RedquorinXS Mutants with Enhanced Calcium Sensitivity and Bioluminescence Output Efficiently Report Cellular and Neuronal Network Activities
Abstract
:1. Introduction
2. Result
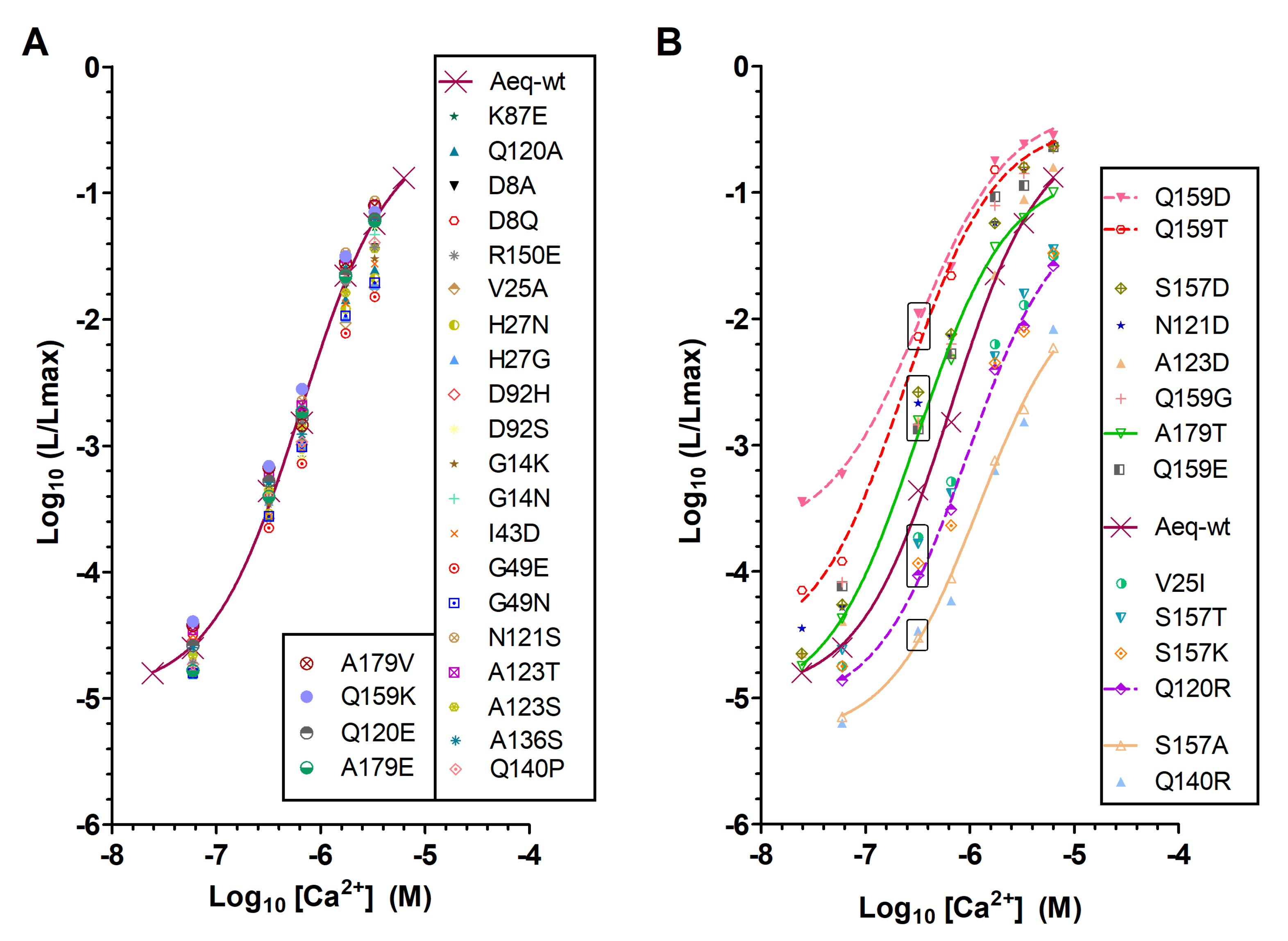
2.1. Identification of New Mutations That Enhance Aequorin Ca2+ Sensitivity
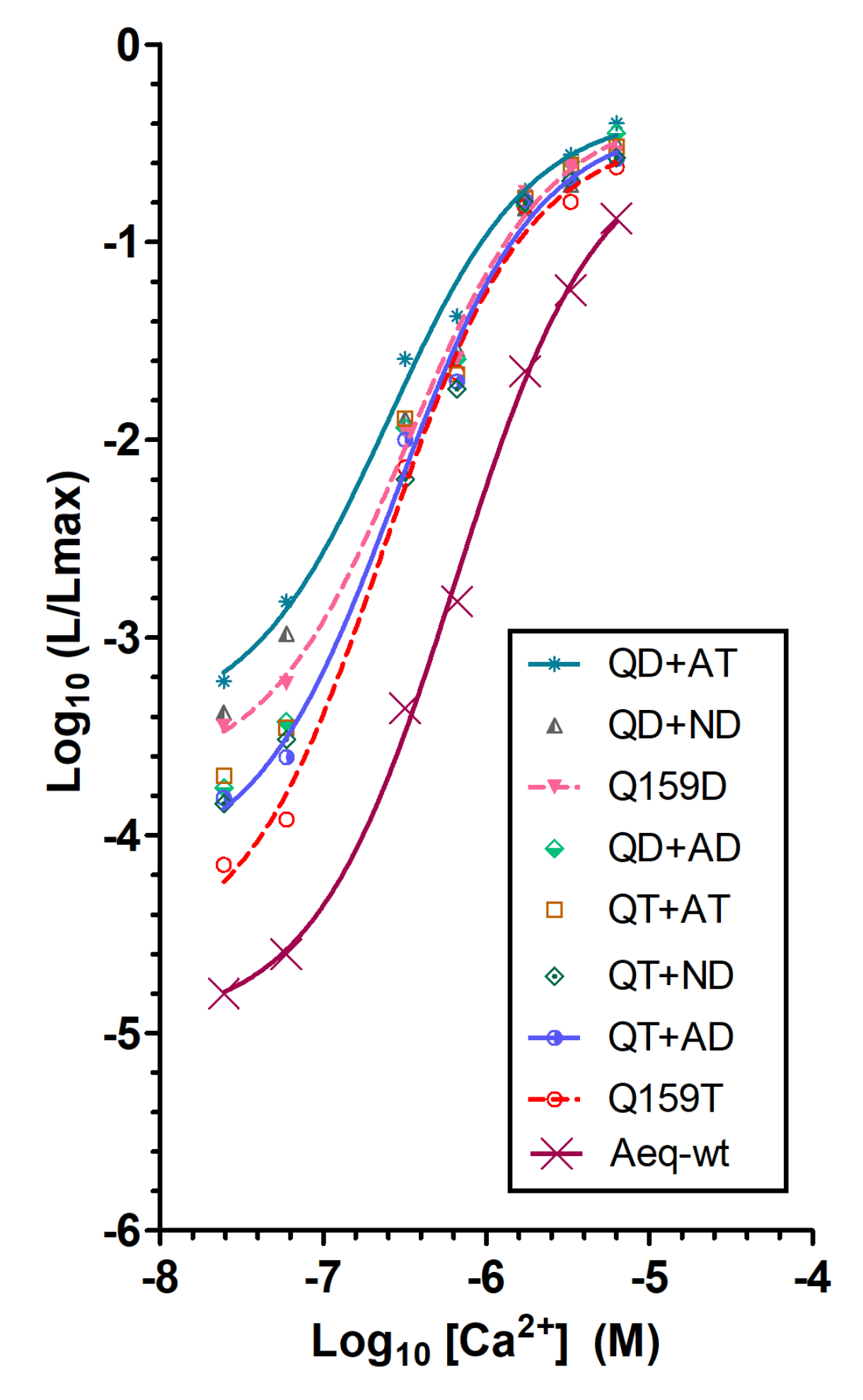
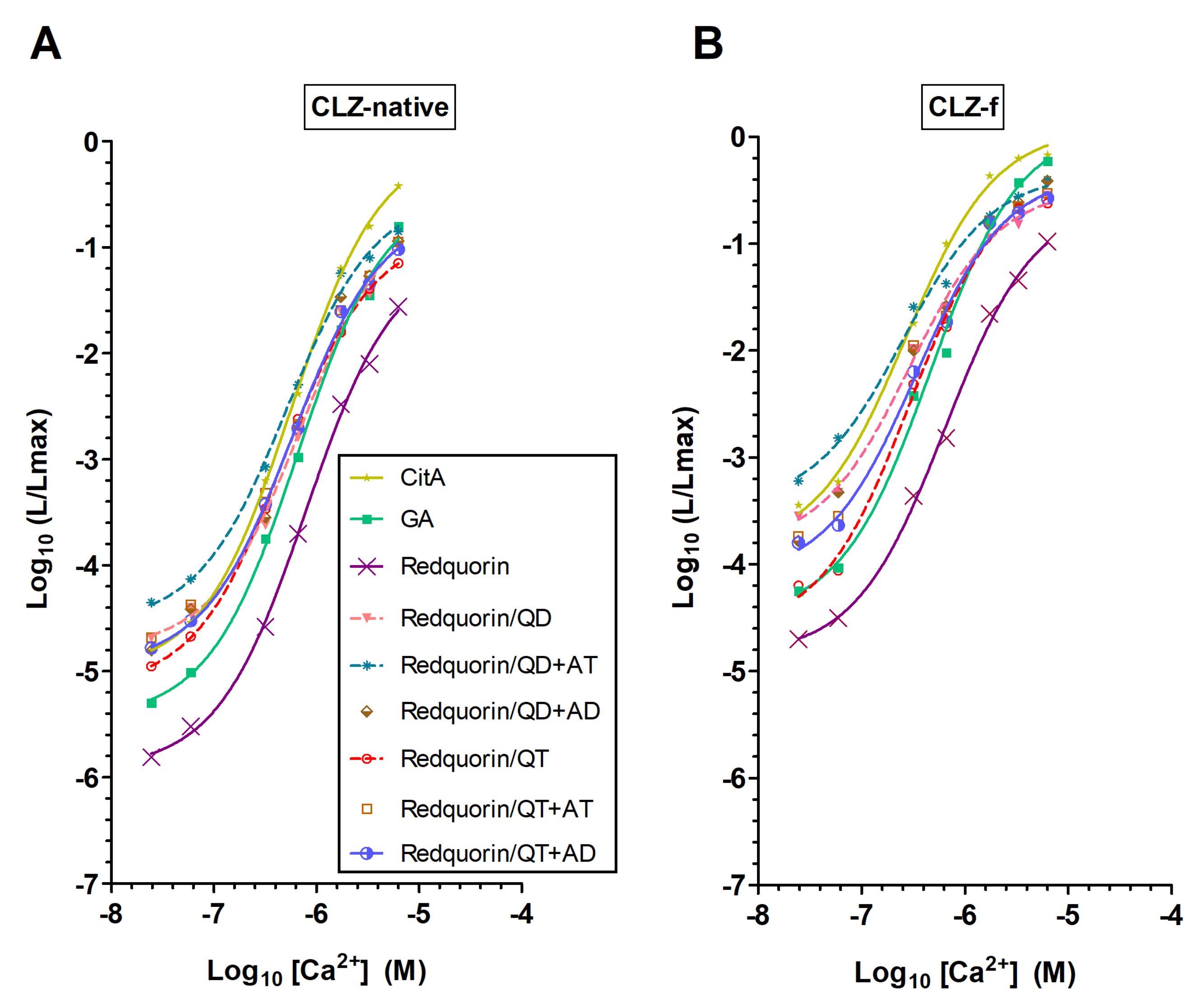
2.2. Generating Redquorin Mutants with Ca2+ Sensitivity and Light Emission Rate That Match Those of GA
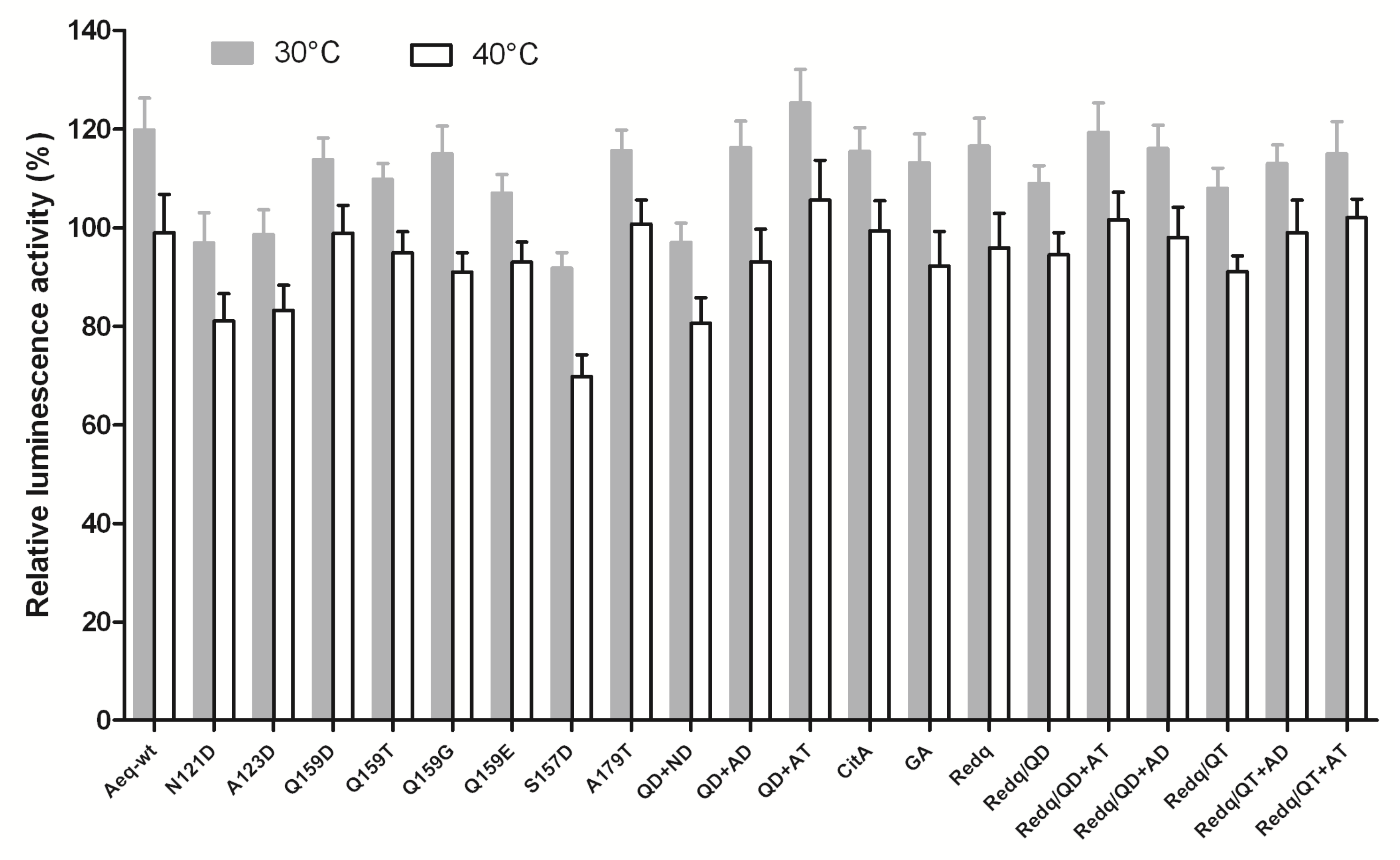
2.3. Effects of Key Amino Acid Mutations on Redquorin Thermostability
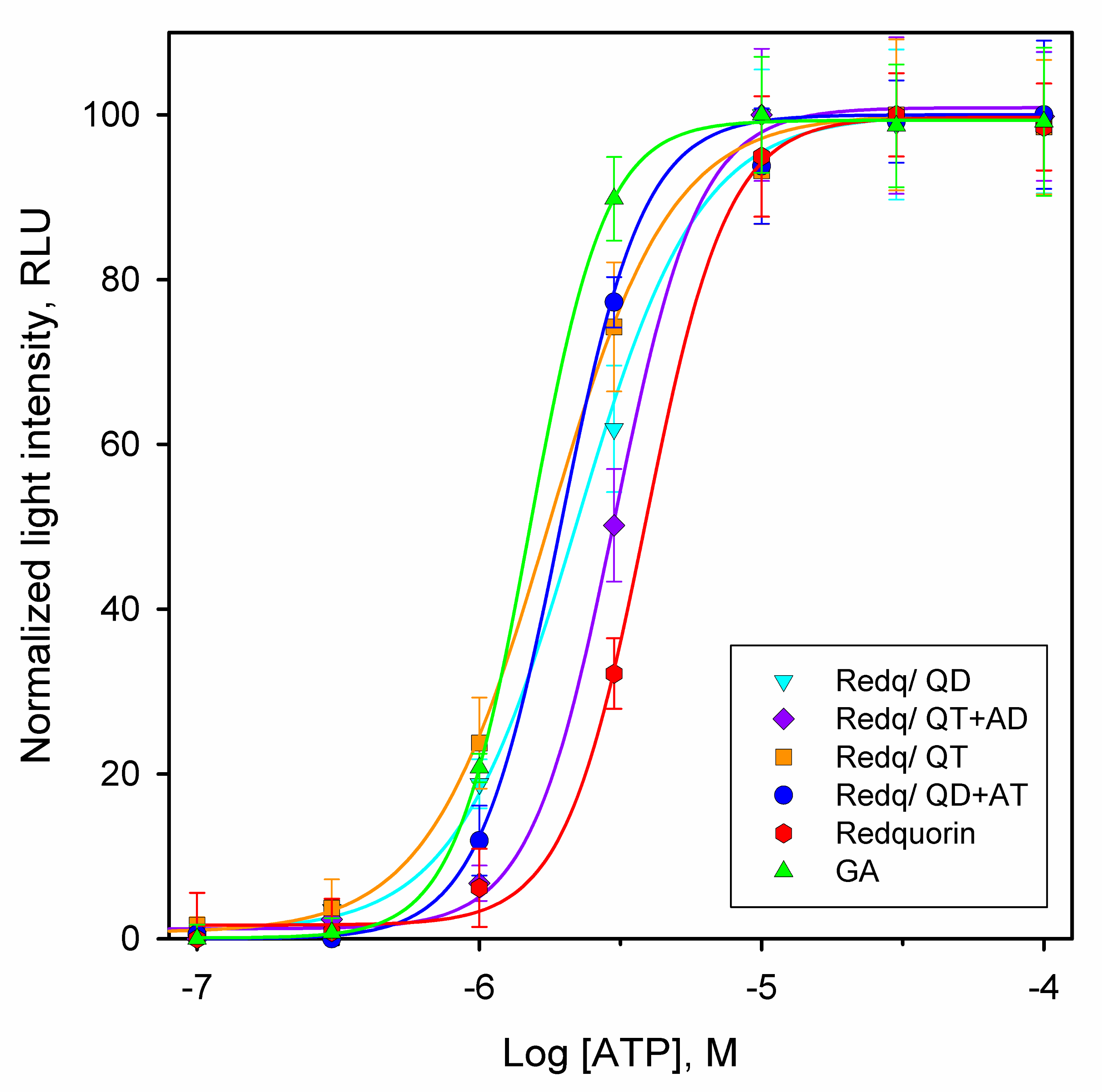
2.4. RedquorinXS Mutants Efficiently Report the Activation of P2Y2 Receptor in CHO Cells
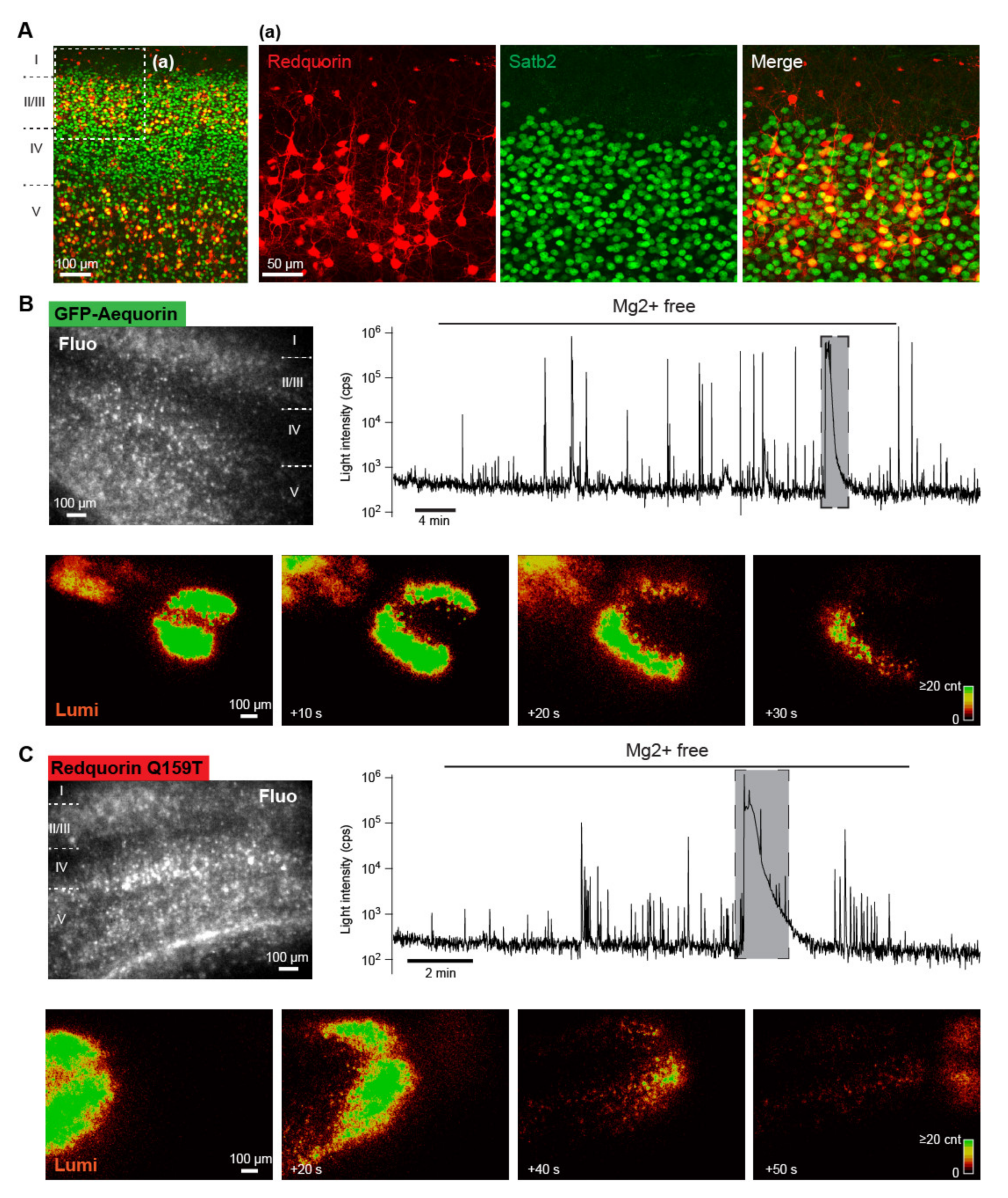
2.5. Imaging Neuronal Network Activities in Brain Slice with a RedquorinXS Mutant
3. Discussion
3.1. Apoaequorin Mutations That Enhance Aequorin Ca2+ Sensitivity
3.2. AequorinXS Mutations Primarily Enhance Redquorin Sensitivity to Low Ca2+
3.3. RedquorinXS Sensors Report Intracellular Ca2+ Signals with High Sensitivity in Cell Lines and in Brain Slice
4. Materials and Methods
4.1. Site-Directed Mutagenesis of Apo-Aequorin and Redquorin
4.2. Production and Purification of Apophotoproteins from Mammalian HEK Cells
4.3. Protein Expression and Purification from E. coli
4.4. Aequorin Reconstitution for In Vitro Assays
4.5. Functional Analysis of Aequorin Mutants and Fusions In Vitro
4.6. Functional Analysis of Photoproteins in CHO Cell Lines
4.7. Bioluminescence Imaging of Photoproteins in Mouse Neocortical Slices
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Russell, J.T. Imaging calcium signals in vivo: A powerful tool in physiology and pharmacology. Br. J. Pharmacol. 2011, 163, 1605–1625. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wilson, T.; Hastings, J.W. Bioluminescence. Annu. Rev. Cell. Dev. Biol. 1998, 14, 197–230. [Google Scholar] [CrossRef] [PubMed]
- Vysotski, E.S.; Lee, J. Ca2+-regulated photoproteins: Structural insight into the bioluminescence mechanism. Acc. Chem. Res. 2004, 37, 405–415. [Google Scholar] [CrossRef] [PubMed]
- Shimomura, O.; Johnson, F.H. Calcium binding, quantum yield, and emitting molecule in aequorin bioluminescence. Nature 1970, 227, 1356–1357. [Google Scholar] [CrossRef] [PubMed]
