Identification and Characterization of Tomato SWI3-Like Proteins: Overexpression of SlSWIC Increases the Leaf Size in Transgenic Arabidopsis
Abstract
:1. Introduction
2. Results
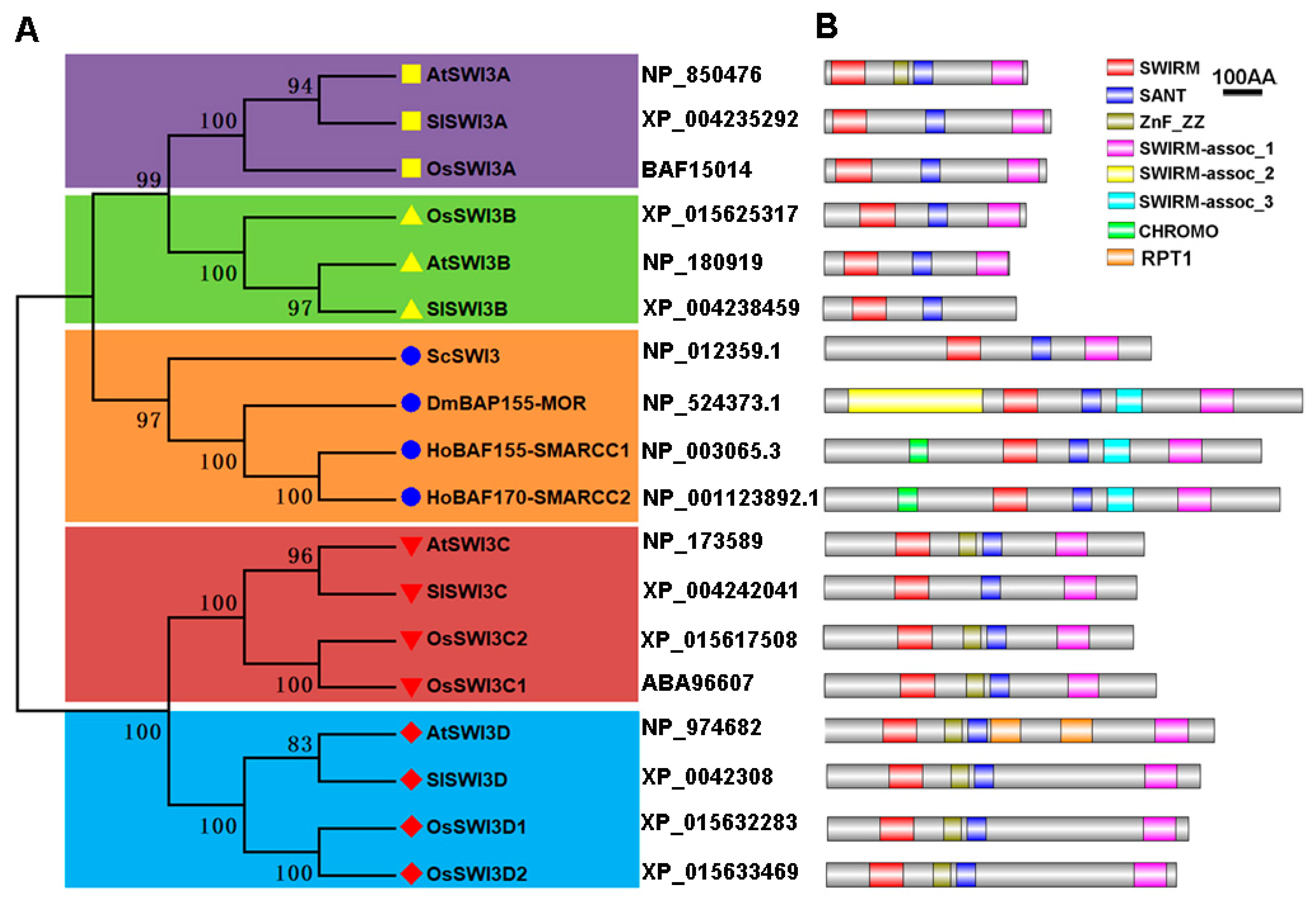
2.1. Identification and Phylogenetic Analysis of Tomato Swi3-Like Proteins
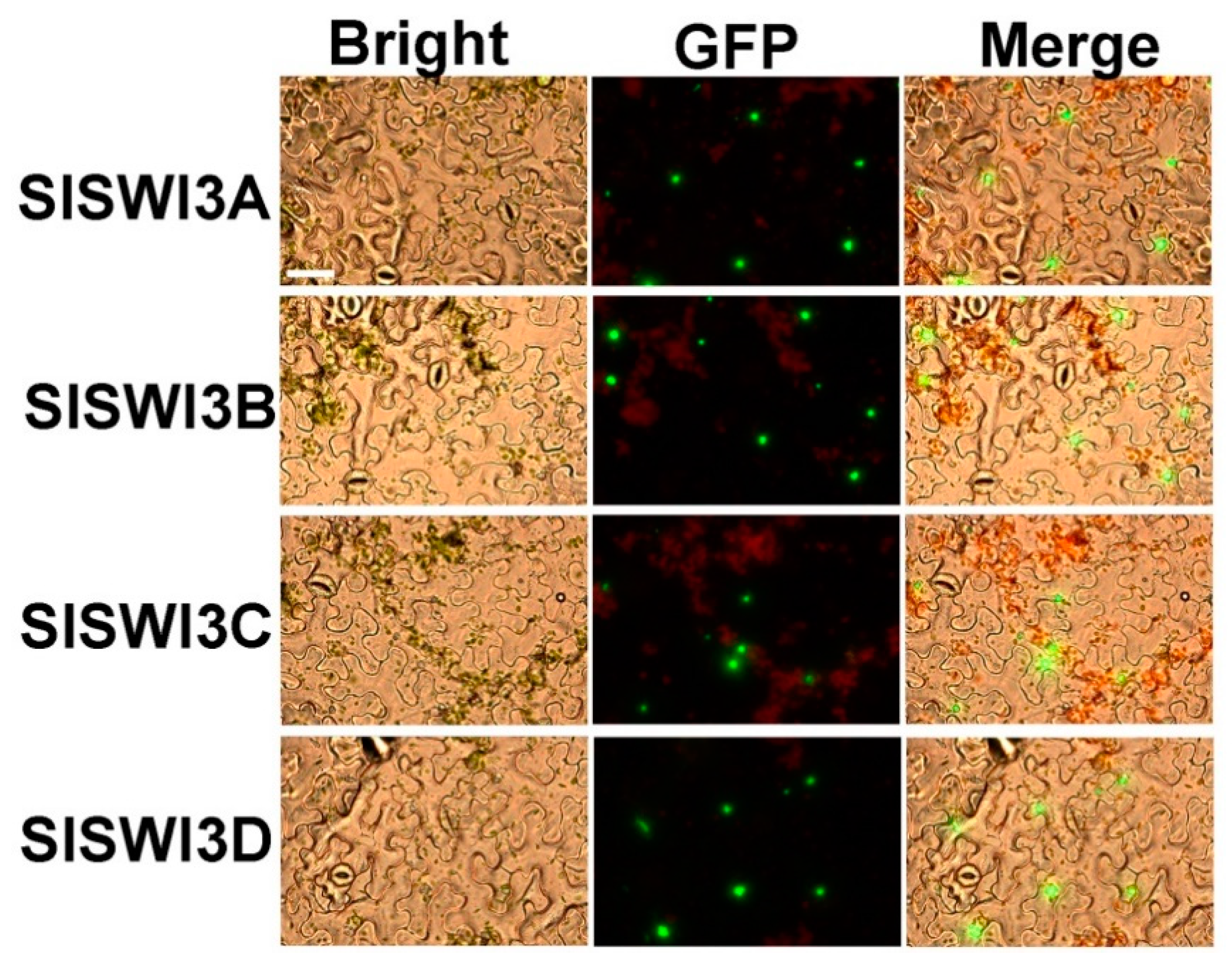
2.2. Subcellular Localization of Tomato Swi3-Like Proteins
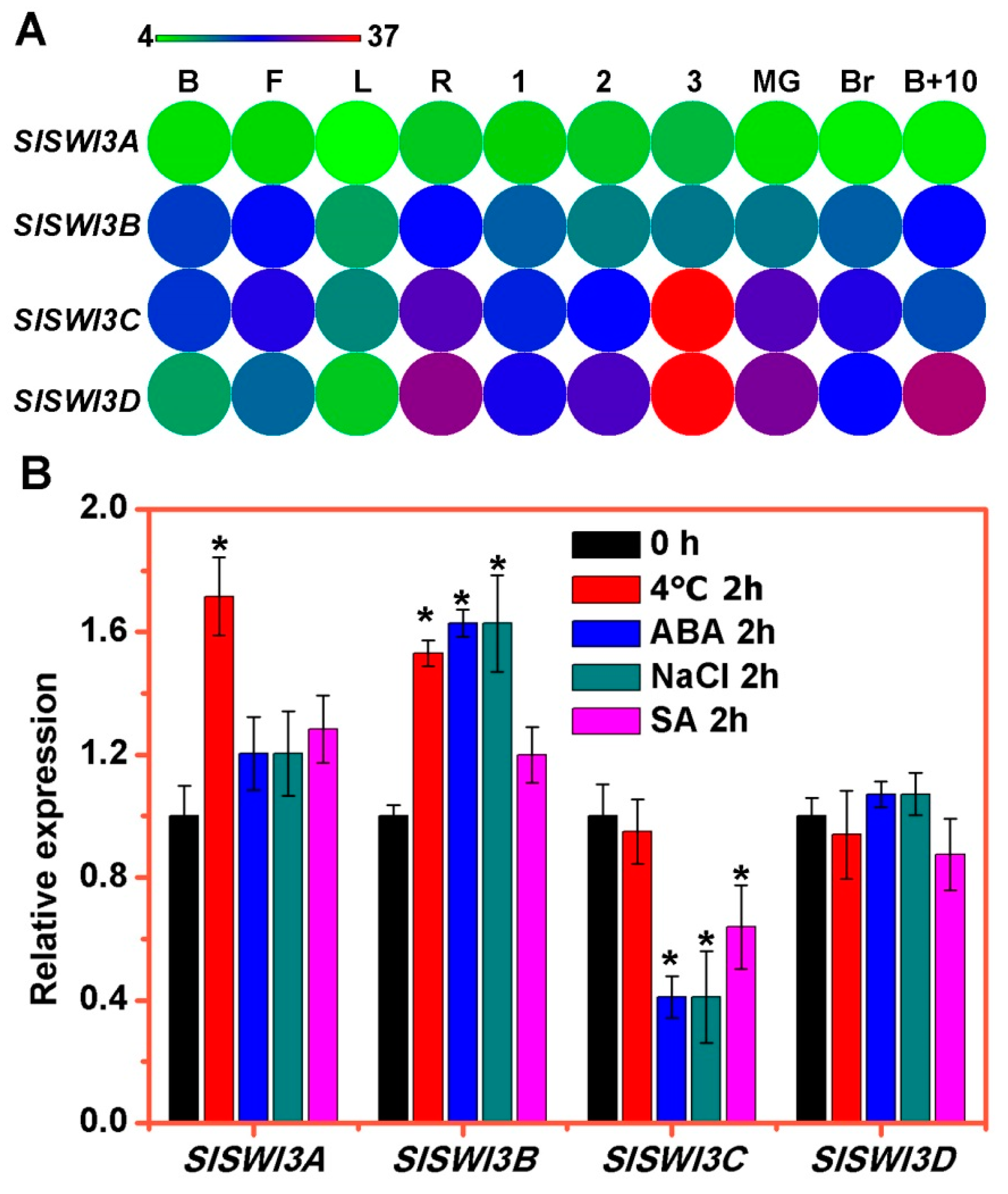
2.3. The Expression Patterns of Tomato Swi3-Like Genes
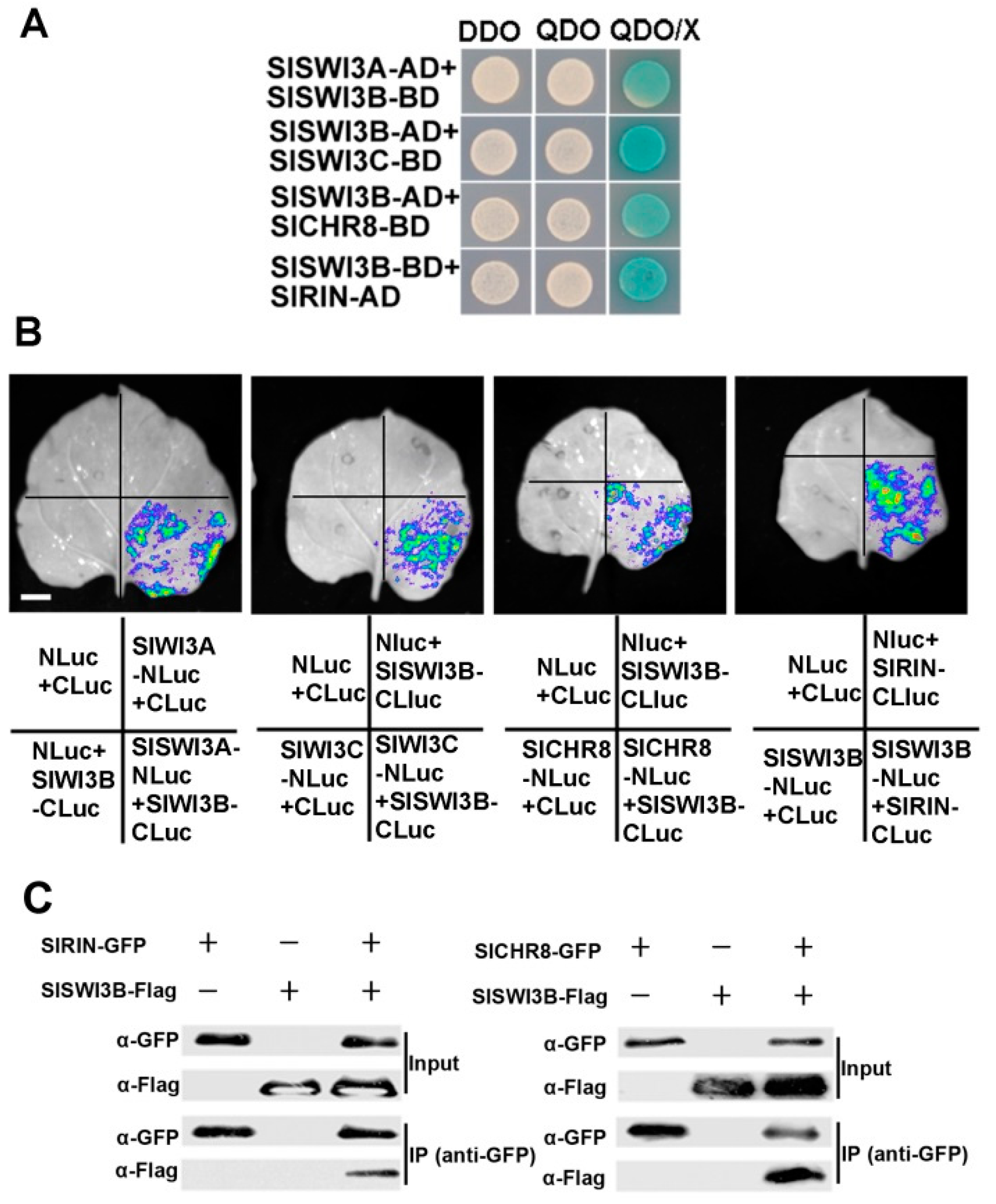
2.4. Members of SlSWI3s Interact with Other Proteins
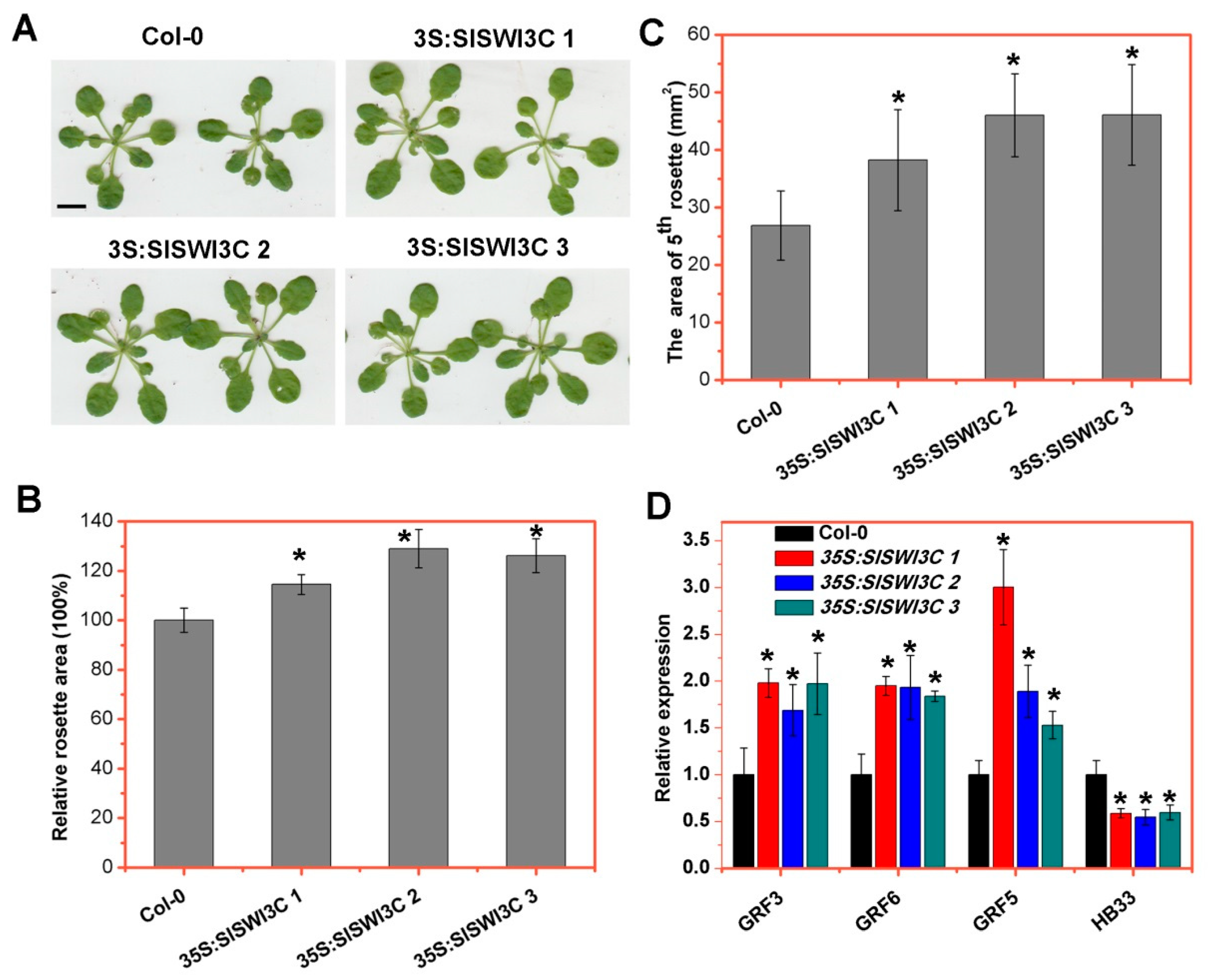
2.5. Ectopic Overexpression of SlSWI3C Enhances Leaf Growth
3. Discussion
4. Materials and Methods
4.1. Plant Materials and Growth Conditions
4.2. Identification of Tomato Swi3-Like Genes and Phylogenetic Tree Construction
4.3. Subcellular Localization Assays
4.4. Yeast-Two-Hybrid Assay
4.5. LCI Assay
4.6. Co-Immunoprecipitation (Co-IP) Assays
4.7. RNA Extraction and Expression Analyses
4.8. Generation of Transgenic Plants
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Luger, K.; Mader, A.W.; Richmond, R.K.; Sargent, D.F.; Richmond, T.J. Crystal structure of the nucleosome core particle at 2.8 A resolution. Nature 1997, 389, 251–260. [Google Scholar] [CrossRef] [PubMed]
- Henikoff, S.; Shilatifard, A. Histone modification: Cause or cog? Trends in Genet. 2011, 27, 389–396. [Google Scholar] [CrossRef] [PubMed]
- Jerzmanowski, A. SWI/SNF chromatin remodeling and linker histones in plants. Biochim. Biophys. Acta 2007, 1769, 330–345. [Google Scholar] [CrossRef] [PubMed]
- Imbalzano, A.N. Energy-dependent chromatin remodelers: Complex complexes and their components. Crit. Rev. Eukaryot. Gene Expr. 1998, 8, 225–255. [Google Scholar] [CrossRef]
- Clapier, C.R.; Cairns, B.R. The Biology of Chromatin Remodeling Complexes. Annu. Rev. Biochem. 2009, 78, 273–304. [Google Scholar] [CrossRef]
