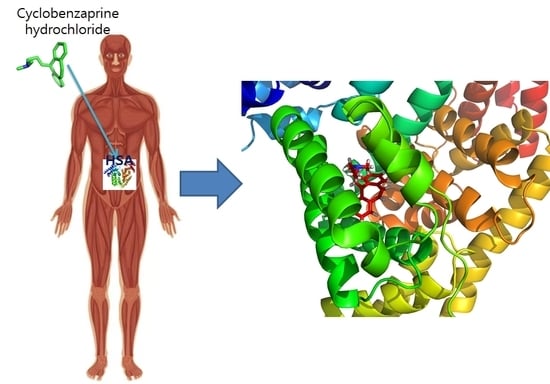
Multi-Spectroscopic Characterization of Human Serum Albumin Binding with Cyclobenzaprine Hydrochloride: Insights from Biophysical and In Silico Approaches
Abstract
1. Introduction
2. Results and Discussion
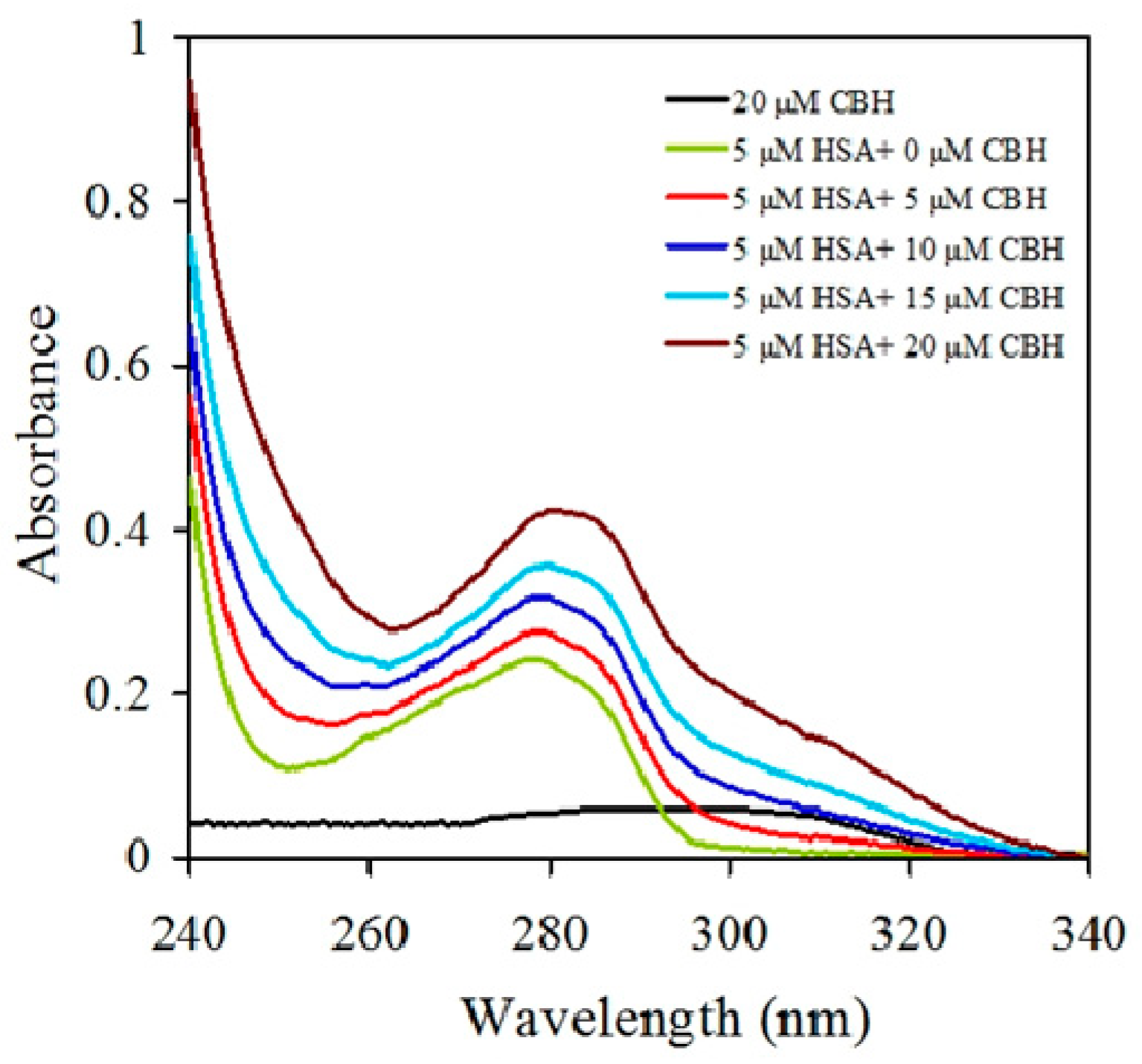
2.1. UV-Visible Spectroscopy of HSA-CBH Complex
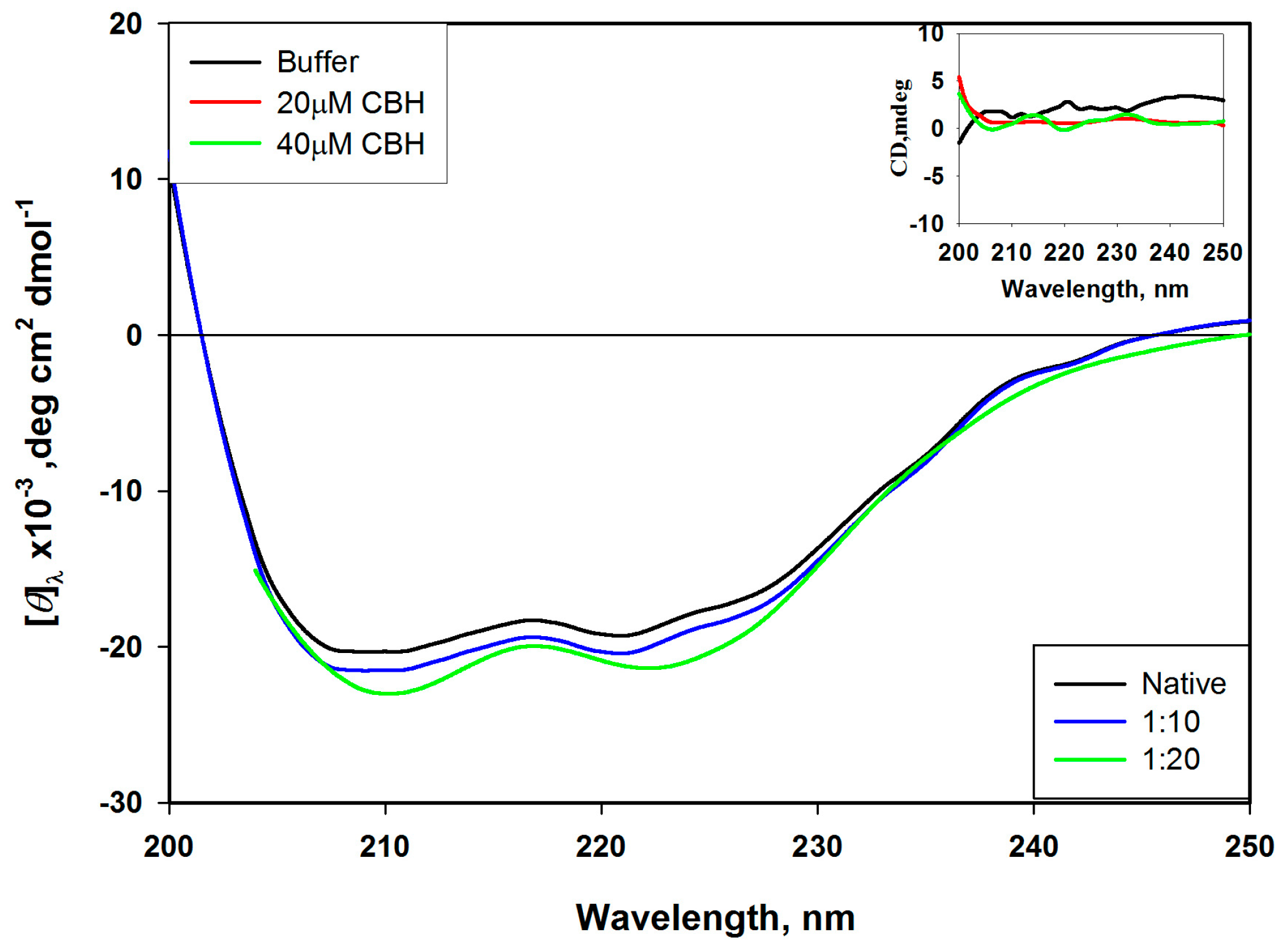
2.2. Alterations in Secondary Structure of HSA
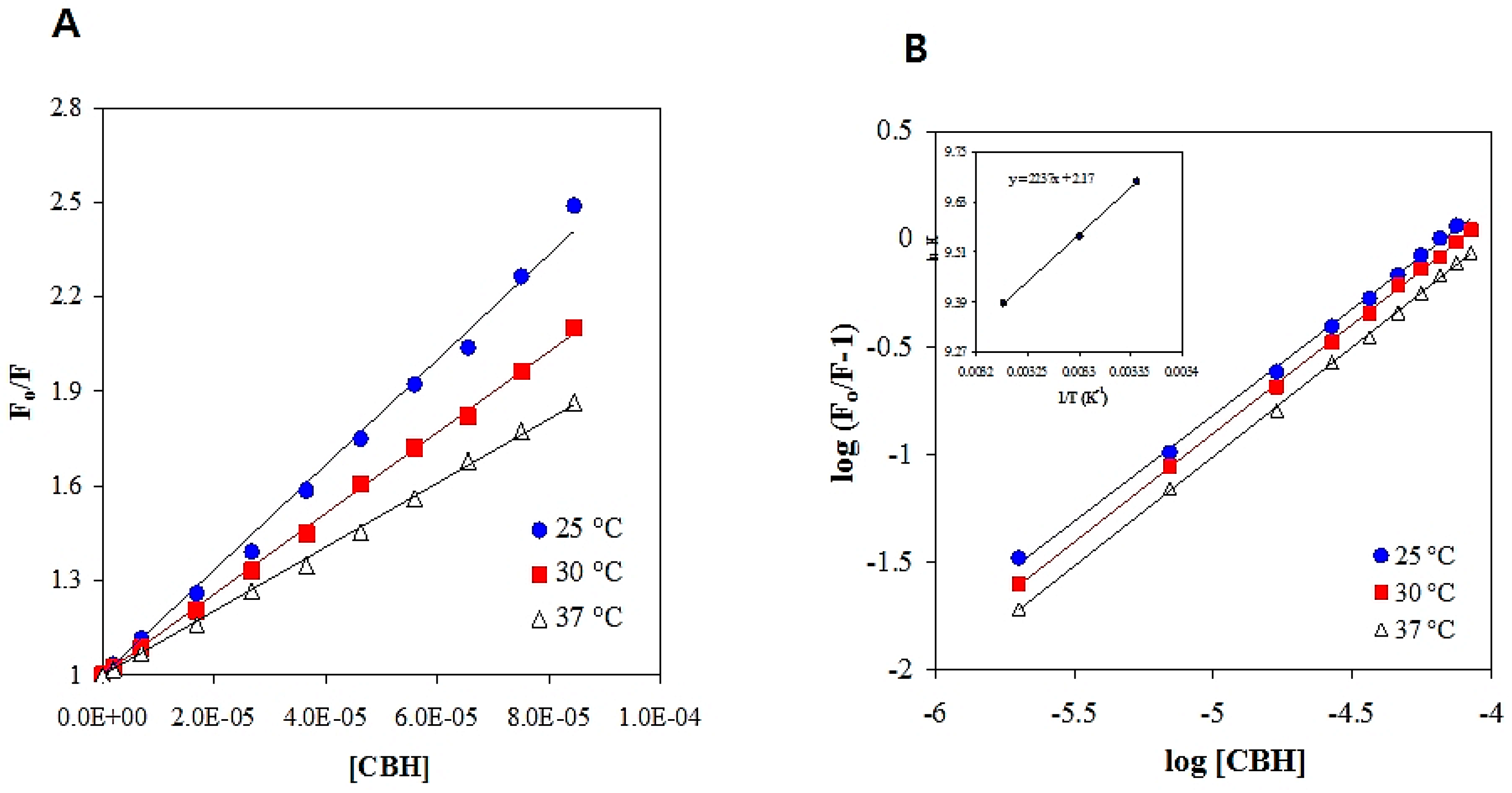
2.3. Fluorescence Quenching Analysis of HSA in the Presence of CBH
2.4. Thermodynamic Parameters and Forces Involved in HSA-CBH Binding
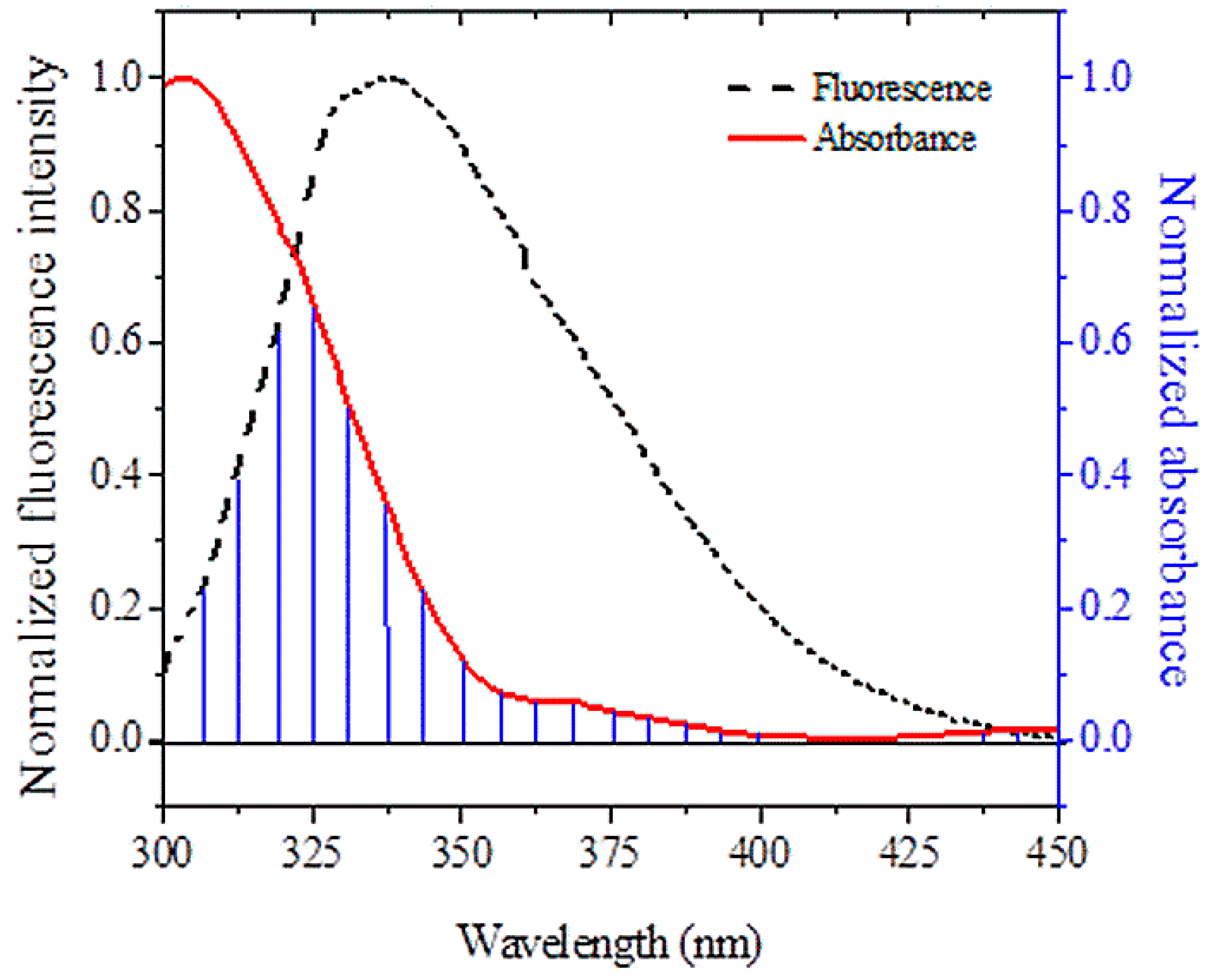
2.5. Energy Transfer Efficiency and HSA and Binding Distance
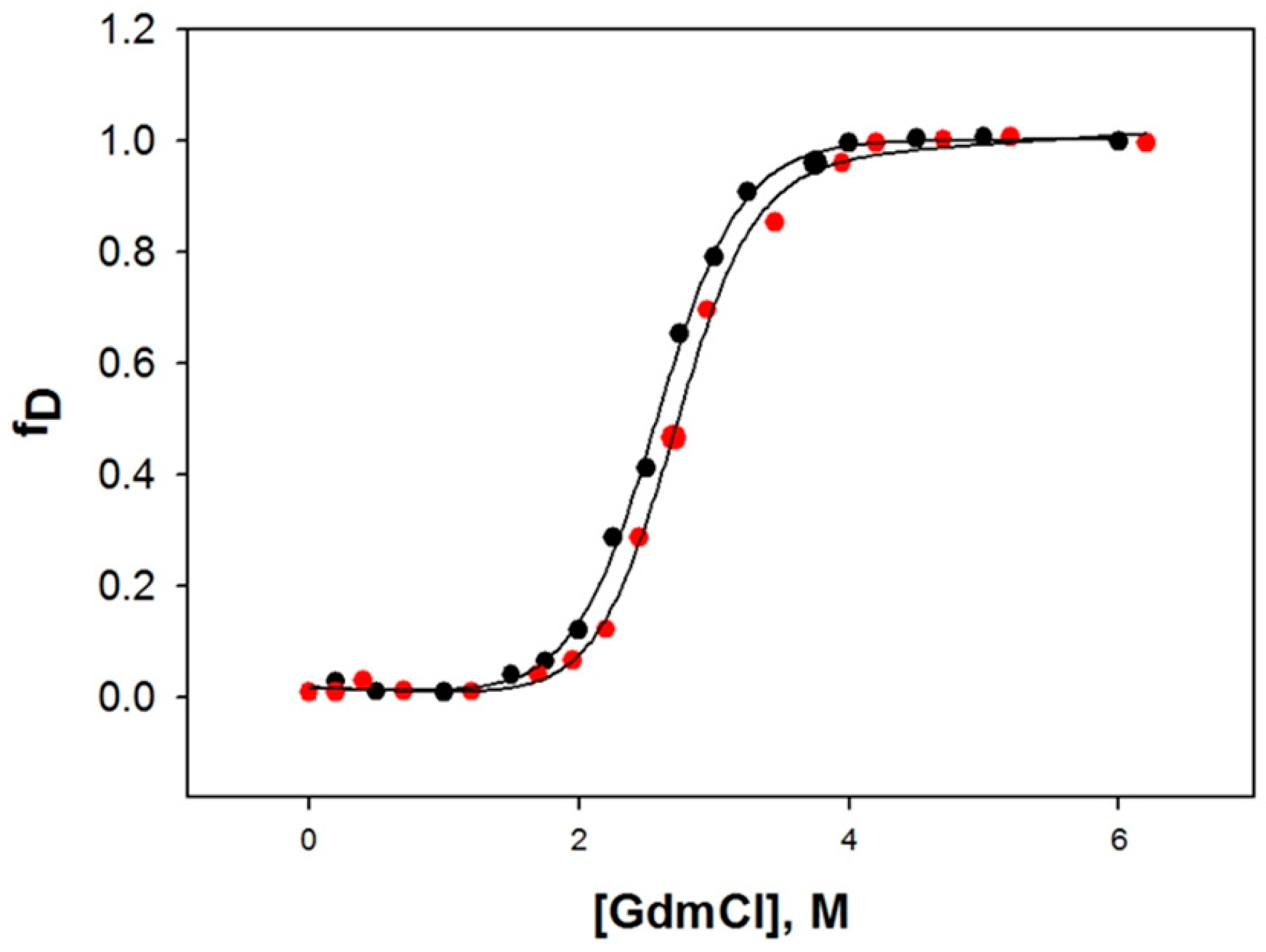
2.6. Denaturation of HSA with Guanidine Hydrochloride (GdmCl)
2.7. Enzyme Activity
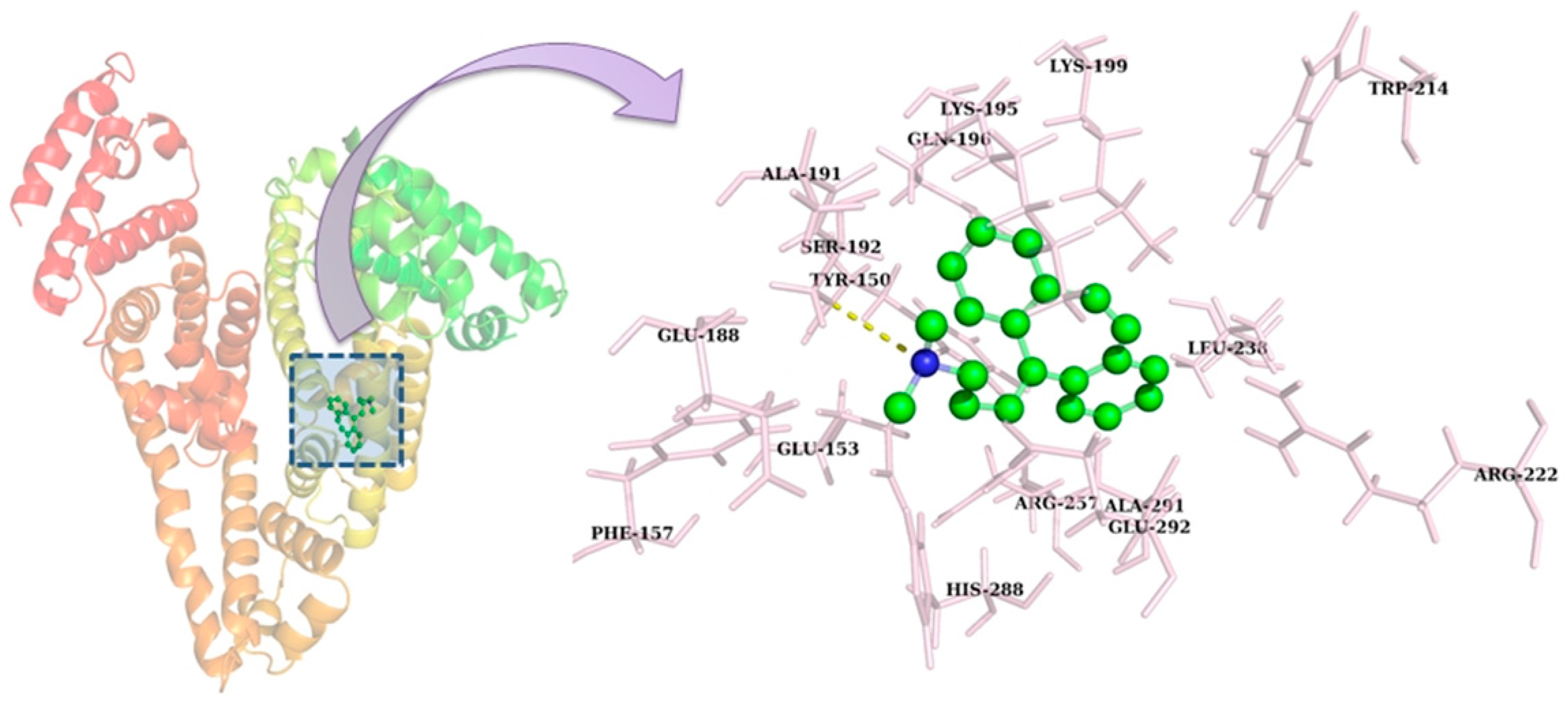
2.8. Molecular Docking
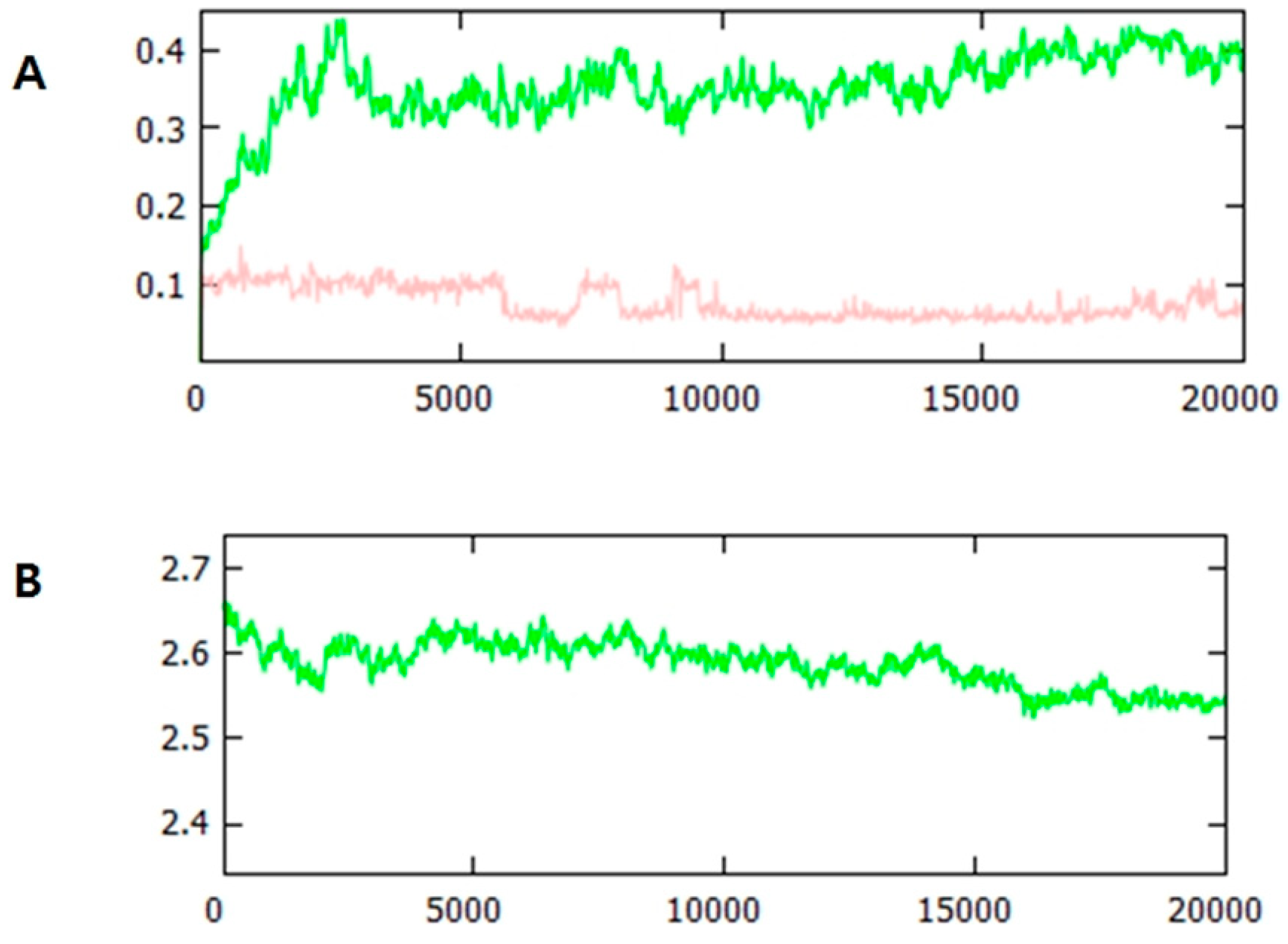
2.9. Molecular Dynamics Simulation
3. Materials and Methods
3.1. Reagents
3.2. UV-Visible Absorption Spectra Measurements
3.3. Fluorescence Quenching and Data Analysis
3.4. Thermodynamic Analysis of the Binding Process
3.5. Far-UV Circular Dichroism (CD) Spectropolarimetry for Structural Change Determination
3.6. Guanidine Hydrochloride-Induced Unfolding Studies in the Absence and Presence of CBH
3.7. Esterase-Like Activity of HSA
3.8. Molecular Docking
3.9. Molecular Dynamics
4. Conclusions
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| CD | Circular dichroism |
| CBH | Cyclobenzaprine hydrochloride |
| ∆H | Enthalpy |
| HSA | Human serum albumin |
| Km | Michaelis-Menten constant |
| λmax | Wavelength maxima |
| MRE | mean residue ellipticity |
| Vmax | maximum velocity |
References
- Nedergaard, A.; Karsdal, M.A.; Sun, S.; Henriksen, K. Serological muscle loss biomarkers: An overview of current concepts and future possibilities. J. Cachexia Sarcopenia Muscle 2013, 4, 1–17. [Google Scholar] [CrossRef]
