A tRNA-mimic Strategy to Explore the Role of G34 of tRNAGly in Translation and Codon Frameshifting
Abstract
1. Introduction
2. Results and Discussion
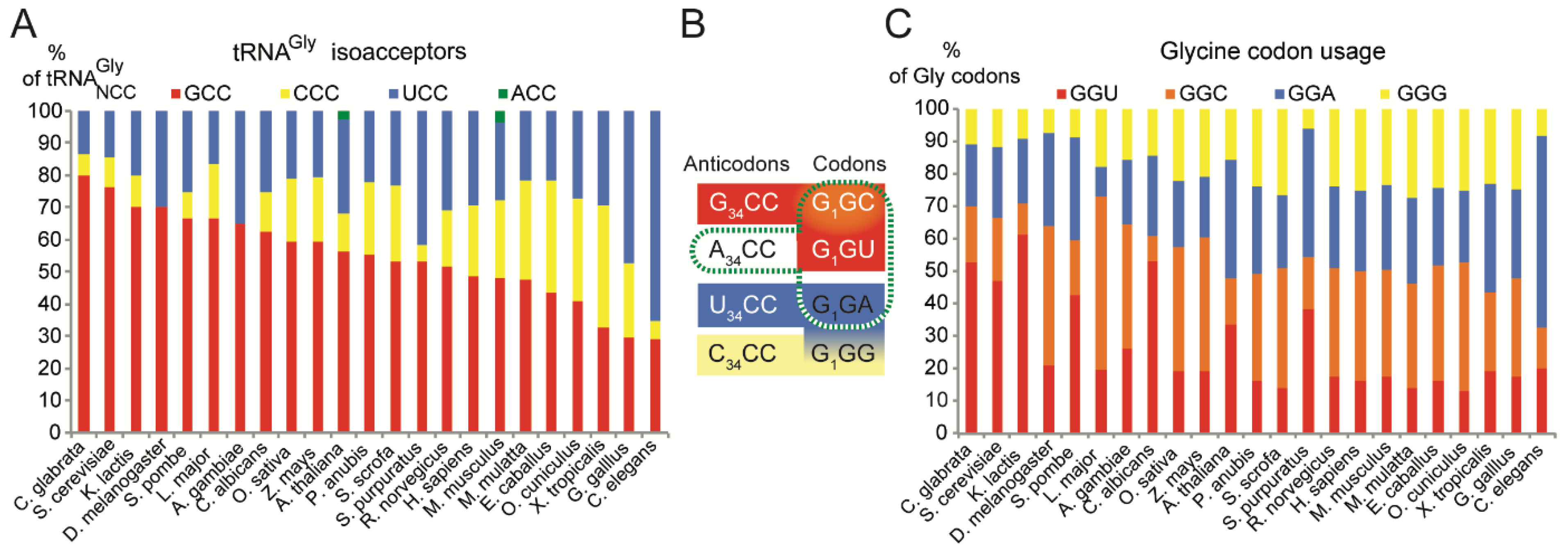
2.1. tRNAsGly Have Unusual Nucleotide 34 Distribution
2.2. Usage of Codons Gly in Eukaryotes Does Not Show Any Bias
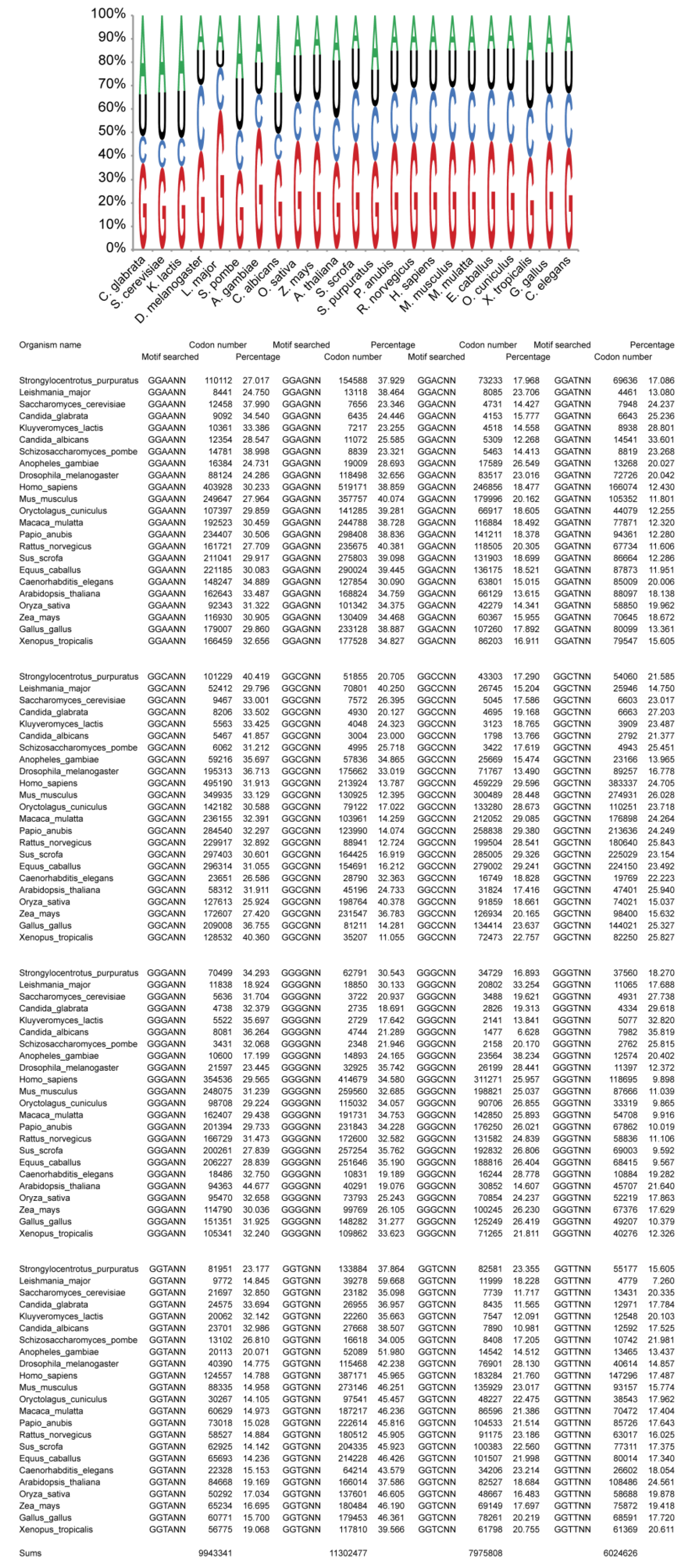
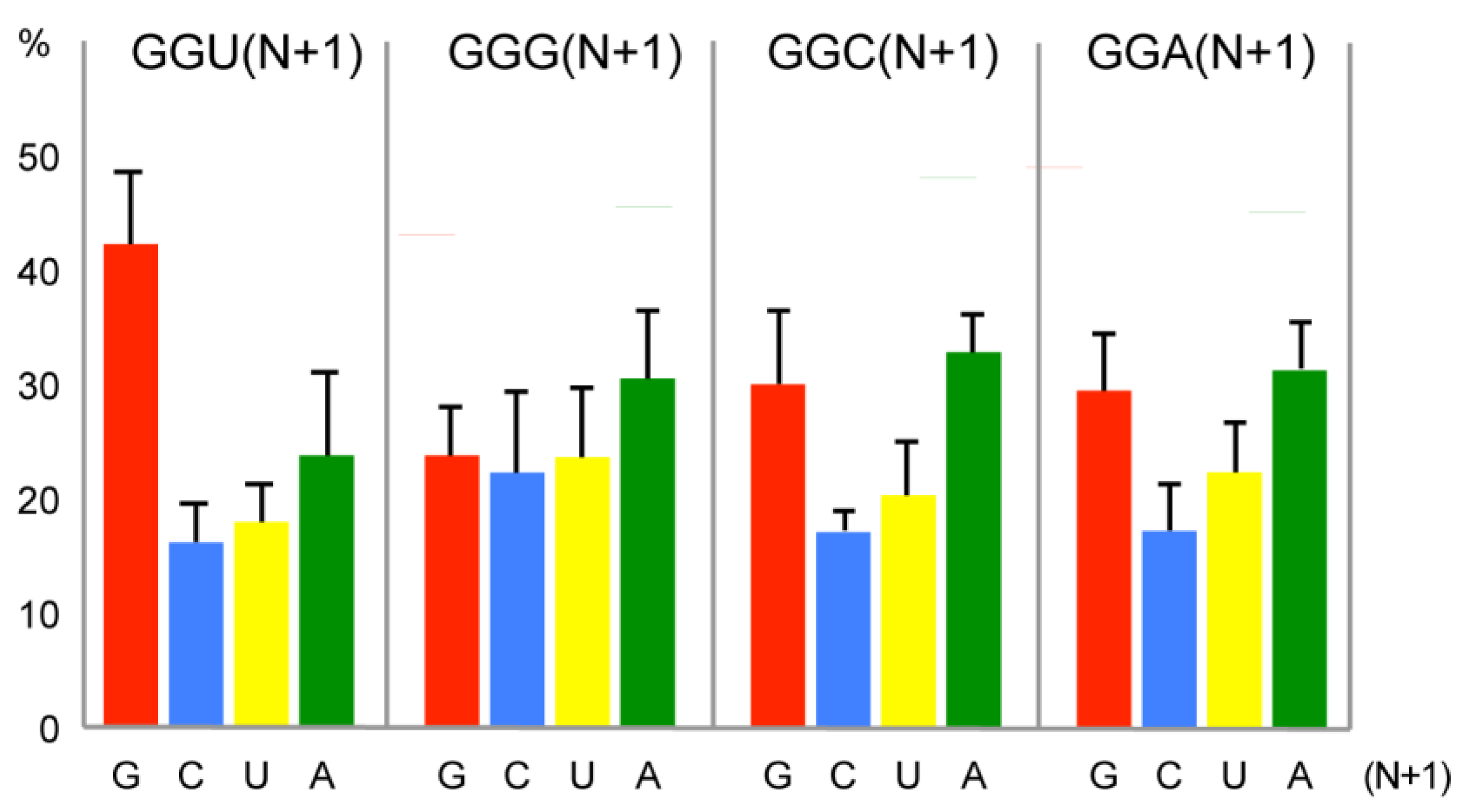
2.3. Bias of Nucleotide +1 after GGU Glycine Codon
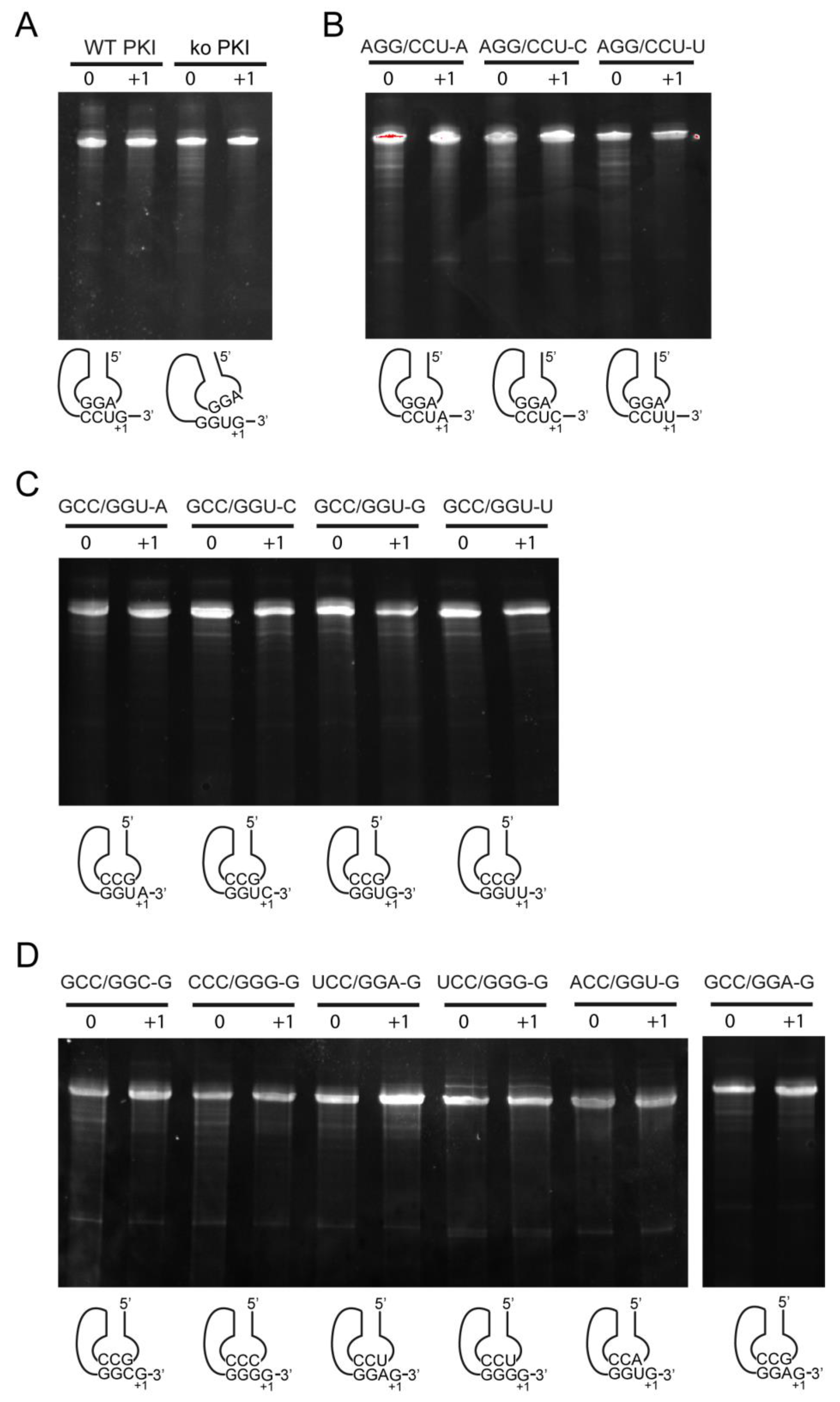
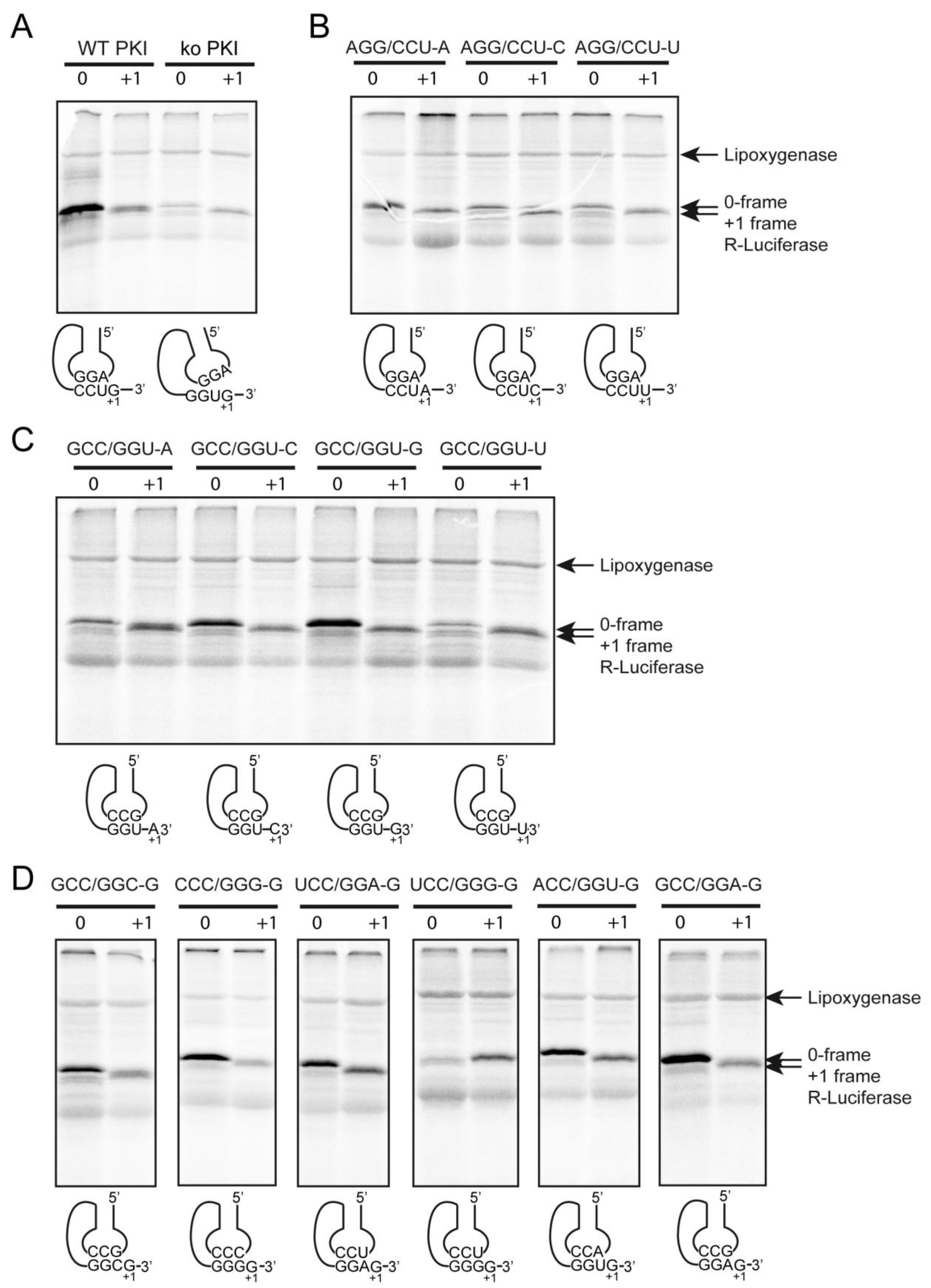
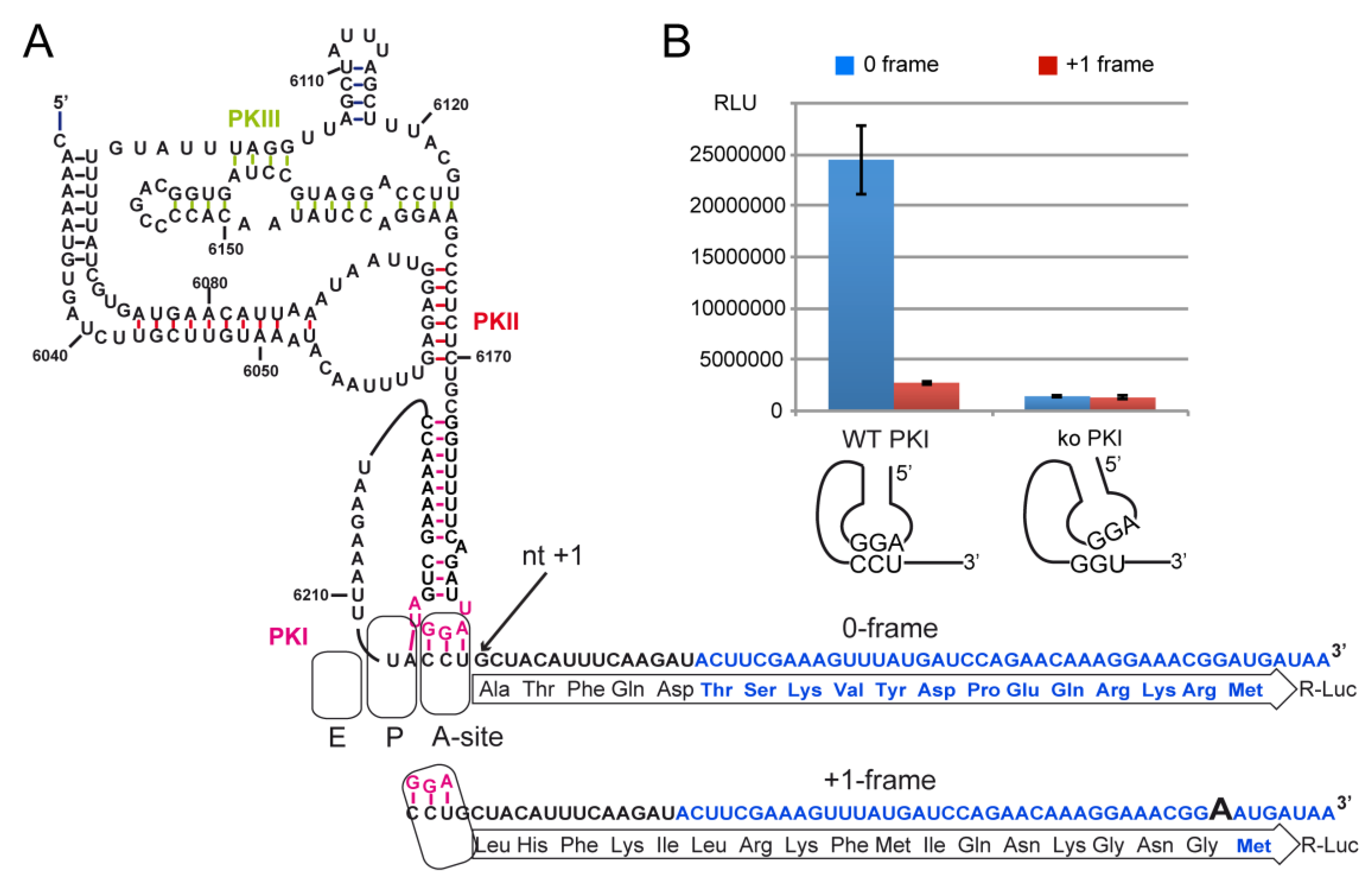
2.4. Decoding and Frameshifting Efficiency on Gly Codons
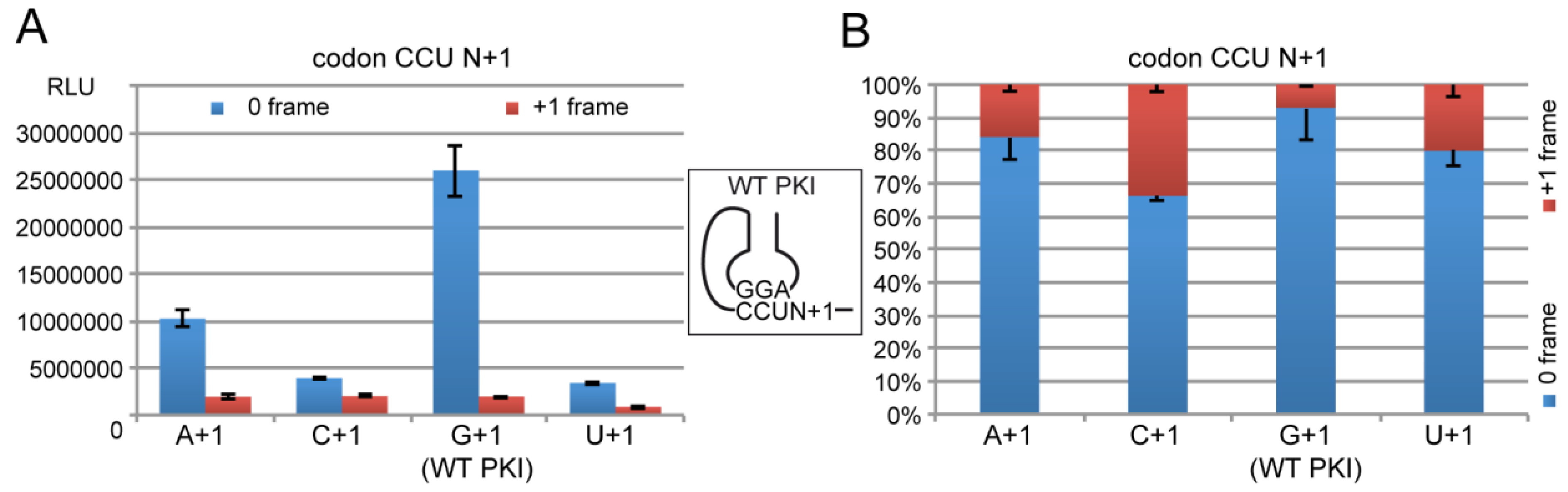
2.4.1. Decoding and Frameshifting Efficiency of the WT IGR
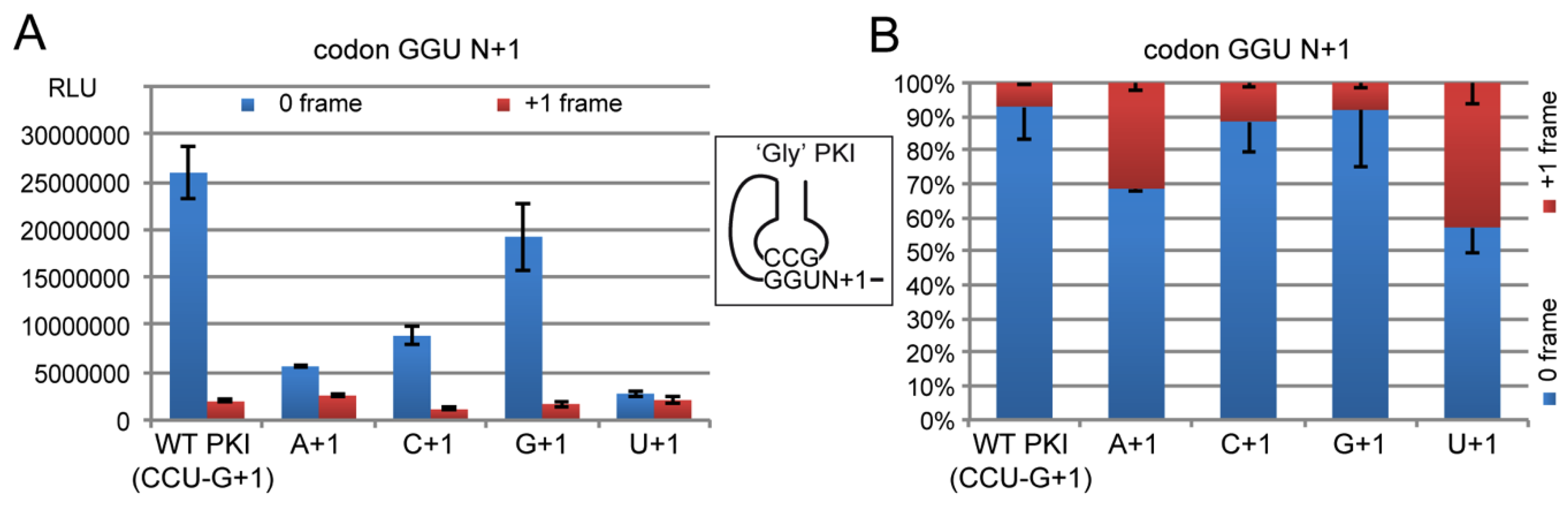
2.4.2. Decoding Efficiency of the IGR Programmed with GGU Gly Codon
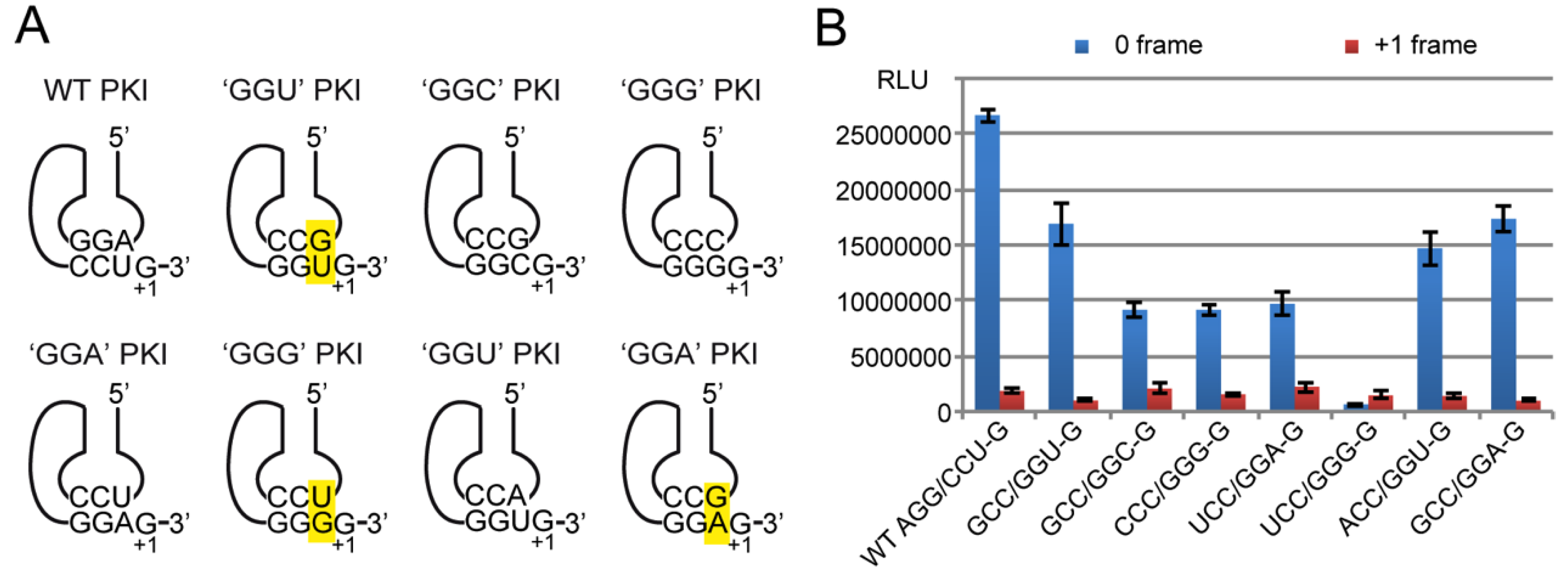
2.4.3. Decoding Efficiency of the IGR Programmed with Other Gly Codons
2.4.4. Frameshifting Activity of the IGR Programmed with GGU Codons
3. Materials and Methods
3.1. Data Processing
3.2. Construction of the IGR-luciferase Reporters
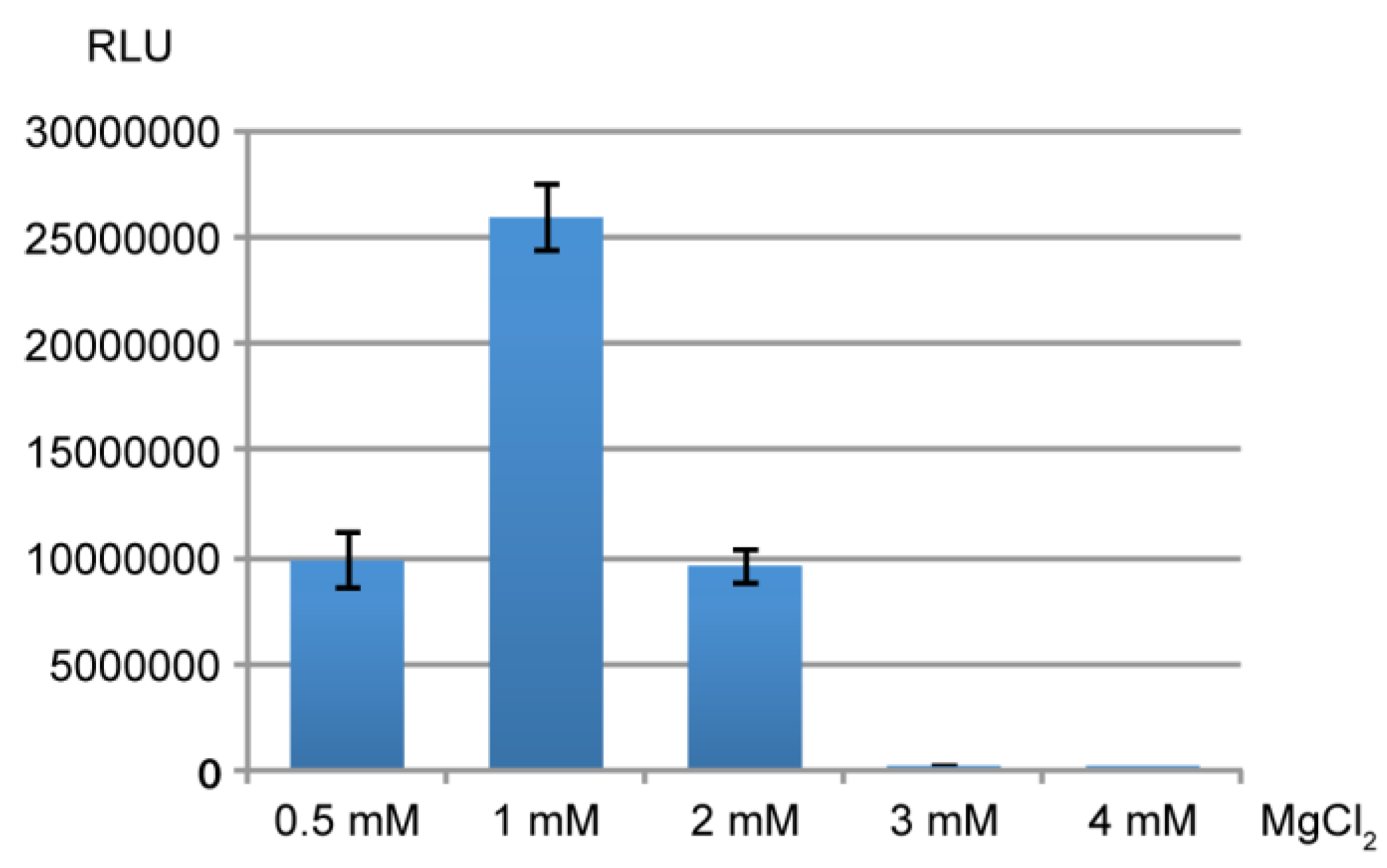
3.3. In vitro Translation in Rabbit Reticulocyte Lysates
4. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| ASL | Anticodon Stem Loop |
| CDS | Coding DNA sequence |
| CrPV | Cricket Paralysis Virus |
| IGR | InterGenic Region |
| IRES | Internal Ribosomal Entry Site |
| nt | nucleotide |
| ORF | Open Reading Frame |
| PKI | Pseudoknot I |
| R-Luc | Renilla Luciferase |
| RRL | Rabbit Reticulocyte Lysate |
Appendix A
References
- Pape, T.; Wintermeyer, W.; Rodnina, M. Induced fit in initial selection and proofreading of aminoacyl-tRNA on the ribosome. EMBO J. 1999, 18, 3800–3807. [Google Scholar] [CrossRef] [PubMed]
- Rodnina, M.V.; Wintermeyer, W. Fidelity of Aminoacyl-tRNA Selection on the Ribosome: Kinetic and Structural Mechanisms. Annu. Rev. Biochem. 2001, 70, 415–435. [Google Scholar] [CrossRef] [PubMed]
- Rodnina, M.V.; Wintermeyer, W. Ribosome fidelity: tRNA discrimination, proofreading and induced fit. Trends Biochem. Sci. 2001, 26, 124–130. [Google Scholar] [CrossRef]
- Sasaki, J.; Nakashima, N. Translation initiation at the CUU codon is mediated by the internal ribosome entry site of an insect picorna-like virus in vitro. J. Virol. 1999, 73, 1219–1226. [Google Scholar] [PubMed]
- Wilson, J.E.; Pestova, T.V.; Hellen, C.U.; Sarnow, P. Initiation of protein synthesis from the A site of the ribosome. Cell 2000, 102, 511–520. [Google Scholar] [CrossRef]
