Association Analysis of Salt Tolerance in Asiatic cotton (Gossypium arboretum) with SNP Markers
Abstract
1. Introduction
2. Results
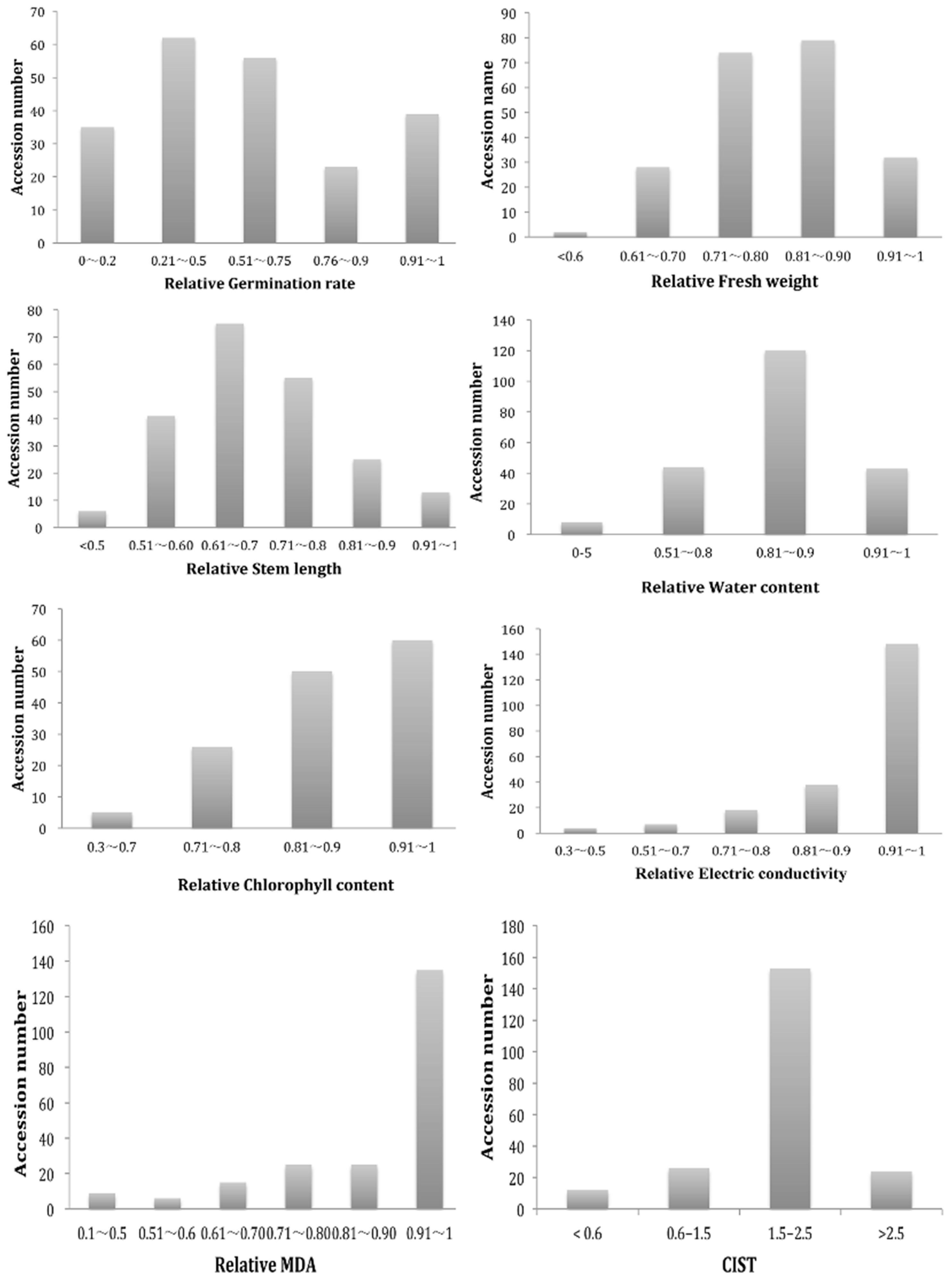
2.1. Phenotypic Diversity of G. arboretum Population
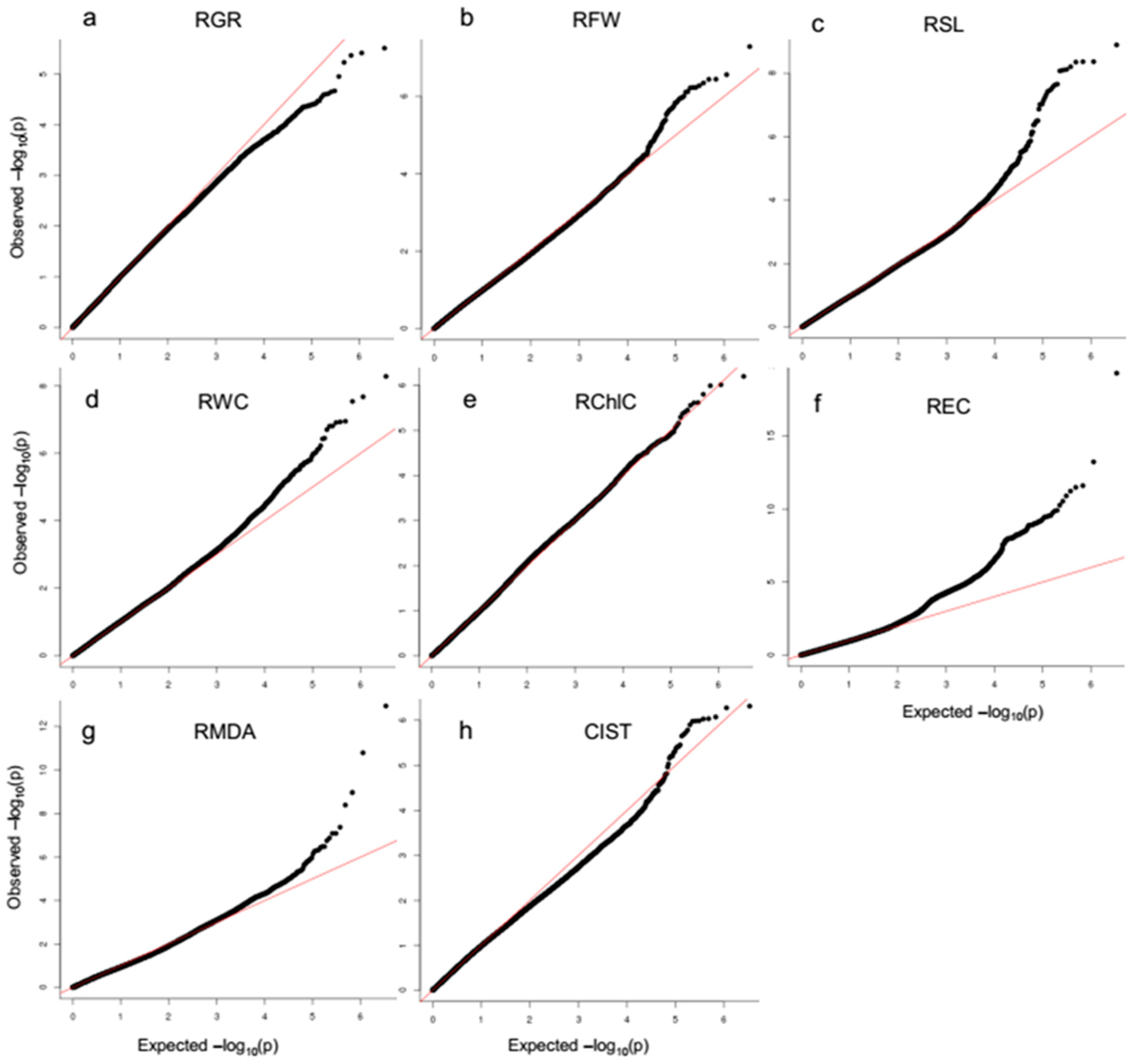
2.2. Association Mapping of Salt Tolerance Related Traits Using SNP Markers
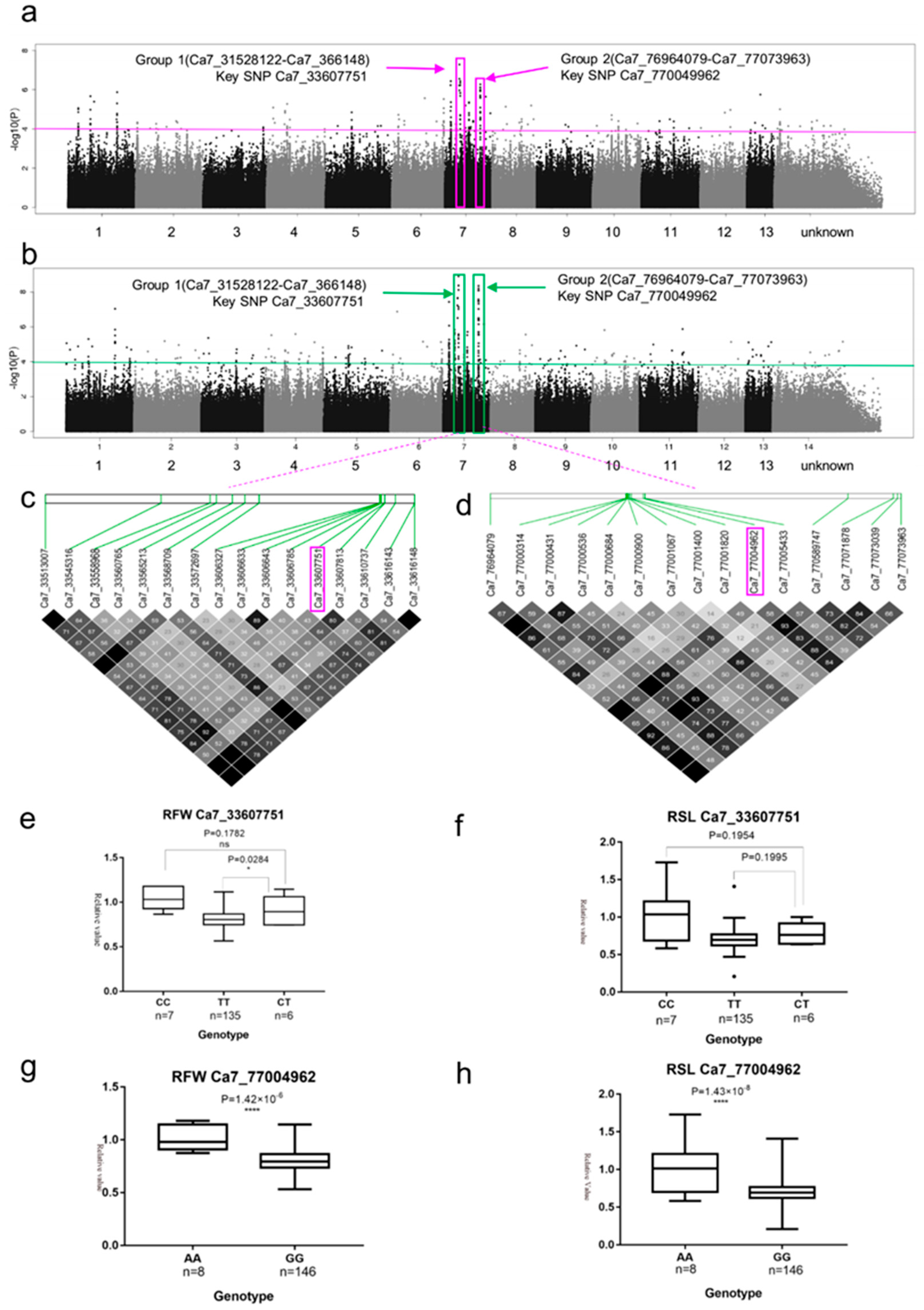
2.3. SNP Rich Regions Associated with RFW and RSL
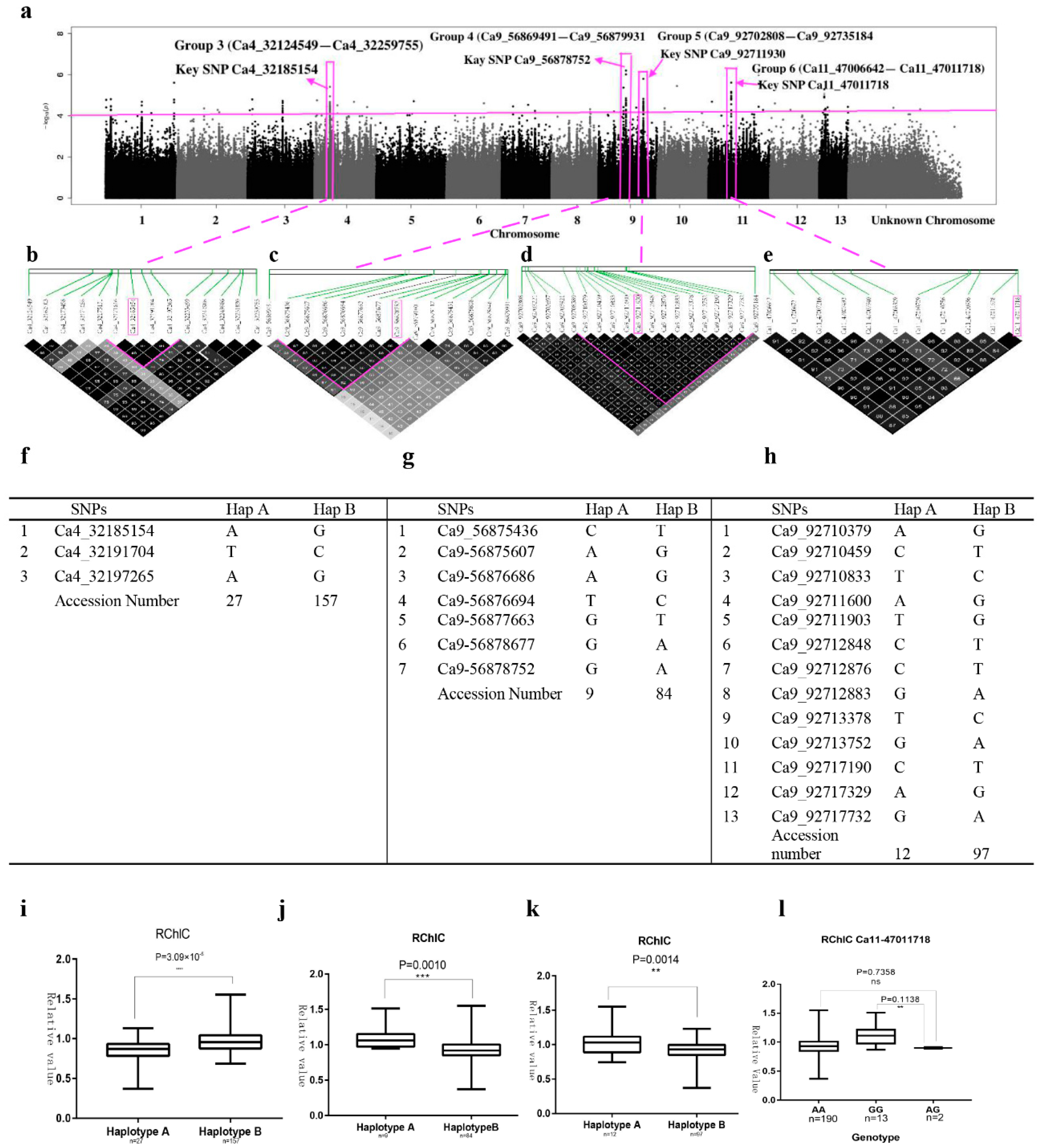
2.4. SNP Rich Regions Associated with RChlC
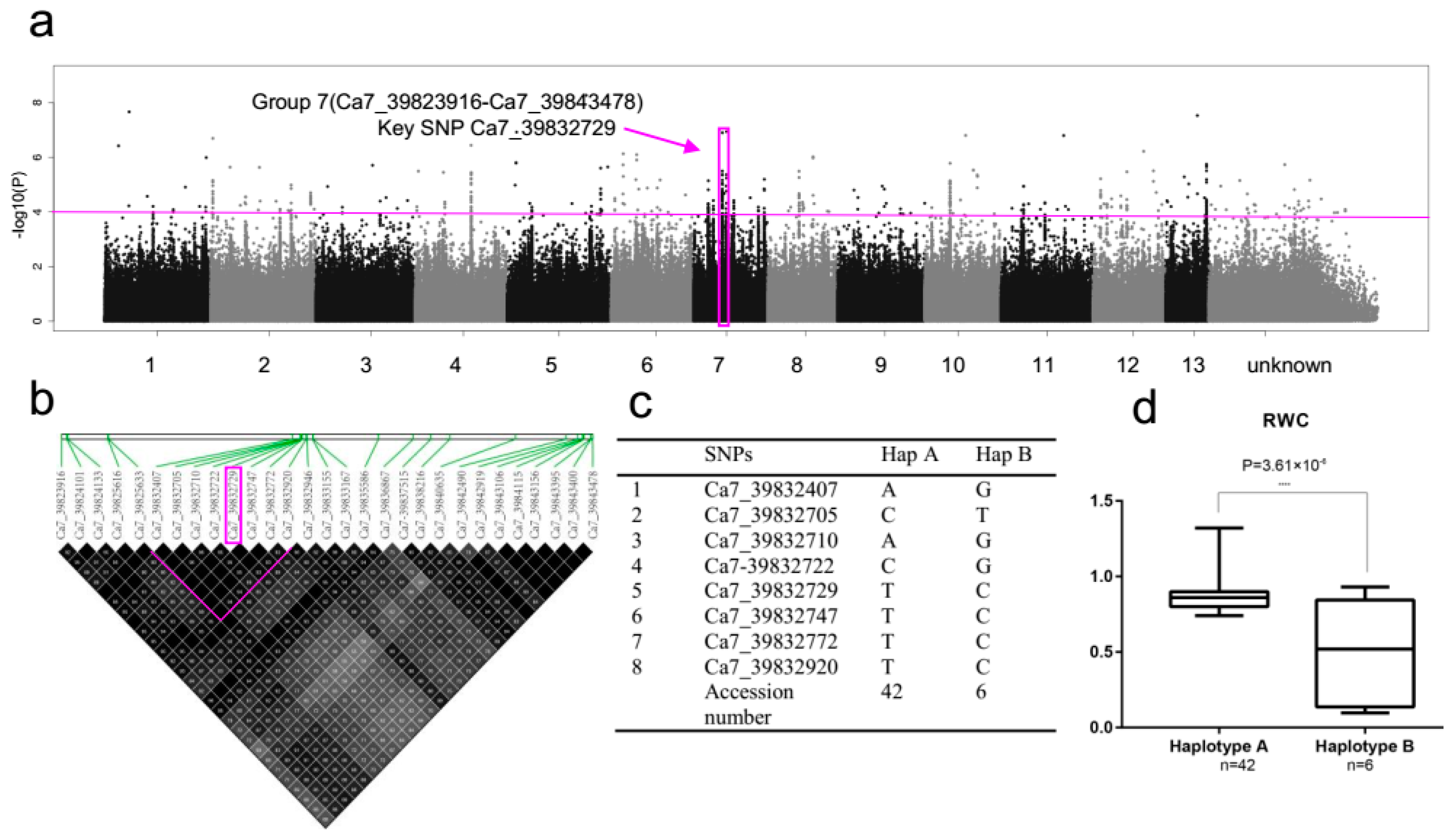
2.5. SNP Rich Region Associated with RWC
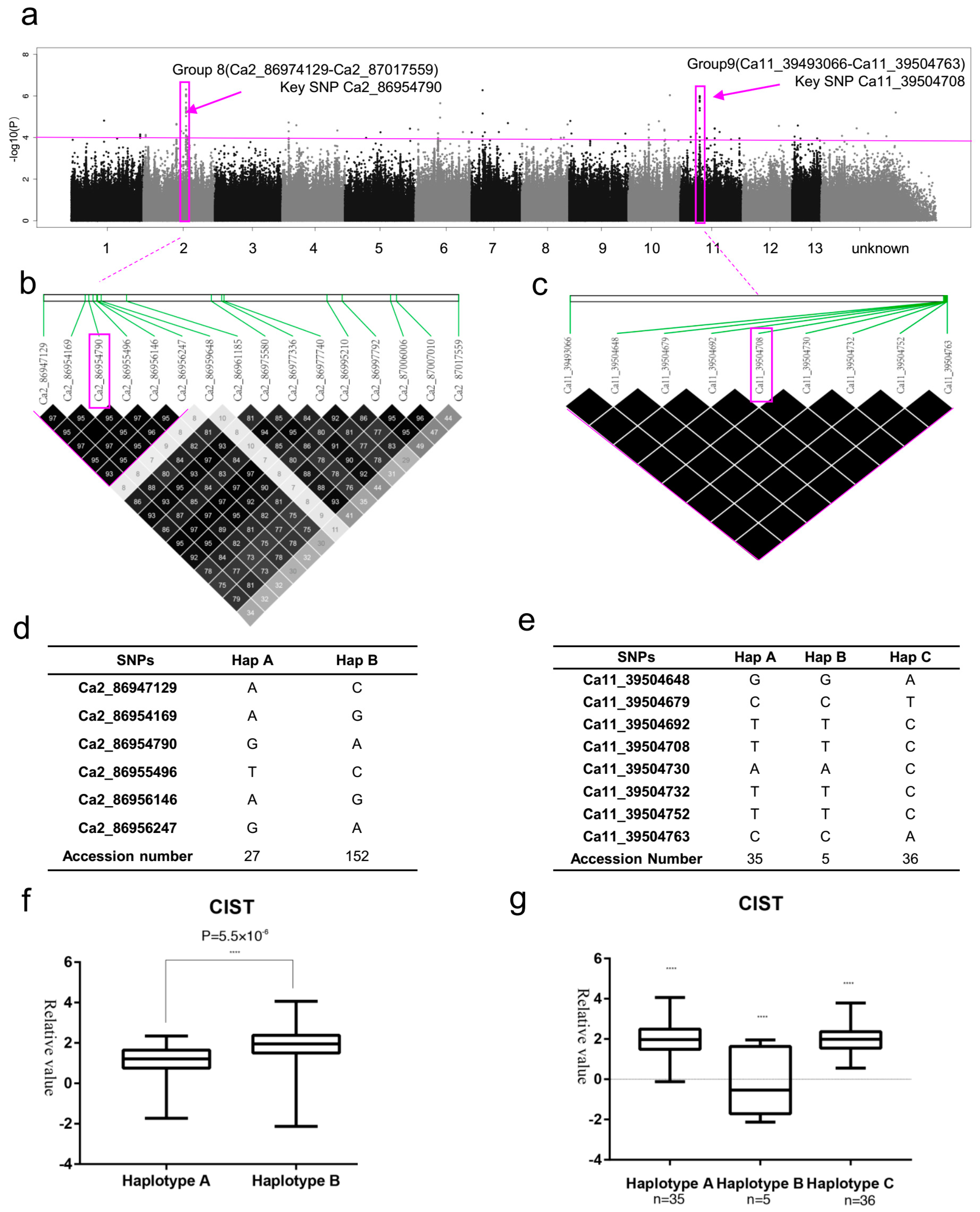
2.6. SNP Rich Regions Associated with CIST
3. Discussion
3.1. Genetic Variation in Salt Tolerance Related Traits of G. arboretum Accessions
3.2. Association Mapping of Salt Tolerance Traits Using SNP Markers
4. Materials and Methods
4.1. Plant Materials and Sample Preparation
4.2. Trait Evaluation
4.3. DNA Extraction
4.4. Phenotypic Diversity
4.5. Genome-Wide Association Analysis
4.6. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| RGR | Relative germination rate |
| RFW | Relative fresh weight |
| RSL | Relative stem length |
| RWC | Relative water content |
| RChlC | Relative chlorophyll content; |
| REC | Relative electric conduct; |
| RMDA | Relative methylene dioxyamphetamine |
| CIST | Comprehensive index of salt tolerance |
| MMLM | Multi-Locus Mixed Model |
| QTL | Quantitative trait loci; |
| SSR | Simple sequence repeats |
| SNP | Single-nucleotide polymorphism |
| TW | Turgid weights; |
| ANOVA | Analysis of variance; |
| Quantile–quantile plots | |
| LD | Linkage disequilibrium; |
| GWAS | Genome-wide association study; |
| HR | Homologous recombination |
| ABA | Abscisic acid; |
| GHs | Glycosyl hydrolases |
References
- Wang, Y.; Deng, C.; Liu, Y.; Niu, Z.; Li, Y. Identifying change in spatial accumulation of soil salinity in an inland river watershed, China. Sci. Total Environ. 2018, 621, 177–185. [Google Scholar] [CrossRef] [PubMed]
- Nematzadeh, G.A. Salt-related Genes Expression in Salt-Tolerant and Salt-Sensitive Cultivars of Cotton (Gossypium sp. L.) under NaCl Stress. J. Plant Mol. Breed. 2018. [Google Scholar] [CrossRef]