- Shimomura, O.; Johnson, F.H. Regeneration of the photoprotein aequorin. Nature 1975, 256, 236–238. [Google Scholar] [CrossRef] [PubMed]
- Allen, D.G.; Blinks, J.R.; Prendergast, F.G. Aequorin luminescence: Relation of light emission to calcium concentration—A calcium-independent component. Science 1977, 195, 996–998. [Google Scholar] [CrossRef] [PubMed]
- Tricoire, L.; Tsuzuki, K.; Courjean, O.; Gibelin, N.; Bourout, G.; Rossier, J.; Lambolez, B. Calcium dependence of aequorin bioluminescence dissected by random mutagenesis. Proc. Natl. Acad. Sci. USA 2006, 103, 9500–9505. [Google Scholar] [CrossRef] [Green Version]
- Hastings, J.W.; Mitchell, G.; Mattingly, P.H.; Blinks, J.R.; Van Leeuwen, M. Response of aequorin bioluminescence to rapid changes in calcium concentration. Nature 1969, 222, 1047–1050. [Google Scholar] [CrossRef]
- Illarionov, B.A.; Frank, L.A.; Illarionova, V.A.; Bondar, V.S.; Vysotski, E.S.; Blinks, J.R. Recombinant obelin: Cloning and expression of cDNA purification, and characterization as a calcium indicator. Methods Enzymol. 2000, 305, 223–249. [Google Scholar]
- Head, J.F.; Inouye, S.; Teranishi, K.; Shimomura, O. The crystal structure of the photoprotein aequorin at 2.3 A resolution. Nature 2000, 405, 372–376. [Google Scholar] [CrossRef]
- Deng, L.; Vysotski, E.S.; Markova, S.V.; Liu, Z.J.; Lee, J.; Rose, J.; Wang, B.C. All three Ca2+-binding loops of photoproteins bind calcium ions: The crystal structures of calcium-loaded apo-aequorin and apo-obelin. Protein Sci. 2005, 14, 663–675. [Google Scholar] [CrossRef] [Green Version]
- Toma, S.; Chong, K.T.; Nakagawa, A.; Teranishi, K.; Inouye, S.; Shimomura, O. The crystal structures of semi-synthetic aequorins. Protein Sci. 2005, 14, 409–416. [Google Scholar] [CrossRef] [Green Version]
- Malikova, N.P.; Burakova, L.P.; Markova, S.V.; Vysotski, E.S. Characterization of hydromedusan Ca2+-regulated photoproteins as a tool for measurement of Ca2+ concentration. Anal. Bioanal. Chem. 2014, 406, 5715–5726. [Google Scholar] [CrossRef]
