Cross-Linked Self-Assembling Peptides and Their Post-Assembly Functionalization via One-Pot and In Situ Gelation System
Abstract
1. Introduction
2. Results and Discussion
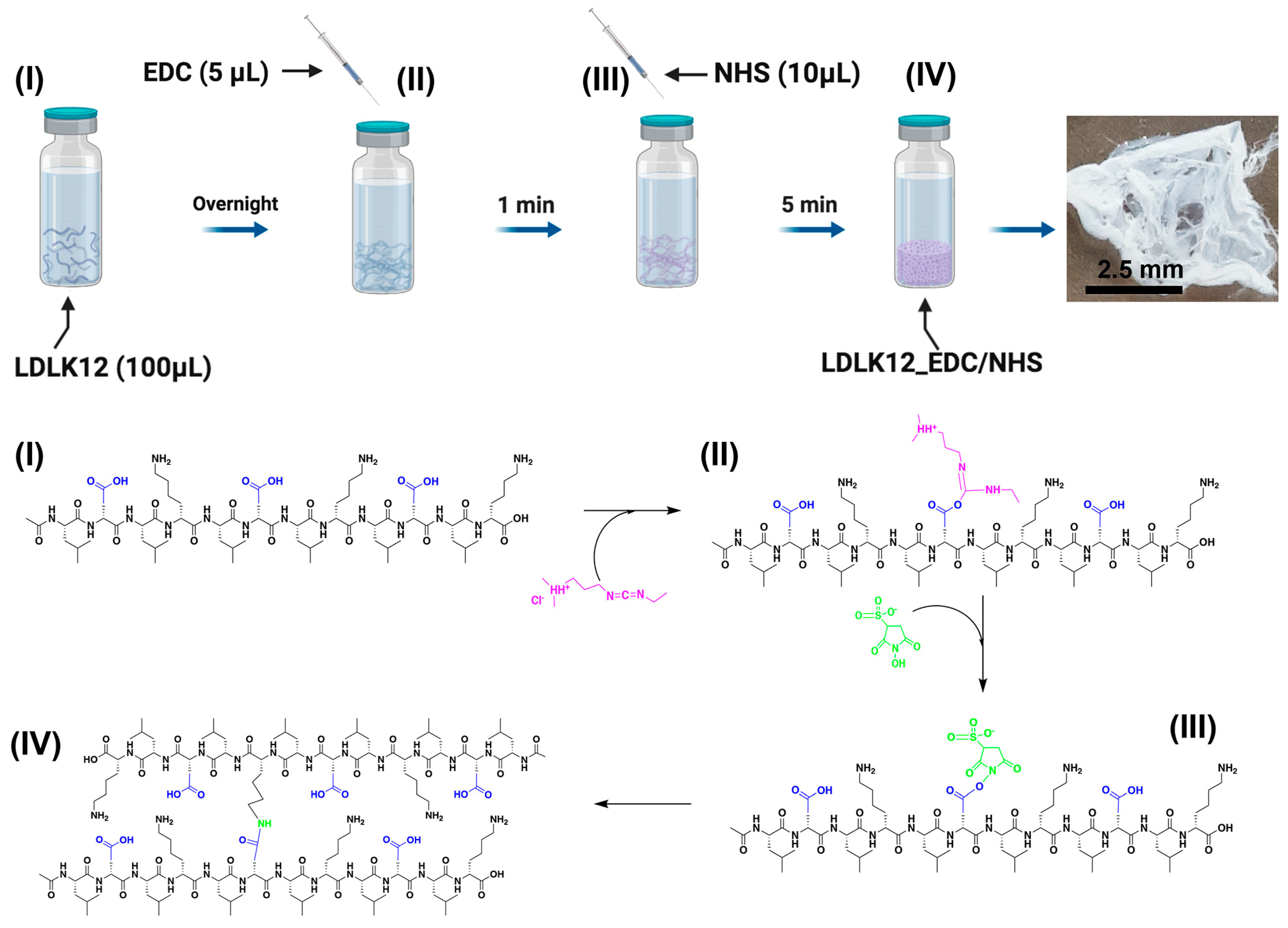
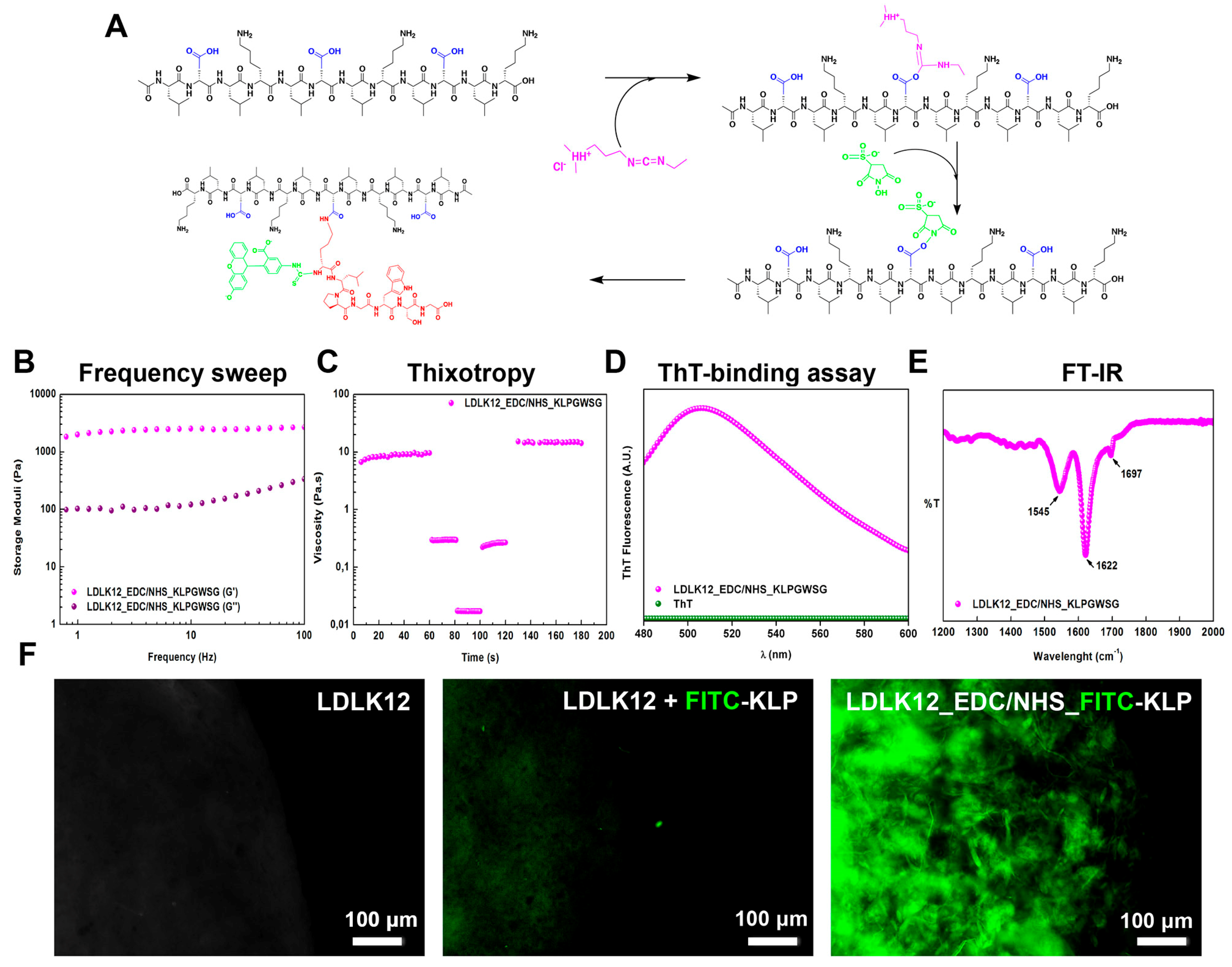
2.1. EDC/Sulfo–NHS Cross-Linked Peptide Preparation
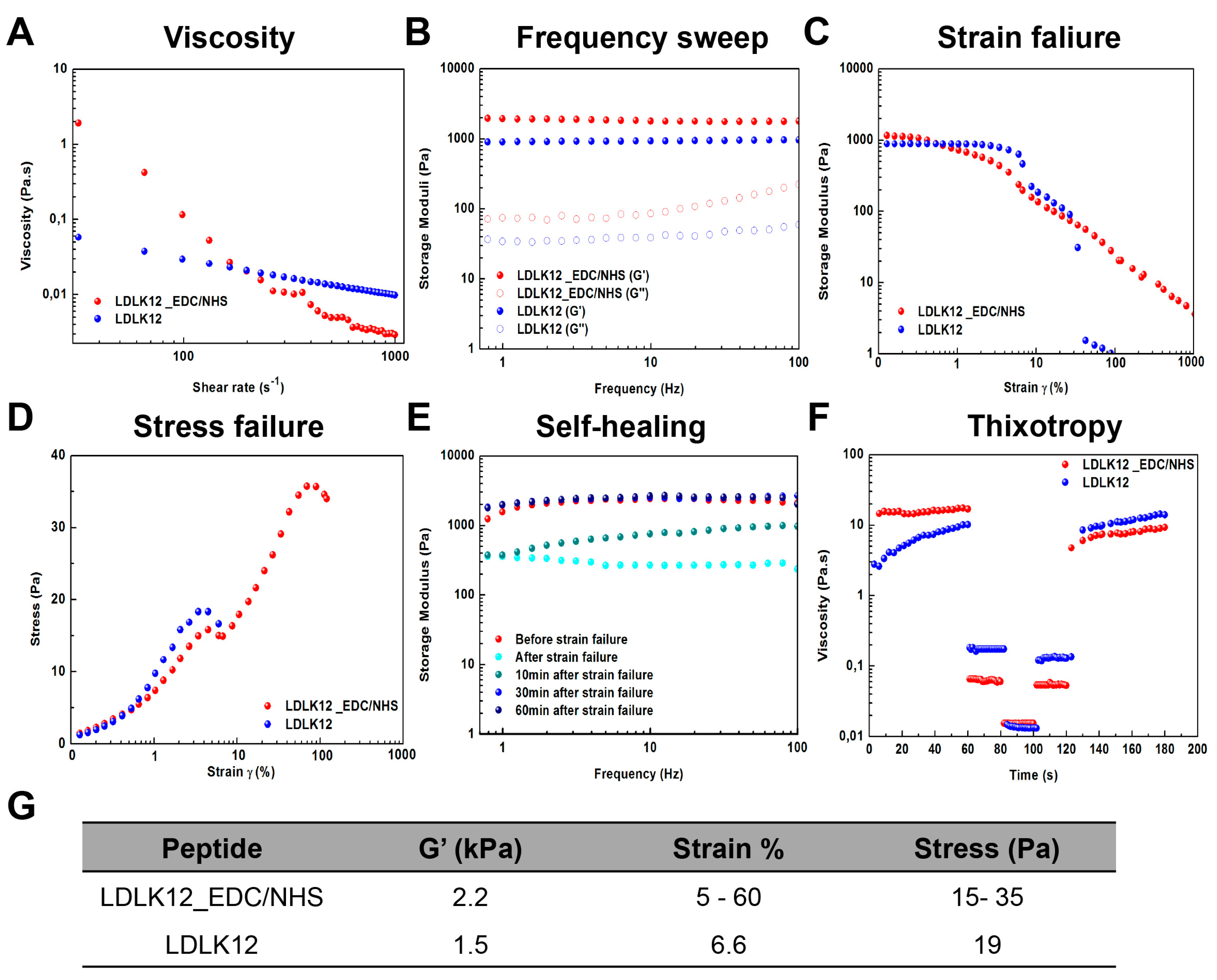
2.2. Mechanical Properties of Peptide Nanostructures
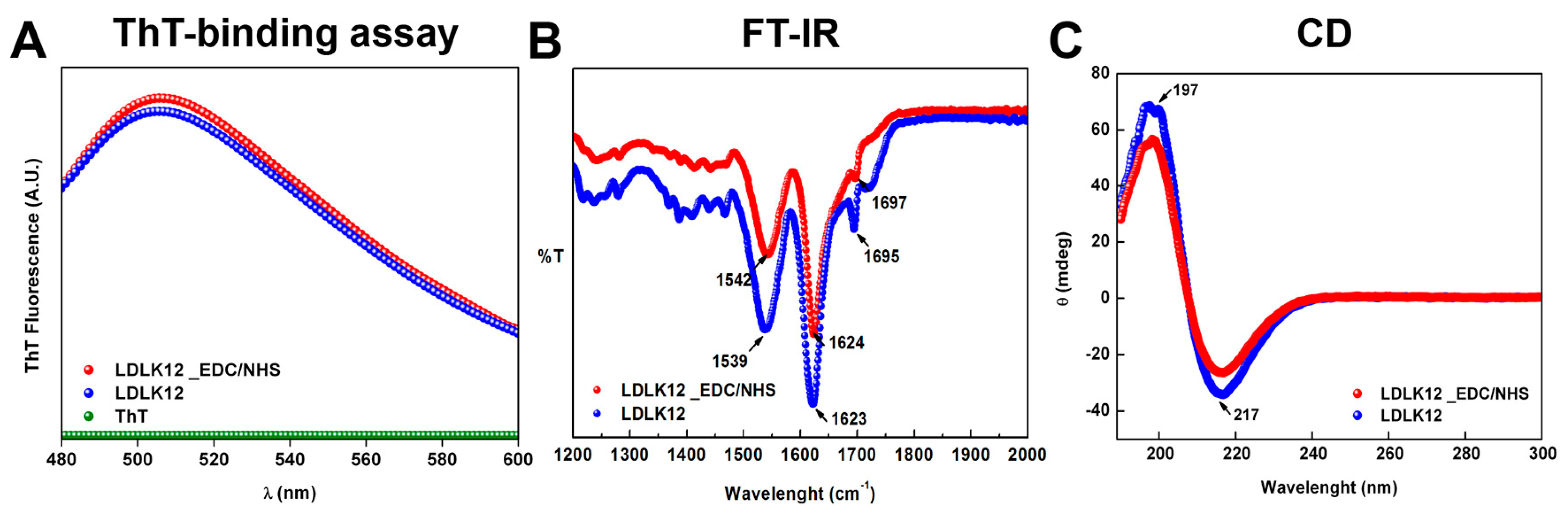
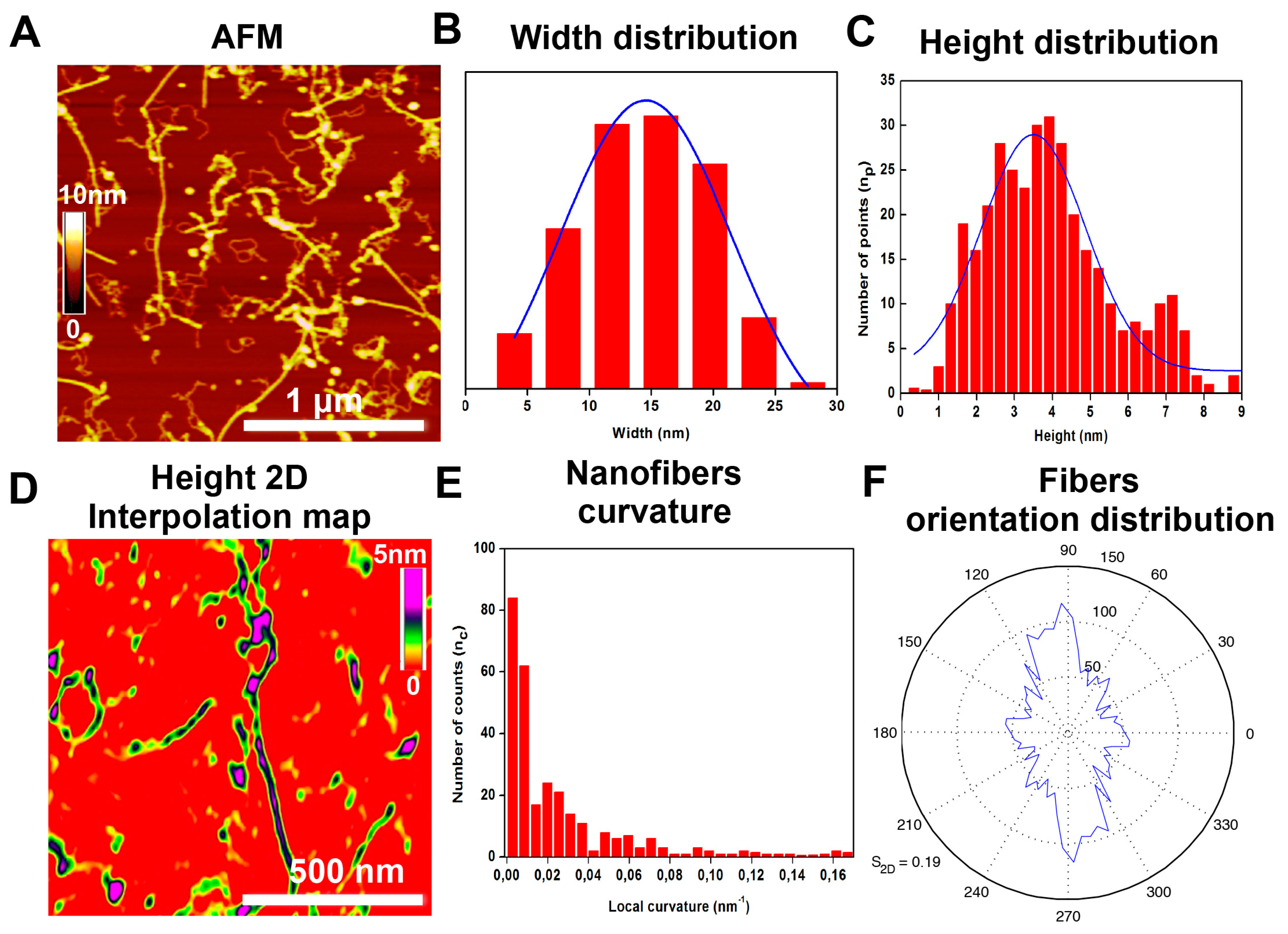
2.3. Supramolecular Organizations of Peptide Nanostructures
2.4. Post-Assembly Functionalization of KLPGWSG Peptide to the Surface of SAP Nanofibers
3. Materials and Methods
3.1. Materials
3.2. Peptide Synthesis
3.3. Fluorescein Isothiocyanate (FITC) Labeling
3.4. Cross-Linked Peptide Preparation
3.5. Mechanical Testing
3.6. Thioflavin T (ThT) Spectroscopy Assay
3.7. Fourier Transform Infrared Spectroscopy (FT-IR)
3.8. Circular Dichroism spectroscopy (CD)
3.9. Atomic Force Microscopy (AFM)
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| SAPs | Self-assembling peptides |
| ECM | Extracellular matrix |
| EDC | 1-ethyl-3-[3-dimethylaminopropyl]carbodiimide |
| Sulfo–NHS | N-hydroxysulfosuccinimide |
| TBI | Traumatic brain injury |
| SCI | Spinal cord injury |
| LVR | Linear viscoelastic regime |
| DMF | Dimethylformamide |
| HBTU | N,N,N′,N′-Tetramethyl-O-(1H-benzotriazol-1-yl)uronium hexafluorophosphate |
| DIEA | N,N-Diisopropylethylamine |
| NMP | N-Methyl-2-pyrrolidone |
| TFA | Trifluoroacetic acid |
| TIS | Triisopropylsilane |
| ThT | Thioflavin-T |
| FT-IR | Fourier transform infrared spectroscopy |
| CD | Circular dichroism |
| AFM | Atomic force microscopy |
| DPBS | Dulbecco’s phosphate-buffered saline |
| FITC | Fluorescein Isothiocyanate |
| DMSO | Dimethyl sulfoxide |
References
- Chaudhuri, O.; Gu, L.; Klumpers, D.; Darnell, M.; Bencherif, S.A.; Weaver, J.C.; Huebsch, N.; Lee, H.P.; Lippens, E.; Duda, G.N.; et al. Hydrogels with tunable stress relaxation regulate stem cell fate and activity. Nat. Mater. 2016, 15, 326–334. [Google Scholar] [CrossRef] [PubMed]
