Diversity of Parallel Guanine Quadruplexes Induced by Guanine Substitutions
Abstract
1. Introduction
2. Results and Discussion
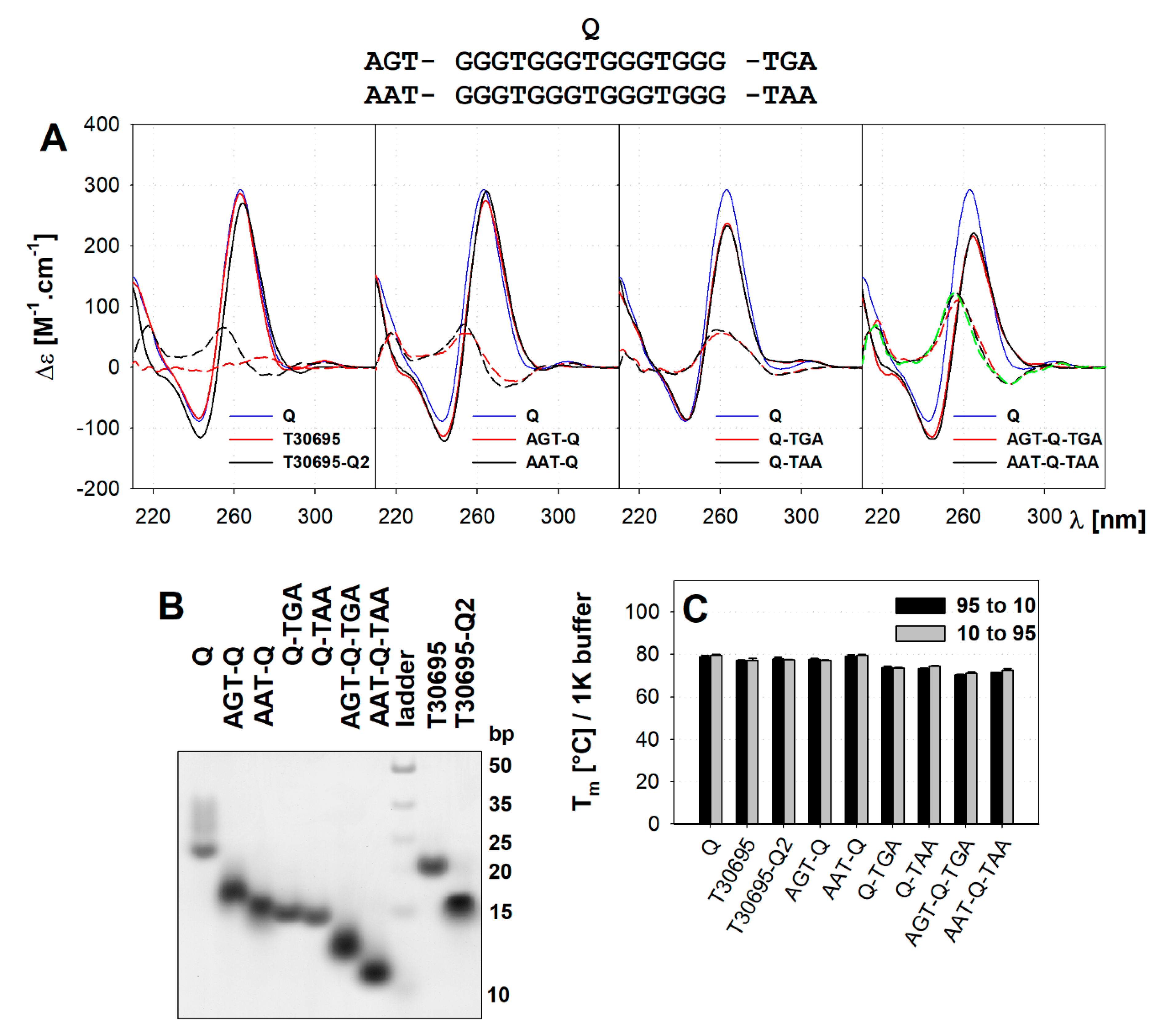
2.1. The Multimerisation of Parallel Quadruplexes
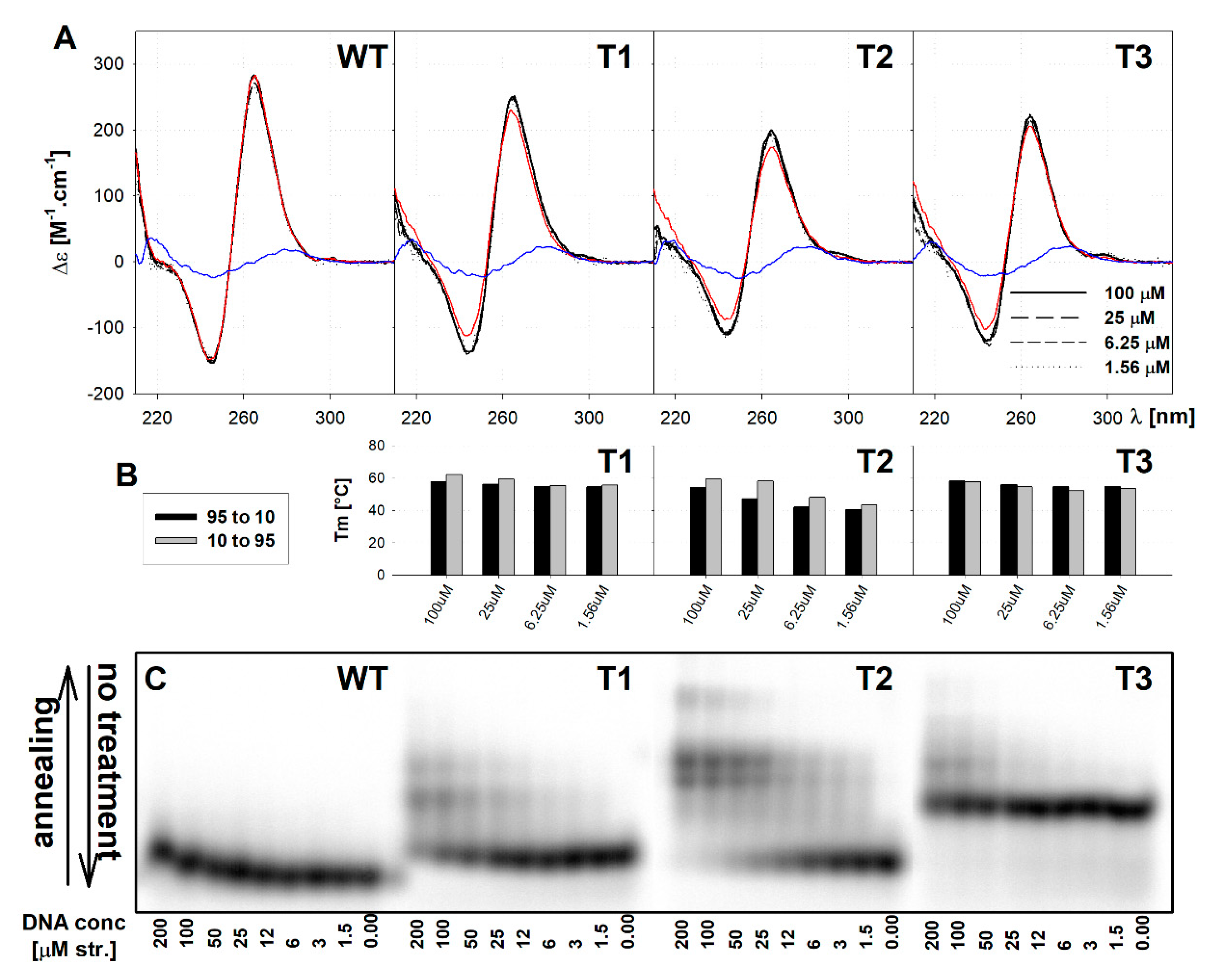
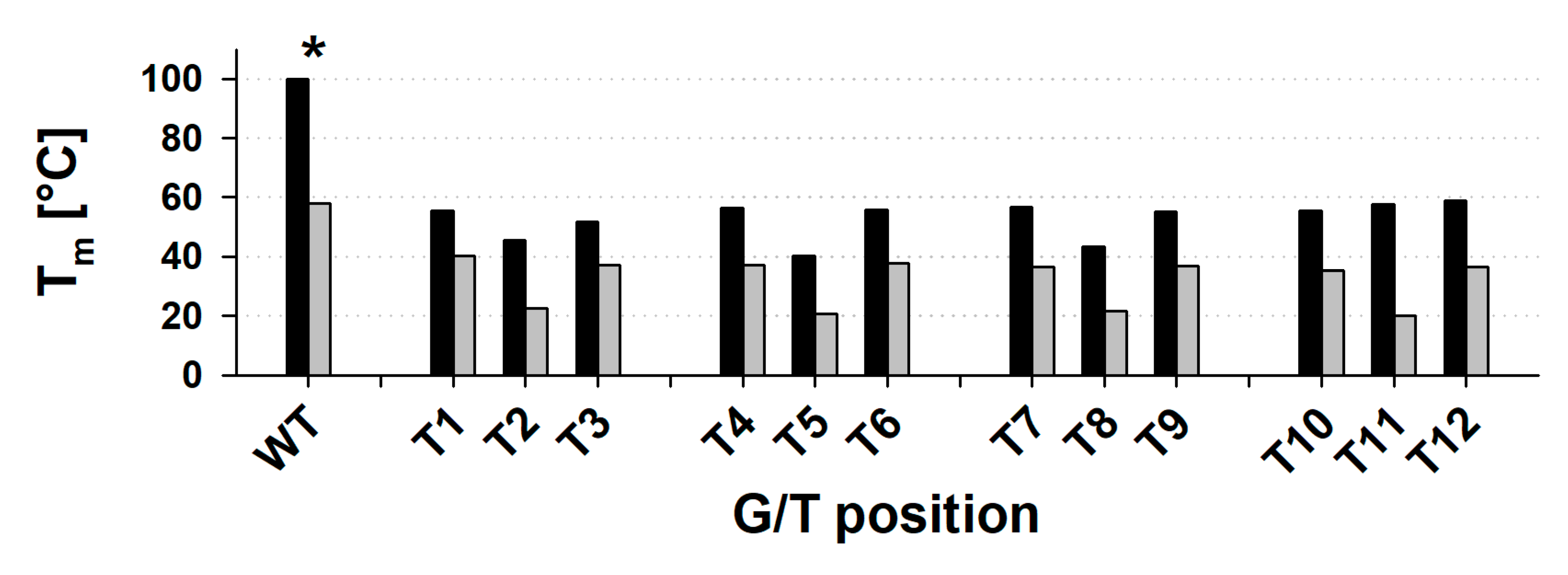
2.2. The Mutated Parallel Variants Are Uniform in Some Aspects and Diverse in Others
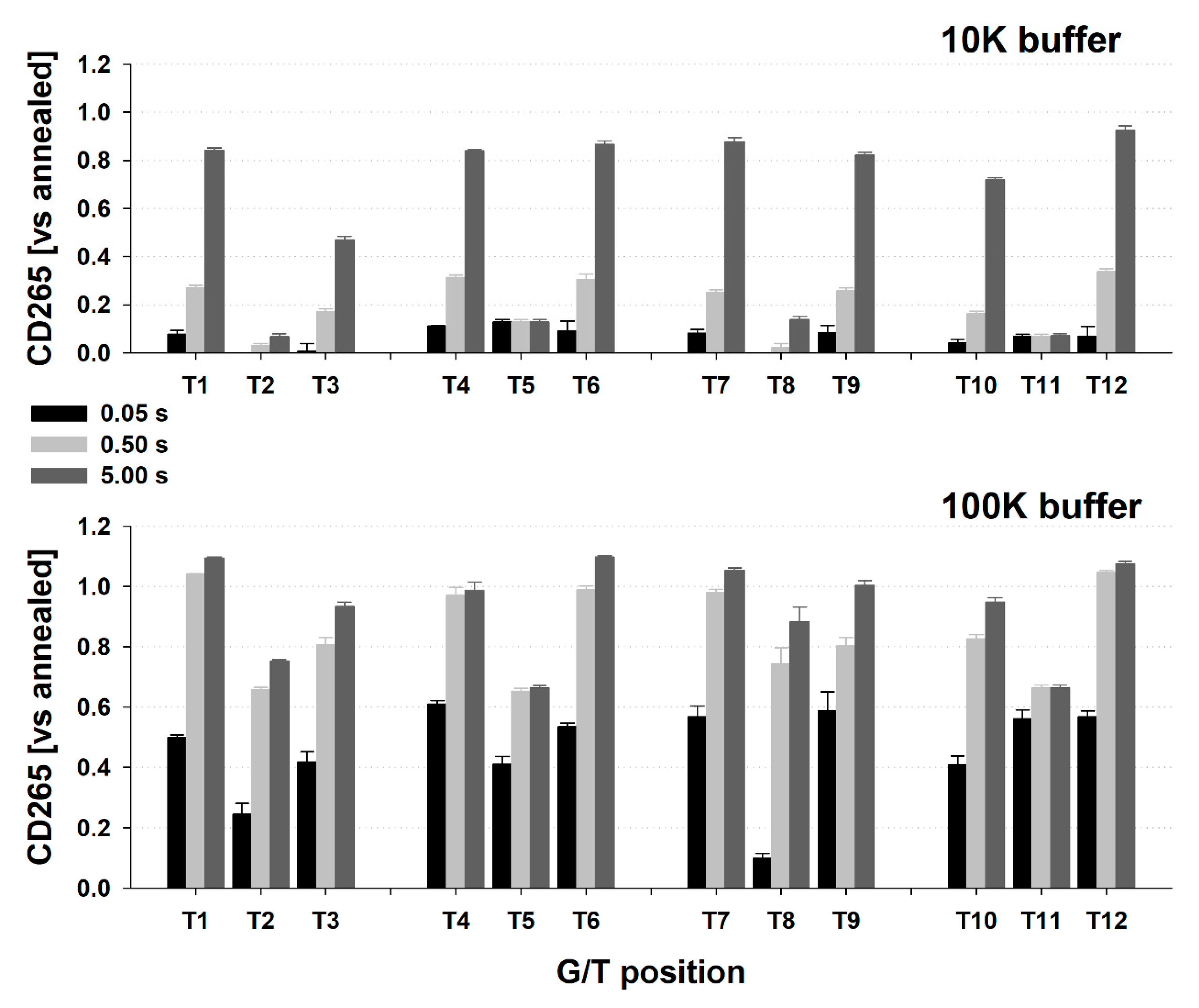
2.3. The DNA Concentration-Dependent Formation of Type 2 Multimolecular Structures Is Not Directly Reflected in CD
2.4. Parallel Quadruplexes Are Significantly More Affected by Guanine Lesions than the Antiparallel Ones
3. Materials and Methods
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| G4 | Guanine quadruplex |
| CD | Circular dichroism |
| PAGE | Polyacrylamide gel electrophoresis |
| SF | Stopped-flow |
References
- Neidle, S.; Balasubramanian, S. Quadruplex Nucleic Acids; Royal Society of Chemistry: London, Cambridge, UK, 2006; pp. 1–3. [Google Scholar]
- Miyoshi, D.; Fujimoto, T.; Sugimoto, N. Molecular Crowding and Hydration Regulating of G-Quadruplex Formation. Quadruplex Nucleic Acids 2012, 330, 87–110. [Google Scholar] [CrossRef]
- Zhou, J.; Tateishi-Karimata, H.; Mergny, J.; Cheng, M.; Feng, Z.; Miyoshi, D.; Sugimoto, N.; Li, C. Reevaluation of the stability of G-quadruplex structures under crowding conditions. Biochimie 2016, 121, 204–208. [Google Scholar] [CrossRef] [PubMed]
