Interaction Analysis of Commercial Graphene Oxide Nanoparticles with Unicellular Systems and Biomolecules
Abstract
1. Introduction
2. Results and Discussion
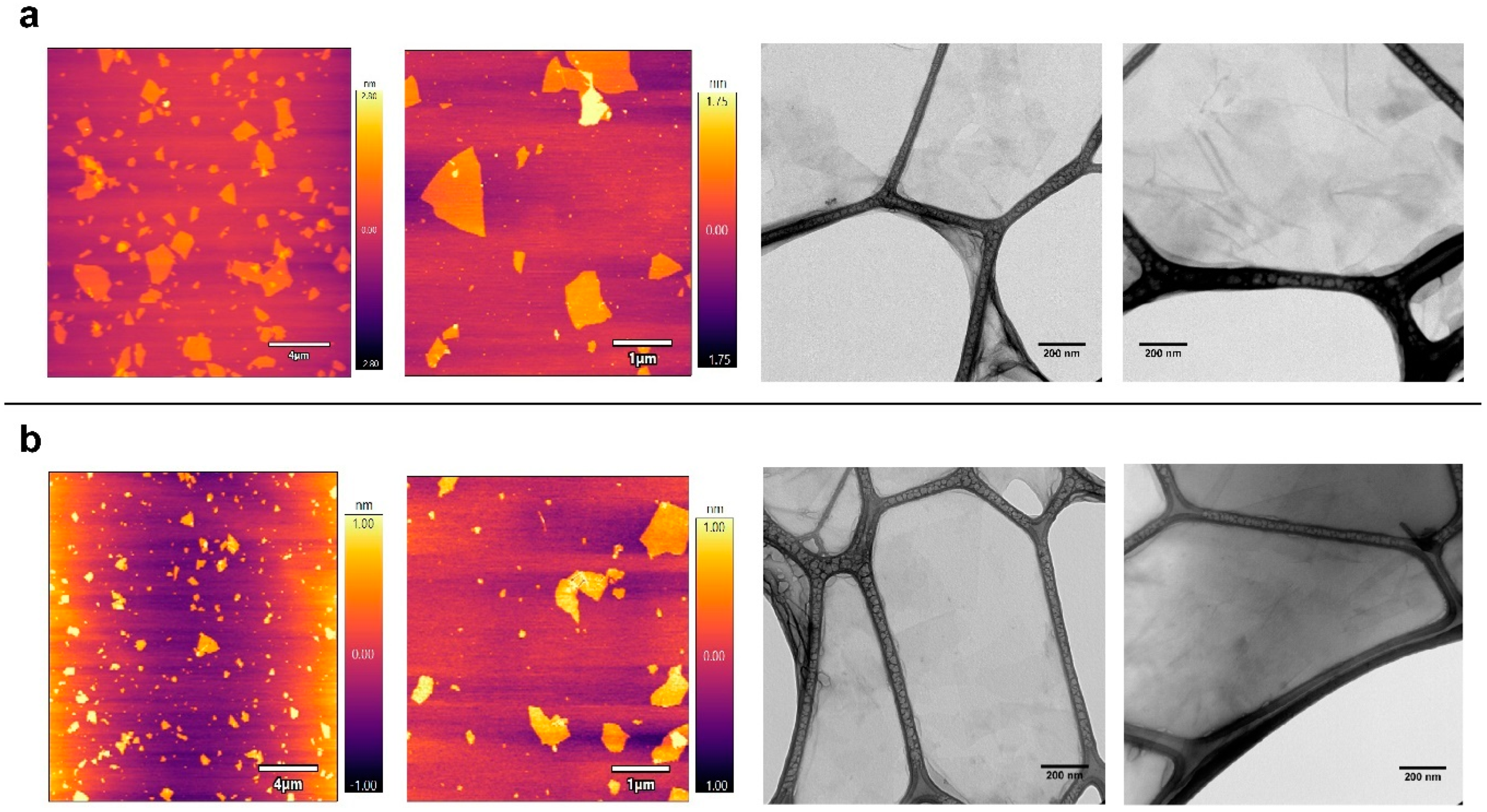
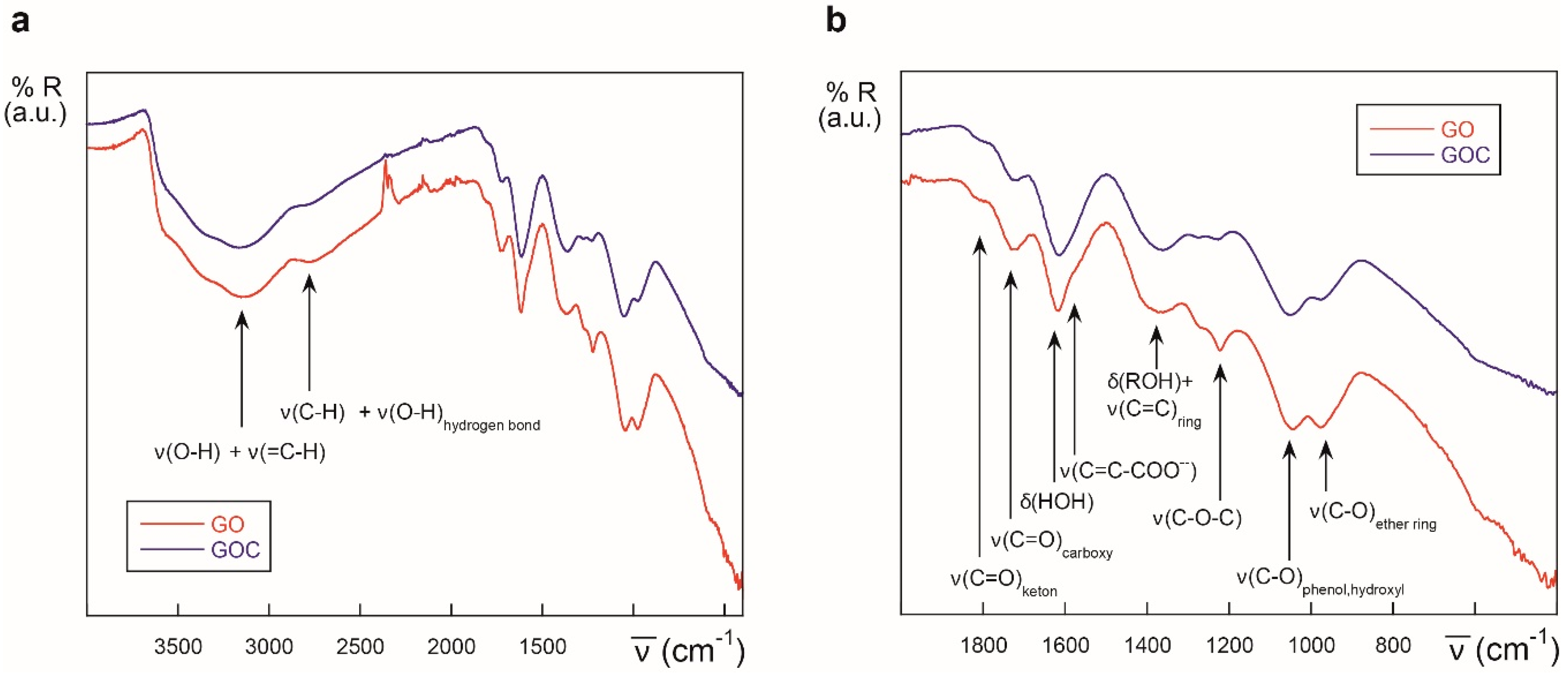
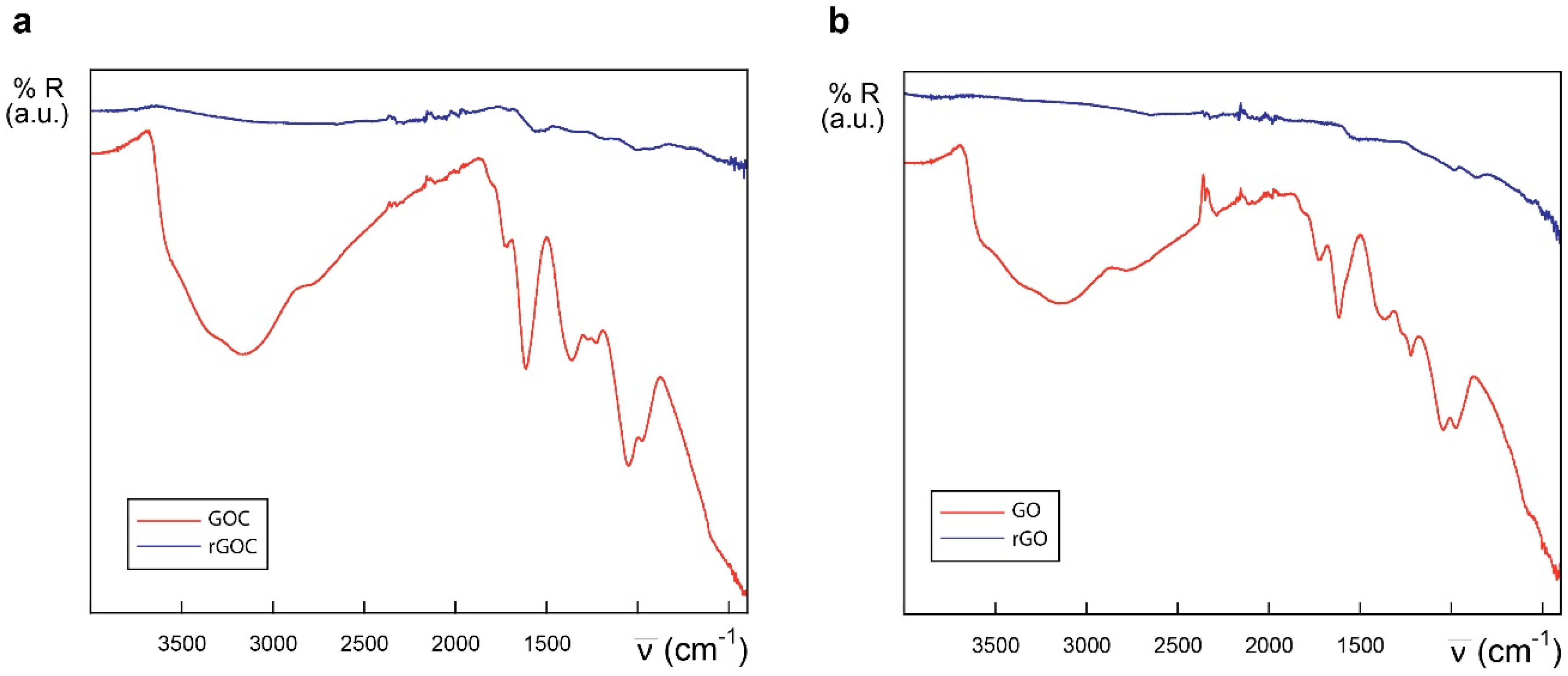
2.1. Characteristics of the Selected Commercial Graphene Oxide Derivatives
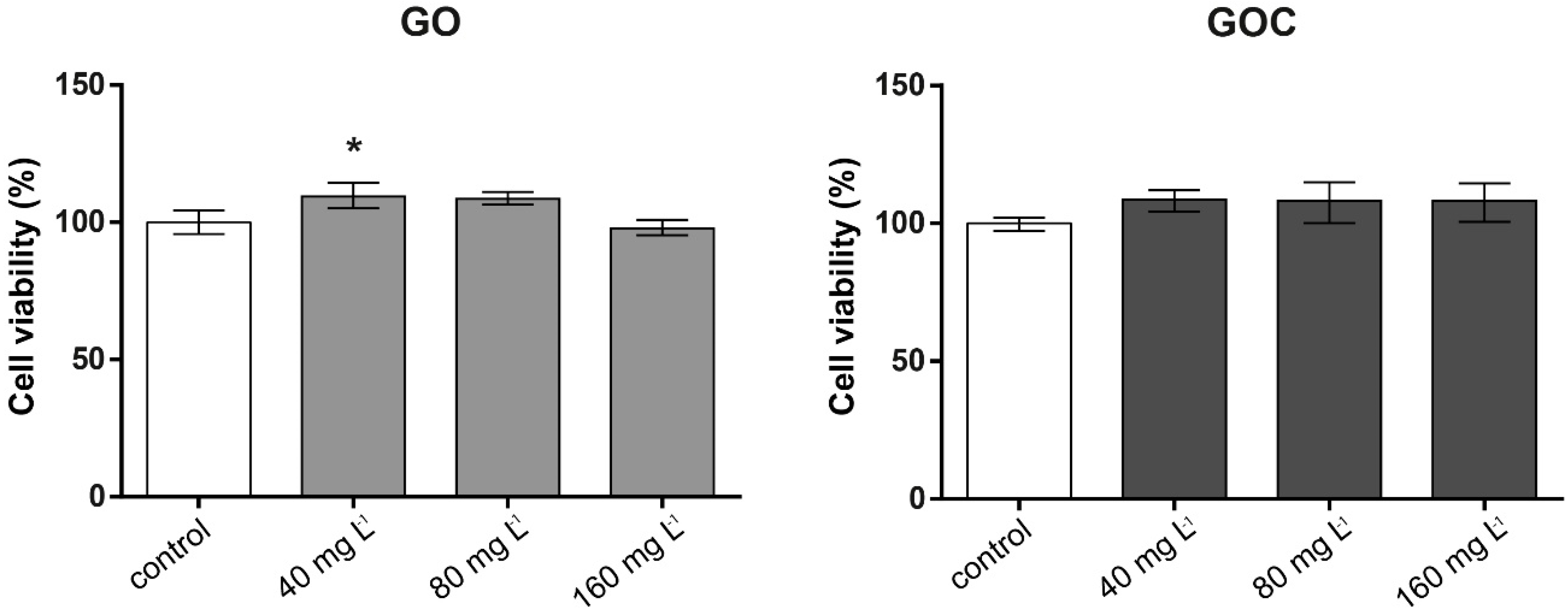
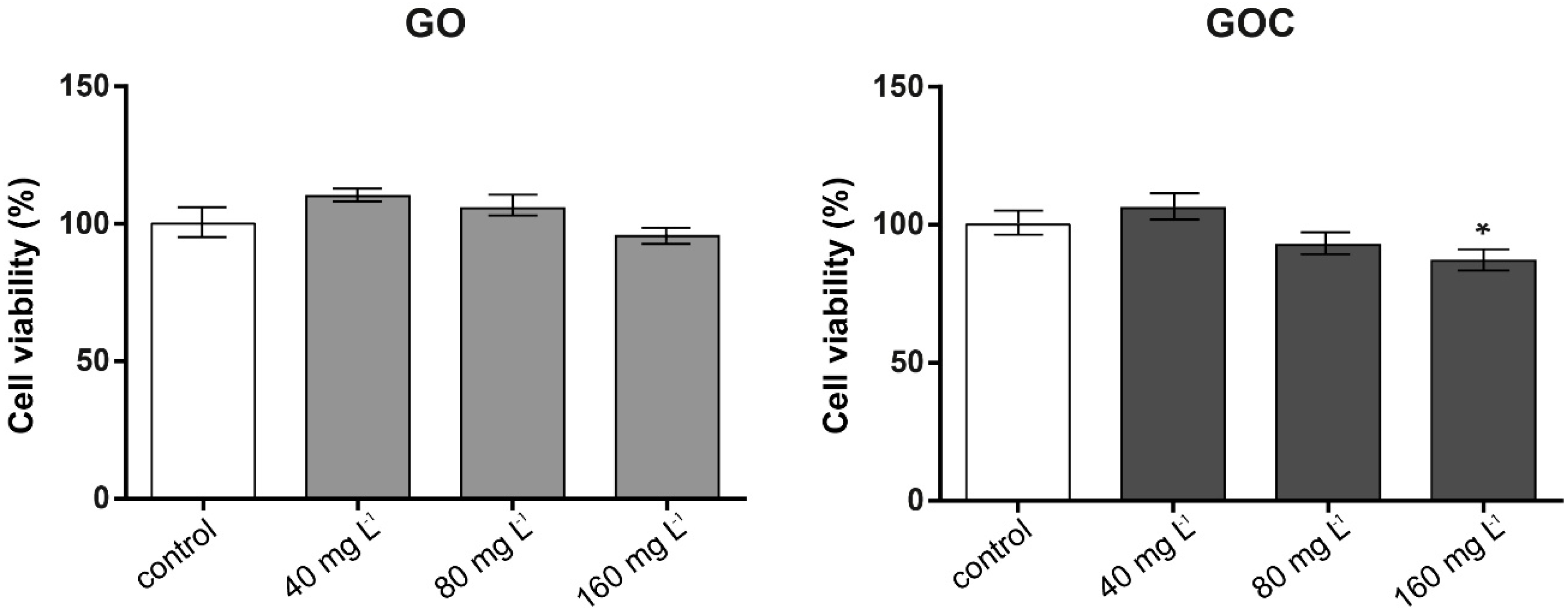
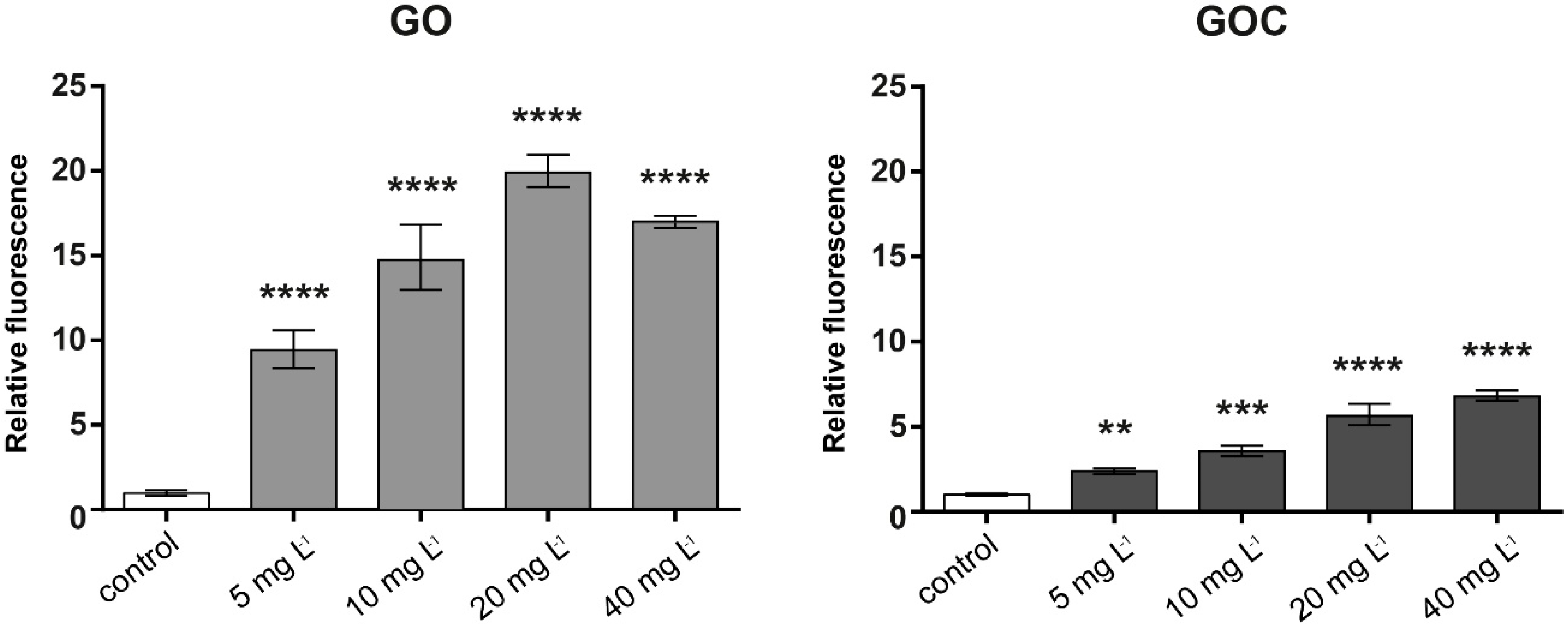
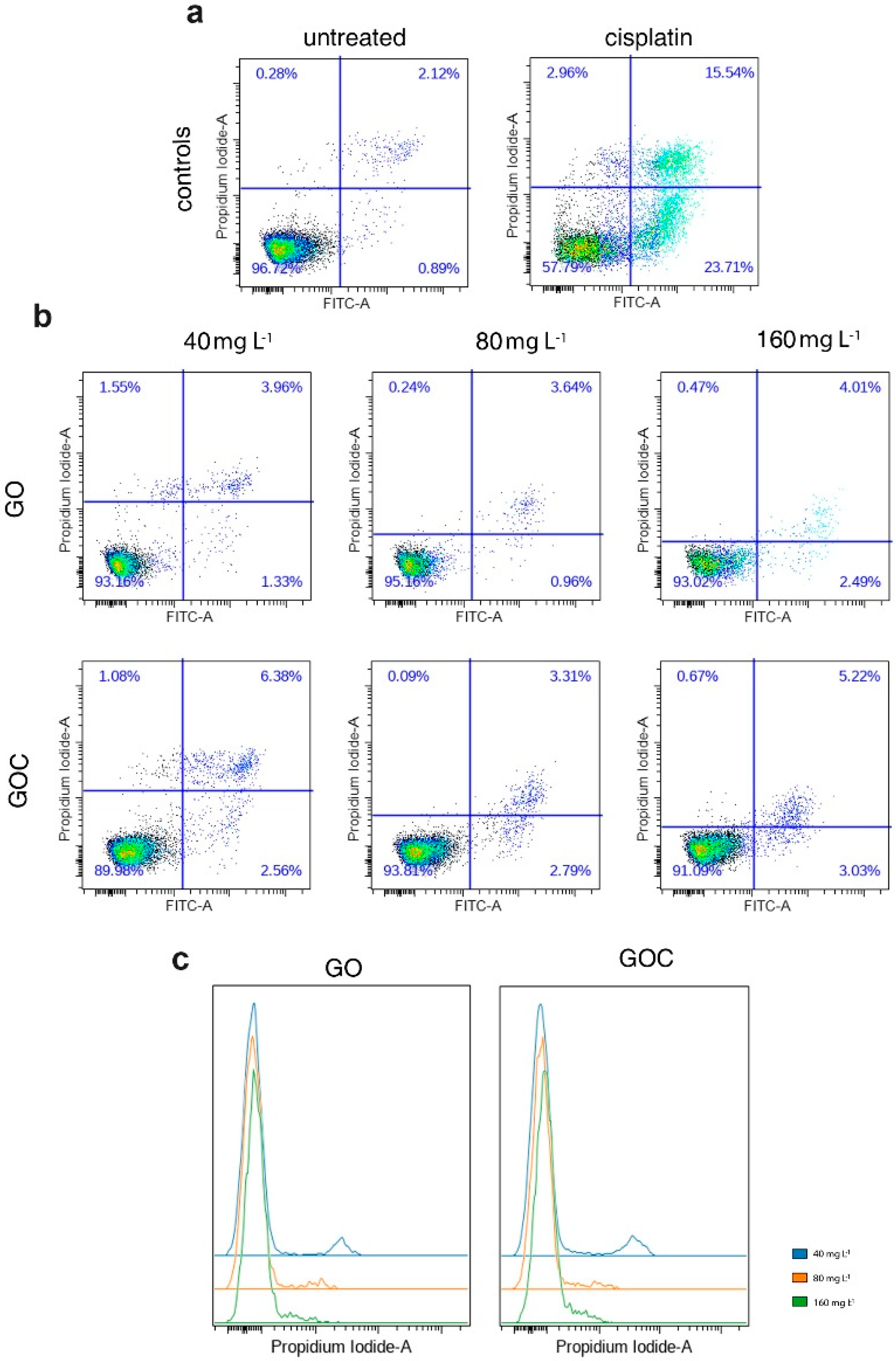
2.2. Determination of Human Cancer Cell Line A549 Response to GO and GOC
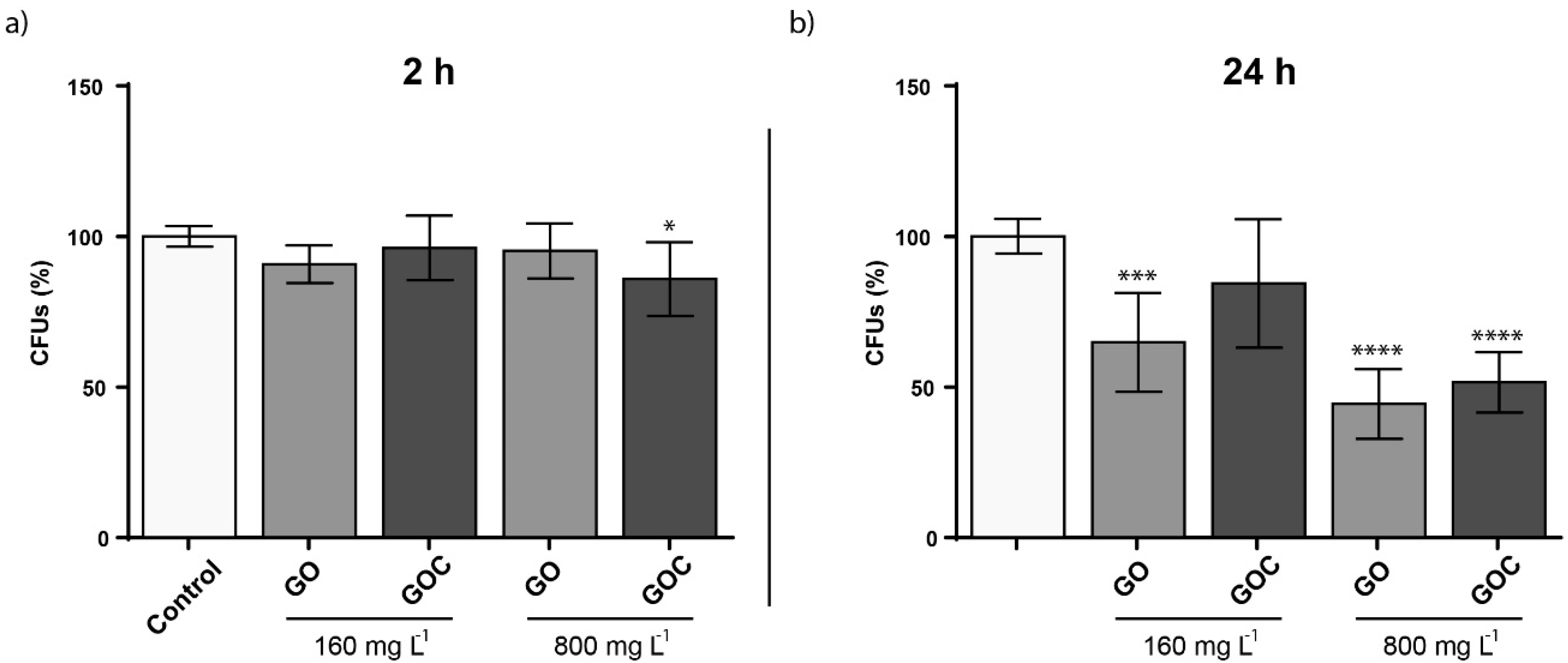
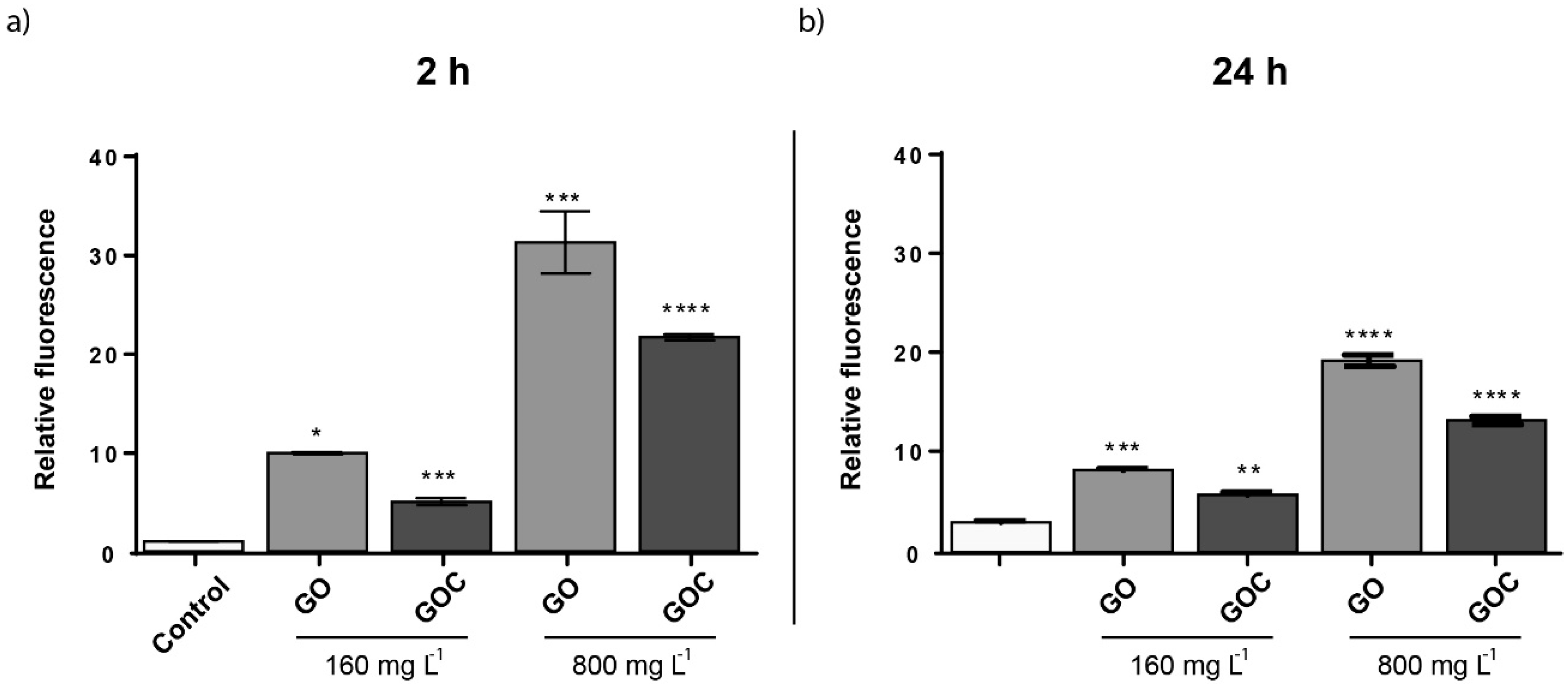
2.3. Determination of Saccharomyces Cerevisiae Cells Response to GO and GOC
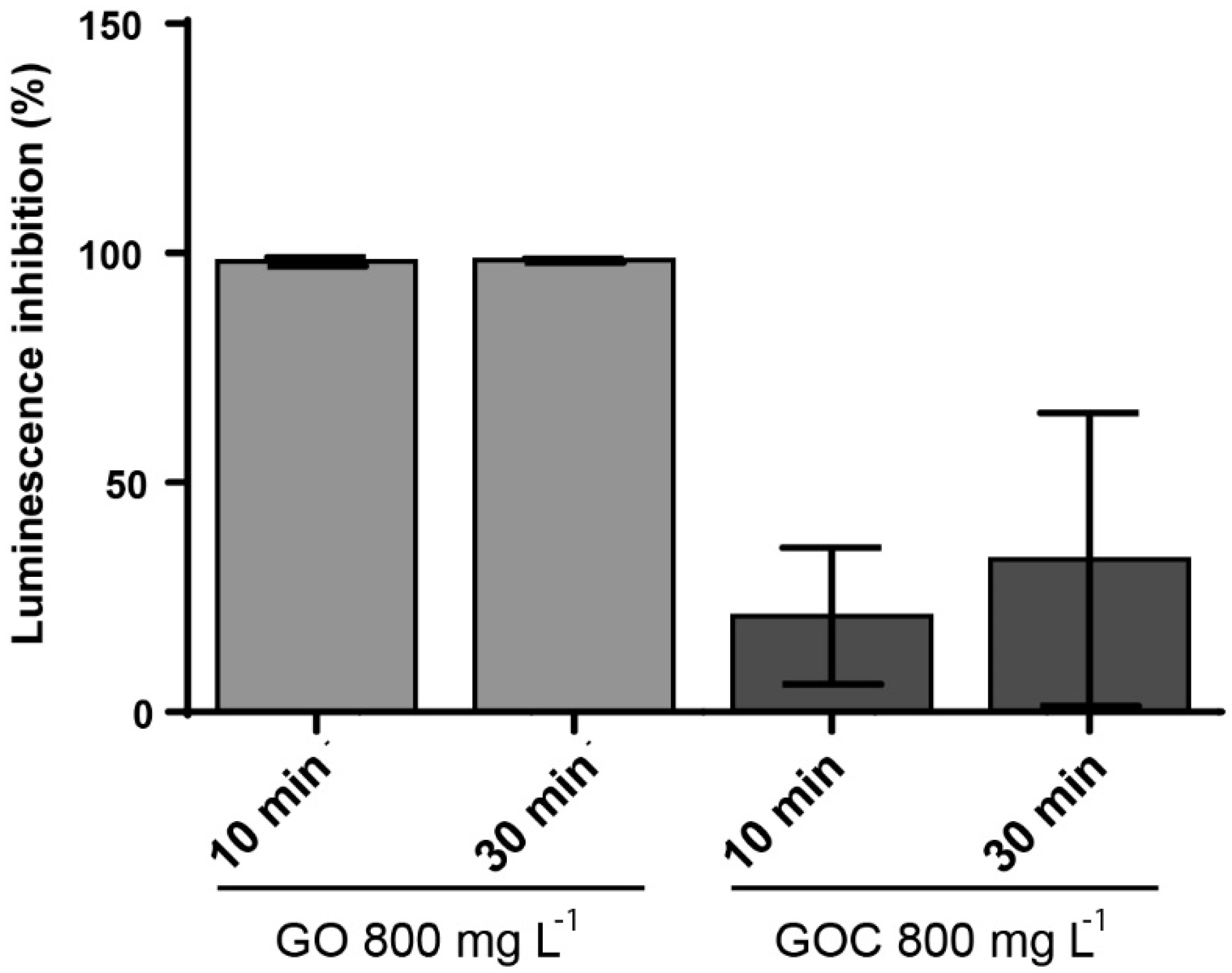
2.4. Determination of Vibrio Fischeri Bioluminescence Inhibition to GO and GOC
2.5. Determination of GO and GOC Binding Efficiency on Different Microbial Enzymes
3. Materials and Methods
3.1. Materials and Reagents
3.2. ATR-FTIR Analysis
3.3. ICP-MS
3.4. XPS Analysis
3.5. AFM and TEM Analysis
3.6. Assays in A549 Cells
3.6.1. Neutral Red Assay
3.6.2. MTT Assay
3.6.3. ROS Determination in Human Cells
3.6.4. Apoptosis Assay
3.7. Assays in Saccharomyces Cerevisiae
3.7.1. Colony Forming Units Determination
3.7.2. ROS Determination in S. cerevisiae
3.8. Vibrio Fischeri Luminescence Inhibition Assay
3.9. Preparation of rGO and rGOC
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| GO | Monolayer graphene oxide |
| GOC | Graphene oxide nanocolloids |
| ROS | Reactive oxygen species |
| AFM | Atomic force microscopy |
| TEM | Transmission electron microscopy |
| FTIR | Fourier-transform infrared spectroscopy |
| ICP-MS | Inductively coupled plasma mass spectrometry |
| ppm | parts-per-million |
| ATP | Adenosine triphosphate |
| SD | Standard deviation |
| PI | Propidium iodide |
| MWCNs | Multiwalled carbon nanotubes |
| O-SWCNTs | Oxidized single-walled carbon nanotubes |
| rGO | Reduced monolayer graphene oxide |
| rGOC | Reduced graphene oxide nanocolloids |
| DMSO | Dimethyl sulfoxide |
References
- Bayat, Z.; Hassanshahian, M.; Cappello, S. Immobilization of microbes for bioremediation of crude oil polluted environments: A mini review. Open Microbiol. J. 2015, 9, 48–54. [Google Scholar] [PubMed]
- Gurung, N.; Ray, S.; Bose, S.; Rai, V. A broader view: Microbial enzymes and their relevance in industries, medicine, and beyond. Biomed. Res. Int. 2013, 2013, 329121. [Google Scholar] [CrossRef] [PubMed]
