Identification and Expression Profiling of the Regulator of Chromosome Condensation 1 (RCC1) Gene Family in Gossypium Hirsutum L. under Abiotic Stress and Hormone Treatments
Abstract
1. Introduction
2. Results
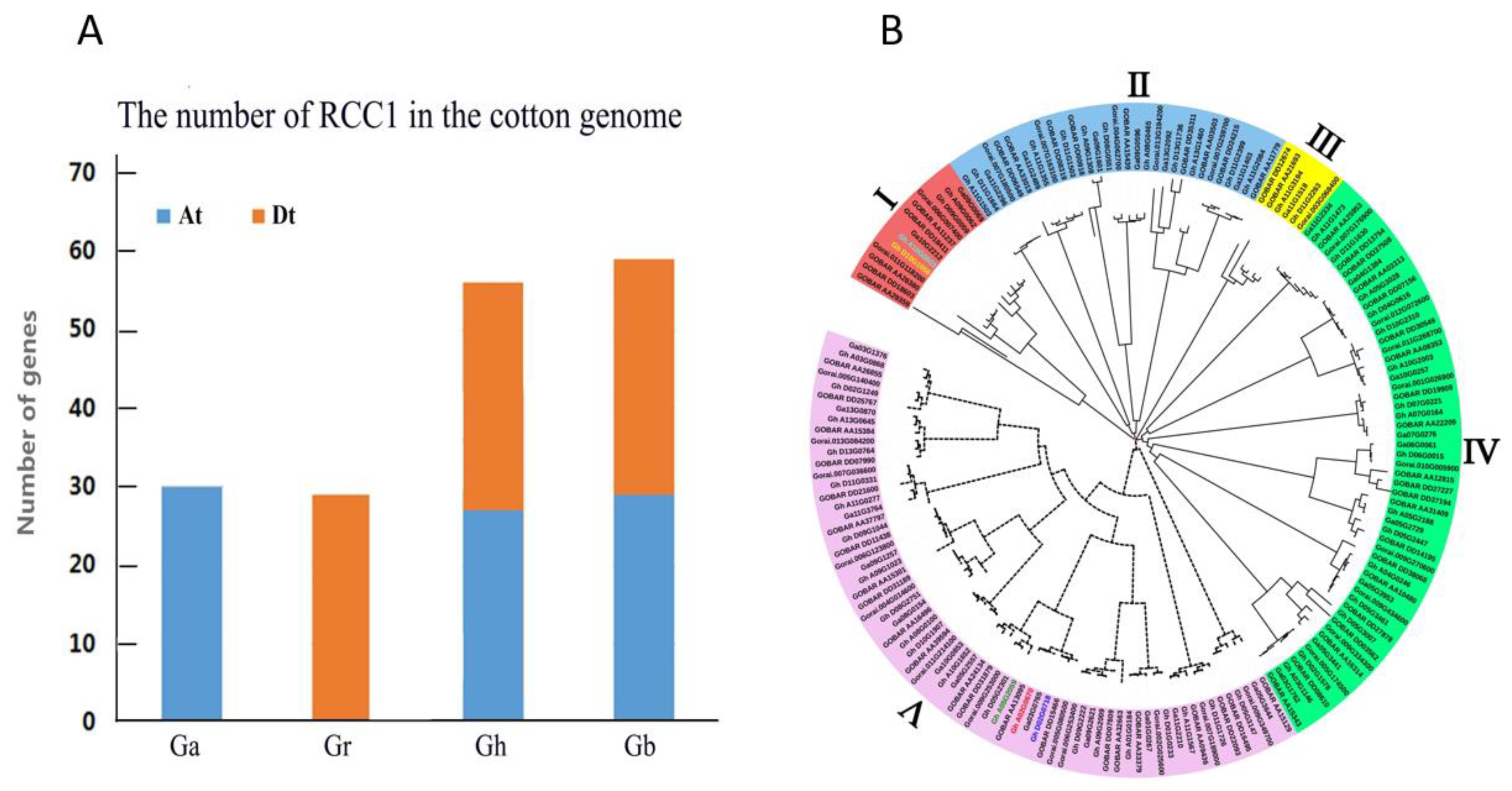
2.1. Comparison of RCC1 Genes Among Four Cotton Species
2.2. Phylogenetic Analysis of the RCC1 Gene Family
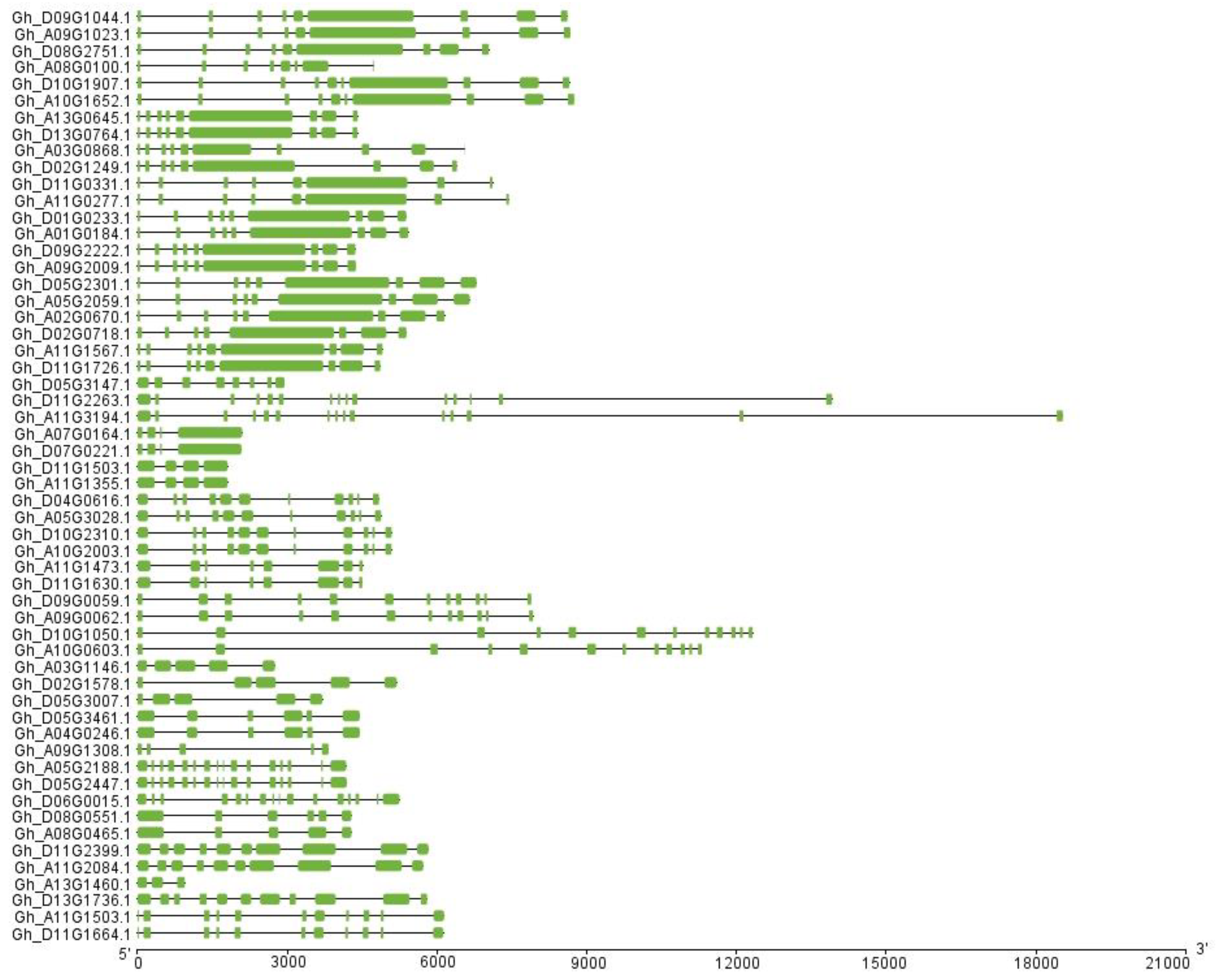
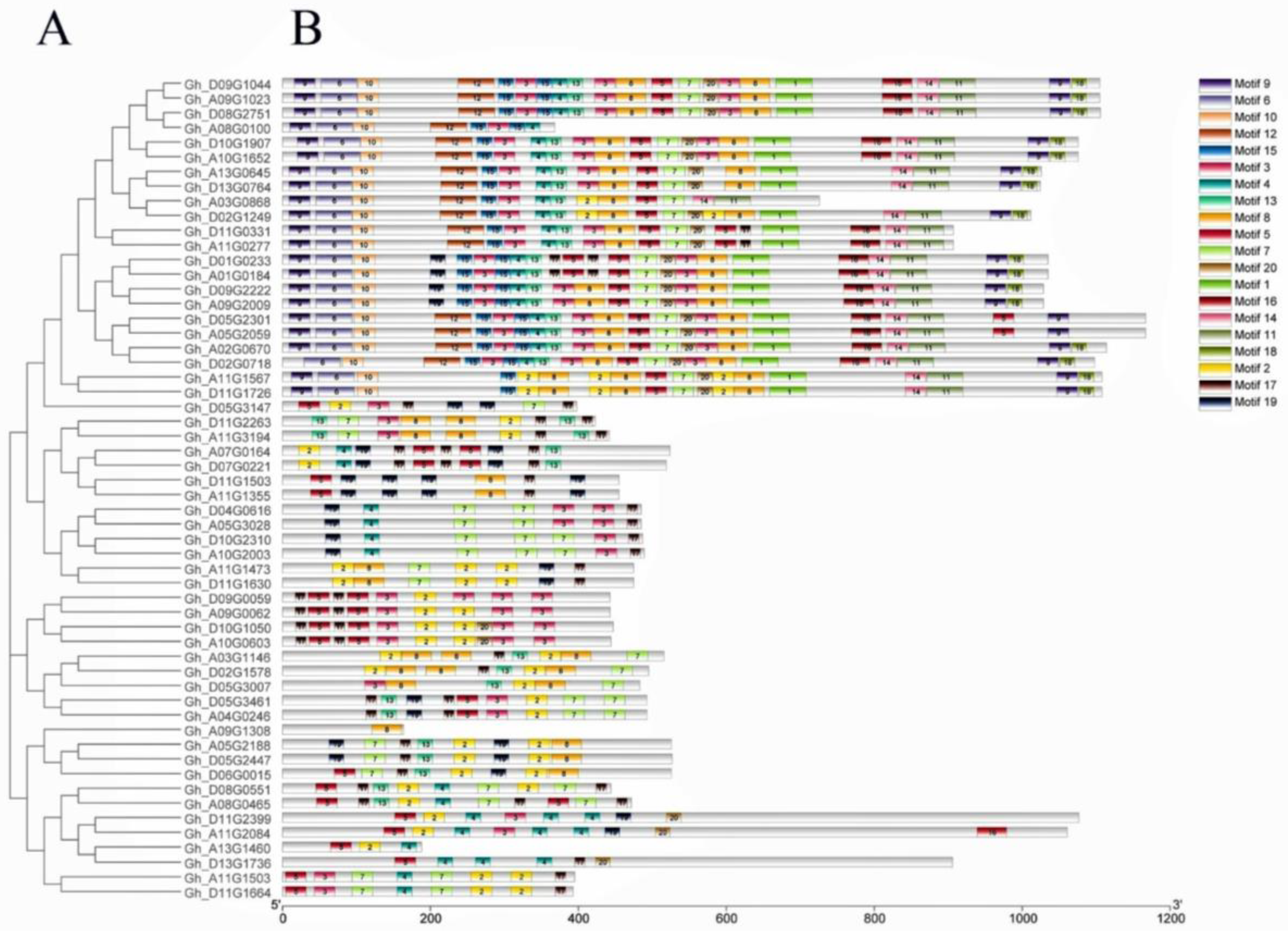
2.3. Exon–Intron Structure, Motif Assay, and Chromosomal Location Analyses
2.4. Analyses of Tissue-Specific Expressions
2.5. Expression of GhRCC1 Genes in Response to Various Hormone Treatments
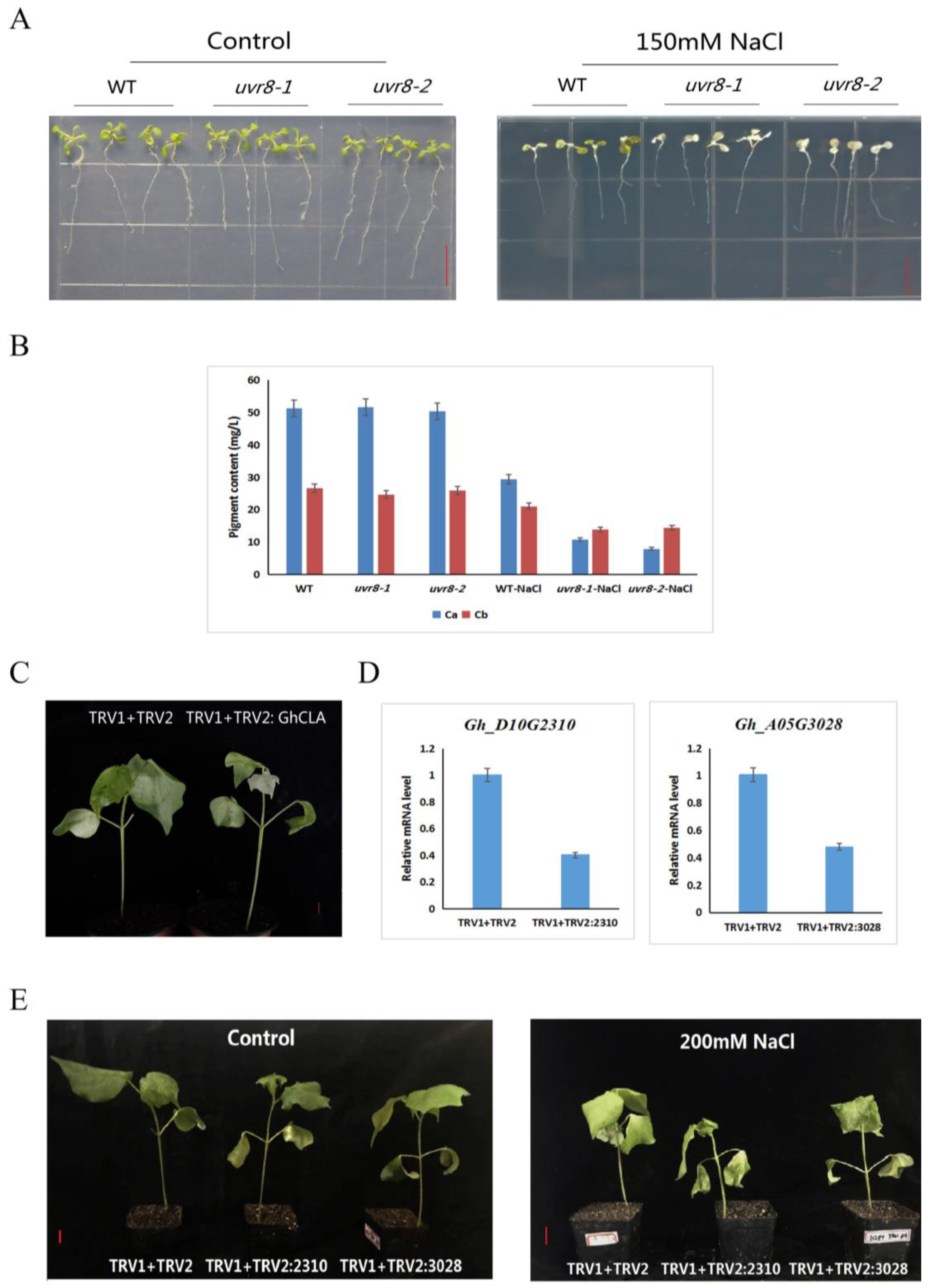
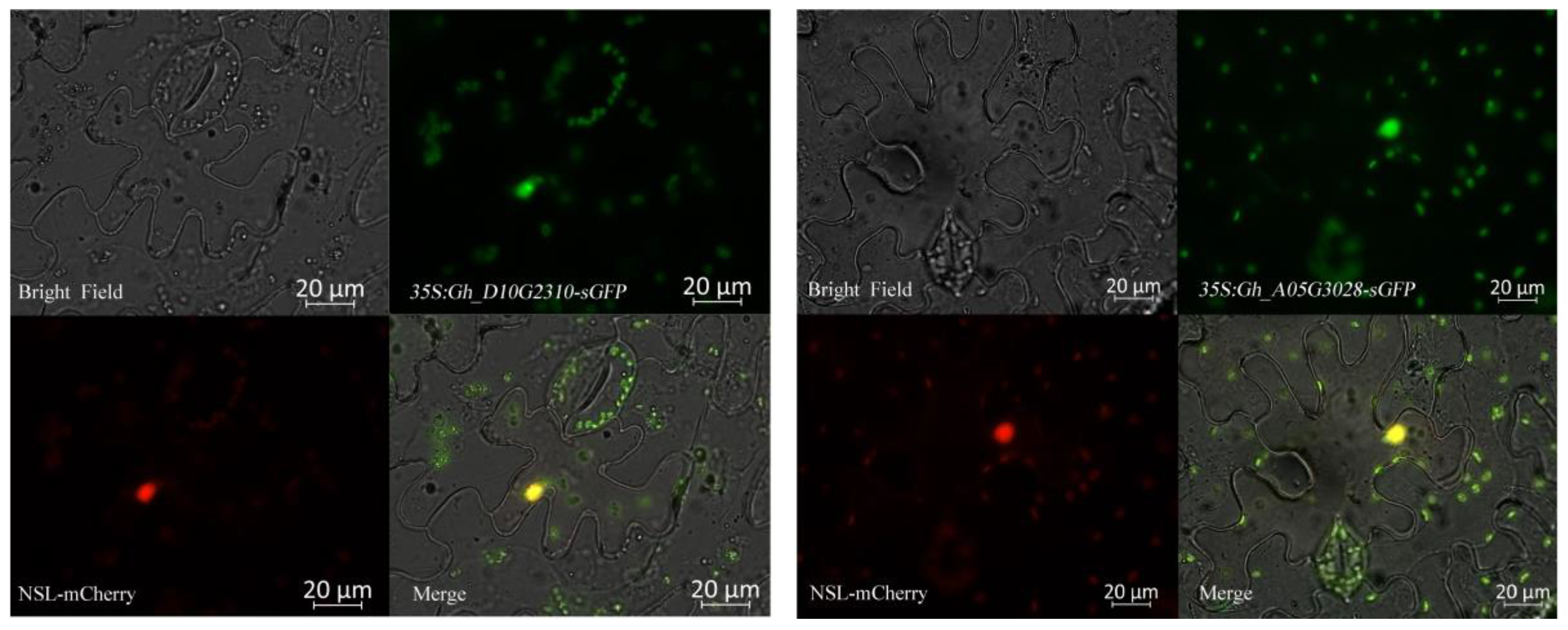
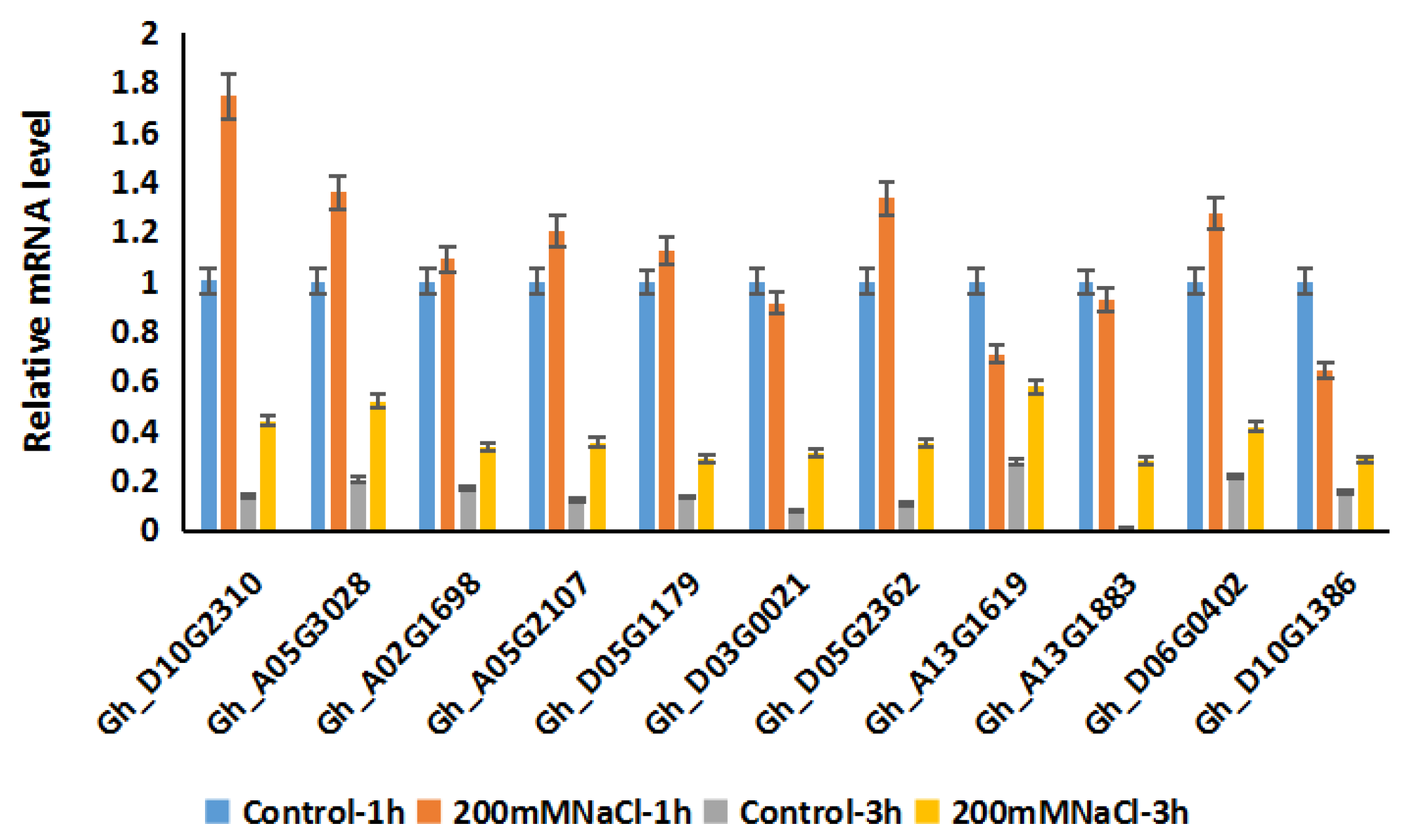
2.6. Expression Patterns of GhRCC1 Genes in Response to Various Abiotic Stress Treatments and Subcellular Location
3. Discussion
4. Materials and Methods
4.1. Plant Materials and Treatment
4.2. Identification of RCC1 Genes in Gossypium spp.
4.3. Phylogenetic Analysis
4.4. Exon/Intron Structure, Motif Assay and Chromosomal Location Analyses
4.5. RNA Isolation and Quantitative Real-Time Polymerase Chain Reaction (qRT-PCR)
4.6. Identification of T-DNA Insertional Mutants
4.7. Subcellular Localization
4.8. Gene Ontology Enrichment and Kyoto Encyclopedia of Genes and Genomes (KEGG) Analysis
4.9. VIGS (Virus-Induced Gene Silencing) of the Gh_A05G3028 and Gh_D10G2310 in Cotton
4.10. Determination of Chlorophyll Content
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Sexton, P.D.; Gerard, C.J. Emergence Force of Cotton Seedlings as Influenced by Salinity1. Agron. J. 1982, 74, 699–702. [Google Scholar] [CrossRef]