- Le Verche, V.; Sunshine, S.; Hammers, D.; Sweeney, H.; Paushkin, S. Skeletal Muscle in Spinal Muscular Atrophy as an Opportunity for Therapeutic Intervention. In Spinal Muscular Atrophy; Elsevier: New York, NY, USA, 2017; pp. 341–356. [Google Scholar]
- Ahmad, K.; Lee, E.J.; Moon, J.S.; Park, S.Y.; Choi, I. Multifaceted Interweaving Between Extracellular Matrix, Insulin Resistance, and Skeletal Muscle. Cells 2018, 7, 148. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, H.; Hasegawa, Y.; Ono, H. Cyclobenzaprine, a centrally acting muscle relaxant, acts on descending serotonergic systems. Eur. J. Pharm. 1996, 311, 29–35. [Google Scholar] [CrossRef]
- Braschi, E.; Garrison, S.; Allan, G.M. Cyclobenzaprine for acute back pain. Can. Fam. Phys. 2015, 61, 1074. [Google Scholar]
- Bulbena-Cabre, A.; Dunn, N.R.; Swift, R.G. Cyclobenzaprine-Induced Hallucinosis. Prim. Care Companion CNS Disord. 2015, 17. [Google Scholar] [CrossRef] [PubMed]
- Lee, P.; Wu, X. Review: Modifications of human serum albumin and their binding effect. Curr. Pharm. Des. 2015, 21, 1862–1865. [Google Scholar] [CrossRef] [PubMed]
- Rabbani, G.; Baig, M.H.; Lee, E.J.; Cho, W.K.; Ma, J.Y.; Choi, I. Biophysical Study on the Interaction between Eperisone Hydrochloride and Human Serum Albumin Using Spectroscopic, Calorimetric, and Molecular Docking Analyses. Mol. Pharm. 2017, 14, 1656–1665. [Google Scholar] [CrossRef]
- Evans, T.W. Review article: Albumin as a drug–biological effects of albumin unrelated to oncotic pressure. Aliment. Pharm. 2002, 16 (Suppl. 5), 6–11. [Google Scholar] [CrossRef]
- Rabbani, G.; Lee, E.J.; Ahmad, K.; Baig, M.H.; Choi, I. Binding of Tolperisone Hydrochloride with Human Serum Albumin: Effects on the Conformation, Thermodynamics, and Activity of HSA. Mol. Pharm. 2018, 15, 1445–1456. [Google Scholar] [CrossRef]
- Sugio, S.; Kashima, A.; Mochizuki, S.; Noda, M.; Kobayashi, K. Crystal structure of human serum albumin at 2.5 A resolution. Protein Eng. 1999, 12, 439–446. [Google Scholar] [CrossRef]
