Herb-Induced Liver Injury: Phylogenetic Relationship, Structure-Toxicity Relationship, and Herb-Ingredient Network Analysis
Abstract
1. Introduction
2. Results and Discussion
2.1. Cluster Pattern of Medicinal Plants with Potential Hepatotoxicity
2.2. Phytochemicals with Potential Hepatotoxicity
2.2.1. Skeleton-Based Cluster Pattern of Hepatotoxic Ingredients
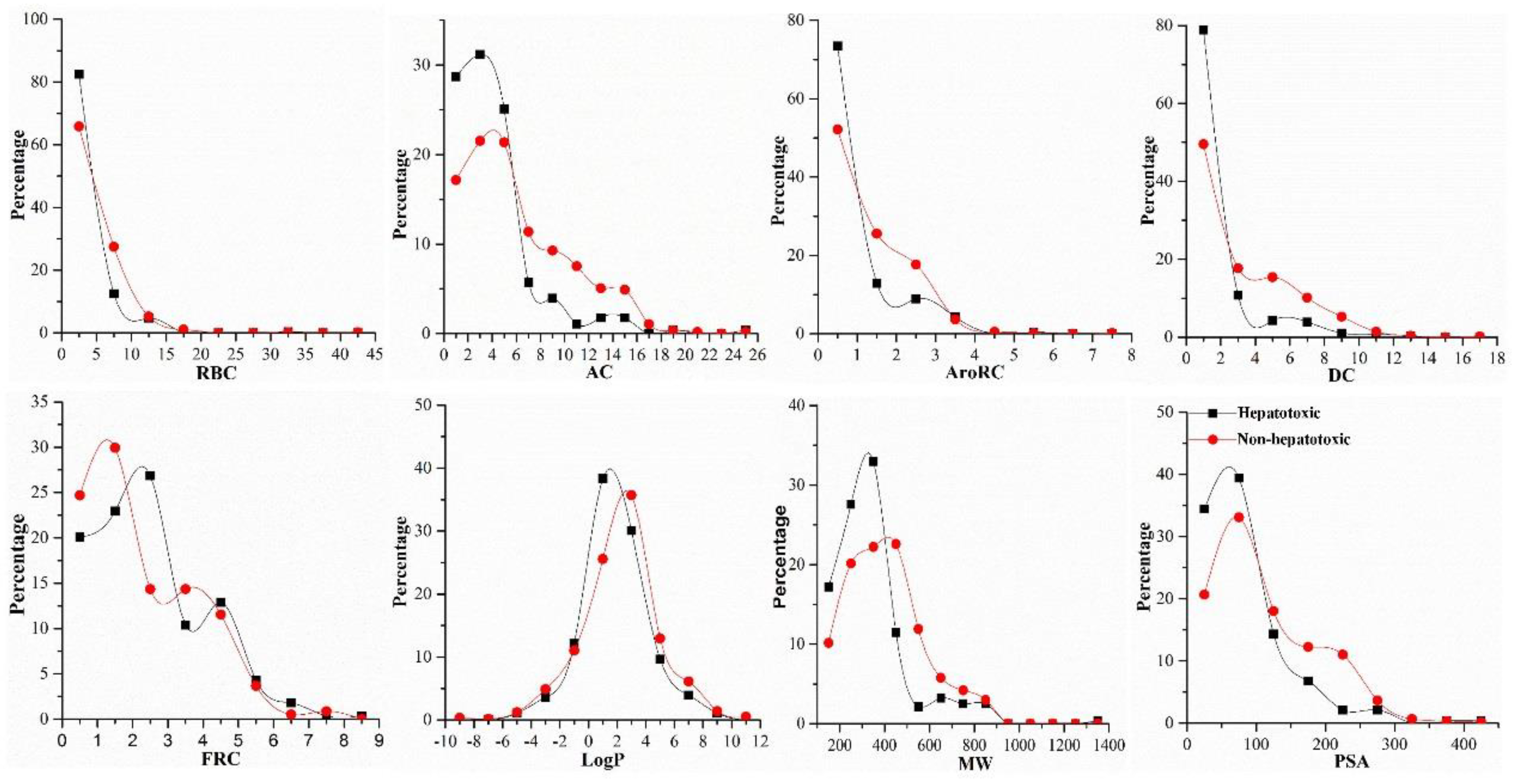
2.2.2. Chemical Space and Drug-Likeness
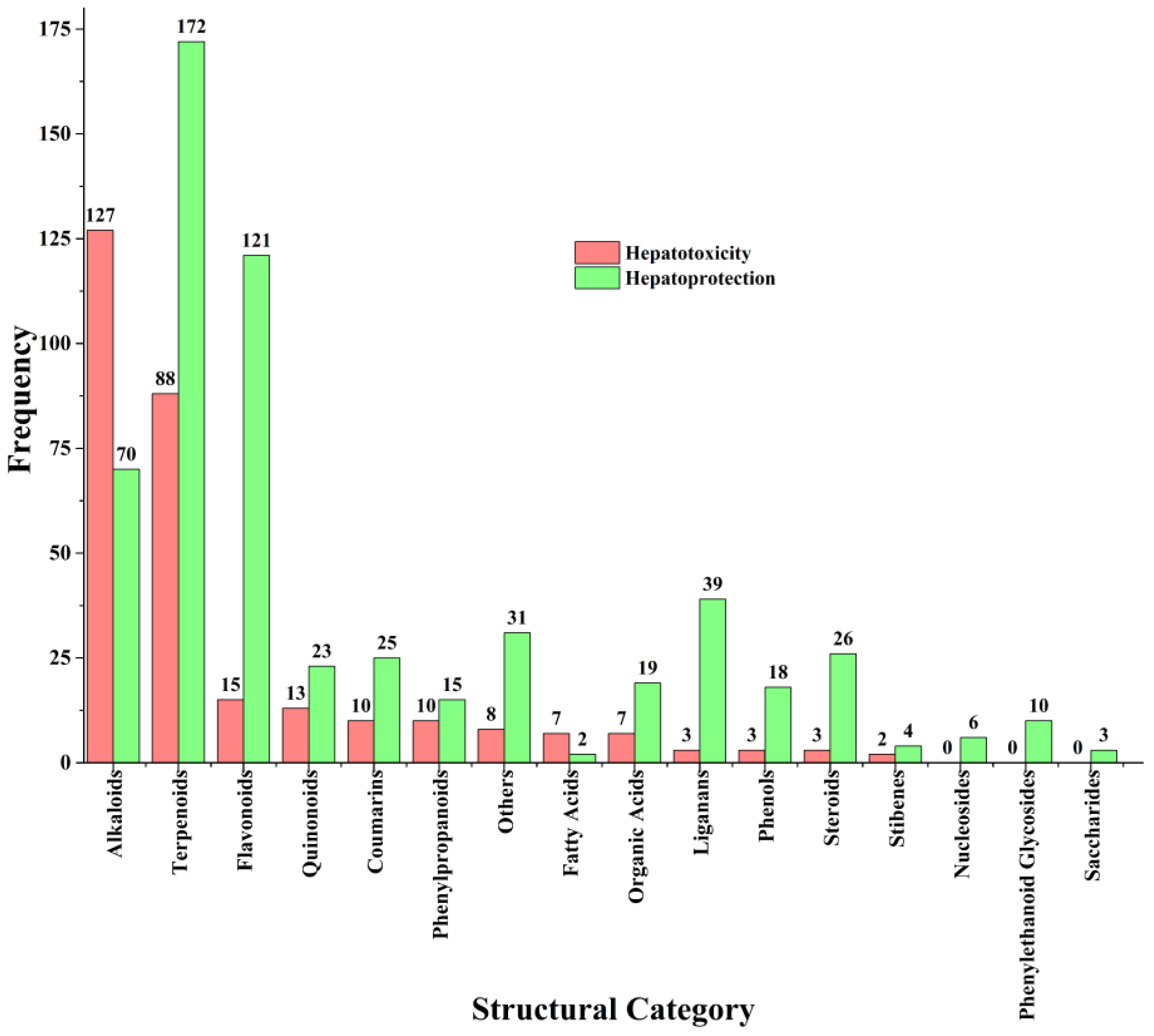
2.3. Eight Structural Categories with Higher Frequency of Hepatotoxicity
2.3.1. Pyrrolizidine Alkaloids
2.3.2. Phenylpropene-Type Simple Phenylpropanooids
2.3.3. Benzophenanthridine Alkaloids
2.3.4. Cadinane-Type Sesquiterpenes
2.3.5. p-Menthane-Type Monoterpenes
2.3.6. Guaiane-Type Sesquiterpenes
2.3.7. Emodin-Type Anthraquinones
2.3.8. Eudesmane-Type Sesquiterpenes
2.4. Structural Alerts for Hepatotoxicity
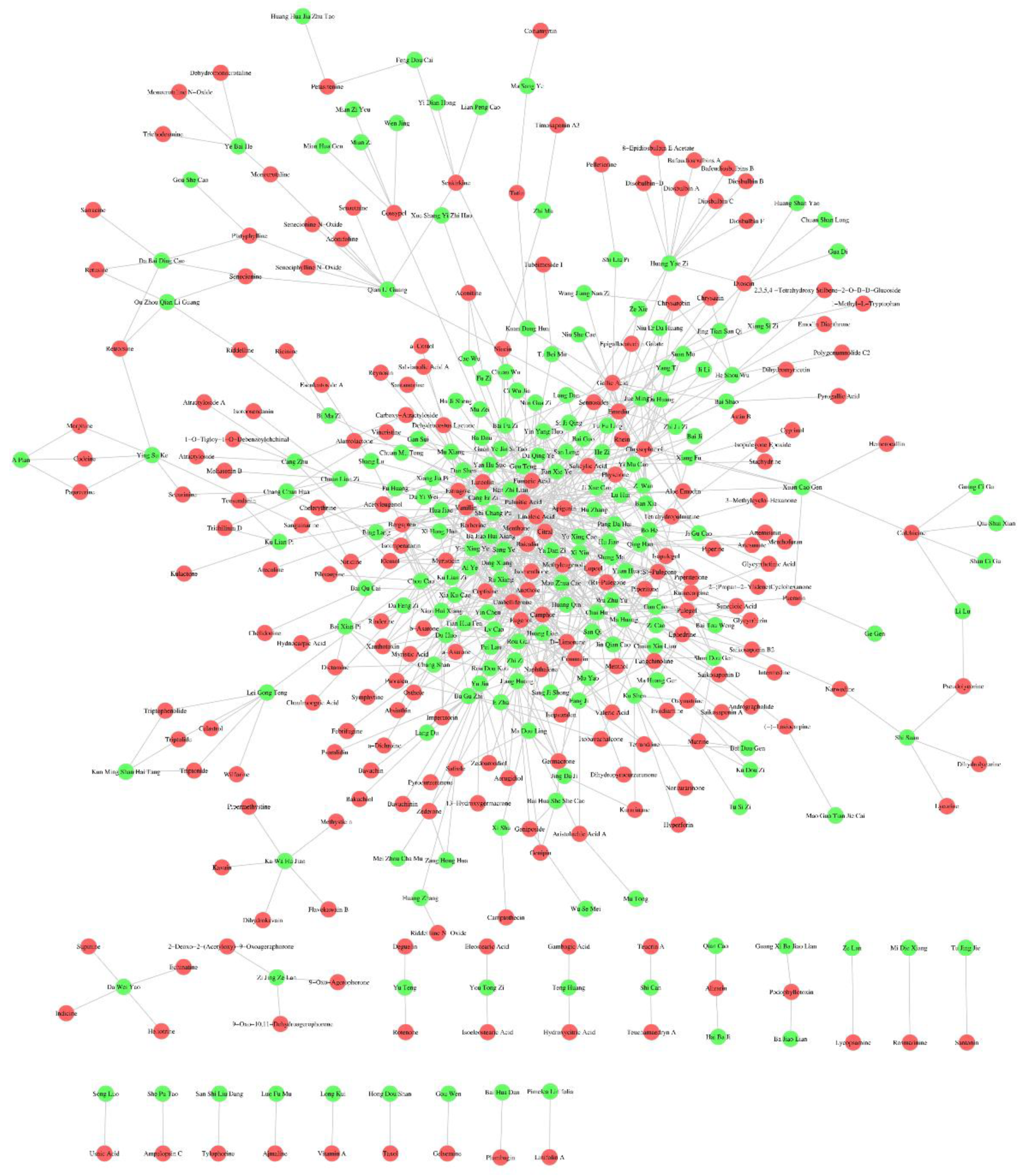
2.5. Construction of Herb-Ingredient Network
3. Materials and Methods
3.1. Search and Identification Terms
3.2. Phylogenetic Tree Construction
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Kaplowitz, N. Idiosyncratic drug hepatotoxicity. Nat. Rev. Drug Discov. 2005, 4, 489–499. [Google Scholar] [CrossRef] [PubMed]
- Navarro, V.J.; Senior, J.R. Drug-related hepatotoxicity. N. Engl. J. Med. 2006, 354, 731–739. [Google Scholar] [CrossRef] [PubMed]
- Hebels, D.G.; Jetten, M.J.; Aerts, H.J.; Herwig, R.; Theunissen, D.H.; Gaj, S.; van Delft, J.H.; Kleinjans, J.C. Evaluation of database-derived pathway development for enabling biomarker discovery for hepatotoxicity. Biomark. Med. 2014, 8, 185–200. [Google Scholar] [CrossRef] [PubMed]
- Hoofnagle, J.H.; Serrano, J.; Knoben, J.E.; Navarro, V.J. LiverTox: A website on drug-induced liver injury. Hepatology 2013, 57, 873–874. [Google Scholar] [CrossRef] [PubMed]
- Russmann, S.; Kullak-Ublick, G.A.; Grattagliano, I. Current concepts of mechanisms in drug-induced hepatotoxicity. Curr. Med. Chem. 2009, 16, 3041–3053. [Google Scholar] [CrossRef] [PubMed]
- Siramshetty, V.B.; Nickel, J.; Omieczynski, C.; Gohlke, B.O.; Drwal, M.N.; Preissner, R. WITHDRAWN—A resource for withdrawn and discontinued drugs. Nucleic Acids Res. 2016, 44, D1080–D1086. [Google Scholar] [CrossRef]
- Prabhakar, A.; Kaiser, J.M.; Novitch, M.B.; Cornett, E.M.; Urman, R.D.; Kaye, A.D. The Role of Complementary and Alternative Medicine Treatments in Fibromyalgia: A Comprehensive Review. Curr. Rheumatol. Rep. 2019, 21, 14. [Google Scholar] [CrossRef]
- Jeong, S.M.; Seo, B.K.; Park, Y.C.; Baek, Y.H. A Review of Complementary and Alternative Medicine Therapies on Muscular Atrophy: A Literature Review of In Vivo/In Vitro Studies. Evid. Based Complement. Altern. Med. Ecam. 2018, 2018, 8654719. [Google Scholar] [CrossRef]
- Terhorst, L.; Schneider, M.J.; Kim, K.H.; Goozdich, L.M.; Stilley, C.S. Complementary and alternative medicine in the treatment of pain in fibromyalgia: A systematic review of randomized controlled trials. J. Manip. Physiol. Ther. 2011, 34, 483–496. [Google Scholar] [CrossRef]
- Seeff, L.B.; Lindsay, K.L.; Bacon, B.R.; Kresina, T.F.; Hoofnagle, J.H. Complementary and alternative medicine in chronic liver disease. Hepatology 2001, 34, 595–603. [Google Scholar] [CrossRef]
- Philippou, Y.; Hadjipavlou, M.; Khan, S.; Rane, A. Complementary and alternative medicine (CAM) in prostate and bladder cancer. BJU Int. 2013, 112, 1073–1079. [Google Scholar] [CrossRef] [PubMed]
- Xu, W.; Zou, Z.; Pei, J.; Huang, L. Longitudinal trend of global artemisinin research in chemistry subject areas (1983–2017). Bioorg. Med. Chem. 2018, 26, 5379–5387. [Google Scholar] [CrossRef] [PubMed]
- Wishart, D.S.; Feunang, Y.D.; Guo, A.C.; Lo, E.J.; Marcu, A.; Grant, J.R.; Sajed, T.; Johnson, D.; Li, C.; Sayeeda, Z.; et al. DrugBank 5.0: A major update to the DrugBank database for 2018. Nucleic Acids Res. 2018, 46, D1074–D1082. [Google Scholar] [CrossRef] [PubMed]
- Afolabi, S.O.; Akindele, A.J.; Awodele, O.; Anunobi, C.C.; Adeyemi, O.O. A 90 day chronic toxicity study of Nigerian herbal preparation DAS-77 in rats. BMC Complement. Altern. Med. 2012, 12, 79. [Google Scholar] [CrossRef] [PubMed]
- Abdualmjid, R.J.; Sergi, C. Hepatotoxic botanicals—An evidence-based systematic review. J. Pharm. Pharm. Sci. 2013, 16, 376–404. [Google Scholar] [CrossRef] [PubMed]
- Shen, T.; Liu, Y.; Shang, J.; Xie, Q.; Li, J.; Yan, M.; Xu, J.; Niu, J.; Liu, J.; Watkins, P.B.; et al. Incidence and Etiology of Drug-Induced Liver Injury in Mainland China. Gastroenterology 2019, 156, 2230–2241. [Google Scholar] [CrossRef] [PubMed]