- Huppert, J.; Balasubramanian, S. Prevalence of quadruplexes in the human genome. Nucleic Acids Res. 2005, 33, 2908–2916. [Google Scholar] [CrossRef] [PubMed]
- Marsico, G.; Chambers, V.S.; Sahakyan, A.B.; McCauley, P.; Boutell, J.M.; Di Antonio, M.; Balasubramanian, S. Whole genome experimental maps of DNA G-quadruplexes in multiple species. Nucleic Acids Res. 2019, 47, 3862–3874. [Google Scholar] [CrossRef]
- Biffi, G.; Tannahill, D.; McCafferty, J.; Balasubramanian, S. Quantitative visualization of DNA G-quadruplex structures in human cells. Nat. Chem. 2013, 5, 182–186. [Google Scholar] [CrossRef]
- Varizhuk, A.M.; Ischenko, D.; Tsvetkov, V.B.; Novikov, R.; Kulemin, N.A.; Kaluzhny, D.; Vlasenok, M.; Naumov, V.; Smirnov, I.; Pozmogova, G. The expanding repertoire of G4 DNA structures. Biochimie 2017, 135, 54–62. [Google Scholar] [CrossRef]
- Lightfoot, H.L.; Hagen, T.; Tatum, N.J.; Hall, J. The diverse structural landscape of quadruplexes. FEBS Lett. 2019, 593, 2083–2102. [Google Scholar] [CrossRef]
- Konvalinová, H.; Dvorakova, Z.; Renčiuk, D.; Bednářová, K.; Kejnovska, I.; Trantírek, L.; Vorlíčková, M.; Sagi, J. Diverse effects of naturally occurring base lesions on the structure and stability of the human telomere DNA quadruplex. Biochimie 2015, 118, 15–25. [Google Scholar] [CrossRef]
- Sagi, J. G-quadruplexes incorporating modified constituents: A review. J. Biomol. Struct. Dyn. 2013, 32, 477–511. [Google Scholar] [CrossRef]
