Formation of HERV-K and HERV-Fc1 Envelope Family Members is Suppressed on Transcriptional and Translational Level
Abstract
1. Introduction
2. Results
2.1. ORFs of HERV-K18, HERV-K113, and HERV-Fc1 form Envelope Proteins
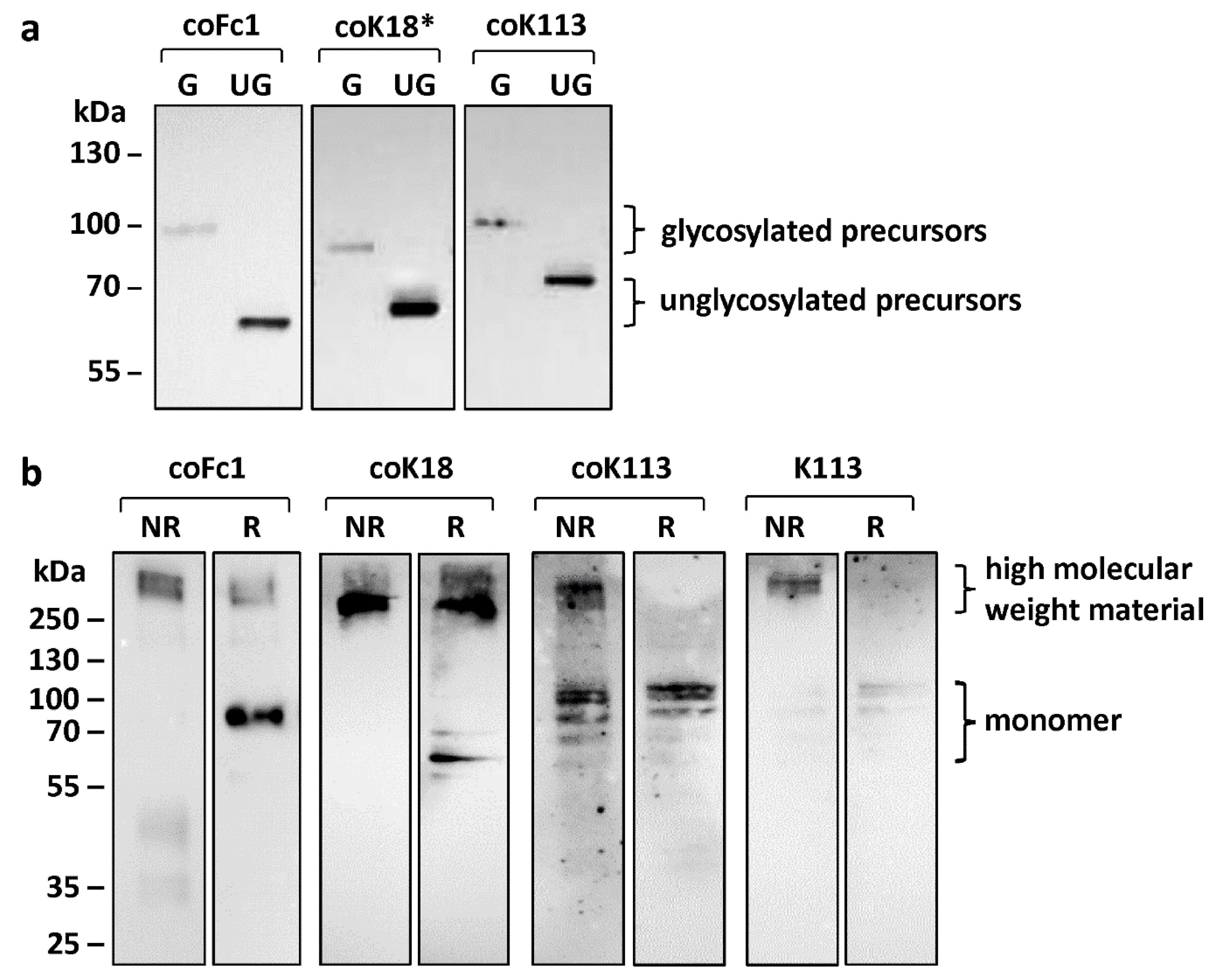
2.2. Analysis of Post-Translational Processing of HERV ENV
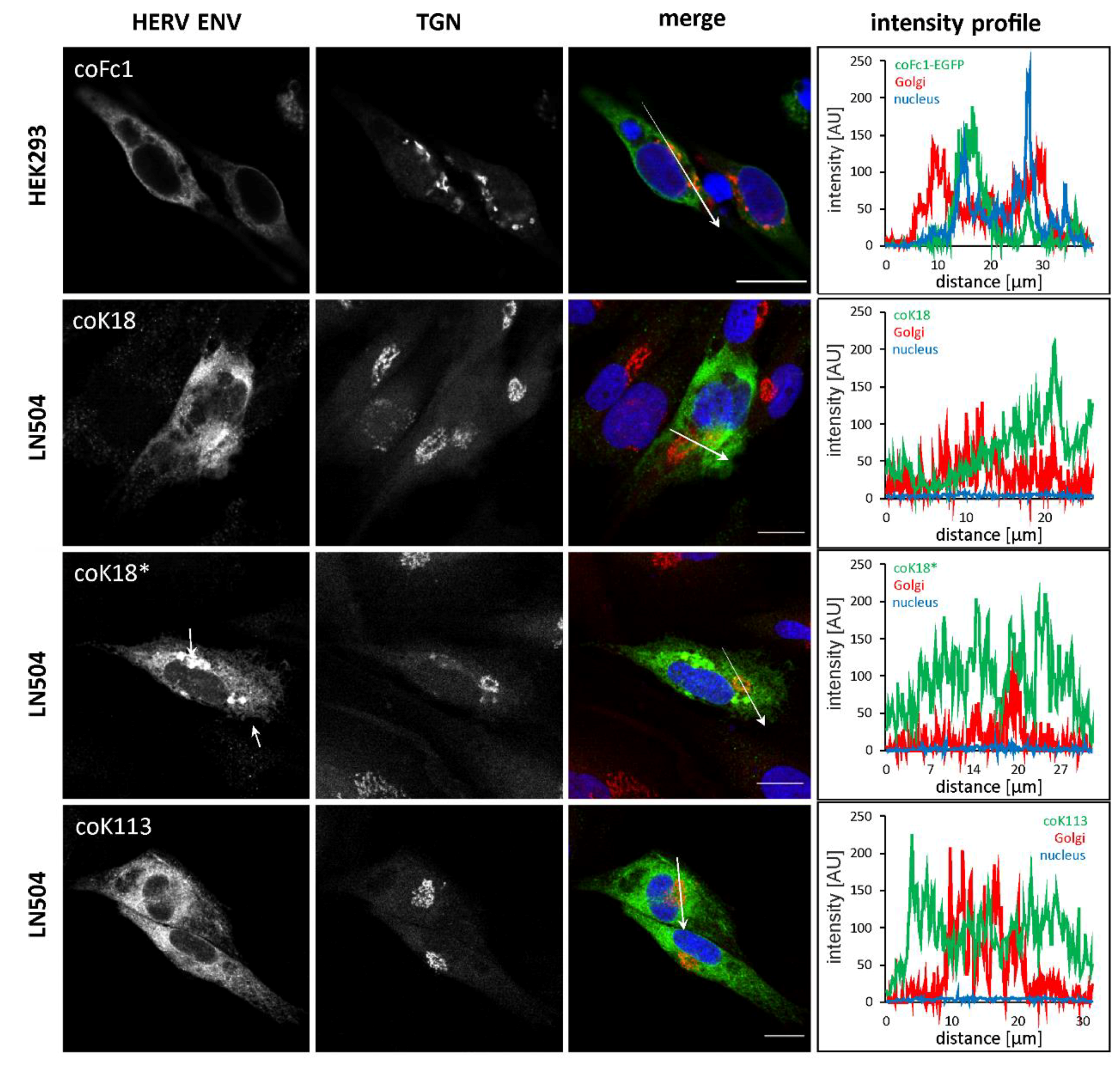
2.3. Analyzed HERV Envelope Proteins are not Transported to the Plasma Membrane but Reside within the Endoplasmic Reticulum
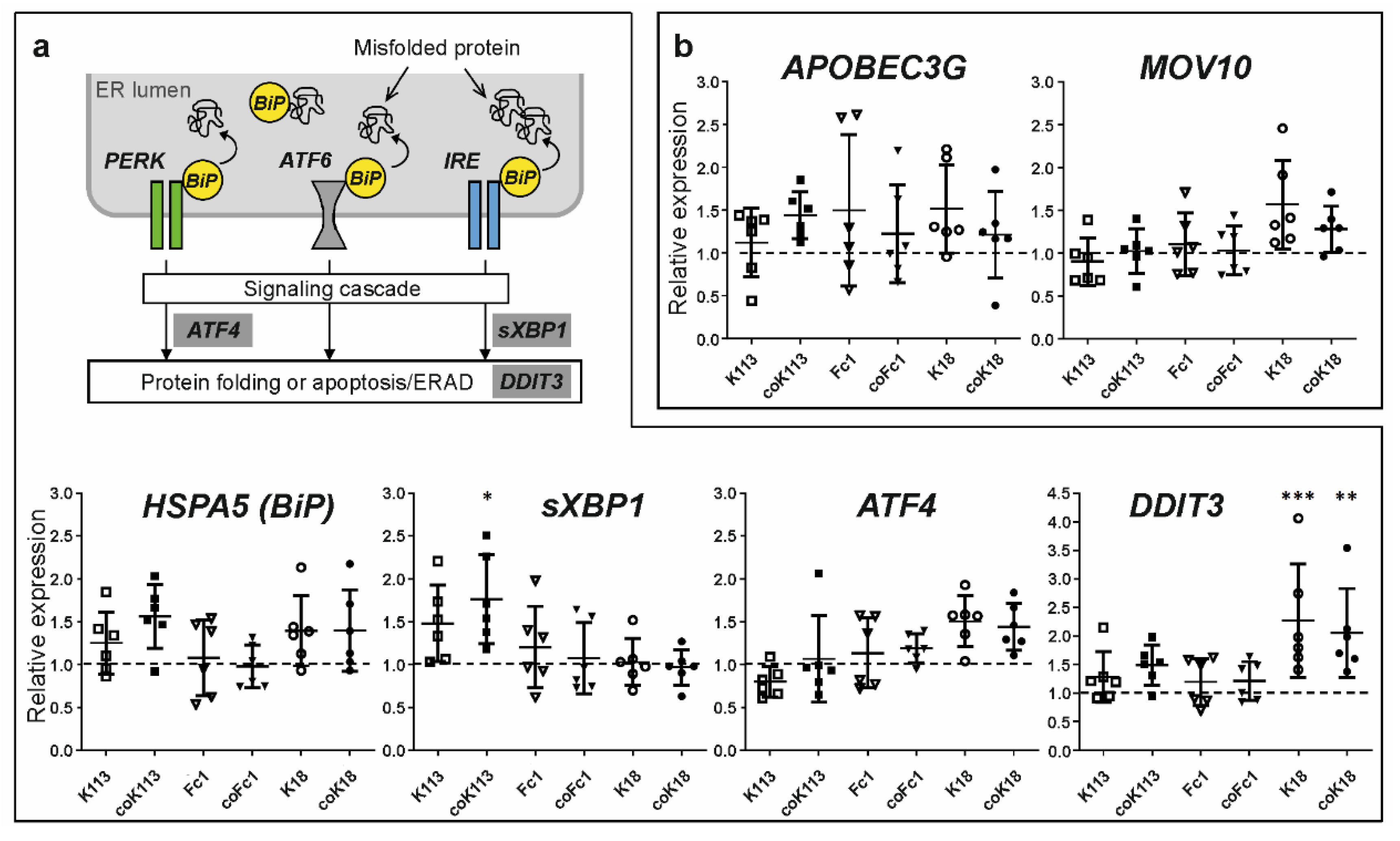
2.4. HERV Envelope Proteins Increase Gene Expression of Markers for Unfolded Protein Response but Trigger no Antiviral Response
2.5. Rare Leucine and Valine Codons Result in Decreased ENV Protein Synthesis in Human Cell Lines
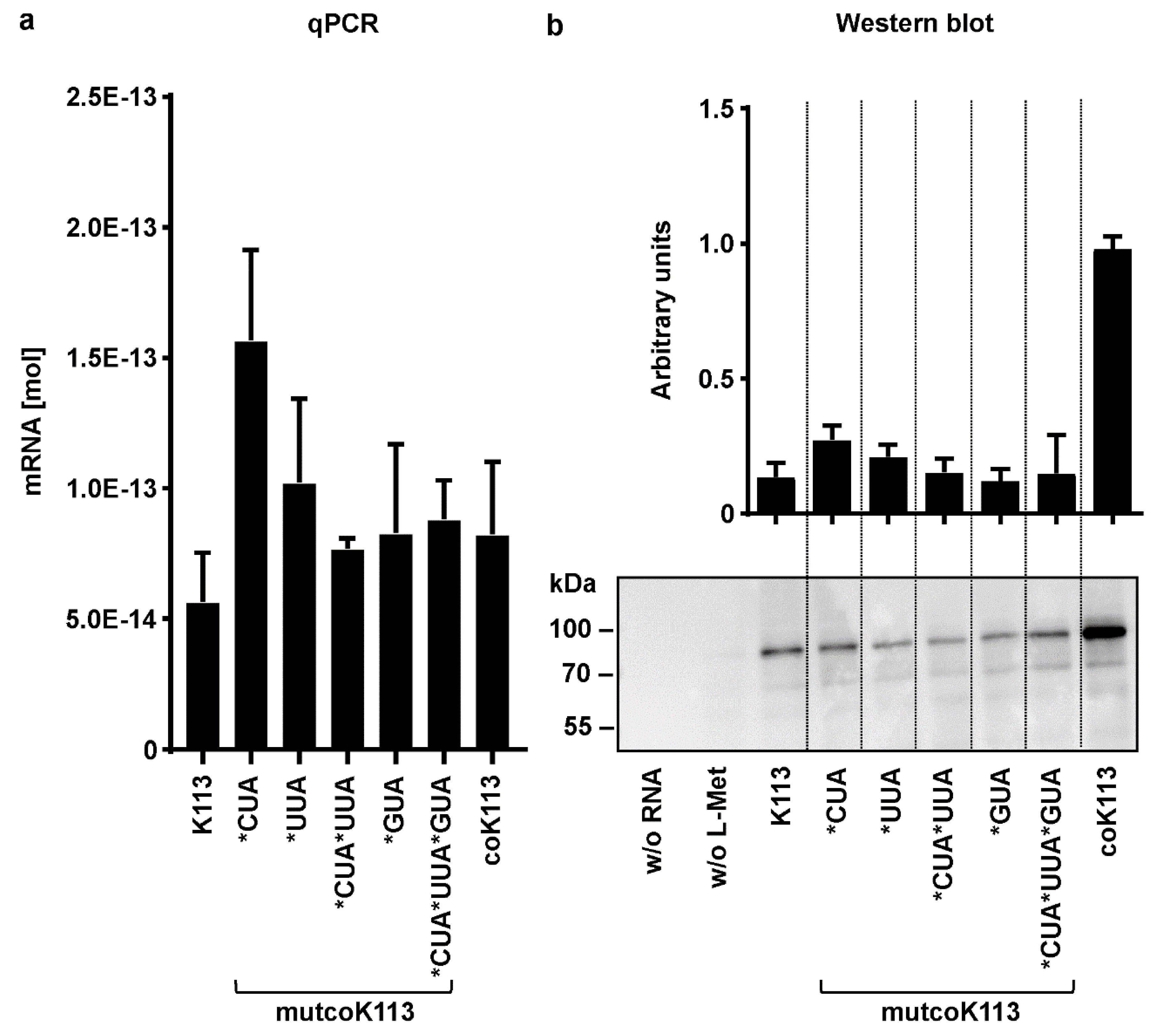
2.6. Rare Codons have a Negative Effect on Expression of HERV-K113 Envelope Protein in a Cell-Free Expression System
2.7. Inefficient Synthesis of Native HERV ENV—A Problem of Secondary RNA Structure?
3. Discussion
4. Materials and Methods
4.1. Generation of Expression Plasmids
4.2. Cell Lines
4.3. Transfection
4.4. Protein Extraction
4.5. Deglycosylation
4.6. SDS-PAGE
4.7. Western Blot Analysis
4.8. Sample Preparation for Mass Spectrometry
4.9. LC-MS/MS Data Acquisition and Analysis
4.10. Flow Cytometry
4.11. Immunocytochemistry
4.12. RNA Extraction and cDNA Generation
4.13. Quantitative Real Time PCR (qRT-PCR)
4.14. Cell-Free In Vitro Transcription and Translation
4.15. In-Silico Determination of mRNA Secondary Structure
4.16. Statistics
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Griffiths, D.J. Endogenous retroviruses in the human genome sequence. Genome Biol. 2001, 2, 1017. [Google Scholar] [CrossRef] [PubMed]
- Belshaw, R.; Pereira, V.; Katzourakis, A.; Talbot, G.; Paces, J.; Burt, A.; Tristem, M. Long-term reinfection of the human genome by endogenous retroviruses. Proc. Natl. Acad. Sci. USA 2004, 101, 4894–4899. [Google Scholar] [CrossRef] [PubMed]
- Christensen, T. HERVs in neuropathogenesis. J. Neuroimmune Pharmacol. 2010, 5, 326–335. [Google Scholar] [CrossRef] [PubMed]