- Neigeborn, L.; Carlson, M. Genes Affecting the Regulation of Suc2 Gene-Expression by Glucose Repression in Saccharomyces-Cerevisiae. Genetics 1984, 108, 845–858. [Google Scholar]
- Peterson, C.L.; Herskowitz, I. Characterization of the Yeast Swi1, Swi2, and Swi3 Genes, Which Encode a Global Activator of Transcription. Cell 1992, 68, 573–583. [Google Scholar] [CrossRef]
- Cairns, B.R.; Kim, Y.J.; Sayre, M.H.; Laurent, B.C.; Kornberg, R.D. A Multisubunit Complex Containing the Swi1/Adr6, Swi2/Snf2, Swi3, Snf5, and Snf6 Gene-Products Isolated from Yeast. Proc. Natl Acad. Sci. USA 1994, 91, 1950–1954. [Google Scholar] [CrossRef]
- Phelan, M.L.; Sif, S.; Narlikar, G.J.; Kingston, R.E. Reconstitution of a core chromatin remodeling complex from SWI/SNF subunits. Mol. Cell 1999, 3, 247–253. [Google Scholar] [CrossRef]
- Johnson, C.N.; Adkins, N.L.; Georgel, P. Chromatin remodeling complexes: ATP-dependent machines in action. Biochem. Cell Biol. 2005, 83, 405–417. [Google Scholar] [CrossRef]
- Yang, X.F.; Zaurin, R.; Beato, M.; Peterson, C.L. Swi3p controls SWI/SNF assembly and ATP-dependent H2A-H2B displacement. Nat. Struct. Mol. Biol. 2007, 14, 540–547. [Google Scholar] [CrossRef] [PubMed]
- Dechassa, M.L.; Zhang, B.; Horowitz-Scherer, R.; Persinger, J.; Woodcock, C.L.; Peterson, C.L.; Bartholomew, B. Architecture of the SWI/SNF-nucleosome complex. Mol. Cell Biol. 2008, 28, 6010–6021. [Google Scholar] [CrossRef] [PubMed]
- Hu, Z.Z.; Killion, P.J.; Iyer, V.R. Genetic reconstruction of a functional transcriptional regulatory network. Nat. Genet. 2007, 39, 683–687. [Google Scholar] [CrossRef] [PubMed]
- Euskirchen, G.M.; Auerbach, R.K.; Davidov, E.; Gianoulis, T.A.; Zhong, G.N.; Rozowsky, J.; Bhardwaj, N.; Gerstein, M.B.; Snyder, M. Diverse Roles and Interactions of the SWI/SNF Chromatin Remodeling Complex Revealed Using Global Approaches. Plos Genet. 2011, 7, e1002008. [Google Scholar] [CrossRef]
