1. Introduction
Venezuelan equine encephalitis virus (VEEV) is a New World alphavirus and belongs to the
Togaviridae family of Group IV viruses. In the past few thousand years, the New World alphaviruses including Eastern equine encephalitis virus (EEEV) and Western equine encephalitis virus (WEEV) have evolved separately from those of the Old World including Sindbis virus (SINV), Semliki forest virus (SFV) and Chikungunya virus (CHIKV) [
1]. VEEV is transferred to birds, horses, humans, and other vertebrates via arbovector (mosquitos), however, it is especially dangerous for equines, with an average fatality rate of 20–80% [
2,
3]. In humans, acute VEEV infections are usually defeated by innate and adaptive immune responses, however, about 14% of the infected humans develop neurological symptoms and about 1% of the infections result in lethal bugoencephalitis [
3,
4]. The inhaled virus can enter the brain via the olfactory neurons and as its viral particles are unusually resistant to desiccation and can be stably freeze-dried and aerosolized, therefore, VEEV has been regarded as potential biological weapon, that has been reportedly developed by several nations including the US and former Soviet Union [
5,
6].
Alphaviruses are small, spherical enveloped viruses and possess an ~11-kb-long ss(+)RNA genome [
7]. The mRNA-like genome consists of two open reading frames (ORFs): (i) the first cistron codes for four non-structural proteins (nsPs) that are essential for viral replication complex formation and are translated in a form of a single polyprotein, nsP123 or nsP1234; while (ii) the second ORF encodes the structural proteins that form viral particles and are translated in the late stage of the infection.
The non-structural protein 2 (nsP2) is a central element of the VEEV life-cycle and has multiple enzyme activities. Its N-terminal region (Gly1-Ile456) is associated with ATP-ase and GTP-ase activity [
8], RNA helicase activity [
9], and RNA 5′-triphosphatase activity [
10], while the C-terminal region (Met457–Cys794) controls 26S subgenomic RNA synthesis [
11], downregulates minus-strand RNA synthesis late in infection [
12,
13], and directs nsP2 for nuclear transport [
14]. Besides the listed activities, nsP2 is also essential in forming the alphavirus non-structural polyprotein replication complex via its proteolytic activity [
15,
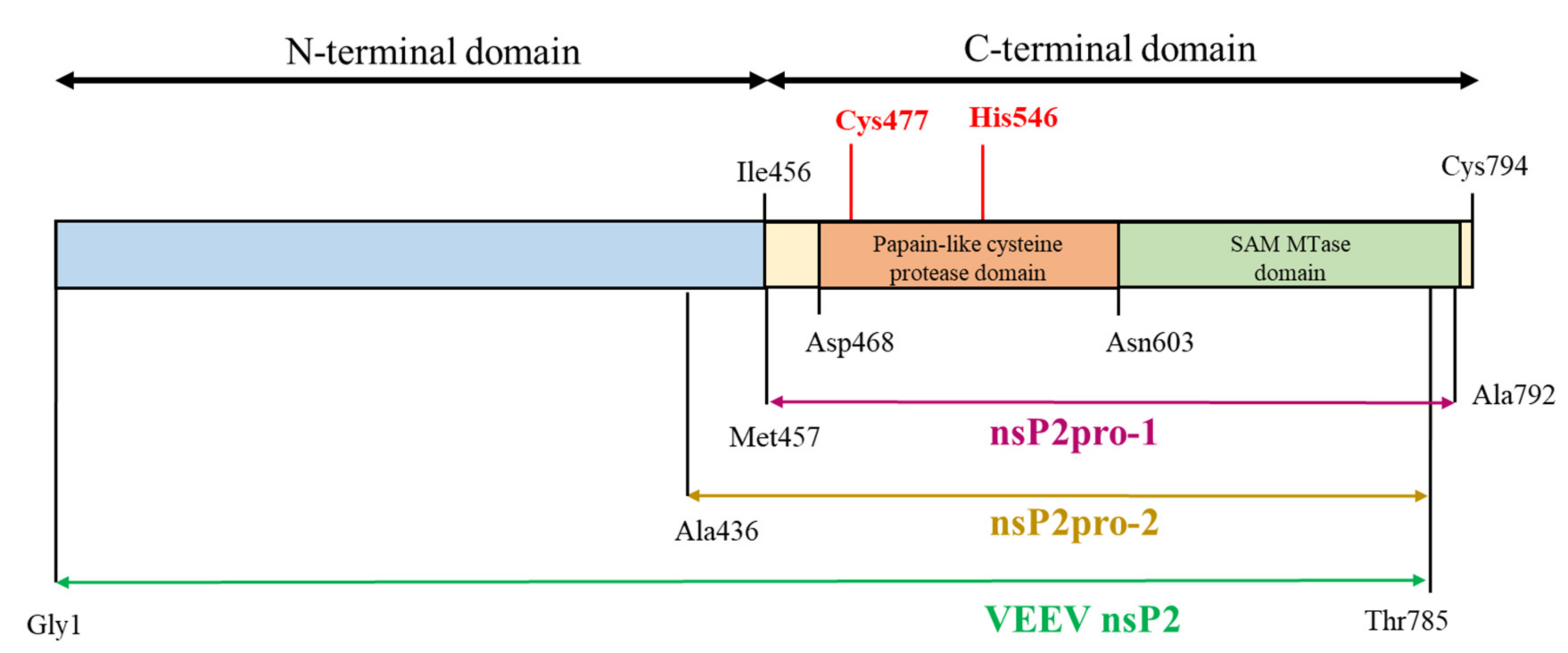
16], as the C-terminal region of nsP2—frequently referred to as nsP2pro—consists of a papain-like cysteine protease domain linked to an S-adenosyl-L-methionine-dependent RNA methyltransferase domain (SAM MTase), for which the function has not been clarified. The active site of the protease (PR) is formed by a catalytic dyad including residues Cys477 and His546, while the substrate binding cleft is positioned between the protease and SAM MTase domains. Most of the residues that contribute to substrate binding are located in the protease domain; however, Hu et al. [
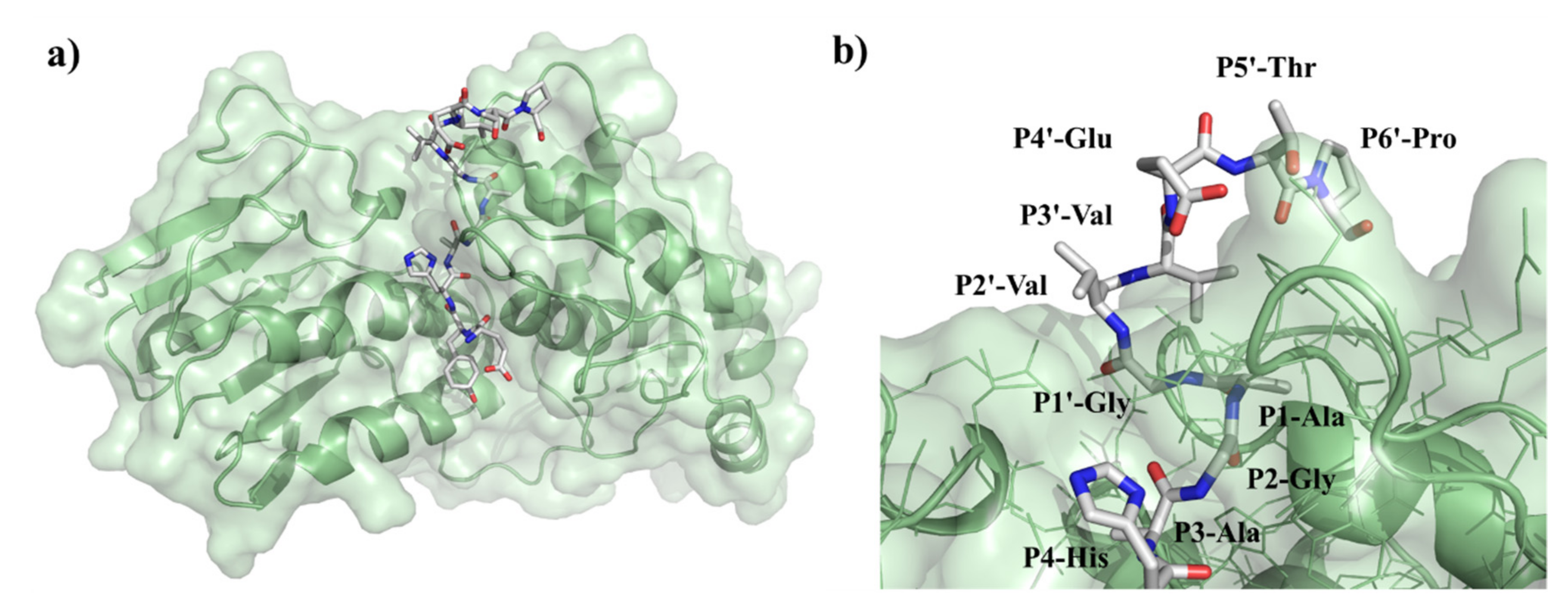
5] have confirmed that at least three residues of the SAM MTase domain—including Arg662, Lys705 and Lys706—are involved in substrate recognition and in the regulation of the substrate-binding cleft structure. NsP2pro processes the non-structural polyprotein at three different sites located between nsP1/nsP2, nsP2/nsP3 and nsP3/nsP4 (referred to as nsP12, nsP23, and nsP34, respectively). All these sites share common sequential properties. According to the alignment of Strauss and Strauss [
17] the residue in the P2 position is Gly in all three sites and the residue in P1 position is always an amino acid with a small side chain. Furthermore, the P3 site is considered as highly conserved owing to the relatively low number of the residues occurring therein. In contrast, P4 residues show greater variability suggesting that the major recognition signal for cleavage is likely to be displayed by residues P3-P2-P1. The amino acid sequence of P1′–P4′ positions are highly similar only within the same type of cleavage sites and differ significantly across them. This indicates that these residues—corresponding to the N terminus of the newly formed nsPs—are likely to be determined by their function in the released nsPs rather than by a requirement for cleavage site recognition.