- Gifford, R.J.; Blomberg, J.; Coffin, J.M.; Fan, H.; Heidmann, T.; Mayer, J.; Stoye, J.; Tristem, M.; Johnson, W.E. Nomenclature for endogenous retrovirus (ERV) loci. Retrovirology 2018, 15, 59. [Google Scholar] [CrossRef]
- de Parseval, N.; Heidmann, T. Human endogenous retroviruses: From infectious elements to human genes. Cytogenet. Genome Res. 2005, 110, 318–332. [Google Scholar] [CrossRef]
- Blond, J.L.; Lavillette, D.; Cheynet, V.; Bouton, O.; Oriol, G.; Chapel-Fernandes, S.; Mandrand, B.; Mallet, F.; Cosset, F.L. An envelope glycoprotein of the human endogenous retrovirus HERV-W is expressed in the human placenta and fuses cells expressing the type D mammalian retrovirus receptor. J. Virol. 2000, 74, 3321–3329. [Google Scholar] [CrossRef]
- Grandi, N.; Tramontano, E. HERV envelope proteins: Physiological role and pathogenic potential in cancer and autoimmunity. Front. Microbiol. 2018, 9, 462. [Google Scholar] [CrossRef]
- Balada, E.; Vilardell-Tarrés, M.; Ordi-Ros, J. Implication of human endogenous retroviruses in the development of autoimmune diseases. Int. Rev. Immunol. 2010, 29, 351–370. [Google Scholar] [CrossRef]
- Lemaître, C.; Tsang, J.; Bireau, C.; Heidmann, T.; Dewannieux, M. A human endogenous retrovirus-derived gene that can contribute to oncogenesis by activating the ERK pathway and inducing migration and invasion. PLoS Pathog. 2017, 13, e1006451. [Google Scholar] [CrossRef] [PubMed]
- Gröger, V.; Cynis, H. Human endogenous retroviruses and their putative role in the development of autoimmune disorders Such as multiple sclerosis. Front. Microbiol. 2018, 9, 265. [Google Scholar] [CrossRef] [PubMed]
- Staege, M.S.; Emmer, A. Editorial: Endogenous viral elements-links between autoimmunity and cancer? Front. Microbiol. 2018, 9, 3171. [Google Scholar] [CrossRef] [PubMed]
- Grabski, D.F.; Hu, Y.; Sharma, M.; Rasmussen, S.K. Close to the bedside: A systematic review of endogenous retroviruses and their impact in oncology. J. Surg. Res. 2019, 240, 145–155. [Google Scholar] [CrossRef] [PubMed]
