Brassinosteroids, the Sixth Class of Phytohormones: A Molecular View from the Discovery to Hormonal Interactions in Plant Development and Stress Adaptation
Abstract
1. Introduction
2. Brassinosteroids: Functions and Signaling Pathway
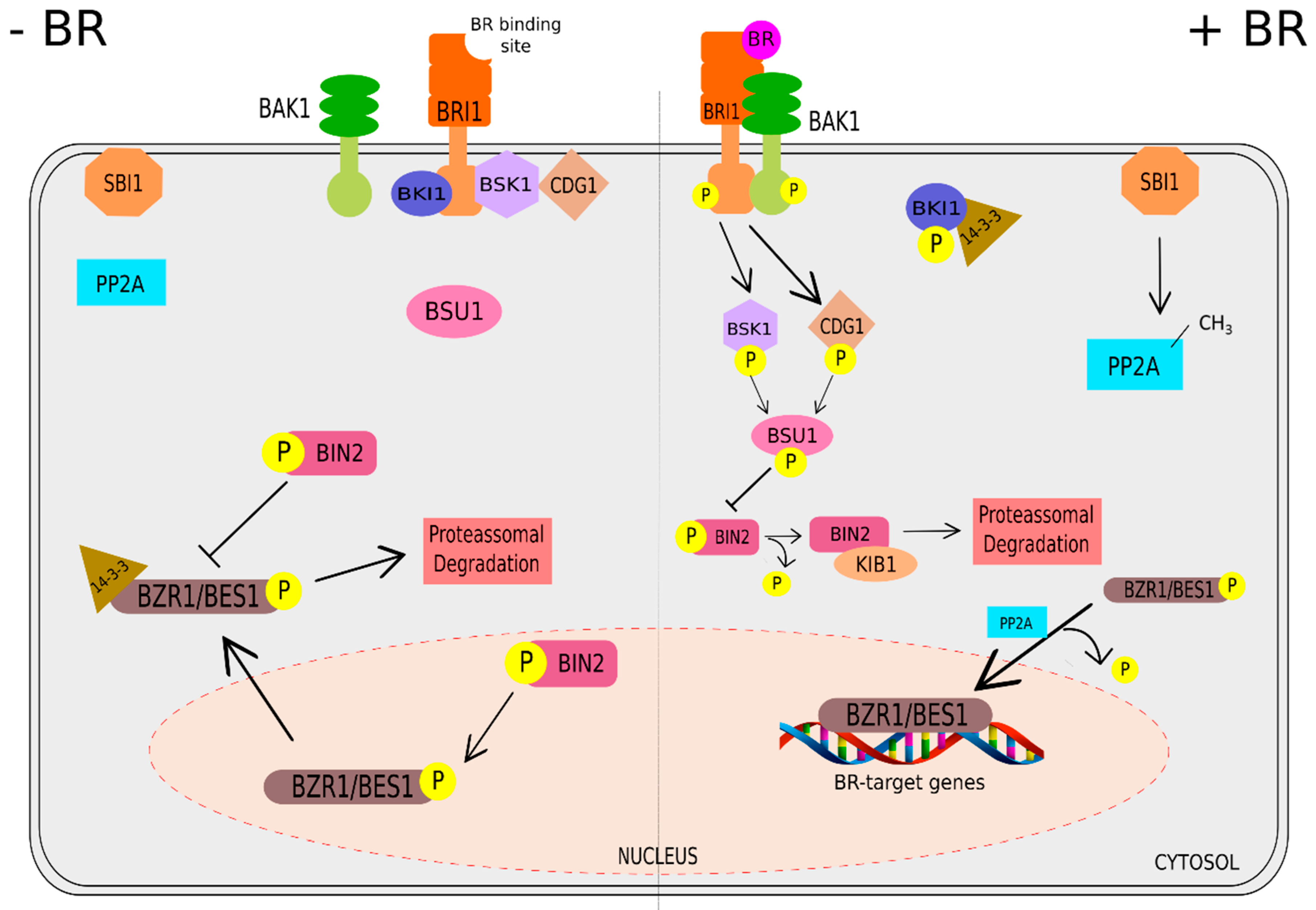
Signaling Pathway
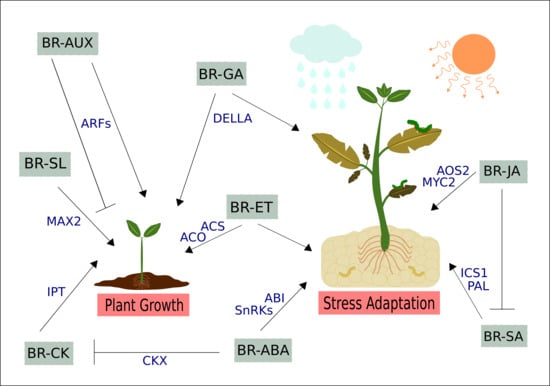
3. Cross-talk between BRs and Other Phytohormones in Plant Growth, Development, and Stress Responses
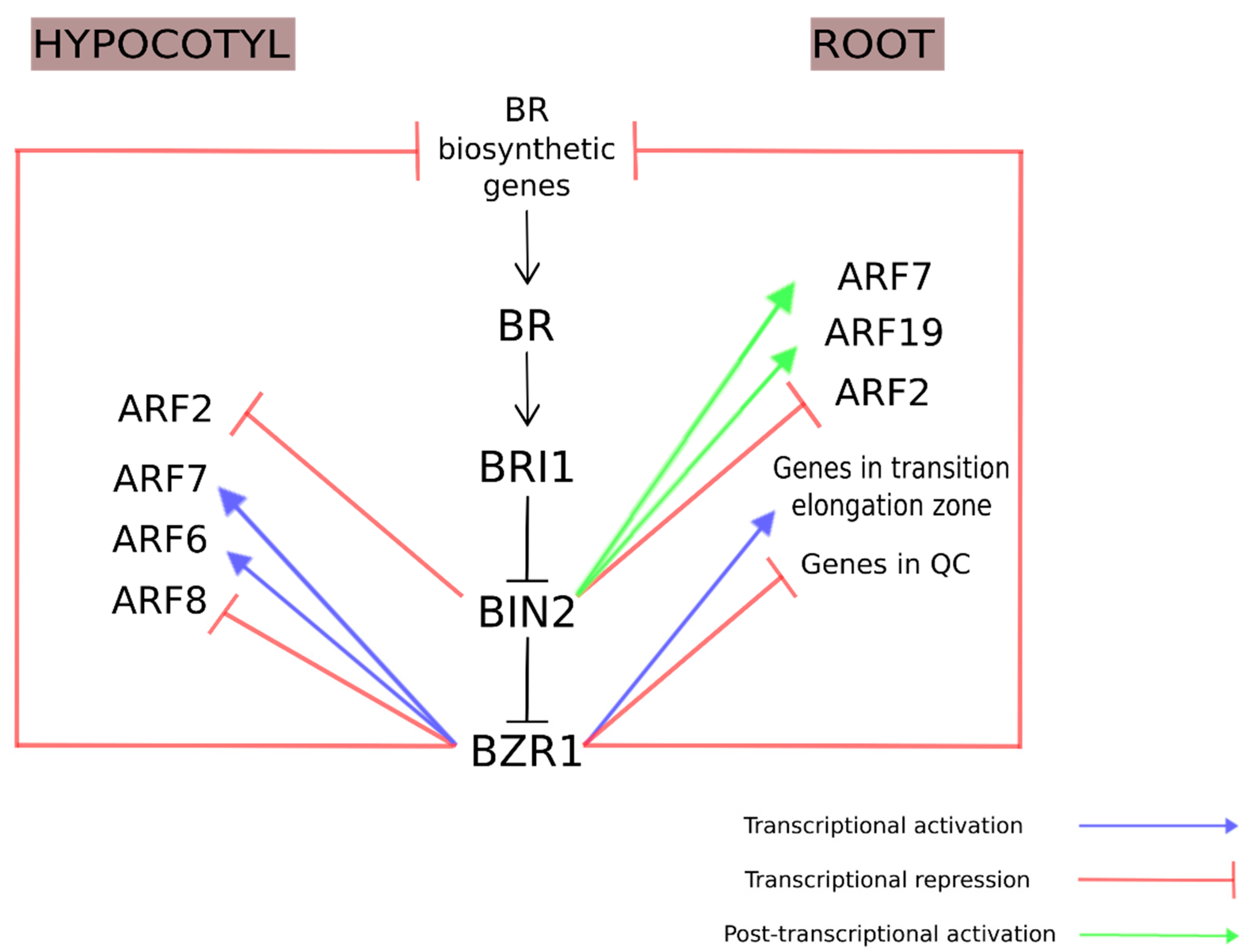
3.1. Brassinosteroids and Auxins
3.2. Brassinosteroids and Gibberellins
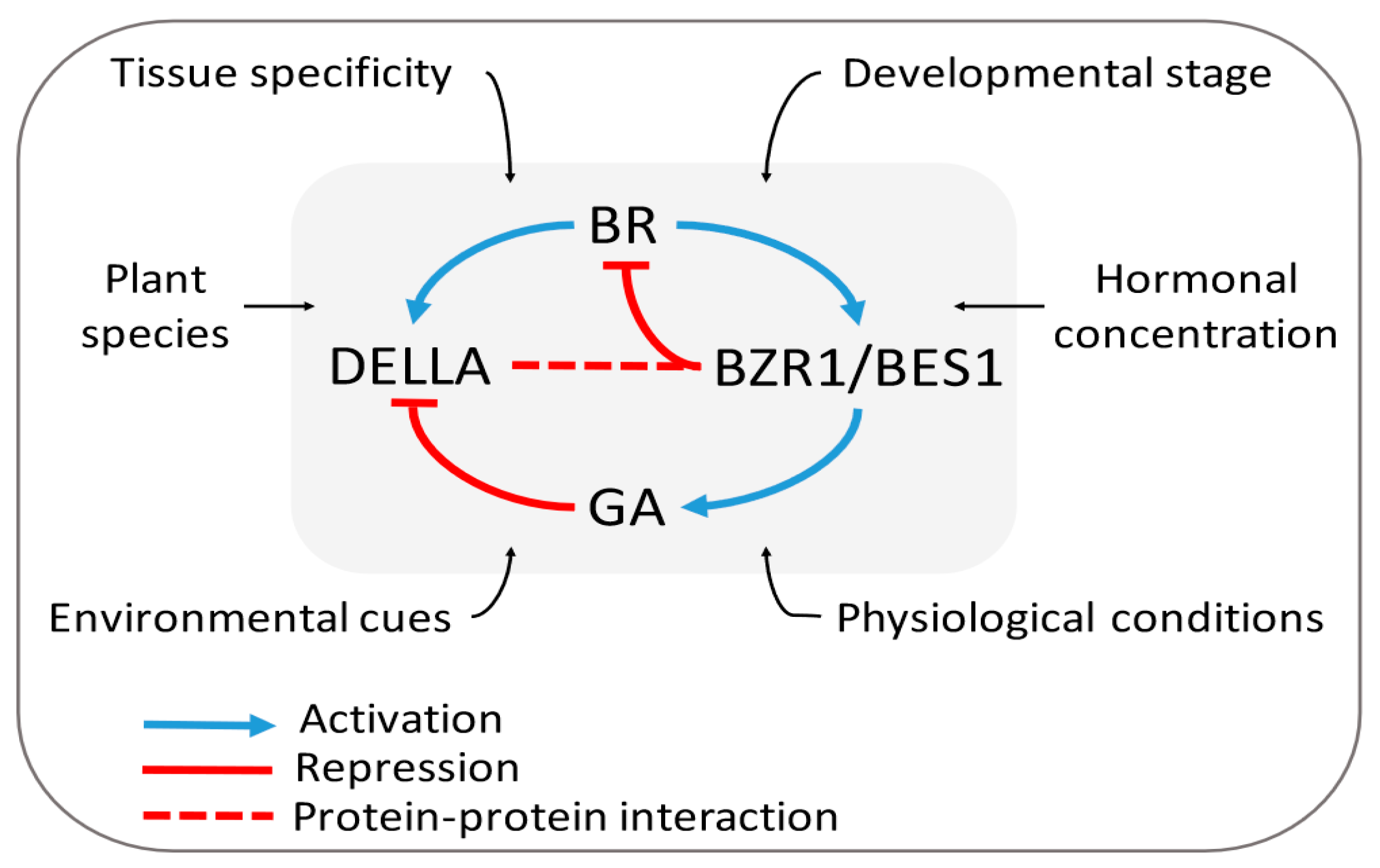
3.2.1. BR–GA Cross-talk: The Signaling Model
3.2.2. The Expanded and Integrated Model
3.2.3. Is the BR–GA Antagonism an Alternate Strategy to Tackle Biotic and Abiotic Stresses?
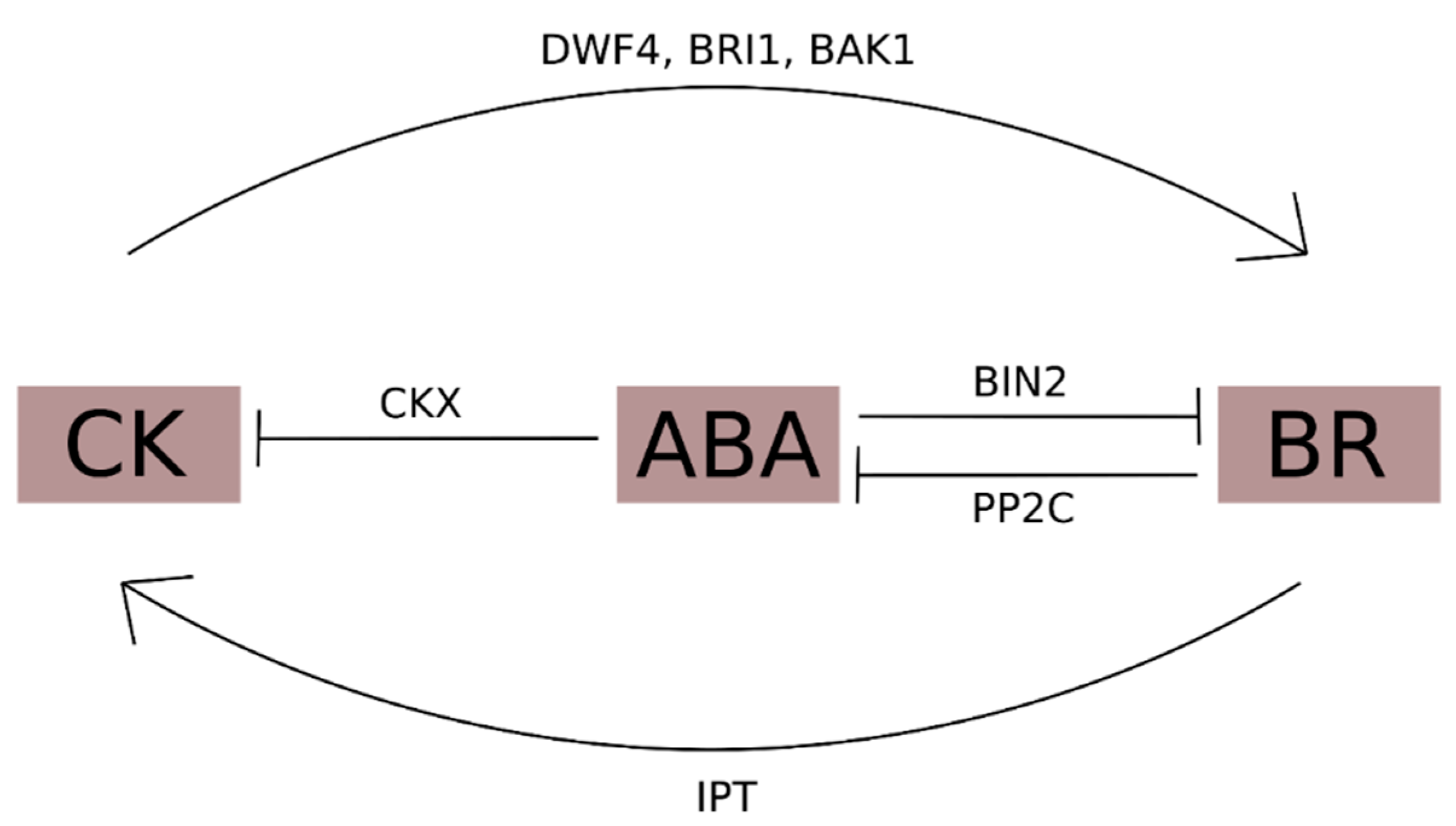
3.3. Brassinosteroids and Cytokinins
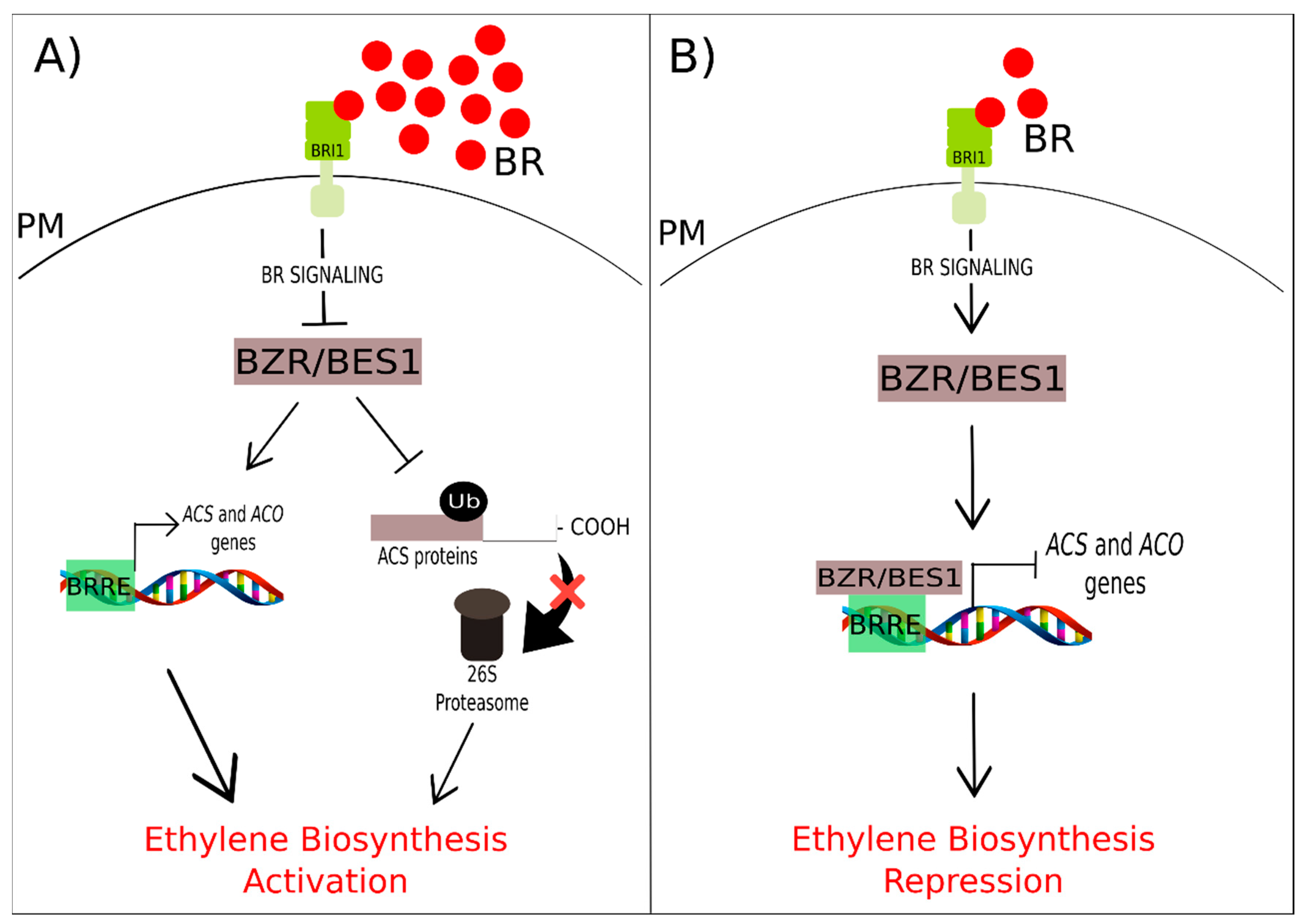
3.4. Brassinosteroids and Ethylene
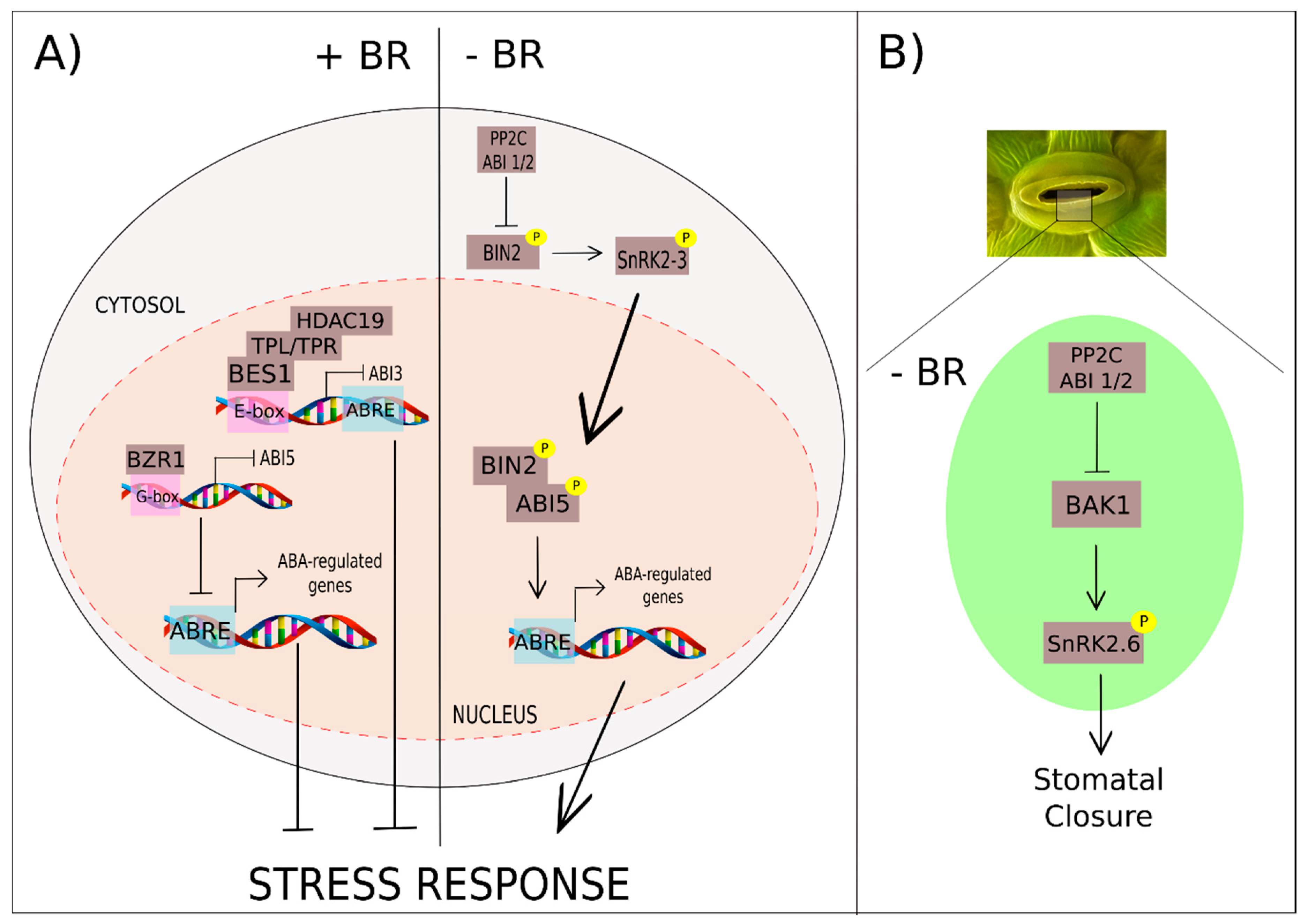
3.5. Brassinosteroids and Abscisic Acid
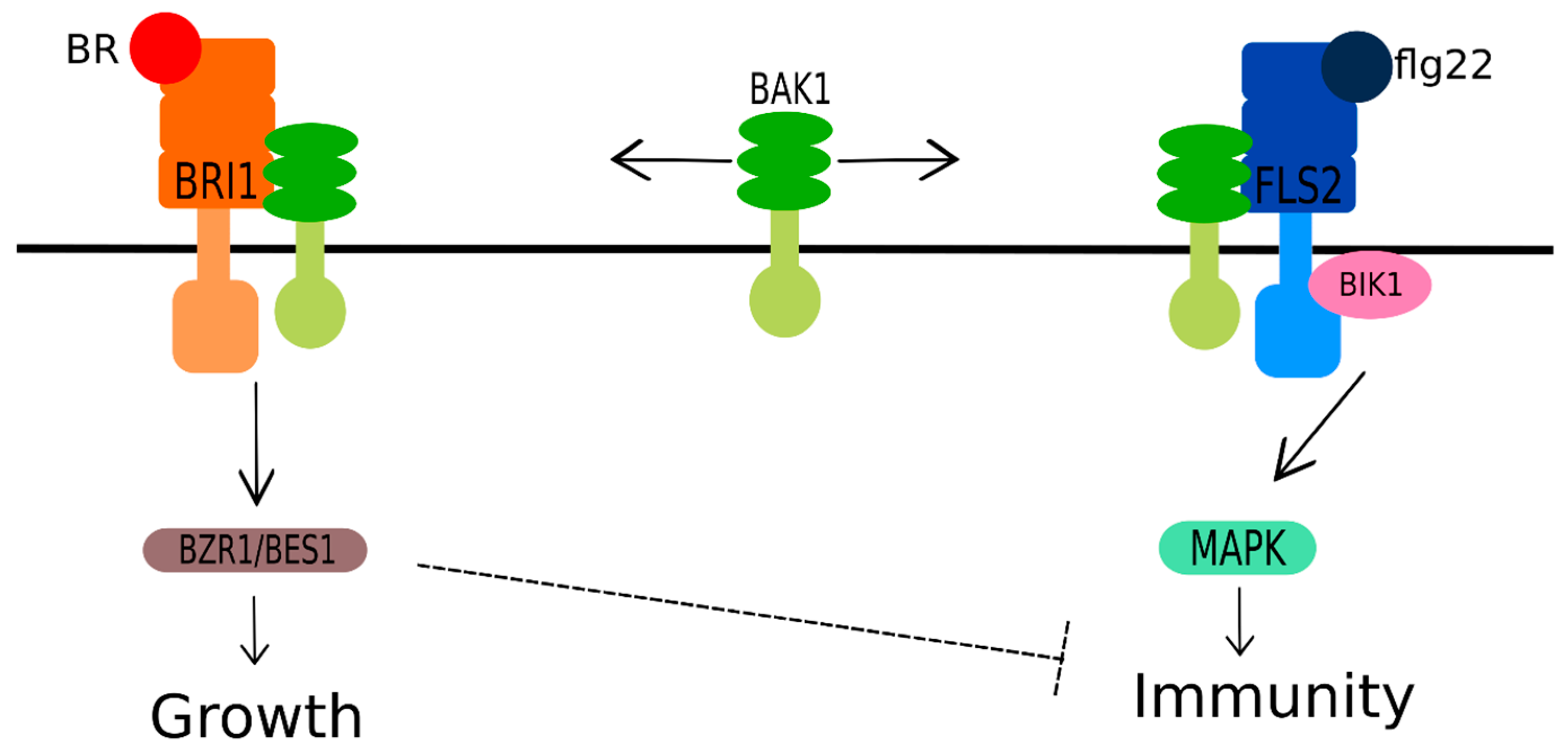
3.6. Brassinosteroids and Jasmonic and Salicylic Acids
3.7. Brassinosteroids and Strigolactones
4. Conclusions and Remarks
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| ABA | Abscisic Acid |
| ABI | ABA Insensitive |
| ACS | ACC-SYNTHASE |
| ACC | 1-aminocyclopropane-1-carboxylicacid |
| ACO | ACC-OXIDASE ABA Insensitive |
| AOS2 | ALLENE OXIDE SYNTHASE 1 |
| ARF | AUXIN RESPONSE FACTOR |
| AUX | Auxin |
| AUX/IAA | AUXIN/INDOLE ACETIC ACID |
| AuxREs | Auxin-responsive DNA Elements |
| BAK1 | BRI1-ASSOCIATED RECEPTOR KINASE 1 |
| BES1/BZR2 | BRI1-EMS SUPPRESSOR 1 |
| BIK1 | BOTRYTIS-INDUCED KINASE 1 |
| BIN2 | BRASSINOSTEROID-INSENSITIVE 2 |
| BL | Brassinolide |
| BPH | Brown Planthopper |
| BRI1 | BRASSINOSTEROID INSENSITIVE 1 |
| BRs | Brassinosteroids |
| BRX | BREVIS RADIX |
| BRZ | Brassinazole |
| BSKs | BR-SIGNALING KINASE 1 |
| BSU1 | BRI1 SUPPRESSOR 1 |
| bZIP | Basic Leucine-Zipper |
| BZR1 | BRASSINAZOLE RESISTANT 1 |
| CDG1 | CONSTITUTIVE DIFFERENTIAL GROWTH 1 |
| ChiP | Chromatin Immunoprecipitation |
| ChiP-seq | Chromatin Immunoprecipitation-sequencing |
| CKs | Cytokinins |
| CKXs | Cytokinin oxidases/dehydrogenases |
| CPD | CONSTITUTIVE PHOTOMORPHISM AND DWARFISM |
| DWF4 | DWARF 4 |
| EGL3 | Enhancer of Glabra 3 |
| ET | Ethylene |
| Fgl22 | Flagellin 22 |
| FLS2 | Flg-sensing 2 |
| GAs | Gibberellins |
| GID1 | GIBBERELLIN INSENSITIVE DWARF1 |
| GL3 | Glabra 3 |
| HDAC19 | HISTONE DEACETYLASE 19 |
| HYD1 | HYDRA 1 |
| IAA | Indole Acetic Acid |
| ICS1 | ISOCHORISMATE SYNTHASE 1 |
| IPT | Isopentenyltransferases |
| JA | Jasmonic Acid |
| KIB1 | KINK SUPPRESSED IN BZR1-1D |
| LOX1 | LIPOXYGENASE 1 |
| LRR | Leucine Rich Repeat |
| MAMPs | Microbial Associated Molecules Patters |
| MAPKs | Mitogen Activated Protein Kinases |
| MAX2 | More Axillary Growth Locus 2 |
| MTI | MAMP-triggered Immunity |
| NPF | NITRATE TRANSPORTER 1/PEPTIDE TRANSPORTER FAMILY |
| OA | Okadaic Acid |
| PAL | PHENYLALANINE AMMONIUM LYASE |
| PAMPs | Pathogen Associated Molecules Patters |
| PAP1 | Anthocyanin Pigment 1 |
| PP2A | PHOSPHATASE 2A |
| PP2AB’ | PP2AB’α and PP2AB’β subunits |
| PP2C | PHOSPHATASE 2C |
| QC | Quiescence Center |
| qRT-PCR | Real-time Quantitative Reverse Transcription-PCR |
| RLK | Receptor-like kinase |
| ROS | Reactive Oxygen Species |
| SA | Salicylic Acid |
| SAM | Shoot Apical Meristem |
| SAM² | S-adenosyl methionine |
| SBI1 | SUPPRESSOR OF BRI1 |
| SLs | Strigolactones |
| SLR1 | SLENDER RICE 1 |
| TIR1/AFB | TRANSPORT INHIBITOR RESPONSE1/AUXIN SIGNALING F-BOX |
| TPL | TOPLESS |
| WT | Wilt-type |
References
- Gustafson, F.G. Parthenocarpy Induced by Pollen Extracts. Am. J. Bot. 1937, 24, 102–107. [Google Scholar] [CrossRef]
- Gustafson, F.G. Inducement of Fruit Development by Growth-Promoting Chemicals. Proc. Natl. Acad. Sci. USA 1936, 22, 628–636. [Google Scholar] [CrossRef] [PubMed]
- Mitchell, J.W.; Whitehead, M.R. Responses of Vegetative Parts of Plants Following Application of Extract of Pollen from Zea mays. Bot. Gaz. 1941, 102, 770–791. [Google Scholar] [CrossRef]
- Mitchell, J.W.; Skraggs, D.P.; Anderson, W.P. Plant Growth-stimulating Hormones in Immature Bean Seeds. Science 1951, 114, 159–161. [Google Scholar] [CrossRef] [PubMed]
- Mitchell, J.W.; Mandava, N.B.; Worley, J.F.; Plimmer, J.R.; Smith, M.V. Brassins—A new family of plant hormones from rape pollen. Nature 1970, 225, 1065–1066. [Google Scholar] [CrossRef] [PubMed]
- Mandava, N.B.; Mitchell, J.W. New plant hormones: Chemical and biological investigations. Indian Agric. 1971, 15, 19–31. [Google Scholar]
- Steffens, G.L. U.S. Department of Agriculture Brassins Projects: 1970–1980. In Brassinosteroids: Chemistry, Bioactivity and Applications; Cutler, H.G., Yokota, T., Adam, G., Eds.; American Chemical Society: Washington, DC, USA, 1991; pp. 2–17. [Google Scholar] [CrossRef]
- Mandava, N.B.; Sidwell, B.A.; Mitchell, J.W.; Worley, J.F. Production of Brassins from Rape Pollen. A Convenient Preparatory Method. Ind. Eng. Chem. Prod. Res. Dev. 1973, 12, 138–139. [Google Scholar] [CrossRef]
- Grove, M.D.; Spencer, G.F.; Rohwedder, W.K.; Mandava, N.; Worley, J.F.; Warthen, J.D.; Steffens, G.L.; Flippen-Anderson, J.L.; Cook, J.C., Jr. Brassinolide, a plant growth-promoting steroid isolated from Brassica napus pollen. Nature 1979, 281, 216–217. [Google Scholar] [CrossRef]
- Fung, S.; Sidall, J.B. Stereoselective Synthesis of Brassinolide: A Plant Growth Promoting Steroidal Lactone. J. Am. Chem. Soc. 1980, 102, 6580–6581. [Google Scholar] [CrossRef]
- Ishiguro, M.; Takatsuto, S.; Morisaki, M.; Ikekawa, N. Synthesis of Brassinolide, a Steroidal Lactone with Plant-growth Promoting Activity. J. Chem. Soc. Chem. Comm. 1980, 20, 962–964. [Google Scholar] [CrossRef]
- Thompson, M.J.; Mandava, N.B.; Flippen-Anderson, J.L.; Worley, J.F.; Dutky, S.R.; Robbins, W.E.; Lusby, W. Synthesis of Brassino Steroids: New Plant-Growth-Promoting Steroids. J. Org. Chem. 1979, 44, 5002–5004. [Google Scholar] [CrossRef]
- Thompson, M.J.; Meudt, W.J.; Mandava, N.B.; Dutky, S.R.; Lusby, W.R.; Spaulding, D.W. Synthesis of brassinosteroids and relationship of structure to plant growth-promoting effects. Steroids 1982, 39, 89–105. [Google Scholar] [CrossRef]
- Thompson, M.J.; Mandava, N.B.; Meudt, W.J.; Lusby, W.R.; Spaulding, D.W. Synthesis and biological activity of brassinolide and its 22β,23β-isomer: Novel plant growth-promoting steroids. Steroids 1981, 38, 567–580. [Google Scholar] [CrossRef]
- Sakakibara, M.; Mori, K. Facile synthesis of (22R,23R)-homobrassinolide. Agric. Biol. Chem. 1982, 46, 2769–2779. [Google Scholar] [CrossRef]
- Takatsuto, S.; Ying, B.; Morisaki, M.; Ikekawa, N. Microanalysis of brassinolide and its analogs by gas chromatography and gas chromatography-mass spectrometry. J. Chromatogr. A 1982, 239, 233–241. [Google Scholar] [CrossRef]
- Takatsuto, S. Brassinosteroids: Distribution in plants, bioassays and microanalysis by gas chromatography-mass spectrometry. J. Chromatogr. A 1994, 658, 3–15. [Google Scholar] [CrossRef]
- Gamoh, K.; Takatsuto, S. Liquid chromatographic assay of brassinosteroids in plants. J. Chromatogr. A 1994, 658, 17–25. [Google Scholar] [CrossRef]
- Mandava, N.B. Plant growth-promoting brassinosteroids. Annu. Rev. Plant Physiol. Plant Mol. Biol. 1988, 39, 23–52. [Google Scholar] [CrossRef]