- Fernández, I.S.; Bai, X.C.; Murshudov, G.; Scheres, S.H.W.; Ramakrishnan, V. Initiation of translation by cricket paralysis virus IRES requires its translocation in the ribosome. Cell 2014, 157, 823–831. [Google Scholar] [CrossRef] [PubMed]
- Costantino, D.A.; Pfingsten, J.S.; Rambo, R.P.; Kieft, J.S. tRNA–mRNA mimicry drives translation initiation from a viral IRES. Nat. Struct. Mol. Biol. 2008, 15, 57–64. [Google Scholar] [CrossRef]
- Koh, C.S.; Brilot, A.F.; Grigorieff, N.; Korostelev, A.A. Taura syndrome virus IRES initiates translation by binding its tRNA-mRNA-like structural element in the ribosomal decoding center. Proc. Natl. Acad. Sci. USA 2014, 111, 9139–9144. [Google Scholar] [CrossRef]
- Murray, J.; Savva, C.G.; Shin, B.S.; Dever, T.E.; Ramakrishnan, V.; Fernández, I.S. Structural characterization of ribosome recruitment and translocation by type IV IRES. Elife 2016, 5, e13567. [Google Scholar] [CrossRef]
- Hussain, T.; Llácer, J.L.; Fernández, I.S.; Munoz, A.; Martin-Marcos, P.; Savva, C.G.; Lorsch, J.R.; Hinnebusch, A.G.; Ramakrishnan, V. Structural changes enable start codon recognition by the eukaryotic translation initiation complex. Cell 2014, 159, 597–607. [Google Scholar] [CrossRef]
- Erzberger, J.P.; Stengel, F.; Pellarin, R.; Zhang, S.; Schaefer, T.; Aylett, C.H.S.; Cimermančič, P.; Boehringer, D.; Sali, A.; Aebersold, R.; et al. Molecular architecture of the 40S⋅eIF1⋅eIF3 translation initiation complex. Cell 2014, 158, 1123–1135. [Google Scholar] [CrossRef] [PubMed]
- Muhs, M.; Hilal, T.; Mielke, T.; Skabkin, M.A.; Sanbonmatsu, K.Y.; Pestova, T.V.; Spahn, C.M.T. Cryo-EM of ribosomal 80s complexes with termination factors reveals the translocated cricket paralysis virus IRES. Mol. Cell 2015, 57, 422–433. [Google Scholar] [CrossRef] [PubMed]
- Pisareva, V.P.; Pisarev, A.V.; Fernández, I.S. Dual tRNA mimicry in the Cricket Paralysis Virus IRES uncovers an unexpected similarity with the Hepatitis C Virus IRES. Elife 2018, 7, e34062. [Google Scholar] [CrossRef] [PubMed]
- Petrov, A.; Grosely, R.; Chen, J.; O’Leary, S.E.; Puglisi, J.D. Multiple Parallel Pathways of Translation Initiation on the CrPV IRES. Mol. Cell 2016, 62, 92–103. [Google Scholar] [CrossRef] [PubMed]
- Kurland, C.G.; Ehrenberg, M. Growth-optimizing accuracy of gene expression. Annu. Rev. Biophys. Biophys. Chem. 1987, 16, 291–317. [Google Scholar] [CrossRef]
- Farabaugh, P.J. Programmed translational frameshifting. Microbiol. Rev. 1996, 60, 103–134. [Google Scholar] [CrossRef]
