The Potential of the FSP1cre-Pparb/d−/− Mouse Model for Studying Juvenile NAFLD
Abstract
:1. Introduction
2. Results
2.1. Deletion of Pparb/d in FSP1+CD11b+ Cells
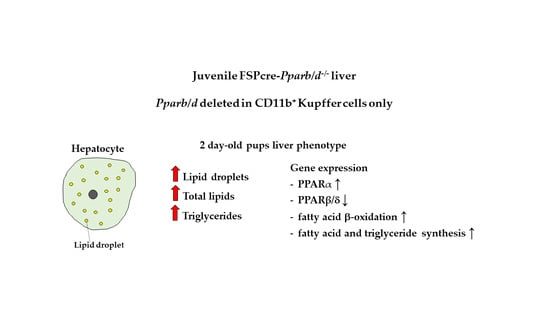
2.2. More and Larger Intracellular Lipid Droplets in FSP1cre-Pparb/d−/− Livers at P2 and Week 4
2.3. Higher Triglyceride Levels in the Liver of FSP1cre-Pparb/d−/− Mice at P2 and Week 4
2.4. Upregulation of Genes Involved in Fatty Acid β-Oxidation, Fatty Acid Synthesis and Triglyceride Synthesis in FSP1cre-Pparb/d−/− Liver at P2 and Week 4
2.5. Increase in Hadha, Phospho-ACLY, Phospho-ACC and GPD2 in the Liver of P2 FSP1cre-Pparb/d−/− Mice
3. Discussion
4. Materials and Methods
4.1. Chemicals and Antibodies
4.2. Animals
4.3. Genotyping Analysis
4.4. Histology
4.5. Double Immunofluorescence Staining
4.6. Isolation of Hepatic Cells
4.7. Flow Cytometry
4.8. Quantification of Total Lipids, Total Cholesterol, and Triglycerides
4.9. Total RNA Extraction and Reverse Transcription
4.10. Quantitative Real-Time PCR
4.11. Western Blot and Densitometric Analysis
4.12. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| ACAA2 | acetyl-CoA acyltransferase 2 |
| ACAT1 | acetyl-CoA acetyltransferase 1 |
| ACC | acetyl-CoA synthetase |
| ACLY | acyl-CoA oxidase 1 |
| ACS | acetyl-CoA synthetase |
| ACOX1 | acyl-CoA oxidase 1 |
| AGPAT3 | 1-acylglycerol-3-phosphate O-acyltransferase 3 |
| AMPK | AMP-activated protein kinase |
| CYP51 | cytochrome P450 family 51 subfamily A member 1 |
| DGAT1 | diacylglycerol O-acyltransferase 1 |
| DHCR7 | 7-dehydrocholesterol reductase |
| FAS | fatty acid synthase |
| FSP1 | fibroblast-specific protein 1 |
| GFAP | glial fibrillary acidic protein |
| GPD1 | glycerol 3-phosphate dehydrogenase 1 |
| GPD2 | glycerol 3-phosphate dehydrogenase 2 |
| H&E | hematoxylin and eosin |
| HADHA | hydroxyacyl-CoA dehydrogenase |
| HMGCS1 | hydroxymethylglutaryl-CoA synthase |
| HSC | hepatic stellate cell |
| IL4 | interleukin 4 |
| IL13 | interleukin 13 |
| IgG | immunoglobulin G |
| LPIN3 | lipin 3 |
| LSEC | liver sinusoidal endothelial cell |
| NAFLD | non-alcoholic fatty liver disease |
| NASH | non-alcoholic steatohepatitis |
| PLIN5 | perilipin 5 |
| PPAR | peroxisome proliferator-activated receptor |
| ROS | reactive oxygen species |
| SCD1 | stearoyl-CoA desaturase 1 |
| SREBP1c | sterol regulatory element-binding protein 1 |
| SREBP2 | sterol regulatory element-binding protein 2 |
| TG | triglycerides |
| TZDs | thiazolidinediones |
Appendix A
| Pparb/dfl/fl Genotyping Protocol | |
| PCR Reaction mixture | PCR program setting |
| 1.2 µL PBX10 | 95 °C 3 min |
| 0.6 µL AB008 | |
| 0.6 µL AB021 | 30 cycles: |
| 8.1 µL Water | 94 °C 15 s |
| 12.5 µL 2× KAPPA2G Fast (HotStart) genotyping mix with dye | 68 °C 10 s |
| 72 °C 20 s | |
| 72 °C 10 min | |
| 4 °C hold | |
| FSP1cre-Pparb/d−/− Genotyping Protocol | |