- Wang, Y.; Wang, S.; Huang, M. Structure and enzymatic activities of human serum albumin. Curr. Pharm. Des. 2015, 21, 1831–1836. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Chen, X. Simple bioconjugate chemistry serves great clinical advances: Albumin as a versatile platform for diagnosis and precision therapy. Chem. Soc. Rev. 2016, 45, 1432–1456. [Google Scholar] [CrossRef] [PubMed]
- Otagiri, M. Study on binding of drug to serum protein. Yakugaku Zasshi 2009, 129, 413–425. [Google Scholar] [CrossRef] [PubMed]
- Meti, M.D.; Nandibewoor, S.T.; Joshi, S.D.; More, U.A.; Chimatadar, S.A. Multi-spectroscopic investigation of the binding interaction of fosfomycin with bovine serum albumin. J. Pharm. Anal. 2015, 5, 249–255. [Google Scholar] [CrossRef]
- Buddanavar, A.T.; Nandibewoor, S.T. Multi-spectroscopic characterization of bovine serum albumin upon interaction with atomoxetine. J. Pharm. Anal. 2017, 7, 148–155. [Google Scholar] [CrossRef] [PubMed]
- Naik, P.N.; Nandibewoor, S.T.; Chimatadar, S.A. Non-covalent binding analysis of sulfamethoxazole to human serum albumin: Fluorescence spectroscopy, UV-vis, FT-IR, voltammetric and molecular modeling. J. Pharm. Anal. 2015, 5, 143–152. [Google Scholar] [CrossRef] [PubMed]
- Rabbani, G.; Baig, M.H.; Jan, A.T.; Ju Lee, E.; Khan, M.V.; Zaman, M.; Farouk, A.E.; Khan, R.H.; Choi, I. Binding of erucic acid with human serum albumin using a spectroscopic and molecular docking study. Int. J. Biol. Macromol. 2017, 105 Pt 3, 1572–1580. [Google Scholar] [CrossRef]
- Shahabadi, N.; Khorshidi, A.; Moghadam, N.H. Study on the interaction of the epilepsy drug, zonisamide with human serum albumin (HSA) by spectroscopic and molecular docking techniques. Spectrochim. Acta A Mol. Biomol. Spectrosc. 2013, 114, 627–632. [Google Scholar] [CrossRef]
- Stan, D.; Matei, I.; Mihailescu, C.; Savin, M.; Hillebrand, M.; Baciu, I.; Matache, M. Spectroscopic investigations of the binding interaction of a new indanedione derivative with human and bovine serum albumins. Molecules 2009, 14, 1614–1626. [Google Scholar] [CrossRef]