According to Russo et al. [
18], S1 pocket is formed by Asn475, Val476, and Ala509 residues that are thought to be either highly conserved or contain similar residues in most sequences (
Figure S1), while backbone amides of Val476 and Cys477 likely form the oxyanion hole. The S2 subsite is mainly defined by the highly conserved Trp547 following the catalytic His546 and, together with the fact that the corresponding P2 site contains a highly conserved Gly, this structure of VEEV nsP2pro highly resembles the so-called “glycine specificity motif” of other cysteine proteases [
19]. The S3 binding pocket is located on the SAM MTase domain, is composed of Ile698 and Met702, and is flanked by highly conserved Ala509 and His510 residues.
The characterization of nsP2pro in vivo and in vitro enzyme activity and specificity has been performed mainly in the context of SINV and SFV, and most of the previously performed mutagenesis studies were focused on activity of SIN and SFV nsP2pro on their cognate substrates mutated at P5-P1′ [
20]. In the case of VEEV nsP2pro, only limited data are available regarding its specificity, although, the crystal structure of both the free and the E-64d-bound form has been solved [
5,
18]. Previously we have tested activities of three alphavirus proteases to investigate whether these enzymes are useful tools for affinity tag removal, similar to potyviral proteases including tobacco etch virus protease (TEV PR) that is already widely utilized for such purposes [
20]. The activity of the recombinant SIN, SFV, and VEEV nsP2pro domains was investigated on different fusion proteins and on artificial oligopeptide substrates designed based on the P6-P6′ residues of their natural recognition sites. As the recognition sites of the three enzymes are highly similar, besides investigating the activity of the proteases on their cognate substrates, their cross-reactivity was also tested [
20].
Kinetic parameters were determined previously on oligopeptide substrates for three enzyme–substrate pairs including SFV protease on its cognate SFV nsP12 and nsP34 cleavage site and VEEV protease on SFV nsP12. In that study [
20], the VEEV protease was constructed based on Met457–Ala792, further referred to as nsP2pro-1, corresponding to the C-terminal domain of the nearly full-length nsP2 that lacks 793–794 terminal residues (
Figure 1). The observed k
cat/K
M values were substantially lower than those of potyviral proteases on their cognate substrates, furthermore, their ability to process fusion protein substrates was also relatively poor. We concluded that despite the fact that alphavirus proteases theoretically exhibit sufficient sequence specificity, in practice their cleavage properties in their investigated form could not offer any advantage over the already successfully utilized potyviral proteases.
Besides the biotechnological considerations, VEEV nsP2pro is also an antiviral drug target owing to (i) its essential function in the viral life-cycle as discussed above, and (ii) its potential central role in antagonizing the interferon response by cleaving host proteins corresponding to the generation of the innate immune response via short stretches of homologous host–pathogen protein sequences (SSHHPS) [
21]. Mechanisms that antagonize early innate immune responses by proteolytic activity of viral nsPs are believed to have important function in other Group IV viruses, e.g.,
Picornaviridae,
Flaviviridae, and
Coronaviridae. Coronaviruses are under “super-intensive” investigation because several new pathogens contributing to recent global pandemics have emerged from these families including Middle East respiratory syndrome- and severe acute respiratory syndrome-associated coronavirus (MERS- and SARS-CoV), and SARS-CoV-2. In accordance with the similar mode of action of
Coronaviridae and
Togaviridae, structural similarities have been suggested by Compton et al. [
22] between VEEV nsP2 and coronaviral papain-like proteases/deubiquitinases of the SARS- and MERS-CoV.
Currently there are no therapeutics approved for the treatment of VEEV infections, therefore, the characterization of the nsP2 proteolytic activity may be important for the development of successful inhibitor candidate molecules. In order to investigate the in vitro proteolytic activity of VEEV nsP2pro, we aimed to design two recombinant VEEV nsP2 constructs and study the effects of an N-terminal sequence elongation of the protease domain. In these new constructs, the sequence of nsP2pro-1 was expanded with (i) the whole N-terminal domain, further referred to as VEEV nsP2 or (ii) with the Ala436-Met457 region of VEEV nsP2, referred as nsP2pro-2 (see
Figure 1). To study catalytic activity and specificity in vitro, we utilized such His
6-MBP-mEYFP fusion protein substrates (His
6, hexahistidine; MBP, maltose binding protein; mEYFP, monomeric enhanced yellow fluorescent protein). The recombinant proteins comprised a wild-type or a modified SFV nsP12 cleavage site sequence and were used as substrates in an Ni-NTA magnetic bead-based protease assay which has been developed in our laboratory [