- Esposito, V.; Martino, L.; Citarella, G.; Virgilio, A.; Mayol, L.; Giancola, C.; Galeone, A. Effects of abasic sites on structural, thermodynamic and kinetic properties of quadruplex structures. Nucleic Acids Res. 2009, 38, 2069–2080. [Google Scholar] [CrossRef]
- Kelley, S.; Boroda, S.; Musier-Forsyth, K.; Kankia, B.I. HIV-integrase aptamer folds into a parallel quadruplex: A thermodynamic study. Biophys. Chem. 2011, 155, 82–88. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.; Amrane, S.; Rosu, F.; Salgado, G.F.; Bian, Y.; Tateishi-Karimata, H.; Largy, E.; Korkut, D.N.; Bourdoncle, A.; Miyoshi, D.; et al. Unexpected Position-Dependent Effects of Ribose G-Quartets in G-Quadruplexes. J. Am. Chem. Soc. 2017, 139, 7768–7779. [Google Scholar] [CrossRef] [PubMed]
- Lombardi, E.P.; Holmes, A.; Verga, D.; Teulade-Fichou, M.-P.; Nicolas, A.; Londoño-Vallejo, A. Thermodynamically stable and genetically unstable G-quadruplexes are depleted in genomes across species. Nucleic Acids Res. 2019, 47, 6098–6113. [Google Scholar] [CrossRef]
- Heddi, B.; Martin-Pintado, N.; Serimbetov, Z.; Kari, T.M.A.; Phan, A.T. G-quadruplexes with (4n − 1) guanines in the G-tetrad core: Formation of a G-triad·water complex and implication for small-molecule binding. Nucleic Acids Res. 2015, 44, 910–916. [Google Scholar] [CrossRef] [PubMed]
- Mukundan, V.T.; Do, N.Q.; Phan, A.T. HIV-1 integrase inhibitor T30177 forms a stacked dimeric G-quadruplex structure containing bulges. Nucleic Acids Res. 2011, 39, 8984–8991. [Google Scholar] [CrossRef]