- Yang, C.; Tibbitt, M.W.; Basta, L.; Anseth, K.S. Mechanical memory and dosing influence stem cell fate. Nat. Mater. 2014, 13, 645–652. [Google Scholar] [CrossRef] [PubMed]
- Baker, E.L.; Bonnecaze, R.T.; Zaman, M.H. Extracellular matrix stiffness and architecture govern intracellular rheology in cancer. Biophys. J. 2009, 97, 1013–1021. [Google Scholar] [CrossRef] [PubMed]
- Gattazzo, F.; Urciuolo, A.; Bonaldo, P. Extracellular matrix: A dynamic microenvironment for stem cell niche. Biochim. Et Biophys. Acta 2014, 1840, 2506–2519. [Google Scholar] [CrossRef] [PubMed]
- Medina, S.H.; Bush, B.; Cam, M.; Sevcik, E.; DelRio, F.W.; Nandy, K.; Schneider, J.P. Identification of a mechanogenetic link between substrate stiffness and chemotherapeutic response in breast cancer. Biomaterials 2019, 202, 1–11. [Google Scholar] [CrossRef]
- Dupont, S. Role of YAP/TAZ in cell-matrix adhesion-mediated signalling and mechanotransduction. Exp. Cell Res. 2016, 343, 42–53. [Google Scholar] [CrossRef]
- Vining, K.H.; Mooney, D.J. Mechanical forces direct stem cell behaviour in development and regeneration. Nat. Rev. Mol. Cell Biol. 2017, 18, 728–742. [Google Scholar] [CrossRef]
- Zajiczek, L.; Shaw, M.; Faruqui, N.; Bella, A.; Pawar, V.M.; Srinivasan, M.A.; Ryadnov, M.G. Nano-mechanical single-cell sensing of cell-matrix contacts. Nanoscale 2016, 8, 18105–18112. [Google Scholar] [CrossRef]
- Engler, A.J.; Sweeney, H.L.; Discher, D.E.; Schwarzbauer, J.E. Extracellular matrix elasticity directs stem cell differentiation. J. Musculoskelet. Neuronal Interact. 2007, 7, 335. [Google Scholar]
- Yu, Z.; Tantakitti, F.; Yu, T.; Palmer, L.C.; Schatz, G.C.; Stupp, S.I. Simultaneous covalent and noncovalent hybrid polymerizations. Science 2016, 351, 497–502. [Google Scholar] [CrossRef]
- Pugliese, R.; Marchini, A.; Saracino, G.; Gelain, F. Functionalization of self-assembling peptides for neural tissue engineering. Self-Assem. Biomater.: Mol. Des. Charact. Appl. Biol. Med. 2018, 475–493. [Google Scholar] [CrossRef]