23,
24,
25].
3. Discussion
This study was made with the aim of characterization of nsP2 cysteine protease of VEEV. First, we compared the expression profiles and solubility of two N-terminally elongated constructs. We studied a nearly full-length VEEV nsP2 (1–785) and a VEEV nsP2pro-2 (436–785) construct (
Figure 1). The VEEV nsP2pro-2 construct contains the C-terminal protease domain of nsP2 extended with the C-terminal segment of the N-terminal domain (436–457). The recombinant proteins were fused either to His
6- or His
6-MBP tags and expressed in
E. coli Rozetta cells. We found a low level of expression and marginal solubility for both the His
6- or His
6-MBP-fused VEEV nsP2 in each tested condition. In the case of the His
6-VEEV nsP2, this may be explained in part by the inherent structural and folding properties of the protein. Based on the findings of Saisawang et al. [
34], the insufficient solubility may be a result of the aggregation of the construct with the bacterial host cell proteins. As they described, the attachment of MBP to the full-length CHIKV nsP2 solved the problem related to inclusion body formation, but the tagged proteins were still aggregated with contaminating
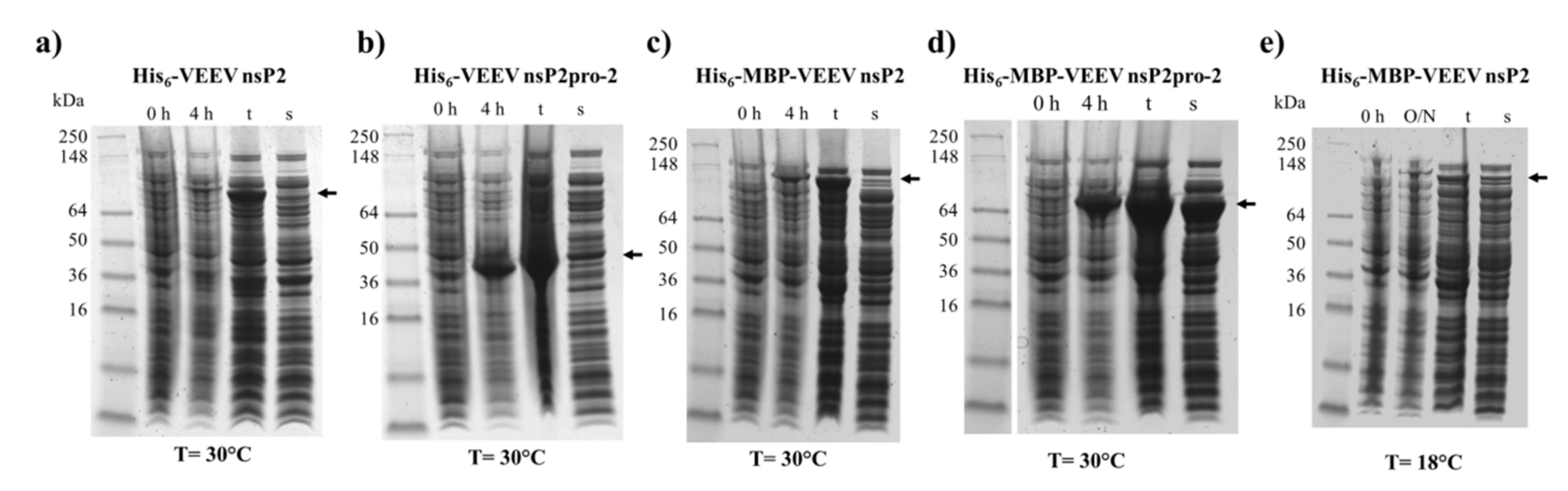
E. coli proteins. In contrast, the expression and solubility level of His
6-MBP-VEEV nsP2pro-2 construct was sufficient (
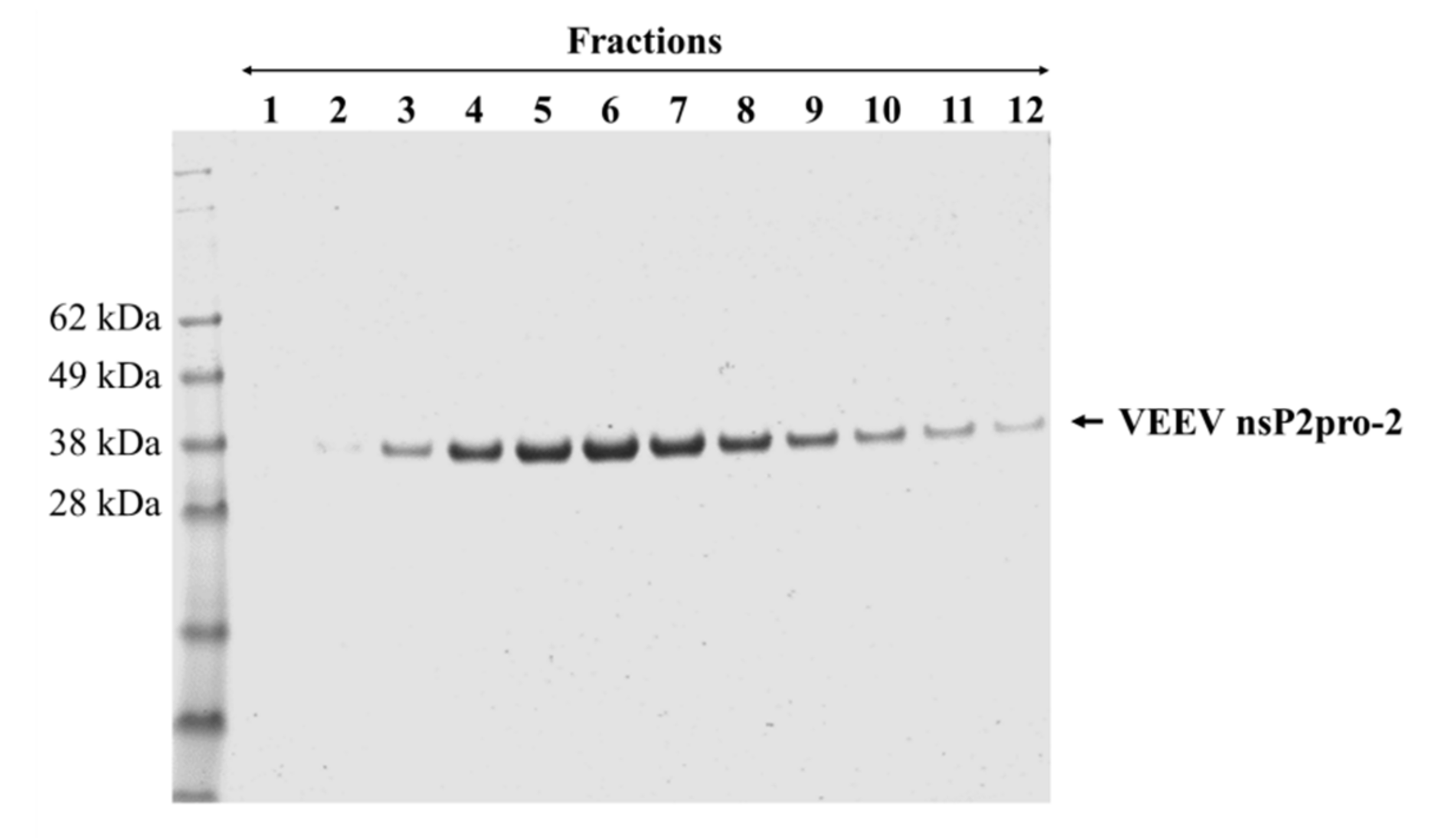
Figure 2). Therefore, we used this construct to prepare VEEV nsP2pro-2 for enzyme specificity studies. The untagged VEEV nsP2pro-2 was produced in high purity using a four-step purification protocol including an enzymatic removal of the dual fusion tag by TEV protease (
Figure 3).
The enzyme activity measurements were performed by using a recombinant fluorescent protein substrate-based protease assay, which has been previously applied successfully to study TEV and HIV-1 proteases [
23], the retroviral-like protease of human paternally expressed gene 10 (PEG10) protein [
35] and yeast retrotransposon Ty1 [
36]. The previously developed method was slightly modified and some improvements were introduced as follows. We reduced the volume of substrate expression from 50 to 15 mL, the small-scale expression was carried out in centrifuge tubes that enabled working with multiple mutants simultaneously. The protease assay was adapted to a 96-well plate-based format, reduced volumes of working samples increased cost-efficiency and throughput of the assay as compared to the microcentrifuge tube-based assay. Random mutagenesis was also used for the generation of the mutants, which has not been applied previously in the case of this assay system.
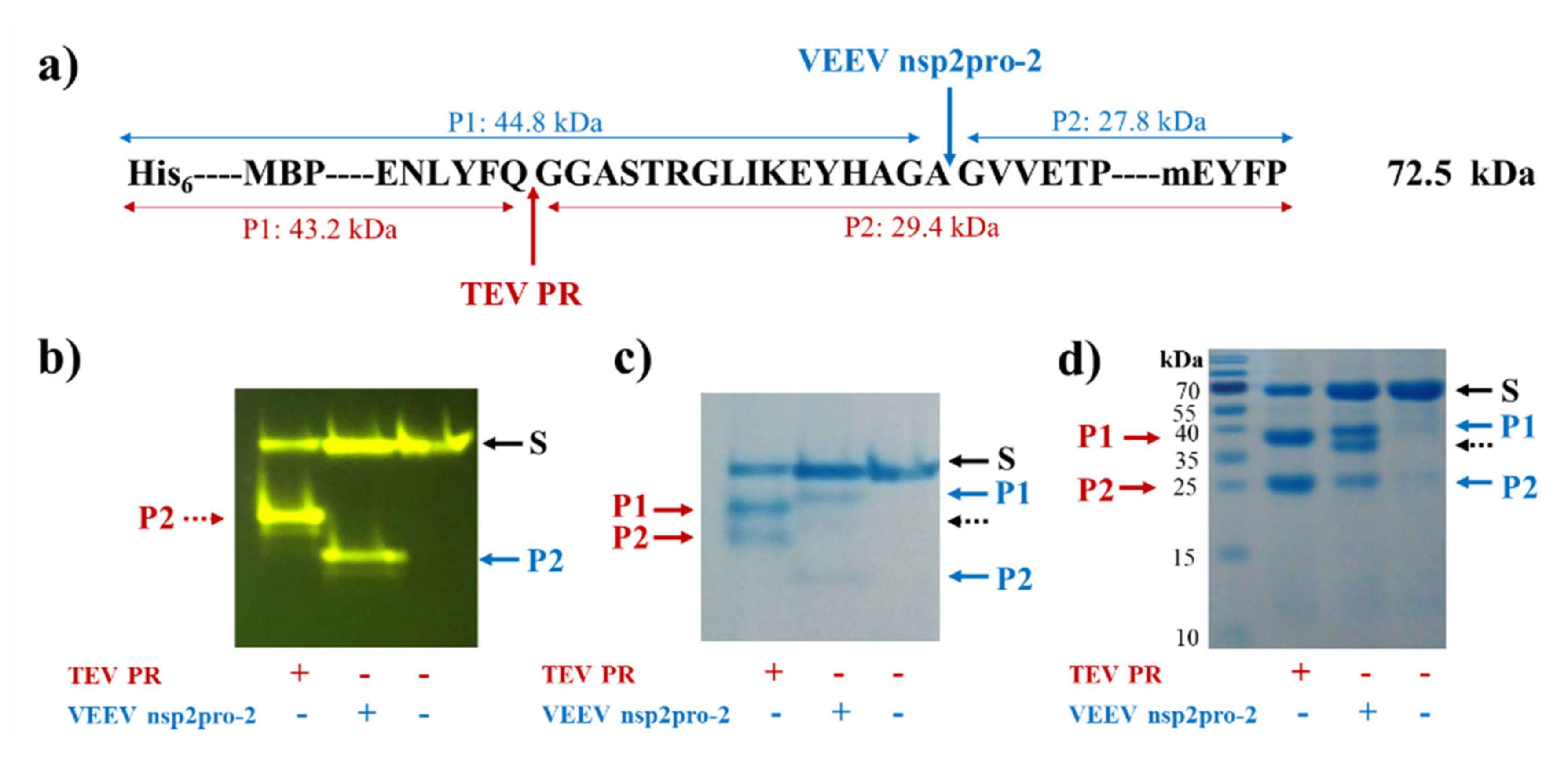
We have adapted the previously designed substrate system to investigate VEEV nsP2 protease, and generated His
6-MBP-mEYFP substrates that contained the wild-type or modified nsP12 cleavage site of SFV (EYHAGA↓GVVETP). The complete series of P1′ variants and other substrate mutants may be applicable to study additional alphavirus proteases, e.g., specificities of SFV and CHIKV PRs. Substrates containing SFV nsP12 recognition sequence were chosen for VEEV PR enzymatic assays because substrates representing the same cleavage site have already been applied successfully for the investigation of VEEV nsP2 protease [
5,
20], which is supported by our earlier specificity results which suggested that the peptide representing SFV nsP12 was a substantially better substrate of VEEV protease than its cognate sequences [
20]. The designed recombinant proteins were proved to be folded properly and are susceptible for cleavage by TEV and VEEV nsP2 proteases (
Figure 4). The generated substrate library can be potentially used to study other alphaviral proteases because, e.g., proteases of SFV, SINV are also known to cleave SFV nsP12 cleavage site [
20].
The kinetic parameters obtained for VEEV nsP2pro-2 on the protein substrate containing nsP1/nsP2 cleavage site of SFV, were compared to the kinetic data determined previously for VEEV nsP2pro-1 using oligopeptide substrates [
20] or CFP/YFP-based recombinant FRET substrates [
5] coding the same cleavage site sequence. The differences between the K
M values may be explained by the conformational differences of the different substrate, in agreement with our previous kinetic studies on TEV PR and HIV-1 PRs [
23,
24]. The k
cat and k
cat/K
M values are not fully comparable with the literature data [
5,
20], because the active enzyme concentration cannot be determined due to the lack of any selective inhibitor. Still, if enzyme activity is considered as 100% in each case, the k
cat/K
M values are at the same order of magnitude, indicating that the catalytic efficiency of VEEV nsP2pro-2 is similar to that of VEEV nsP2pro-1. This implies that N-terminal extension of VEEV nsP2pro-1 (with 436–457 region of nsP2) has no substantial effect on the activity of the VEEV protease domain in vitro.