- Herve, C.A.; Lugli, E.B.; Brand, A.; Griffiths, D.J.; Venables, P.J.W. Autoantibodies to human endogenous retrovirus-K are frequently detected in health and disease and react with multiple epitopes. Clin. Exp. Immunol. 2002, 128, 75–82. [Google Scholar] [CrossRef] [PubMed]
- Mameli, G.; Erre, G.L.; Caggiu, E.; Mura, S.; Cossu, D.; Bo, M.; Cadoni, M.L.; Piras, A.; Mundula, N.; Colombo, E.; et al. Identification of a HERV-K env surface peptide highly recognized in rheumatoid arthritis (RA) patients: A cross-sectional case-control study. Clin. Exp. Immunol. 2017, 189, 127–131. [Google Scholar] [CrossRef]
- Giebler, M.; Staege, M.S.; Blauschmidt, S.; Ohm, L.I.; Kraus, M.; Würl, P.; Taubert, H.; Greither, T. Elevated HERV-K expression in soft tissue sarcoma is associated with worsened relapse-free survival. Front. Microbiol. 2018, 9, 211. [Google Scholar] [CrossRef]
- Johnston, J.B.; Silva, C.; Holden, J.; Warren, K.G.; Clark, A.W.; Power, C. Monocyte activation and differentiation augment human endogenous retrovirus expression: Implications for inflammatory brain diseases. Ann. Neurol. 2001, 50, 434–442. [Google Scholar] [CrossRef]
- Moyes, D.L.; Martin, A.; Sawcer, S.; Temperton, N.; Worthington, J.; Griffiths, D.J.; Venables, P.J. The distribution of the endogenous retroviruses HERVK113 and HERV-K115 in health and disease. Genomics 2005, 86, 337–341. [Google Scholar] [CrossRef]
- de la Hera, B.; Varadé, J.; García-Montojo, M.; Lamas, J.R.; de la Encarnación, A.; Arroyo, R.; Fernández-Gutiérrez, B.; Alvarez-Lafuente, R.; Urcelay, E. Role of the human endogenous retrovirus HERV-K18 in autoimmune disease susceptibility: Study in the Spanish population and meta-analysis. PLoS ONE 2013, 8, e62090. [Google Scholar] [CrossRef]
- Nexø, B.A.; Villesen, P.; Nissen, K.K.; Lindegaard, H.M.; Rossing, P.; Petersen, T.; Tarnow, L.; Hansen, B.; Lorenzen, T.; Hørslev-Petersen, K.; et al. Are human endogenous retroviruses triggers of autoimmune diseases? Unveiling associations of three diseases and viral loci. Immunol. Res. 2016, 64, 55–63. [Google Scholar] [CrossRef]
