Proteomic Analysis of the Effect of Inorganic and Organic Chemicals on Silver Nanoparticles in Wheat
Abstract
:1. Introduction
2. Results
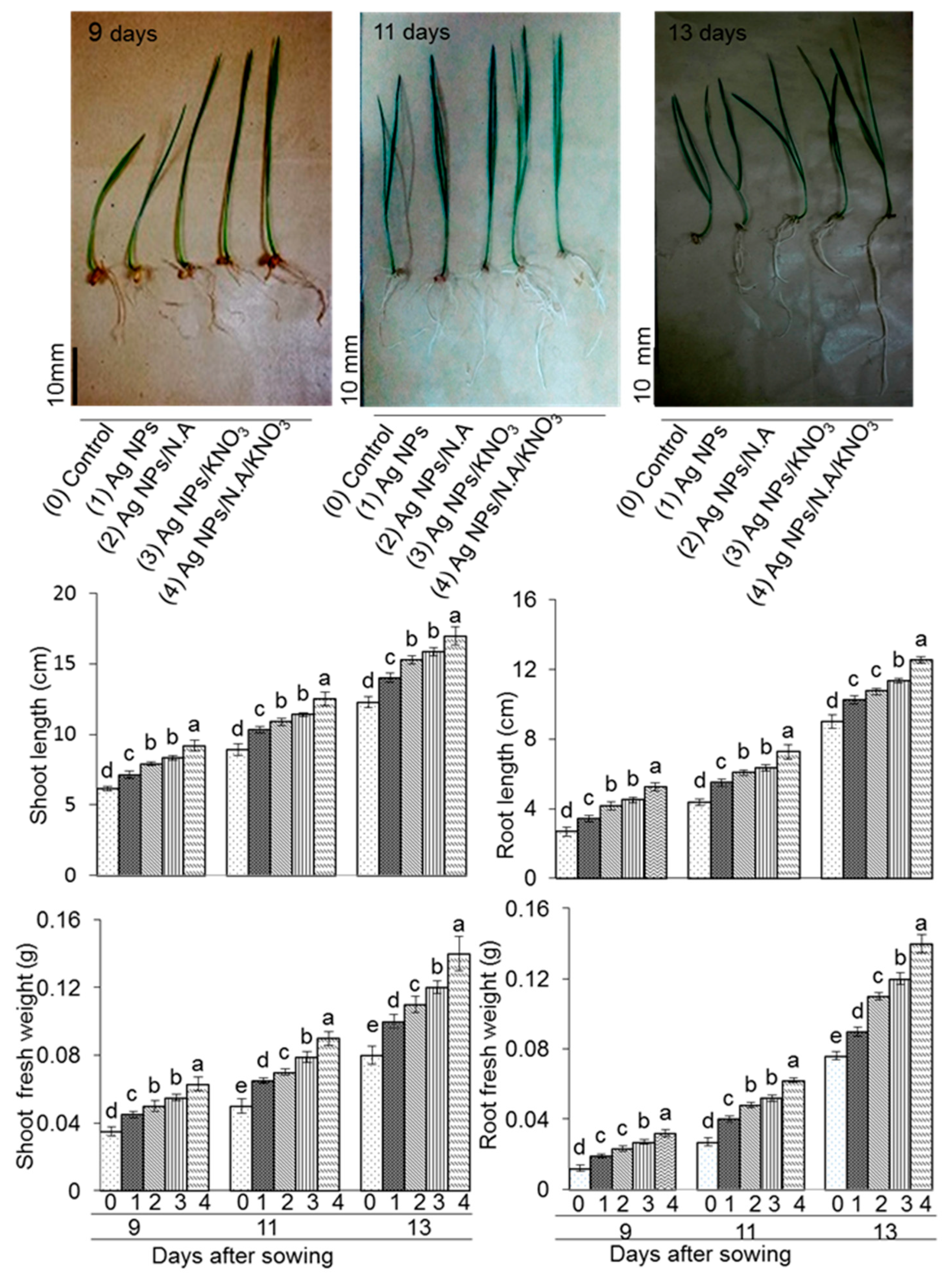
2.1. Growth Response of Wheat to Ag NPs Mixed with Organic and Inorganic Chemicals
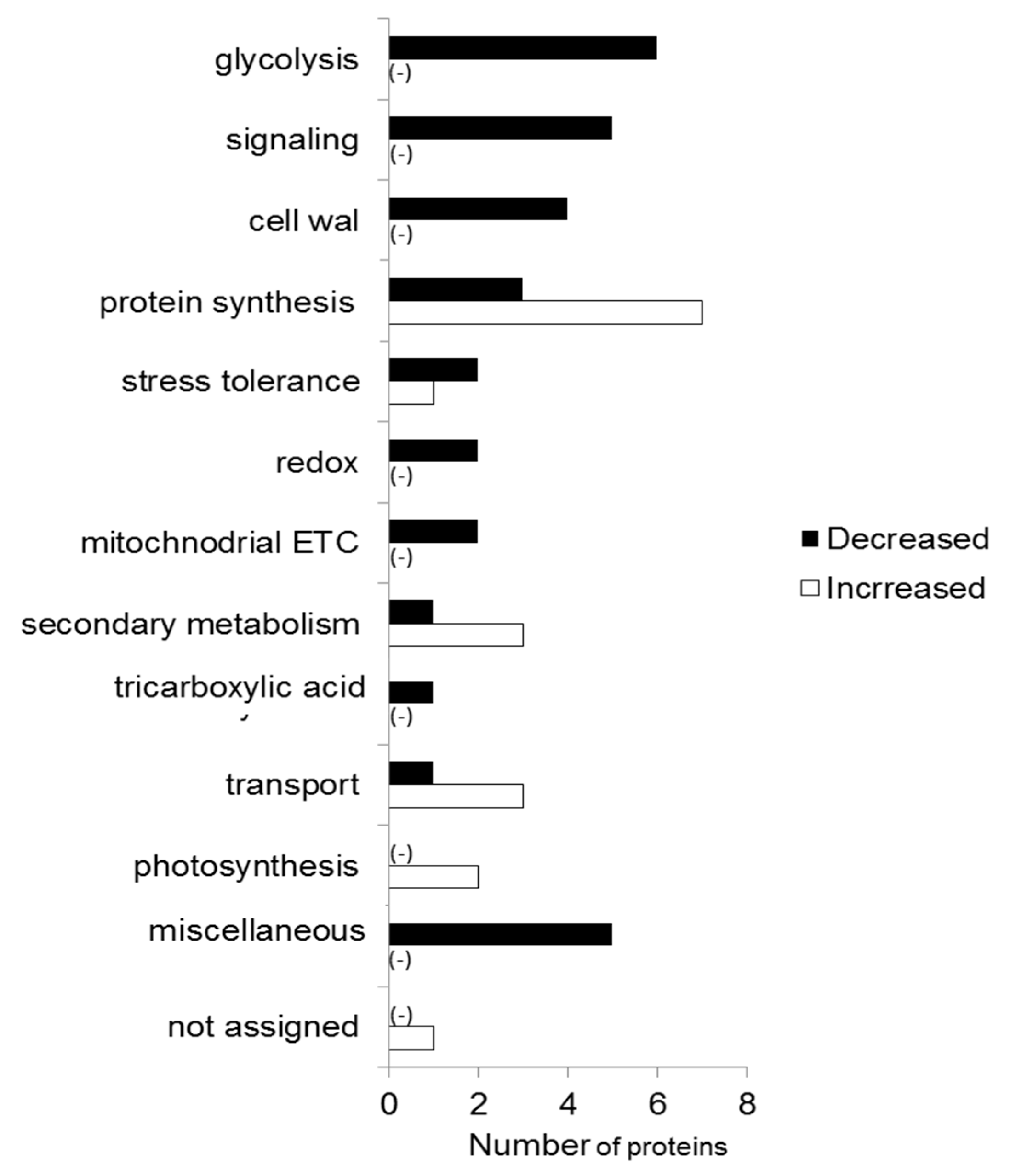
2.2. Proteomic Analysis
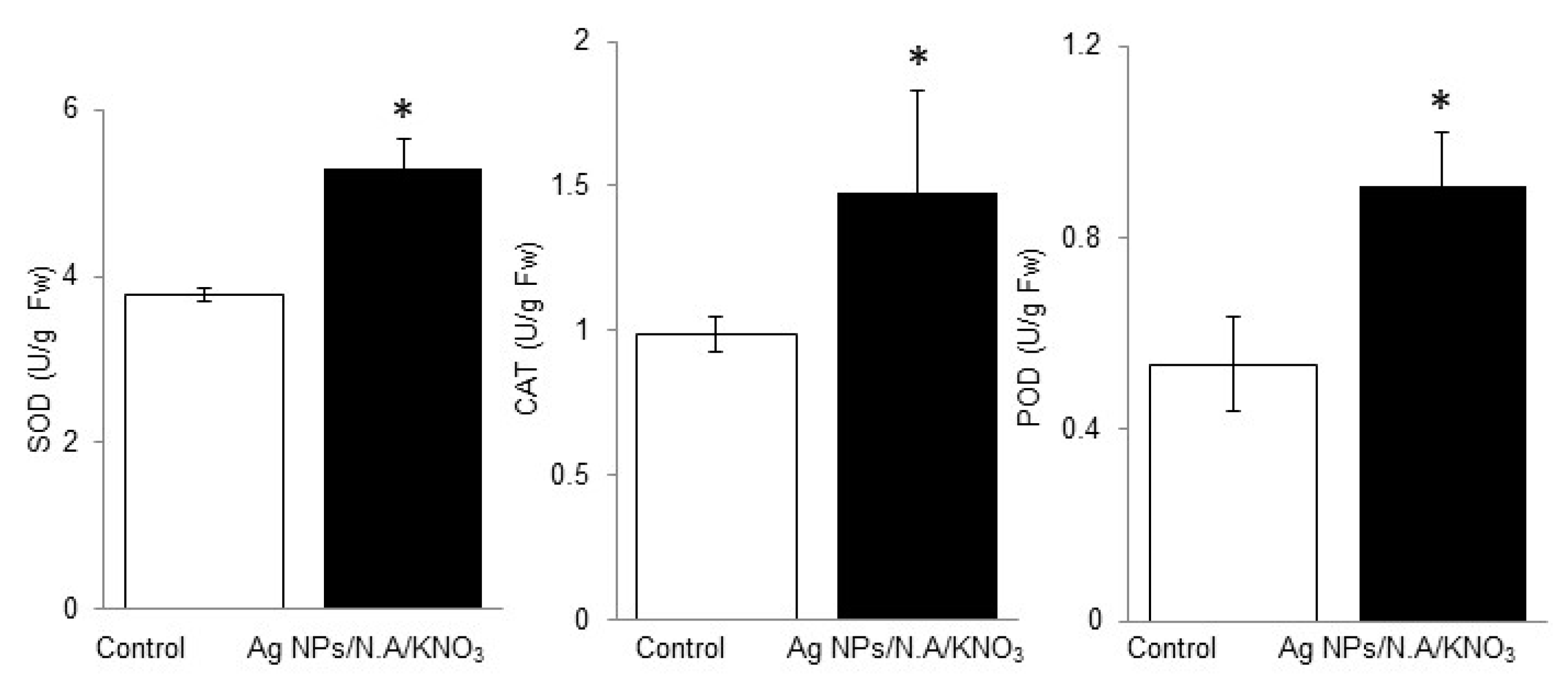
2.3. Effect of Chemo-Blended Ag NPs on Antioxidant Enzyme Activity Analysis of Wheat
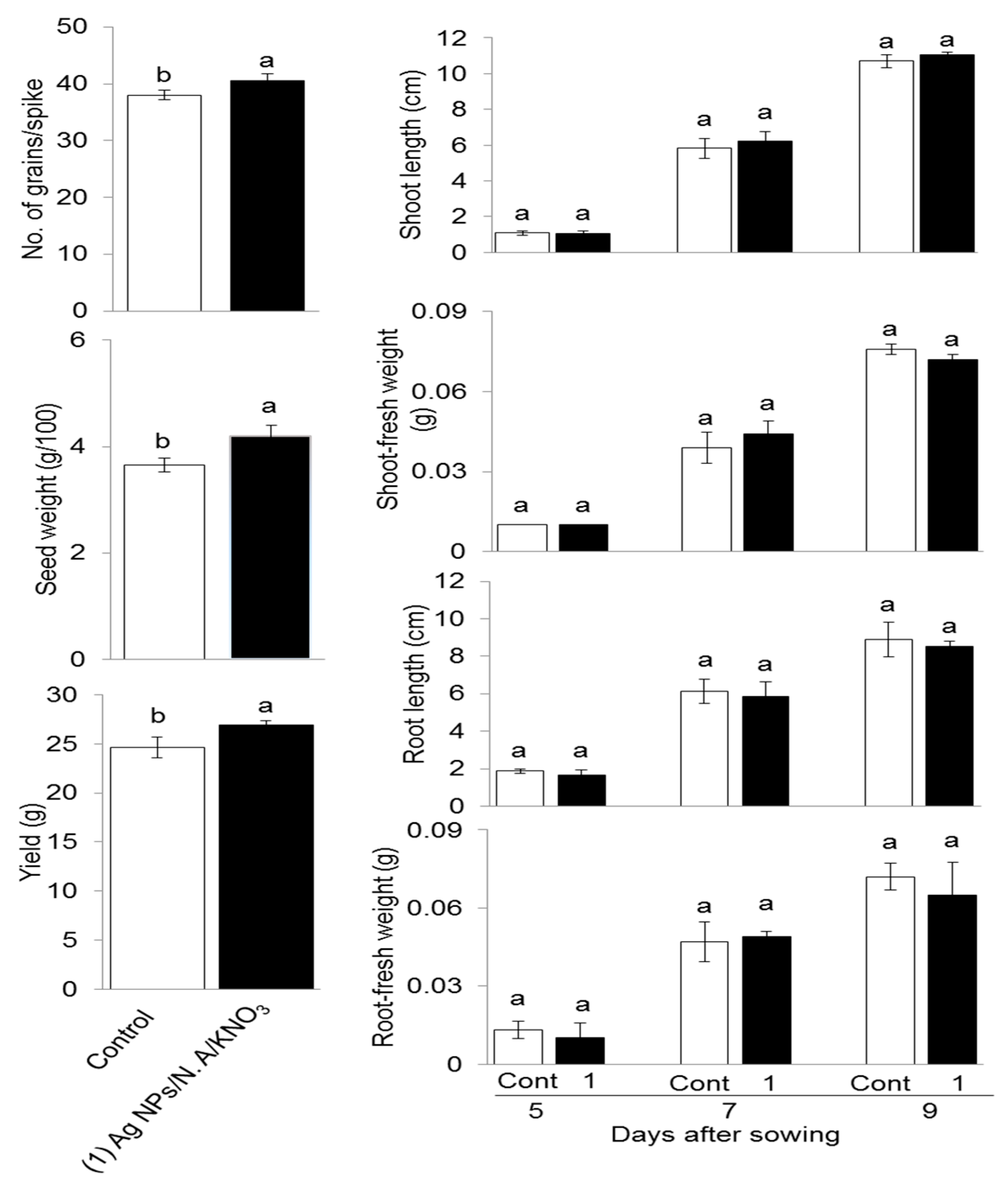
2.4. Effect of Chemo-Blended Ag NPs on Yield and Lifecycle of Wheat
3. Discussion
3.1. Effect of Chemo-Blended Ag NPs on Morphological Attributes of Wheat
3.2. Chemo-Blended Ag NPs Affect Protein Metabolism of Wheat
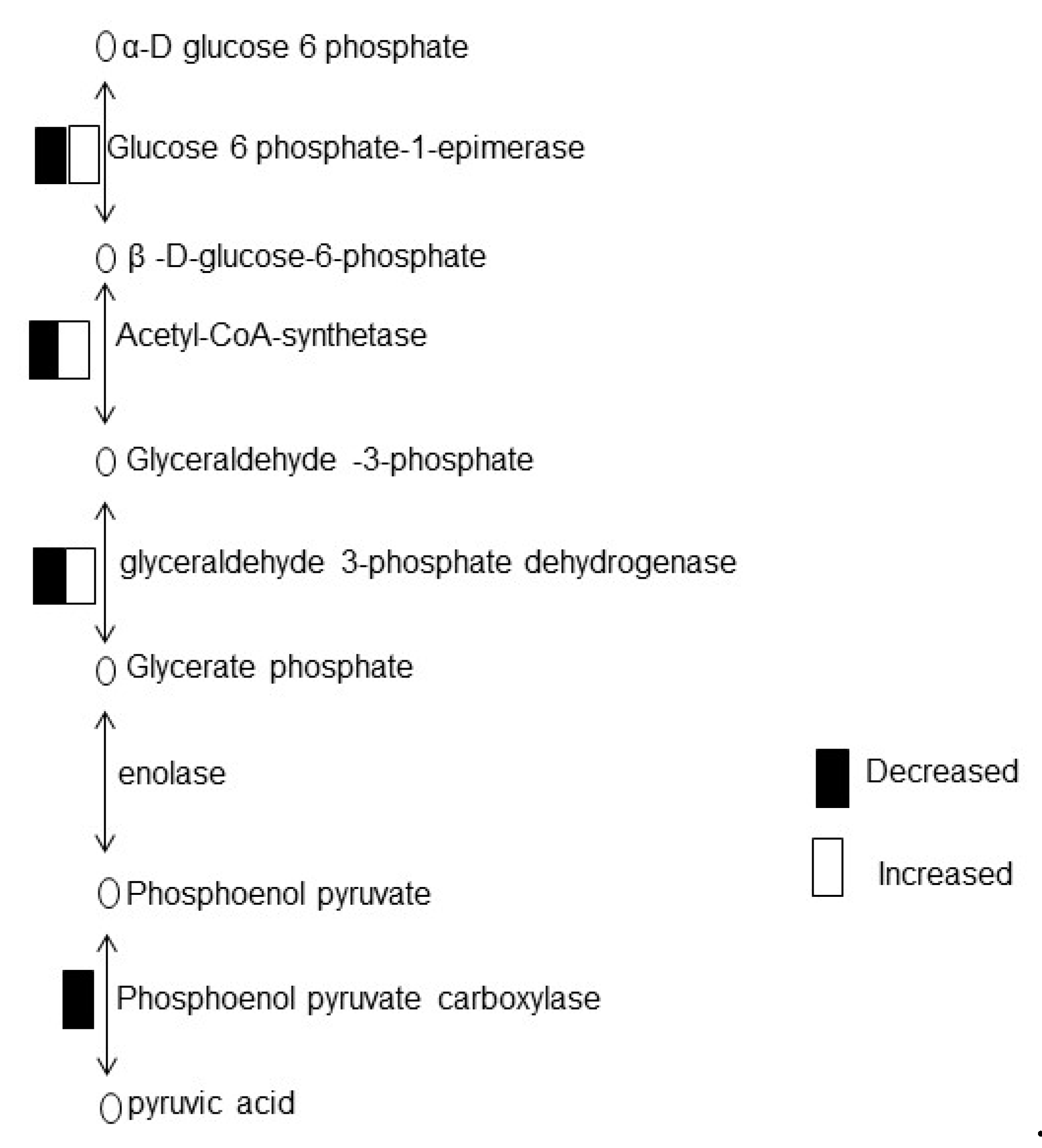
3.3. Chemo-Blended Ag NPs Affect Glycolysis of Wheat
3.4. Impact of Chemo-Blended Ag NPs on Scavenging Activity of SOD, CAT, and POD
3.5. Effect of Chemo-Blended Ag NPs on Yield and Growth of Next Generation
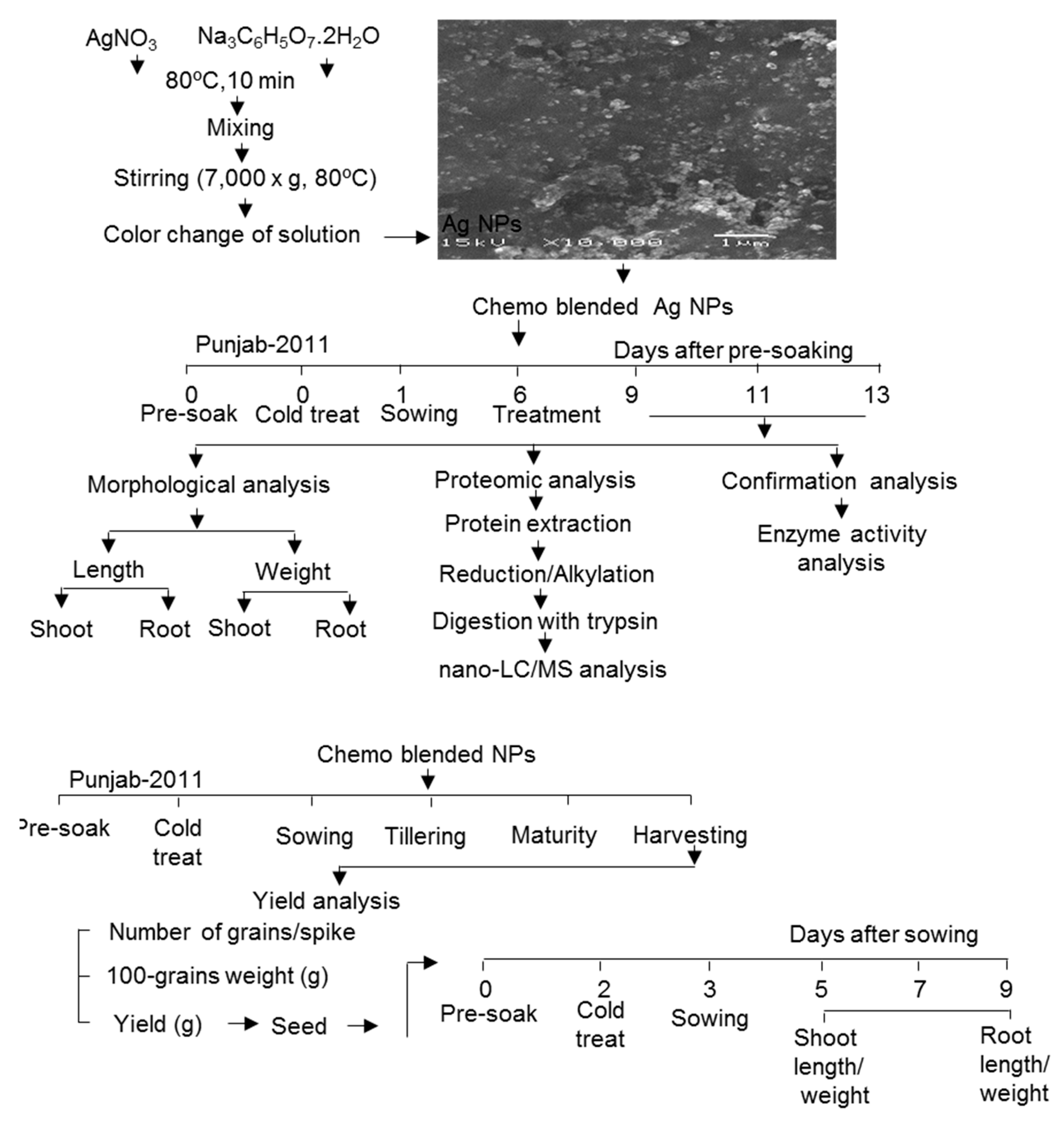
4. Materials and Methods
4.1. Preparation of Chemo-Blended Nanoparticles
4.2. Plant Material and Treatment
4.3. Protein Extraction
4.4. Protein Enrichment, Reduction, Alkylation and Digestion
4.5. Measurement of Protein and Peptide Concentrations
4.6. Protein Identification Using Nano LC-MS/MS
4.7. MS Data Analysis
4.8. Differential Analysis of Proteins Using MS Data
4.9. Functional Categorization
4.10. Analysis of Superoxide Dismutase, Catalase Activity, and Peroxidase in Response to Chemo-Blended Ag NPs
4.11. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| NPs | Nanoparticles |
| LC | Liquid Chromatography |
| ROS | Reactive oxygen species |
| MS | Mass Spectrometry |
| SOD | Superoxide dismutase |
| POD | Peroxidase |
| CAT | Catalase |
References
- Peterson, E.J.; Henery, T.B.; Zhao, J.; MacCuspie, R.I.; Kirshling, T.L.; Dobrovolskaia, M.A.; White, J.C. Identification and avoidance of potential artifacts and misinterpretations in nano material toxicity measurements. Environ. Sci. Technol. 2014, 48, 4226–4246. [Google Scholar] [CrossRef] [PubMed]
- Carlos, A.; Batista, S.; Larson, R.G.; Kotov, N.A. Nonadditivity of nanoparticle interactions. Science 2015, 350, 176–187. [Google Scholar]
- Nel, A.; Xia, T.; Madler, L.; Li, N. Toxic potential of materials at the novel. Science 2006, 311, 622–627. [Google Scholar] [CrossRef] [PubMed]
- Cox, A.; Venkatachalam, P.; Sohi, S.; Sharma, N. Silver and TiO2 NPs toxicity in plants: A review of current research. Plant Physiol. Biochem. 2016, 107, 147–163. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Lambi, E.; Zhao, F.J.; Kopihke, P.M. Nanotechnology: A new opportunity in plant sciences. Trends Plant Sci. 2016, 21, 699–712. [Google Scholar] [CrossRef]
- Rastogi, A.; Marek, Z.; Oksana, S.; Hazem, M.K.; He, X.; Sonia, M.; Marian, B. Impact of metal and metal oxide nanoparticles on plant: A critical review. Front. Chem. 2017, 5, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Cvjetko, P.; Milosic, A.; Domijao, A.M.; Vinkovic, V.I.; Tolic, S.; Pehrec, S.P.; Letofsky, P.I.; Tkalec, M.; Balen, B. Toxicity of silver ions and differently coated silver nanoparticles in Allium cepa roots. Ecotoxicol. Environ. Saf. 2017, 137, 18–28. [Google Scholar] [CrossRef]