- Robatjazi, S.M.; Shojaosadati, S.A.; Khalilzadeh, R.; Farahani, E.V.; Balochi, N. Immobilization of magnetic modified Flavobacterium ATCC 27551 using magnetic field and evaluation of the enzyme stability of immobilized bacteria. Bioresour. Technol. 2012, 104, 6–11. [Google Scholar] [CrossRef] [PubMed]
- Antón-Millán, N.; García-Tojal, J.; Marty-Roda, M.; Garroni, S.; Cuesta-López, S.; Tamayo-Ramos, J.A. Influence of three commercial graphene derivatives on the catalytic properties of a Lactobacillus plantarum α-l-rhamnosidase when used as immobilization matrices. ACS Appl. Mater. Interfaces 2018, 10, 18170–18182. [Google Scholar] [CrossRef]
- Tamayo-Ramos, J.A.; Rumbo, C.; Caso, F.; Rinaldi, A.; Garroni, S.; Notargiacomo, A.; Romero-Santacreu, L.; Cuesta-López, S. Analysis of polycaprolactone microfibers as biofilm carriers for biotechnologically relevant bacteria. ACS Appl. Mater. Interfaces 2018, 10, 32773–32781. [Google Scholar] [CrossRef]
- Li, Y.-G.; Gao, H.-S.; Li, W.-L.; Xing, J.-M.; Liu, H.-Z. In situ magnetic separation and immobilization of dibenzothiophene-desulfurizing bacteria. Bioresour. Technol. 2009, 100, 5092–5096. [Google Scholar] [CrossRef]
- Kurečič, M.; Rijavec, T.; Hribernik, S.; Lapanje, A.; Kleinschek, K.S.; Maver, U. Novel electrospun fibers with incorporated commensal bacteria for potential preventive treatment of the diabetic foot. Nanomedicine 2018, 13, 1583–1594. [Google Scholar] [CrossRef]
- Zhang, J.; Zhao, W.; Zhang, H.; Wang, Z.; Fan, C.; Zang, L. Recent achievements in enhancing anaerobic digestion with carbon-based functional materials. Bioresour. Technol. 2018, 266, 555–567. [Google Scholar] [CrossRef]
- Jesionowski, T.; Zdarta, J.; Krajewska, B. Enzyme immobilization by adsorption: A review. Adsorption 2014, 20, 801–821. [Google Scholar] [CrossRef]
- Putzbach, W.; Ronkainen, N.J. Immobilization techniques in the fabrication of nanomaterial-based electrochemical biosensors: A review. Sensors 2013, 13, 4811–4840. [Google Scholar] [CrossRef]
- Yang, S.H.; Lee, T.; Seo, E.; Ko, E.H.; Choi, I.S.; Kim, B.-S. Interfacing living yeast cells with graphene oxide nanosheaths. Macromol. Biosci. 2012, 12, 61–66. [Google Scholar] [CrossRef] [PubMed]
- Bilal, M.; Zhao, Y.; Rasheed, T.; Iqbal, H.M.N. Magnetic nanoparticles as versatile carriers for enzymes immobilization: A review. Int. J. Biol. Macromol. 2018, 120, 2530–2544. [Google Scholar] [CrossRef] [PubMed]
- Lin, L.-H.; Shih, J.-S. Immobilized fullerene C60-enzyme-based electrochemical glucose sensor. J. Chin. Chem. Soc. 2011, 58, 228–235. [Google Scholar] [CrossRef]