- Markova, S.V.; Vysotski, E.S.; Blinks, J.R.; Burakova, L.P.; Wang, B.C.; Lee, J. Obelin from the bioluminescent marine hydroid Obelia geniculata: Cloning, expression, and comparison of some properties with those of other Ca2+-regulated photoproteins. Biochemistry 2002, 41, 2227–2236. [Google Scholar] [CrossRef]
- Shimomura, O.; Musicki, B.; Kishi, Y.; Inouye, S. Light-emitting properties of recombinant semi-synthetic aequorins and recombinant fluorescein-conjugated aequorin for measuring cellular calcium. Cell Calcium 1993, 14, 373–378. [Google Scholar] [CrossRef]
- Rowe, L.; Rothert, A.; Logue, C.; Ensor, C.M.; Deo, S.K.; Daunert, S. Spectral tuning of photoproteins by partnering site-directed mutagenesis strategies with the incorporation of chromophore analogs. Protein Eng. Des. Sel. 2008, 21, 73–81. [Google Scholar] [CrossRef] [Green Version]
- Dikici, E.; Qu, X.; Rowe, L.; Millner, L.; Logue, C.; Deo, S.K.; Ensor, M.; Daunert, S. Aequorin variants with improved bioluminescence properties. Protein Eng. Des. Sel. 2009, 22, 243–248. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gealageas, R.; Malikova, N.P.; Picaud, S.; Borgdorff, A.J.; Burakova, L.P.; Brûlet, P.; Vysotski, E.S.; Dodd, R.H. Bioluminescent properties of obelin and aequorin with novel coelenterazine analogues. Anal. Bioanal. Chem. 2014, 406, 2695–2707. [Google Scholar] [CrossRef] [PubMed]
- Bakayan, A.; Domingo, B.; Vaquero, C.F.; Peyriéras, N.; Llopis, J. Fluorescent Protein-photoprotein Fusions and Their Applications in Calcium Imaging. Photochem. Photobiol. 2017, 93, 448–465. [Google Scholar] [CrossRef] [Green Version]
- Brini, M.; Pinton, P.; Pozzan, T.; Rizzuto, R. Targeted recombinant aequorins: Tools for monitoring [Ca2+] in the various compartments of a living cell. Microsc. Res. Tech. 1999, 46, 380–389. [Google Scholar] [CrossRef]
- Kendall, J.M.; Sala-Newby, G.; Ghalaut, V.; Dormer, R.L.; Campbell, A.K. Engineering the Ca2+-activated photoprotein aequorin with reduced affinity for calcium. Biochem. Biophys. Res. Commun. 1992, 187, 1091–1097. [Google Scholar] [CrossRef]
- Montero, M.; Brini, M.; Marsault, R.; Alvarez, J.; Sitia, R.; Pozzan, T.; Rizzuto, R. Monitoring dynamic changes in free Ca2+ concentration in the endoplasmic reticulum of intact cells. EMBO J. 1995, 14, 5467–5475. [Google Scholar] [CrossRef] [PubMed]