- Sharma, P.; Bhatt, D.; Zaidi, M.G.; Saradhi, P.P.; Khanna, P.K.; Arora, S. Silver nanoparticle-mediated enhancement in growth and antioxidant status of Brassica juncea. Appl. Biochem. Biotechnol. 2012, 167, 2225–2233. [Google Scholar] [CrossRef]
- Stampoulis, D.; Sinha, S.K.; White, J.C. Assay- dependent phytotoxicity of nanoparticles on plants. Environ. Sci. Technol. 2009, 43, 9473–9479. [Google Scholar] [CrossRef]
- Krishnaraj, C.; Jagan, E.G.; Ramchandran, R.; Abirami, S.M.; Mohan, N.; Kalaichelvan, P.T. Effect of biologically synthesized silver nanoparticles on Baccopamonnieri (Linn.) wettst. Plant growth metabolism. J. Process Biochem. 2012, 47, 651–658. [Google Scholar] [CrossRef]
- Song, U.; Jun, H.; Waldman, B.; Roh, J.; Kim, Y.; Yi, J.; Lee, E.J. Functional analysis of nanoparticle toxicity: A comparative study of the effects of TiO2 and Ag on tomatoes (Lycopersiconescolentum). Ecotoxicol. Environ. Saf. 2013, 93, 60–67. [Google Scholar] [CrossRef] [PubMed]
- Sriram, T.; Pandidurai, V. In vitro growth analysis of Zea mays L. using Ag NPs. Int. J. Pharma Bio. Sci. 2017, 8, 30–37. [Google Scholar]
- Bello-Bello, J.J.; Chavez-Santoscoy, R.A.; Lecona-Guzman, C.A.; Bogdanchikova, N.; Salinase-Ruiz, J.; Gornez-Merino, F.C.; Pestryakov, A. Harmetic response of Ag NPs on in vitro multiplication of sugarcane using a temperorary immersion system. Dose Response 2017, 15, 1–9. [Google Scholar] [CrossRef]
- Syu, Y.Y.; Hung, J.H.; Chen, J.C.; Chuang, H.W. Impact of size and shape of silver NPs on Arabidopsis plant growth and gene expression. Plant Physiol. Biochem. 2014, 83, 57–64. [Google Scholar] [CrossRef] [PubMed]
- Sajjad, Y.; Jaskani, M.; Ashraf, M.Y.; Ahamd, R. Response of morphological and physiological growth attributes to foliar application of plant growth regulators in gladiolous ‘white prosperity’. Pak. J. Agric. Sci. 2014, 51, 123–129. [Google Scholar]
- Noctor, G.; Queval, G.; Gakiere, B. NADP synthesis and pyridine nucleotide cycling in plants and their potential importance in stress conditions. J. Exp. Bot. 2006, 57, 1603–1620. [Google Scholar] [CrossRef] [PubMed]
- Berglund, T.; Wallstrom, A.; Nguyen, T.V.; Laurell, C.; Ohlson, B.A. Nicotinamide, antioxidative and DNA hypomethylation effects in plant cells. Plant Physi. Biochem. 2017, 118, 551–560. [Google Scholar] [CrossRef]
- Wang, M.; Zheng, Q.; Shen, Q.; Guo, S. The critical role of potassium in plant stress response. Int. J. Mol. Sci. 2013, 14, 370–390. [Google Scholar] [CrossRef]
- Singh, H.; Singh, M.; Kang, J.S. Effect of potassium nitrate on yield and yield attributes of spring maize (Zea mays L.) under different dates of planting. Int. J. Curr. Microbiol. Appl. Sci. 2017, 6, 1581–1590. [Google Scholar] [CrossRef]
- Ahmad, P.; Abdel Latef, A.A.; Abd Allah, E.F.; Hashem, A.; Sarwat, M.; Anjum, N.A.; Gucel, S. Calcium and potassium supplementation enhanced growth, osmolyte secondary metabolite production and enzymatic antioxidant machinery in cadmium exposed chickpea. Front. Plant Sci. 2016, 27, 513. [Google Scholar] [CrossRef]
- Vanini, C.; Domingo, G.; Onelli, E.; DeMattia, F.; Brani, I.; Marsomi, M.; Bracale, M. Phytotoxic and genotoxic effects of Ag NPs exposure on germinating wheat seedlings. J. Plant Physiol. 2014, 171, 1142–1148. [Google Scholar] [CrossRef] [PubMed]