- Pugliese, R.; Gelain, F. Peptidic Biomaterials: From Self-Assembling to Regenerative Medicine. Trends Biotechnol. 2017, 35, 145–158. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S. Discovery and design of self-assembling peptides. Interface Focus 2017, 7, 20170028. [Google Scholar] [CrossRef] [PubMed]
- Gelain, F.; Bottai, D.; Vescovi, A.; Zhang, S. Designer self-assembling peptide nanofiber scaffolds for adult mouse neural stem cell 3-dimensional cultures. PLoS ONE 2006, 1, e119. [Google Scholar] [CrossRef]
- Gelain, F.; Horii, A.; Zhang, S. Designer self-assembling peptide scaffolds for 3-d tissue cell cultures and regenerative medicine. Macromol. Biosci. 2007, 7, 544–551. [Google Scholar] [CrossRef]
- Genove, E.; Shen, C.; Zhang, S.; Semino, C.E. The effect of functionalized self-assembling peptide scaffolds on human aortic endothelial cell function. Biomaterials 2005, 26, 3341–3351. [Google Scholar] [CrossRef]
- Kisiday, J.; Jin, M.; Kurz, B.; Hung, H.; Semino, C.; Zhang, S.; Grodzinsky, A.J. Self-assembling peptide hydrogel fosters chondrocyte extracellular matrix production and cell division: Implications for cartilage tissue repair. Proc. Natl. Acad. Sci. USA 2002, 99, 9996–10001. [Google Scholar] [CrossRef]
- Sun, J.; Zheng, Q. Experimental study on self-assembly of KLD-12 peptide hydrogel and 3-D culture of MSC encapsulated within hydrogel in vitro. J. Huazhong Univ. Sci. Technol Med. Sci. 2009, 29, 512–516. [Google Scholar] [CrossRef]
- Pugliese, R.; Fontana, F.; Marchini, A.; Gelain, F. Branched peptides integrate into self-assembled nanostructures and enhance biomechanics of peptidic hydrogels. Acta Biomater. 2018, 66, 258–271. [Google Scholar] [CrossRef]
- Taraballi, F.; Natalello, A.; Campione, M.; Villa, O.; Doglia, S.M.; Paleari, A.; Gelain, F. Glycine-spacers influence functional motifs exposure and self-assembling propensity of functionalized substrates tailored for neural stem cell cultures. Front. Neuroeng. 2010, 3, 1. [Google Scholar] [CrossRef]