- Cutler, H.G.; Yokota, T.; Adam, G. Brassinosteroids: Chemistry, Bioactivity and Applications; American Chemical Society: Washington, DC, USA, 1991; ISBN 0-8412-2126-X. [Google Scholar]
- Khripach, V.A.; Zhabinskii, V.N.; de Groot, A.E. Brassinosteroids—A New Class of Plant Hormones, 1st ed.; Academic Press: San Diego, CA, USA, 1999; ISBN 0-12-406360-8. [Google Scholar]
- Sakurai, A.; Yokota, T.; Clouse, S.D. Brassinosteroids-Steroidal Plant Hormones, 1st ed.; Springer: Tokyo, Japan, 1999; ISBN 978-4-431-70214-6. [Google Scholar]
- Hayat, S.; Ahmad, A. Brassinosteroids: Bioactivity and Crop Productivity, 1st ed.; Kluwer Academic: Dordrecht, The Netherlands, 2003; ISBN 1-4020-1710-3. [Google Scholar]
- Pereira-Netto, A.B. Brassinosteroids Practical Applications in Agriculture and Human Health, 1st ed.; Bentham Science: Sharjah, UAE, 2012; ISBN 978-1-60805-298-1. [Google Scholar]
- Zullo, M.A.T. Brassinosteroids and Related Compounds, 1st ed.; Lambert Academic Publishing: Beau Bassin, Mauritius, 2018; ISBN 978-3-330-34627-7. [Google Scholar]
- Zullo, M.A.T.; Adam, G. Brassinosteroid phytohormones-Structure, bioactivity and applications. Braz. J. Plant Physiol. 2002, 14, 143–181. [Google Scholar] [CrossRef]
- Mitchell, J.W.; Gregory, L.E. Enhancement of overall plant growth, a new response to brassins. Nat. New Biol. 1972, 239, 253–254. [Google Scholar] [CrossRef]
- Gregory, L.E. Acceleration of Plant Growth through Seed Treatment with Brassins. Am. J. Bot. 1981, 68, 586–588. [Google Scholar] [CrossRef]
- Mitchell, J.W.; Mandava, N.B.; Worley, J.F.; Drowne, M.E. Fatty Hormones in Pollen and Immature Seeds of Bean. J. Agric. Food Chem. 1971, 19, 391–393. [Google Scholar] [CrossRef]
- Yopp, J.H.; Colclasure, G.C.; Mandava, N.B. Effects of Brassin-Complex on Auxin and Gibberellin Mediated Events in the Morphogenesis of the Etiolated Bean Hypocotyl. Physiol. Plant. 1979, 46, 247–254. [Google Scholar] [CrossRef]
- Krizek, D.T.; Worley, L.J.F. The influence of spectral quality on the internodal response of intact bean plants to brassins. Physiol. Plant. 1981, 51, 259–264. [Google Scholar] [CrossRef]
- Yopp, J.H.; Mandava, N.B.; Sasse, J.M. Brassinolide, a growth-promoting steroidal lactone. I. Activity in selected auxin bioassays. Physiol. Plant. 1981, 53, 445–452. [Google Scholar] [CrossRef]
- Yopp, J.H.; Mandava, N.B.; Thompson, M.J.; Sasse, J.M. Activity of brassinosteroid in selected bioassays in combination with chemicals known to synergize or retard responses to auxin and gibberellin [Dwarf bean, sunflower, adsuki bean, mung bean]. Proc. Plant Growth Regul. Soc. Am. 1981, 8, 138–145. [Google Scholar]
- Mandava, N.B.; Sasse, J.M.; Yopp, J.H. Brassinolide, a growth-promoting steroidal lactone: II. Activity in selected gibberellin and cytokinin bioassays. Physiol. Plant. 1981, 53, 453–461. [Google Scholar] [CrossRef]
- Gregory, L.E.; Mandava, N.B. The activity and interaction of brassinolide and gibberellic acid in mung bean epicotyls. Physiol. Plant. 1982, 54, 239–243. [Google Scholar] [CrossRef]
- Schlagnhaufer, C.; Arteca, R.N.; Yopp, J.H. A brassinosteroid-cytokinin interaction on ethylene production by etiolated mung bean segments. Physiol. Plant. 1984, 60, 347–350. [Google Scholar] [CrossRef]
- Schlagnhaufer, C.; Arteca, R.N.; Yopp, J.H. Evidence that brassinosteroid stimulates auxin-induced ethylene synthesis in mung bean hypocotyls between S-adenosylmethionine and 1-aminocyclopropane-1-carboxylic acid. Physiol. Plant. 1984, 61, 555–558. [Google Scholar] [CrossRef]
- Cohen, J.D.; Meudt, W.J. Investigations on the mechanism of the brassinosteroid response. 1. Indole-3-acetic-acid metabolism and transport. Plant Physiol. 1983, 72, 691–694. [Google Scholar] [CrossRef] [PubMed]
- Meudt, W.J.; Thompson, M.J.; Bennett, H.W. Investigations on the mechanism of the brassinosteroid response. III. Techniques for potential enhancement of crop production. Proc. Annu. Meet. Plant Growth Regul. Soc. Am. 1983, 10, 312–318. [Google Scholar]
- Meudt, W.J.; Thompson, M.J. Investigations on the mechanism of the brassinosteroid response. II. A modulation of auxin action. Proc. Annu. Meet. Plant Growth Regul. Soc. Am. 1983, 10, 306–311. [Google Scholar]
- Sasse, J.M. The place of brassinolide in the sequential response to plant growth regulators in elongating tissue. Physiol. Plant. 1985, 63, 303–308. [Google Scholar] [CrossRef]
- Hewitt, F.R.; Hough, T.; O’Neill, P.; Sasse, J.M.; Williams, E.G.; Rowan, K.S. Effect of brassinolide and other growth regulators on the germination and growth of pollen tubes of Prunus avium using a multiple hanging-drop assay. Aust. J. Plant Physiol. 1985, 12, 201–211. [Google Scholar] [CrossRef]
- Clouse, S.D.; Zurek, D. Molecular analysis of brassinolide action in plant growth and development. ACS Symp. Ser. Am. Chem. Soc. 1991, 122–140. [Google Scholar] [CrossRef]
- Nomura, T.; Nakayama, M.; Reid, J.B.; Takeuchi, Y.; Yokota, T. Blockage of Brassinosteroid Biosynthesis and Sensitivity Causes Dwarfism in Garden Pea. Plant Physiol. 1997, 113, 31–37. [Google Scholar] [CrossRef]
- Li, J.; Chory, J. A Putative Leucine-Rich Repeat Receptor Kinase Involved in Brassinosteroid Signal Transduction. Cell. 1997, 90, 929–938. [Google Scholar] [CrossRef]
- Asami, T.; Min, Y.K.; Nagata, N.; Yamagishi, K.; Takatsuto, S.; Fujioka, S.; Murofushi, N.; Yamaguchi, I.; Yoshida, S. Characterization of brassinazole, a triazole-type brassinosteroid biosynthesis inhibitor. Plant Physiol. 2000, 123, 93–99. [Google Scholar] [CrossRef]
- Yoshida, S. Regulators of Plant Signal Transduction for Healthy Growth. J. Pestic. Sci. 2000, 25, 441–445. [Google Scholar] [CrossRef]
- Hothorn, M.; Belkhadir, Y.; Dreux, M.; Dabi, T.; Noel, J.P.; Wilson, I.A.; Chory, J. Structural basis of steroid hormone perception by the receptor kinase BRI1. Nature 2011, 474, 467–471. [Google Scholar] [CrossRef] [PubMed]
- She, J.; Han, Z.; Kim, T.-W.; Wang, J.; Cheng, W.; Chang, J.; Shi, S.; Wang, J.; Yang, M.; Wang, Z.-Y.; et al. Structural insight into brassinosteroid perception by BRI1. Nature 2011, 474, 472–476. [Google Scholar] [CrossRef]
- Clouse, S.D. A History of Brassinosteroid Research from 1970 through 2005: Thirty-Five Years of Phytochemistry, Physiology, Genes, and Mutants. J. Plant Growth Regul. 2015, 34, 828–844. [Google Scholar] [CrossRef]
- Bajguz, A. Metabolism of brassinosteroids in plants. Plant Physiol. Biochem. 2007, 45, 95–107. [Google Scholar] [CrossRef]
- Clouse, S.D. Brassinosteroid Signal Transduction: From Receptor Kinase Activation to Transcriptional Networks Regulating Plant Development. Plant Cell 2011, 23, 1219–1230. [Google Scholar] [CrossRef] [PubMed]
- Vriet, C.; Russinova, E.; Reuzeau, C. Boosting Crop Yields with Plant Steroids. Plant Cell 2012, 24, 842–857. [Google Scholar] [CrossRef] [PubMed]
- Verhoef, N.; Yokota, T.; Shibata, K.; De Boer, G.J.; Gerats, T.; Vandenbussche, M.; Koes, R.; Souer, E. Brassinosteroid biosynthesis and signalling in Petunia hybrida. J. Exp. Bot. 2013, 64, 2435–2448. [Google Scholar] [CrossRef]
- Sun, Y.; Fan, X.Y.; Cao, D.M.; Tang, W.; He, K.; Zhu, J.Y.; He, J.X.; Bai, M.Y.; Zhu, S.; Oh, E.; et al. Integration of Brassinosteroid Signal Transduction with the Transcription Network for Plant Growth Regulation in Arabidopsis. Dev. Cell 2010, 19, 765–777. [Google Scholar] [CrossRef]