- Mustafa, G.; Skata, K.; Komatsu, S. Proteomic analysis of soybean roots exposed to varying sizes of silver nanoparticles on the effects of silver nanoparticles under flooding stress. J. Proteom. 2016, 148, 113–125. [Google Scholar] [CrossRef] [PubMed]
- Mirzajani, F.; Askari, H.; Hamzelou, S.; Schober, Y.; Rompp, A.; Ghassempour, A.; Spengler, B. Proteomic study of silver nanoparticles toxicity on Oryza sativa L. Ecotoxico. Environ. Saf. 2014, 108, 335–339. [Google Scholar] [CrossRef] [PubMed]
- Mahakhalm, W.; Srmah, A.K.; Meensiri, S.; Theerakulpisut, P. Nanopriming technology for enhancing germination and starch metabolism of aged rice seeds using phytosynthesized Ag NPs. Sci. Rep. 2017, 15, 1–21. [Google Scholar]
- Vishwakarma, K.; Shweta, U.N.; Singh, J.; Liu, S.; Singh, V.P.; Parasad, S.M.; Chauhan, D.K.; Sharma, S. Differential phototoxic impact of plant mediated Ag NPs and AgNO3 on brassica sp. Front. Plant Sci. 2017, 12, 1501. [Google Scholar] [CrossRef] [PubMed]
- Prasad, T.N.V.K.V.; Adam, S.; Visweswara, R.; Ravindra, R.B.; Gridhara, K.T. Size dependent effects of antifungal phytogenic silver NPs on germination, growth and biochemical parameters of rice, maize and peanut. IET Nanobiotechnol. 2017, 11, 277–285. [Google Scholar] [CrossRef] [PubMed]
- Liu, G.; Zhang, M.; Jin, Y.; Fan, X.; Xu, J.; Zhu, Y.; Fu, Z.; Pan, X.; Qian, H. The effects of low concentration of Ag NPs on wheat growth, seed quality and soil microbial communities. J. Water Air Soil Pollut. 2017, 228, 1–12. [Google Scholar] [CrossRef]
- Barabanov, P.V.; Gerasimov, A.V.; Bilnov, A.V.; Kravtsov, V.A. Influence of nanosilver on the efficiency of Pisumsativum crops germination. Ecotoxicol. Environ. Saf. 2017, 147, 715–719. [Google Scholar] [CrossRef]
- Tripathi, D.K.; Singh, S.; Singh, S.; Srivastava, P.K.; Singh, V.P.; Singh, S.; Parasad, S.M.; Singh, P.K.; Dubey, N.K.; Panday, A.C.; et al. Nitric oxide alleviates (AgNPs)- induced phytotoxicity in Pisumsativum seedlings. Plant Physiol. Biochem. 2017, 110, 167–177. [Google Scholar] [CrossRef]
- El-Tamsah, Y.S.; Joner, E.J. Impact of Fe and Ag NPs on seed germination and differences in bioavailability during exposure in aqueous suspension and soil. Environ. Toxicol. 2010, 27, 42–49. [Google Scholar] [CrossRef]
- Hojjat, S.S.; Hojjat, H. Effect of Ag NPs exposure on germination of lentil. Int. J. Farm. Allied Sci. 2016, 5, 248–252. [Google Scholar]
- Mustafa, G.; Skata, K.; Hossain, Z.; Komatsu, S. Proteomic study on the effects of silver nanoparticles on soybean under flooding stress. J. Proteom. 2015, 122, 100–118. [Google Scholar] [CrossRef] [PubMed]
- Ferryra, M.L.F.; Pezza, A.; Biarc, J.; Burlingame, A.L.; Cast, P. Plant L10 ribosomal proteins have different roles during development and translation under ultraviolet-B stress. Plant Phsysiol. 2010, 153, 1878–1894. [Google Scholar] [CrossRef] [PubMed]
- Komatsu, S.; Kuji, R.; Nanjo, Y.; Hiraga, S.; Furakuwa, K. Comprehensive analysis of endoplasmic reticulum-enriched fraction in root tips of soybean under flooding stress using proteomic techniques. J. Proteom. 2012, 77, 531–562. [Google Scholar] [CrossRef] [PubMed]