- Byeon, J.H.; Kil, J.H.; Ahn, Y.C.; Son, C.G. Systematic review of published data on herb induced liver injury. J. Ethnopharmacol. 2019, 233, 190–196. [Google Scholar] [CrossRef]
- Wu, L.; Han, W.; Chen, Y.; Zhang, T.; Liu, J.; Zhong, S.; Liu, H.; Han, C.; Zhang, Z.; Liu, S.; et al. Gender Differences in the Hepatotoxicity and Toxicokinetics of Emodin: The Potential Mechanisms Mediated by UGT2B7 and MRP2. Mol. Pharm. 2018, 15, 3931–3945. [Google Scholar] [CrossRef]
- George, N.; Chen, M.; Yuen, N.; Hunt, C.M.; Suzuki, A. Interplay of gender, age and drug properties on reporting frequency of drug-induced liver injury. Regul. Toxicol. Pharmacol. 2018, 94, 101–107. [Google Scholar] [CrossRef]
- Vaughan, N.; Tweed, J.; Greenwell, C.; Notrica, D.M.; Langlais, C.S.; Peter, S.D.; Leys, C.M.; Ostlie, D.J.; Maxson, R.T.; Ponsky, T.; et al. The impact of morbid obesity on solid organ injury in children using the ATOMAC protocol at a pediatric level I trauma center. J. Pediatr. Surg. 2017, 52, 345–348. [Google Scholar] [CrossRef]
- Ahmad, J.; Odin, J.A. Epidemiology and Genetic Risk Factors of Drug Hepatotoxicity. Clin. Liver Dis. 2017, 21, 55–72. [Google Scholar] [CrossRef] [PubMed]
- Devarbhavi, H.; Singh, R.; Patil, M.; Sheth, K.; Adarsh, C.K.; Balaraju, G. Outcome and determinants of mortality in 269 patients with combination anti-tuberculosis drug-induced liver injury. J. Gastroenterol. Hepatol. 2013, 28, 161–167. [Google Scholar] [CrossRef] [PubMed]
- Chen, M.; Borlak, J.; Tong, W. High lipophilicity and high daily dose of oral medications are associated with significant risk for drug-induced liver injury. Hepatology 2013, 58, 388–396. [Google Scholar] [CrossRef] [PubMed]
- Sasaki, E.; Yokoi, T. Role of cytochrome P450-mediated metabolism and involvement of reactive metabolite formations on antiepileptic drug-induced liver injuries. J. Toxicol. Sci. 2018, 43, 75–87. [Google Scholar] [CrossRef] [PubMed]
- Corsini, A.; Bortolini, M. Drug-induced liver injury: The role of drug metabolism and transport. J. Clin. Pharmacol. 2013, 53, 463–474. [Google Scholar] [CrossRef] [PubMed]
- Wai, C.T.; Tan, B.H.; Chan, C.L.; Sutedja, D.S.; Lee, Y.M.; Khor, C.; Lim, S.G. Drug-induced liver injury at an Asian center: A prospective study. Liver Int. 2007, 27, 465–474. [Google Scholar] [CrossRef] [PubMed]
- Philips, C.A.; Augustine, P.; Rajesh, S.; John, S.K.; Valiathan, G.C.; Mathew, J.; Phalke, S.; Antony, K.L. Slimming to the Death: Herbalife(R)-Associated Fatal Acute Liver Failure-Heavy Metals, Toxic Compounds, Bacterial Contaminants and Psychotropic Agents in Products Sold in India. J. Clin. Exp. Hepatol. 2019, 9, 268–272. [Google Scholar] [CrossRef]
- Philips, C.A.; Paramaguru, R.; Joy, A.K.; Antony, K.L.; Augustine, P. Clinical outcomes, histopathological patterns, and chemical analysis of Ayurveda and herbal medicine associated with severe liver injury—A single-center experience from southern India. Indian J. Gastroenterol. 2018, 37, 9–17. [Google Scholar] [CrossRef]
- Wang, J.; Ma, Z.; Niu, M.; Zhu, Y.; Liang, Q.; Zhao, Y.; Song, J.; Bai, Z.; Zhang, Y.; Zhang, P.; et al. Evidence chain-based causality identification in herb-induced liver injury: Exemplification of a well-known liver-restorative herb Polygonum multiflorum. Front. Med. 2015, 9, 457–467. [Google Scholar] [CrossRef]
- Tseng, A.; Wong, D.K. Hepatotoxicity and potential drug interaction with ledipasvir/sofosbuvir in HIV/HCV infected patients. J. Hepatol. 2016, 65, 651–653. [Google Scholar] [CrossRef]
- Parvez, M.K.; Rishi, V. Herb-Drug Interactions and Hepatotoxicity. Curr. Drug Metab. 2019, 20, 275–282. [Google Scholar] [CrossRef] [PubMed]
- Vinken, M.; Maes, M.; Vanhaecke, T.; Rogiers, V. Drug-induced liver injury: Mechanisms, types and biomarkers. Curr. Med. Chem. 2013, 20, 3011–3021. [Google Scholar] [CrossRef] [PubMed]
- Mulliner, D.; Schmidt, F.; Stolte, M.; Spirkl, H.P.; Czich, A.; Amberg, A. Computational Models for Human and Animal Hepatotoxicity with a Global Application Scope. Chem. Res. Toxicol. 2016, 29, 757–767. [Google Scholar] [CrossRef] [PubMed]
- Holt, M.P.; Ju, C. Mechanisms of drug-induced liver injury. AAPS J. 2006, 8, E48–E54. [Google Scholar] [CrossRef] [PubMed]
- Maddrey, W.C. Hepatotoxicity: The adverse effects of drugs and other chemicals on the liver. Gastroenterology 1999, 118, 984–985. [Google Scholar] [CrossRef]