- Aravind, L.; Iyer, L.M. The SWIRM domain: A conserved module found in chromosomal proteins points to novel chromatin-modifying activities. Genome Bio. 2002, 3. [Google Scholar] [CrossRef]
- Aasland, R.; Stewart, A.F.; Gibson, T. The SANT domain: A putative DNA-binding domain in the SWI-SNF and ADA complexes, the transcriptional corepressor N-CoR and TFIIIB. Trends Biochem. Sci. 1996, 21, 87–88. [Google Scholar] [CrossRef]
- Boyer, L.A.; Langer, M.R.; Crowley, K.A.; Tan, S.; Denu, J.M.; Peterson, C.L. Essential role for the SANT domain in the functioning of multiple chromatin remodeling enzymes. Mol. Cell 2002, 10, 935–942. [Google Scholar] [CrossRef]
- Sarnowski, T.J.; Swiezewski, S.; Pawlikowska, K.; Kaczanowski, S.; Jerzmanowski, A. AtSWI3B, an Arabidopsis homolog of SWI3, a core subunit of yeast Swi/Snf chromatin remodeling complex, interacts with FCA, a regulator of flowering time. Nucleic Acids Res. 2002, 30, 3412–3421. [Google Scholar] [CrossRef] [Green Version]
- Zhou, C.H.; Miki, B.; Wu, K.Q. CHB2, a member of the SWI3 gene family, is a global regulator in Arabidopsis. Plant Mol. Biol. 2003, 52, 1125–1134. [Google Scholar] [CrossRef]
- Sarnowski, T.J.; Rios, G.; Jasik, J.; Swiezewski, S.; Kaczanowski, S.; Li, Y.; Kwiatkowska, A.; Pawlikowska, K.; Kozbial, M.; Kozbial, P.; et al. SWI3 subunits of putative SWI/SNF chromatin-remodeling complexes play distinct roles during Arabidopsis development. Plant Cell 2005, 17, 2454–2472. [Google Scholar] [CrossRef]
- Saez, A.; Rodrigues, A.; Santiago, J.; Rubio, S.; Rodriguez, P.L. HAB1-SWI3B Interaction Reveals a Link between Abscisic Acid Signaling and Putative SWI/SNF Chromatin-Remodeling Complexes in Arabidopsis. Plant Cell 2008, 20, 2972–2988. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.Y.; Rowley, M.J.; Bohmdorfer, G.; Wierzbicki, A.T. A SWI/SNF Chromatin-Remodeling Complex Acts in Noncoding RNA-Mediated Transcriptional Silencing. Mol. Cell 2013, 49, 298–309. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, Z.W.; Zhou, J.X.; Huang, H.W.; Li, Y.Q.; Shao, C.R.; Li, L.; Cai, T.; Chen, S.; He, X.J. Two Components of the RNA-Directed DNA Methylation Pathway Associate with MORC6 and Silence Loci Targeted by MORC6 in Arabidopsis. Plos Genet. 2016, 12, e1006026. [Google Scholar] [CrossRef] [PubMed]
- Han, W.X.; Han, D.L.; He, Z.P.; Hu, H.; Wu, Q.; Zhang, J.J.; Jiang, J.M.; Qin, G.J.; Cui, Y.H.; Lai, J.B.; et al. The SWI/SNF subunit SWI3B regulates IAMT1 expression via chromatin remodeling in Arabidopsis leaf development. Plant Sci. 2018, 271, 127–132. [Google Scholar] [CrossRef] [PubMed]
- Sarnowska, E.A.; Rolicka, A.T.; Bucior, E.; Cwiek, P.; Tohge, T.; Fernie, A.R.; Jikumaru, Y.; Kamiya, Y.; Franzen, R.; Schmelzer, E.; et al. DELLA-Interacting SWI3C Core Subunit of Switch/Sucrose Nonfermenting Chromatin Remodeling Complex Modulates Gibberellin Responses and Hormonal Cross Talk in Arabidopsis. Plant Physiol. 2013, 163, 305–317. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vercruyssen, L.; Verkest, A.; Gonzalez, N.; Heyndrickx, K.S.; Eeckhout, D.; Han, S.K.; Jegu, T.; Archacki, R.; Van Leene, J.; Andriankaja, M.; et al. ANGUSTIFOLIA3 Binds to SWI/SNF Chromatin Remodeling Complexes to Regulate Transcription during Arabidopsis Leaf Development. Plant Cell 2014, 26, 210–229. [Google Scholar] [CrossRef] [PubMed]
- Yu, X.; Jiang, L.; Wu, R.; Meng, X.; Zhang, A.; Li, N.; Xia, Q.; Qi, X.; Pang, J.; Xu, Z.Y.; et al. The Core Subunit of a Chromatin-Remodeling Complex, ZmCHB101, Plays Essential Roles in Maize Growth and Development. Sci. Rep. 2016, 6, 38504. [Google Scholar] [CrossRef]
- Gao, Y.; Yang, S.G.; Yuan, L.Y.; Cui, Y.H.; Wu, K.Q. Comparative Analysis of SWIRM Domain-Containing Proteins in Plants. Comp. Funct. Genom. 2012, 2012, 310402. [Google Scholar] [CrossRef]