- Brugnoli, E.; Lauteri, M. Effects of Salinity on Stomatal Conductance, Photosynthetic Capacity, and Carbon Isotope Discrimination of Salt-Tolerant (Gossypium hirsutum L.) and Salt-Sensitive (Phaseolus vulgaris L.) C(3) Non-Halophytes. Plant Physiol. 1991, 95, 628–635. [Google Scholar] [CrossRef] [PubMed]
- Gerard, C.J. Emergence force by cotton seedlings. Agron. J. 1980, 72, 473–476. [Google Scholar] [CrossRef]
- Park, H.J.; Kim, W.Y.; Yun, D.J. A New Insight of Salt Stress Signaling in Plant. Mol. Cells 2016, 39, 447–459. [Google Scholar] [CrossRef] [PubMed]
- Peng, Z.; He, S.; Gong, W.; Xu, F.; Pan, Z.; Jia, Y.; Geng, X.; Du, X. Integration of proteomic and transcriptomic profiles reveals multiple levels of genetic regulation of salt tolerance in cotton. BMC Plant Biol. 2018, 18, 128. [Google Scholar] [CrossRef]
- Tanveer, M.; Shah, A.N. An insight into salt stress tolerance mechanisms of Chenopodium album. Environ. Sci. Pollut. Res. Int. 2017, 24, 16531–16535. [Google Scholar] [CrossRef] [PubMed]
- Allakhverdiev, S.I.; Kinoshita, M.; Inaba, M.; Suzuki, I.; Murata, N. Unsaturated fatty acids in membrane lipids protect the photosynthetic machinery against salt-induced damage in Synechococcus. Plant Physiol. 2001, 125, 1842–1853. [Google Scholar] [CrossRef]
- Bischoff, F.R.; Ponstingl, H. Catalysis of guanine nucleotide exchange on Ran by the mitotic regulator RCC1. Nature 1991, 354, 80–82. [Google Scholar] [CrossRef] [PubMed]
- Moore, M.S.; Blobel, G. The GTP-binding protein Ran/TC4 is required for protein import into the nucleus. Nature 1993, 365, 661–663. [Google Scholar] [CrossRef]
- Newport, J. A role for Ran-GTP and Crm1 in blocking re-replication. Cell 2004, 119, 145. [Google Scholar] [CrossRef]