- Namy, O.; Moran, S.J.; Stuart, D.I.; Gilbert, R.J.; Brierley, I. A mechanical explanation of RNA pseudoknot function in programmed ribosomal frameshifting. Nature 2006, 441, 244–247. [Google Scholar] [CrossRef]
- Atkins, J.F.; Loughran, G.; Bhatt, P.R.; Firth, A.E.; Baranov, P.V. Ribosomal frameshifting and transcriptional slippage: From genetic steganography and cryptography to adventitious use. Nucleic Acids Res. 2016, 44, 7007–7078. [Google Scholar] [CrossRef]
- Atkins, J.F.; Gesteland, R.F.; Reid, B.R.; Anderson, C.W. Normal tRNAs promote ribosomal frameshifting. Cell 1979, 18, 1119–1131. [Google Scholar] [CrossRef]
- Björk, G.R.; Wikström, P.M.; Byström, A.S. Prevention of translational frameshifting by the modified nucleoside 1-methylguanosine. Science 1989, 244, 986–989. [Google Scholar] [CrossRef]
- Hagervall, T.G.; Tuohy, T.M.; Atkins, J.F.; Björk, G.R. Deficiency of 1-methylguanosine in tRNA from Salmonella typhimurium induces frameshifting by quadruplet translocation. J. Mol. Biol. 1993, 232, 756–765. [Google Scholar] [CrossRef] [PubMed]
- Qian, Q.; Björk, G.R. Structural alterations far from the anticodon of the tRNAProGGG of Salmonella typhimurium induce +1 frameshifting at the peptidyl-site. J. Mol. Biol. 1997, 273, 978–992. [Google Scholar] [CrossRef] [PubMed]
- Hong, S.; Sunita, S.; Maehigashi, T.; Hoffer, E.D.; Dunkle, J.A.; Dunham, C.M. Mechanism of tRNA-mediated +1 ribosomal frameshifting. Proc. Natl. Acad. Sci. USA 2018, 115, 11226–11231. [Google Scholar] [CrossRef] [PubMed]
- Maehigashi, T.; Dunkle, J.A.; Miles, S.J.; Dunham, C.M. Structural insights into +1 frameshifting promoted by expanded or modification-deficient anticodon stem loops. Proc. Natl. Acad. Sci. USA 2014, 111, 12740–12745. [Google Scholar] [CrossRef] [PubMed]
- Ledoux, S.; Olejniczak, M.; Uhlenbeck, O.C. A sequence element that tunes Escherichia coli tRNA(Ala)(GGC) to ensure accurate decoding. Nat. Struct. Mol. Biol. 2009, 16, 359–364. [Google Scholar] [CrossRef] [PubMed]
- Murakami, H.; Ohta, A.; Suga, H. Bases in the anticodon loop of tRNAAlaGGC prevent misreading. Nat. Struct. Mol. Biol. 2009, 16, 353–358. [Google Scholar] [CrossRef] [PubMed]
- Olejniczak, M.; Uhlenbeck, O.C. tRNA residues that have coevolved with their anticodon to ensure uniform and accurate codon recognition. Biochimie 2006, 88, 943–950. [Google Scholar] [CrossRef]