- Li, J.; Jin, H. Regulation of brassinosteroid signaling. Trends Plant Sci. 2007, 12, 37–41. [Google Scholar] [CrossRef]
- Kim, T.W.; Wang, Z.Y. Brassinosteroid Signal Transduction from Receptor Kinases to Transcription Factors. Annu. Rev. Plant Biol. 2009, 61, 681–704. [Google Scholar] [CrossRef]
- Yang, C.J.; Zhang, C.; Lu, Y.N.; Jin, J.Q.; Wang, X.L. The mechanisms of brassinosteroids’ action: From signal transduction to plant development. Mol. Plant 2011, 4, 588–600. [Google Scholar] [CrossRef] [PubMed]
- Shiu, S.; Karlowski, W.M.; Pan, R.; Tzeng, Y.; Mayer, K.F.X.; Li, W. Comparative Analysis of the Receptor-Like Kinase Family in Arabidopsis and Rice. Plant Cell 2004, 16, 1220–1234. [Google Scholar] [CrossRef]
- Belkhadir, Y.; Chory, J. Brassinosteroid signaling: A paradigm for steroid hormone signaling from the cell surface. Science 2006, 314, 1410–1411. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Wen, J.; Lease, K.A.; Doke, J.T.; Tax, F.E.; Walker, J.C. BAK1, an Arabidopsis LRR receptor-like protein kinase, interacts with BRI1 and modulates brassinosteroid signaling. Cell 2002, 110, 213–222. [Google Scholar] [CrossRef]
- Li, J.; Nam, K.H. Regulation of Brassinosteroid Signaling by a GSK3/SHAGGY-like Kinase. Science 2002, 295, 1299–1301. [Google Scholar] [CrossRef]
- Russinova, E.; Borst, J.-W.; Kwaaitaal, M.; Caño-Delgado, A.; Yin, Y.; Chory, J.; De Vries, S.C. Heterodimerization and Endocytosis of Arabidopsis Brassinosteroid Receptors BRI1 and AtSERK3 (BAK1). Plant Cell Online 2004, 16, 3216–3229. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Goshe, M.B.; Soderblom, E.J.; Phinney, B.S.; Kuchar, J.A.; Li, J.; Asami, T.; Yoshida, S.; Huber, S.C.; Clouse, S.D. Identification and functional analysis of in vivo phosphorylation sites of the 723 Arabidopsis BRASSINOSTEROID-INSENSITIVE1 receptor kinase. Plant Cell 2005, 17, 1685–1703. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Kota, U.; He, K.; Blackburn, K.; Li, J.; Goshe, M.B.; Huber, S.C.; Clouse, S.D. Sequential Transphosphorylation of the BRI1/BAK1 Receptor Kinase Complex Impacts Early Events in Brassinosteroid Signaling. Dev. Cell 2008, 15, 220–235. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Chory, J. Brassinosteroids Regulate Dissociation of BKI1, a Negative Regulator of BRI1 Signaling, from the Plasma Membrane. Science 2006, 313, 1118–1122. [Google Scholar] [CrossRef]
- Jaillais, Y.; Hothorn, M.; Belkhadir, Y.; Jaillais, Y.; Hothorn, M.; Belkhadir, Y.; Dabi, T.; Nimchuk, Z.L.; Meyerowitz, E.M.; Chory, J. Tyrosine phosphorylation controls brassinosteroid receptor activation by triggering membrane release of its kinase inhibitor service Tyrosine phosphorylation controls brassinosteroid receptor activation by triggering membrane release of its kinase inhibit. Genes Dev. 2011, 25, 232–237. [Google Scholar] [CrossRef] [PubMed]
- Bai, M.-Y.; Zhang, L.-Y.; Gampala, S.S.; Zhu, S.-W.; Song, W.-Y.; Chong, K.; Wang, Z.-Y. Functions of OsBZR1 and 14-3-3 proteins in brassinosteroid signaling in rice. Proc. Natl. Acad. Sci. USA 2007, 104, 13839–13844. [Google Scholar] [CrossRef] [PubMed]
- Gampala, S.S.; Kim, T.W.; He, J.X.; Tang, W.; Deng, Z.; Bai, M.Y.; Guan, S.; Lalonde, S.; Sun, Y.; Gendron, J.M.; et al. An Essential Role for 14-3-3 Proteins in Brassinosteroid Signal Transduction in Arabidopsis. Dev. Cell 2007, 13, 177–189. [Google Scholar] [CrossRef]
- Ryu, H.; Kim, K.; Cho, H.; Park, J.; Choe, S.; Hwang, I. Nucleocytoplasmic Shuttling of BZR1 Mediated by Phosphorylation Is Essential in Arabidopsis Brassinosteroid Signaling. Plant Cell Online 2007, 19, 2749–2762. [Google Scholar] [CrossRef] [PubMed]
- Ryu, H.; Cho, H.; Kim, K.; Hwang, I. Phosphorylation dependent nucleocytoplasmic shuttling of BES1 is a key regulatory event in brassinosteroid signaling. Mol. Cells 2010, 29, 283–290. [Google Scholar] [CrossRef]
- Ryu, H.; Kim, K.; Cho, H.; Hwang, I. Predominant actions of cytosolic BSU1 and nuclear BIN2 regulate subcellular localization of bes1 in brassinosteroid signaling. Mol. Cells 2010, 29, 291–296. [Google Scholar] [CrossRef]
- Tang, W.; Kim, T.-W.; Oses-prieto, J.A.; Sun, Y.; Deng, Z.; Zhu, S.; Wang, R.; Burlingame, A.L.; Wang, Z.-Y. Brassinosteroid-Signaling Kinases (BSKs) mediate signal transduction from the receptor kinase BRI1 in Arabidopsis. Science 2008, 321, 557–560. [Google Scholar] [CrossRef] [PubMed]
- Kim, T.W.; Guan, S.; Burlingame, A.L.; Wang, Z.Y. The CDG1 Kinase Mediates Brassinosteroid Signal Transduction from BRI1 Receptor Kinase to BSU1 Phosphatase and GSK3-like Kinase BIN2. Mol. Cell 2011, 43, 561–571. [Google Scholar] [CrossRef] [PubMed]
- Mora-Garcia, S.; Vert, G.; Yin, Y.H.; Cano-Delgado, A.; Cheong, H.; Chory, J. Nuclear protein phosphatases with Kelch-repeat domains modulate the response to bras sino steroids in Arabidopsis. Genes Dev. 2004, 18, 448–460. [Google Scholar] [CrossRef]
- Zhu, J.Y.; Li, Y.; Cao, D.M.; Yang, H.; Oh, E.; Bi, Y.; Zhu, S.; Wang, Z.Y. The F-box Protein KIB1 Mediates Brassinosteroid-Induced Inactivation and Degradation of GSK3-like Kinases in Arabidopsis. Mol. Cell 2017, 66, 648–657. [Google Scholar] [CrossRef]
- Wang, Z.Y.; Nakano, T.; Gendron, J.; He, J.; Chen, M.; Vafeados, D.; Yang, Y.; Fujioka, S.; Yoshida, S.; Asami, T.; et al. Nuclear-localized BZR1 mediates brassinosteroid-induced growth and feedback suppression of brassinosteroid biosynthesis. Dev. Cell 2002, 2, 505–513. [Google Scholar] [CrossRef]
- Wu, G.; Wang, X.; Li, X.; Kamiya, Y.; Otegui, M.S. Methylation of a Phosphatase Specifies Dephosphorylation and Degradation of Activated Brassinosteroid Receptors. Sci. Signal. 2011, 4, 1–25. [Google Scholar] [CrossRef]
- Wei, Z.; Li, J. Brassinosteroids Regulate Root Growth, Development, and Symbiosis. Mol. Plant 2015, 9, 86–100. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y. Auxin Biosynthesis and Its Role in Plant Development. Annu. Rev. Plant Biol. 2010, 61, 49–64. [Google Scholar] [CrossRef]
- Nemhauser, J.L.; Mockler, T.C.; Chory, J. Interdependency of brassinosteroid and auxin signaling in Arabidopsis. PLoS Biol. 2004, 2, 1460–1471. [Google Scholar] [CrossRef] [PubMed]
- Vert, G.; Walcher, C.L.; Chory, J.; Nemhauser, J.L. Integration of auxin and brassinosteroid pathways by Auxin Response Factor 2. Proc. Natl. Acad. Sci. USA 2008, 105, 9829–9834. [Google Scholar] [CrossRef] [PubMed]
- Ljung, K.; Bhalerao, R.P. Sites and homeostatic control of auxin biosynthesis in Arabidopsis during vegetative growth. Plant J. 2001, 28, 465–474. [Google Scholar] [CrossRef] [PubMed]
- Ljung, K.; Hull, A.K.; Celenza, J.; Yamada, M.; Estelle, M.; Normanly, J. Sites and Regulation of Auxin Biosynthesis in Arabidopsis Roots. Plant Cell 2005, 17, 1090–1104. [Google Scholar] [CrossRef] [PubMed]
- Tian, H.; Lv, B.; Ding, T.; Bai, M.; Ding, Z. Auxin-BR Interaction Regulates Plant Growth and Development. Front. Plant Sci. 2018, 8, 2256. [Google Scholar] [CrossRef]
- Wang, D.; Pei, K.; Fu, Y.; Sun, Z.; Li, S.; Liu, H.; Tang, K.; Han, B.; Tao, Y. Genome-wide analysis of the auxin response factors (ARF) gene family in rice (Oryza sativa). Gene 2007, 394, 13–24. [Google Scholar] [CrossRef]