- Ohtsubo, M.; Kai, R.; Furuno, N.; Sekiguchi, T.; Sekiguchi, M.; Hayashida, H.; Kuma, K.; Miyata, T.; Fukushige, S.; Murotsu, T.; et al. Isolation and characterization of the active cDNA of the human cell cycle gene (RCC1) involved in the regulation of onset of chromosome condensation. Genes Dev. 1987, 1, 585–593. [Google Scholar] [CrossRef]
- Lin, Y.L.; Chen, H.L.; Cheng, S.B.; Yeh, D.C.; Huang, C.C.; P’Eng, F.K.; Tsai, T.C.; Wu, C.C.; Chen, C.M. Methylation-silencing RCC1 expression is associated with tumorigenesis and depth of invasion in gastric cancer. Int. J. Clin. Exp. Pathol. 2015, 8, 14257–14269. [Google Scholar]
- Kliebenstein, D.J.; Lim, J.E.; Landry, L.G.; Last, R.L. Arabidopsis UVR8 regulates ultraviolet-B signal transduction and tolerance and contains sequence similarity to human regulator of chromatin condensation 1. Plant Physiol. 2002, 130, 234–243. [Google Scholar] [CrossRef]
- Brown, B.A.; Cloix, C.; Jiang, G.H.; Kaiserli, E.; Herzyk, P.; Kliebenstein, D.J.; Jenkins, G.I. A UV-B-specific signaling component orchestrates plant UV protection. Proc. Natl. Acad. Sci. USA 2005, 102, 18225–18230. [Google Scholar] [CrossRef]
- Favory, J.J.; Stec, A.; Gruber, H.; Rizzini, L.; Oravecz, A.; Funk, M.; Albert, A.; Cloix, C.; Jenkins, G.I.; Oakeley, E.J.; et al. Interaction of COP1 and UVR8 regulates UV-B-induced photomorphogenesis and stress acclimation in Arabidopsis. EMBO J. 2009, 28, 591–601. [Google Scholar] [CrossRef]
- Kaiserli, E.; Jenkins, G.I. UV-B promotes rapid nuclear translocation of the Arabidopsis UV-B specific signaling component UVR8 and activates its function in the nucleus. Plant Cell 2007, 19, 2662–2673. [Google Scholar] [CrossRef]
- Rizzini, L.; Favory, J.J.; Cloix, C.; Faggionato, D.; O’Hara, A.; Kaiserli, E.; Baumeister, R.; Schafer, E.; Nagy, F.; Jenkins, G.I.; et al. Perception of UV-B by the Arabidopsis UVR8 protein. Science 2011, 332, 103–106. [Google Scholar] [CrossRef]