- Grosjean, H.; Westhof, E. An integrated, structure- and energy-based view of the genetic code. Nucleic Acids Res. 2016, 44, 8020–8040. [Google Scholar] [CrossRef]
- Chan, P.P.; Lowe, T.M. GtRNAdb 2.0: An expanded database of transfer RNA genes identified in complete and draft genomes. Nucleic Acids Res. 2016, 44, D184–D189. [Google Scholar] [CrossRef]
- Rogalski, M.; Karcher, D.; Bock, R. Superwobbling facilitates translation with reduced tRNA sets. Nat. Struct. Mol. Biol. 2008, 15, 192–198. [Google Scholar] [CrossRef]
- Curran, J.F. Decoding with the A:I wobble pair is inefficient. Nucleic Acids Res. 1995, 23, 683–688. [Google Scholar] [CrossRef]
- Pak, D.; Kim, Y.; Burton, Z.F. Aminoacyl-tRNA synthetase evolution and sectoring of the genetic code. Transcription 2018, 9, 1–15. [Google Scholar] [CrossRef]
- Gaber, R.F.; Mathison, L.; Edelman, I.; Culbertson, M.R. Frameshift Suppression in Saccharomyces cerevisiae VI. Complete Genetic Map of Twenty-Five Suppressor Genes. Genetics 1983, 103, 389–407. [Google Scholar]
- Mathew, S.F.; Crowe-McAuliffe, C.; Graves, R.; Cardno, T.S.; McKinney, C.; Poole, E.S.; Tate, W.P. The highly conserved codon following the slippery sequence supports—1 frameshift efficiency at the HIV-1 frameshift site. PLoS ONE 2015, 10, e0122176. [Google Scholar] [CrossRef][Green Version]
- Shehu-Xhilaga, M.; Crowe, S.M.; Mak, J. Maintenance of the Gag/Gag-Pol Ratio Is Important for Human Immunodeficiency Virus Type 1 RNA Dimerization and Viral Infectivity. J. Virol. 2001, 75, 1834–1841. [Google Scholar] [CrossRef]
- Belew, A.T.; Meskauskas, A.; Musalgaonkar, S.; Advani, V.M.; Sulima, S.O.; Kasprzak, W.K.; Shapiro, B.A.; Dinman, J.D. Ribosomal frameshifting in the CCR5 mRNA is regulated by miRNAs and the NMD pathway. Nature 2014, 512, 265–269. [Google Scholar] [CrossRef]
- Advani, V.M.; Belew, A.T.; Dinman, J.D. Yeast telomere maintenance is globally controlled by programmed ribosomal frameshifting and the nonsense-mediated mRNA decay pathway. Translation 2013, 1, e24418. [Google Scholar] [CrossRef]
- Sharma, V.; Prere, M.F.; Canal, I.; Firth, A.E.; Atkins, J.F.; Baranov, P.V.; Fayet, O. Analysis of tetra- and hepta-nucleotides motifs promoting -1 ribosomal frameshifting in Escherichia coli. Nucleic Acids Res. 2014, 42, 7210–7225. [Google Scholar] [CrossRef]
- Nakamura, Y.; Gojobori, T.; Ikemura, T. Codon usage tabulated from international DNA sequence databases: Status for the year 2000. Nucleic Acids Res. 2000, 28, 292. [Google Scholar] [CrossRef]