- Goda, H.; Sawa, S.; Asami, T.; Fujioka, S.; Shimada, Y.; Yoshida, S. Comprehensive Comparison of 785 Auxin-Regulated and Brassinosteroid-Regulated Genes in Arabidopsis. Plant Physiol. 2004, 134, 1555–1573. [Google Scholar] [CrossRef]
- Bargmann, B.O.R.; Vanneste, S.; Krouk, G.; Nawy, T.; Efroni, I.; Shani, E.; Choe, G.; Friml, J.; Bergmann, D.C.; Estelle, M.; et al. A map of cell type-specific auxin responses. Mol. Syst. Biol. 2013, 9. [Google Scholar] [CrossRef]
- Jung, J.; Lee, M.; Park, C. A Transcriptional Feedback Loop Modulating Signaling Crosstalks between Auxin and Brassinosteroid in Arabidopsis. Mol. Cells 2010, 449–456. [Google Scholar] [CrossRef] [PubMed]
- Tian, C.; Muto, H.; Higuchi, K.; Matamura, T.; Tatematsu, K.; Koshiba, T. Disruption and overexpression of auxin response factor 8 gene of Arabidopsis affect hypocotyl elongation and root growth habit, indicating its possible involvement in auxin homeostasis in light condition. Plant J. 2004, 3, 333–343. [Google Scholar] [CrossRef]
- Oh, E.; Zhu, J.; Bai, M.; Arenhart, R.A.; Sun, Y.; Wang, Z. Cell elongation is regulated through a central circuit of interacting transcription factors in the Arabidopsis hypocotyl. eLife 2014, 3, e03031. [Google Scholar] [CrossRef] [PubMed]
- Liu, K.; Li, Y.; Chen, X.; Li, L.; Liu, K.; Zhao, H.; Wang, Y.; Han, S. ERF72 interacts with ARF6 and BZR1 to regulate hypocotyl elongation in Arabidopsis. J. Exp. Bot. 2018, 69, 3933–3947. [Google Scholar] [CrossRef] [PubMed]
- Zhou, X.Y.; Song, L.; Xue, H.W. Brassinosteroids regulate the differential growth of Arabidopsis hypocotyls through auxin signaling components IAA19 and ARF7. Mol. Plant 2013, 6, 887–904. [Google Scholar] [CrossRef]
- Zhang, Y.; Liu, Z.; Wang, J.; Chen, Y.; Bi, Y.; He, J. Brassinosteroid is required for sugar promotion of hypocotyl elongation in Arabidopsis in darkness. Planta 2015, 242, 881–893. [Google Scholar] [CrossRef] [PubMed]
- Cho, H.; Ryu, H.; Rho, S.; Hill, K.; Smith, S.; Audenaert, D.; Park, J.; Han, S.; Beeckman, T.; Bennett, M.J.; et al. A secreted peptide acts on BIN2-mediated phosphorylation of ARFs to potentiate auxin response during lateral root development. Nat. Cell Biol. 2014, 16, 66–76. [Google Scholar] [CrossRef]
- Clouse, S.D.; Langford, M.; McMorris, T.C. A Brassinosteroid-Insensitive Mutant in Arabidopsis thaliana Exhibits Multiple Defects in Growth and Development. Plant Physiol. 1996, 111, 671–678. [Google Scholar] [CrossRef]
- Chaiwanon, J.; Wang, Z.Y. Spatiotemporal brassinosteroid signaling and antagonism with auxin pattern stem cell dynamics in Arabidopsis roots. Curr. Biol. 2015, 25, 1031–1042. [Google Scholar] [CrossRef]
- González-García, M.-P.; Vilarrasa-Blasi, J.; Zhiponova, M.; Divol, F.; Mora-García, A.; Russinova, E.; Caño-Delgado, A. Brassinosteroids control meristem size by promoting cell cycle progression in Arabidopsis roots. Stem Cells Dev. 2011, 138, 849–859. [Google Scholar] [CrossRef] [PubMed]
- Yin, Y.; Wang, Z.Y.; Mora-Garcia, S.; Li, J.; Yoshida, S.; Asami, T.; Chory, J. BES1 accumulates in the nucleus in response to brassinosteroids to regulate gene expression and promote stem elongation. Cell 2002, 109, 181–191. [Google Scholar] [CrossRef]
- Walcher, C.L.; Nemhauser, J.L. Bipartite promoter element required for auxin response. Plant Physiol. 2012, 158, 273–282. [Google Scholar] [CrossRef] [PubMed]
- Paponov, I.A.; Teale, W.D.; Trebar, M.; Blilou, I.; Palme, K. The PIN auxin efflux facilitators: Evolutionary and functional perspectives. Trends Plant Sci. 2005, 10, 170–177. [Google Scholar] [CrossRef] [PubMed]
- Mouchel, C.F.; Osmont, K.S.; Hardtke, C.S. BRX mediates feedback between brassinosteroid levels and auxin signalling in root growth. Nature 2006, 443, 458–461. [Google Scholar] [CrossRef] [PubMed]
- Depuydt, S.; Hardtke, C.S. Hormone Signalling Cross-talk in Plant Growth Regulation. Curr. Biol. 2011, 21, R365–R373. [Google Scholar] [CrossRef] [PubMed]
- Binenbaum, J.; Weinstain, R.; Shani, E. Gibberellin Localization and Transport in Plants. Trends Plant Sci. 2018, 23, 410–421. [Google Scholar] [CrossRef] [PubMed]
- Hedden, P.; Sponsel, V. A Century of Gibberellin Research. J. Plant Growth Regul. 2015, 34, 740–760. [Google Scholar] [CrossRef] [PubMed]
- Yamaguchi, S. Gibberellin Metabolism and its Regulation. Annu. Rev. Plant Biol. 2008, 59, 225–251. [Google Scholar] [CrossRef] [PubMed]
- Tal, I.; Zhang, Y.; Jørgensen, M.E.; Pisanty, O.; Barbosa, I.C.R.; Zourelidou, M.; Regnault, T.; Crocoll, C.; Erik Olsen, C.; Weinstain, R.; et al. The Arabidopsis NPF3 protein is a GA transporter. Nat. Commun. 2016, 7. [Google Scholar] [CrossRef]
- Kanno, Y.; Oikawa, T.; Chiba, Y.; Ishimaru, Y.; Shimizu, T.; Sano, N.; Koshiba, T.; Kamiya, Y.; Ueda, M.; Seo, M. AtSWEET13 and AtSWEET14 regulate gibberellin-mediated physiological processes. Nat. Commun. 2016, 7. [Google Scholar] [CrossRef] [PubMed]
- Rizza, A.; Walia, A.; Lanquar, V.; Frommer, W.B.; Jones, A.M. In vivo gibberellin gradients visualized in rapidly elongating tissues. Nat. Plants 2017, 3, 803–813. [Google Scholar] [CrossRef] [PubMed]
- Shani, E.; Weinstain, R.; Zhang, Y.; Castillejo, C.; Kaiserli, E.; Chory, J.; Tsien, R.Y.; Estelle, M. Gibberellins accumulate in the elongating endodermal cells of Arabidopsis root. Proc. Natl. Acad. Sci. USA 2013, 110, 4834–4839. [Google Scholar] [CrossRef] [PubMed]
- Davière, J.-M.; Achard, P. Gibberellin signaling in plants. Development 2013, 140, 1147–1151. [Google Scholar] [CrossRef] [PubMed]
- Okada, K.; Ito, T.; Fukazawa, J.; Takahashi, Y. Gibberellin Induces an Increase in Cytosolic Ca2+ via a DELLA-Independent Signaling Pathway. Plant Physiol. 2017, 175, 1536–1542. [Google Scholar] [CrossRef] [PubMed]
- Koornneef, M.; van der Veen, J.H. Induction and analysis of gibberellin sensitive mutants in Arabidopsis thaliana (L.) heynh. Theor. Appl. Genet. 1980, 58, 257–263. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Nagpal, P.; Vitart, V.; McMorris, T.C.; Chory, J. A role for brassinosteroids in light-dependent development of Arabidopsis. Science 1996, 272, 398–401. [Google Scholar] [CrossRef]
- Szekeres, M.; Németh, K.; Koncz-Kálmán, Z.; Mathur, J.; Kauschmann, A.; Altmann, T.; Rédei, G.P.; Nagy, F.; Schell, J.; Koncz, C. Brassinosteroids rescue the deficiency of CYP90, a cytochrome P450, controlling cell elongation and de-etiolation in Arabidopsis. Cell 1996, 85, 171–182. [Google Scholar] [CrossRef]
- Talon, M.; Koornneef, M.; Zeevaart, J.A. Endogenous gibberellins in Arabidopsis thaliana and possible steps blocked in the biosynthetic pathways of the semidwarf ga4 and ga5 mutants. Proc. Natl. Acad. Sci. USA 1990, 87, 7983–7987. [Google Scholar] [CrossRef]
- Wilson, R.N.; Somerville, C.R. Phenotypic Suppression of the Gibberellin-Insensitive Mutant (gai) of Arabidopsis. Plant Physiol. 1995, 108, 495–502. [Google Scholar] [CrossRef]
- Jager, C.E.; Symons, G.M.; Ross, J.J.; Smith, J.J.; Reid, J.B. The brassinosteroid growth response in pea is not mediated by changes in gibberellin content. Planta 2005, 221, 141–148. [Google Scholar] [CrossRef] [PubMed]
- Katsumi, M. Interaction of a Brassinosteroid with IAA and GA3 in the Elongation of Cucumber Hypocotyl Sections. Plant Cell Physiol. 1985, 26, 615–625. [Google Scholar] [CrossRef]