- Ponting, C.P.; Blake, D.J.; Davies, K.E.; KendrickJones, J.; Winder, S.J. ZZ and TAZ: New putative zinc fingers in dystrophin and other proteins. Trends Biochem. Sci. 1996, 21, 11–13. [Google Scholar] [CrossRef]
- Legge, G.B.; Martinez-Yamout, M.A.; Hambly, D.M.; Trinh, T.; Lee, B.M.; Dyson, H.J.; Wright, P.E. ZZ domain of CBP: An unusual zinc finger fold in a protein interaction module. J. Mol. Biol. 2004, 343, 1081–1093. [Google Scholar] [CrossRef]
- Sparkes, I.A.; Runions, J.; Kearns, A.; Hawes, C. Rapid, transient expression of fluorescent fusion proteins in tobacco plants and generation of stably transformed plants. Nat. Protoc. 2006, 1, 2019–2025. [Google Scholar] [CrossRef] [PubMed]
- Sato, S.; Tabata, S.; Hirakawa, H.; Asamizu, E.; Shirasawa, K.; Isobe, S.; Kaneko, T.; Nakamura, Y.; Shibata, D.; Aoki, K.; et al. The tomato genome sequence provides insights into fleshy fruit evolution. Nature 2012, 485, 635–641. [Google Scholar] [Green Version]
- Efroni, I.; Han, S.K.; Kim, H.J.; Wu, M.F.; Steiner, E.; Birnbaum, K.D.; Hong, J.C.; Eshed, Y.; Wagner, D. Regulation of Leaf Maturation by Chromatin-Mediated Modulation of Cytokinin Responses. Dev. Cell 2013, 24, 438–445. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hurtado, L.; Farrona, S.; Reyes, J.C. The putative SWI/SNF complex subunit BRAHMA activates flower homeotic genes in Arabidopsis thaliana. Plant. Mol. Biol. 2006, 62, 291–304. [Google Scholar] [CrossRef] [PubMed]
- Vrebalov, J.; Ruezinsky, D.; Padmanabhan, V.; White, R.; Medrano, D.; Drake, R.; Schuch, W.; Giovannoni, J.A. MADS-box gene necessary for fruit ripening at the tomato ripening-inhibitor (Rin) locus. Science 2002, 296, 343–346. [Google Scholar] [CrossRef]
- Zhang, D.D.; Gao, S.J.; Yang, P.; Yang, J.; Yang, S.G.; Wu, K.Q. Identification and Expression Analysis of Snf2 Family Proteins in Tomato (Solanum lycopersicum). Int. J. Genom. 2019. [Google Scholar] [CrossRef]