- Tayyab, S.; Izzudin, M.M.; Kabir, M.Z.; Feroz, S.R.; Tee, W.V.; Mohamad, S.B.; Alias, Z. Binding of an anticancer drug, axitinib to human serum albumin: Fluorescence quenching and molecular docking study. J. Photochem. Photobiol. B 2016, 162, 386–394. [Google Scholar] [CrossRef]
- Kang, J.; Liu, Y.; Xie, M.X.; Li, S.; Jiang, M.; Wang, Y.D. Interactions of human serum albumin with chlorogenic acid and ferulic acid. Biochim. Biophys. Acta 2004, 1674, 205–214. [Google Scholar] [CrossRef] [PubMed]
- Tunc, S.; Duman, O.; Soylu, I.; Kanci Bozoglan, B. Study on the bindings of dichlorprop and diquat dibromide herbicides to human serum albumin by spectroscopic methods. J. Hazard. Mater. 2014, 273, 36–43. [Google Scholar] [CrossRef] [PubMed]
- Whitmore, L.; Wallace, B.A. Protein secondary structure analyses from circular dichroism spectroscopy: Methods and reference databases. Biopolymers 2008, 89, 392–400. [Google Scholar] [CrossRef] [PubMed]
- Gao, H.; Lei, L.; Liu, J.; Kong, Q.; Chen, X.; Hu, Z. The study on the interaction between human serum albumin and a new reagent with antitumour activity by spectrophotometric methods. J. Photochem. Photobiol. A Chem. 2004, 167, 213–221. [Google Scholar] [CrossRef]
- Rabbani, G.; Ahmad, E.; Zaidi, N.; Khan, R.H. pH-dependent conformational transitions in conalbumin (ovotransferrin), a metalloproteinase from hen egg white. Cell Biochem. Biophys. 2011, 61, 551–560. [Google Scholar] [CrossRef]
- Ross, P.D.; Subramanian, S. Thermodynamics of protein association reactions: Forces contributing to stability. Biochemistry 1981, 20, 3096–3102. [Google Scholar] [CrossRef]
- Hu, Y.J.; Liu, Y.; Xiao, X.H. Investigation of the interaction between Berberine and human serum albumin. Biomacromolecules 2009, 10, 517–521. [Google Scholar] [CrossRef]
- Bekale, L.; Chanphai, P.; Sanyakamdhorn, S.; Agudelo, D.; Tajmir-Riahi, H. Microscopic and thermodynamic analysis of PEG–β-lactoglobulin interaction. RSC Adv. 2014, 4, 31084–31093. [Google Scholar] [CrossRef]
- Főrster, T. 10th Spiers Memorial Lecture. Transfer mechanisms of electronic excitation. Discuss. Faraday Soc. 1959, 27, 7–17. [Google Scholar] [CrossRef]