To study enzyme specificity of VEEV nsP2pro-2 in the context of nsP12 cleavage site of SFV, we designed His
6-MBP-mEYFP substrates modified in P5, P4, P2, P1, P1′, and P2′ positions. The screening studies were performed by the 96-well plate-adapted format of the previously designed microtube-based assay [
23,
24,
25]. As the amount of substrate needed for the plate-based format was much smaller than that of the microvial-based setup, we optimized the expression of the substrates to a smaller scale (15 mL), accordingly. Following the microplate-based specificity studies, kinetic parameters were determined for some selected variants. The activities determined by the 96-well plate-based assay were in agreement with the values obtained by the microvial-based measurements. This suggests the utility of the plate-based setup in ranking the hits according to their catalytic efficiency if the applied substrate concentration is in the dynamic range of the enzyme, although the results of the applied statistical tests on the specific activity and k
cat/K
M data are contradictory in the case of P4-Arg, P4-Gly, P1-Gly, and P2-Ser.
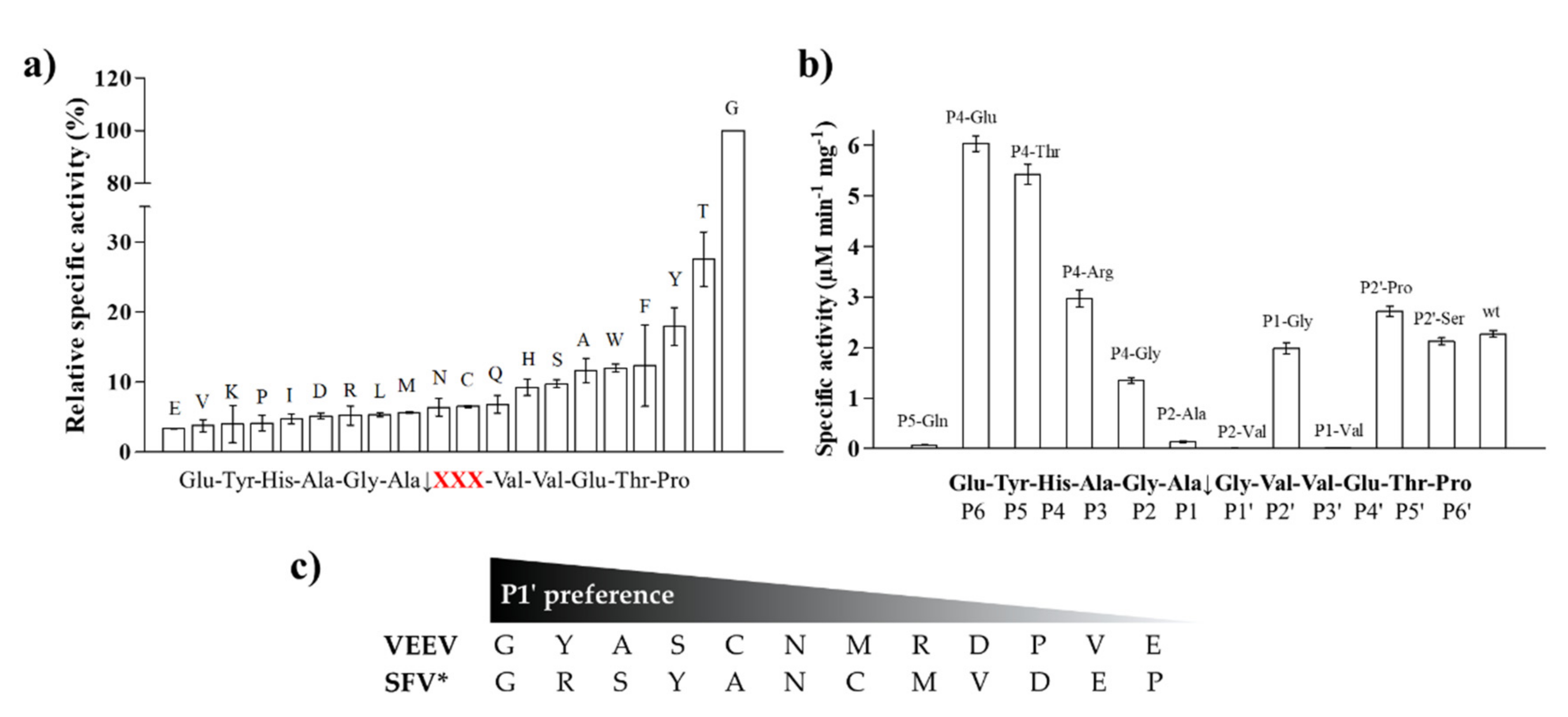
To support the interpretation of the in vitro enzymatic measurements, modeled structure of the VEEV nsP2pro (469–767) complexed with the studied oligopeptide substrate was prepared, and the effects of mutations on enzyme–substrate interactions were investigated in silico. Screening of P1′ specificity showed the highest preference of VEEV nsP2pro for a P1′-Gly residue, while significantly lower preference was determined for all P1′ variants in the context of EYHAGA↓GVVETP cleavage site (
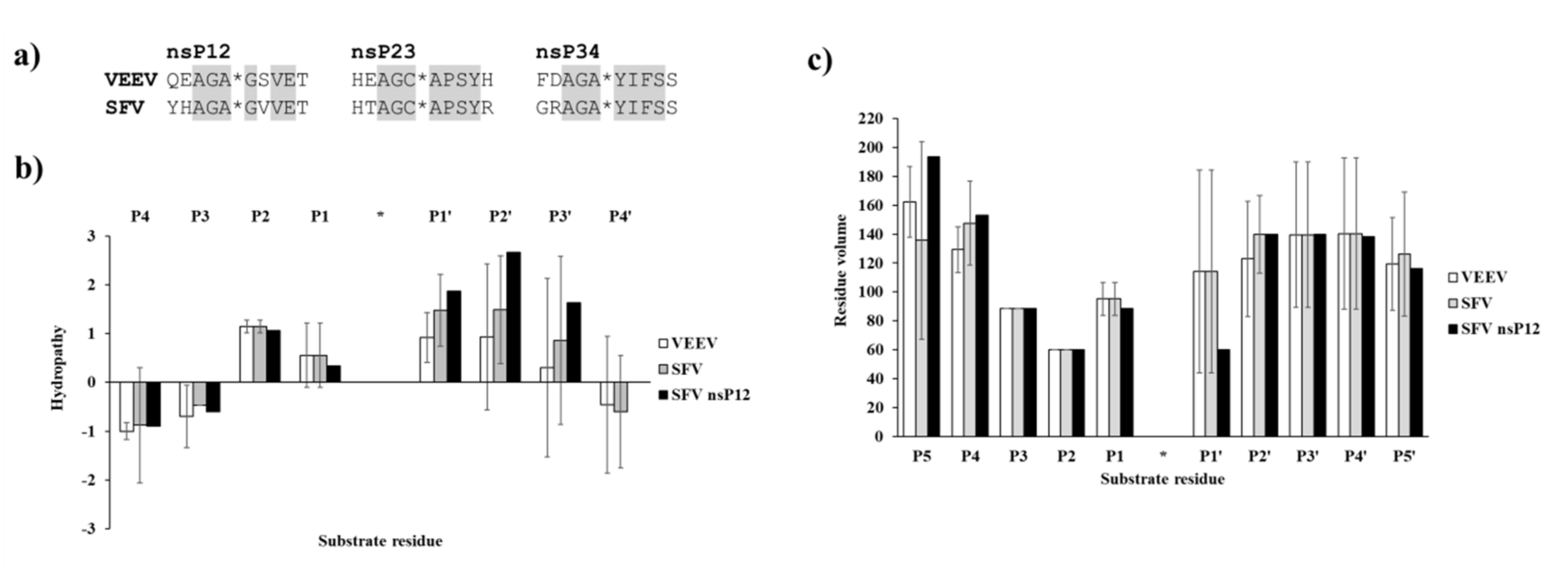
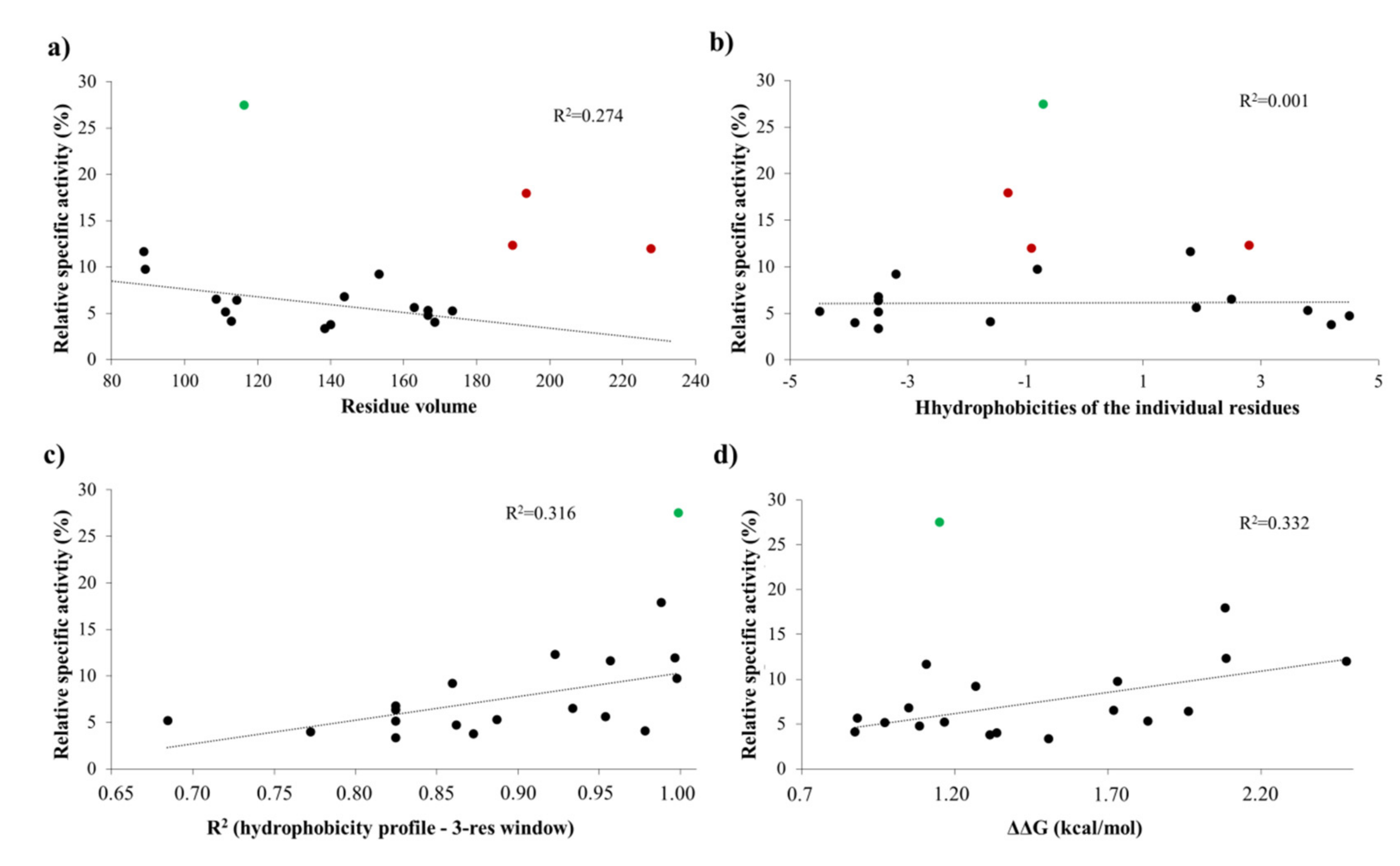
Figure 5). Structural analysis showed no correlation of the obtained activities with hydrophobicities and volumes of the individual P1′ residues, or with the overall hydrophobicity profiles of cleavage sites, furthermore, predicted stability changes upon the mutations also showed no strong correlation with the in vitro data (