- Melo-Filho, C.C.; Braga, R.C.; Andrade, C.H. Advances in methods for predicting phase I metabolism of polyphenols. Curr. Drug Metab. 2014, 15, 120–126. [Google Scholar] [CrossRef] [PubMed]
- Lo, Y.C.; Rensi, S.E.; Torng, W.; Altman, R.B. Machine learning in chemoinformatics and drug discovery. Drug Discov. Today 2018, 23, 1538–1546. [Google Scholar] [CrossRef]
- Kar, S.; Leszczynski, J. Exploration of Computational Approaches to Predict the Toxicity of Chemical Mixtures. Toxics 2019, 7, 15. [Google Scholar] [CrossRef]
- Zhang, L.; Zhang, H.; Ai, H.; Hu, H.; Li, S.; Zhao, J.; Liu, H. Applications of Machine Learning Methods in Drug Toxicity Prediction. Curr. Top. Med. Chem. 2018, 18, 987–997. [Google Scholar] [CrossRef]
- Martin, T.M.; Harten, P.; Young, D.M.; Muratov, E.N.; Golbraikh, A.; Zhu, H.; Tropsha, A. Does rational selection of training and test sets improve the outcome of QSAR modeling? J. Chem. Inf. Model. 2012, 52, 2570–2578. [Google Scholar] [CrossRef]
- Kawakami, K. Modification of physicochemical characteristics of active pharmaceutical ingredients and application of supersaturatable dosage forms for improving bioavailability of poorly absorbed drugs. Adv. Drug Deliv. Rev. 2012, 64, 480–495. [Google Scholar] [CrossRef] [PubMed]
- Tyzack, J.D.; Kirchmair, J. Computational methods and tools to predict cytochrome P450 metabolism for drug discovery. Chem. Biol. Drug Des. 2019, 93, 377–386. [Google Scholar] [CrossRef] [PubMed]
- Johnston, H.J.; Hutchison, G.R.; Christensen, F.M.; Aschberger, K.; Stone, V. The biological mechanisms and physicochemical characteristics responsible for driving fullerene toxicity. Toxicol. Sci. 2010, 114, 162–182. [Google Scholar] [CrossRef] [PubMed]
- Reimann, A.; Nurhayati, N.; Backenkohler, A.; Ober, D. Repeated evolution of the pyrrolizidine alkaloid-mediated defense system in separate angiosperm lineages. Plant. Cell. 2004, 16, 2772–2784. [Google Scholar] [CrossRef] [PubMed]
- Schoental, R. Hepatotoxic action of pyrrolizidine (Senecio) alkaloids in relation to their structure. Nature 1957, 179, 361–363. [Google Scholar] [CrossRef] [PubMed]
- Mattocks, A.R.; White, I.N. Pyrrolic metabolites from non-toxic pyrrolizidine alkaloids. Nat. New Biol. 1971, 231, 114–115. [Google Scholar] [CrossRef] [PubMed]
- McLean, E.K. The toxic actions of pyrrolizidine (Senecio) alkaloids. Pharm. Rev. 1970, 22, 429–483. [Google Scholar]
- Tu, M.; Li, L.; Lei, H.; Ma, Z.; Chen, Z.; Sun, S.; Xu, S.; Zhou, H.; Zeng, S.; Jiang, H. Involvement of organic cation transporter 1 and CYP3A4 in retrorsine-induced toxicity. Toxicology 2014, 322, 34–42. [Google Scholar] [CrossRef]
- Yang, M.; Ruan, J.; Fu, P.P.; Lin, G. Cytotoxicity of pyrrolizidine alkaloid in human hepatic parenchymal and sinusoidal endothelial cells: Firm evidence for the reactive metabolites mediated pyrrolizidine alkaloid-induced hepatotoxicity. Chem. Biol. Interact. 2016, 243, 119–126. [Google Scholar] [CrossRef]
- Zhu, L.; Xue, J.; Xia, Q.; Fu, P.P.; Lin, G. The long persistence of pyrrolizidine alkaloid-derived DNA adducts in vivo: Kinetic study following single and multiple exposures in male ICR mice. Arch. Toxicol. 2017, 91, 949–965. [Google Scholar] [CrossRef]
- Yang, M.; Ruan, J.; Gao, H.; Li, N.; Ma, J.; Xue, J.; Ye, Y.; Fu, P.P.; Wang, J.; Lin, G. First evidence of pyrrolizidine alkaloid N-oxide-induced hepatic sinusoidal obstruction syndrome in humans. Arch. Toxicol. 2017, 91, 3913–3925. [Google Scholar] [CrossRef]
- Yan, X.; Kang, H.; Feng, J.; Yang, Y.; Tang, K.; Zhu, R.; Yang, L.; Wang, Z.; Cao, Z. Identification of Toxic Pyrrolizidine Alkaloids and Their Common Hepatotoxicity Mechanism. Int. J. Mol. Sci. 2016, 17, 318. [Google Scholar] [CrossRef] [PubMed]
- Neuman, M.G.; Cohen, L.; Opris, M.; Nanau, R.M.; Hyunjin, J. Hepatotoxicity of Pyrrolizidine Alkaloids. J. Pharm. Pharm. Sci. 2015, 18, 825–843. [Google Scholar] [CrossRef] [PubMed]
- Pramod, K.; Ansari, S.H.; Ali, J. Eugenol: A natural compound with versatile pharmacological actions. Nat. Prod. Commun. 2010, 5, 1999–2006. [Google Scholar] [CrossRef] [PubMed]
- Mizutani, T.; Satoh, K.; Nomura, H.; Nakanishi, K. Hepatotoxicity of eugenol in mice depleted of glutathione by treatment with DL-buthionine sulfoximine. Res. Commun. Chem. Pathol. Pharmacol. 1991, 71, 219–230. [Google Scholar] [PubMed]
- Bodell, W.J.; Ye, Q.; Pathak, D.N.; Pongracz, K. Oxidation of eugenol to form DNA adducts and 8-hydroxy-2’-deoxyguanosine: Role of quinone methide derivative in DNA adduct formation. Carcinogenesis 1998, 19, 437–443. [Google Scholar] [CrossRef] [PubMed]