- Garcia-Montojo, M.; Doucet-O’Hare, T.; Henderson, L.; Nath, A. Human endogenous retrovirus-K (HML-2): A comprehensive review. Crit. Rev. Microbiol. 2018, 44, 715–738. [Google Scholar] [CrossRef]
- Buzdin, A.; Ustyugova, S.; Khodosevich, K.; Mamedov, I.; Lebedev, Y.; Hunsmann, G.; Sverdlov, E. Human-specific subfamilies of HERV-K (HML-2) long terminal repeats: Three master genes were active simultaneously during branching of hominoid lineages. Genomics 2003, 81, 149–156. [Google Scholar] [CrossRef]
- Wildschutte, J.H.; Williams, Z.H.; Montesion, M.; Subramanian, R.P.; Kidd, J.M.; Coffin, J.M. Discovery of unfixed endogenous retrovirus insertions in diverse human populations. Proc. Natl. Acad. Sci. USA 2016, 113, E2326–E2334. [Google Scholar] [CrossRef] [PubMed]
- Tönjes, R.R.; Czauderna, F.; Kurth, R. Genome-wide screening, cloning, chromosomal assignment, and expression of full-length human endogenous retrovirus type K. J. Virol. 1999, 73, 9187–9195. [Google Scholar] [CrossRef] [PubMed]
- Stauffer, Y.; Marguerat, S.; Meylan, F.; Ucla, C.; Sutkowski, N.; Huber, B.; Pelet, T.; Conrad, B. Interferon-alpha-induced endogenous superantigen. a model linking environment and autoimmunity. Immunity 2001, 15, 591–601. [Google Scholar] [CrossRef]
- Sutkowski, N.; Conrad, B.; Thorley-Lawson, D.A.; Huber, B.T. Epstein-Barr virus transactivates the human endogenous retrovirus HERV-K18 that encodes a superantigen. Immunity 2001, 15, 579–589. [Google Scholar] [CrossRef]
- Thorley-Lawson, D.A.; Schooley, R.T.; Bhan, A.K.; Nadler, L.M. Epstein-Barr virus superinduces a new human B cell differentiation antigen (B-LAST 1) expressed on transformed lymphoblasts. Cell 1982, 30, 415–425. [Google Scholar] [CrossRef]
- Barth, M.; Gröger, V.; Cynis, H.; Staege, M.S. Identification of human endogenous retrovirus transcripts in Hodgekin Lymphoma cells. Mol. Biol. Rep. 2019, 46, 1885. [Google Scholar] [CrossRef]