- Mukundan, V.T.; Phan, A.T. Bulges in G-Quadruplexes: Broadening the Definition of G-Quadruplex-Forming Sequences. J. Am. Chem. Soc. 2013, 135, 5017–5028. [Google Scholar] [CrossRef]
- An, N.; Fleming, A.M.; Burrows, C.J. Human Telomere G-Quadruplexes with Five Repeats Accommodate 8-Oxo-7,8-dihydroguanine by Looping out the DNA Damage. ACS Chem. Boil. 2015, 11, 500–507. [Google Scholar] [CrossRef]
- Beckett, J.; Burns, J.; Broxson, C.; Tornaletti, S. Spontaneous DNA Lesions Modulate DNA Structural Transitions Occurring at Nuclease Hypersensitive Element III1 of the Human c-myc Proto-Oncogene. Biochemie 2012, 51, 5257–5268. [Google Scholar] [CrossRef]
- Dvořáková, Z.; Vorlíčková, M.; Renčiuk, D. Spectroscopic insights into quadruplexes of five-repeat telomere DNA sequences upon G-block damage. Biochim. Biophys. Acta (BBA) Gen. Subj. 2017, 1861, 2750–2757. [Google Scholar] [CrossRef]
- Fleming, A.M.; Zhou, J.; Wallace, S.S.; Burrows, C.J. A Role for the Fifth G-Track in G-Quadruplex Forming Oncogene Promoter Sequences during Oxidative Stress: Do These “Spare Tires” Have an Evolved Function? ACS Central Sci. 2015, 1, 226–233. [Google Scholar] [CrossRef]
- Islam, B.; Stadlbauer, P.; Vorlickova, M.; Mergny, J.; Otyepka, M.; Sponer, J. Stability of Two-Quartet G-Quadruplexes and Their Dimers in Atomistic Simulations. J. Chem. Theory Comput. 2020, 16, 3447–3463. [Google Scholar] [CrossRef] [PubMed]