- Naik, D.; Moorthy, P.; Priyadarsini, K. Nonradiative energy transfer from 7-amino coumarin dyes to thiazine dyes in methanolic solutions. Chem. Phys. Lett. 1990, 168, 533–538. [Google Scholar] [CrossRef]
- Farruggia, B.; Garcia, G.; D’Angelo, C.; Pico, G. Destabilization of human serum albumin by polyethylene glycols studied by thermodynamical equilibrium and kinetic approaches. Int. J. Biol. Macromol. 1997, 20, 43–51. [Google Scholar] [CrossRef]
- Farruggia, B.; Pico, G.A. Thermodynamic features of the chemical and thermal denaturations of human serum albumin. Int. J. Biol. Macromol. 1999, 26, 317–323. [Google Scholar] [CrossRef]
- Watanabe, H.; Tanase, S.; Nakajou, K.; Maruyama, T.; Kragh-Hansen, U.; Otagiri, M. Role of arg-410 and tyr-411 in human serum albumin for ligand binding and esterase-like activity. Biochem. J. 2000, 349 Pt 3, 813–819. [Google Scholar] [CrossRef]
- Nerli, B.; Pico, G. Evidence of human serum albumin beta-lactamase activity. Biochem. Mol. Biol. Int. 1994, 32, 789–795. [Google Scholar] [PubMed]
- Kragh-Hansen, U. Molecular and practical aspects of the enzymatic properties of human serum albumin and of albumin-ligand complexes. Biochim. Biophys. Acta 2013, 1830, 5535–5544. [Google Scholar] [CrossRef]
- Walker, J.E. Lysine residue 199 of human serum albumin is modified by acetylsalicyclic acid. FEBS Lett. 1976, 66, 173–175. [Google Scholar] [CrossRef]
- Phuangsawai, O.; Hannongbua, S.; Gleeson, M.P. Elucidating the origin of the esterase activity of human serum albumin using QM/MM calculations. J. Phys. Chem. B 2014, 118, 11886–11894. [Google Scholar] [CrossRef]
- Baig, M.H.; Ahmad, K.; Roy, S.; Ashraf, J.M.; Adil, M.; Siddiqui, M.H.; Khan, S.; Kamal, M.A.; Provaznik, I.; Choi, I. Computer Aided Drug Design: Success and Limitations. Curr. Pharm. Des. 2016, 22, 572–581. [Google Scholar] [CrossRef]
- Kragh-Hansen, U. Structure and ligand binding properties of human serum albumin. Dan. Med. Bull. 1990, 37, 57–84. [Google Scholar]
- Peters, T., Jr. Serum albumin. Adv. Protein Chem. 1985, 37, 161–245. [Google Scholar]