| PCR Reaction mixture | PCR program setting |
| 0.375 µL Fsp1 Ctrl Forward | 94 °C 3 min |
| 0.375 µL Fsp1 Ctrl Reverse | |
| 1.25 µL Fsp1 Forward | 30 cycles: |
| 1.25 µL Fsp1 Reverse | 95 °C 15 s |
| 8.25 µL Water | 59.2 °C 30 s |
| 12.5 µL 2× KAPPA2G Fast (HotStart) genotyping mix with dye | 72 °C 10 s |
| 1.0 µL template | |
| 72 °C 10 min | |
| 4 °C hold | |
| Sry Genotyping Protocol | |
| PCR Reaction mixture | PCR program setting |
| 1 µL Sry Forward | 94 °C 5 min |
| 1 µL Sry Reverse | |
| 3.5 µL Water | 33 cycles: |
| 7.5 µL 2× KAPPA2G Fast (HotStart) genotyping mix with dye | 94 °C 30 s |
| 1.0 µL template | 60 °C 30 s |
| 72 °C 45 s | |
| 72 °C 10 min | |
| 4 °C hold |
Appendix B
| Gene | Primer Sequence |
| Acaa2 | Forward (5′-3′) TCTTGACCCCAGCAAAACCA |
| Reverse (5′-3′) CCACCTCGACGCCTTAACTC | |
| Acc | Forward (5′-3′) GACAACACCTGTGTGGTGGA |
| Reverse (5′-3′) GAGGTTGGAGGCAAAGGACA | |
| Acat1 | Forward (5′-3′) AATGCTGGAGATTGACCCCC |
| Reverse (5′-3′) CGGGCTCCAGACATCCCAAT | |
| Acly | Forward (5′-3′) GTCCCAAGTCCAAGATCCCTG |
| Reverse (5′-3′) TGTGATCCCCAGTGAAAGGG | |
| Acox1 | Forward (5′-3′) GAGCAGCAGGAGCGTTTCTT |
| Reverse (5′-3′) CAGGACTATCGCATGATTGGAAG | |
| Agpat3 | Forward (5′-3′) TACTGCGAAGGAACACGCTT |
| Reverse (5′-3′) TCTTGCAGGGCATCCTTCTC | |
| Cd36 | Forward (5′-3′) TGATACTATGCCCGCCTCTCC |
| Reverse (5′-3′) TTTCCCACACTCCTTTCTCCTCTA | |
| Cyp51 | Forward (5′-3′) CGTCTACCTGTTCCGTCTCG |
| Reverse (5′-3′) ATGTGGTGGACTTTTCGCTC | |
| Dgat1 | Forward (5′-3′) TAGAAGAGGACGAGGTGCGAG |
| Reverse (5′-3′) GTAGAGACAGCTTTGGCCTTGA | |
| Dhcr7 | Forward (5′-3′) ATGGCTTCGAAATCCCAGCA |
| Reverse (5′-3′) GAACCAGTCCACTTCCCAGG | |
| Fas | Forward (5′-3′) AGTGTCCACCAACAAGCG |
| Reverse (5′-3′) GATGCCGTCAGGTTTCAG | |
| Fsp1 | Forward (5′-3′) CTCTCTTGGTCTGGTCTCAAC |
| Reverse (5′-3′) AACTTGTCACCCTCTTTGCCT | |
| Gpd1 | Forward (5′-3′) CATCACGACCTGCTATGGGG |
| Reverse (5′-3′) GCTGCTCAATGGACTTTCCAG | |
| Gpd2 | Forward (5′-3′) ACTCCGTTCTGGCTGGAGGAT |
| Reverse (5′-3′) CATATGCCAGGCTCACTTGCTTC | |
| Hadha | Forward (5′-3′) TGCTCAAGATGGTGGCGTCC |
| Reverse (5′-3′) AATGCAGCCTCTGGAGCGTAG | |
| Hmgcs1 | Forward (5′-3′) TCCCCTTTGGCTCTTTCACC |
| Reverse (5′-3′) GGGCAACGATTCCCACATCT | |
| Lpin3 | Forward (5′-3′) ACAGCTCATTGTACGTGGCA |
| Reverse (5′-3′) ATCTCCACATGGCTTTCCCG | |
| Plin5 | Forward (5′-3′) GCCATCAGACATGGTGGTGA |
| Reverse (5′-3′) CCTCAGTCATGGGCAGGAAG | |
| Ppara | Forward (5′-3′) TACTGCCGTTTTCACAAGTGC |
| Reverse (5′-3′) AGGTCGTGTTCACAGGTAAGA | |
| Pparb/d | Forward (5′-3′) CGGCAGCCTCAACATGG |
| Reverse (5′-3′) AGATCCGATCGCACTTCTCATAC | |
| Pparg1 | Forward (5′-3′) TGTGAGACCAACAGCCTGAC |
| Reverse (5′-3′) CCGCTTCTTTCAAATCTTGTCTGT | |
| Scd1 | Forward (5′-3′) GCCCACATGCTCCAAGAGAT |
| Reverse (5′-3′) GGGCACTGTCTTCACCTTCT | |
| Srebp1c | Forward (5′-3′) AAGCGCTACCGGTCTTCTATC |
| Reverse (5′-3′) TGTGCACTTCGTAGGGTCAG | |
| Srebp2 | Forward (5′-3′) TGGGCGATGGATGAGAGCAG |