- Murphy, F.V.; Ramakrishnan, V. Structure of a purine-purine wobble base pair in the decoding center of the ribosome. Nat. Struct. Mol. Biol. 2004, 11, 1251–1252. [Google Scholar] [CrossRef]
- Grosjean, H.J.; de Henau, S.; Crothers, D.M. On the physical basis for ambiguity in genetic coding interactions. Proc. Natl. Acad. Sci. USA 1978, 75, 610–614. [Google Scholar] [CrossRef]
- Grosjean, H.; de Crécy-Lagard, V.; Marck, C. Deciphering synonymous codons in the three domains of life: Co-evolution with specific tRNA modification enzymes. FEBS Lett. 2010, 584, 252–264. [Google Scholar] [CrossRef]
- Komine, Y.; Kitabatake, M.; Yokogawa, T.; Nishikawa, K.; Inokuchi, H. A tRNA-like structure is present in 10Sa RNA, a small stable RNA from Escherichia coli. Proc. Natl. Acad. Sci. USA 1994, 91, 9223–9227. [Google Scholar] [CrossRef]
- Kerr, C.H.; Wang, Q.S.; Moon, K.M.; Keatings, K.; Allan, D.W.; Foster, L.J.; Jan, E. IRES-dependent ribosome repositioning directs translation of a +1 overlapping ORF that enhances viral infection. Nucleic Acids Res. 2018, 46, 11952–11967. [Google Scholar] [CrossRef]
- Ren, Q.; Au, H.H.T.; Wang, Q.S.; Lee, S.; Jan, E. Structural determinants of an internal ribosome entry site that direct translational reading frame selection. Nucleic Acids Res. 2014, 42, 9366–9382. [Google Scholar] [CrossRef]
- Au, H.H.; Cornilescu, G.; Mouzakis, K.D.; Ren, Q.; Burke, J.E.; Lee, S.; Butcher, S.E.; Jan, E. Global shape mimicry of tRNA within a viral internal ribosome entry site mediates translational reading frame selection. Proc. Natl. Acad. Sci. USA 2015, 112, E6446–E6455. [Google Scholar] [CrossRef]
- Ren, Q.; Wang, Q.S.; Firth, A.E.; Chan, M.M.Y.; Gouw, J.W.; Guarna, M.M.; Foster, L.J.; Atkins, J.F.; Jan, E. Alternative reading frame selection mediated by a tRNA-like domain of an internal ribosome entry site. Proc. Natl. Acad. Sci. USA 2012, 109, E630–E639. [Google Scholar] [CrossRef]
- Kieft, J.S. Comparing the three-dimensional structures of Dicistroviridae IGR IRES RNAs with other viral RNA structures. Virus Res. 2009, 139, 148–156. [Google Scholar] [CrossRef]
- Varshavsky, A. The N-end rule. Cell 1992, 69, 725–735. [Google Scholar] [CrossRef]
- Westhof, E. Isostericity and tautomerism of base pairs in nucleic acids. FEBS Lett. 2014, 588, 2464–2469. [Google Scholar] [CrossRef]
- Westhof, E.; Yusupov, M.; Yusupova, G. Recognition of Watson-Crick base pairs: Constraints and limits due to geometric selection and tautomerism. F1000Prime Rep. 2014, 6, 19. [Google Scholar] [CrossRef]