- Tai, A.K.; O’Reilly, E.J.; Alroy, K.A.; Munger, K.L.; Huber, B.T.; Ascherio, A. Human endogenous retrovirus-K18 Env as a risk factor in multiple sclerosis. Mult. Scler. 2008, 14, 1175–1180. [Google Scholar] [CrossRef] [PubMed]
- Beimforde, N.; Hanke, K.; Ammar, I.; Kurth, R.; Bannert, N. Molecular cloning and functional characterization of the human endogenous retrovirus K113. Virology 2008, 371, 216–225. [Google Scholar] [CrossRef]
- Turner, G.; Barbulescu, M.; Su, M.; Jensen-Seaman, M.I.; Kidd, K.K.; Lenz, J. Insertional polymorphisms of full-length endogenous retroviruses in humans. Curr. Biol. 2001, 11, 1531–1535. [Google Scholar] [CrossRef]
- Jern, P.; Sperber, G.O.; Blomberg, J. Definition and variation of human endogenous retrovirus H. Virology 2004, 327, 93–110. [Google Scholar] [CrossRef]
- Jern, P.; Sperber, G.O.; Blomberg, J. Use of endogenous retroviral sequences (ERVs) and structural markers for retroviral phylogenetic interference and taxonomy. Retrovirology 2005, 2, 50. [Google Scholar] [CrossRef] [PubMed]
- Hansen, D.T.; Petersen, T.; Christensen, T. Retroviral envelope proteins: Involvement in neuropathogenesis. J. Neurol. Sci. 2017, 380, 151–163. [Google Scholar] [CrossRef]
- Laska, M.J.; Brudek, T.; Nissen, K.K.; Christensen, T.; Møller-Larsen, A.; Petersen, T.; Nexø, B.A. Expression of HERV-Fc1, a human endogenous retrovirus, is increased in patients with active multiple sclerosis. J. Virol. 2012, 86, 3713–3722. [Google Scholar] [CrossRef] [PubMed]
- Benit, L.; Calteau, A.; Heidmann, T. Characterization of the low-copy HERV-Fc family: Evidence for recent integrations in primates of elements with coding envelope genes. Virology 2003, 312, 159–168. [Google Scholar] [CrossRef][Green Version]
- Hanke, K.; Kramer, P.; Seeher, S.; Beimforde, N.; Kurth, R.; Bannert, N. Reconstitution of the ancestral glycoprotein of human endogenous retrovirus k and modulation of its functional activity by truncation of the cytoplasmic domain. J. Virol. 2009, 83, 12790–12800. [Google Scholar] [CrossRef]