- Sudlow, G.; Birkett, D.J.; Wade, D.N. The characterization of two specific drug binding sites on human serum albumin. Mol. Pharm. 1975, 11, 824–832. [Google Scholar]
- Yeggoni, D.P.; Kuehne, C.; Rachamallu, A.; Subramanyam, R. Elucidating the binding interaction of andrographolide with the plasma proteins: Biophysical and computational approach. RSC Adv. 2017, 7, 5002–5012. [Google Scholar] [CrossRef]
- Rabbani, G.; Khan, M.J.; Ahmad, A.; Maskat, M.Y.; Khan, R.H. Effect of copper oxide nanoparticles on the conformation and activity of beta-galactosidase. Colloids Surf. B Biointerfaces 2014, 123, 96–105. [Google Scholar] [CrossRef] [PubMed]
- Masters, B.R. Principles of fluorescence spectroscopy. J. Biomed. Opt. 2008, 13, 029901. [Google Scholar]
- Chen, Y.H.; Yang, J.T.; Martinez, H.M. Determination of the secondary structures of proteins by circular dichroism and optical rotatory dispersion. Biochemistry 1972, 11, 4120–4131. [Google Scholar] [CrossRef] [PubMed]
- Santoro, M.M.; Bolen, D.W. Unfolding free energy changes determined by the linear extrapolation method. 1. Unfolding of phenylmethanesulfonyl alpha-chymotrypsin using different denaturants. Biochemistry 1988, 27, 8063–8068. [Google Scholar] [CrossRef] [PubMed]
- Goodsell, D.S.; Morris, G.M.; Olson, A.J. Automated docking of flexible ligands: Applications of AutoDock. J. Mol. Recognit. 1996, 9, 1–5. [Google Scholar] [CrossRef]
- Rose, P.W.; Prlic, A.; Bi, C.; Bluhm, W.F.; Christie, C.H.; Dutta, S.; Green, R.K.; Goodsell, D.S.; Westbrook, J.D.; Woo, J.; et al. The RCSB Protein Data Bank: Views of structural biology for basic and applied research and education. Nucleic Acids Res. 2015, 43, D345–D356. [Google Scholar] [CrossRef]
- Van Der Spoel, D.; Lindahl, E.; Hess, B.; Groenhof, G.; Mark, A.E.; Berendsen, H.J. GROMACS: Fast, flexible, and free. J. Comput. Chem. 2005, 26, 1701–1718. [Google Scholar] [CrossRef]