- Yasmeen, F.; Raja, N.I.; Mustafa, G.; Sakata, K.; Komatsu, S. Quantitative proteomic analysis of post flooding recovery in soybean roots exposed to Aluminum oxide NPs. J. Proteom. 2016, 143, 136–150. [Google Scholar] [CrossRef] [PubMed]
- Yasmeen, F.; Raja, N.I.; Razzaq, A.; Komatsu, S. Gel-free/label-free proteomic analysis of wheat shoot in stress tolerant varieties under iron nanoparticles exposure. Biochim. Biophys. Acta 2016, 1864, 1586–1598. [Google Scholar] [CrossRef] [PubMed]
- Vanini, C.; Domingo, G.; Onelli, E.; Prinsi, B.; Marsoni, M.; Espen, L.; Barcale, M. Morphological and proteomic responses of Eruca sativa exposed to AgNPs or AgNO3. PLoS ONE 2013, 8, 68752. [Google Scholar] [CrossRef]
- Plaxton, W.C. The organization and regulation of plant glycolysis. Annu. Rev. Plant Physiol. Plant Mol. Biol. 1996, 47, 185–214. [Google Scholar] [CrossRef]
- Banti, V.; Giuntoli, B.; Gonzoli, S.; Loreti, E.; Magneshchi, L.; Novi, G. Low oxygen response mechanism in green organism. Int. J. Mol. Sci. 2013, 14, 4734–4761. [Google Scholar] [CrossRef]
- Khan, M.; Jan, A.; Karibe, H.; Komatsu, S. Identification of phosphor proteins regulated by gibberellins in rice leaf sheath. Plant Mol. Biol. 2005, 58, 27–40. [Google Scholar] [CrossRef]
- Yasmeen, F.; Raja, N.I.; Ilyas, N.; Komatsu, S. Quantitative proteomic analysis of shoot in stress tolerant wheat varieties on copper nanoparticle exposure. Plant Mol. Biol. Rep. 2018, 36, 326–340. [Google Scholar] [CrossRef]
- Gorrini, C.; Harris, I.S.; Mak, T.W. Modulation of oxidative stress as an anticancer strategy. Nat. Rev. Drug Discov. 2013, 12, 931–947. [Google Scholar] [CrossRef] [PubMed]
- Miller, G.; Suzuki, N.; Ciftic-Yilmaz, S.; Mittler, R. Reactive oxygen species homeostasis and signaling during drought and salinity stress. Plant Cell Environ. 2010, 33, 453–467. [Google Scholar] [CrossRef] [PubMed]
- Du, W.; Tan, W.; Peralta-Videa, J.R.; Gardea-Torresdey, J.L.; Ji, R.; Guo, H. Interaction of metal oxide nanoparticles with higher terrestrial plants: Physiological and biochemical aspects. Plant Physiol. Biochem. 2017, 110, 210–225. [Google Scholar] [CrossRef] [PubMed]
- Verma, S.; Dubey, R. Lead toxicity induces lipid peroxidation and alters the activities of antioxidant enzymes in growing rice plants. Plant Sci. 2003, 164, 645–655. [Google Scholar] [CrossRef]
- Jiang, H.S.; Yin, L.Y.; Ren, N.N.; Zhao, S.T.; Li, Z.; Zhi, Y.; Shao, H.; Li, W.; Gontero, B. Silver nanoparticles induced accumulation of reactive oxygen species and alteration of antioxidant systems in the aquatic plant Spirodelapolyrhiza. Nanotoxicology 2017, 11, 1–42. [Google Scholar] [CrossRef] [PubMed]
- Ma, C.; Liu, H.; Guo, H.; Musante, C.; Coskun, S.H.; Nelson, B.C.; White, J.C.; Xing, B.; Dhankher, O.P. Defense mechanisms and nutrient displacement in Arabidopsis thaliana upon exposure to CeO2 and In2O3 nanoparticles. Environ. Sci. Nano 2016, 3, 1369–1379. [Google Scholar] [CrossRef]
- Rani, P.U.; Yasur, J.y.; Loke, K.S.; Dutta, D. Effect of synthetic and biosynthesized silver nanoparticles on growth, physiology and oxidative stress of water hyacinth: Eichhorniacrassipes (Mart) Solms. Acta Physiol. Plant 2016, 38, 58. [Google Scholar] [CrossRef]