Figure 11). Nevertheless, our results are in agreement with the literature data, as previous studies also found that enzyme–substrate interactions at this site are mediated mainly by the main-chain atoms of the substrate [
31]. The most preferred variants for which the obtained activity was ≥10% contained Phe, Tyr, Trp, Ser, Thr, or Ala residues in P1′ position (
Figure 5). Some P5, P4, P2, P1, and P2′ mutants were also tested in vitro and in silico (
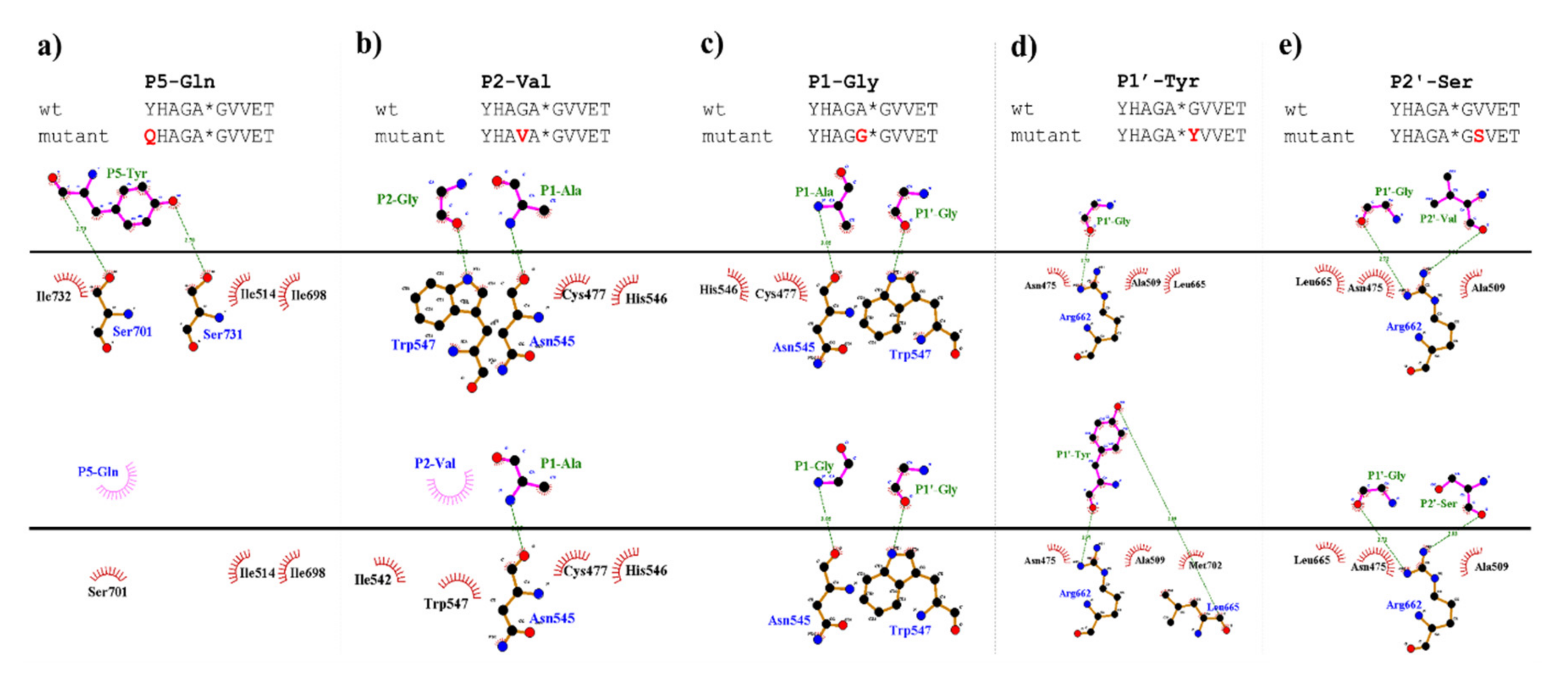
Table 1a) (
Figure 5). We found significantly lower preference for P5-Gln mutant as compared to the wild-type substrate; we assume that the side chain of the wild-type P5-Tyr forms H-bonds with S5 residues, while the same interactions are not enabled in the case of the P5-Gln variant.
The highest catalytic efficiency was obtained for VEEV nsP2pro-2 by using P4-Glu mutant substrate, in which Glu represents the wild-type P4 residue of VEEV nsP1/nsP2 and nsP2/nsP3 cleavage sites, and its side chain is supposed to connect to Lys-706 of the S4 subsite via a salt bridge [
5,
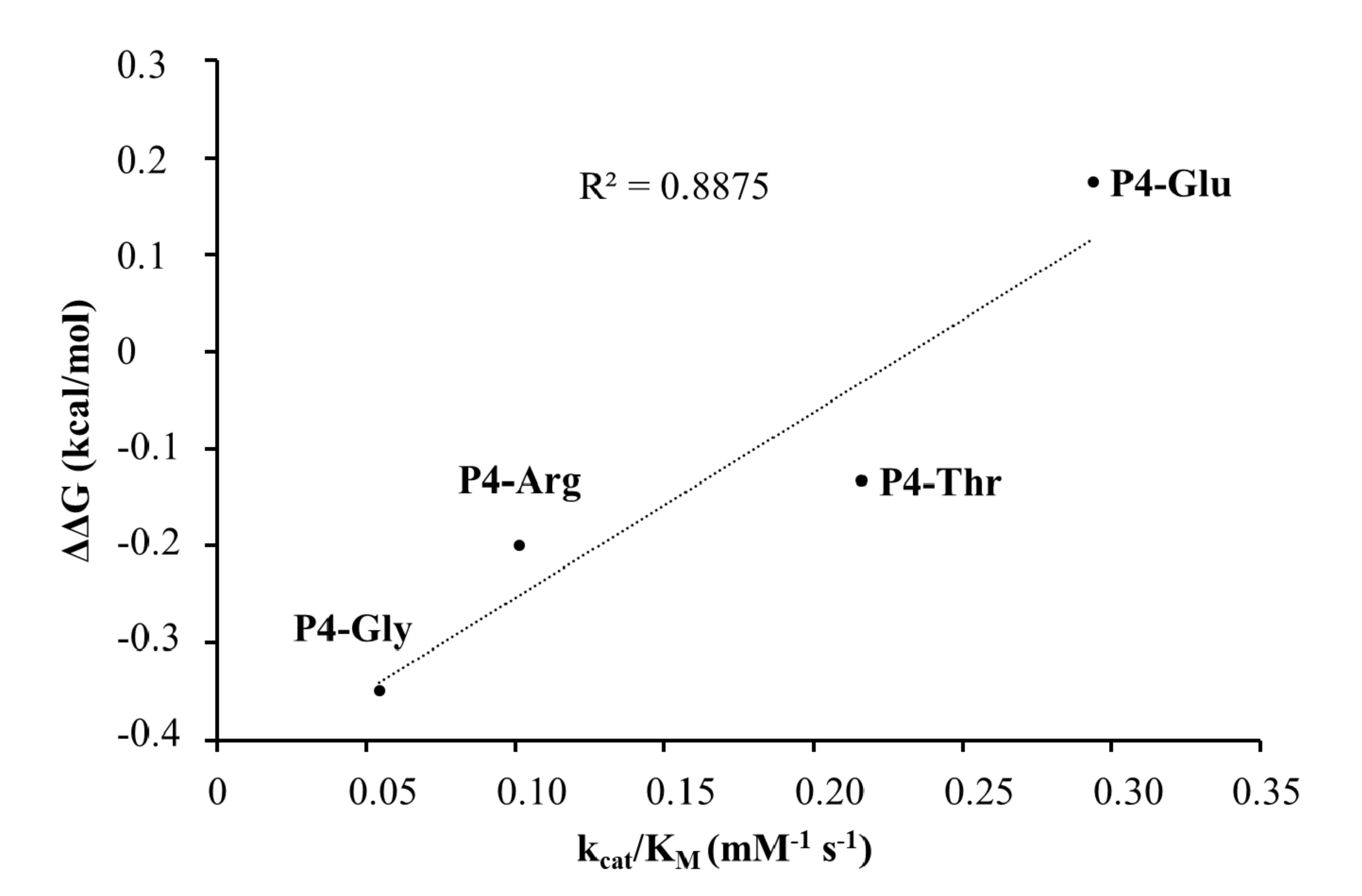
31]. For the other tested P4 variants (Thr, Arg, and Gly) the specific activity was significantly lower as compared to P4-Glu, most probably due to the lack of side-chain-mediated polar interactions at S4 site. Nevertheless, P4-Thr variant was processed with significantly higher efficiency in contrast to the wild type, while for P4-Arg and P4-Gly this difference was not significant. The in vitro determined k
cat/K
M values of the different P4 variants were in agreement with the previously reported dominance of polar interactions at this site [
31], and showed good correlation with the in silico predicted changes of energy (
Figure 10). SFV PR was found previously to be highly specific for P4-Arg which cannot be replaced by other residues in this position [
32]. While a P4-Thr residue prevented substrate processing by SFV PR [
32], we found that VEEV nsP2pro can cleave a substrate containing Thr in P4 position, thus, our results reveal that S4 site of VEEV nsP2pro has a wider tolerance as compared to that of SFV protease.