- Gelain, F.; Cigognini, D.; Caprini, A.; Silva, D.; Colleoni, B.; Donega, M.; Antonini, S.; Cohen, B.E.; Vescovi, A. New bioactive motifs and their use in functionalized self-assembling peptides for NSC differentiation and neural tissue engineering. Nanoscale 2012, 4, 2946–2957. [Google Scholar] [CrossRef] [PubMed]
- Caprini, A.; Silva, D.; Zanoni, I.; Cunha, C.; Volonte, C.; Vescovi, A.; Gelain, F. A novel bioactive peptide: Assessing its activity over murine neural stem cells and its potential for neural tissue engineering. New Biotechnol. 2013, 30, 552–562. [Google Scholar] [CrossRef] [PubMed]
- Raspa, A.; Saracino, G.A.A.; Pugliese, R.; Silva, D.; Cigognini, D.; Vescovi, A.; Gelain, F. Complementary Co-assembling Peptides: From In Silico Studies to In Vivo Application. Adv. Funct. Mater. 2014, 24, 6317–6328. [Google Scholar] [CrossRef]
- Ji, W.; Alvarez, Z.; Edelbrock, A.N.; Sato, K.; Stupp, S.I. Bioactive Nanofibers Induce Neural Transdifferentiation of Human Bone Marrow Mesenchymal Stem Cells. ACS Appl. Mater. Interfaces 2018, 10, 41046–41055. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Li, W.; Wu, X.; Zhang, N.; Zhang, Y.; Ouyang, S.; Song, X.; Fang, X.; Seeram, R.; Xue, W.; et al. Functional Self-Assembling Peptide Nanofiber Hydrogels Designed for Nerve Degeneration. ACS Appl. Mater. Interfaces 2016, 8, 2348–2359. [Google Scholar] [CrossRef] [PubMed]
- Lee, K.M.; Kim, J.H.; Choi, E.S.; Kim, E.; Choi, S.K.; Jeon, W.B. RGD-containing elastin-like polypeptide improves islet transplantation outcomes in diabetic mice. Acta Biomater. 2019, 94, 351–360. [Google Scholar] [CrossRef] [PubMed]
- Tysseling, V.M.; Sahni, V.; Pashuck, E.T.; Birch, D.; Hebert, A.; Czeisler, C.; Stupp, S.I.; Kessler, J.A. Self-assembling peptide amphiphile promotes plasticity of serotonergic fibers following spinal cord injury. J. Neurosci. Res. 2010, 88, 3161–3170. [Google Scholar] [CrossRef]
- Castelletto, V.; Gouveia, R.M.; Connon, C.J.; Hamley, I.W. New RGD-peptide amphiphile mixtures containing a negatively charged diluent. Faraday Discuss. 2013, 166, 381–397. [Google Scholar] [CrossRef]