- Heinrich, S.; Cameron, A.; Bourenkov, G.P.; Kiefersauer, R.; Huber, R.; Lindberg, I.; Bode, W.; Than, M.E. The crystal structure of the proprotein processing proteinase furin explains its stringent specificity. Nat. Struct. Mol. Biol. 2003, 10, 520–526. [Google Scholar] [CrossRef] [PubMed]
- Apte, S.; Sanders, D.A. Effects of retroviral envelope-protein cleavage upon trafficking, incorporation, and membrane fusion. Virology 2010, 405, 214–224. [Google Scholar] [CrossRef] [PubMed]
- Dewannieux, M.; Blaise, S.; Heidmann, T. Identification of a functional envelope protein from the HERV-K family of human endogenous retroviruses. J. Virol. 2005, 79, 15573–15577. [Google Scholar] [CrossRef] [PubMed]
- Michaud, H.-A.; de Mulder, M.; SenGupta, D.; Deeks, S.G.; Martin, J.N.; Pilcher, C.D.; Hecht, F.M.; Sacha, J.B.; Nixon, D.F. Trans-activation, post-transcriptional maturation, and induction of antibodies to HERV-K (HML-2) envelope transmembrane protein in HIV-1 infection. Retrovirology 2014, 11, 10. [Google Scholar] [CrossRef] [PubMed]
- Hanson, G.; Coller, J. Codon optimality, bias and usage in translation and mRNA decay. Nat. Rev. Mol. Cell. Biol. 2018, 19, 20–30. [Google Scholar] [CrossRef]
- Nakaya, Y.; Takayuki, M. Dysfunction of bovine endogenous retrovirus K2 envelope glycoprotein is related to unsuccessful intracellular trafficking. J. Virol. 2014, 88, 6869–6905. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.J.; Pan, H.Y.; Gao, S.J. Reverse transcription slippage over the mRNA secondary structure of the LIP1 gene. Biotechniques 2001, 31, 1286–1294. [Google Scholar] [CrossRef]
- Forrest, M.E.; Pinkard, O.; Martin, S.; Sweet, T.J.; Hanson, G.; Coller, J. Codon and amino acid content are associated with mRNA stability in mammalian cells. PLoS ONE 2020, 15, e0228730. [Google Scholar] [CrossRef]
- Torrent, M.; Chalancon, G.; de Groot, N.S.; Wuster, A.; Babu, M.M. Cells alter their TRNA abundance to selectively regulate protein synthesis during stress conditions. Sci. Signal. 2018, 11, eaat6409. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Z.; Dang, Y.; Zhou, M.; Li, L.; Yu, C.H.; Fu, J.; Chen, S.; Liu, Y. Codon usage is an important determinant of gene expression levels largely through its effect on transcription. Proc. Natl. Acad. Sci. USA 2016, 113, E6117–E6125. [Google Scholar] [CrossRef] [PubMed]