In the P2 position, wild-type VEEV cleavage sites contain a Gly residue, which is highly conserved across alphavirus species [
21] and is recognized by the “glycine specificity motif” of VEEV nsP2pro, similarly to the mechanism described for other cysteine proteases [
19]. The in vitro-specific activity rates for both P2-Val and P2-Ala variants were marginal, most probably owing to (i) the relatively larger sizes of Ala and Val residues and (ii) the lack of a H-bond formed by the backbone atom of Gly.
The specific activity of the P1-Gly variant was half that of the P1-Ala, that may be explained by the preference of hydrophobic S1-P1 interaction. Val has a larger volume and higher hydrophobicity compared to Ala, and accordingly, in silico analysis predicted more favorable interactions for this variant than for Gly; however, the in vitro-measured specific activity on this variant was marginal. This can be explained by the findings of Russo et al. [
31], that branched side chains have lower flexibility that is likely to cause less favorable binding to the S1 site. Additionally, they observed that covariances of residues suggest relatively high tolerance for substitutions at P1 site. In accordance with this, the in vitro-determined k
cat/K
M values did not differ significantly between the wild-type and the P1-Gly variants of SFV nsP12 cleavage site.
Both Lulla et al. [
32] and Russo et al. [
31] described that the substrate is mainly recognized by the S4-S1’ subsites of VEEV nsP2 protease, therefore, previous specificity studies focused mainly on the P4-P1′ preferences. Despite the importance of these sites, the complete screening of P1′ specificity has not been performed experimentally to date. Additionally, the data on the amino acid preferences of S2’ site are also limited. Based on the modeled complex, S2’ site is not a less well-defined pocket as P2′–P6′ substrate residues are more exposed to the solvent. The specific activities determined for P2′-Ser and P2′-Pro variants in vitro were comparable with the values obtained for the wild-type substrates containing P2′-Val residue, and the
kcat/K
M constants determined for the wild-type and P2′-Ser substrates were practically identical, despite differences of the hydrophobicities of these residues. This is in agreement with the results of Hu et al. [
5] who proposed favorable backbone-mediated interactions for the P1′–P6′ sites.
In summary, our results are in agreement with the literature data and complement knowledge about in vitro specificity of VEEV nsP2pro. While former studies used different approaches to study specificity (e.g., substrate-binding sites of the protease were targeted by mutagenesis [
5] or information about specificity was obtained based on different cleavage efficiencies of natural cleavage sites [
20]), here we prepared series of modified substrates for position screening. Furthermore, structural backgrounds of enzyme–substrate interactions were studied previously by others [
31], and the present study provided experimental evidence for the former findings. Considering the importance of alphaviruses from both economic and military aspects and the lack of any effective therapy against their infections, obviously there is a great demand for the further investigation of these viruses. Thus, our results may provide valuable information about amino acid preferences that may support future studies on inhibitor development against alphaviruses or other important Group IV viruses.
4. Materials and Methods
All materials were purchased from Sigma-Aldrich (St. Louis, MO, USA), unless otherwise indicated.
4.1. Cloning of VEEV nsP2 and VEEV nsP2pro-2 Expression Vectors
Plasmid encoding the cDNA of non-structural proteins of VEEV was a gift from Dr. Christine L. Pugh (United States Army Research Institute of Infectious Diseases). The ORFs of VEEV nsP2 coding for residues 1–785, and VEEV nsP2pro-2 encoding residues 436–785, have been cloned into both pDEST17 and pDEST-His
6-MBP destination vectors by Gateway Cloning Technology (Thermo Fisher Scientific, Waltham, MA, USA) according to Tropea et al. [
27]. The linear DNA sequences were amplified as follows: (i) The ORF of VEEV nsP2 was amplified using PE2685 and PE2686 primers for PCR according to the following reaction setups: 10 ng template, 5 nmol PE2685, 5 nmol PE2686, 10 µL Phusion Flash High Fidelity PCR Master Mix (New England Biolabs, Ipswich, MA, USA) and nuclease-free water up to 20 µL using the following PCR protocol: cycle1 (1×): 60 s at 98 °C; cycle2 (30×): 15 s at 98 °C, 75 s at 72 °C, and 60 s at 72 °C; cycle3 (1×): store at 4 °C. (ii) The ORF of VEEV nsp2pro-2 was amplified using PE2687 and PE2686 primers for PCR, according to the following reaction setups: 10 ng template, 5 nmol PE2685, 5 nmol PE2686, 10 µL Phusion Flash High Fidelity PCR Master Mix and nuclease-free water up to 20 µL using the following PCR protocol: cycle1 (1×): 60 s at 98 °C; cycle2 (30×): 10 s at 98 °C, 30 s at 72 °C, and 60 s at 72 °C; cycle3 (1×): store at 4 °C. The PCR products were purified by MinElute (Qiagen, Hilden, Germany) and 60–80 ng of the resulting PCR amplicons were subsequently used as the templates for another PCR with 5 nmol PE277, 5 nmol PE2686, 10 µL Phusion Flash High Fidelity PCR Master Mix and nuclease-free water up to 20 µL using the following PCR protocol (i) for VEEV nsP2: cycle1 (1×): 60 s at 98 °C; cycle2 (30×): 15 s at 98 °C, 75 s at 72 °C, and 60 s at 72 °C; cycle3 (1×): store at 4 °C; (ii) for VEEV nsp2pro-2: cycle1 (1×): 60 s at 98 °C; cycle2 (30×): 10 s at 98 °C, 30 s at 72 °C, and 60 s at 72 °C; cycle3 (1×): store at 4 °C. The linear DNA sequences were transferred via pDON221 donor vector (Invitrogen, Thermo Fisher Scientific, Waltham, MA, USA) into pDEST-His6-MBP (Invitrogen, Thermo Fisher Scientific, Waltham, MA, USA) and pDEST17 (Invitrogen, Thermo Fisher Scientific, Waltham, MA, USA) as described by Tropea et al. [
27]. The sequence the of entry clones was verified by capillary sequencing. The oligonucleotide primers applied during PCRs and sequencing reactions are listed in
Table S1. Expression vectors pBB2546 (pDEST-His
6-VEEVnsP2), pBB2547 (pDEST-His
6-VEEVnsP2pro-2), pBB2549 (pDEST-His
6-MBP-VEEVnsP2), and pBB2550 (pDEST-His
6-MBP-VEEVnsP2pro-2) were purified by Qiagen Miniprep Kit (Qiagen, Hilden, Germany).
4.2. Sequence Alignment
The primary sequences of VEEV nsP2 (NCBI Ref. Seq.: NP_740697.1), SINV (NCBI Ref. Seq: NP_740671.1), CHIKV (GenBank: ADZ47896.1), Sagiyama virus (GenBank: BAA92845.1) and SFV (NCBI Reference Sequence: NP_740666.1) were aligned using multiple sequence alignment tool Clustal Omega (EMBL-EBI, Hinxton, UK).
4.3. Generation of the Substrate Expression Plasmids
The generation of the pDEST-His
6-MBP-mEYFP empty plasmid and the insertion of the cleavage sites of interest to its cloning cassette were performed as described by Bozóki et al. [
23,
24] using pmEYFP-N1 vector as a template. Plasmid pmEYFP-N1 was generated previously in our laboratory by the modification of EYFP-N1, which was a kind gift of Prof. Thomas Jovin (Göttingen). The modification included the elimination of the EYFP dimerization surface by introducing A208K mutation [
37].