- Folta, A.; Bargsten, J.W.; Bisseling, T.; Nap, J.P.; Mlynarova, L. Compact tomato seedlings and plants upon overexpression of a tomato chromatin remodelling ATPase gene. Plant. Biotechnol. J. 2016, 14, 581–591. [Google Scholar] [CrossRef]
- Kim, J.H.; Choi, D.S.; Kende, H. The AtGRF family of putative transcription factors is involved in leaf and cotyledon growth in Arabidopsis. Plant. J. 2003, 36, 94–104. [Google Scholar] [CrossRef]
- Horiguchi, G.; Kim, G.T.; Tsukaya, H. The transcription factor AtGRF5 and the transcription coactivator AN3 regulate cell proliferation in leaf primordia of Arabidopsis thaliana. Plant. J. 2005, 43, 68–78. [Google Scholar] [CrossRef]
- Kim, J.H.; Lee, B.H. GROWTH-REGULATING FACTOR4 of Arabidopsis thaliana is required for development of leaves, cotyledons, and shoot apical meristem. J. Plant. Biol. 2006, 49, 463–468. [Google Scholar] [CrossRef]
- Kim, S.S.; Kim, M.S.; Yoo, N.J.; Lee, S.H. Frameshift mutations of a chromatin-remodeling gene SMARCC2 in gastric and colorectal cancers with microsatellite instability. Apmis 2013, 121, 168–169. [Google Scholar] [CrossRef] [PubMed]
- Kadoch, C.; Hargreaves, D.C.; Hodges, C.; Elias, L.; Ho, L.; Ranish, J.; Crabtree, G.R. Proteomic and bioinformatic analysis of mammalian SWI/SNF complexes identifies extensive roles in human malignancy. Nat. Genet. 2013, 45, 592–601. [Google Scholar] [CrossRef] [PubMed]
- Boyer, L.A.; Latek, R.R.; Peterson, C.L. The SANT domain: A unique histone-tail-binding module? Nat. Rev. Mol. Cell Biol. 2004, 5, 158–163. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.D.; Xue, Y.T.; Zhou, S.; Kuo, A.; Cairns, B.R.; Crabtree, G.R. Diversity and specialization of mammalian SWI/SNF complexes. Gene Dev. 1996, 10, 2117–2130. [Google Scholar] [CrossRef]
- Winter, D.; Vinegar, B.; Nahal, H.; Ammar, R.; Wilson, G.V.; Provart, N.J. An “Electronic Fluorescent Pictograph” Browser for Exploring and Analyzing Large-Scale Biological Data Sets. PLoS ONE 2007, 2, e718. [Google Scholar] [CrossRef]
