | 1.0 | 0 | 0 |
| Adenylate kinase 3 | Q9VGU6 | 0 | 0 | 1.3 | 1.4 | 1.2 | 1.3 | 1.2 | 1.3 | 0 | 1.0 | 0 | 1.1 |
| UDP-glucose 6-dehydrogenase | O02373 | 0 | 0 | 1.8 | 0 | 0 | 0 | 0 | 0 | 0 | 1.0 | 0 | 0 |
| Aquaporin | Q9V5Z7 | 0 | 0 | 1.7 | 0 | 1.1 | 0 | 1.3 | 1.4 | 0.1 | 1.0 | 0 | 0 |
| Receptor-type guanylate cyclase | Q7JQ32 | 0 | 0 | 0 | 0 | 1.3 | 0 | 0 | 0 | 0.1 | 0 | 0 | 0 |
| Odorant receptor co-receptor (Orco) | Q9VNB5 | 0.6 | 0 | 1.5 | 1.5 | 1.2 | 1.4 | 1.3 | 1.6 | 0 | 1.4 | 0 | 0 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ruiz-May, E.; Altúzar-Molina, A.; Elizalde-Contreras, J.M.; Arellano-de los Santos, J.; Monribot-Villanueva, J.; Guillén, L.; Vázquez-Rosas-Landa, M.; Ibarra-Laclette, E.; Ramírez-Vázquez, M.; Ortega, R.; et al. A First Glimpse of the Mexican Fruit Fly Anastrepha ludens (Diptera: Tephritidae) Antenna Morphology and Proteome in Response to a Proteinaceous Attractant. Int. J. Mol. Sci. 2020, 21, 8086. https://doi.org/10.3390/ijms21218086
Ruiz-May E, Altúzar-Molina A, Elizalde-Contreras JM, Arellano-de los Santos J, Monribot-Villanueva J, Guillén L, Vázquez-Rosas-Landa M, Ibarra-Laclette E, Ramírez-Vázquez M, Ortega R, et al. A First Glimpse of the Mexican Fruit Fly Anastrepha ludens (Diptera: Tephritidae) Antenna Morphology and Proteome in Response to a Proteinaceous Attractant. International Journal of Molecular Sciences. 2020; 21(21):8086. https://doi.org/10.3390/ijms21218086
Chicago/Turabian StyleRuiz-May, Eliel, Alma Altúzar-Molina, José M. Elizalde-Contreras, Jiovanny Arellano-de los Santos, Juan Monribot-Villanueva, Larissa Guillén, Mirna Vázquez-Rosas-Landa, Enrique Ibarra-Laclette, Mónica Ramírez-Vázquez, Rafael Ortega, and et al. 2020. "A First Glimpse of the Mexican Fruit Fly Anastrepha ludens (Diptera: Tephritidae) Antenna Morphology and Proteome in Response to a Proteinaceous Attractant" International Journal of Molecular Sciences 21, no. 21: 8086. https://doi.org/10.3390/ijms21218086
APA StyleRuiz-May, E., Altúzar-Molina, A., Elizalde-Contreras, J. M., Arellano-de los Santos, J., Monribot-Villanueva, J., Guillén, L., Vázquez-Rosas-Landa, M., Ibarra-Laclette, E., Ramírez-Vázquez, M., Ortega, R., & Aluja, M. (2020). A First Glimpse of the Mexican Fruit Fly Anastrepha ludens (Diptera: Tephritidae) Antenna Morphology and Proteome in Response to a Proteinaceous Attractant. International Journal of Molecular Sciences, 21(21), 8086. https://doi.org/10.3390/ijms21218086