- Murphy, F.V.; Ramakrishnan, V.; Malkiewicz, A.; Agris, P.F. The role of modifications in codon discrimination by tRNALysUUU. Nat. Struct. Mol. Biol. 2004, 11, 1186–1191. [Google Scholar] [CrossRef]
- Vendeix, F.A.P.; Murphy, F.V.; Cantara, W.A.; Leszczyńska, G.; Gustilo, E.M.; Sproat, B.; Malkiewicz, A.; Agris, P.F. Human tRNA(Lys3)(UUU) is pre-structured by natural modifications for cognate and wobble codon binding through keto-enol tautomerism. J. Mol. Biol. 2012, 416, 467–485. [Google Scholar] [CrossRef]
- Kurata, S.; Weixlbaumer, A.; Ohtsuki, T.; Shimazaki, T.; Wada, T.; Kirino, Y.; Takai, K.; Watanabe, K.; Ramakrishnan, V.; Suzuki, T. Modified Uridines with C5-methylene Substituents at the First Position of the tRNA Anticodon Stabilize U·G Wobble Pairing during Decoding. J. Biol. Chem. 2008, 283, 18801–18811. [Google Scholar] [CrossRef]
- Weixlbaumer, A.; Murphy, F.V.; Dziergowska, A.; Malkiewicz, A.; Vendeix, F.A.P.; Agris, P.F.; Ramakrishnan, V. Mechanism for expanding the decoding capacity of transfer RNAs by modification of uridines. Nat. Struct. Mol. Biol. 2007, 14, 498–502. [Google Scholar] [CrossRef]
- Ito, Y.; Sone, Y.; Mizutani, T. Stability of non-Watson-Crick G-A/A-G base pair in synthetic DNA and RNA oligonucleotides. Mol. Biol. Rep. 2004, 31, 31–36. [Google Scholar] [CrossRef]
- Olson, W.K.; Li, S.; Kaukonen, T.; Colasanti, A.V.; Xin, Y.; Lu, X.J. Effects of Noncanonical Base Pairing on RNA Folding: Structural Context and Spatial Arrangements of G·A Pairs. Biochemistry 2019, 58, 2474–2487. [Google Scholar] [CrossRef]
- Fernández, I.S.; Ng, C.L.; Kelley, A.C.; Wu, G.; Yu, Y.T.; Ramakrishnan, V. Unusual base pairing during the decoding of a stop codon by the ribosome. Nature 2013, 500, 107–110. [Google Scholar] [CrossRef]
- Ogle, J.M.; Ramakrishnan, V. Structural Insights into translational fidelity. Annu. Rev. Biochem. 2005, 74, 129–177. [Google Scholar] [CrossRef]
- Chen, J.L.; Dishler, A.L.; Kennedy, S.D.; Yildirim, I.; Liu, B.; Turner, D.H.; Serra, M.J. Testing the nearest neighbor model for canonical RNA base pairs: Revision of GU parameters. Biochemistry 2012, 51, 3508–3522. [Google Scholar] [CrossRef]
- Wohlgemuth, I.; Pohl, C.; Mittelstaet, J.; Konevega, A.L.; Rodnina, M.V. Evolutionary optimization of speed and accuracy of decoding on the ribosome. Philos. Trans. R. Soc. B Biol. Sci. 2011, 366, 2979–2986. [Google Scholar] [CrossRef]
- Wohlgemuth, I.; Pohl, C.; Rodnina, M.V. Optimization of speed and accuracy of decoding in translation. EMBO J. 2010, 29, 3701–3709. [Google Scholar] [CrossRef]