- Yin, R.; Skvortsova, M.Y.; Loubery, S.; Ulm, R. COP1 is required for UV-B-induced nuclear accumulation of the UVR8 photoreceptor. Proc. Natl. Acad. Sci. USA 2016, 113, E4415–E4422. [Google Scholar] [CrossRef]
- Kuhn, K.; Carrie, C.; Giraud, E.; Wang, Y.; Meyer, E.H.; Narsai, R.; des Francs-Small, C.C.; Zhang, B.; Murcha, M.W.; Whelan, J. The RCC1 family protein RUG3 is required for splicing of nad2 and complex I biogenesis in mitochondria of Arabidopsis thaliana. Plant. J. 2011, 67, 1067–1080. [Google Scholar] [CrossRef]
- Ji, H.; Wang, Y.; Cloix, C.; Li, K.; Jenkins, G.I.; Wang, S.; Shang, Z.; Shi, Y.; Yang, S.; Li, X. The Arabidopsis RCC1 Family Protein TCF1 Regulates Freezing Tolerance and Cold Acclimation through Modulating Lignin Biosynthesis. PLoS Genet. 2015, 11, e1005471. [Google Scholar] [CrossRef]
- Wywial, E.; Singh, S.M. Identification and structural characterization of FYVE domain-containing proteins of Arabidopsis thaliana. BMC Plant. Biol 2010, 10, 157. [Google Scholar]
- Hopkins, J.; Pierre, O.; Kazmierczak, T.; Gruber, V.; Frugier, F.; Clement, M.; Frendo, P.; Herouart, D.; Boncompagni, E. MtZR1, a PRAF protein, is involved in the development of roots and symbiotic root nodules in Medicago truncatula. Plant Cell Environ. 2014, 37, 658–669. [Google Scholar] [CrossRef]
- Zhang, T.; Hu, Y.; Jiang, W.; Fang, L.; Guan, X.; Chen, J.; Zhang, J.; Saski, C.A.; Scheffler, B.E.; Stelly, D.M.; et al. Sequencing of allotetraploid cotton (Gossypium hirsutum L. acc. TM-1) provides a resource for fiber improvement. Nat. Biotechnol. 2015, 33, 531–537. [Google Scholar] [CrossRef]