- Drobac, E.; Tricoire, L.; Chaffotte, A.-F.; Guiot, E.; Lambolez, B. Ca2+ imaging in single neurons from brain slices using bioluminescent reporters. J. Neurosci. Res. 2010, 88, 695–711. [Google Scholar]
- Baubet, V.; Le Mouellic, H.; Campbell, A.K.; Lucas-Meunier, E.; Fossier, P.; Brûlet, P. Chimeric green fluorescent protein-aequorin as bioluminescent Ca2+ reporters at the single-cell level. Proc. Natl. Acad. Sci. USA 2000, 97, 7260–7265. [Google Scholar] [CrossRef] [Green Version]
- Rogers, K.L.; Stinnakre, J.; Agulhon, C.; Jublot, D.; Shorte, S.L.; Kremer, E.J.; Brûlet, P. Visualization of local Ca2+ dynamics with genetically encoded bioluminescent reporters. Eur. J. Neurosci. 2005, 21, 597–610. [Google Scholar] [CrossRef] [PubMed]
- Rogers, K.L.; Picaud, S.; Roncali, E.; Boisgard, R.; Colasante, C.; Stinnakre, J.; Tavitian, B.; Brûlet, P. Non-invasive in vivo imaging of Ca2+ signaling in mice. PLoS ONE 2007, 2, e974. [Google Scholar] [CrossRef] [PubMed]
- Agulhon, C.; Platel, J.C.; Kolomiets, B.; Forster, V.; Picaud, S.; Brocard, J.; Faure, P.; Brûlet, P. Bioluminescent imaging of Ca2+ activity reveals spatiotemporal dynamics in glial networks of dark-adapted mouse retina. J. Physiol. 2007, 583, 945–958. [Google Scholar] [CrossRef] [PubMed]
- Martin, J.R.; Rogers, K.L.; Chagneau, C.; Brûlet, P. In vivo bioluminescence imaging of Ca signalling in the brain of Drosophila. PLoS ONE 2007, 2, e275. [Google Scholar] [CrossRef]
- Naumann, E.A.; Kampff, A.R.; Prober, D.A.; Schier, A.F.; Engert, F. Monitoring neural activity with bioluminescence during natural behavior. Nat. Neurosci. 2010, 13, 513–520. [Google Scholar] [CrossRef] [Green Version]
- Tricoire, L.; Drobac, E.; Tsuzuki, K.; Gallopin, T.; Picaud, S.; Cauli, B.; Rossier, J.; Lambolez, B. Bioluminescence calcium imaging of network dynamics and their cholinergic modulation in slices of cerebral cortex from male rats. J. Neurosci. Res. 2019, 97, 414–432. [Google Scholar] [CrossRef]
- Curie, T.; Rogers, K.L.; Colasante, C.; Brûlet, P. Red-shifted aequorin-based bioluminescent reporters for in vivo imaging of Ca2 signaling. Mol. Imaging 2007, 6, 30–42. [Google Scholar] [CrossRef] [PubMed]