- Pagán, M.; Suazo, D.; Del Toro, N.; Griebenow, K. A comparative study of different protein immobilization methods for the construction of an efficient nano-structured lactate oxidase-SWCNT-biosensor. Biosens. Bioelectron. 2015, 64, 138–146. [Google Scholar] [CrossRef]
- Mubarak, N.M.; Wong, J.R.; Tan, K.W.; Sahu, J.N.; Abdullah, E.C.; Jayakumar, N.S.; Ganesan, P. Immobilization of cellulase enzyme on functionalized multiwall carbon nanotubes. J. Mol. Catal. B Enzym. 2014, 107, 124–131. [Google Scholar] [CrossRef]
- Fraga-García, P.; Kubbutat, P.; Brammen, M.; Schwaminger, S.; Berensmeier, S. Bare iron oxide nanoparticles for magnetic harvesting of microalgae: From interaction behavior to process realization. Nanomaterials 2018, 8, 292. [Google Scholar] [CrossRef]
- Ranmadugala, D.; Ebrahiminezhad, A.; Manley-Harris, M.; Ghasemi, Y.; Berenjian, A. Magnetic immobilization of bacteria using iron oxide nanoparticles. Biotechnol. Lett. 2018, 40, 237–248. [Google Scholar] [CrossRef]
- Banerjee, P.; Barman, S.R.; Swarnakar, S.; Mukhopadhyay, A.; Das, P. Treatment of textile effluent using bacteria-immobilized graphene oxide nanocomposites: Evaluation of effluent detoxification using Bellamya bengalensis. Clean Technol. Environ. Policy 2018, 20, 2287–2298. [Google Scholar] [CrossRef]
- Durán, N.; Martinez, D.S.T.; Silveira, C.P.; Durán, M.; De Moraes, A.C.M.; Simões, M.B.; Alves, O.L.; Fávaro, W.J. Graphene oxide: A carrier for pharmaceuticals and a scaffold for cell interactions. Curr. Top. Med. Chem. 2015, 15, 309–327. [Google Scholar] [CrossRef]
- Poulin, P.; Jalili, R.; Neri, W.; Nallet, F.; Divoux, T.; Colin, A.; Aboutalebi, S.H.; Wallace, G.; Zakri, C. Superflexibility of graphene oxide. Proc. Natl. Acad. Sci. USA 2016, 113, 11088–11093. [Google Scholar] [CrossRef]
- Liao, C.; Li, Y.; Tjong, S. Graphene nanomaterials: Synthesis, biocompatibility, and cytotoxicity. Int. J. Mol. Sci. 2018, 19, 3564. [Google Scholar] [CrossRef] [PubMed]
- Lammel, T.; Boisseaux, P.; Fernández-Cruz, M.-L.; Navas, J.M. Internalization and cytotoxicity of graphene oxide and carboxyl graphene nanoplatelets in the human hepatocellular carcinoma cell line Hep G2. Part. Fibre Toxicol. 2013, 10, 27. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.; Ruan, J.; Song, H.; Zhang, J.; Wo, Y.; Guo, S.; Cui, D. Biocompatibility of graphene oxide. Nanoscale Res. Lett. 2011, 6, 8. [Google Scholar] [CrossRef] [PubMed]
- Ge, L.; Chen, A.; Pei, J.; Zhao, L.; Fang, X.; Ding, G.; Wang, Z.; Xiao, W.; Tang, F. Enhancing the thermostability of α-l-rhamnosidase from Aspergillus terreus and the enzymatic conversion of rutin to isoquercitrin by adding sorbitol. BMC Biotechnol. 2017, 17, 21. [Google Scholar] [CrossRef]