- Pronk, S.; Pall, S.; Schulz, R.; Larsson, P.; Bjelkmar, P.; Apostolov, R.; Shirts, M.R.; Smith, J.C.; Kasson, P.M.; van der Spoel, D.; et al. GROMACS 4.5: A high-throughput and highly parallel open source molecular simulation toolkit. Bioinformatics 2013, 29, 845–854. [Google Scholar] [CrossRef]
- Oostenbrink, C.; Villa, A.; Mark, A.E.; van Gunsteren, W.F. A biomolecular force field based on the free enthalpy of hydration and solvation: The GROMOS force-field parameter sets 53A5 and 53A6. J. Comput. Chem. 2004, 25, 1656–1676. [Google Scholar] [CrossRef] [PubMed]
- Toukan, K.; Rahman, A. Molecular-dynamics study of atomic motions in water. Phys. Rev. B Condens. Matter 1985, 31, 2643–2648. [Google Scholar] [CrossRef] [PubMed]
- Darden, T.; York, D.; Pedersen, L. Particle mesh Ewald: An N log (N) method for Ewald sums in large systems. J. Chem. Phys. 1993, 98, 10089–10092. [Google Scholar] [CrossRef]
- Hess, B. P-LINCS: A Parallel Linear Constraint Solver for Molecular Simulation. J. Chem. Theory Comput. 2008, 4, 116–122. [Google Scholar] [CrossRef] [PubMed]
- Eslami, H.; Mojahedi, F.; Moghadasi, J. Molecular dynamics simulation with weak coupling to heat and material baths. J. Chem. Phys. 2010, 133, 084105. [Google Scholar] [CrossRef] [PubMed]
- Humphrey, W.; Dalke, A.; Schulten, K. VMD: Visual molecular dynamics. J. Mol. Graph. 1996, 14, 33–38. [Google Scholar] [CrossRef]


| Parameter | 25 °C | 30 °C | 37 °C |
|---|---|---|---|
| n (binding stoichiometry, HSA:CBH) | 0.99 ± 0.11 | 1.01 ± 0.13 | 1.101 ± 0.12 |
| KSV (Stern-Volmer constant, M−1) | 1.7 ± 0.21 × 104 | 1.3 ± 0.24 × 104 | 1.0 ± 0.31 × 104 |
| Kb (binding constant, M−1) | 1.6 ± 0.22 × 104 | 1.4 ± 0.19 × 104 | 1.2 ± 0.23 × 104 |
| kq (bimolecular quenching rate constant, M−1 s−1) | 2.6 ± 0.12×1012 | 2.2 ± 0.15 × 1012 | 1.7 ± 0.19 × 1012 |
| ΔH° (binding enthalpy, kcal mol−1) | - | −4.44±0.16 | - |
| TΔS° (entropy change, kcal mol-1) | 1.28 ± 0.18 | 1.30 ± 0.15 | 1.33 ± 0.16 |
| ΔG° (Gibbs free energy change, kcal mol-1) | −5.72 ± 0.12 | −5.75 ± 0.15 | −5.78 ± 0.14 |
| Parameter | Value |
|---|---|
| J (cm−3 M−1) | 3.08 × 10−15 |
| Ro (nm) | 2.18 |
| r (nm) | 2.34 |
| EFRET | 0.39 |
| Fo | 2546 |
| F | 1829 |