- Bakayan, A.; Vaquero, C.F.; Picazo, F.; Llopis, J. Red fluorescent protein-aequorin fusions as improved bioluminescent Ca2+ reporters in single cells and mice. PLoS ONE. 2011, 6, e19520. [Google Scholar] [CrossRef]
- Bakayan, A.; Domingo, B.; Miyawaki, A.; Llopis, J. Imaging Ca2+ activity in mammalian cells and zebrafish with a novel red-emitting aequorin variant. Pflügers Archiv Eur. J. Physiol. 2015, 467, 2031–2042. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Stepanyuk, G.A.; Golz, S.; Markova, S.V.; Frank, L.A.; Lee, J.; Vysotski, E.S. Interchange of aequorin and obelin bioluminescence color is determined by substitution of one active site residue of each photoprotein. FEBS Lett. 2005, 579, 1008–1014. [Google Scholar] [CrossRef] [Green Version]
- Tsuzuki, K.; Tricoire, L.; Courjean, O.; Gibelin, N.; Rossier, J.; Lambolez, B. Thermostable mutants of the photoprotein aequorin obtained by in vitro evolution. J. Biol. Chem. 2005, 280, 34324–34331. [Google Scholar] [CrossRef] [Green Version]
- Stables, J.; Green, A.; Marshall, F.; Fraser, N.; Knight, E.; Sautel, M.; Milligan, G.; Lee, M.; Rees, S. A bioluminescent assay for agonist activity at potentially any G-protein-coupled receptor. Anal. Biochem. 1997, 252, 115–126. [Google Scholar] [CrossRef]
- Ungrin, M.D.; Singh, L.M.; Stocco, R.; Sas, D.E. An automated aequorin luminescence-based functional calcium assay for G-protein-coupled receptors. Anal. Biochem. 1999, 272, 34–42. [Google Scholar] [CrossRef]
- Zhang, J.H.; Chung, T.D.Y.; Oldenburg, K.R. A simple statistical parameter for use in evaluation and validation of high throughput screening assays. J. Biomol. Screen. 1999, 4, 67–73. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Song, N.N.; Lan, W.; Hu, L.; Su, C.J.; Ding, Y.Q.; Zhang, L. Expression of transcription factor Satb2 in adult mouse brain. Anat. Rec. 2013, 296, 452–461. [Google Scholar] [CrossRef]
- Holmgren, C.; Harkany, T.; Svennenfors, B.; Zilberter, Y. Pyramidal cell communication within local networks in layer 2/3 of rat neocortex. J. Physiol. 2003, 551, 139–153. [Google Scholar] [CrossRef]
- Gutnick, M.J.; Connors, B.W.; Prince, D.A. Mechanisms of neocortical epileptogenesis in vitro. J. Neurophysiol. 1982, 48, 1321–1335. [Google Scholar] [CrossRef] [PubMed]