- Li, R.; Guiney, L.M.; Chang, C.H.; Mansukhani, N.D.; Ji, Z.; Wang, X.; Liao, Y.-P.; Jiang, W.; Sun, B.; Hersam, M.C.; et al. Surface oxidation of graphene oxide determines membrane damage, lipid peroxidation, and cytotoxicity in macrophages in a pulmonary toxicity model. ACS Nano 2018, 12, 1390–1402. [Google Scholar] [CrossRef]
- Pelin, M.; Fusco, L.; Martín, C.; Sosa, S.; Frontiñán-Rubio, J.; González-Domínguez, J.M.; Durán-Prado, M.; Vázquez, E.; Prato, M.; Tubaro, A. Graphene and graphene oxide induce ROS production in human HaCaT skin keratinocytes: The role of xanthine oxidase and NADH dehydrogenase. Nanoscale 2018, 10, 11820–11830. [Google Scholar] [CrossRef]
- Malina, T.; Maršálková, E.; Holá, K.; Tuček, J.; Scheibe, M.; Zbořil, R.; Maršálek, B. Toxicity of graphene oxide against algae and cyanobacteria: Nanoblade-morphology-induced mechanical injury and self-protection mechanism. Carbon N. Y. 2019, 155, 386–396. [Google Scholar] [CrossRef]
- Zhang, M.; Yu, Q.; Liang, C.; Liu, Z.; Zhang, B.; Li, M. Graphene oxide induces plasma membrane damage, reactive oxygen species accumulation and fatty acid profiles change in Pichia pastoris. Ecotoxicol. Environ. Saf. 2016, 132, 372–378. [Google Scholar] [CrossRef]
- Arrais, A.; Diana, E.; Boccaleri, E. A study on the carbon soot derived from the wood combustion and on the relative alkali-extractable fraction. J. Mater. Sci. 2006, 41, 6035–6045. [Google Scholar] [CrossRef]
- Wong, C.H.A.; Sofer, Z.; Kubešová, M.; Kučera, J.; Matějková, S.; Pumera, M. Synthetic routes contaminate graphene materials with a whole spectrum of unanticipated metallic elements. Proc. Natl. Acad. Sci. USA 2014, 111, 13774–13779. [Google Scholar] [CrossRef]
- Eigler, S.; Dotzer, C.; Hof, F.; Bauer, W.; Hirsch, A. Sulfur species in graphene oxide. Chem. A Eur. J. 2013, 19, 9490–9496. [Google Scholar] [CrossRef] [PubMed]
- Seabra, A.B.; Paula, A.J.; De Lima, R.; Alves, O.L.; Durán, N. Nanotoxicity of graphene and graphene oxide. Chem. Res. Toxicol. 2014, 27, 159–168. [Google Scholar] [CrossRef] [PubMed]
- Gies, V.; Zou, S. Systematic toxicity investigation of graphene oxide: Evaluation of assay selection, cell type, exposure period and flake size. Toxicol. Res. (Camb.) 2018, 7, 93–101. [Google Scholar] [CrossRef] [PubMed]
- Chang, Y.; Yang, S.-T.; Liu, J.-H.; Dong, E.; Wang, Y.; Cao, A.; Liu, Y.; Wang, H. In vitro toxicity evaluation of graphene oxide on A549 cells. Toxicol. Lett. 2011, 200, 201–210. [Google Scholar] [CrossRef]