- Yang, A.H.; Zhang, L.; Zhi, D.X.; Liu, W.L.; Gao, X.; He, X. Identification and analysis of the reactive metabolites related to the hepatotoxicity of safrole. Xenobiotica 2018, 48, 1164–1172. [Google Scholar] [CrossRef]
- Chung, Y.T.; Chen, C.L.; Wu, C.C.; Chan, S.A.; Chi, C.W.; Liu, T.Y. Safrole-DNA adduct in hepatocellular carcinoma associated with betel quid chewing. Toxicol. Lett. 2008, 183, 21–27. [Google Scholar] [CrossRef]
- Murray, M. Toxicological actions of plant-derived and anthropogenic methylenedioxyphenyl-substituted chemicals in mammals and insects. J. Toxicol. Environ. Health B Crit. Rev. 2012, 15, 365–395. [Google Scholar] [CrossRef]
- Chellian, R.; Pandy, V.; Mohamed, Z. Pharmacology and toxicology of alpha- and beta-Asarone: A review of preclinical evidence. Phytomedicine 2017, 32, 41–58. [Google Scholar] [CrossRef]
- Lopez, M.L.; Hernandez, A.; Chamorro, G.; Mendoza-Figueroa, T. alpha-Asarone toxicity in long-term cultures of adult rat hepatocytes. Planta Med. 1993, 59, 115–120. [Google Scholar] [CrossRef] [PubMed]
- Patel, D.N.; Ho, H.K.; Tan, L.L.; Tan, M.M.; Zhang, Q.; Low, M.Y.; Chan, C.L.; Koh, H.L. Hepatotoxic potential of asarones: In vitro evaluation of hepatotoxicity and quantitative determination in herbal products. Front. Pharmacol. 2015, 6, 25. [Google Scholar] [CrossRef] [PubMed]
- Choy, C.S.; Cheah, K.P.; Chiou, H.Y.; Li, J.S.; Liu, Y.H.; Yong, S.F.; Chiu, W.T.; Liao, J.W.; Hu, C.M. Induction of hepatotoxicity by sanguinarine is associated with oxidation of protein thiols and disturbance of mitochondrial respiration. J. Appl. Toxicol. 2008, 28, 945–956. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Tu, M.; Yang, X.; Sun, S.; Wu, X.; Zhou, H.; Zeng, S.; Jiang, H. The contribution of human OCT1, OCT3, and CYP3A4 to nitidine chloride-induced hepatocellular toxicity. Drug Metab. Dispos. 2014, 42, 1227–1234. [Google Scholar] [CrossRef] [PubMed]
- Ouyang, C.B.; Liu, X.M.; Liu, Q.; Bai, J.; Li, H.Y.; Li, Y.; Wang, Q.X.; Yan, D.D.; Mao, L.G.; Cao, A.; et al. Toxicity Assessment of Cadinene Sesquiterpenes from Eupatorium adenophorum in Mice. Nat. Prod. Bioprospect. 2014, 5, 29–36. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Lei, H.P. Hepatotoxicity of gossypol in rats. J. Ethnopharmacol. 1987, 20, 53–64. [Google Scholar] [PubMed]
- Tang, C.; Zhang, K.; Zhan, T.; Zhao, Q.; Zhang, J. Metabolic Characterization of Dairy Cows Treated with Gossypol by Blood Biochemistry and Body Fluid Untargeted Metabolome Analyses. J. Agric. Food Chem. 2017, 65, 9369–9378. [Google Scholar] [CrossRef] [PubMed]
- Manabe, S.; Nuber, D.C.; Lin, Y.C. Zone-specific hepatotoxicity of gossypol in perfused rat liver. Toxicon 1991, 29, 787–790. [Google Scholar] [CrossRef]
- Fonseca, N.B.; Gadelha, I.C.; Oloris, S.C.; Soto-Blanco, B. Effectiveness of albumin-conjugated gossypol as an immunogen to prevent gossypol-associated acute hepatotoxicity in rats. Food Chem. Toxicol. 2013, 56, 149–153. [Google Scholar] [CrossRef]
- Gordon, W.P.; Forte, A.J.; McMurtry, R.J.; Gal, J.; Nelson, S.D. Hepatotoxicity and pulmonary toxicity of pennyroyal oil and its constituent terpenes in the mouse. Toxicol. Appl. Pharmacol. 1982, 65, 413–424. [Google Scholar] [CrossRef]
- Gordon, W.P.; Huitric, A.C.; Seth, C.L.; McClanahan, R.H.; Nelson, S.D. The metabolism of the abortifacient terpene, (R)-(+)-pulegone, to a proximate toxin, menthofuran. Drug Metab. Dispos. 1987, 15, 589–594. [Google Scholar] [PubMed]
- Zarybnicky, T.; Matouskova, P.; Lancosova, B.; Subrt, Z.; Skalova, L.; Bousova, I. Inter-Individual Variability in Acute Toxicity of R-Pulegone and R-Menthofuran in Human Liver Slices and Their Influence on miRNA Expression Changes in Comparison to Acetaminophen. Int. J. Mol. Sci. 2018, 19, 1805. [Google Scholar] [CrossRef] [PubMed]
- Jodynis-Liebert, J.; Murias, M.; Bloszyk, E. Effect of several sesquiterpene lactones on lipid peroxidation and glutathione level. Planta Med. 1999, 65, 320–324. [Google Scholar] [CrossRef] [PubMed]
- Jodynis-Liebert, J.; Murias, M.; Bloszyk, E. Effect of sesquiterpene lactones on antioxidant enzymes and some drug-metabolizing enzymes in rat liver and kidney. Planta Med. 2000, 66, 199–205. [Google Scholar] [CrossRef] [PubMed]
- Merrill, J.C.; Kim, H.L.; Safe, S.; Murray, C.A.; Hayes, M.A. Role of glutathione in the toxicity of the sesquiterpene lactones hymenoxon and helenalin. J. Toxicol. Environ. Health 1988, 23, 159–169. [Google Scholar] [CrossRef]