- Guédin, A.; De Cian, A.; Gros, J.; Lacroix, L.; Mergny, J. Sequence effects in single-base loops for quadruplexes. Biochimie 2008, 90, 686–696. [Google Scholar] [CrossRef] [PubMed]
- Kolesnikova, S.; Hubálek, M.; Bednárová, L.; Cvačka, J.; Curtis, E.A. Multimerization rules for G-quadruplexes. Nucleic Acids Res. 2017, 45, 8684–8696. [Google Scholar] [CrossRef]
- Kolesnikova, S.; Curtis, E.A. Structure and Function of Multimeric G-Quadruplexes. Molecules 2019, 24, 3074. [Google Scholar] [CrossRef]
- Kogut, M.; Kleist, C.; Czub, J. Why do G-quadruplexes dimerize through the 5′-ends? Driving forces for G4 DNA dimerization examined in atomic detail. PLoS Comput. Biol. 2019, 15, e1007383. [Google Scholar] [CrossRef] [PubMed]
- Krishnan-Ghosh, Y.; Liu, D.; Balasubramanian, S. Formation of an Interlocked Quadruplex Dimer by d(GGGT). J. Am. Chem. Soc. 2004, 126, 11009–11016. [Google Scholar] [CrossRef]
- Do, N.Q.; Lim, K.W.; Teo, M.H.; Heddi, B.; Phan, A.T. Stacking of G-quadruplexes: NMR structure of a G-rich oligonucleotide with potential anti-HIV and anticancer activity†. Nucleic Acids Res. 2011, 39, 9448–9457. [Google Scholar] [CrossRef]
- Do, N.Q.; Phan, A.T. Monomer–Dimer Equilibrium for the 5′–5′ Stacking of Propeller-Type Parallel-Stranded G-Quadruplexes: NMR Structural Study. Chem. Eur. J. 2012, 18, 14752–14759. [Google Scholar] [CrossRef]
- Renčiuk, D.; Ryneš, J.; Kejnovská, I.; Foldynová-Trantírková, S.; Andäng, M.; Trantírek, L.; Vorlíčková, M. G-quadruplex formation in the Oct4 promoter positively regulates Oct4 expression. Biochim. Biophys. Acta (BBA) Bioenerg. 2017, 1860, 175–183. [Google Scholar] [CrossRef]