| Reverse (5′-3′) AACTGTAGCATCTCGTCGATGTC |
References
- Angulo, P. Obesity and Nonalcoholic Fatty Liver Disease. Nutr. Rev. 2007, 65, 57–63. [Google Scholar] [CrossRef]
- Lazo, M.; Clark, J. The Epidemiology of Nonalcoholic Fatty Liver Disease: A Global Perspective. Semin. Liver Dis. 2008, 28, 339–350. [Google Scholar] [CrossRef] [PubMed]
- Williams, C.D.; Stengel, J.; Asike, M.I.; Torres, D.M.; Shaw, J.; Contreras, M.; Landt, C.L.; Harrison, S.A. Prevalence of Nonalcoholic Fatty Liver Disease and Nonalcoholic Steatohepatitis Among a Largely Middle-Aged Population Utilizing Ultrasound and Liver Biopsy: A Prospective Study. Gastroenterology 2011, 140, 124–131. [Google Scholar] [CrossRef] [PubMed]
- Wieckowska, A.; Papouchado, B.G.; Li, Z.; Lopez, R.; Zein, N.N.; Feldstein, A.E. Increased Hepatic and Circulating Interleukin-6 Levels in Human Nonalcoholic Steatohepatitis. Am. J. Gastroenterol. 2008, 103, 1372–1379. [Google Scholar] [CrossRef] [PubMed]
- Dowman, J.K.; Tomlinson, J.W.; Newsome, P.N. Systematic review: The diagnosis and staging of non-alcoholic fatty liver disease and non-alcoholic steatohepatitis. Aliment. Pharmacol. Ther. 2010, 33, 525–540. [Google Scholar] [CrossRef]
- McCarthy, E.M.; Rinella, M.E. The Role of Diet and Nutrient Composition in Nonalcoholic Fatty Liver Disease. J. Acad. Nutr. Diet. 2012, 112, 401–409. [Google Scholar] [CrossRef]
- Promrat, K.; Kleiner, D.E.; Niemeier, H.M.; Jackvony, E.; Kearns, M.; Wands, J.R.; Fava, J.L.; Wing, R.R. Randomized controlled trial testing the effects of weight loss on nonalcoholic steatohepatitis. Hepatology 2010, 51, 121–129. [Google Scholar] [CrossRef]
- St George, A.; Bauman, A.; Johnston, A.; Farrell, G.; Chey, T.; George, J. Effect of a lifestyle intervention in patients with abnormal liver enzymes and metabolic risk factors. J. Gastroenterol. Hepatol. 2008, 24, 399–407. [Google Scholar] [CrossRef]
- Haukeland, J.W.; Konopski, Z.; Eggesbø, H.B.; Von Volkmann, H.L.; Raschpichler, G.; Bjøro, K.; Haaland, T.; Løberg, E.M.; Birkeland, K. Metformin in patients with non-alcoholic fatty liver disease: A randomized, controlled trial. Scand. J. Gastroenterol. 2009, 44, 853–860. [Google Scholar] [CrossRef]
- Chen, J.; Montagner, A.; Tan, N.S.; Wahli, W. Insights into the Role of PPARβ/δ in NAFLD. Int. J. Mol. Sci. 2018, 19, 1893. [Google Scholar] [CrossRef]
- Seki, S.; Habu, Y.; Kawamura, T.; Takeda, K.; Dobashi, H.; Ohkawa, T.; Hiraide, H. The liver as a crucial organ in the first line of host defense: The roles of Kupffer cells, natural killer (NK) cells and NK1.1 Ag+ T cells in T helper 1 immune responses. Immunol. Rev. 2000, 174, 35–46. [Google Scholar] [CrossRef] [PubMed]
- Thomson, A.W.; Knolle, P.A. Antigen-presenting cell function in the tolerogenic liver environment. Nat. Rev. Immunol. 2010, 10, 753–766. [Google Scholar] [CrossRef]