- Rozov, A.; Demeshkina, N.; Westhof, E.; Yusupov, M.; Yusupova, G. New Structural Insights into Translational Miscoding. Trends Biochem. Sci. 2016, 41, 798–814. [Google Scholar] [CrossRef]
- Ogle, J.M.; Brodersen, D.E.; Clemons, W.M.; Tarry, M.J.; Carter, A.P.; Ramakrishnan, V. Recognition of cognate transfer RNA by the 30S ribosomal subunit. Science 2001, 292, 897–902. [Google Scholar] [CrossRef]
- Jenner, L.B.; Demeshkina, N.; Yusupova, G.; Yusupov, M. Structural aspects of messenger RNA reading frame maintenance by the ribosome. Nat. Struct. Mol. Biol. 2010, 17, 555–560. [Google Scholar] [CrossRef]
- Selmer, M.; Dunham, C.M.; Murphy, F.V., 4th; Weixlbaumer, A.; Petry, S.; Kelley, A.C.; Weir, J.R.; Ramakrishnan, V. Structure of the 70S Ribosome Complexed with mRNA and tRNA. Science 2006, 313, 1935–1942. [Google Scholar] [CrossRef]
- Demeshkina, N.; Jenner, L.; Yusupova, G.; Yusupov, M. Interactions of the ribosome with mRNA and tRNA. Curr. Opin. Struct. Biol. 2010, 20, 325–332. [Google Scholar] [CrossRef]
- Murgola, E.J.; Hijazi, K.A.; Goringer, H.U.; Dahlberg, A.E. Mutant 16S ribosomal RNA: A codon-specific translational suppressor. Proc. Natl. Acad. Sci. USA 1988, 85, 4162–4165. [Google Scholar] [CrossRef]
- Prescott, C.D.; Dahlberg, A.E. A single base change at 726 in 16S rRNA radically alters the pattern of proteins synthesized in vivo. EMBO J. 1990, 9, 289–294. [Google Scholar] [CrossRef]
- Moine, H.; Dahlberg, A.E. Mutations in Helix 34 of Escherichia coli 16 S Ribosomal RNA Have Multiple Effects on Ribosome Function and Synthesis. J. Mol. Biol. 1994, 243, 402–412. [Google Scholar] [CrossRef]
- Ledoux, S.; Uhlenbeck, O.C. Different aa-tRNAs are selected uniformly on the ribosome. Mol. Cell 2008, 31, 114–123. [Google Scholar] [CrossRef]
- Gross, L.; Schaeffer, L.; Alghoul, F.; Hayek, H.; Allmang, C.; Eriani, G.; Martin, F. Tracking the m 7 G-cap during translation initiation by crosslinking methods. Methods 2018, 137, 3–10. [Google Scholar] [CrossRef]
- Martin, F.; Ménétret, J.F.; Simonetti, A.; Myasnikov, A.G.; Vicens, Q.; Prongidi-Fix, L.; Natchiar, S.K.; Klaholz, B.P.; Eriani, G. Ribosomal 18S rRNA base pairs with mRNA during eukaryotic translation initiation. Nat. Commun. 2016, 7, 12622. [Google Scholar] [CrossRef]
- Martin, F.; Barends, S.; Jaeger, S.; Schaeffer, L.; Prongidi-Fix, L.; Eriani, G. Cap-assisted internal initiation of translation of histone h4. Mol. Cell 2011, 41, 197–209. [Google Scholar] [CrossRef]