- Peng, Z.; He, S.; Sun, J.; Pan, Z.; Gong, W.; Lu, Y.; Du, X. Na+ compartmentalization related to salinity stress tolerance in upland cotton (Gossypium hirsutum) seedlings. Sci. Rep. 2016, 6, 34548. [Google Scholar] [CrossRef] [PubMed]
- Paul, D.; Lade, H. Plant-growth-promoting rhizobacteria to improve crop growth in saline soils: A review. Agron. Sustain. Dev. 2014, 34, 737–752. [Google Scholar] [CrossRef]
- Gao, W.; Xu, F.C.; Guo, D.D.; Zhao, J.R.; Liu, J.; Guo, Y.W.; Singh, P.K.; Ma, X.N.; Long, L.; Botella, J.R. Calcium-dependent protein kinases in cotton: Insights into early plant responses to salt stress. BMC Plant Biol. 2018, 18, 15. [Google Scholar] [CrossRef] [PubMed]
- Munns, R.; Tester, M. Mechanisms of Salinity Tolerance. Annu. Rev. Plant Biol. 2008, 59, 651–681. [Google Scholar] [CrossRef] [PubMed]
- Gray, S.B.; Brady, S.M. Plant Developmental Responses to Climate Change. Dev. Biol. 2016, 419, 64–77. [Google Scholar] [CrossRef]
- Ahmad, S.; Ashraf, M.; Khan, N. Genetic basis of salt-tolerance in cotton (Gossypium hirsutum L.). Sci. Technol. Dev. 2004, 23, 45–50. [Google Scholar]
- Frouin, J.; Languillaume, A.; Mas, J.; Mieulet, D.; Boisnard, A.; Labeyrie, A.; Bettembourg, M.; Bureau, C.; Lorenzini, E.; Portefaix, M. Tolerance to mild salinity stress in japonica rice: A genome-wide association mapping study highlights calcium signaling and metabolism genes. PLoS ONE 2018, 13, e0190964. [Google Scholar] [CrossRef] [PubMed]
- Peng, Z.; He, S.; Gong, W.; Sun, J.; Pan, Z.; Xu, F.; Lu, Y.; Du, X. Comprehensive analysis of differentially expressed genes and transcriptional regulation induced by salt stress in two contrasting cotton genotypes. BMC Genom. 2014, 15, 760. [Google Scholar] [CrossRef]
- Parida, A.K.; Das, A.B. Salt tolerant and salinity effects on plants. Ecotoxicol. Environ. Saf. 2005, 60, 324–349. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; Gao, S.; Xu, S.; Zhang, Z.; Prasanna, B.M.; Li, L.; Li, J.; Yan, J. Characterization of a global germplasm collection and its potential utilization for analysis of complex quantitative traits in maize. Mol. Breed. 2011, 28, 511–526. [Google Scholar] [CrossRef]
- Abbasi, Z.; Majidi, M.M.; Arzani, A.; Rajabi, A.; Mashayekhi, P.; Bocianowski, J. Association of SSR markers and morpho-physiological traits associated with salinity tolerance in sugar beet (Beta vulgaris L.). Euphytica 2015, 205, 785–797. [Google Scholar] [CrossRef]