- Matsuda, H.; Morikawa, T.; Toguchida, I.; Ninomiya, K.; Yoshikawa, M. Medicinal foodstuffs. XXVIII. Inhibitors of nitric oxide production and new sesquiterpenes, zedoarofuran, 4-epicurcumenol, neocurcumenol, gajutsulactones A and B, and zedoarolides A and B, from Zedoariae Rhizoma. Chem. Pharm. Bull. 2001, 49, 1558–1566. [Google Scholar] [CrossRef]
- Matsuda, H.; Ninomiya, K.; Morikawa, T.; Yoshikawa, M. Inhibitory effect and action mechanism of sesquiterpenes from Zedoariae Rhizoma on D-galactosamine/lipopolysaccharide-induced liver injury. Bioorg. Med. Chem. Lett. 1998, 8, 339–344. [Google Scholar] [CrossRef]
- Yang, X.; Zhang, Y.; Liu, Y.; Chen, C.; Xu, W.; Xiao, H. Emodin induces liver injury by inhibiting the key enzymes of FADH/NADPH transport in rat liver. Toxicol. Res. 2018, 7, 888–896. [Google Scholar] [CrossRef]
- Dong, X.; Ni, B.; Fu, J.; Yin, X.; You, L.; Leng, X.; Liang, X.; Ni, J. Emodin induces apoptosis in human hepatocellular carcinoma HepaRG cells via the mitochondrial caspasedependent pathway. Oncol. Rep. 2018, 40, 1985–1993. [Google Scholar]
- Jiang, L.L.; Jiang, Y.; Zhao, D.S.; Fan, Y.X.; Yu, Q.; Li, P.; Li, H.J. CYP3A Activation and Glutathione Depletion Aggravate Emodin-Induced Liver Injury. Chem. Res. Toxicol. 2018, 31, 1052–1060. [Google Scholar] [CrossRef]
- Lin, L.; Liu, Y.; Fu, S.; Qu, C.; Li, H.; Ni, J. Inhibition of Mitochondrial Complex Function-The Hepatotoxicity Mechanism of Emodin Based on Quantitative Proteomic Analyses. Cells 2019, 8, 263. [Google Scholar] [CrossRef] [PubMed]
- He, L.N.; Yang, A.H.; Cui, T.Y.; Zhai, Y.R.; Zhang, F.L.; Chen, J.X.; Jin, C.H.; Fan, Y.W.; Wu, Z.J.; Wang, L.L.; et al. Reactive metabolite activation by CYP2C19-mediated rhein hepatotoxicity. Xenobiotica 2015, 45, 361–372. [Google Scholar] [CrossRef] [PubMed]
- KoraMagazi, A.; Wang, D.; Yousef, B.; Guerram, M.; Yu, F. Rhein triggers apoptosis via induction of endoplasmic reticulum stress, caspase-4 and intracellular calcium in primary human hepatic HL-7702 cells. Biochem. Biophys. Res. Commun. 2016, 473, 230–236. [Google Scholar] [CrossRef] [PubMed]
- You, L.; Dong, X.; Yin, X.; Yang, C.; Leng, X.; Wang, W.; Ni, J. Rhein Induces Cell Death in HepaRG Cells through Cell Cycle Arrest and Apoptotic Pathway. Int. J. Mol. Sci. 2018, 19, 1060. [Google Scholar] [CrossRef] [PubMed]
- Kang, L.; Si, L.; Rao, J.; Li, D.; Wu, Y.; Wu, S.; Wu, M.; He, S.; Zhu, W.; Wu, Y.; et al. Polygoni Multiflori Radix derived anthraquinones alter bile acid disposition in sandwich-cultured rat hepatocytes. Toxicol. In Vitro 2017, 40, 313–323. [Google Scholar] [CrossRef] [PubMed]
- Dong, X.; Fu, J.; Yin, X.; Qu, C.; Yang, C.; He, H.; Ni, J. Induction of Apoptosis in HepaRG Cell Line by Aloe-Emodin through Generation of Reactive Oxygen Species and the Mitochondrial Pathway. Cell. Physiol. Biochem. 2017, 42, 685–696. [Google Scholar] [CrossRef]
- Dong, X.; Fu, J.; Yin, X.; Yang, C.; Ni, J. Aloe-emodin Induces Apoptosis in Human Liver HL-7702 Cells through Fas Death Pathway and the Mitochondrial Pathway by Generating Reactive Oxygen Species. Phytother Res. 2017, 31, 927–936. [Google Scholar] [CrossRef]
- Quan, Y.; Gong, L.; He, J.; Zhou, Y.; Liu, M.; Cao, Z.; Li, Y.; Peng, C. Aloe emodin induces hepatotoxicity by activating NF-kappaB inflammatory pathway and P53 apoptosis pathway in zebrafish. Toxicol. Lett. 2019, 306, 66–79. [Google Scholar] [CrossRef]
- Lim, S.; Lee, S.J.; Nam, K.W.; Kim, K.H.; Mar, W. Hepatoprotective effects of reynosin against thioacetamide-induced apoptosis in primary hepatocytes and mouse liver. Arch. Pharm. Res. 2013, 36, 485–494. [Google Scholar] [CrossRef]
- Qin, C.Z.; Lv, Q.L.; Wu, N.Y.; Cheng, L.; Chu, Y.C.; Chu, T.Y.; Hu, L.; Cheng, Y.; Zhang, X.; Zhou, H.H. Mechanism-based inhibition of Alantolactone on human cytochrome P450 3A4 in vitro and activity of hepatic cytochrome P450 in mice. J. Ethnopharmacol. 2015, 168, 146–149. [Google Scholar] [CrossRef]
- Wang, L.; He, X.; Jin, C.; Ondieki, G. Mechanism-Based Inhibitors from Phytomedicine: Risks of Hepatotoxicity and their Potential Hepatotoxic Substructures. Curr. Drug Metab. 2016, 17, 971–991. [Google Scholar] [CrossRef] [PubMed]
- Pizzo, F.; Lombardo, A.; Manganaro, A.; Benfenati, E. A New Structure-Activity Relationship (SAR) Model for Predicting Drug-Induced Liver Injury, Based on Statistical and Expert-Based Structural Alerts. Front. Pharmacol. 2016, 7, 442. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Ding, L.; Zou, Y.; Hu, S.Q.; Huang, H.G.; Kong, W.B.; Zhang, J. Predicting drug-induced liver injury in human with Naive Bayes classifier approach. J. Comput. Aided Mol. Des. 2016, 30, 889–898. [Google Scholar] [CrossRef] [PubMed]