- Goldberger, J.E.; Berns, E.J.; Bitton, R.; Newcomb, C.J.; Stupp, S.I. Electrostatic control of bioactivity. Angew. Chem. 2011, 50, 6292–6295. [Google Scholar] [CrossRef]
- Taraballi, F.; Campione, M.; Sassella, A.; Vescovi, A.; Paleari, A.; Hwang, W.; Gelain, F. Effect of functionalization on the self-assembling propensity of β-sheet forming peptides. Soft Matter 2009, 5, 660–668. [Google Scholar] [CrossRef]
- Marchini, A.; Raspa, A.; Pugliese, R.; El Malek, M.A.; Pastori, V.; Lecchi, M.; Vescovi, A.L.; Gelain, F. Multifunctionalized hydrogels foster hNSC maturation in 3D cultures and neural regeneration in spinal cord injuries. Proc. Natl. Acad. Sci. USA 2019, 116, 7483–7492. [Google Scholar] [CrossRef] [PubMed]
- Horii, A.; Wang, X.; Gelain, F.; Zhang, S. Biological designer self-assembling peptide nanofiber scaffolds significantly enhance osteoblast proliferation, differentiation and 3-D migration. PLoS ONE 2007, 2, e190. [Google Scholar] [CrossRef] [PubMed]
- Pugliese, R.; Marchini, A.; Saracino, G.A.A.; Zuckermann, R.N.; Gelain, F. Cross-linked self-assembling peptide scaffolds. Nano Res. 2018, 11, 586–602. [Google Scholar] [CrossRef]
- Pugliese, R.; Maleki, M.; Zuckermann, R.N.; Gelain, F. Self-assembling peptides cross-linked with genipin: Resilient hydrogels and self-standing electrospun scaffolds for tissue engineering applications. Biomater. Sci. 2018, 7, 76–91. [Google Scholar] [CrossRef] [PubMed]
- Greenfield, M.A.; Hoffman, J.R.; de la Cruz, M.O.; Stupp, S.I. Tunable mechanics of peptide nanofiber gels. Langmuir: ACS J. Surf. Colloids 2010, 26, 3641–3647. [Google Scholar] [CrossRef]
- Li, I.C.; Hartgerink, J.D. Covalent Capture of Aligned Self-Assembling Nanofibers. J. Am. Chem. Soc. 2017, 139, 8044–8050. [Google Scholar] [CrossRef]
- Li, I.C.; Hulgan, S.A.H.; Walker, D.R.; Farndale, R.W.; Hartgerink, J.D.; Jalan, A.A. Covalent Capture of a Heterotrimeric Collagen Helix. Org. Lett. 2019, 21, 5480–5484. [Google Scholar] [CrossRef]
- Fischer, M.J. Amine coupling through EDC/NHS: A practical approach. Methods Mol. Biol. 2010, 627, 55–73. [Google Scholar]