- Kantartzi, S.; Stewart, J.M. Association analysis of fibre traits in Gossypium arboreum accessions. Plant Breed. 2008, 127, 173–179. [Google Scholar] [CrossRef]
- Du, L.; Cai, C.; Wu, S.; Zhang, F.; Hou, S.; Guo, W. Evaluation and exploration of favorable QTL alleles for salt stress related traits in cotton cultivars (G. hirsutum L.). PLoS ONE 2016, 11, e0151076. [Google Scholar] [CrossRef]
- Cai, C.; Zhu, G.; Zhang, T.; Guo, W. High-density 80 K SNP array is a powerful tool for genotyping G. hirsutum accessions and genome analysis. BMC Genom. 2017, 18, 654. [Google Scholar] [CrossRef]
- Sun, Z.; Li, H.; Zhang, Y.; Li, Z.; Ke, H.; Wu, L.; Zhang, G.; Wang, X.; Ma, Z. Identification of SNPs and Candidate Genes Associated with Salt Tolerance at the Seedling Stage in Cotton (Gossypium hirsutum L.). Front. Plant Sci. 2018, 9, 1011. [Google Scholar] [CrossRef] [PubMed]
- Cai, C.; Wu, S.; Niu, E.; Cheng, C.; Guo, W. Identification of genes related to salt stress tolerance using intron-length polymorphic markers, association mapping and virus-induced gene silencing in cotton. Sci. Rep. 2017, 7, 528. [Google Scholar] [CrossRef]
- Wang, S.; Chen, J.; Zhang, W.; Hu, Y.; Chang, L.; Fang, L.; Wang, Q.; Lv, F.; Wu, H.; Si, Z. Sequence-based ultra-dense genetic and physical maps reveal structural variations of allopolyploid cotton genomes. Genome Biol. 2015, 16, 108. [Google Scholar] [CrossRef] [PubMed]
- Tan, Z.; Zhang, Z.; Sun, X.; Li, Q.; Sun, Y.; Yang, P.; Wang, W.; Liu, X.; Chen, C.; Liu, D. Genetic Map Construction and Fiber Quality QTL Mapping Using the CottonSNP80K Array in Upland Cotton. Front. Plant Sci. 2018, 9, 225. [Google Scholar] [CrossRef]
- Atwell, S.; Huang, Y.S.; Vilhjálmsson, B.J.; Willems, G.; Horton, M.; Li, Y.; Meng, D.; Platt, A.; Tarone, A.M.; Hu, T.T. Genome-wide association study of 107 phenotypes in Arabidopsis thaliana inbred lines. Nature 2010, 465, 627. [Google Scholar] [CrossRef] [PubMed]
- Xue, Y.; Warburton, M.L.; Sawkins, M.; Zhang, X.; Setter, T.; Xu, Y.; Grudloyma, P.; Gethi, J.; Ribaut, J.-M.; Li, W. Genome-wide association analysis for nine agronomic traits in maize under well-watered and water-stressed conditions. Theor. Appl. Genet. 2013, 126, 2587–2596. [Google Scholar] [CrossRef]