- Yang, G.X.; Jan, A.; Shen, S.H.; Yazaki, J.; Ishikawa, M.; Shimatani, Z.; Kishimoto, N.; Kikuchi, S.; Matsumoto, H.; Komatsu, S. Microarray analysis of brassinosteroids- and gibberellin-regulated gene expression in rice seedlings. Mol. Genet. Genom. 2004, 271, 468–478. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, K.; Nakamura, Y.; Asami, T.; Yoshida, S.; Matsuo, T.; Okamoto, S. Physiological Roles of Brassinosteroids in Early Growth of Arabidopsis: Brassinosteroids Have a Synergistic Relationship with Gibberellin as well as Auxin in Light-Grown Hypocotyl Elongation. J. Plant Growth Regul. 2003, 22, 259–271. [Google Scholar] [CrossRef]
- Bai, M.-Y.; Shang, J.-X.; Oh, E.; Fan, M.; Bai, Y.; Zentella, R.; Sun, T.; Wang, Z.-Y. Brassinosteroid, gibberellin and phytochrome impinge on a common transcription module in Arabidopsis. Nat. Cell Biol. 2012, 14, 810–817. [Google Scholar] [CrossRef] [PubMed]
- Gallego-Bartolome, J.; Minguet, E.G.; Grau-Enguix, F.; Abbas, M.; Locascio, A.; Thomas, S.G.; Alabadi, D.; Blazquez, M.A. Molecular mechanism for the interaction between gibberellin and brassinosteroid signaling pathways in Arabidopsis. Proc. Natl. Acad. Sci. USA 2012, 109, 13446–13451. [Google Scholar] [CrossRef]
- Li, Q.F.; Wang, C.; Jiang, L.; Li, S.; Sun, S.S.M.; He, J.X. An interaction between BZR1 and DELLAs mediates direct signaling crosstalk between brassinosteroids and gibberellins in Arabidopsis. Sci. Signal. 2012, 5, ra72. [Google Scholar] [CrossRef]
- Stewart Lilley, J.L.; Gan, Y.; Graham, I.A.; Nemhauser, J.L. The effects of DELLAs on growth change with developmental stage and brassinosteroid levels. Plant J. 2013, 76, 165–173. [Google Scholar] [CrossRef]
- Unterholzner, S.J.; Rozhon, W.; Papacek, M.; Ciomas, J.; Lange, T.; Kugler, K.G.; Mayer, K.F.; Sieberer, T.; Poppenberger, B. Brassinosteroids Are Master Regulators of Gibberellin Biosynthesis in Arabidopsis. Plant Cell 2015, 27, 2261–2272. [Google Scholar] [CrossRef]
- Tong, H.; Xiao, Y.; Liu, D.; Gao, S.; Liu, L.; Yin, Y.; Jin, Y.; Qian, Q.; Chu, C. Brassinosteroid Regulates Cell Elongation by Modulating Gibberellin Metabolism in Rice. Plant Cell Online 2014, 26, 4376–4393. [Google Scholar] [CrossRef]
- Ross, J.J.; Quittenden, L.J. Interactions between Brassinosteroids and Gibberellins: Synthesis or Signaling? Plant Cell 2016, 28, 829–832. [Google Scholar] [CrossRef] [PubMed]
- Tong, H.; Chu, C. Reply: Brassinosteroid Regulates Gibberellin Synthesis to Promote Cell Elongation in Rice: Critical Comments on Ross and Quittenden’s Letter. Plant Cell 2016, 28, 833–835. [Google Scholar] [CrossRef]
- Unterholzner, S.J.; Rozhon, W.; Poppenberger, B. REPLY: Interaction Between Brassinosteroids and Gibberellins: Synthesis or Signaling? In Arabidopsis, Both! Plant Cell 2016, 28, 836–839. [Google Scholar] [CrossRef] [PubMed]
- Allen, H.R.; Ptashnyk, M. Mathematical modelling and analysis of the brassinosteroid and gibberellin signalling pathways and their interactions. J. Theor. Biol. 2017, 432, 109–131. [Google Scholar] [CrossRef] [PubMed]
- De Vleesschauwer, D.; Van Buyten, E.; Satoh, K.; Balidion, J.; Mauleon, R.; Choi, I.-R.; Vera-Cruz, C.; Kikuchi, S.; Hofte, M. Brassinosteroids Antagonize Gibberellin- and Salicylate-Mediated Root Immunity in Rice. Plant Physiol. 2012, 158, 1833–1846. [Google Scholar] [CrossRef] [PubMed]
- Schmitz, A.J.; Folsom, J.J.; Jikamaru, Y.; Ronald, P.; Walia, H. SUB1A-mediated submergence tolerance response in rice involves differential regulation of the brassinosteroid pathway. New Phytol. 2013, 198, 1060–1070. [Google Scholar] [CrossRef] [PubMed]
- Achard, P.; Renou, J.P.; Berthomé, R.; Harberd, N.P.; Genschik, P. Plant DELLAs Restrain Growth and Promote Survival of Adversity by Reducing the Levels of Reactive Oxygen Species. Curr. Biol. 2008, 18, 656–660. [Google Scholar] [CrossRef]
- Werner, T.; Schmülling, T. Cytokinin action in plant development. Curr. Opin. Plant Biol. 2009, 12, 527–538. [Google Scholar] [CrossRef]
- Werner, T.; Nehnevajova, E.; Köllmer, I.; Novák, O.; Strnad, M.; Krämer, U.; Schmülling, T. Root-Specific Reduction of Cytokinin Causes Enhanced Root Growth, Drought Tolerance, and Leaf Mineral Enrichment in Arabidopsis and Tobacco. Plant Cell 2010, 22, 3905–3920. [Google Scholar] [CrossRef]
- Nishiyama, R.; Watanabe, Y.; Fujita, Y.; Le, D.T.; Kojima, M.; Werner, T.; Vankova, R.; Yamaguchi-Shinozaki, K.; Shinozaki, K.; Kakimoto, T.; et al. Analysis of Cytokinin Mutants and Regulation of Cytokinin Metabolic Genes Reveals Important Regulatory Roles of Cytokinins in Drought, Salt and Abscisic Acid Responses, and Abscisic Acid Biosynthesis. Plant Cell 2011, 23, 2169–2183. [Google Scholar] [CrossRef]
- Vercruyssen, L.; Gonzalez, N.; Werner, T.; Schmulling, T.; Inze, D. Combining Enhanced Root and Shoot Growth Reveals Cross Talk between Pathways That Control Plant Organ Size in Arabidopsis. Plant Physiol. 2011, 155, 1339–1352. [Google Scholar] [CrossRef] [PubMed]
- Yuan, L.B.; Peng, Z.H.; Zhi, T.T.; Zho, Z.; Liu, Y.; Zhu, Q.; Xiong, X.Y.; Ren, C.M. Brassinosteroid enhances cytokinin-induced anthocyanin biosynthesis in Arabidopsis seedlings. Biol. Plant. 2014, 59, 99–105. [Google Scholar] [CrossRef]
- Tran, L.-S.P.; Urao, T.; Qin, F.; Maruyama, K.; Kakimoto, T.; Shinozaki, K.; Yamaguchi-Shinozaki, K. Functional analysis of AHK1/ATHK1 and cytokinin receptor histidine kinases in response to abscisic acid, drought, and salt stress in Arabidopsis. Proc. Natl. Acad. Sci. USA 2007, 104, 20623–20628. [Google Scholar] [CrossRef] [PubMed]
- Havlová, M.; Dobrev, P.I.; Motyka, V.; Štorchová, H.; Libus, J.; Dobrá, J.; Malbeck, J.; Gaudinová, A.; Vanková, R. The role of cytokinins in responses to water deficit in tobacco plants over-expressing trans-zeatin O-glucosyltransferase gene under 35S or SAG12 promoters. Plant Cell Environ. 2008, 31, 341–353. [Google Scholar] [CrossRef] [PubMed]
- Argueso, C.T.; Ferreira, F.J.; Kieber, J.J. Environmental perception avenues: The interaction of cytokinin and environmental response pathways. Plant Cell Environ. 2009, 32, 1147–1160. [Google Scholar] [CrossRef] [PubMed]
- Nishiyama, R.; Le, D.T.; Watanabe, Y.; Matsui, A.; Tanaka, M.; Seki, M.; Yamaguchi-Shinozaki, K.; Shinozaki, K.; Tran, L.S.P. Transcriptome analyses of a salt-tolerant cytokinin-deficient mutant reveal differential regulation of salt stress response by cytokinin deficiency. PLoS ONE 2012, 7, e32124. [Google Scholar] [CrossRef] [PubMed]
- Peleg, Z.; Reguera, M.; Tumimbang, E.; Walia, H.; Blumwald, E. Cytokinin-mediated source/sink modifications improve drought tolerance and increase grain yield in rice under water-stress. Plant Biotechnol. J. 2011, 9, 747–758. [Google Scholar] [CrossRef] [PubMed]
- Roitsch, T.; Ehneß, R. Regulation of source/sink relations by cytokinins. Plant Growth Regul. 2000, 32, 359–367. [Google Scholar] [CrossRef]
- Vahala, J.; Schlagnhaufer, C.D.; Pell, E.J. Induction of an ACC synthase cDNA by ozone in light-grown Arabidopsis thaliana leaves. Physiol. Plant. 1998, 103, 45–50. [Google Scholar] [CrossRef]
- Khripach, V.; Zhabinskii, V.; De Groot, A. Twenty years of brassinosteroids: Steroidal plant hormones warrant better crops for the XXI century. Ann. Bot. 2000, 86, 441–447. [Google Scholar] [CrossRef]
- Choe, S.; Fujioka, S.; Noguchi, T.; Takatsuto, S.; Yoshida, S.; Feldmann, K.A. Overexpression of DWARF4 in the brassinosteroid biosynthetic pathway results in increased vegetative growth and seed yield in Arabidopsis. Plant J. 2001, 26, 573–582. [Google Scholar] [CrossRef] [PubMed]