- Reshma, S.C.; Syama, S.; Mohanan, P.V. Nano-biointeractions of PEGylated and bare reduced graphene oxide on lung alveolar epithelial cells: A comparative in vitro study. Colloids Surf. B Biointerfaces 2016, 140, 104–116. [Google Scholar] [CrossRef]
- Mittal, S.; Kumar, V.; Dhiman, N.; Chauhan, L.K.S.; Pasricha, R.; Pandey, A.K. Physico-chemical properties based differential toxicity of graphene oxide/reduced graphene oxide in human lung cells mediated through oxidative stress. Sci. Rep. 2016, 6, 39548. [Google Scholar] [CrossRef]
- Hu, W.; Peng, C.; Luo, W.; Lv, M.; Li, X.; Li, D.; Huang, Q.; Fan, C. Graphene-based antibacterial paper. ACS Nano 2010, 4, 4317–4323. [Google Scholar] [CrossRef]
- Chng, E.L.K.; Pumera, M. The toxicity of graphene oxides: Dependence on the oxidative methods used. Chem. A Eur. J. 2013, 19, 8227–8235. [Google Scholar] [CrossRef]
- Das, S.; Singh, S.; Singh, V.; Joung, D.; Dowding, J.M.; Reid, D.; Anderson, J.; Zhai, L.; Khondaker, S.I.; Self, W.T.; et al. Oxygenated functional group density on graphene oxide: Its effect on cell toxicity. Part. Part. Syst. Charact. 2013, 30, 148–157. [Google Scholar] [CrossRef]
- Wu, W.; Yan, L.; Wu, Q.; Li, Y.; Li, Q.; Chen, S.; Yang, Y.; Gu, Z.; Xu, H.; Yin, Z.Q. Evaluation of the toxicity of graphene oxide exposure to the eye. Nanotoxicology 2016, 10, 1329–1340. [Google Scholar] [CrossRef]
- Mendonça, M.C.P.; Soares, E.S.; De Jesus, M.B.; Ceragioli, H.J.; Batista, Â.G.; Nyúl-Tóth, Á.; Molnár, J.; Wilhelm, I.; Maróstica, M.R.; Krizbai, I.; et al. PEGylation of reduced graphene oxide induces toxicity in cells of the blood–brain barrier: An in vitro and in vivo study. Mol. Pharm. 2016, 13, 3913–3924. [Google Scholar] [CrossRef] [PubMed]
- Pustovit, V.N.; Shahbazyan, T.V. Fluorescence quenching near small metal nanoparticles. J. Chem. Phys. 2012, 136, 204701. [Google Scholar] [CrossRef] [PubMed]
- Kasry, A.; Ardakani, A.A.; Tulevski, G.S.; Menges, B.; Copel, M.; Vyklicky, L. Highly efficient fluorescence quenching with graphene. J. Phys. Chem. C 2012, 116, 2858–2862. [Google Scholar] [CrossRef]
- Wu, X.; Xing, Y.; Zeng, K.; Huber, K.; Zhao, J.X. Study of fluorescence quenching ability of graphene oxide with a layer of rigid and tunable silica spacer. Langmuir 2018, 34, 603–611. [Google Scholar] [CrossRef]
- Ou, L.; Lin, S.; Song, B.; Liu, J.; Lai, R.; Shao, L. The mechanisms of graphene-based materials-induced programmed cell death: A review of apoptosis, autophagy, and programmed necrosis. Int. J. Nanomed. 2017, 12, 6633–6646. [Google Scholar] [CrossRef]
- Zhang, Y.; Ali, S.F.; Dervishi, E.; Xu, Y.; Li, Z.; Casciano, D.; Biris, A.S. Cytotoxicity effects of graphene and single-wall carbon nanotubes in neural phaeochromocytoma-derived PC12 cells. ACS Nano 2010, 4, 3181–3186. [Google Scholar] [CrossRef]