- Bai, W.Q.; Xiao, Y.H.; Zhao, J.; Song, S.Q.; Hu, L.; Zeng, J.Y.; Li, X.B.; Hou, L.; Luo, M.; Li, D.M.; et al. Gibberellin overproduction promotes sucrose synthase expression and secondary cell wall deposition in cotton fibers. PLoS ONE 2014, 9, e96537. [Google Scholar] [CrossRef]
- Beasley, C.A.; Ting, I.P. Effects of Plant Growth Substances on in Vitro Fiber Development from Unfertilized Cotton Ovules. Am. J. Bot. 1974, 61, 188–194. [Google Scholar] [CrossRef]
- Gialvalis, S.; Seagull, R.W. Plant Hormones Alter Fiber Initiation in Unfertilized, Cultured Ovules of Gossypium hirsutum. J. Cotton Sci. 2001, 5, 252–258. [Google Scholar]
- Olszewski, N.; Sun, T.P.; Gubler, F. Gibberellin signaling: Biosynthesis, catabolism, and response pathways. Plant Cell 2002, 14 (Suppl. 1), S61–S80. [Google Scholar] [CrossRef]
- Qin, Y.M.; Hu, C.Y.; Pang, Y.; Kastaniotis, A.J.; Hiltunen, J.K.; Zhu, Y.X. Saturated very-long-chain fatty acids promote cotton fiber and Arabidopsis cell elongation by activating ethylene biosynthesis. Plant Cell 2007, 19, 3692–3704. [Google Scholar] [CrossRef]
- Seagull, R.W.; Giavalis, S.; Pre- Seagull, R.W.; Giavalis, S. Pre- and post-anthesis application of exogenous hormones alters fiber production in Gossypium hirsutum L. cultivar Maxxa GTO. J. Cotton Sci. 2004, 8, 105–111. [Google Scholar]
- Xiao, Y.H.; Li, D.M.; Yin, M.H.; Li, X.B.; Zhang, M.; Wang, Y.J.; Dong, J.; Zhao, J.; Luo, M.; Luo, X.Y.; et al. Gibberellin 20-oxidase promotes initiation and elongation of cotton fibers by regulating gibberellin synthesis. J. Plant Physiol. 2010, 167, 829–837. [Google Scholar] [CrossRef]
- Zhang, M.; Zheng, X.; Song, S.; Zeng, Q.; Hou, L.; Li, D.; Zhao, J.; Wei, Y.; Li, X.; Luo, M.; et al. Spatiotemporal manipulation of auxin biosynthesis in cotton ovule epidermal cells enhances fiber yield and quality. Nat. Biotechnol. 2011, 29, 453–458. [Google Scholar] [CrossRef]