- Chen, G.; Zhang, H.; Deng, Z.; Wu, R.; Li, D.; Wang, M.; Tian, J. Genome-wide association study for kernel weight-related traits using SNPs in a Chinese winter wheat population. Euphytica 2016, 212, 173–185. [Google Scholar] [CrossRef]
- McCouch, S.R.; Wright, M.H.; Tung, C.-W.; Maron, L.G.; McNally, K.L.; Fitzgerald, M.; Singh, N.; DeClerck, G.; Agosto-Perez, F.; Korniliev, P. Open access resources for genome-wide association mapping in rice. Nat. Commun. 2016, 7, 10532. [Google Scholar] [CrossRef]
- Abdelraheem, A.; Fang, D.D.; Zhang, J. Quantitative trait locus mapping of drought and salt tolerance in an introgressed recombinant inbred line population of Upland cotton under the greenhouse and field conditions. Euphytica 2018, 214, 8. [Google Scholar] [CrossRef]
- Zhao, Y.L.; Wang, H.M.; Shao, B.X. SSR-based association mapping of salt tolerance in cotton (Gossypium hirsutum L.). Genet. Mol. Res. 2016, 15, gmr-15027370. [Google Scholar] [CrossRef] [PubMed]
- Saeed, M.; Guo, W.Z.; Zhang, T.Z. Association mapping for salinity tolerance in cotton (Gossypium hirsutum L.) germplasm from US and diverse regions of China. Aust. J. Crop Sci. 2014, 8, 338–346. [Google Scholar]
- Jia, Y.H.; Sun, J.L.; Wang, X.W.; Zhou, Z.L.; Pan, Z.E.; He, S.P.; Pang, B.Y. Molecular Diversity and Association Analysis of Drought and Salt Tolerance in Gossypium hirsutum L. Germplasm. J. Integr. Agric. 2014, 13, 1845–1853. [Google Scholar] [CrossRef]
- Liu, F.; Zhou, Z.L.; Wang, C.Y.; Wang, Y.H.; Cai, X.Y.; Wang, X.X.; Zhang, Z.S.; Wang, K.B. Genetic diversity and relationship analysis of Gossypium arboreum accessions. Genet. Mol. Res. 2015, 14, gmr-14522. [Google Scholar] [CrossRef] [PubMed]
- Du, X.; Huang, G.; He, S.; Yang, Z.; Sun, G.; Ma, X.; Li, N.; Zhang, X.; Sun, J.; Liu, M. Resequencing of 243 diploid cotton accessions based on an updated A genome identifies the genetic basis of key agronomic traits. Nat. Genet. 2018, 50, 796. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Fan, G.; Wang, K.; Sun, F.; Yuan, Y.; Song, G.; Li, Q.; Ma, Z.; Lu, C.; Zou, C. Genome sequence of the cultivated cotton Gossypium arboreum. Nat. Genet. 2014, 46, 567. [Google Scholar] [CrossRef]
- Mehetre, S.S.; Aher, A.R.; Gawande, V.L.; Patil, V.R.; Mokate, A.S. Induced polyploidy in Gossypium: A tool to overcome interspecific incompatibility of cultivated tetraploid and diploid cottons. Curr. Sci. 2003, 84, 1510–1512. [Google Scholar]