- Markovic, Z.M.; Ristic, B.Z.; Arsikin, K.M.; Klisic, D.G.; Harhaji-Trajkovic, L.M.; Todorovic-Markovic, B.M.; Kepic, D.P.; Kravic-Stevovic, T.K.; Jovanovic, S.P.; Milenkovic, M.M.; et al. Graphene quantum dots as autophagy-inducing photodynamic agents. Biomaterials 2012, 33, 7084–7092. [Google Scholar] [CrossRef]
- Tabish, T.A.; Pranjol, M.Z.I.; Jabeen, F.; Abdullah, T.; Latif, A.; Khalid, A.; Ali, M.; Hayat, H.; Winyard, P.G.; Whatmore, J.L.; et al. Investigation into the toxic effects of graphene nanopores on lung cancer cells and biological tissues. Appl. Mater. Today 2018, 12, 389–401. [Google Scholar] [CrossRef]
- Adil, S.; Khan, M.; Khan, M.; Al-Marri, A.; Al-Warthan, A.; Alkhathlan, H.; Siddiqui, M.; Nayak, V.; Kamal, A. Apoptosis inducing ability of silver decorated highly reduced graphene oxide nanocomposites in A549 lung cancer. Int. J. Nanomed. 2016, 11, 873. [Google Scholar] [CrossRef]
- Mbeh, D.A.; Akhavan, O.; Javanbakht, T.; Mahmoudi, M.; Yahia, L. Cytotoxicity of protein corona-graphene oxide nanoribbons on human epithelial cells. Appl. Surf. Sci. 2014, 320, 596–601. [Google Scholar] [CrossRef]
- Yuan, X.; Liu, Z.; Guo, Z.; Ji, Y.; Jin, M.; Wang, X. Cellular distribution and cytotoxicity of graphene quantum dots with different functional groups. Nanoscale Res. Lett. 2014, 9, 108. [Google Scholar] [CrossRef] [PubMed]
- Zhu, S.; Luo, F.; Zhu, B.; Wang, G.-X. Toxicological effects of graphene oxide on Saccharomyces cerevisiae. Toxicol. Res. (Camb.) 2017, 6, 535–543. [Google Scholar] [CrossRef]
- Zhu, S.; Zhu, B.; Huang, A.; Hu, Y.; Wang, G.; Ling, F. Toxicological effects of multi-walled carbon nanotubes on Saccharomyces cerevisiae: The uptake kinetics and mechanisms and the toxic responses. J. Hazard. Mater. 2016, 318, 650–662. [Google Scholar] [CrossRef]
- Zhu, S.; Luo, F.; Li, J.; Zhu, B.; Wang, G.-X. Biocompatibility assessment of single-walled carbon nanotubes using Saccharomyces cerevisiae as a model organism. J. Nanobiotechnol. 2018, 16, 44. [Google Scholar] [CrossRef] [PubMed]
- Oliveira, R.; Johansson, B. Quantitative DNA damage and repair measurement with the yeast comet assay. Methods Mol. Biol. 2012, 920, 101–109. [Google Scholar] [PubMed]
- Mortimer, M.; Kasemets, K.; Heinlaan, M.; Kurvet, I.; Kahru, A. High throughput kinetic Vibrio fischeri bioluminescence inhibition assay for study of toxic effects of nanoparticles. Toxicol. Vitr. 2008, 22, 1412–1417. [Google Scholar] [CrossRef] [PubMed]