- Benigni, R.; Bossa, C. Structure alerts for carcinogenicity, and the Salmonella assay system: A novel insight through the chemical relational databases technology. Mutat. Res. 2008, 659, 248–261. [Google Scholar] [CrossRef] [PubMed]
- Ferrari, T.; Cattaneo, D.; Gini, G.; Golbamaki Bakhtyari, N.; Manganaro, A.; Benfenati, E. Automatic knowledge extraction from chemical structures: The case of mutagenicity prediction. SAR QSAR Environ. Res. 2013, 24, 365–383. [Google Scholar] [CrossRef] [PubMed]
- Lombardo, A.; Pizzo, F.; Benfenati, E.; Manganaro, A.; Ferrari, T.; Gini, G. A new in silico classification model for ready biodegradability, based on molecular fragments. Chemosphere 2014, 108, 10–16. [Google Scholar] [CrossRef] [PubMed]
- Teschke, R.; Genthner, A.; Wolff, A. Kava hepatotoxicity: Comparison of aqueous, ethanolic, acetonic kava extracts and kava-herbs mixtures. J. Ethnopharmacol. 2009, 123, 378–384. [Google Scholar] [CrossRef]
- Gordon, P.; Khojasteh, S.C. A decades-long investigation of acute metabolism-based hepatotoxicity by herbal constituents: A case study of pennyroyal oil. Drug Metab. Rev. 2015, 47, 12–20. [Google Scholar] [CrossRef]
- Wei, J.; Zhang, F.; Zhang, Y.; Cao, C.; Li, X.; Li, D.; Liu, X.; Yang, H.; Huang, L. Proteomic investigation of signatures for geniposide-induced hepatotoxicity. J. Proteome Res. 2014, 13, 5724–5733. [Google Scholar] [CrossRef]
- Yamano, T.; Tsujimoto, Y.; Noda, T.; Shimizu, M.; Ohmori, M.; Morita, S.; Yamada, A. Hepatotoxicity of geniposide in rats. Food Chem. Toxicol. 1990, 28, 515–519. [Google Scholar] [CrossRef]
- Garcia-Cortes, M.; Robles-Diaz, M.; Ortega-Alonso, A.; Medina-Caliz, I.; Andrade, R.J. Hepatotoxicity by Dietary Supplements: A Tabular Listing and Clinical Characteristics. Int. J. Mol. Sci. 2016, 17, 537. [Google Scholar] [CrossRef] [PubMed]
- Zarybnicky, T.; Bousova, I.; Ambroz, M.; Skalova, L. Hepatotoxicity of monoterpenes and sesquiterpenes. Arch. Toxicol. 2018, 92, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Federhen, S. The NCBI Taxonomy database. Nucleic Acids Res. 2012, 40, D136–D143. [Google Scholar] [CrossRef] [PubMed]
- Letunic, I.; Bork, P. Interactive Tree Of Life (iTOL) v4: Recent updates and new developments. Nucleic Acids Res. 2019, 47, W256–W259. [Google Scholar] [CrossRef] [PubMed]
- Newman, D.J.; Cragg, G.M. Natural Products as Sources of New Drugs from 1981 to 2014. J. Nat. Prod. 2016, 79, 629–661. [Google Scholar] [CrossRef] [PubMed]
- Newman, D.J.; Cragg, G.M. Natural products as sources of new drugs over the 30 years from 1981 to 2010. J. Nat. Prod. 2012, 75, 311–335. [Google Scholar] [CrossRef] [PubMed]
- Newman, D.J.; Cragg, G.M. Natural products as sources of new drugs over the last 25 years. J. Nat. Prod. 2007, 70, 461–477. [Google Scholar] [CrossRef]
- Zhu, F.; Qin, C.; Tao, L.; Liu, X.; Shi, Z.; Ma, X.; Jia, J.; Tan, Y.; Cui, C.; Lin, J.; et al. Clustered patterns of species origins of nature-derived drugs and clues for future bioprospecting. Proc. Natl. Acad. Sci. USA 2011, 108, 12943–12948. [Google Scholar] [CrossRef]
- Saslis-Lagoudakis, C.H.; Klitgaard, B.B.; Forest, F.; Francis, L.; Savolainen, V.; Williamson, E.M.; Hawkins, J.A. The use of phylogeny to interpret cross-cultural patterns in plant use and guide medicinal plant discovery: An example from Pterocarpus (Leguminosae). PLoS ONE 2011, 6, e22275. [Google Scholar] [CrossRef]
- Saslis-Lagoudakis, C.H.; Savolainen, V.; Williamson, E.M.; Forest, F.; Wagstaff, S.J.; Baral, S.R.; Watson, M.F.; Pendry, C.A.; Hawkins, J.A. Phylogenies reveal predictive power of traditional medicine in bioprospecting. Proc. Natl. Acad. Sci. USA 2012, 109, 15835–15840. [Google Scholar] [CrossRef]
- Fu, X.; Song, X.; Li, X.; Wong, K.K.; Li, J.; Zhang, F.; Wang, C.; Wang, Z. Phylogenetic Tree Analysis of the Cold-Hot Nature of Traditional Chinese Marine Medicine for Possible Anticancer Activity. Evid. Based Complement. Altern. Med. Ecam. 2017, 2017, 4365715. [Google Scholar] [CrossRef] [PubMed]
- Brown, A.C. Kidney toxicity related to herbs and dietary supplements: Online table of case reports. Part 3 of 5 series. Food Chem. Toxicol. 2017, 107, 502–519. [Google Scholar] [CrossRef] [PubMed]
- Brown, A.C. Liver toxicity related to herbs and dietary supplements: Online table of case reports. Part 2 of 5 series. Food Chem. Toxicol. 2017, 107, 472–501. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.Y.; Jun, S.A.; Hong, S.S.; Ahn, Y.C.; Lee, D.S.; Son, C.G. Systematic Review of Adverse Effects from Herbal Drugs Reported in Randomized Controlled Trials. Phytother Res. 2016, 30, 1412–1419. [Google Scholar] [CrossRef] [PubMed]