- Peng, Z.; He, S.; Sun, J.; Xu, F.; Jia, Y.; Pan, Z.; Wang, L. An Efficient Approach to Identify Salt Tolerance of Upland Cotton at Seedling Stage. Acta Agron. Sin. 2014, 40, 476–486. [Google Scholar] [CrossRef]
- Zhao, Y.; Wang, H.; Wei, C.; Li, Y. Genetic Structure, Linkage Disequilibrium and Association Mapping of Verticillium Wilt Resistance in Elite Cotton (Gossypium hirsutum L.) Germplasm Population. PLoS ONE 2014, 9, e86308. [Google Scholar] [CrossRef]
- Sun, Z.; Wang, X.; Liu, Z.; Gu, Q.; Zhang, Y.; Li, Z.; Ke, H.; Yang, J.; Wu, J.; Wu, L. Genome-wide association study discovered genetic variation and candidate genes of fibre quality traits in Gossypium hirsutum L. Plant Biotechnol. J. 2017, 15, 982–996. [Google Scholar] [CrossRef] [PubMed]
- Song, H.M.; Zhao, R.M.; Fan, P.X.; Wang, X.C.; Chen, X.Y.; Li, Y.X. Overexpression of AtHsp90.2, AtHsp90.5 and AtHsp90.7 in Arabidopsis thaliana enhances plant sensitivity to salt and drought stresses. Planta 2009, 229, 955–964. [Google Scholar] [CrossRef] [PubMed]
- Schopf, F.H.; Biebl, M.M.; Buchner, J. The HSP90 chaperone machinery. Nat. Rev. Mol. Cell Biol. 2017, 18, 345. [Google Scholar] [CrossRef] [PubMed]
- Zhou, X.H.; Li, X.S.; Wang, P.; Yan, B.L.; Teng, Y.J.; Yi, L.F. Molecular cloning and expression analysis of HSP90 gene from Porphyra yezoensis Ueda (Bangiales, Rhodophyta). J. Fish. China 2010, 34, 1844–1852. [Google Scholar]
- Ding, Z.; Yang, G.; Wu, J. Effect of Heat Shock at Germinating Period on Growing Developmental of Cotton. J. Wuhan Bot. Res. 2006, 24, 579–582. [Google Scholar]
- Sangster, T.A.; Bahrami, A.; Wilczek, A.; Watanabe, E.; Schellenberg, K.; Mclellan, C.; Kelley, A.; Kong, S.W.; Queitsch, C.; Lindquist, S. Phenotypic Diversity and Altered Environmental Plasticity in Arabidopsis thaliana with Reduced Hsp90 Levels. PLoS ONE 2007, 2, e648. [Google Scholar] [CrossRef] [PubMed]
- Busch, W.; Wunderlich, H.F. Identification of novel heat shock factor-dependent genes and biochemical pathways in Arabidopsis thaliana. Plant J. Cell Mol. Biol. 2010, 41, 1–14. [Google Scholar] [CrossRef]
- Juan, G.; Yan, Z.; Wangbin, Z.; Jean, M.; Aiwu, D.; Wen-Hui, S. NAP1 family histone chaperones are required for somatic homologous recombination in Arabidopsis. Plant Cell 2012, 24, 1437–1447. [Google Scholar]