- Connors, B.W. Initiation of synchronized neuronal bursting in neocortex. Nature 1984, 310, 685–687. [Google Scholar] [CrossRef]
- Gloveli, T.; Albrecht, D.; Heinemann, U. Properties of low Mg2+ induced epileptiform activity in rat hippocampal and entorhinal cortex slices during adolescence. Brain Res. Dev. Brain Res. 1995, 87, 145–152. [Google Scholar] [CrossRef]
- Köhling, R.; Koch, U.R.; Hagemann, G.; Redecker, C.; Straub, H.; Speckmann, E.J. Differential sensitivity to induction of spreading depression by partial disinhibition in chronically epileptic human and rat as compared to native rat neocortical tissue. Brain Res. 2003, 975, 129–134. [Google Scholar] [CrossRef]
- Ayata, C.; Lauritzen, M. Spreading Depression, Spreading Depolarizations, and the Cerebral Vasculature. Physiol. Rev. 2015, 95, 953–993. [Google Scholar] [CrossRef] [Green Version]
- Ohashi, W.; Inouye, S.; Yamazaki, T.; Hirota, H. NMR analysis of the Mg2+-binding properties of aequorin, a Ca2+-binding photoprotein. J. Biochem. 2005, 138, 613–620. [Google Scholar] [CrossRef]
- Xu, C.; Peter, M.; Bouquier, N.; Ollendorff, V.; Villamil, I.; Liu, J.; Fagni, L.; Perroy, J. REV, a BRET-Based Sensor of ERK Activity. Front. Endocrinol. 2013, 4, 95. [Google Scholar] [CrossRef] [Green Version]
- Suzuki, K.; Nagai, T. Recent progress in expanding the chemiluminescent toolbox for bioimaging. Curr. Opin. Biotechnol. 2017, 48, 135–141. [Google Scholar] [CrossRef]
- Namkung, Y.; LeGouill, C.; Kumar, S.; Cao, Y.; Teixeira, L.B.; Lukasheva, V.; Giubilaro, J.; Simões, S.C.; Longpré, J.M.; Devost, D.; et al. Functional selectivity profiling of the angiotensin II type 1 receptor using pathway-wide BRET signaling sensors. Sci. Signal. 2018, 11, eaat1631. [Google Scholar] [CrossRef] [Green Version]
- Valkovic, A.L.; Leckey, M.B.; Whitehead, A.R.; Hossain, M.A.; Inoue, A.; Kocan, M.; Bathgate, R.A.D. Real-time examination of cAMP activity at relaxin family peptide receptors using a BRET-based biosensor. Pharmacol. Res. Perspect. 2018, 6, e00432. [Google Scholar] [CrossRef] [PubMed]
- Noda, N.; Ishimoto, T.; Mori, H.; Ozawa, T. Enhanced bioluminescent sensor for longitudinal detection of CREB activation in living cells. Photochem. Photobiol. Sci. 2019, 18, 2740–2747. [Google Scholar] [CrossRef]