- Bednářová, K.; Kejnovská, I.; Vorlíčková, M.; Renčiuk, D. Guanine Substitutions Prevent Conformational Switch from Antiparallel to Parallel G-Quadruplex. Chem. Eur. J. 2019, 25, 13422–13428. [Google Scholar] [CrossRef]
- Jing, N.; Rando, R.F.; Pommier, Y.; Hogan, M.E. Ion Selective Folding of Loop Domains in a Potent Anti-HIV Oligonucleotide. Biochemie 1997, 36, 12498–12505. [Google Scholar] [CrossRef] [PubMed]
- Varizhuk, A.M.; Protopopova, A.D.; Tsvetkov, V.B.; Barinov, N.; Podgorsky, V.; Tankevich, M.V.; A Vlasenok, M.; Severov, V.V.; Smirnov, I.P.; Dubrovin, E.V.; et al. Polymorphism of G4 associates: From stacks to wires via interlocks. Nucleic Acids Res. 2018, 46, 8978–8992. [Google Scholar] [CrossRef] [PubMed]
- Kejnovská, I.; Bednárová, K.; Renciuk, D.; Dvoráková, Z.; Školáková, P.; Trantírek, L.; Fiala, R.; Vorlícková, M.; Sagi, J. Clustered abasic lesions profoundly change the structure and stability of human telomeric G-quadruplexes. Nucleic Acids Res. 2017, 45, 4294–4305. [Google Scholar] [CrossRef] [PubMed]
- Material not intended for publication: Bednářová, K. and Renčiuk, D. Institute of Biophysics of the Czech Academy of Sciences, Královopolská 135, Brno 612 65, Czech Republic. Stopped-flow-assisted circular dichroism and UV absorption measurements of formation kinetics of guanine quadruplexes. 2019.