- Huang, Z.; He, K.; Song, Z.; Zeng, G.; Chen, A.; Yuan, L.; Li, H.; Hu, L.; Guao, Z.; Chen, G. Antioxidative response of Phanerochaetechrysosporium against silver nanoparticle-induced toxicity and its potential mechanism. Chemosphere 2018, 211, 573–583. [Google Scholar] [CrossRef]
- Das, P.; Barua, S.; Sarkar, S.; Karak, N.; Bhattacharyya, P.; Raza, N.; Kim, K.H.; Bhattacharyya, S.S. Plant extract- mediated green SNPs: Efficacy as soil conditioner and plant growth promoter. J. Hazad. Mater. 2017, 15, 62–72. [Google Scholar]
- Yasmeen, F.; Raja, N.I.; Razzaq, A.; Komatsu, S. Proteomic and physiological analyses of wheat seeds exposed to copper and iron nanoparticles. Biochim. Biophys. Acta 2017, 1865, 28–42. [Google Scholar] [CrossRef]
- Wang, Q.; Ebbs, S.D.; Chen, Y.; Ma, X. Trans-generational impact of Cerium Oxide NPs on tomato plants. Metallomics 2013, 5, 753–759. [Google Scholar] [CrossRef] [PubMed]
- Zhao, L.; Sun, Y.; Hernandez-Viezcas, J.A.; Servin, A.D.; Hong, J.N.G.; Peralta-Videa, J.R.; Duarte-Gardea, M.; Gardea-Torresdey, J.L. Influence of CeO2 and ZnO nanoparticles on cucumber physiological markers and bioaccumulation of Ce and Zn: A life cycle study. J. Agric. Food Chem. 2014, 61, 11945–11951. [Google Scholar] [CrossRef]
- Hafeez, A.; Razzaq, A.; Mahmood, T.; Jhanzab, H.M. Potential of copper nanoparticles to increase growth and yield of wheat. J. Nanosci. Adv. Technol. 2015, 1, 6–11. [Google Scholar]
- Shams, G.; Ranjbar, M.; Amiri, A. Effect of Ag NPs on concentrations of silver heavy element and growth indexes in cucumber (Cucumissativus. L. negeen). J. Nanopart. Res. 2013, 15, 1630–1635. [Google Scholar] [CrossRef]
- Sheykhbaglou, R.; Sedghi, M.; Shishevan, M.T.; Sharifi, R.F. Effects of nano-iron oxide particles on agronomic traits of soybean. Not. Sci. Biol. 2010, 2, 112–113. [Google Scholar] [CrossRef]
- Feizi, H.; Pour, S.J.; Rad, K.H. Biological response of muskmelon to magnetic field and silver nanoparticles. Ann. Rev. Res. Biol. 2013, 3, 794–804. [Google Scholar]
- Venkatachalam, P.; Priyanka, N.; Manikandan, K.; Ganeshbabu, I.; Indiraarulselvi, P.; Geetha, N.; Muralikrishna, K.; Bhattacharya, R.C.; Tiwari, M.; Sharma, N.; et al. Enhanced plant growth promoting role of phycomolecules coated zinc oxide nanoparticles with P supplementation in cotton (Gossypium hirsutum L.). Plant Physiol. Biochem. 2017, 110, 118–127. [Google Scholar] [CrossRef] [PubMed]
- Zhu, H.; Han, J.; Xiao, J.Q.; Jin, Y. Uptake, translocation, accumulation of manufactured Iron oxide nanoparticles by pumpkin plants. J. Environ. Monit. 2008, 10, 713–717. [Google Scholar] [CrossRef]
- Ajirloo, A.R.; Shaaban, M.; Motlagh, Z.R. Effect of K nano-fertilizer and N bio-fertilizer on yield and yield components of tomato (Lycopersicon Esculentum L.). Int. J. Adv. Biol. Biom. Res. 2015, 3, 138–143. [Google Scholar]
- Razzaq, A.; Ammara, R.; Jhanzab, H.M.; Mahmood, T.; Hafeez, A.; Hussain, S. A novel nanomaterial to enhance growth and yield of wheat. J. Nanosci. Technol. 2016, 2, 55–58. [Google Scholar]
- Komatsu, S.; Han, C.; Nanjo, Y.; Altaf-Un-Nahar, M.; Wang, K.; He, D.; Yang, P. Label-free quantitative proteomic analysis of abscissic acid effect in early-stage soybean under flooding. J. Proteome Res. 2013, 12, 4769–4784. [Google Scholar] [CrossRef] [PubMed]