The inserted linear short dsDNA sequences coding for wild-type, P5, P4, P1, P2, and P1′ variants of SFV nsP12 cleavage site were generated either by annealing of chemically synthetized complement oligonucleotides primers (
Table S2) as described by Bozóki et al. [
23,
24] or by random mutagenesis on P1′ site as follows. pT7-Blue-3 plasmid (Merck-Millipore, Burlington, MA, USA) was linearized by BamHI and NheI restriction endonucleases (New England Biolabs, Ipswich, MA, USA). Oligonucleotide primers coding for wild-type SFV nsP12 (EYHAGA↓GVVETP) cleavage site sequence that were flanked by BamH I and NheI sticky ends, were inserted into pT7-Blue-3 plasmid. For ligation, 40 ng linearized pT7- Blue-3 plasmid was incubated with 800 ng SFV nsP12 BamHI forward primer (5′-GATCCTTAATTAAAGAGTACCATGCTGGTGCTGGTGTGGTGGAGACACCGG-3′) and 800 ng SFV nsP12 BamHI reverse primer (5′-CTAGCCGGTGTCTCCACCACACCAGCACCAGCATGGTACTCTTTAATTAAG-3′) for 2 min at 65 °C, then for 2 min at 4 °C. Hereafter, T4 DNA Ligase Reaction Buffer (10×) and T4 DNA ligase (New England Biolabs, Ipswich, MA, USA) were added, followed by incubation overnight at 16 °C. An amount of 100 µL of
E. coli DH5α-competent cells (New England Biolabs, Ipswich, MA, USA) was transformed by 5 µL of the ligation reaction mixture by heat shock (at 42 °C). The transformants were spread on Luria-Bertani (LB) agar plates containing 100 µg/mL ampicillin and were grown at 37 °C overnight. The transformed cells were then cultured in LB medium containing 100 µg/mL ampicillin followed by preparation of pT7-Blue-3-SFV nsP12 plasmid by Qiaprep Spin Mini Prep Kit (Qiagen, Hilden, Germany). Random mutagenesis of the triplets coding for the P1′ residue of SFV nsP12 was performed by QuickChange Lightning Multi-Site Directed Mutagenesis Kit (Agilent Technologies, Santa Clara, CA, USA) according to the manufacturer’s protocol using SFV nsP12 P1′ DEG oligonucleotide primer (5′-GAGTACCATGCTGGTGCTNNNGTGGTGGAGACACCGGCTAGC-3′). P1′-mutagenized pT7-Blue-3 SFV nsP12 plasmids were recovered from the cultures of grown colonies using Qiagen Plasmid Midi Kit (Qiagen, Hilden, Germany) and were linearized using NheI and PacI (New England Biolabs, Ipswich, MA, USA). The short dsDNA sequences coding for SFV nsP12 P1′ variants were separated on polyacrylamide gels containing 15% urea and were purified by Qiaex II Gel Extraction Kit (Qiagen, Hilden, Germany). The mutagenized dsDNA fragment (100 ng) was mixed with 200 ng linearized pDEST-His
6-MBP-mEYFP plasmid during the ligation reaction described above.
4.4. Expression of VEEV nsP2pro-2 and VEEV nsP2
For expression, freezer stocks of E. coli Rozetta cells were used that were transformed previously by pBB2546, pBB2547, pBB2549, or pBB2550 plasmids. An amount of 100 µL of freezer stock was added to 100 mL LB medium and incubated 16 h at 37 °C. Hereafter 30 mL of the starter culture was measured to 1000 mL LB medium containing 125 µg/mL ampicillin, 30 µg/mL chloramphenicol and 0.2% glucose. Cells were grown up to an OD600 = 0.5–0.6 at 37 °C, and expression was induced by 1 mM isopropyl β-d-1-thiogalactopyranoside IPTG (final concentration) and incubated for 4 h at 30 °C. Cells were harvested at 2000 g for 10 min using a tabletop centrifuge. After the removal of the medium, the cell pellets were stored at −80 °C. Other conditions were also tested to improve the expression and solubility of His6-MBP-VEEV nsP2, using pBB2549 construct. In this setup, when the 1000-mL culture reached OD600 = 0.5, the temperature was shifted to 18 °C, the culture was further incubated for 20 min and after induction and culture was incubated overnight at 18 °C.
To examine protein expression profile of the different constructs, cells were sampled right before induction (0 h), and prior to harvest (4 h or overnight). One ml from each sample was spun at 10,000 g for 5 min on a tabletop centrifuge and pellets were stored at −80 °C. The next day after thawing, pellets were re-suspended in distilled water.
For solubility profile analysis, 10 mL suspension right before cell harvest was centrifuged at 5000 g for 10 min in a tabletop centrifuge and the pellets were stored at −80 °C. The next day after thawing, cells were lysed with B-PER buffer (1.5 mL B-PER (Pierce, Thermo Fisher Scientific, Waltham, MA, USA), 3 µL lysozyme (2 mg/mL) and 3 µL DNase (New England Biolabs, Ipswich, MA, USA) for 15 min at room temperature. Samples taken from the crude cell lysate represented the total protein fraction, while cleared lysate represented the soluble protein fraction.
Samples were analyzed on 14% SDS-PAGE at reducing conditions.
4.5. Purification of VEEV nsP2pro-2
Bacterial cell pellet expressing His
6-MBP-VEEV nsP2pro-2 construct were suspended in 50 mL of ice-cold buffer A (50 mM sodium phosphate, 150 mM NaCl, 25 mM imidazole, pH 7.5). The cells were disrupted by APV Gaulin Model G homogenizer (Invensys, Albertslund, Denmark) at 10,000 psi and centrifuged at 30,000
g for 30 min at 4 °C. The supernatant was filtered through a 0.22-µm polyethersulfone membrane and was applied to a 15-mL (3 × 5 mL) HisTrap FF crude affinity column (GE Healthcare, Piscataway, NJ, USA) equilibrated in buffer A. The elution was performed with a linear gradient from 5–50% buffer B (50 mM sodium phosphate, 150 mM NaCl, 500 mM imidazole, pH 7.5) and eluate fractions containing the His
6-MBP-VEEV nsP2pro-2 were analyzed by reducing SDS-PAGE. Fractions with the highest concentration were pooled and concentrated with Amicon YM30 membrane (Millipore, Billerica, MA, USA), then diluted 6-fold with 50 mM sodium phosphate buffer (containing 150 mM NaCl, pH 7.5) to reduce the imidazole concentration to 25 mM. The fusion protein was then processed with a 5 mg/mL stock solution of the His-tagged TEV PR (70:1
v/
v) overnight at 4 °C. Hereafter, the cleavage products were applied to a 20-mL (4 × 5 mL) HisTrap FF crude affinity column equilibrated in buffer A. Both flow-through, containing the VEEV nsP2pro protease, and eluate fractions were collected. Flow-through fractions were analyzed on SDS-PAGE at reducing conditions and those that contained a high amount of VEEV nsP2pro-2 were pooled. The sample was concentrated to 10 mg/mL using an Amicon YM30 membrane (Millipore, Burlington, MA, USA) and applied to a HiPrep 26/60 Sephacryl S200 gel filtration column (GE Healthcare, Chicago, IL, USA) equilibrated with 50 mM Tris-HCl buffer (containing 150 mM NaCl, 2 mM Tris (2-carboxyethyl) phosphine hydrochloride (TCEP), pH 7.5). The peak fractions comprising VEEV nsp2pro-2 protease were analyzed on reducing SDS-PAGE, pooled and concentrated to 1–5 mg/mL. The concentration of VEEV nsp2pro-2 was determined as 2.9 mg/mL by A = 280 method. The theoretical extinction coefficient and molecular weight were calculated based on the primary structure of the proteins by ProtParam tool of ExPASy [
38]. Aliquots were flash-frozen with liquid nitrogen and stored at −80 °C until further use.
4.6. Expression and Purification of the Substrates
The procedure of recombinant substrate expression and purification described by Bozóki et al. [
23,
24] was optimized to a small-scale to support microplate-based specificity screening measurements. In this setup, the following parameters were changed compared to the original protocol: (i) 2.5 mL starter culture was added to 15 mL LB containing 100 µg/mL ampicillin in a 50-mL centrifuge tube; (ii) cells were grown at OD
600 = 0.5–0.6; (iii) after IPTG induction, cells were incubated for 4 h at 37 °C; (iv) the volume of the lysis buffer was reduced to 1 mL.
His
6-MBP-mEYFP substrates representing the wild-type, P4-Glu, P4-Thr, P4-Arg, P4-Gly, P1-Gly, P1′-Thr, and P2′-Ser variants of SFV nsP12 cleavage site were expressed and purified based on the previously described protocol. After the induction of expression by IPTG, the incubation was followed by addition of tetracycline to the cell suspensions (200 µg/mL final concentration) to arrest protein translation and enable improved folding of protein substrates [
24]. After the treatment cell were incubated for 2 h at 37°C, followed by harvesting the cells.
4.7. Gel Electrophoresis
Wild-type His
6-MBP-EYHAGA↓GVVETP-mEYFP substrates attached to Ni-NTA magnetic agarose beads (SAMBs) were suspended in elution buffer (50 mM NaH
2PO
4, 300 mM NaCl, 500 mM imidazole, 0.05% Tween 20, pH 8.0) or cleavage buffer (50 mM NaH
2PO
4, 300 mM NaCl, 0.05% Tween 20, pH 7.5). SAMBs in cleavage buffer were digested by TEV PR (at 7.1 µM final concentration) and VEEV nsP2pro-2 (at 5.2 µM final concentration) on 30 °C overnight. TEV (S219V) PR stock solution was purified according to Kapust et al. [
28].
For native PAGE analysis, samples were prepared from 10 µL of each reaction with 2× loading buffer (62.5 mM Tris-HCI, pH 6.8, 25% glycerol, 0.01% bromophenol Blue). After electrophoresis, the gels were illuminated by using Dark Reader Blue Transilluminator (Labgene Scientific, Châtel-Saint-Denis, Switzerland), and subsequently proteins were detected by PageBlue Protein Staining Solution (Thermo Fisher Scientific, Waltham, MA, USA).
For reducing SDS-PAGE, samples were supplemented with 6x loading buffer (300 mM Tris, pH 6.8, 20% glycerol, 0.05% bromophenol blue, 12% SDS, 100 mM β-mercaptoethanol), denatured at 95 °C for 10 min, followed by electrophoresis. Proteins were detected in the gels by PageBlue Protein Staining Solution (Thermo Fisher Scientific, Waltham, MA, USA).