- Šponer, J.; Bussi, G.; Stadlbauer, P.; Kührová, P.; Banas, P.; Islam, B.; Haider, S.M.; Neidle, S.; Otyepka, M. Folding of guanine quadruplex molecules–funnel-like mechanism or kinetic partitioning? An overview from MD simulation studies. Biochim. Biophys. Acta (BBA) Gen. Subj. 2017, 1861, 1246–1263. [Google Scholar] [CrossRef]
- Gray, R.D.; Trent, J.O.; Arumugam, S.; Chaires, J.B. Folding Landscape of a Parallel G-Quadruplex. J. Phys. Chem. Lett. 2019, 10, 1146–1151. [Google Scholar] [CrossRef]
- Mergny, J.; Phan, A.T.; Lacroix, L. Following G-quartet formation by UV-spectroscopy. FEBS Lett. 1998, 435, 74–78. [Google Scholar] [CrossRef]
- Zhang, A.Y.Q.; Balasubramanian, S. The Kinetics and Folding Pathways of Intramolecular G-Quadruplex Nucleic Acids. J. Am. Chem. Soc. 2012, 134, 19297–19308. [Google Scholar] [CrossRef]
- Yett, A.; Lin, L.Y.; Beseiso, D.; Miao, J.; Yatsunyk, L.A. N-methyl mesoporphyrin IX as a highly selective light-up probe for G-quadruplex DNA. J. Porphyrins Phthalocyanines 2019, 23, 1195–1215. [Google Scholar] [CrossRef]
- Nicoludis, J.M.; Miller, S.T.; Jeffrey, P.D.; Barrett, S.P.; Rablen, P.R.; Lawton, T.J.; Yatsunyk, L.A. Optimized End-Stacking Provides Specificity of N-Methyl Mesoporphyrin IX for Human Telomeric G-Quadruplex DNA. J. Am. Chem. Soc. 2012, 134, 20446–20456. [Google Scholar] [CrossRef]
- Harkness, R.W.; Mittermaier, A.K. G-register exchange dynamics in guanine quadruplexes. Nucleic Acids Res. 2016, 44, 3481–3494. [Google Scholar] [CrossRef]
- Guédin, A.; Gros, J.; Alberti, P.; Mergny, J. How long is too long? Effects of loop size on G-quadruplex stability. Nucleic Acids Res. 2010, 38, 7858–7868. [Google Scholar] [CrossRef] [PubMed]
- Hazel, P.; Huppert, J.; Balasubramanian, S.; Neidle, S. Loop-Length-Dependent Folding of G-Quadruplexes. J. Am. Chem. Soc. 2004, 126, 16405–16415. [Google Scholar] [CrossRef] [PubMed]
- Kejnovská, I.; Renčiuk, D.; Palacký, J.; Vorlíčková, M. CD Study of the G-Quadruplex Conformation. In Methods in Molecular Biology; Springer Science and Business Media LLC: Berlin, Germany, 2019; Volume 2035, pp. 25–44. [Google Scholar]
- Mergny, J.; Lacroix, L. Analysis of Thermal Melting Curves. Oligonucleotides 2003, 13, 515–537. [Google Scholar] [CrossRef] [PubMed]

© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bednářová, K.; Vorlíčková, M.; Renčiuk, D. Diversity of Parallel Guanine Quadruplexes Induced by Guanine Substitutions. Int. J. Mol. Sci. 2020, 21, 6123. https://doi.org/10.3390/ijms21176123
Bednářová K, Vorlíčková M, Renčiuk D. Diversity of Parallel Guanine Quadruplexes Induced by Guanine Substitutions. International Journal of Molecular Sciences. 2020; 21(17):6123. https://doi.org/10.3390/ijms21176123
Chicago/Turabian StyleBednářová, Klára, Michaela Vorlíčková, and Daniel Renčiuk. 2020. "Diversity of Parallel Guanine Quadruplexes Induced by Guanine Substitutions" International Journal of Molecular Sciences 21, no. 17: 6123. https://doi.org/10.3390/ijms21176123
APA StyleBednářová, K., Vorlíčková, M., & Renčiuk, D. (2020). Diversity of Parallel Guanine Quadruplexes Induced by Guanine Substitutions. International Journal of Molecular Sciences, 21(17), 6123. https://doi.org/10.3390/ijms21176123