- Bradford, M.M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 1976, 72, 248–254. [Google Scholar] [CrossRef]
- Li, X.; Rehman, S.U.; Yamaguchi, H.; Hitachi, K.; Tsuchida, K.; Yamaguchi, T.; Sunohara, Y.; Matsumoto, H.; Komatsu, S. Proteomic analysis of the effect of plant-derived smokeon soybean during recovery from flooding stress. J. Proteom. 2018, 181, 238–248. [Google Scholar] [CrossRef] [PubMed]
- Tyanova, S.; Temu, T.; Siniteyn, P.; Carlson, A.; Hein, Y.; Gieger, T.; Mann, M.; Cox, J. The Perseus computational platform for comprehensive analysis of proteomics data. Nat. Methods 2016, 13, 731–740. [Google Scholar] [CrossRef] [PubMed]
- Usadel, B.; Nagel, A.; Thimm, O.; Redestig, H.; Blaesing, O.E.; Rofas, N.P.; Selbig, J.; Hannemann, J.; Piques, M.C.; Steinhauser, D.; et al. Extension of the visualization tool MapMan to allow statistical analysis of arrays, display of corresponding genes and comparison with known responses. Plant Physiol. 2005, 138, 1195–1204. [Google Scholar] [CrossRef] [PubMed]
- Usadel, B.; Poree, F.; Nagel, A.; Loshe, M.; Czedik-Eysenberg, A.; Sitt, M. Aguide to using MapMan to visualize and compare omics in plants: A case study in the crop species, maize. Plant Cell Environ. 2009, 32, 1211–1229. [Google Scholar] [CrossRef] [PubMed]
- Kanehisa, M.; Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 2000, 28, 27–30. [Google Scholar] [CrossRef] [PubMed]
- Beauchamp, C.; Fridovich, I. Superoxide dismutase: Improved assays and an assay applicable to acrylamide gels. Anal. Biochem. 1971, 44, 276–287. [Google Scholar] [CrossRef]
- Aebi, H.E. Catalase. In Methods of Enzymatic Analysis; Bergmeyer, H.U., Ed.; VerlagChemie: Weinhem, Germany, 1983; pp. 273–286. [Google Scholar]
- Li, S.; Yan, T.; Yang, J.Q.; Oberley, T.D.; Oberley, W. The role of cellular glutathione peroxidase redox regulation in the suppression of tumor cell growth by manganese superoxide dismutase. Cancer Res. 2000, 60, 3927–3939. [Google Scholar] [PubMed]
| No | Accession | Description | Difference | Functional Category | Biological Process | Cellular Component | Molecular Function |
|---|---|---|---|---|---|---|---|
| 1 | W5AYF4 | Putative SNAP receptor protein | 3.56 | Transport | Cell organization and biogenesis | membrane | protein binding |
| 2 | A0A1D5WZM5 | ER membrane protein complex | 3.2 | Protein | Transport | membrane | metal ion binding |
| 3 | A0A1D6B0Y2 | At4g14100-like | 2.91 | Transport | Not assigned | membrane | catalytic activity |
| 4 | W5DZQ3 | Ribosomal protein S1 | 2.85 | Protein | Response to stimulus | chloroplast | RNA binding |
| 5 | Q5G1T9 | Gamma-glutamylcysteinesynthetase | 2.57 | Protein | Metabolic process | Chloroplast, cytosol | catalytic activity |
| 6 | W5EA17 | D-ribose 5-phosphate | 2.41 | secondary metabolism | Metabolic process | cytosol | catalytic activity |
| 7 | A0A1D5UQX6 | Unknown | 2.27 | not assigned | Not assigned | not assigned | not assigned |
| 8 | W5DL10 | Glutamate--tRNA ligase | 1.96 | Protein | Metabolic process | cytoplasm | catalytic activity |
| 9 | A0A1D5YQ15 | Ferritin | 1.86 | photosynthesis | Cellular homeostasis | cytosol | catalytic activity |
| 10 | W5APX0 | GrpE protein homolog | 1.81 | Transport | Metabolic process | mitochondrion | enzyme regulator activity |