- Toth, C.A.; Thomas, D. Liver endocytosis and Kupffer cells. Hepatology 1992, 16, 255–266. [Google Scholar] [CrossRef] [PubMed]
- Lanthier, N.; Molendi-Coste, O.; Horsmans, Y.; Van Rooijen, N.; Cani, P.D.; Leclercq, I.A. Kupffer cell activation is a causal factor for hepatic insulin resistance. Am. J. Physiol. Liver Physiol. 2010, 298, G107–G116. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kinoshita, M.; Uchida, T.; Sato, A.; Nakashima, M.; Nakashima, H.; Shono, S.; Habu, Y.; Miyazaki, H.; Hiroi, S.; Seki, S. Characterization of two F4/80-positive Kupffer cell subsets by their function and phenotype in mice. J. Hepatol. 2010, 53, 903–910. [Google Scholar] [CrossRef] [PubMed]
- Girroir, E.E.; Hollingshead, H.E.; He, P.; Zhu, B.; Perdew, G.H.; Peters, J.M. Quantitative expression patterns of peroxisome proliferator-activated receptor-beta/delta (PPARbeta/delta) protein in mice. Biochem. Biophys. Res. Commun. 2008, 371, 456–461. [Google Scholar]
- Hoekstra, M.; Kruijt, J.K.; Van Eck, M.; Van Berkel, T.J.C. Specific Gene Expression of ATP-binding Cassette Transporters and Nuclear Hormone Receptors in Rat Liver Parenchymal, Endothelial, and Kupffer Cells. J. Boil. Chem. 2003, 278, 25448–25453. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, Y.X.; Lee, C.H.; Tiep, S.; Yu, R.T.; Ham, J.; Kang, H.; Evans, R.M. Peroxisome-proliferator-activated receptor delta activates fat metabolism to prevent obesity. Cell 2003, 113, 159–170. [Google Scholar] [CrossRef]
- Berger, J.; Leibowitz, M.D.; Doebber, T.W.; Elbrecht, A.; Zhang, B.; Zhou, G.; Biswas, C.; A Cullinan, C.; Hayes, N.S.; Li, Y.; et al. Novel peroxisome proliferator-activated receptor (PPAR) gamma and PPARdelta ligands produce distinct biological effects. J. Boil. Chem. 1999, 274, 6718–6725. [Google Scholar] [CrossRef]
- Odegaard, J.I.; Ricardo-Gonzalez, R.R.; Eagle, A.R.; Vats, D.; Morel, C.R.; Goforth, M.H.; Subramanian, V.; Mukundan, L.; Ferrante, A.W.; Chawla, A. Alternative M2 activation of Kupffer cells by PPARdelta ameliorates obesity-induced insulin resistance. Cell Metab. 2008, 7, 496–507. [Google Scholar] [CrossRef]
- Peters, J.M.; Gonzalez, F.J.; Müller, R. Establishing the Role of PPARβ/δ in Carcinogenesis. Trends Endocrinol. Metab. 2015, 26, 595–607. [Google Scholar] [CrossRef]
- Wang, D.; Dubois, R.N. PPARδ and PGE2 signaling pathways communicate and connect inflammation to colorectal cancer. Inflamm. Cell Signal. 2014, 1. [Google Scholar] [CrossRef]
- Vitale, S.G.; Laganà, A.S.; Nigro, A.; La Rosa, V.L.; Rossetti, P.; Rapisarda, A.M.C.; La Vignera, S.; Condorelli, R.A.; Corrado, F.; Buscema, M.; et al. Peroxisome Proliferator-Activated Receptor Modulation during Metabolic Diseases and Cancers: Master and Minions. PPAR Res. 2016, 2016, 1–9. [Google Scholar] [CrossRef] [Green Version]
- Boye, K.; Maelandsmo, G.M. S100A4 and metastasis: A small actor playing many roles. Am. J. Pathol 2010, 176, 528–535. [Google Scholar] [CrossRef]