| Protein | ΔGDo, kcal mol−1 | mg, kcal mol−1 M−1 | Cm, M |
|---|---|---|---|
| 0.0 μM CBH | 4.92 ± 0.24 | 1.98 ± 0.14 | 2.48 |
| 100 μM CBH | 5.42 ± 0.19 | 2.01 ± 0.18 | 2.69 |
| HSA:CBH | Vmax (mM min−1) | Km (mM) | kcat (min−1) | kcat/Km (mM−1 min−1) |
|---|---|---|---|---|
| 1:0 | 10.2 × 10−4 | 34.8 × 10−2 | 12.8 × 10−2 | 0.36 |
| 1:5 | 10.2 × 10−4 | 50.6 × 10−2 | 12.8 × 10−2 | 0.25 |
| 1:10 | 10.0 × 10−4 | 60.0 × 10−2 | 12.5 × 10−2 | 0.20 |
| Complex | Binding Free Energy (ΔG) (kcal mol−1) | Residues Involved | |
|---|---|---|---|
| Hydrogen Bond | Hydrophobic Interaction | ||
| HSA-CBH | −7.55 | S192 | Y150, E153, S192, K195, Q196, K199, L238, R257, H288, A291, E292 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Baig, M.H.; Rahman, S.; Rabbani, G.; Imran, M.; Ahmad, K.; Choi, I. Multi-Spectroscopic Characterization of Human Serum Albumin Binding with Cyclobenzaprine Hydrochloride: Insights from Biophysical and In Silico Approaches. Int. J. Mol. Sci. 2019, 20, 662. https://doi.org/10.3390/ijms20030662
Baig MH, Rahman S, Rabbani G, Imran M, Ahmad K, Choi I. Multi-Spectroscopic Characterization of Human Serum Albumin Binding with Cyclobenzaprine Hydrochloride: Insights from Biophysical and In Silico Approaches. International Journal of Molecular Sciences. 2019; 20(3):662. https://doi.org/10.3390/ijms20030662
Chicago/Turabian StyleBaig, Mohammad Hassan, Safikur Rahman, Gulam Rabbani, Mohd Imran, Khurshid Ahmad, and Inho Choi. 2019. "Multi-Spectroscopic Characterization of Human Serum Albumin Binding with Cyclobenzaprine Hydrochloride: Insights from Biophysical and In Silico Approaches" International Journal of Molecular Sciences 20, no. 3: 662. https://doi.org/10.3390/ijms20030662
APA StyleBaig, M. H., Rahman, S., Rabbani, G., Imran, M., Ahmad, K., & Choi, I. (2019). Multi-Spectroscopic Characterization of Human Serum Albumin Binding with Cyclobenzaprine Hydrochloride: Insights from Biophysical and In Silico Approaches. International Journal of Molecular Sciences, 20(3), 662. https://doi.org/10.3390/ijms20030662