| 11 | A0A1D6AQL7 | PPIasecyclophilin-type | 1.77 | Stress | Metabolic process | cytosol | catalytic activity; protein binding |
| 12 | A0A1D5Y2E6 | Reverse transcriptase | 1.77 | Protein | Metabolic process | Cytoplasm | catalytic activity |
| 13 | A0A1D5SUT9 | Peptidase A1 | 1.76 | Protein | Metabolic process | Membrane | catalytic activity |
| 14 | O21432 | Ribosomal protein S2 | 1.65 | Protein | Metabolic process | mitochondrion | structural molecule activity |
| 15 | A0A1D6AKZ2 | ATP-dependent Clp protease proteolytic subunit | 1.16 | photosynthesis | Metabolic process | chloroplast | catalytic activity |
| 16 | A0A1D5V5A5 | Bifunctional inhibitor/plant lipid transfer | 1.12 | secondary metabolism | Transport | Membrane | catalytic activity; metal ion binding |
| 17 | A0A1D5YXC6 | Plant lipid transfer protein | 1.01 | secondary metabolism | Metabolic process | Membrane | catalytic activity |
| 18 | W5DX10 | Allene oxide synthase-lipoxygenase | 0.89 | Stress | Metabolic process | chloroplast | catalytic activity; metal ion binding |
| 19 | A0A1D5YZ95 | Synaptotagmin-like mitochondrial lipid-binding proteins | 0.88 | Transport | Metabolic process | Membrane | metal ion binding |
| 20 | A0A1D6DJK9 | Tyrosine--tRNA ligase | 0.86 | Protein | Metabolic process | Cytoplasm | catalytic activity |
| 21 | A0A1D6RMY5 | Short-chain dehydrogenase/reductase2 | 0.74 | Stress | Metabolic process | cytosol | catalytic activity |
| 22 | W5QKZ0 | Chalcone-flavonone isomerase | 0.7 | secondary Metabolism | Metabolic process | not assigned | catalytic activity |
| 23 | A0A1D5SIK2 | NAD(P)H-quinone oxidoreductase subunit I, | 0.7 | ETC | metabolic process | membrane | catalytic activity; metal ion binding |
| 24 | A0A1D5S4W8 | glyeraldehyde dehydrogenase | 0.69 | Glycolysis | metabolic process | membrane | catalytic activity |
| 25 | W5AV30 | Acetyl-CoA synthetase | 0.66 | Glycolysis | not assigned | membrane | protein binding |
| 26 | Q06I94 | Fasciclin-like protein FLA12 | 0.59 | Cell wall | response to stimulus | membrane | not assigned |
| 27 | A0A1D5UQL1 | Glucose-6-phosphate 1-epimerase | 0.57 | Glycolysis | metabolic process | chloroplast | catalytic activity |
| 28 | A0A1D6A8Y7]\ | 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase | 0.53 | Primary metabolism | metabolic process | cytoplasm | catalytic activity |
| 29 | A0A1D6RZJ3 | N-acetyltransferase | 0.4 | Signaling | metabolic process | not assigned | catalytic activity |
| No | Accession | Description | Difference | Functional Category | Biological Process | Cellular Component | Molecular Function |
|---|---|---|---|---|---|---|---|
| 1 | A0A1D6C3J5 | Copper transport protein | −0.65 | Redox | Transport | membrane | metal ion binding |
| 2 | A0A1D6S991 | Isocitrate dehydrogenase | −0.66 | TCA | Metabolic process | Mitochondrion | catalytic activity |
| 3 | A0A1D5UMA5 | 3-Oxoacyl- synthase III | −0.71 | lipid metabolism | Metabolic process | Chloroplast | catalytic activity |
| 4 | W5BFA5 | glyceraldehyde-3-phosphate dehydrogenase | −0.72 | Glycolysis | Metabolic process | Cytosol; Golgi | catalytic activity |
| 5 | A0A1D5V012 | NADH dehydrogenase ubiquinone Fe-S protein 4 | −0.74 | Redox | Metabolic process | mitochondrion | catalytic activity |