- Sakamoto, T.; Morinaka, Y.; Ohnishi, T.; Sunohara, H.; Fujioka, S.; Ueguchi-Tanaka, M.; Mizutani, M.; Sakata, K.; Takatsuto, S.; Yoshida, S.; et al. Erect leaves caused by brassinosteroid deficiency increase biomass production and grain yield in rice. Nat. Biotechnol. 2006, 24, 105–109. [Google Scholar] [CrossRef]
- Morinaka, Y.; Sakamoto, T.; Inukai, Y.; Agetsuma, M.; Kitano, H.; Ashikari, M.; Matsuoka, M. Morphological alteration caused by brassinosteroid insensitivity increases the biomass and grain production of rice. Plant Physiol. 2006, 141, 924–931. [Google Scholar] [CrossRef] [PubMed]
- Sahni, S.; Prasad, B.D.; Liu, Q.; Grbic, V.; Sharpe, A.; Singh, S.P.; Krishna, P. Overexpression of the brassinosteroid biosynthetic gene DWF4 in Brassica napus simultaneously increases seed yield and stress tolerance. Sci. Rep. 2016, 6, 28298. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.-F.; Yu, J.-W.; Lu, J.; Fei, H.-Y.; Luo, M.; Cao, B.-W.; Huang, L.-C.; Zhang, C.-Q.; Liu, Q.-Q. Seed-Specific Expression of OsDWF4, a Rate-Limiting Gene Involved in Brassinosteroids Biosynthesis, Improves Both Grain Yield and Quality in Rice. J. Agric. Food Chem. 2018, 66, 3759–3772. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Cai, Z.; Wang, X. The primary signaling outputs of brassinosteroids are regulated by abscisic acid signaling. Proc. Natl. Acad. Sci. USA 2009, 106, 4543–4548. [Google Scholar] [CrossRef] [PubMed]
- Abeles, F.B.; Morgan, P.W.; Salveit, M.E. Ethylene in Plant Biology, 2nd ed.; Academic Press, Inc.: San Diego, CA, USA, 1992; ISBN 0-12-041451-1. [Google Scholar]
- Yang, S.F.; Hoffman, N.E. Ethylene biosynthesis and its regulation inhi gher plants. Annu. Rev. Plant Physiol. 1984, 35, 155–189. [Google Scholar] [CrossRef]
- Hansen, M.; Chae, H.S.; Kieber, J.J. Regulation of ACS protein stability by cytokinin and brassinosteroid. Plant J. 2009, 57, 606–614. [Google Scholar] [CrossRef]
- Zimmermann, P. GENEVESTIGATOR. Arabidopsis Microarray Database and Analysis Toolbox. Plant Physiol. 2004, 136, 2621–2632. [Google Scholar] [CrossRef]
- Lv, B.; Tian, H.; Zhang, F.; Liu, J.; Lu, S.; Bai, M.; Li, C.; Ding, Z. Brassinosteroids regulate root growth by controlling reactive oxygen species homeostasis and dual effect on ethylene synthesis in Arabidopsis. PLoS Genet. 2018, 14, e1007144. [Google Scholar] [CrossRef]
- Guo, Y.; Shan, W.; Liang, S.; Wu, C.; Wei, W.; Chen, J.; Lu, W.; Kuang, J. MaBZR1/2 act as transcriptional repressors of ethylene biosynthetic genes in banana fruit. Physiol. Plant. 2018. [Google Scholar] [CrossRef] [PubMed]
- Zhu, T.; Tan, W.R.; Deng, X.G.; Zheng, T.; Zhang, D.W.; Lin, H.H. Effects of brassinosteroids on quality attributes and ethylene synthesis in postharvest tomato fruit. Postharvest Biol. Technol. 2015, 100, 196–204. [Google Scholar] [CrossRef]
- Li, Q.F.; He, J.X. BZR1 Interacts with HY5 to Mediate Brassinosteroid- and Light-Regulated Cotyledon Opening in Arabidopsis in Darkness. Mol. Plant 2016, 9, 113–125. [Google Scholar] [CrossRef]
- Nie, S.; Huang, S.; Wang, S.; Cheng, D.; Liu, J.; Lv, S.; Li, Q.; Wang, X. Enhancing Brassinosteroid Signaling via Overexpression of Tomato (Solanum lycopersicum) SlBRI1 Improves Major Agronomic Traits. Front. Plant Sci. 2017, 8, 1386. [Google Scholar] [CrossRef]
- Zhu, T.; Deng, X.; Zhou, X.; Zhu, L.; Zou, L.; Li, P.; Zhang, D.; Lin, H. Ethylene and hydrogen peroxide are involved in brassinosteroid-induced salt tolerance in tomato. Sci. Rep. 2016, 6, 35392. [Google Scholar] [CrossRef]
- Larkindale, J.; Hall, J.D.; Knight, M.R.; Vierling, E.; Larkindale, J.; Hall, J.D.; Knight, M.R.; Vierling, E. Heat Stress Phenotypes of Arabidopsis Mutants Implicate Multiple Signaling Pathways in the Acquisition of Thermotolerance. Plant Physiol. 2018, 138, 882–897. [Google Scholar] [CrossRef]
- Bari, R.; Jones, J.D.G. Role of plant hormones in plant defence responses. Plant Mol. Biol. 2009, 69, 473–488. [Google Scholar] [CrossRef]
- Divi, U.K.; Rahman, T.; Krishna, P. Brassinosteroid-mediated stress tolerance in Arabidopsis shows interactions with abscisic acid, ethylene and salicylic acid pathways. BMC Plant Biol. 2010, 10, 151. [Google Scholar] [CrossRef]
- Serna, M.; Coll, Y.; Zapata, P.J.; Botella, M.Á.; Pretel, M.T.; Amorós, A. A brassinosteroid analogue prevented the effect of salt stress on ethylene synthesis and polyamines in lettuce plants. Sci. Hortic. 2015, 185, 105–112. [Google Scholar] [CrossRef]
- Mayak, S.; Tirosh, T.; Glick, B.R. Plant growth-promoting bacteria confer resistance in tomato plants to salt stress. Plant Physiol. Biochem. 2004, 42, 565–572. [Google Scholar] [CrossRef]
- Mustilli, A.-C.; Merlot, S.; Vavasseur, A.; Fenzi, F.; Giraudat, J. Arabidopsis OST1 Protein Kinase Mediates the Regulation of Stomatal Aperture by Abscisic Acid and Acts Upstream of Reactive Oxygen Species Production. Plant Cell Online 2002, 14, 3089–3099. [Google Scholar] [CrossRef]
- Yoshida, R.; Hobo, T.; Ichimura, K.; Mizoguchi, T.; Takahashi, F.; Aronso, J.; Ecker, J.R.; Shinozaki, K. ABA-activated SnRK2 protein kinase is required for dehydration stress signaling in Arabidopsis. Plant Cell Physiol. 2002, 43, 1473–1483. [Google Scholar] [CrossRef]
- Yoshida, R.; Umezawa, T.; Mizoguchi, T.; Takahashi, S.; Takahashi, F.; Shinozaki, K. The regulatory domain of SRK2E/OST1/SnRK2.6 interacts with ABI1 and integrates abscisic acid (ABA) and osmotic stress signals controlling stomatal closure in Arabidopsis. J. Biol. Chem. 2006, 281, 5310–5318. [Google Scholar] [CrossRef] [PubMed]
- Fujii, H.; Verslues, P.E.; Zhu, J.-K. Identification of Two Protein Kinases Required for Abscisic Acid Regulation of Seed Germination, Root Growth, and Gene Expression in Arabidopsis. Plant Cell Online 2007, 19, 485–494. [Google Scholar] [CrossRef] [PubMed]
- Fujii, H.; Zhu, J.-K. Arabidopsis mutant deficient in 3 abscisic acid-activated protein kinases reveals critical roles in growth, reproduction, and stress. Proc. Natl. Acad. Sci. USA 2009, 106, 8380–8385. [Google Scholar] [CrossRef]
- Umezawa, T.; Nakashima, K.; Miyakawa, T.; Kuromori, T.; Tanokura, M.; Shinozaki, K.; Yamaguchi-Shinozaki, K. Molecular basis of the core regulatory network in ABA responses: Sensing, signaling and transport. Plant Cell Physiol. 2010, 51, 1821–1839. [Google Scholar] [CrossRef] [PubMed]
- Fujii, H.; Chinnusamy, V.; Rodrigues, A.; Rubio, S.; Antoni, R.; Park, S.-Y.; Cutler, S.R.; Sheen, J.; Rodriguez, P.L.; Zhu, J.-K. In vitro reconstitution of an abscisic acid signalling pathway. Nature 2009, 462, 660–664. [Google Scholar] [CrossRef] [PubMed]
- Fujita, Y.; Nakashima, K.; Yoshida, T.; Katagiri, T.; Kidokoro, S.; Kanamori, N.; Umezawa, T.; Fujita, M.; Maruyama, K.; Ishiyama, K.; et al. Three SnRK2 protein kinases are the main positive regulators of abscisic acid signaling in response to water stress in Arabidopsis. Plant Cell Physiol. 2009, 50, 2123–2132. [Google Scholar] [CrossRef] [PubMed]
- Nakashima, K.; Fujita, Y.; Kanamori, N.; Katagiri, T.; Umezawa, T.; Kidokoro, S.; Maruyama, K.; Yoshida, T.; Ishiyama, K.; Kobayashi, M.; et al. Three Arabidopsis SnRK2 protein kinases, SRK2D/SnRK2.2, SRK2E/SnRK2.6/OST1 and SRK2I/SnRK2.3, involved in ABA signaling are essential for the control of seed development and dormancy. Plant Cell Physiol. 2009, 50, 1345–1363. [Google Scholar] [CrossRef] [PubMed]