- Brown, A.C. Heart Toxicity Related to Herbs and Dietary Supplements: Online Table of Case Reports. Part 4 of 5. J. Diet. Suppl. 2018, 15, 516–555. [Google Scholar] [CrossRef]
- Brown, A.C. Cancer Related to Herbs and Dietary Supplements: Online Table of Case Reports. Part 5 of 5. J. Diet. Suppl. 2018, 15, 556–581. [Google Scholar] [CrossRef] [PubMed]
- Saini, N.; Bakshi, S.; Sharma, S. In-silico approach for drug induced liver injury prediction: Recent advances. Toxicol. Lett. 2018, 295, 288–295. [Google Scholar] [CrossRef]
- Chen, M.; Bisgin, H.; Tong, L.; Hong, H.; Fang, H.; Borlak, J.; Tong, W. Toward predictive models for drug-induced liver injury in humans: Are we there yet? Biomark. Med. 2014, 8, 201–213. [Google Scholar] [CrossRef]
- Wetzel, S.; Schuffenhauer, A.; Roggo, S.; Ertl, P.; Waldmann, H. Cheminformatic Analysis of Natural Products and their Chemical Space. CHIMIA Int. J. Chem. 2007, 61, 355–360. [Google Scholar] [CrossRef]
- Stratton, C.F.; Newman, D.J.; Tan, D.S. Cheminformatic comparison of approved drugs from natural product versus synthetic origins. Bioorg. Med. Chem. Lett. 2015, 25, 4802–4807. [Google Scholar] [CrossRef]
- Zhu, Y.L.; Zu-Guang, Y.E. Computational toxicology and its application in toxicity study of traditional Chinese medicine. Chin. J. New Drugs 2011, 20, 2424–2429. [Google Scholar]
- Ai, H.; Chen, W.; Zhang, L.; Huang, L.; Yin, Z.; Hu, H.; Zhao, Q.; Zhao, J.; Liu, H. Predicting Drug-Induced Liver Injury Using Ensemble Learning Methods and Molecular Fingerprints. Toxicol. Sci. 2018, 165, 100–107. [Google Scholar] [CrossRef] [PubMed]
- Zhao, P.; Liu, B.; Wang, C. Hepatotoxicity evaluation of traditional Chinese medicines using a computational molecular model. Clin. Toxicol. 2017, 55, 996–1000. [Google Scholar] [CrossRef] [PubMed]
- Villa, F.A.; Gerwick, L. Marine natural product drug discovery: Leads for treatment of inflammation, cancer, infections, and neurological disorders. Immunopharmacol. Immunotoxicol. 2010, 32, 228–237. [Google Scholar] [CrossRef] [PubMed]
- Baker, D.D.; Chu, M.; Oza, U.; Rajgarhia, V. The value of natural products to future pharmaceutical discovery. Nat. Prod. Rep. 2007, 24, 1225–1244. [Google Scholar] [CrossRef] [PubMed]
- Mishra, B.B.; Tiwari, V.K. Natural products: An evolving role in future drug discovery. Eur. J. Med. Chem. 2011, 46, 4769–4807. [Google Scholar] [CrossRef]
- Kitada, S.; Kress, C.L.; Krajewska, M.; Jia, L.; Pellecchia, M.; Reed, J.C. Bcl-2 antagonist apogossypol (NSC736630) displays single-agent activity in Bcl-2-transgenic mice and has superior efficacy with less toxicity compared with gossypol (NSC19048). Blood 2008, 111, 3211–3219. [Google Scholar] [CrossRef]
- Feng, Z.; Zhou, C.; Dong, S.; Liu, Z.; Liu, T.; Zhou, L.; Zhou, X. Catalpol and panax notoginseng saponins synergistically alleviate triptolide-induced hepatotoxicity through Nrf2/ARE pathway. Toxicol. In Vitro 2019, 56, 141–149. [Google Scholar] [CrossRef]
- Li, W.; Yin, H.; Bardelang, D.; Xiao, J.; Zheng, Y.; Wang, R. Supramolecular formulation of nitidine chloride can alleviate its hepatotoxicity and improve its anticancer activity. Food Chem. Toxicol. 2017, 109, 923–929. [Google Scholar] [CrossRef]
- Liu, C.; Fan, H.; Li, Y.; Xiao, X. Research Advances on Hepatotoxicity of Herbal Medicines in China. Biomed. Res. Int. 2016, 2016, 7150391. [Google Scholar] [CrossRef]
- Stine, J.G.; Lewis, J.H. Current and future directions in the treatment and prevention of drug-induced liver injury: A systematic review. Expert Rev. Gastroenterol. Hepatol. 2016, 10, 517–536. [Google Scholar] [CrossRef] [PubMed]


| Rank | Order | Size | Frequency of Hepatotoxic Herb | Percentage |
|---|---|---|---|---|
| 1 | Ranunculales | 2830 | 33 | 1.1661% |
| 2 | Boraginales | 2700 | 12 | 0.4444% |
| 3 | Piperales | 4170 | 11 | 0.2638% |
| 4 | Sapindales | 5700 | 15 | 0.2632% |
| 5 | Apiales | 5489 | 12 | 0.2186% |
| 6 | Gentianales | 17000 | 23 | 0.1353% |
| 7 | Malpighiales | 16000 | 20 | 0.1250% |
| 8 | Caryophyllales | 12000 | 15 | 0.1250% |
| 9 | Asterales | 26870 | 31 | 0.1154% |
| 10 | Fabales | 20000 | 21 | 0.1050% |
| 11 | Lamiales | 23800 | 23 | 0.0966% |
| 12 | Asparagales | 36200 | 14 | 0.0387% |
| Rank | Family | Size | Frequency of Hepatotoxic Herb | Percentage |
|---|---|---|---|---|
| 1 | Aristolochiaceae | 590 | 8 | 1.3559% |