- Grabarek, Z.; Gergely, J. Zero-length crosslinking procedure with the use of active esters. Anal. Biochem. 1990, 185, 131–135. [Google Scholar] [CrossRef]
- Tripathi, J.K.; Pal, S.; Awasthi, B.; Kumar, A.; Tandon, A.; Mitra, K.; Chattopadhyay, N.; Ghosh, J.K. Variants of self-assembling peptide, KLD-12 that show both rapid fracture healing and antimicrobial properties. Biomaterials 2015, 56, 92–103. [Google Scholar] [CrossRef]
- Cigognini, D.; Silva, D.; Paloppi, S.; Gelain, F. Evaluation of mechanical properties and therapeutic effect of injectable self-assembling hydrogels for spinal cord injury. J Biomed. Nanotechnol. 2014, 10, 309–323. [Google Scholar] [CrossRef] [PubMed]
- Ma, X.; Agas, A.; Siddiqui, Z.; Kim, K.; Iglesias-Montoro, P.; Kalluru, J.; Kumar, V.; Haorah, J. Angiogenic peptide hydrogels for treatment of traumatic brain injury. Bioact. Mater. 2020, 5, 124–132. [Google Scholar] [CrossRef] [PubMed]
- Sahab Negah, S.; Oliazadeh, P.; Jahanbazi Jahan-Abad, A.; Eshaghabadi, A.; Samini, F.; Ghasemi, S.; Asghari, A.; Gorji, A. Transplantation of human meningioma stem cells loaded on a self-assembling peptide nanoscaffold containing IKVAV improves traumatic brain injury in rats. Acta Biomater. 2019, 92, 132–144. [Google Scholar] [CrossRef] [PubMed]
- Cigognini, D.; Satta, A.; Colleoni, B.; Silva, D.; Donega, M.; Antonini, S.; Gelain, F. Evaluation of early and late effects into the acute spinal cord injury of an injectable functionalized self-assembling scaffold. PLoS ONE 2011, 6, e19782. [Google Scholar] [CrossRef] [PubMed]
- Raspa, A.; Marchini, A.; Pugliese, R.; Mauri, M.; Maleki, M.; Vasita, R.; Gelain, F. A biocompatibility study of new nanofibrous scaffolds for nervous system regeneration. Nanoscale 2016, 8, 253–265. [Google Scholar] [CrossRef] [PubMed]
- Raspa, A.; Pugliese, R.; Maleki, M.; Gelain, F. Recent therapeutic approaches for spinal cord injury. Biotechnol. Bioeng. 2016, 113, 253–259. [Google Scholar] [CrossRef]
- Hsu, B.B.; Conway, W.; Tschabrunn, C.M.; Mehta, M.; Perez-Cuevas, M.B.; Zhang, S.; Hammond, P.T. Clotting Mimicry from Robust Hemostatic Bandages Based on Self-Assembling Peptides. ACS Nano 2015, 9, 9394–9406. [Google Scholar] [CrossRef]