- Houhoula, D.P.; Oreopoulou, V.; Tzia, C. The effect of process time and temperature on the accumulation of polar compounds in cottonseed oil during deep-fat frying. J. Sci. Food Agric. 2003, 83, 314–319. [Google Scholar] [CrossRef]
- Abe, H.; Kamai, T.; Shirataki, H.; Oyama, T.; Arai, K.; Yoshida, K. High expression of Ran GTPase is associated with local invasion and metastasis of human clear cell renal cell carcinoma. Int. J. Cancer 2008, 122, 2391–2397. [Google Scholar] [CrossRef]
- Cekan, P.; Hasegawa, K.; Pan, Y.; Tubman, E.; Odde, D.; Chen, J.Q.; Herrmann, M.A.; Kumar, S.; Kalab, P. RCC1-dependent activation of Ran accelerates cell cycle and DNA repair, inhibiting DNA damage-induced cell senescence. Mol. Biol. Cell 2016, 27, 1346–1357. [Google Scholar] [CrossRef]
- Kurisetty, V.V.; Johnston, P.G.; Johnston, N.; Erwin, P.; Crowe, P.; Fernig, D.G.; Campbell, F.C.; Anderson, I.P.; Rudland, P.S.; El-Tanani, M.K. RAN GTPase is an effector of the invasive/metastatic phenotype induced by osteopontin. Oncogene 2008, 27, 7139–7149. [Google Scholar] [CrossRef]
- Ly, T.K.; Wang, J.; Pereira, R.; Rojas, K.S.; Peng, X.; Feng, Q.; Cerione, R.A.; Wilson, K.F. Activation of the Ran GTPase is subject to growth factor regulation and can give rise to cellular transformation. J. Biol. Chem. 2010, 285, 5815–5826. [Google Scholar] [CrossRef]
- Xia, F.; Lee, C.W.; Altieri, D.C. Tumor cell dependence on Ran-GTP-directed mitosis. Cancer Res. 2008, 68, 1826–1833. [Google Scholar] [CrossRef]
- Yuen, H.F.; Chan, K.K.; Grills, C.; Murray, J.T.; Platt-Higgins, A.; Eldin, O.S.; O’Byrne, K.; Janne, P.; Fennell, D.A.; Johnston, P.G.; et al. Ran is a potential therapeutic target for cancer cells with molecular changes associated with activation of the PI3K/Akt/mTORC1 and Ras/MEK/ERK pathways. Clin. Cancer Res. 2012, 18, 380–391. [Google Scholar] [CrossRef]