- Saint-Léger, A.; Bello, C.; Dans, P.D.; Torres, A.G.; Novoa, E.M.; Camacho, N.; Orozco, M.; Kondrashov, F.A.; Ribas de Pouplana, L. Saturation of recognition elements blocks evolution of new tRNA identities. Sci. Adv. 2016, 2, e1501860. [Google Scholar] [CrossRef]
- Chang, A.T.; Nikonowicz, E.P. Solution NMR determination of hydrogen bonding and base pairing between the glyQS T box riboswitch Specifier domain and the anticodon loop of tRNA(Gly). FEBS Lett. 2013, 587, 3495–3499. [Google Scholar] [CrossRef]
- Grundy, F.J.; Winkler, W.C.; Henkin, T.M. tRNA-mediated transcription antitermination in vitro: Codon-anticodon pairing independent of the ribosome. Proc. Natl. Acad. Sci. USA 2002, 99, 11121–11126. [Google Scholar] [CrossRef]
- Niyomporn, B.; Dahl, J.L.; Strominger, J.L. Biosynthesis of the peptidoglycan of bacterial cell walls. IX. Purification and properties of glycyl transfer ribonucleic acid synthetase from Staphylococcus aureus. J. Biol. Chem. 1968, 243, 773–778. [Google Scholar]
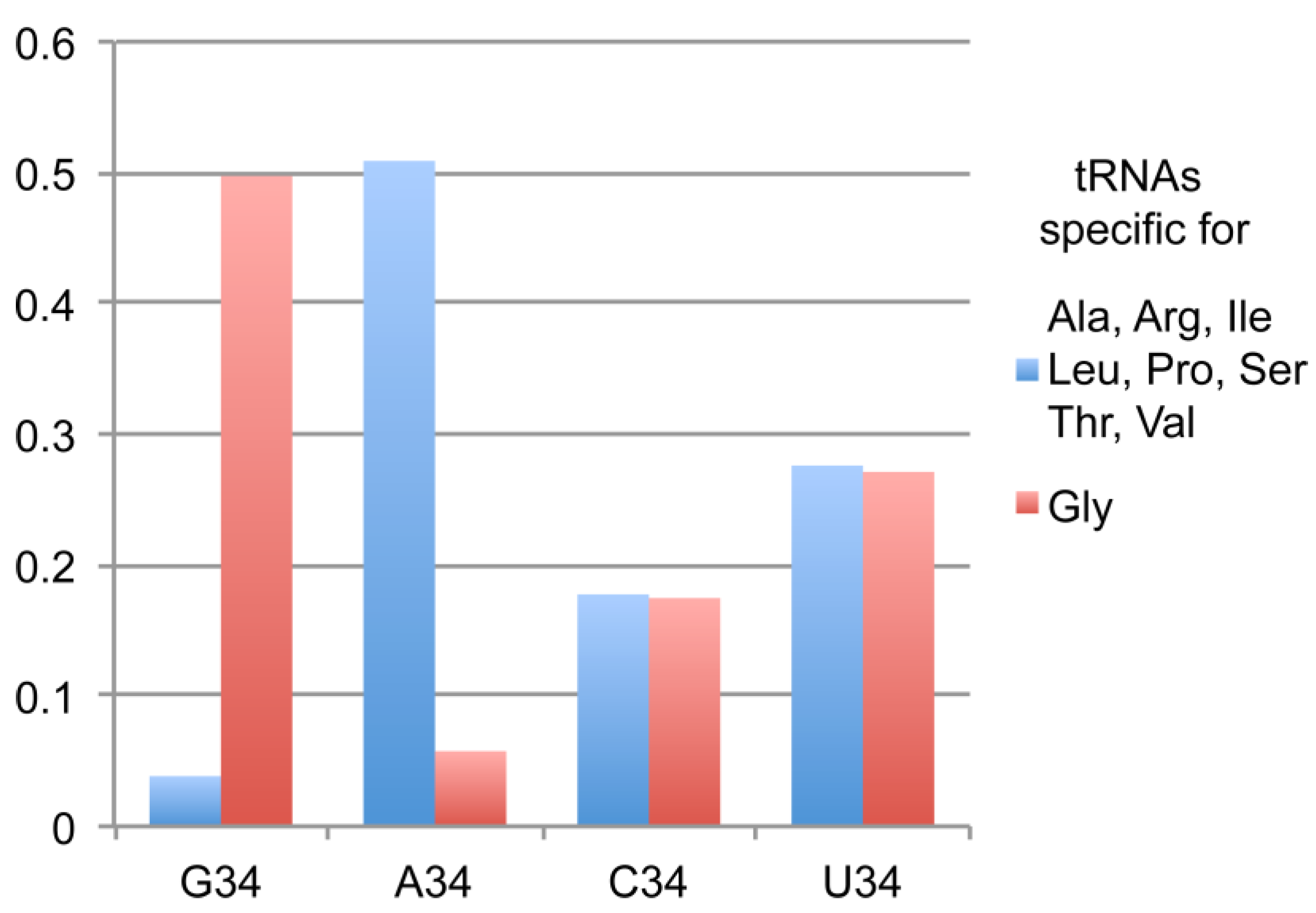
| tRNAs Specific for | Total Number in Eukarya 1 | With G34 | With A34 | With C34 | With U34 |
|---|---|---|---|---|---|
| Ala, Arg, Ile, Leu, Pro, Ser, Thr and Val | 80,060 | 3063 | 40,726 | 14,194 | 22,077 |
| Gly | 13,347 | 6636 | 767 | 2328 | 3616 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Janvier, A.; Despons, L.; Schaeffer, L.; Tidu, A.; Martin, F.; Eriani, G. A tRNA-mimic Strategy to Explore the Role of G34 of tRNAGly in Translation and Codon Frameshifting. Int. J. Mol. Sci. 2019, 20, 3911. https://doi.org/10.3390/ijms20163911
Janvier A, Despons L, Schaeffer L, Tidu A, Martin F, Eriani G. A tRNA-mimic Strategy to Explore the Role of G34 of tRNAGly in Translation and Codon Frameshifting. International Journal of Molecular Sciences. 2019; 20(16):3911. https://doi.org/10.3390/ijms20163911
Chicago/Turabian StyleJanvier, Aurélie, Laurence Despons, Laure Schaeffer, Antonin Tidu, Franck Martin, and Gilbert Eriani. 2019. "A tRNA-mimic Strategy to Explore the Role of G34 of tRNAGly in Translation and Codon Frameshifting" International Journal of Molecular Sciences 20, no. 16: 3911. https://doi.org/10.3390/ijms20163911
APA StyleJanvier, A., Despons, L., Schaeffer, L., Tidu, A., Martin, F., & Eriani, G. (2019). A tRNA-mimic Strategy to Explore the Role of G34 of tRNAGly in Translation and Codon Frameshifting. International Journal of Molecular Sciences, 20(16), 3911. https://doi.org/10.3390/ijms20163911