- Madl, C.M.; LeSavage, B.L.; Dewi, R.E.; Dinh, C.B.; Stowers, R.S.; Khariton, M.; Lampe, K.J.; Nguyen, D.; Chaudhuri, O.; Enejder, A.; et al. Maintenance of neural progenitor cell stemness in 3D hydrogels requires matrix remodelling. Nat. Mater. 2017, 16, 1233–1242. [Google Scholar] [CrossRef]
- Huebsch, N.; Lippens, E.; Lee, K.; Mehta, M.; Koshy, S.T.; Darnell, M.C.; Desai, R.M.; Madl, C.M.; Xu, M.; Zhao, X.; et al. Matrix elasticity of void-forming hydrogels controls transplanted-stem-cell-mediated bone formation. Nat. Mater. 2015, 14, 1269–1277. [Google Scholar] [CrossRef]
- Zhao, X.; Huebsch, N.; Mooney, D.J.; Suo, Z. Stress-relaxation behavior in gels with ionic and covalent crosslinks. J. Appl. Phys. 2010, 107, 63509. [Google Scholar] [CrossRef]
- Rosales, A.M.; Anseth, K.S. The design of reversible hydrogels to capture extracellular matrix dynamics. Nat. Rev. Mater. 2016, 1, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Madl, C.M.; LeSavage, B.L.; Dewi, R.E.; Lampe, K.J.; Heilshorn, S.C. Matrix Remodeling Enhances the Differentiation Capacity of Neural Progenitor Cells in 3D Hydrogels. Adv. Sci. 2019, 6, 1801716. [Google Scholar] [CrossRef] [PubMed]
- Pugliese, R.; Gelain, F. Characterization of elastic, thermo-responsive, self-healable supramolecular hydrogel made of self-assembly peptides and guar gum. Mater. Des. 2020, 186, 108370. [Google Scholar] [CrossRef]
- Guvendiren, M.; Lu, H.D.; Burdick, J.A. Shear-thinning hydrogels for biomedical applications. Soft Matter 2012, 8, 260–272. [Google Scholar] [CrossRef]
- Chen, M.H.; Wang, L.L.; Chung, J.J.; Kim, Y.H.; Atluri, P.; Burdick, J.A. Methods To Assess Shear-Thinning Hydrogels for Application As Injectable Biomaterials. ACS Biomater. Sci. Eng. 2017, 3, 3146–3160. [Google Scholar] [CrossRef]
- Mealy, J.E.; Chung, J.J.; Jeong, H.H.; Issadore, D.; Lee, D.; Atluri, P.; Burdick, J.A. Injectable Granular Hydrogels with Multifunctional Properties for Biomedical Applications. Adv. Mater. 2018, 30, e1705912. [Google Scholar] [CrossRef]