- Heijde, M.; Ulm, R. UV-B photoreceptor-mediated signalling in plants. Trends Plant Sci. 2012, 17, 230–237. [Google Scholar] [CrossRef]
- Jenkins, G.I. The UV-B photoreceptor UVR8: From structure to physiology. Plant Cell 2014, 26, 21–37. [Google Scholar] [CrossRef]
- Li, J.; Yang, L.; Jin, D.; Nezames, C.D.; Terzaghi, W.; Deng, X.W. UV-B-induced photomorphogenesis in Arabidopsis. Protein Cell 2013, 4, 485–492. [Google Scholar] [CrossRef]
- Tilbrook, K.; Arongaus, A.B.; Binkert, M.; Heijde, M.; Yin, R.; Ulm, R. The UVR8 UV-B Photoreceptor: Perception, Signaling and Response. Arabidopsis Book 2013, 11, e0164. [Google Scholar] [CrossRef]
- Allen, V.G.; Baker, M.T.; Segarra, E.; Brown, C.P. Integrated Irrigated Crop—Livestock Systems in Dry Climates. Agron. J. 2007, 99, 346–360. [Google Scholar] [CrossRef]
- Xiong, L.; Schumaker, K.S.; Zhu, J.K. Cell Signaling during Cold, Drought, and Salt Stress. Plant Cell 2002, 14, S165. [Google Scholar] [CrossRef]
- Huala, E.; Dickerman, A.W.; Garcia-Hernandez, M.; Weems, D.; Reiser, L.; LaFond, F.; Hanley, D.; Kiphart, D.; Zhuang, M.; Huang, W.; et al. The Arabidopsis Information Resource (TAIR): A comprehensive database and web-based information retrieval, analysis, and visualization system for a model plant. Nucleic Acids Res. 2001, 29, 102–105. [Google Scholar] [CrossRef]
- Sievers, F.; Higgins, D.G. Clustal omega. Curr. Protoc. Bioinform. 2014, 48, 3–13. [Google Scholar]
- Letunic, I.; Bork, P. Interactive Tree of Life (iTOL): An online tool for phylogenetic tree display and annotation. Bioinformatics 2007, 23, 127–128. [Google Scholar] [CrossRef]