| 2 | Polygonaceae | 1100 | 11 | 1.0000% |
| 3 | Ranunculaceae | 2252 | 16 | 0.7105% |
| 4 | Boraginaceae | 2700 | 9 | 0.3333% |
| 5 | Lamiaceae | 7000 | 17 | 0.2429% |
| 6 | Apiaceae | 3700 | 8 | 0.2162% |
| 7 | Euphorbiaceae | 6745 | 11 | 0.1631% |
| 8 | Asteraceae | 26870 | 31 | 0.1154% |
| 9 | Fabaceae | 20000 | 21 | 0.1050% |
| 10 | Rubiaceae | 13150 | 11 | 0.0837% |
| ID | Genus | Frequency of Hepatotoxicity Herb |
|---|---|---|
| 1 | Aristolochia | 6 |
| 2 | Senecio | 5 |
| 3 | Uncaria | 5 |
| 4 | Rumex | 5 |
| 5 | Aconitum | 4 |
| 6 | Actaea | 4 |
| 7 | Epimedium | 4 |
| 8 | Senna | 4 |
| 9 | Gentiana | 4 |
| 10 | Euphorbia | 4 |
| 11 | Symphytum | 4 |
| 12 | Gossypium | 4 |
| 13 | Curcuma | 4 |
| 14 | Dioscorea | 4 |
| 15 | Eupatorium | 4 |
| ID | Description | Representative Compounds | Hepatotoxic/Total (Percentage) | |
|---|---|---|---|---|
| Name | Structure | |||
| 1 | Pyrrolizidine alkaloids | Senecionine |  | 77/77(100.00%) |
| 2 | Phenylpropene-type simple phenylpropanooids | Eugenol |  | 9/9(100.00%) |
| 3 | Benzophenanthridine Alkaloids | Nitidine | 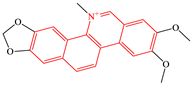 | 4/5(80.00%) |
| 4 | Cadinane-type sesquiterpenes | 9-Oxo-agerophorone | 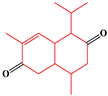 | 4/5(80.00%) |
| 5 | P-menthane-type monoterpenes | Pulegone | 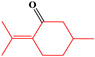 | 13/17(76.47%) |
| 6 | Guaiane-type sesquiterpenes | Atractyloside A | 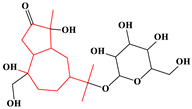 | 9/14(64.29%) |
| 7 | Emodin-type anthraquinones | Rhein | 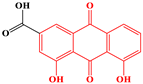 | 6/11(54.55%) |
| 8 | Eudesmane-type sesquiterpenes | Alpha-costol | 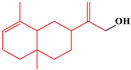 | 7/17(41.18%) |
| ID | SAs | LR | Hepatotoxic/Total (Percentage) | Distribution of SAs |
|---|---|---|---|---|
| 1 | 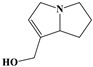 | inf | 47/47 (100.00%) | Pyrrolizidine alkaloids |
| 2 | 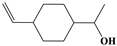 | inf | 7/7 (100.00%) | Sesquiterpenes |
| 3 | 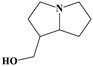 | inf | 7/7 (100.00%) | Pyrrolizidine alkaloids |
| 4 | 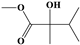 | 112.77 | 55/56 (98.21%) | Pyrrolizidine alkaloids |
| 5 | 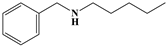 | 12.30 | 6/7 (85.71%) | Alkaloids |
| 6 | 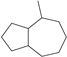 | 12.30 | 6/7 (85.71%) | Terpenoids dominated by Guaiane sesquiterpenes |
| 7 | 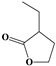 | 11.62 | 17/20 (85.00%) | Terpenoids |
| 8 |  | 8.71 | 68/84 (80.95%) | Pyrrolizidine alkaloids and Sesquiterpenes |
| 9 | 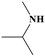 | 6.66 | 78/102 (76.47%) | Alkaloids |
| 10 | 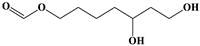 | 5.33 | 13/18 (72.22%) | Terpenoids |
| 11 | 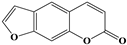 | 5.13 | 5/7 (71.43%) | Furanocoumarins |
| 12 | 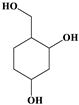 | 5.13 | 10/14 (71.43%) | Terpenoids |
| 13 | 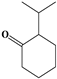 | 4.10 | 10/15 (66.67%) | Terpenoids dominated by tetracyclic triterpenoids |
| 14 | 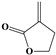 | 4.10 | 12/18 (66.67%) | Sesquiterpenes |
| 15 | 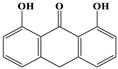 | 3.69 | 9/14 (64.29%) | Emodin anthraquinones and their derivatives |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
He, S.; Zhang, C.; Zhou, P.; Zhang, X.; Ye, T.; Wang, R.; Sun, G.; Sun, X. Herb-Induced Liver Injury: Phylogenetic Relationship, Structure-Toxicity Relationship, and Herb-Ingredient Network Analysis. Int. J. Mol. Sci. 2019, 20, 3633. https://doi.org/10.3390/ijms20153633
He S, Zhang C, Zhou P, Zhang X, Ye T, Wang R, Sun G, Sun X. Herb-Induced Liver Injury: Phylogenetic Relationship, Structure-Toxicity Relationship, and Herb-Ingredient Network Analysis. International Journal of Molecular Sciences. 2019; 20(15):3633. https://doi.org/10.3390/ijms20153633
Chicago/Turabian StyleHe, Shuaibing, Chenyang Zhang, Ping Zhou, Xuelian Zhang, Tianyuan Ye, Ruiying Wang, Guibo Sun, and Xiaobo Sun. 2019. "Herb-Induced Liver Injury: Phylogenetic Relationship, Structure-Toxicity Relationship, and Herb-Ingredient Network Analysis" International Journal of Molecular Sciences 20, no. 15: 3633. https://doi.org/10.3390/ijms20153633
APA StyleHe, S., Zhang, C., Zhou, P., Zhang, X., Ye, T., Wang, R., Sun, G., & Sun, X. (2019). Herb-Induced Liver Injury: Phylogenetic Relationship, Structure-Toxicity Relationship, and Herb-Ingredient Network Analysis. International Journal of Molecular Sciences, 20(15), 3633. https://doi.org/10.3390/ijms20153633