- Patel, S.K.S.; Choi, S.H.; Kang, Y.C.; Lee, J.-K. Eco-friendly composite of Fe3O4-reduced graphene oxide particles for efficient enzyme immobilization. ACS Appl. Mater. Interfaces 2017, 9, 2213–2222. [Google Scholar] [CrossRef] [PubMed]
- Contreras Rodríguez, A.R.; Saiz-Poseu, J.; García-Pardo, J.; García, B.; Lorenzo, J.; Ojea-Jiménez, I.; Komilis, D.; Sedó, J.; Busqué, F.; Sánchez, A.; et al. Biocompatible polydopamine-like particles for the removal of heavy metals at extremely low concentrations. RSC Adv. 2016, 6, 40058–40066. [Google Scholar] [CrossRef]
- Ríos, F.; Fernández-Arteaga, A.; Fernández-Serrano, M.; Jurado, E.; Lechuga, M. Silica micro-and nanoparticles reduce the toxicity of surfactant solutions. J. Hazard. Mater. 2018, 353, 436–443. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Zhang, J.; Huang, X.; Zhou, X.; Wu, H.; Guo, S. Assembly of graphene oxide-enzyme conjugates through hydrophobic interaction. Small 2012, 8, 154–159. [Google Scholar] [CrossRef] [PubMed]
- Barbero, F.; Russo, L.; Vitali, M.; Piella, J.; Salvo, I.; Borrajo, M.L.; Busquets-Fité, M.; Grandori, R.; Bastús, N.G.; Casals, E.; et al. Formation of the protein corona: The interface between nanoparticles and the immune system. Semin. Immunol. 2017, 34, 52–60. [Google Scholar] [CrossRef] [PubMed]
- Chong, Y.; Ge, C.; Yang, Z.; Garate, J.A.; Gu, Z.; Weber, J.K.; Liu, J.; Zhou, R. Reduced cytotoxicity of graphene nanosheets mediated by blood-protein coating. ACS Nano 2015, 9, 5713–5724. [Google Scholar] [CrossRef] [PubMed]
- Wei, X.-Q.; Hao, L.-Y.; Shao, X.-R.; Zhang, Q.; Jia, X.-Q.; Zhang, Z.-R.; Lin, Y.-F.; Peng, Q. Insight into the interaction of graphene oxide with serum proteins and the impact of the degree of reduction and concentration. ACS Appl. Mater. Interfaces 2015, 7, 13367–13374. [Google Scholar] [CrossRef] [PubMed]
- Qi, Y.; Chen, W.; Liu, F.; Liu, J.; Zhang, T.; Chen, W. Aggregation morphology is a key factor determining protein adsorption on graphene oxide and reduced graphene oxide nanomaterials. Environ. Sci. Nano 2019, 6, 1303–1309. [Google Scholar] [CrossRef]
- Zhang, Y.; Wu, C.; Guo, S.; Zhang, J. Interactions of graphene and graphene oxide with proteins and peptides. Nanotechnol. Rev. 2013, 2, 27–45. [Google Scholar] [CrossRef]
- Ou, L.; Song, B.; Liang, H.; Liu, J.; Feng, X.; Deng, B.; Sun, T.; Shao, L. Toxicity of graphene-family nanoparticles: A general review of the origins and mechanisms. Part. Fibre Toxicol. 2016, 13, 57. [Google Scholar] [CrossRef]
- James, J.; Fiji, N.; Roy, D.; Andrew MG, D.; Shihabudeen, M.S.; Chattopadhyay, D.; Thirumurugan, K. A rapid method to assess reactive oxygen species in yeast using H2DCF-DA. Anal. Methods 2015, 7, 8572–8575. [Google Scholar] [CrossRef]