- Bera, S.; Mondal, S.; Xue, B.; Shimon, L.J.W.; Cao, Y.; Gazit, E. Rigid helical-like assemblies from a self-aggregating tripeptide. Nat. Mater. 2019, 18, 503–509. [Google Scholar] [CrossRef]
- Liu, Y.; Yang, Y.; Wang, C.; Zhao, X. Stimuli-responsive self-assembling peptides made from antibacterial peptides. Nanoscale 2013, 5, 6413–6421. [Google Scholar] [CrossRef]
- Biancalana, M.; Makabe, K.; Koide, A.; Koide, S. Molecular mechanism of thioflavin-T binding to the surface of beta-rich peptide self-assemblies. J. Mol. Biol. 2009, 385, 1052–1063. [Google Scholar] [CrossRef]
- Jordens, S.; Schwenke, K.; Usov, I.; Del Gado, E.; Mezzenga, R. Nematic field transfer in a two-dimensional protein fibril assembly. Soft Matter 2016, 12, 1830–1835. [Google Scholar] [CrossRef]
- Usov, I.; Mezzenga, R. FiberApp: An Open-Source Software for Tracking and Analyzing Polymers, Filaments, Biomacromolecules, and Fibrous Objects. Macromolecules 2015, 48, 1269–1280. [Google Scholar] [CrossRef]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pugliese, R.; Gelain, F. Cross-Linked Self-Assembling Peptides and Their Post-Assembly Functionalization via One-Pot and In Situ Gelation System. Int. J. Mol. Sci. 2020, 21, 4261. https://doi.org/10.3390/ijms21124261
Pugliese R, Gelain F. Cross-Linked Self-Assembling Peptides and Their Post-Assembly Functionalization via One-Pot and In Situ Gelation System. International Journal of Molecular Sciences. 2020; 21(12):4261. https://doi.org/10.3390/ijms21124261
Chicago/Turabian StylePugliese, Raffaele, and Fabrizio Gelain. 2020. "Cross-Linked Self-Assembling Peptides and Their Post-Assembly Functionalization via One-Pot and In Situ Gelation System" International Journal of Molecular Sciences 21, no. 12: 4261. https://doi.org/10.3390/ijms21124261
APA StylePugliese, R., & Gelain, F. (2020). Cross-Linked Self-Assembling Peptides and Their Post-Assembly Functionalization via One-Pot and In Situ Gelation System. International Journal of Molecular Sciences, 21(12), 4261. https://doi.org/10.3390/ijms21124261