- Bouquier, N.; Moutin, E.; Tintignac, L.A.; Reverbel, A.; Jublanc, E.; Sinnreich, M.; Chastagnier, Y.; Averous, J.; Fafournoux, P.; Verpelli, C.; et al. AIMTOR, a BRET biosensor for live imaging, reveals subcellular mTOR signaling and dysfunctions. BMC Biol. 2020, 18, 81. [Google Scholar] [CrossRef] [PubMed]
- Tricoire, L.; Lambolez, B. Neuronal network imaging in acute slices using Ca2+ sensitive bioluminescent reporter. Methods Mol. Biol. 2014, 1098, 33–45. [Google Scholar]


| Mutation | Ca2+ Sensitivity, EC50 (nM) a | Relative Intensity a | Decay Kinetics, t1/2 (ms) b | Emission Peak Wavelength (nm) | ||
|---|---|---|---|---|---|---|
| at pCa 6.5 | at pCa 7.2 | |||||
| Aeq-wt | 659 ± 23 | 1 | 1 | 906 ± 53 | 476 | |
| Single mutants with medium-high to high Ca2+ sensitivity | Q159D | 330 ± 8 | 25.1 | 23.4 | 794 ± 42 | 478 |
| Q159T | 310 ± 13 | 16.6 | 4.8 | 866 ± 23 | 477 | |
| S157D | 322 ± 20 | 6.0 | 2.2 | 835 ± 47 | 475 | |
| N121D | 400 ± 14 | 4.9 | 2.1 | 957 ± 30 | 477 | |
| A123D | 374 ± 9 | 3.6 | 1.6 | 862 ± 48 | 476 | |
| Q159G | 492 ± 10 | 3.3 | 3.2 | 830 ± 37 | 478 | |
| A179T | 330 ± 19 | 3.0 | 1.3 | 779 ± 33 | 477 | |
| Double mutants with high Ca2+ sensitivity | QD+AT | 216 ± 26 | 58.5 | 60.8 | 750 ± 50 | 478 |
| QD+ND | 295 ± 17 | 29.0 | 41.7 | 738 ± 28 | 477 | |
| QD+AD | 271 ± 15 | 26.2 | 15.0 | 612 ± 26 | 475 | |
| QT+AT | 281 ± 25 | 29.0 | 14.0 | 650 ± 68 | 478 | |
| QT+ND | 337 ± 10 | 14.4 | 12.2 | 680 ± 55 | 478 | |
| QT+AD | 279 ± 16 | 22.8 | 10.0 | 620 ± 33 | 476 | |
| Mutation | Ca2+ Sensitivity, EC50 (nM) a | Relative Intensity a | Decay Kinetics, t1/2 (ms) b | Emission Peak Wavelength (nm) | ||
|---|---|---|---|---|---|---|
| at pCa 6,5 | at pCa 7,2 | |||||
| CLZ-native | Redq | 859 ± 45 | 1.0 | 1.0 | 1 203 ± 70 | 582 |
| Redq/Q159D | 680 ± 33 | 9.1 | 13.2 | 980 ± 66 | 582 | |
| Redq/QD+AT | 515 ± 20 | 32.1 | 24.5 | 913 ± 39 | 582 | |
| Redq/QD+AD | 621 ± 44 | 10.8 | 12.8 | 1 120 ± 46 | 582 | |
| Redq/Q159T | 478 ± 39 | 13.0 | 7.1 | 1 103 ± 50 | 582 | |
| Redq/QT+AT | 605 ± 16 | 18.3 | 14.1 | 990 ± 87 | 582 | |
| Redq/QT+AD | 577 ± 33 | 14.4 | 10.0 | 950 ± 56 | 582 | |
| CitA | 573 ± 45 | 24 | 9.5 | 794 ± 46 | 529 | |
| GA | 630 ± 29 | 6.8 | 3.2 | 852 ± 64 | 509 | |
| CLZ-f | Redq | 600 ± 35 | 1.0 | 1.0 | 880 ± 68 | 582 |