- O’Malley, R.C.; Barragan, C.C.; Ecker, J.R. A User’s Guide to the Arabidopsis T-DNA Insertion Mutant Collections. In Plant Functional Genomics; Humana Press: New York, NY, USA, 2015. [Google Scholar]
- Fradin, E.F.; Zhang, Z.; Juarez Ayala, J.C.; Castroverde, C.D.; Nazar, R.N.; Robb, J.; Liu, C.M.; Thomma, B.P. Genetic dissection of Verticillium wilt resistance mediated by tomato Ve1. Plant Physiol. 2009, 150, 320–332. [Google Scholar] [CrossRef]




© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, X.; Wu, X.; Sun, C.; Rong, J. Identification and Expression Profiling of the Regulator of Chromosome Condensation 1 (RCC1) Gene Family in Gossypium Hirsutum L. under Abiotic Stress and Hormone Treatments. Int. J. Mol. Sci. 2019, 20, 1727. https://doi.org/10.3390/ijms20071727
Liu X, Wu X, Sun C, Rong J. Identification and Expression Profiling of the Regulator of Chromosome Condensation 1 (RCC1) Gene Family in Gossypium Hirsutum L. under Abiotic Stress and Hormone Treatments. International Journal of Molecular Sciences. 2019; 20(7):1727. https://doi.org/10.3390/ijms20071727
Chicago/Turabian StyleLiu, Xiao, Xingchen Wu, Chendong Sun, and Junkang Rong. 2019. "Identification and Expression Profiling of the Regulator of Chromosome Condensation 1 (RCC1) Gene Family in Gossypium Hirsutum L. under Abiotic Stress and Hormone Treatments" International Journal of Molecular Sciences 20, no. 7: 1727. https://doi.org/10.3390/ijms20071727
APA StyleLiu, X., Wu, X., Sun, C., & Rong, J. (2019). Identification and Expression Profiling of the Regulator of Chromosome Condensation 1 (RCC1) Gene Family in Gossypium Hirsutum L. under Abiotic Stress and Hormone Treatments. International Journal of Molecular Sciences, 20(7), 1727. https://doi.org/10.3390/ijms20071727