| 6 | A0A1D5U7K0 | F-box associated interaction domain | −0.89 | Signaling | Regulation of biological process | cytoplasm | protein binding |
| 7 | A0A1D5VAF5 | glycine-tyrosine-phenylalanine | −0.94 | Stress | Metabolic process | mitochondrion | protein binding |
| 8 | A0A1D5XH81 | carboxy peptidase | −0.97 | Glycolysis | Metabolic process | chloroplast | catalytic activity |
| 9 | A0A1D6AAU8 | acetyltransferase-superfamily | −1.11 | Signaling | Metabolic process | not assigned | catalytic activity |
| 10 | A0A1D6AVB7 | Ubiquitin-associated domain | −1.17 | Protein | Response to stimulus | cytosol | protein binding |
| 11 | A0A1D6D1I9 | Glutaredoxin | −1.18 | ETC | Cellular homeostasis | mitochondrion | catalytic activity |
| 12 | A0A1D6AQL9 | Rho termination factor, N-terminal domain superfamily | −1.26 | primary metabolism | Metabolic process | mitochondrion | catalytic activity |
| 13 | A0A1D5ZVW9 | La-type RNA-binding | −1.63 | RNA | Metabolic process | cytosol | RNA binding |
| 14 | A0A096ULK3 | Small GTPase superfamily | −1.76 | Signaling | Regulation of biological process | cytosol | catalytic activity |
| 15 | A0A1D6RV75 | peptidyl-prolyl cis-trans isomerase | −1.92 | Protein | Metabolic process | mitochondrion; | catalytic activity |
| 16 | W5E0G5 | Proteasome component | −2.34 | Protein | Metabolic process | cytosol | protein binding |
| 17 | A0A1D6AAV9 | Carbon-nitrogen hydrolase | −2.52 | Signaling | Metabolic process | not assigned | catalytic activity |
| 18 | A0A1D5TDD8 | Plant invertase/pectin methylesterase inhibitor | −2.9 | cell wall | Regulation of biological process | membrane | enzyme regulator activity |
| 19 | W5E5N6 | LysM Domain (Peptidoglycan binding) | −2.9 | cell wall | Response to stimulus | membrane | protein binding |
| 20 | A0A1D6DBZ9 | Polygalacturonase | −3.58 | cell wall | Cell organization and biogenesis | extracellular | catalytic activity |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jhanzab, H.M.; Razzaq, A.; Bibi, Y.; Yasmeen, F.; Yamaguchi, H.; Hitachi, K.; Tsuchida, K.; Komatsu, S. Proteomic Analysis of the Effect of Inorganic and Organic Chemicals on Silver Nanoparticles in Wheat. Int. J. Mol. Sci. 2019, 20, 825. https://doi.org/10.3390/ijms20040825
Jhanzab HM, Razzaq A, Bibi Y, Yasmeen F, Yamaguchi H, Hitachi K, Tsuchida K, Komatsu S. Proteomic Analysis of the Effect of Inorganic and Organic Chemicals on Silver Nanoparticles in Wheat. International Journal of Molecular Sciences. 2019; 20(4):825. https://doi.org/10.3390/ijms20040825
Chicago/Turabian StyleJhanzab, Hafiz Muhammad, Abdul Razzaq, Yamin Bibi, Farhat Yasmeen, Hisateru Yamaguchi, Keisuke Hitachi, Kunihiro Tsuchida, and Setsuko Komatsu. 2019. "Proteomic Analysis of the Effect of Inorganic and Organic Chemicals on Silver Nanoparticles in Wheat" International Journal of Molecular Sciences 20, no. 4: 825. https://doi.org/10.3390/ijms20040825
APA StyleJhanzab, H. M., Razzaq, A., Bibi, Y., Yasmeen, F., Yamaguchi, H., Hitachi, K., Tsuchida, K., & Komatsu, S. (2019). Proteomic Analysis of the Effect of Inorganic and Organic Chemicals on Silver Nanoparticles in Wheat. International Journal of Molecular Sciences, 20(4), 825. https://doi.org/10.3390/ijms20040825