- Zhu, Y.; Dong, A.; Meyer, D.; Pichon, O.; Renou, J.P.; Cao, K.; Shen, W.H. Arabidopsis NRP1 and NRP2 encode histone chaperones and are required for maintaining postembryonic root growth. Plant Cell 2006, 18, 2879–2892. [Google Scholar] [CrossRef]
- Waśkiewicz, A.; Beszterda, M.; Goliński, P. ABA: Role in Plant Signaling Under Salt Stress. Salt Stress Plants 2013, 175–196. [Google Scholar] [CrossRef]
- Asaoka, R.; Uemura, T.; Nishida, S.; Fujiwara, T.; Ueda, T.; Nakano, A. New insights into the role of Arabidopsis RABA1 GTPases in salinity stress tolerance. Plant Signal. Behav. 2013, 8, e25377. [Google Scholar] [CrossRef][Green Version]
- Park, M.Y.; Chung, M.S.; Koh, H.S.; Lee, D.J.; Ahn, S.J.; Kim, C.S. Isolation and functional characterization of the Arabidopsis salt-tolerance 32 (AtSAT32) gene associated with salt tolerance and ABA signaling. Physiol. Plant. 2010, 135, 426–435. [Google Scholar] [CrossRef]
- Wang, T.Z.; Xia, X.Z.; Zhao, M.G.; Tian, Q.Y.; Zhang, W.H. Expression of a Medicago falcata small GTPase gene, MfARL1 enhanced tolerance to salt stress in Arabidopsis thaliana. Plant Physiol. Biochem. 2013, 63, 227–235. [Google Scholar] [CrossRef]
- Sripinyowanich, S.; Klomsakul, P.; Boonburapong, B.; Bangyeekhun, T.; Asami, T.; Gu, H.; Buaboocha, T.; Chadchawan, S. Exogenous ABA induces salt tolerance in indica rice (Oryza sativa L.): The role of OsP5CS1 and OsP5CR gene expression during salt stress. Environ. Exp. Bot. 2013, 86, 94–105. [Google Scholar] [CrossRef]
- Noaman, M.M.; Dvorak, J.; Dong, J.M. Genes inducing salt tolerance in wheat, Lophopyrum elongatum and amphiploid and their responses to ABA under salt stress. In Prospects for Saline Agriculture; Springer: Dordrecht, The Netherlands, 2002; pp. 139–144. [Google Scholar]
- Zhao, K.F.; Fan, H.; Harris, P. Effect of exogenous aba on the salt tolerance of corn seedlings under salt stress. Acta Bot. Sin. 1995, 37, 295–300. [Google Scholar]
- O’Mahony, P.J.; Oliver, M.J. Characterization of a desiccation-responsive small GTP-binding protein (Rab2) from the desiccation-tolerant grass Sporobolus stapfianus. Plant Mol. Biol. 1999, 39, 809–821. [Google Scholar] [CrossRef]
- Nahm, M.Y.; Kim, S.W.; Yun, D.; Lee, S.Y.; Cho, M.J.; Bahk, J.D. Molecular and biochemical analyses of OsRab7, a rice Rab7 homolog. Plant Cell Physiol. 2003, 44, 1341–1349. [Google Scholar] [CrossRef] [PubMed]