- Strutz, F.; Okada, H.; Lo, C.W.; Danoff, T.; Carone, R.L.; Tomaszewski, J.E.; Neilson, E.G. Identification and characterization of a fibroblast marker: FSP1. J. Cell Biol 2005, 130, 393–405. [Google Scholar] [CrossRef]
- Lawson, W.E.; Polosukhin, V.V.; Zoia, O.; Stathopoulos, G.T.; Han, W.; Plieth, D.; Loyd, J.E.; Neilson, E.G.; Blackwell, T.S. Characterization of Fibroblast-specific Protein 1 in Pulmonary Fibrosis. Am. J. Respir. Crit. Care Med. 2005, 171, 899–907. [Google Scholar] [CrossRef]
- Schneider, M.; Kostin, S.; Grigorian, M.; Andersen, C.B.; Lukanidin, E.; Lerchehansen, J.; Strøm, C.C.; Aplin, M.; Lyngbæk, S.; Theilade, J.; et al. S100A4 is upregulated in injured myocardium and promotes growth and survival of cardiac myocytes. Cardiovasc. Res. 2007, 75, 40–50. [Google Scholar] [CrossRef] [Green Version]
- Grigorian, M.S.; Tulchinsky, E.M.; Zain, S.; Ebralidze, A.K.; Kramerov, D.A.; Kriajevska, M.V.; Georgiev, G.P.; Lukanidin, E.M. The mts1 gene and control of tumor metastasis. Gene 1993, 135, 229–238. [Google Scholar] [CrossRef]
- Takenaga, K.; Nakamura, Y.; Sakiyama, S. Cellular Localization of pEL98 Protein, An S100-related Calcium Binding Protein, in Fibroblasts and Its Tissue Distribution Analyzed by Monoclonal Antibodies. Cell Struct. Funct. 1994, 19, 133–141. [Google Scholar] [CrossRef]
- Gibbs, F.E.; Barraclough, R.; Platt-Higgins, A.; Rudland, P.S.; Wilkinson, M.C.; Parry, E.W. Immunocytochemical distribution of the calcium-binding protein p9Ka in normal rat tissues: Variation in the cellular location in different tissues. J. Histochem. Cytochem. 1995, 43, 169–180. [Google Scholar] [CrossRef]
- Österreicher, C.H.; Penz-Österreicher, M.; Grivennikov, S.I.; Guma, M.; Koltsova, E.K.; Datz, C.; Sasik, R.; Hardiman, G.; Karin, M.; Brenner, D.A. Fibroblast-specific protein 1 identifies an inflammatory subpopulation of macrophages in the liver. Proc. Natl Acad. Sci. USA 2011, 108, 308–313. [Google Scholar] [CrossRef]
- Chen, L.; Li, J.; Zhang, J.; Dai, C.; Liu, X.; Wang, J.; Gao, Z.; Guo, H.; Wang, R.; Lu, S.; et al. S100A4 promotes liver fibrosis via activation of hepatic stellate cells. J. Hepatol. 2015, 62, 156–164. [Google Scholar] [CrossRef]
- Tan, E.H.P.; Sng, M.K.; How, I.S.B.; Chan, J.S.K.; Chen, J.; Tan, C.K.; Wahli, W.; Tan, N.S. ROS release by PPARβ/δ-null fibroblasts reduces tumor load through epithelial antioxidant response. Oncogene 2018, 37, 2067–2078. [Google Scholar] [CrossRef]
- Sng, M.K.; Chan, J.S.K.; Teo, Z.; Phua, T.; Tan, E.H.P.; Wee, J.W.K.; Koh, N.J.N.; Tan, C.K.; Chen, J.P.; Pal, M.; et al. Selective deletion of PPARβ/δ in fibroblasts causes dermal fibrosis by attenuated LRG1 expression. Cell Discov. 2018, 4, 15. [Google Scholar] [CrossRef]
- Altin, J.G.; Sloan, E.K. The role of CD45 and CD45-associated molecules in T cell activation. Immunol. Cell Boil. 1997, 75, 430–445. [Google Scholar] [CrossRef]