- Ren, J.; Wen, L.P.; Gao, X.J.; Jin, C.J.; Xue, Y.; Yao, X.B. DOG 1.0: Illustrator of protein domain structures. Cell Res. 2009, 19, 271–273. [Google Scholar] [CrossRef]
- Tamura, K.; Peterson, D.; Peterson, N.; Stecher, G.; Nei, M.; Kumar, S. MEGA5: Molecular Evolutionary Genetics Analysis Using Maximum Likelihood, Evolutionary Distance, and Maximum Parsimony Methods. Mol. Biol. Evol. 2011, 28, 2731–2739. [Google Scholar] [CrossRef] [Green Version]
- Peyret, H.; Lomonossoff, G.P. The pEAQ vector series: The easy and quick way to produce recombinant proteins in plants. Plant. Mol. Biol. 2013, 83, 51–58. [Google Scholar] [CrossRef]
- Chen, H.M.; Zou, Y.; Shang, Y.L.; Lin, H.Q.; Wang, Y.J.; Cai, R.; Tang, X.Y.; Zhou, J.M. Firefly luciferase complementation imaging assay for protein-protein interactions in plants. Plant. Physiol. 2008, 146, 368–376. [Google Scholar] [CrossRef]
- Zhao, M.; Yang, S.; Chen, C.Y.; Li, C.; Shan, W.; Lu, W.; Cui, Y.; Liu, X.; Wu, K. Arabidopsis BREVIPEDICELLUS interacts with the SWI2/SNF2 chromatin remodeling ATPase BRAHMA to regulate KNAT2 and KNAT6 expression in control of inflorescence architecture. Plos Genet. 2015, 11, e1005125. [Google Scholar] [CrossRef]
- Pavlidis, P.; Noble, W.S. Matrix2png: A utility for visualizing matrix data. Bioinformatics 2003, 19, 295–296. [Google Scholar] [CrossRef] [PubMed]
- Mao, J.; Zhang, Y.C.; Sang, Y.; Li, Q.H.; Yang, H.Q. From The Cover: A role for Arabidopsis cryptochromes and COP1 in the regulation of stomatal opening. Proc. Natl. Acad. Sci. USA 2005, 102, 12270–12275. [Google Scholar] [CrossRef] [PubMed]
| SWI3 Gene Family | Gene Name | Gene Code | Accession Number a | ORF Length b | Protein Length | Localization c | Number of Exons | Nuclear Localization Signal d |
|---|---|---|---|---|---|---|---|---|
| SWI3A | SlSWI3A | Solyc03g097450 | XP_004235292 | 1772 | 574 | nucl, cyto | 7 | Yes |
| AtSWI3A | AT2G47620 | NP_850476 | 1539 | 513 | nucl, cyto | 7 | Yes | |