4.8. Microplate-Based Specificity Studies of VEEV nsp2pro-2
The specificity of VEEV nsP2pro-2 was studied by recombinant substrates containing the cleavage site sequences listed in
Table 1a. We applied a 96-well microplate-based adaptation of the Ni-NTA magnetic bead-based assay platform described previously by Bozóki et al. [
24]. For cleavage reactions, VEEV nsP2pro-2 (2.1–6.0 µM final concentration) was incubated with His
6-MBP-mEYFP substrates (1–5 µM final concentration) at 30 °C while continuously shaking at 600 rpm. Incubation time was 40 min in the case of P5, P4, P1, and P2′ variants, while it was 20 h for P1′-modified substrates. The pH of the applied buffers was 7.5. Flat-bottom 96-well plates were applied, using 96-Well Magnet Type A magnetic particle concentrator (Qiagen, Hilden, Germany). The final volume of the assay samples was 70 µL per well. During incubation and washing of the beads, plates were shaken by a digital shaker (IKA MS3). The detection of the fluorescence the products and substrates were performed by using either Victor
2 Wallac 1420 device with 544/15 nm excitation and 590/10 emission filters or Biotek Synergy H1 device at 510 nm excitation and 540 nm emission wavelengths.
Statistical analysis of differences was performed by GraphPad unpaired
t-test (
https://www.graphpad.com/quickcalcs/ttest1/). With the exception of P1′ variants, one-way analysis of variances (ANOVA) was also applied. In the case of an unpaired t-test,
p-value < 0.05 was considered to be statistically significant, while in the case of one-way ANOVA, the null hypothesis (H
0), namely that the means of the tested populations (μ
1 and μ
2) are equal (H
0: μ
1 = μ
2), was rejected if F > F
critical. 4.9. Kinetic Studies of VEEV nsP2pro-2
Kinetic measurements were carried out using the microtube-based Ni-NTA magnetic bead-based assay platform developed previously by Bozóki et al. [
24].
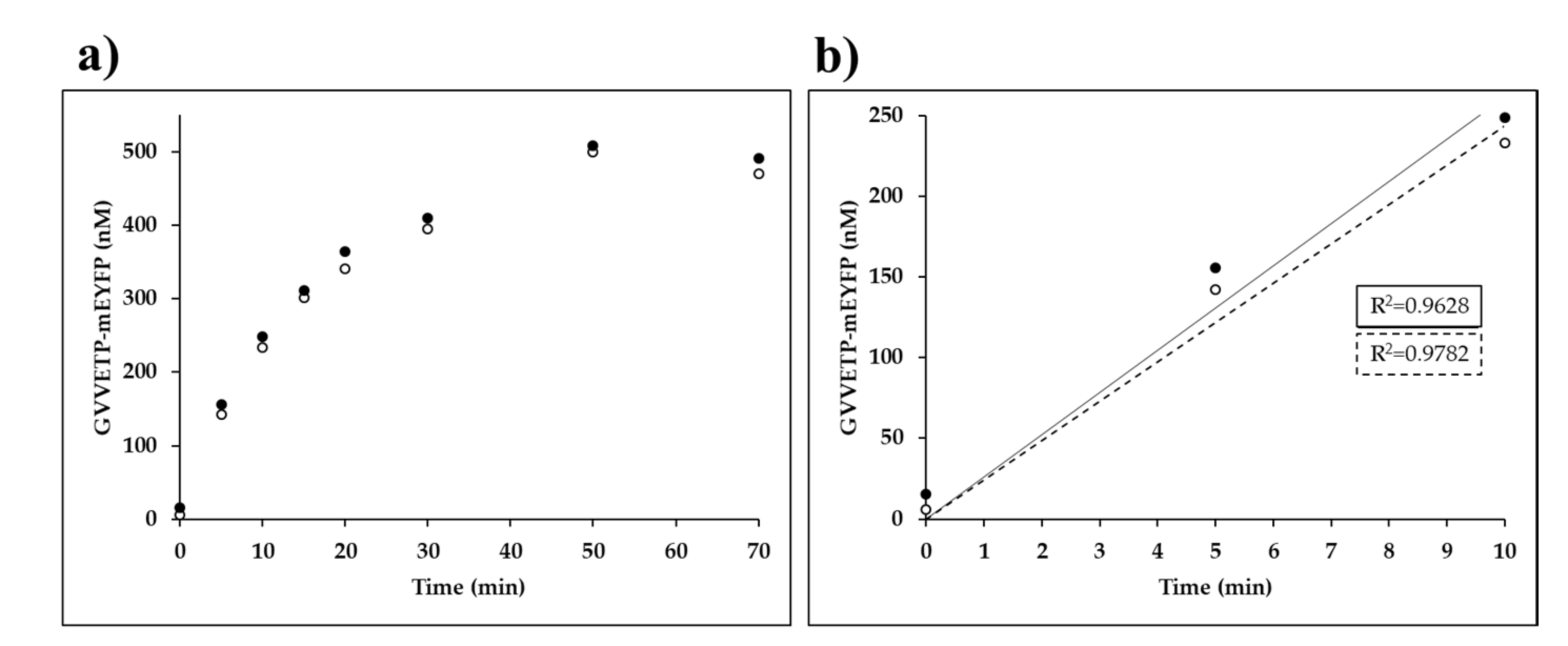
Time-course measurements were performed on His6-MBP-EYHAGA↓GVVETP-mEYFP substrate (at 0.02 mM final concentration) processed by VEEV nsP2pro-2 (at 3.0 µM final total concentration). The product formation was followed by measuring fluorescence after for 0, 5, 10, 15, 20, 30, 50, 70, and 90 min at 30 °C, while continuously shaking at 600 rpm. Molar concentration of the C-terminal fluorescent cleavage product in the reaction samples were plotted against time (min). Linear regression was performed, and the parameters of the fitted lines were determined by GraphPad Prism version 5.00.
Substrate-dependent kinetic studies were performed on substrates at VEEVnsP2pro-2 different enzyme concentrations listed in
Table 2. The samples were incubated for 10 min at 30 °C, while continuously shaking at 600 rpm. Kinetic parameters were determined at <20% substrate turnover by Michaelis–Menten non-linear regression analysis using GraphPad Prism version 5.00 for Windows (GraphPad Software, La Jolla, CA, USA,
www.graphpad.com). The standard deviation values for the k
cat/K
M vales were calculated based on Boross et al. [
39]. Significance of the differences between the k
cat/K
M values of the tested variants and the wild-type substrate were determined by GraphPad unpaired
t-test (
https://www.graphpad.com/quickcalcs/ttest1/).
p-value < 0.05 was considered to be statistically significant.
4.10. In Silico Structural Analyses
The structural analyses were performed based on a modeled complex of VEEV nsP2 protease and LQEAGA↓GSVETP substrate [
5]; the coordinate file was kindly provided by Patricia M. Legler (Center for Bio/molecular Science and Engineering, U.S. Naval Research Laboratory).
The sequence of the original model and the experimentally examined protein was aligned with Clustal Omega [
40]. Both the model protein and substrate found in the crystal structure were mutated to reproduce the in vitro structures, by applying the FoldX software [
41]. To the N-termini of both molecules, an N-acetyl group was connected, while a methylamide group was added to the C-termini. The protonation states were attributed by the Chimera software [
42], the active site contained a deprotonated cysteine and a protonated histidine residue. Prior to the simulations, the systems were immersed in water. For the simulations, the Amber16 software [
43] was used. The protein and the substrate were parametrized by the FF14SB force field [
44], and the surrounding water molecules were described by the TIP3P model [
45]. For the simulations, the system was first minimized via the steepest descent method for 5000 steps, followed by the conjugate gradient method for a further 5000 steps. The system was gradually heated to 300 K during a 2-ns-long simulation, and equilibrated for 2 ns at 300 K, applying 1-fs timesteps. For temperature regulation, the Langevin dynamics method was used. For bonds involving hydrogen atoms, SHAKE was performed. The system was further heated to 400 K during a 0.5-ns-long simulation, and equilibrated for 0.5 ns at 400 K. A simulated annealing cycle was applied to the system, cooling it to 5 K during a 1-ns-long simulation, followed by heating to 400 K during 0.5 ns, and equilibrating at that temperature for 0.5 ns, under NPT conditions. The cycle was repeated two times, heating the system to 300 K in the second cycle. A further 0.5-ns-long equilibration was run at 300 K. During the simulations, harmonic restraints of 50 kcal mol
−1 Å
−2 were applied between the sulfur atom of the catalytic cysteine and the substrate carbonyl oxygen of the Ala residue at the cleavage site, and also between the donor nitrogen atom of the catalytic histidine and the backbone nitrogen of the Gly residue at the cleavage site. Convergence during the last equilibration simulation was monitored with RMSD calculations, performed by the cpptraj [
46] program of Amber16, with the average of coordinates over the equilibration trajectory used as reference.
DynaMut web server was applied to prepare mutant complexes and predict energy changes upon point mutations of substrate residues, the ≥0 kcal/mol change in folding free energy (ΔΔG) was considered to be stabilizing, while ΔΔG < 0 was viewed as destabilizing [
47]. Hydropathy plots were prepared for the cleavage site sequences (P5–P5′) using the ProtScale module of ExPASy web server (
https://web.expasy.org/protscale) [
38], setting a 3-residue window for calculation and using the Kyte and Doolittle hydropathicity scale [
48]. Residue volumes were retrieved from the literature [
49]. LigPlot+_v2.1 program was used for 2D visualization of interaction [
50]. Structural figures were prepared by using PyMOL Molecular Graphics System (Version 1.3 Schrödinger, LLC).