- Umezawa, T.; Sugiyama, N.; Mizoguchi, M.; Hayashi, S.; Myouga, F.; Yamaguchi-Shinozaki, K.; Ishihama, Y.; Hirayama, T.; Shinozaki, K. Type 2C protein phosphatases directly regulate abscisic acid-activated protein kinases in Arabidopsis. Proc. Natl. Acad. Sci. USA 2009, 106, 17588–17593. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.; Szostkiewicz, I.; Korte, A.; Moes, D.; Yang, Y.; Christmann, A.; Grill, E. Regulators of PP2C phosphatase activity function as abscisic acid sensors. Science 2009, 324, 1064–1068. [Google Scholar] [CrossRef] [PubMed]
- Park, S.Y.; Fung, P.; Nishimura, N.; Jensen, D.R.; Fujii, H.; Zhao, Y.; Lumba, S.; Santiago, J.; Rodrigues, A.; Chow, T.F.F.; et al. Abscisic acid inhibits type 2C protein phosphatases via the PYR/PYL family of START proteins. Science 2009, 324, 1068–1071. [Google Scholar] [CrossRef] [PubMed]
- Miyazono, K.; Miyakawa, T.; Sawano, Y.; Kubota, K.; Kang, H.-J.; Asano, A.; Miyauchi, Y.; Takahashi, M.; Zhi, Y.; Fujita, Y.; et al. Structural basis of abscisic acid signalling. Nature 2009, 462, 609–614. [Google Scholar] [CrossRef] [PubMed]
- Melcher, K.; Ng, L.M.; Zhou, X.E.; Soon, F.F.; Xu, Y.; Suino-Powell, K.M.; Park, S.Y.; Weiner, J.J.; Fujii, H.; Chinnusamy, V.; et al. A gate-latch-lock mechanism for hormone signalling by abscisic acid receptors. Nature 2009, 462, 602–608. [Google Scholar] [CrossRef] [PubMed]
- Yin, P.; Fan, H.; Hao, Q.; Yuan, X.; Wu, D.; Pang, Y.; Yan, C.; Li, W.; Wang, J.; Yan, N. Structural insights into the mechanism of abscisic acid signaling by PYL proteins. Nat. Struct. Mol. Biol. 2009, 16, 1230–1236. [Google Scholar] [CrossRef] [PubMed]
- Arc, E.; Sechet, J.; Corbineau, F.; Rajjou, L.; Marion-Poll, A. ABA crosstalk with ethylene and nitric oxide in seed dormancy and germination. Front. Plant Sci. 2013, 4. [Google Scholar] [CrossRef]
- Daszkowska-Golec, A.; Szarejko, I. Open or Close the Gate—Stomata Action under the Control of Phytohormones in Drought Stress Conditions. Front. Plant Sci. 2013, 4, 138. [Google Scholar] [CrossRef] [PubMed]
- O’Brien, J.A.; Benková, E. Cytokinin cross-talking during biotic and abiotic stress responses. Front. Plant Sci. 2013, 4, 451. [Google Scholar] [CrossRef]
- Steber, C.M.; McCourt, P. A role for brassinosteroids in germination in Arabidopsis. Plant Physiol. 2001, 125, 763–769. [Google Scholar] [CrossRef]
- Rodrigues, A.; Santiago, J.; Rubio, S.; Saez, A.; Osmont, K.S.; Gadea, J.; Hardtke, C.S.; Rodriguez, P.L. The Short-Rooted Phenotype of the brevis radix Mutant Partly Reflects Root Abscisic Acid Hypersensitivity. Plant Physiol. 2009, 149, 1917–1928. [Google Scholar] [CrossRef]
- Li, J.; Nam, K.H.; Vafeados, D.; Chory, J. BIN2, a new brassinosteroid-insensitive locus in Arabidopsis. Plant Physiol. 2001, 127, 14–22. [Google Scholar] [CrossRef] [PubMed]
- Belkhadir, Y.; Jaillais, Y. The molecular circuitry of brassinosteroid signaling. New Phytol. 2015, 206, 522–540. [Google Scholar] [CrossRef] [PubMed]
- Cai, Z.; Liu, J.; Wang, H.; Yang, C.; Chen, Y.; Li, Y.; Pan, S.; Dong, R.; Tang, G.; de Dios Barajas-Lopez, J. et al. GSK3-like kinases positively modulate abscisic acid signaling through phosphorylating subgroup III SnRK2s in Arabidopsis. Proc. Natl. Acad. Sci. USA 2014, 111, 9651–9656. [Google Scholar] [CrossRef] [PubMed]
- Lopez-Molina, L.; Mongrand, S.; McLachlin, D.T.; Chait, B.T.; Chua, N.H. ABI5 acts downstream of ABI3 to execute an ABA-dependent growth arrest during germination. Plant J. 2002, 32, 317–328. [Google Scholar] [CrossRef] [PubMed]
- Lopez-Molina, L.; Mongrand, S.; Chua, N.-H. A postgermination developmental arrest checkpoint is mediated by abscisic acid and requires the ABI5 transcription factor in Arabidopsis. Proc. Natl. Acad. Sci. USA 2001, 98, 4782–4787. [Google Scholar] [CrossRef] [PubMed]
- Finkelstein, R.R. The Arabidopsis Abscisic Acid Response Gene ABI5 Encodes a Basic Leucine Zipper Transcription Factor. Plant Cell Online 2000, 12, 599–610. [Google Scholar] [CrossRef]
- Hu, Y.; Yu, D. BRASSINOSTEROID INSENSITIVE2 Interacts with ABSCISIC ACID INSENSITIVE5 to Mediate the Antagonism of Brassinosteroids to Abscisic Acid during Seed Germination in Arabidopsis. Plant Cell Online 2014, 26, 4394–4408. [Google Scholar] [CrossRef]
- Wang, H.; Tang, J.; Liu, J.; Hu, J.; Liu, J.; Chen, Y.; Cai, Z.; Wang, X. Abscisic Acid Signaling Inhibits Brassinosteroid Signaling through Dampening the Dephosphorylation of BIN2 by ABI1 and ABI2. Mol. Plant 2018, 11, 315–325. [Google Scholar] [CrossRef]
- Shang, Y.; Dai, C.; Lee, M.M.; Kwak, J.M.; Nam, K.H. BRI1-Associated Receptor Kinase 1 Regulates Guard Cell ABA Signaling Mediated by Open Stomata 1 in Arabidopsis. Mol. Plant 2016, 9, 447–460. [Google Scholar] [CrossRef]
- Ryu, H.; Cho, H.; Bae, W.; Hwang, I. Control of early seedling development by BES1/TPL/HDA19-mediated epigenetic regulation of ABI3. Nat. Commun. 2014, 5, 4138. [Google Scholar] [CrossRef]
- Yang, X.; Bai, Y.; Shang, J.; Xin, R.; Tang, W. The antagonistic regulation of abscisic acid-inhibited root growth by brassinosteroids is partially mediated via direct suppression of ABSCISIC ACID INSENSITIVE 5 expression by BRASSINAZOLE RESISTANT 1. Plant Cell Environ. 2016, 39, 1994–2003. [Google Scholar] [CrossRef] [PubMed]
- Ng, L.-M.; Soon, F.-F.; Zhou, X.E.; West, G.M.; Kovach, A.; Suino-Powell, K.M.; Chalmers, M.J.; Li, J.; Yong, E.-L.; Zhu, J.-K.; et al. Structural basis for basal activity and autoactivation of abscisic acid (ABA) signaling SnRK2 kinases. Proc. Natl. Acad. Sci. USA 2011, 108, 21259–21264. [Google Scholar] [CrossRef]
- Belin, C.; de Franco, P.-O.; Bourbousse, C.; Chaignepain, S.; Schmitter, J.-M.; Vavasseur, A.; Giraudat, J.; Barbier-Brygoo, H.; Thomine, S. Identification of features regulating OST1 kinase activity and OST1 function in guard cells. Plant Physiol. 2006, 141, 1316–1327. [Google Scholar] [CrossRef]
- Yunta, C.; Martínez-Ripoll, M.; Zhu, J.K.; Albert, A. The structure of Arabidopsis thaliana OST1 provides insights into the kinase regulation mechanism in response to osmotic stress. J. Mol. Biol. 2011, 414, 135–144. [Google Scholar] [CrossRef]
- Wang, Z.-Y. Brassinosteroids modulate plant immunity at multiple levels. Proc. Natl. Acad. Sci. USA 2012, 109, 7–8. [Google Scholar] [CrossRef] [PubMed]
- Tena, G.; Boudsocq, M.; Sheen, J. Protein kinase signaling networks in plant innate immunity. Curr. Opin. Plant Biol. 2011, 14, 519–529. [Google Scholar] [CrossRef] [PubMed]
- McCann, H.C.; Nahal, H.; Thakur, S.; Guttman, D.S. Identification of innate immunity elicitors using molecular signatures of natural selection. Proc. Natl. Acad. Sci. USA 2012, 109, 4215–4220. [Google Scholar] [CrossRef]
- Spoel, S.H.; Dong, X. How do plants achieve immunity? Defence without specialized immune cells. Nat. Rev. Immunol. 2012, 12, 89–100. [Google Scholar] [CrossRef]
- Nakashita, H.; Yasuda, M.; Nitta, T.; Asami, T.; Fujioka, S.; Arai, Y.; Sekimata, K.; Takatsuto, S.; Yamaguchi, I.; Yoshida, S. Brassinosteriod functions in a broad range of disease resistance in tobacco and rice. Plant J. 2003, 33, 887–898. [Google Scholar] [CrossRef]