- Jarque, S.; Masner, P.; Klánová, J.; Prokeš, R.; Bláha, L. Bioluminescent Vibrio fischeri assays in the assessment of seasonal and spatial patterns in toxicity of contaminated river sediments. Front. Microbiol. 2016, 7, 1738. [Google Scholar] [CrossRef]

| GO (ppm) | GOC (ppm) | |
|---|---|---|
| Al | 0.160 ± 0.113 | 1.445 ± 0.106 |
| B | <0.004 | 1.600 ± 0.255 |
| Ba | 0.006 ± 0.008 | 0.214 ± 0.006 |
| Ca | 0.063 ± 0.088 | 0.835 ± 0.035 |
| Cu | 0.052 ± 0.039 | 0.581 ± 0.030 |
| Fe | 0.379 ± 0.067 | 1.899 ± 0.033 |
| Ga | 0.004 ± 0.006 | 0.047 ± 0.000 |
| K | 3.770 ± 0.184 | 2.628 ± 0.252 |
| Mg | 0.350 ± 0.028 | 2.000 ± 0.113 |
| Mn | 34.700 ± 0.156 | 62.405 ± 0.233 |
| Mo | 0.029 ± 0.002 | 0.017 ± 0.001 |
| Na | 1.240 ± 0.509 | 4.810 ± 0.057 |
| Ni | 0.027 ± 0.020 | 0.027 ± 0.007 |
| Pb | 0.054 ± 0.023 | 0.152 ± 0.009 |
| S | 43.200 ± 2.786 | 5.084 ± 2.752 |
| Sn | 0.003 ± 0.003 | 0.034 ± 0.001 |
| Sr | 0.008 ± 0.001 | 0.034 ± 0.001 |
| V | <0.0001 | 0.006 ± 0.001 |
| W | 0.004 ± 0.001 | 0.006 ± 0.001 |
| Zn | 0.068 ± 0.061 | 1.069 ± 0.740 |
| Carbon Nanomaterial | RhaB1 Binding (mg mg−1) | AgB Binding (mg mg−1) |
|---|---|---|
| GO | 4.88 ± 0.17 | 1.65 ± 0.04 |
| GOC | 5.90 ± 0.11 | 1.22 ± 0.14 |
| rGO | 1.98 ± 0.11 | 1.00 ± 0.03 |
| rGOC | 1.99 ± 0.23 | 0.70 ± 0.08 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Domi, B.; Rumbo, C.; García-Tojal, J.; Elena Sima, L.; Negroiu, G.; Tamayo-Ramos, J.A. Interaction Analysis of Commercial Graphene Oxide Nanoparticles with Unicellular Systems and Biomolecules. Int. J. Mol. Sci. 2020, 21, 205. https://doi.org/10.3390/ijms21010205
Domi B, Rumbo C, García-Tojal J, Elena Sima L, Negroiu G, Tamayo-Ramos JA. Interaction Analysis of Commercial Graphene Oxide Nanoparticles with Unicellular Systems and Biomolecules. International Journal of Molecular Sciences. 2020; 21(1):205. https://doi.org/10.3390/ijms21010205
Chicago/Turabian StyleDomi, Brixhilda, Carlos Rumbo, Javier García-Tojal, Livia Elena Sima, Gabriela Negroiu, and Juan Antonio Tamayo-Ramos. 2020. "Interaction Analysis of Commercial Graphene Oxide Nanoparticles with Unicellular Systems and Biomolecules" International Journal of Molecular Sciences 21, no. 1: 205. https://doi.org/10.3390/ijms21010205
APA StyleDomi, B., Rumbo, C., García-Tojal, J., Elena Sima, L., Negroiu, G., & Tamayo-Ramos, J. A. (2020). Interaction Analysis of Commercial Graphene Oxide Nanoparticles with Unicellular Systems and Biomolecules. International Journal of Molecular Sciences, 21(1), 205. https://doi.org/10.3390/ijms21010205