- Mazel, A.; Leshem, Y.; Tiwari, B.S.; Levine, A. Induction of salt and osmotic stress tolerance by overexpression of an intracellular vesicle trafficking protein AtRab7 (AtRabG3e). Plant Physiol. 2004, 134, 118–128. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.J.; Tang, C.L.; Lin, D.; Cai, G.L.; Liu, X.Y.; Bo, L.; Han, Q.M.; Buchenauer, H.; Wei, G.R.; Han, D.J. Characterization of a pathogenesis-related thaumatin-like protein gene TaPR5 from wheat induced by stripe rust fungus. Physiol. Plant. 2010, 139, 27–38. [Google Scholar] [CrossRef]
- Liu, J.J.; Sturrock, R.; Ekramoddoullah AK, M. The superfamily of thaumatin-like proteins: Its origin, evolution, and expression towards biological function. Plant Cell Rep. 2010, 29, 419–436. [Google Scholar] [CrossRef]
- Vashisht, A.A.; Tuteja, N. Stress responsive DEAD-box helicases: A new pathway to engineer plant stress tolerance. J. Photochem. Photobiol. B Biol. 2006, 84, 150–160. [Google Scholar] [CrossRef]
- Opassiri, R.; Pomthong, B.; Akiyama, T.; Nakphaichit, M.; Onkoksoong, T.; Cairns, M.K.; Cairns, J.R.K. A stress-induced rice (Oryza sativa L.) β-glucosidase represents a new subfamily of glycosyl hydrolase family 5 containing a fascin-like domain. Biochem. J. 2007, 408, 241–249. [Google Scholar] [CrossRef] [PubMed]
- Kang, H.M.; Sul, J.H.; Service, S.K.; Zaitlen, N.A.; Kong, S.-Y.; Freimer, N.B.; Sabatti, C.; Eskin, E. Variance component model to account for sample structure in genome-wide association studies. Nat. Genet. 2010, 42, 348. [Google Scholar] [CrossRef] [PubMed]
| Traits 1 | Mean | SD | Min | Max | CV | Mean Square | F | P > F |
|---|---|---|---|---|---|---|---|---|
| GR | 19.936 | 10.77 | 0 | 40 | 54.02 | 338.55 | 8.16 | <0.0001 |
| FW | 0.371 | 0.089 | 0.02 | 0.77 | 24.2 | 0.128 | 34.16 | <0.0001 |
| SL | 4.969 | 1.481 | 0.40 | 10.2 | 29.84 | 40.77 | 49.76 | <0.0001 |
| WC | 1.112 | 3.481 | 0.36 | 96.97 | 312.9 | 31.83 | 4.01 | <0.0001 |
| ChlC | 39.47 | 9.96 | 3.50 | 50.2 | 25.53 | 6763.5 | 8.31 | <0.0001 |
| EC | 31.47 | 42.54 | 0.116 | 844.4 | 135.19 | 541.2 | 6.49 | <0.0001 |
| MDA | 0.007 | 0.0046 | 0 | 0.04 | 59.8 | 0.000095 | 14.36 | <0.0001 |
| Trait 1 | GR | FW | SL | WC | ChlC | EC | MDA |
|---|---|---|---|---|---|---|---|
| GR | 1 | 0.22 *** | 0.295 *** | −0.0178 | 0.112 ** | 0.0841 | 0.012 |