- Rando, G.; Tan, C.K.; Khaled, N.; Montagner, A.; Leuenberger, N.; Bertrand-Michel, J.; Paramalingam, E.; Guillou, H.; Wahli, W. Glucocorticoid receptor-PPARα axis in fetal mouse liver prepares neonates for milk lipid catabolism. eLife 2016, 5, 11853. [Google Scholar] [CrossRef]
- Montagner, A.; Polizzi, A.; Fouché, E.; Ducheix, S.; Lippi, Y.; Lasserre, F.; Barquissau, V.; Régnier, M.; Lukowicz, C.; Benhamed, F.; et al. Liver PPARα is crucial for whole-body fatty acid homeostasis and is protective against NAFLD. Gut 2016, 65, 1202–1214. [Google Scholar] [CrossRef]
- Horton, J.D.; Goldstein, J.L.; Brown, M.S. SREBPs: Activators of the complete program of cholesterol and fatty acid synthesis in the liver. J. Clin. Investig. 2002, 109, 1125–1131. [Google Scholar] [CrossRef]
- Barroso, E.; Rodríguez-Calvo, R.; Serrano-Marco, L.; Astudillo, A.M.; Balsinde, J.; Palomer, X.; Vázquez-Carrera, M. The PPARβ/δ Activator GW501516 Prevents the Down-Regulation of AMPK Caused by a High-Fat Diet in Liver and Amplifies the PGC-1α-Lipin 1-PPARα Pathway Leading to Increased Fatty Acid Oxidation. Endocrinology 2011, 152, 1848–1859. [Google Scholar] [CrossRef]
- Temple, J.L.; Cordero, P.; Li, J.; Nguyen, V.; Oben, J.A. A Guide to Non-Alcoholic Fatty Liver Disease in Childhood and Adolescence. Int. J. Mol. Sci. 2016, 17, 947. [Google Scholar] [CrossRef]
- Schuler, M.; Ali, F.; Chambon, C.; Duteil, D.; Bornert, J.M.; Tardivel, A.; Desvergne, B.; Wahli, W.; Chambon, P.; Metzger, D. PGC1alpha expression is controlled in skeletal muscles by PPARbeta, whose ablation results in fiber-type switching, obesity, and type 2 diabetes. Cell Metab. 2006, 4, 407–414. [Google Scholar] [CrossRef]
- Henneberger, C.; Grantyn, R.; Rothe, T. Rapid genotyping of newborn gene mutant mice. J. Neurosci. Methods 2000, 100, 123–126. [Google Scholar] [CrossRef]








© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chen, J.; Zhuang, Y.; Sng, M.K.; Tan, N.S.; Wahli, W. The Potential of the FSP1cre-Pparb/d−/− Mouse Model for Studying Juvenile NAFLD. Int. J. Mol. Sci. 2019, 20, 5115. https://doi.org/10.3390/ijms20205115
Chen J, Zhuang Y, Sng MK, Tan NS, Wahli W. The Potential of the FSP1cre-Pparb/d−/− Mouse Model for Studying Juvenile NAFLD. International Journal of Molecular Sciences. 2019; 20(20):5115. https://doi.org/10.3390/ijms20205115
Chicago/Turabian StyleChen, Jiapeng, Yan Zhuang, Ming Keat Sng, Nguan Soon Tan, and Walter Wahli. 2019. "The Potential of the FSP1cre-Pparb/d−/− Mouse Model for Studying Juvenile NAFLD" International Journal of Molecular Sciences 20, no. 20: 5115. https://doi.org/10.3390/ijms20205115
APA StyleChen, J., Zhuang, Y., Sng, M. K., Tan, N. S., & Wahli, W. (2019). The Potential of the FSP1cre-Pparb/d−/− Mouse Model for Studying Juvenile NAFLD. International Journal of Molecular Sciences, 20(20), 5115. https://doi.org/10.3390/ijms20205115