| Redq/Q159D | 290 ± 18 | 23.9 | 14.9 | 770 ± 88 | 582 | |
| Redq/QD+AT | 252 ± 40 | 58.5 | 48.3 | 740 ± 55 | 582 | |
| Redq/QD+AD | 300 ± 26 | 22.8 | 15.0 | 750 ± 66 | 582 | |
| Redq/Q159T | 296 ± 13 | 11.0 | 2.8 | 705 ± 36 | 582 | |
| Redq/QT+AT | 284 ± 30 | 25.6 | 9.0 | 810 ± 61 | 582 | |
| Redq/QT+AD | 336 ± 11 | 14.4 | 7.4 | 670 ± 50 | 582 | |
| CitA | 266 ± 18 | 41.1 | 18.7 | 680 ± 72 | 529 | |
| GA | 463 ± 46 | 8.6 | 2.9 | 700 ± 48 | 509 | |
| Ca2+ Sensor Variant | [ATP] CL (nM) | EC50 (µM) | Z-Factor |
|---|---|---|---|
| Redq/Q159T | 310 | 1.7 ± 0.2 | 0.76 |
| Redq/Q159D | 320 | 2.2 ± 0.1 | 0.68 |
| Redq/QD+AT | 548 | 2.0 ± 0.2 | 0.67 |
| Redq/QT+AD | 1025 | 3.0 ± 0.4 | 0.82 |
| Redq | 2033 | 4.0 ± 0.3 | 0.56 |
| GA | 640 | 1.5 ± 0.2 | 0.69 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bakayan, A.; Picaud, S.; Malikova, N.P.; Tricoire, L.; Lambolez, B.; Vysotski, E.S.; Peyriéras, N. RedquorinXS Mutants with Enhanced Calcium Sensitivity and Bioluminescence Output Efficiently Report Cellular and Neuronal Network Activities. Int. J. Mol. Sci. 2020, 21, 7846. https://doi.org/10.3390/ijms21217846
Bakayan A, Picaud S, Malikova NP, Tricoire L, Lambolez B, Vysotski ES, Peyriéras N. RedquorinXS Mutants with Enhanced Calcium Sensitivity and Bioluminescence Output Efficiently Report Cellular and Neuronal Network Activities. International Journal of Molecular Sciences. 2020; 21(21):7846. https://doi.org/10.3390/ijms21217846
Chicago/Turabian StyleBakayan, Adil, Sandrine Picaud, Natalia P. Malikova, Ludovic Tricoire, Bertrand Lambolez, Eugene S. Vysotski, and Nadine Peyriéras. 2020. "RedquorinXS Mutants with Enhanced Calcium Sensitivity and Bioluminescence Output Efficiently Report Cellular and Neuronal Network Activities" International Journal of Molecular Sciences 21, no. 21: 7846. https://doi.org/10.3390/ijms21217846
APA StyleBakayan, A., Picaud, S., Malikova, N. P., Tricoire, L., Lambolez, B., Vysotski, E. S., & Peyriéras, N. (2020). RedquorinXS Mutants with Enhanced Calcium Sensitivity and Bioluminescence Output Efficiently Report Cellular and Neuronal Network Activities. International Journal of Molecular Sciences, 21(21), 7846. https://doi.org/10.3390/ijms21217846