- Fu, J.; Dang, Y.; Counter, C.; Liu, Y. Codon usage regulates human KRAS expression at both transcriptional and translational levels. J. Biol. Chem. 2018, 293, 17929–17940. [Google Scholar] [CrossRef] [PubMed]
- Tsai, K.; Jaguva Vasudevan, A.A.; Martinez Campos, C.; Emery, A.; Swanstrom, R.; Cullen, B.R. Acetylation of cytidine residues boosts HIV-1 gene expression by increasing viral RNA stability. Cell Host Microbe. 2020, 28, 306–312. [Google Scholar] [CrossRef]
- Tsai, K.; Cullen, B.R. Epigenetic and epitranscriptomic regulation of viral replication. Nat. Rev. Microbiol. 2020, 18, 559–570. [Google Scholar] [CrossRef]
- Caspar, J.; Schmoll, H.J.; Schnaidt, U.; Fonatsch, C. Cell lines of human germinal cancer. Int. J. Androl. 1987, 10, 105–113. [Google Scholar] [CrossRef]
- Graham, F.L.; Smiley, J.; Russell, W.C.; Nairn, R.J. Characteristics of a human cell line transformed by DNA from human adenovirus type 5. Gen. Virol. 1977, 36, 59–74. [Google Scholar] [CrossRef] [PubMed]
- Gluzman, Y. SV40-transformed simian cells support the replication of early SV40 mutants. Cell 1981, 23, 175–182. [Google Scholar] [CrossRef]
- Bodmer, S.; Strommer, K.; Frei, K.; Siepl, C.; de Tribolet, N.; Heid, I.; Fontana, A. Immunosuppression and transforming growth factor-beta in glioblastoma. Preferential production of transforming growth factor-beta 2. J. Immunol. 1989, 143, 3222–3229. [Google Scholar] [PubMed]
- Giard, D.J.; Aaronson, S.A.; Todaro, G.J.; Arnstein, P.; Kersey, J.H.; Dosik, H.; Parks, W.P. In vitro cultivation of human tumors: Establishment of cell lines derived from a series of solid tumors. J. Natl. Cancer. Inst. 1973, 51, 1417–1423. [Google Scholar] [CrossRef] [PubMed]
- Laemmli, U.K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 1970, 227, 680–685. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Zuker, M. Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res. 2003, 31, 3406–3415. [Google Scholar] [CrossRef]