- Chinchilla, D.; Shan, L.; He, P.; de Vries, S.; Kemmerling, B. One for all: The receptor-associated kinase BAK1. Trends Plant Sci. 2009, 14, 535–541. [Google Scholar] [CrossRef]
- Chinchilla, D.; Zipfel, C.; Robatzek, S.; Kemmerling, B.; Nürnberger, T.; Jones, J.D.G.; Felix, G.; Boller, T. A flagellin-induced complex of the receptor FLS2 and BAK1 initiates plant defence. Nature 2007, 448, 497–500. [Google Scholar] [CrossRef]
- Heese, A.; Hann, D.R.; Gimenez-Ibanez, S.; Jones, A.M.E.; He, K.; Li, J.; Schroeder, J.I.; Peck, S.C.; Rathjen, J.P. The receptor-like kinase SERK3/BAK1 is a central regulator of innate immunity in plants. Proc. Natl. Acad. Sci. USA 2007, 104, 12217–12222. [Google Scholar] [CrossRef]
- Segonzac, C.; Zipfel, C. Activation of plant pattern-recognition receptors by bacteria. Curr. Opin. Microbiol. 2011, 14, 54–61. [Google Scholar] [CrossRef]
- Lu, D.; Wu, S.; Gao, X.; Zhang, Y.; Shan, L.; He, P. A receptor-like cytoplasmic kinase, BIK1, associates with a flagellin receptor complex to initiate plant innate immunity. Proc. Natl. Acad. Sci. USA 2010, 107, 496–501. [Google Scholar] [CrossRef]
- Zhang, J.; Li, W.; Xiang, T.; Liu, Z.; Laluk, K.; Ding, X.; Zou, Y.; Gao, M.; Zhang, X.; Chen, S.; et al. Receptor-like cytoplasmic kinases integrate signaling from multiple plant immune receptors and are targeted by a Pseudomonas syringae effector. Cell Host Microbe 2010, 7, 290–301. [Google Scholar] [CrossRef]
- Albrecht, C.; Boutrot, F.; Segonzac, C.; Schwessinger, B.; Gimenez-Ibanez, S.; Chinchilla, D.; Rathjen, J.P.; de Vries, S.C.; Zipfel, C. Brassinosteroids inhibit pathogen-associated molecular pattern-triggered immune signaling independent of the receptor kinase BAK1. Proc. Natl. Acad. Sci. USA 2012, 109, 303–308. [Google Scholar] [CrossRef]
- Belkhadir, Y.; Jaillais, Y.; Epple, P.; Balsemao-Pires, E.; Dangl, J.L.; Chory, J. Brassinosteroids modulate the efficiency of plant immune responses to microbe-associated molecular patterns. Proc. Natl. Acad. Sci. USA 2012, 109, 297–302. [Google Scholar] [CrossRef]
- Grant, M.R.; Jones, J.D.G.; Science, S.; Series, N.; May, N. Plant-Microbe Interactions Hormone (Dis) Harmony Moulds Plant Health and Disease. Science 2009, 324, 750–752. [Google Scholar] [CrossRef]
- Robert-Seilaniantz, A.; Grant, M.; Jones, J.D.G. Hormone Cross-talk in Plant Disease and Defense: More Than Just JASMONATE-SALICYLATE Antagonism. Annu. Rev. Phytopathol. 2011, 49, 317–343. [Google Scholar] [CrossRef]
- Koornneef, A.; Pieterse, C.M.J. Cross Talk in Defense Signaling. Plant Physiol. 2008, 146, 839–844. [Google Scholar] [CrossRef]
- Pieterse, C.M.J.; Leon-Reyes, A.; Van Der Ent, S.; Van Wees, S.C.M. Networking by small-molecule hormones in plant immunity. Nat. Chem. Biol. 2009, 5, 308–316. [Google Scholar] [CrossRef]
- Thaler, J.S.; Humphrey, P.T.; Whiteman, N.K. Evolution of jasmonate and salicylate signal. Trends Plant Sci. 2012, 17, 260–270. [Google Scholar] [CrossRef]
- Zhou, G.; Qi, J.; Ren, N.; Cheng, J.; Erb, M.; Mao, B.; Lou, Y. Silencing OsHI-LOX makes rice more susceptible to chewing herbivores, but enhances resistance to a phloem feeder. Plant J. 2009, 60, 638–648. [Google Scholar] [CrossRef]
- Zhou, G.; Ren, N.; Qi, J.; Lu, J.; Xiang, C.; Ju, H.; Cheng, J.; Lou, Y. The 9-lipoxygenase Osr9-LOX1 interacts with the 13-lipoxygenase-mediated pathway to regulate resistance to chewing and piercing-sucking herbivores in rice. Physiol. Plant. 2014, 152, 59–69. [Google Scholar] [CrossRef]
- Tamaoki, D.; Seo, S.; Yamada, S.; Kano, A.; Miyamoto, A.; Shishido, H.; Miyoshi, S.; Taniguch, S.; Akimitsu, K.; Gomi, K. Jasmonic acid and salicylic acid activate a common defense system in rice. Plant Signal. Behav. 2013, 8, e24260. [Google Scholar] [CrossRef]
- Pan, G.; Liu, Y.; Ji, L.; Zhang, X.; He, J.; Huang, J.; Qiu, Z.; Liu, D.; Sun, Z.; Xu, T.; et al. Brassinosteroids Mediate Susceptibility to Brown Planthopper by Integrating with Salicylic Acid and Jasmonic Acid Pathways in Rice. J. Exp. Bot. 2018, 69, 4433–4442. [Google Scholar] [CrossRef]
- Campos, M.L.; De Almeida, M.; Rossi, M.L.; Martinelli, A.P.; Litholdo Junior, C.G.; Figueira, A.; Rampelotti-Ferreira, F.T.; Vendramim, J.D.; Benedito, V.A.; Pereira Peres, L.E. Brassinosteroids interact negatively with jasmonates in the formation of anti-herbivory traits in tomato. J. Exp. Bot. 2009, 60, 4347–4361. [Google Scholar] [CrossRef]
- He, Y.; Zhang, H.; Sun, Z.; Li, J.; Hong, G.; Zhu, Q.; Zhou, X.; MacFarlane, S.; Yan, F.; Chen, J. Jasmonic acid-mediated defense suppresses brassinosteroid-mediated susceptibility to Rice black streaked dwarf virus infection in rice. New Phytol. 2017, 214, 388–399. [Google Scholar] [CrossRef]
- Miyaji, T.; Yamagami, A.; Kume, N.; Sakuta, M.; Osada, H.; Asami, T.; Arimoto, Y.; Nakano, T. Brassinosteroid-related transcription factor BIL1/BZR1 increases plant resistance to insect feeding. Biosci. Biotechnol. Biochem. 2014, 78, 960–968. [Google Scholar] [CrossRef]
- Yang, D.H.; Baldwin, I.T.; Wu, J. Silencing Brassinosteroid Receptor BRI1 Impairs Herbivory-elicited Accumulation of Jasmonic Acid-isoleucine and Diterpene Glycosides, but not Jasmonic Acid and Trypsin Proteinase Inhibitors in Nicotiana attenuata. J. Integr. Plant Biol. 2013, 55, 514–526. [Google Scholar] [CrossRef]
- Huot, B.; Yao, J.; Montgomery, B.L.; He, S.Y. Growth-defense tradeoffs in plants: A balancing act to optimize fitness. Mol. Plant 2014, 7, 1267–1287. [Google Scholar] [CrossRef]
- Beveridge, C.A.; Kyozuka, J. New genes in the strigolactone-related shoot branching pathway. Curr. Opin. Plant Biol. 2010, 13, 34–39. [Google Scholar] [CrossRef]
- Wang, Y.; Sun, S.; Zhu, W.; Jia, K.; Yang, H.; Wang, X. Strigolactone/MAX2-Induced Degradation of Brassinosteroid Transcriptional Effector BES1 Regulates Shoot Branching. Dev. Cell 2013, 27, 681–688. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Peres, A.L.G.L.; Soares, J.S.; Tavares, R.G.; Righetto, G.; Zullo, M.A.T.; Mandava, N.B.; Menossi, M. Brassinosteroids, the Sixth Class of Phytohormones: A Molecular View from the Discovery to Hormonal Interactions in Plant Development and Stress Adaptation. Int. J. Mol. Sci. 2019, 20, 331. https://doi.org/10.3390/ijms20020331
Peres ALGL, Soares JS, Tavares RG, Righetto G, Zullo MAT, Mandava NB, Menossi M. Brassinosteroids, the Sixth Class of Phytohormones: A Molecular View from the Discovery to Hormonal Interactions in Plant Development and Stress Adaptation. International Journal of Molecular Sciences. 2019; 20(2):331. https://doi.org/10.3390/ijms20020331
Chicago/Turabian StylePeres, Ana Laura G. L., José Sérgio Soares, Rafael G. Tavares, Germanna Righetto, Marco A. T. Zullo, N. Bhushan Mandava, and Marcelo Menossi. 2019. "Brassinosteroids, the Sixth Class of Phytohormones: A Molecular View from the Discovery to Hormonal Interactions in Plant Development and Stress Adaptation" International Journal of Molecular Sciences 20, no. 2: 331. https://doi.org/10.3390/ijms20020331
APA StylePeres, A. L. G. L., Soares, J. S., Tavares, R. G., Righetto, G., Zullo, M. A. T., Mandava, N. B., & Menossi, M. (2019). Brassinosteroids, the Sixth Class of Phytohormones: A Molecular View from the Discovery to Hormonal Interactions in Plant Development and Stress Adaptation. International Journal of Molecular Sciences, 20(2), 331. https://doi.org/10.3390/ijms20020331