| OsSWI3A | LOC_Os04g40420 | BAF15014 | 1683 | 561 | nucl, cyto, plas | 7 | Yes | |
| SWI3B | SlSWI3B | Solyc04g082760 | XP_004238459 | 1470 | 490 | nucl, cyto | 6 | Yes |
| AtSWI3B | AT2G33610 | NP_180919 | 1410 | 469 | nucl, cyto | 6 | Yes | |
| OsSWI3B | LOC_Os02g10060 | XP_015625317 | 1536 | 512 | nucl | 6 | Yes | |
| SWI3C | SlSWI3C | Solyc06g060120 | XP_004242041 | 2376 | 792 | nucl, chlo, mito | 8 | Yes |
| AtSWI3C | AT1G21700 | NP_173589 | 2424 | 807 | nucl, chlo, plas | 9 | Yes | |
| OsSWI3C1 | LOC_Os12g07730 | ABA96607 | 2520 | 840 | nucl, chlo, mito | 9 | Yes | |
| OsSWI3C2 | LOC_Os11g08080 | XP_015617508 | 2355 | 785 | nucl, chlo, mito | 9 | Yes | |
| SWI3D | SlSWI3D | Solyc01g109510 | XP_004230866 | 2838 | 946 | nucl | 6 | Yes |
| AtSWI3D | AT4G34430 | NP_974682 | 2961 | 986 | nucl | 7 | Yes | |
| OsSWI3D1 | LOC_Os03g51220 | XP_015632283 | 2745 | 915 | nucl, cyto | 8 | Yes | |
| OsSWI3D2 | LOC_Os04g01970 | XP_015633469 | 2661 | 887 | nucl, chlo, mito | 7 | Yes |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhao, Z.; Li, T.; Peng, X.; Wu, K.; Yang, S. Identification and Characterization of Tomato SWI3-Like Proteins: Overexpression of SlSWIC Increases the Leaf Size in Transgenic Arabidopsis. Int. J. Mol. Sci. 2019, 20, 5121. https://doi.org/10.3390/ijms20205121
Zhao Z, Li T, Peng X, Wu K, Yang S. Identification and Characterization of Tomato SWI3-Like Proteins: Overexpression of SlSWIC Increases the Leaf Size in Transgenic Arabidopsis. International Journal of Molecular Sciences. 2019; 20(20):5121. https://doi.org/10.3390/ijms20205121
Chicago/Turabian StyleZhao, Zhongyi, Tao Li, Xiuling Peng, Keqiang Wu, and Songguang Yang. 2019. "Identification and Characterization of Tomato SWI3-Like Proteins: Overexpression of SlSWIC Increases the Leaf Size in Transgenic Arabidopsis" International Journal of Molecular Sciences 20, no. 20: 5121. https://doi.org/10.3390/ijms20205121