| Sample | Protein | Sequence Coverage (%) | Proteolytic Peptides # |
|---|---|---|---|
| Control | K113, Fc1 envelope proteins | n.d. | n.d. |
| ER chaperone BiP | 20 | 7 | |
| K113 | K113 envelope protein | 4 | 3 |
| ER chaperone BiP | 47 | 31 | |
| coK113 | K113 envelope protein | 20 | 10 |
| ER chaperone BiP | 48 | 27 | |
| Fc1 | Fc1 envelope protein | n.d. | n.d. |
| ER chaperone BiP | 53 | 36 | |
| coFc1 | Fc1 envelope protein | 39 | 25 |
| ER chaperone BiP | 54 | 38 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gröger, V.; Wieland, L.; Naumann, M.; Meinecke, A.-C.; Meinhardt, B.; Rossner, S.; Ihling, C.; Emmer, A.; Staege, M.S.; Cynis, H. Formation of HERV-K and HERV-Fc1 Envelope Family Members is Suppressed on Transcriptional and Translational Level. Int. J. Mol. Sci. 2020, 21, 7855. https://doi.org/10.3390/ijms21217855
Gröger V, Wieland L, Naumann M, Meinecke A-C, Meinhardt B, Rossner S, Ihling C, Emmer A, Staege MS, Cynis H. Formation of HERV-K and HERV-Fc1 Envelope Family Members is Suppressed on Transcriptional and Translational Level. International Journal of Molecular Sciences. 2020; 21(21):7855. https://doi.org/10.3390/ijms21217855
Chicago/Turabian StyleGröger, Victoria, Lisa Wieland, Marcel Naumann, Ann-Christin Meinecke, Beate Meinhardt, Steffen Rossner, Christian Ihling, Alexander Emmer, Martin S. Staege, and Holger Cynis. 2020. "Formation of HERV-K and HERV-Fc1 Envelope Family Members is Suppressed on Transcriptional and Translational Level" International Journal of Molecular Sciences 21, no. 21: 7855. https://doi.org/10.3390/ijms21217855
APA StyleGröger, V., Wieland, L., Naumann, M., Meinecke, A.-C., Meinhardt, B., Rossner, S., Ihling, C., Emmer, A., Staege, M. S., & Cynis, H. (2020). Formation of HERV-K and HERV-Fc1 Envelope Family Members is Suppressed on Transcriptional and Translational Level. International Journal of Molecular Sciences, 21(21), 7855. https://doi.org/10.3390/ijms21217855