| FW | 1 | 0.575 *** | −0.043 | 0.070 *** | 0.0613 | 0.135 *** | |
| SL | 1 | −0.017 | 0.115 *** | −0.051 | 0.030 | ||
| WC | 1 | 0.048 | 0.002 | 0.059 * | |||
| ChlC | 1 | 0.1 ** | −0.073 * | ||||
| EC | 1 | 0.079 * | |||||
| MDA | 1 |
| Chromosome | Total | RGR | RFW | RSL | RWC | RChlC | REC | RMDA | CIST |
|---|---|---|---|---|---|---|---|---|---|
| Chr-1 | 163 | 1 | 26 | 22 | 12 | 15 | 14 | 70 | 3 |
| Chr-2 | 152 | 5 | 10 | 10 | 39 | 3 | 12 | 49 | 24 |
| Chr-3 | 332 | 1 | 3 | 8 | 7 | 11 | 278 | 24 | 0 |
| Chr-4 | 201 | 3 | 12 | 26 | 27 | 51 | 59 | 19 | 4 |
| Chr-5 | 155 | 2 | 7 | 17 | 21 | 4 | 94 | 8 | 2 |
| Chr-6 | 112 | 10 | 16 | 11 | 22 | 2 | 21 | 23 | 7 |
| Chr-7 | 295 | 2 | 67 | 108 | 67 | 6 | 24 | 14 | 7 |
| Chr-8 | 99 | 15 | 10 | 8 | 27 | 4 | 9 | 24 | 2 |
| Chr-9 | 104 | 4 | 2 | 8 | 9 | 47 | 7 | 24 | 3 |
| Chr-10 | 80 | 1 | 7 | 10 | 34 | 4 | 11 | 7 | 6 |
| Chr-11 | 112 | 5 | 5 | 8 | 17 | 20 | 24 | 18 | 15 |
| Chr-12 | 57 | 8 | 4 | 2 | 31 | 3 | 5 | 3 | 1 |
| Chr-13 | 100 | 1 | 4 | 10 | 29 | 16 | 6 | 32 | 2 |
| Chr-UN | 100 | 3 | 14 | 7 | 28 | 4 | 19 | 20 | 5 |
| Total | 2062 | 61 | 187 | 255 | 370 | 190 | 583 | 335 | 81 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dilnur, T.; Peng, Z.; Pan, Z.; Palanga, K.K.; Jia, Y.; Gong, W.; Du, X. Association Analysis of Salt Tolerance in Asiatic cotton (Gossypium arboretum) with SNP Markers. Int. J. Mol. Sci. 2019, 20, 2168. https://doi.org/10.3390/ijms20092168
Dilnur T, Peng Z, Pan Z, Palanga KK, Jia Y, Gong W, Du X. Association Analysis of Salt Tolerance in Asiatic cotton (Gossypium arboretum) with SNP Markers. International Journal of Molecular Sciences. 2019; 20(9):2168. https://doi.org/10.3390/ijms20092168
Chicago/Turabian StyleDilnur, Tussipkan, Zhen Peng, Zhaoe Pan, Koffi Kibalou Palanga, Yinhua Jia, Wenfang Gong, and Xiongming Du. 2019. "Association Analysis of Salt Tolerance in Asiatic cotton (Gossypium arboretum) with SNP Markers" International Journal of Molecular Sciences 20, no. 9: 2168. https://doi.org/10.3390/ijms20092168
APA StyleDilnur, T., Peng, Z., Pan, Z., Palanga, K. K., Jia, Y., Gong, W., & Du, X. (2019). Association Analysis of Salt Tolerance in Asiatic cotton (Gossypium arboretum) with SNP Markers. International Journal of Molecular Sciences, 20(9), 2168. https://doi.org/10.3390/ijms20092168





